Abstract
Yeast Hsp104 is a ring-forming, ATP-dependent protein disaggregase that, together with the cognate Hsp70 chaperone system, has the remarkable ability to rescue stress damaged proteins from a previously aggregated state. Both up-stream and down-stream functions for the Hsp70 system have been reported, but it remains unclear how Hsp70/Hsp40 is coupled to the Hsp104 protein remodeling activity.
Hsp104 is a multi-domain protein that possesses an N-terminal domain, an M-domain, and two tandem AAA+ domains. The M-domain forms an 85-Å long coiled-coil and is a hallmark of the Hsp104 chaperone family. While the 3D structure of Hsp104 has been determined, the function of the M-domain is unclear. Here, we demonstrate that the M-domain is essential for protein disaggregation but dispensable for the Hsp104 ATPase and substrate translocating activities. Remarkably, replacing the Hsp104 M-domain against that of bacterial ClpB, and vice versa, switches the species-specificity so that our chimeras now cooperate with the non-cognate Hsp70/DnaK chaperone system. Our results demonstrate that the M-domain controls the Hsp104 protein remodeling activities in an Hsp70/Hsp40-dependent manner, which is required to unleash the Hsp104 protein disaggregating activity.
Keywords: Molecular chaperone; AAA+ machine; protein disaggregase; Hsp104, ClpB
Yeast Hsp104 is a ring-forming AAA+ machine that together with the cognate Hsp70 system (Hsp70/Hsp40) rescues proteins from a previously aggregated state.1; 2; 3 In addition to its essential role in the yeast stress response, Hsp104 is also required for the maintenance of all amyloid-based yeast prions,4 and for the elimination of [PSI+], a yeast prion that increases translational read-through of nonsense codons.5; 6 The ability to remodel prions is not shared with the orthologous bacterial ClpB,6; 7 and might be reflected in its 3D structure.8; 9 However, we recently demonstrated that Hsp104 and ClpB share a remarkable structural conservation.10; 11; 12 Our Hsp104 structure is consistent with a common mechanism for protein disaggregation and prion propagation,13; 14 and confirms the essential role of the Hsp104 pore loops in substrate translocation.15; 16
Hsp104 and ClpB are multi-domain proteins consisting of an N-terminal domain (NTD), an M-domain, and two Walker-type ATPase domains (AAA+).10; 12 The M-domain is a hallmark of the Hsp104/ClpB chaperone family and is inserted within the D1-small domain of the first AAA+ domain (AAA-1), similar to the β-domain of the distantly related Vps4 ATPase.17 In solution, Hsp104 forms a homo-hexamer with the M-domains on the outside of the hexamer ring,12 as they are in ClpB.10; 11 While the exact function of the Hsp104 M-domain remains unknown, its location on the Hsp104 exterior is consistent with a potential role in contacting large aggregated proteins and prion fibrils, as well as components of the bi-chaperone system.
It is widely accepted that the ability to solubilize protein aggregates requires the synergistic interaction between Hsp104 and the Hsp70 system in a species-specific manner.18; 19 Consistent with this notion, a direct interaction between bacterial ClpB and DnaK has been reported,20; 21 but its functional implication for protein disaggregation remains unclear. Since other Clp/Hsp100 proteins that lack an M-domain do not cooperate with the DnaK/Hsp70 system in protein disaggregation,22 it is conceivable that the M-domain might mediate the synergistic interaction between Hsp104 and Hsp70/Hsp40. To test this hypothesis, we used a biochemical approach to determine the functional role of the M-domain for chaperone activity. We demonstrate that the M-domain is essential for the solubilization of protein aggregates, but dispensable for the ATPase and protein translocating activities of unfolded substrates. Remarkably, we found that replacing the yeast Hsp104 M-domain against that of bacterial ClpB, and vice versa, switches the species-specificity of the bi-chaperone system. Our results demonstrate that the M-domain controls Hsp104 function through direct interaction with the cognate Hsp70 system that is required to unleash the Hsp104 protein remodeling activity.
The Hsp104-T4L chimera solubilizes a variety of heat-aggregated substrates
We previously reported the fitted cryoEM structure of an engineered Hsp104 variant (Hsp104T4L) that harbors T4 lysozyme (T4L) within the M-domain helix 2.12 This Hsp104T4L variant is fully functional biochemically and, like wild-type Hsp104, cooperates with the Hsp70 system to reactivate heat-aggregated β-galactosidase (β-gal) and chemically denatured firefly luciferase (FFL).12 Notably, Hsp104T4L has gained the ability to solubilize heat-aggregated β-gal in the absence of the Hsp70 system (Fig. 1a).12 To determine whether Hsp104T4L can also recognize other aggregated proteins, we tested two additional heat-aggregated model substrates, namely enhanced green fluorescent protein (GFP) and malate dehydrogenase (MDH).
Fig. 1.
Disaggregation of model substrates by Hsp104-T4L fusion proteins. (a-c) Reactivation of protein aggregates by Hsp104, Hsp104T4L, ΔN-Hsp104, or ΔN-Hsp104T4L (1 μM each) alone and together with Hsp70 and Ydj-1 (1 μM each). Assays contained an ATP-regenerating system (20 mM phosphoenolpyruvate and 1 μM pyruvate kinase) and 4 mM ATP. Proteins were produced as described in the supplementary material. (a) Reactivation of heat-aggregated β-gal was determined as described12 and is shown over time as percentage of a native control. Standard errors of three independent assays are shown. (b) 4.5 μM enhanced GFP (BioVision, Mountain View, CA) was denatured at 80 °C in buffer A (25 mM HEPES pH 7.5, 150 mM KOAc, 10 mM Mg(OAc)2, and 10 mM DTT) for 10 min and diluted 1:10 with buffer A containing chaperones as indicated. The increase in fluorescence was measured with a Synergy HT microplate reader (BioTek, Winooski, VT) in 1 min intervals. A representative data set of three experiments is shown. (c) 2 μM MDH (Roche Diagnostics, Indianapolis, IN) was incubated in buffer A at 47 °C for 30 min and diluted 2.5-fold into buffer A at 22 °C with chaperones as indicated. MDH activity was determined after 180 min as described33 and is shown as percentage of a native control. Standard errors of three independent assays are shown.
We found that Hsp104T4L can reactivate all three model substrates in the absence of the Hsp70 system (Fig. 1a-c). Notably, Hsp104T4L alone was able to reactivate heat-aggregated proteins perhaps even more efficiently than the wild-type bi-chaperone system (Fig. 1a-c). The higher efficiency of Hsp104T4L contrasts previously reported activities,12 and is due to an improved protein purification procedure (details provided in the supplementary material). No further enhancement in substrate reactivation was observed in the presence of the Hsp70 system, demonstrating that Hsp104T4L alone can solubilize a wide variety of heat-aggregated substrates.
The Hsp104 N-terminal domain is dispensable for protein disaggregation
Our Hsp104T4L variant provides the necessary means to investigate the Hsp104 protein-remodeling activity free of other molecular chaperones and additional ATPases. Since the ATPase domains are conserved among AAA+ machines, we reasoned that additional domains present in Hsp104 must confer the unique protein disaggregating function. While the Hsp104 NTD is dispensable for thermotolerance and luciferase refolding in vivo,13; 23 it is necessary for [PSI+] prion inheritance13 and curing by Hsp104 overexpression,23 suggesting that the NTD confers some substrate-specificity. To determine whether the Hsp104 NTD is important for protein disaggregation in the absence of the Hsp70 system, we engineered an Hsp104T4L variant lacking the NTD (ΔN-Hsp104T4L). We found that ΔN-Hsp104T4L alone can solubilize all three heat-aggregated model substrates with similar efficiencies as Hsp104T4L (Fig. 1a-c). Our findings confirm that the Hsp104 NTD is dispensable for the disaggregation of heat-aggregated substrates, even in the absence of the Hsp70 system.
The Hsp104 M-domain is essential for protein disaggregation
To determine whether the Hsp104 M-domain is essential for protein disaggregation, we engineered an Hsp104 variant that lacks the M-domain in both wild-type (Hsp104ΔM) and trap mutant background (Hsp104ΔMTrap) (Fig. 2a). We found that Hsp104ΔM forms a hexamer (Fig. 2b), and hydrolyzes ATP, which is stimulated in the presence of κ-casein (Fig. 2c). However, unlike wild-type Hsp104, Hsp104ΔM could not solubilize heat-aggregated model substrates in the presence of Hsp70/Hsp40 (Fig. 2d), indicating that Hsp104ΔM lost the ability to disaggregate amorphous aggregates. Notably, Hsp104ΔMTrap still bound FITC-casein, a soluble model substrate, with similar binding affinity as Hsp104Trap (Fig. 2e). Thus, our results demonstrate that the Hsp104 M-domain is essential for the disaggregation of heat-aggregated proteins but is not required for binding of soluble model substrates.
Fig. 2.
Biochemical characterization of Hsp104 variants. (a) Domain arrangements of Hsp104, Hsp104ΔM, and Hsp104-ClpB chimeras. The AAA-1 domain is colored blue, deletion of the M-domain is indicated by a grey line, and domains that were replaced by corresponding ClpB sequences are colored orange. Hsp104ΔM and Hsp104-ClpB chimeras were constructed by cassette mutagenesis, expressed, and purified as described in the supplementary material. (b) Hsp104, Hsp104ΔM, and Hsp104-ClpB chimeras (0.6 mg/ml) were incubated for 20 min at 22 °C in buffer A containing 2 mM ATPγS and 0.2 mg/ml κ-casein. Hexamer assembly was analyzed by size-exclusion chromatography on a Superdex 200 10/300 GL column (GE Healthcare, Piscataway, NJ) in buffer A plus 10 μM ATPγS. (c) ATPase rates of Hsp104, Hsp104ΔM, and Hsp104-ClpB chimeras in the absence or presence of 0.2 mg/ml κ-casein were determined using a coupled ATP-regenerating system at 30 °C in buffer A containing 2 mM ATP as described.12 Standard errors of three independent assays are shown. (d) Reactivation of aggregated model substrates by Hsp70 and Ydj-1 (1 μM each) alone and together with either Hsp104 (1 μM) or Hsp104ΔM (1 μM). Disaggregation assays were performed as described in Fig. 1a-c. Standard errors of three independent assays are shown. (e) Fluorescence polarization binding isotherms of FITC-casein to Hsp104Trap, Hsp104ΔMTrap, and Hsp104BMTrap. Equilibrium binding constants were determined at 22 °C by adding 0.15 µg/ml FITC-casein (Sigma-Aldrich, St. Louis, MO) to Trap hexamers that were preassembled with 2 mM ATP in buffer A at 22 °C. Standard errors of three independent assays are shown.
Hsp104-ClpB chimeras are functional biochemically
To understand the function of the Hsp104 M-domain in protein disaggregation, we engineered Hsp104 chimeras that feature one or more domains of the orthologous Escherichia coli ClpB. It has previously been shown that Hsp104-ClpB chimeras can provide yeast thermotolerance in vivo as long as the AAA-1 domain of yeast Hsp104 is present,13 suggesting that the AAA-2 domain is dispensable for the Hsp104-specific chaperone activity.
The X-ray structure of ClpB showed that the AAA-1 domain is composed of a D1-large domain and a D1-small domain that features the M-domain inserted between helix C3 and β-strand c2.10 To determine which segment of the AAA-1 domain may confer species-specificity, we replaced the M-domain (Hsp104BM), the D1-small domain (Hsp104BDS) or the entire AAA-1 domain of Hsp104 (Hsp104BDLS) against the analogous domains of E. coli ClpB (Fig. 2a). All of our Hsp104-ClpB chimeras form hexamers as determined by size-exclusion chromatography (Fig. 2b). We also observed additional shoulders and smaller peaks in the chromatograms, indicating that our engineered chimeras are somewhat less stable than wild-type Hsp104 and Hsp104ΔM. However, our biochemical analysis showed that the ATP hydrolysis rates of Hsp104BDS and Hsp104BDLS are similar to those of wild-type Hsp104 (Fig. 2c), suggesting that the lower stability did not affect significantly the basal ATPase activities of those chimeras. Moreover, like wild-type Hsp104 and Hsp104ΔM, the ATPase activities of all three Hsp104-ClpB chimeras are further stimulated by κ-casein (Fig. 2c), albeit to a different extent. The complex nature of the basal and stimulated ATPase activities observed with our engineered chimeras is in line with the proposed role of the M-domain in coupling the ATPase activities of the AAA-1 and AAA-2 domains, 24 which is effected by the conformation and stability of the long coiled-coil.10; 25; 26; 27
Replacing the M-domain switches the species-specificity of the bi-chaperone system
To identify the AAA-1 segment that confers the species-specificity of the bi-chaperone system, we asked whether any of our Hsp104 chimeras could synergize with the bacterial DnaK system (DnaK/DnaJ/GrpE) to rescue aggregated proteins. To do so, we set up chaperone activity assays using either chemically denatured FFL (Fig. 3a) or heat-aggregated MDH (Fig. 3b) as substrate. Both aggregated FFL and MDH have been shown previously to function as model substrates for the yeast (Fig. 1c and Ref. 18) and the bacterial bi-chaperone system.28; 29
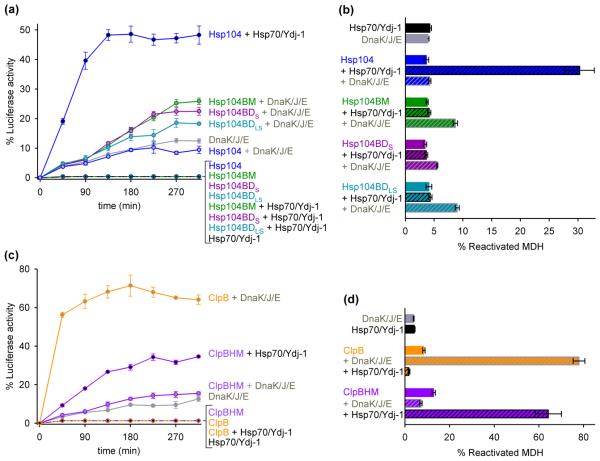
Fig. 3.
M-domain replacement switches the species-specificity of the bi-chaperone system. (a) Reactivation of chemically denatured FFL over time by Hsp104 or Hsp104-ClpB chimeras (1 μM each) alone and together with Hsp70 and Ydj-1 (1 μM each) or DnaK (0.86 μM), DnaJ (0.43 μM), and GrpE (0.25 μM). DnaK, DnaJ, and GrpE were purified as described in the supplementary material. 10 μM FFL (Promega, Madison, WI) was denatured in 7 M Urea, 25 mM HEPES pH 7.5, 50 mM KCl, 15 mM MgCl2, 10 mM DTT, and 5 mM ATP at 22 °C,34 and diluted 125-fold into 25 mM HEPES pH 7.5, 150 mM KCl, 15 mM MgCl2, 2 mM DTT and chaperones as indicated. Luminescence was measured at 22 °C as described.12 (b) Reactivation of heat-aggregated MDH by Hsp104 or Hsp104-ClpB chimeras (1 μM each) alone and together with Hsp70 and Ydj-1 (1 μM each) or DnaK (0.86 μM), DnaJ (0.43 μM), and GrpE (0.25 μM). Assay conditions were the same as in Fig. 1c. (c) Reactivation of chemically denatured FFL over time by ClpB or ClpBHM (1 μM each) alone and together with DnaK (0.86 μM), DnaJ (0.43 μM), and GrpE (0.25 μM) or Hsp70 and Ydj-1 (1 μM each). ClpB and ClpBHM were purified as described in the supplementary material. Assay conditions were the same as in Fig. 3a. (d) Reactivation of heat-aggregated MDH by ClpB or ClpBHM (1 μM each) alone and together with DnaK (0.86 μM), DnaJ (0.43 μM), and GrpE (0.25 μM) or Hsp70 and Ydj-1 (1 μM each). Assay conditions were the same as in Fig. 1c. Standard errors of three independent assays are shown in Fig. 3a-d.
Remarkably, all three Hsp104-ClpB chimeras cooperate synergistically with the DnaK system to solubilize aggregated model substrates but lost their ability to function together with Hsp70/Hsp40 (Fig. 3a and b). Notably, wild-type Hsp104 cooperates only with Hsp70/Hsp40 and not with the bacterial DnaK system (Fig. 3a and b). Moreover, some substrate-specific differences were also observed. For instance, DnaK/DnaJ/GrpE can at least partially solubilize chemically denatured FFL, whereas the contributions of Hsp70/Hsp40 are negligible (Fig. 3a and c). It is conceivable that differences in substrate-specificity together with the reduced hexamer stability of our engineered chimeras (Fig. 2b) could explain the lower efficiency of the chimeric bi-chaperone system compared to native Hsp104/Hsp70/Hsp40 (Fig. 3a and b). Nevertheless, in our assays, the chimeric bi-chaperone system was always more efficient than the Hsp70 or DnaK system alone (Fig. 3a and b).
To confirm whether the M-domain is responsible for switching the species-specificity of the bi-chaperone system, we asked whether the same is true for ClpB also. To address this issue, we replaced the ClpB M-domain against that of Hsp104 (ClpBHM) and analyzed the chimera using our chaperone activity assays. As anticipated, we found that ClpBHM cooperates synergistically with Hsp70/Hsp40 but not with DnaK/DnaJ/GrpE, whereas wild-type ClpB cooperates only with the bacterial DnaK system (Fig. 3c and d). While the reactivation efficiency of denatured FFL by ClpBHM/Hsp70/Hsp40 (Fig. 3c) was similar to that observed with the Hsp104 chimeras together with the DnaK system (Fig. 3a), reactivation of heat-aggregated MDH by ClpBHM/Hsp70/Hsp40 was nearly as efficient as the native bacterial bi-chaperone network (Fig. 3d). The latter is consistent with a different substrate preference for the yeast and the bacterial bi-chaperone system.
Taken together, our results demonstrate that the M-domain is essential for the synergistic interaction with the Hsp70/DnaK chaperone system, and represents the minimal region required to switch the species-specificity of the bi-chaperone network.
The M-domain couples Hsp70/Hsp40 to the Hsp104 protein remodeling activity
To determine whether the M-domain controls the ATP-driven Hsp104 motor activity required for substrate translocation through the Hsp104 hexamer, we generated Hsp104, Hsp104Trap, Hsp104ΔM, and all of our Hsp104-ClpB chimeras as HAP variants by replacing Gly739 to Lys741 with Ile-Leu-Phe. We found that all of our engineered HAP variants, including HAPΔM, can translocate and degrade FITC-casein in the presence of ClpP (Fig. 4a). Neither ClpP alone nor HAPTrap are able to do so, indicating that substrate translocation is ATP hydrolysis driven, but does not require an M-domain or the presence of the Hsp70/Hsp40 system.
Fig. 4.
Disaggregation requires M-domain mediated collaboration with the Hsp70/DnaK system. (a) Degradation of FITC-casein (0.12 mg/ml) by ClpP (2 μM) alone and together with HAPTrap, HAP, HAPΔM, or HAP-ClpB chimeras (1 μM each). FITC-casein was incubated in buffer A at 22 °C for 60 min with chaperones as indicated, together with an ATP-regenerating system (6 mM phosphoenolpyruvate and 0.5 μM pyruvate kinase), and 3 mM ATP. Reactions were analyzed as described35 by measuring the fluorescence signal of acid-soluble peptides using a LS55 fluorescence spectrometer (Perkin Elmer, Waltham, MA). Standard errors of three independent assays are shown. (b) Degradation of heat-aggregated FAM-labeled MDH (1.5 μM; labeling is described in the supplementary material) by ClpP (2 μM) and HAP variants (1 μM) alone and together with DnaK (0.86 μM), DnaJ (0.43 μM), and GrpE (0.25 μM) or Hsp70 and Ydj-1 (1 μM each). Reactions were set up and analyzed the same way as described for the FITC-casein degradation assay (Fig. 4a). Standard errors of three independent assays are shown.
Unlike soluble substrates, it was previously shown that the degradation of protein aggregates by HAP/ClpP is Hsp70/Hsp40-dependent.13; 14 Consistently, we found that the HAP/ClpP-mediated degradation of heat-aggregated FAM-labeled MDH requires the M-domain, and is strictly dependent on the cooperation with the cognate Hsp70 system (Fig. 4b). Notably, we found that our Hsp104-ClpB chimeras alter the species-specificity of the bi-chaperone network so that our HAP chimeras now cooperate with the bacterial DnaK system, but no longer with eukaryotic Hsp70/Hsp40.
Together, our results demonstrate that the M-domain controls the Hsp104 protein-disaggregating activity through functional interaction with the Hsp70 system in a species-specific manner.
Mechanistic model for protein disaggregation
Our results show that the Hsp104 M-domains are responsible for conferring the species-specificity of the bi-chaperone network, presumably through direct interaction with the Hsp70 system. What might be the purpose of such an interaction? It has been proposed that Hsp104 recognizes substrates or is recruited to substrates in an Hsp70/Hsp40-dependent manner followed by substrate translocation through the Hsp104 hexamer.13; 14; 30 Alternatively, although not mutually exclusive, it has been proposed that the Hsp104 M-domains are required to break apart large aggregates into smaller ones, followed by substrate translocation and hand-over to the Hsp70 system.10; 18 Consistent with both activities, we found that the M-domains are located on the Hsp104 exterior.12 Notably, however, an Hsp70/Hsp40-dependent Hsp104-substrate interaction appears non-essential for protein disaggregation.31; 32 The latter is supported by our Hsp104-T4L chimeras that can solubilize heat-aggregated substrates in the absence of the Hsp70 system (Fig. 1a-c).
What might be the function of an Hsp40/Hsp70/Hsp104 M-domain interaction? We previously reported that the ability of Hsp104T4L to solubilize heat-aggregated β-gal required the presence of the T4L moiety within the M-domain helix 2.12 It is therefore tempting to speculate that the T4L moiety mimics a biologically relevant interaction which is required for protein disaggregation. Consistent with our results obtained with the altered specificity chimeras, our findings point at a potential role of Hsp70/Hsp40 in activating the Hsp104 protein remodeling activity by binding to the M-domain. The latter is reminiscent to the known function of Vta1 in modulating the Vps4 AAA+ ATPase through binding to the β-domain that is inserted in the same place of Vps417 as is the M-domain in Hsp104.
Together, our results provide an explanation for the up-stream role of Hsp70/Hsp40 in Hsp104 function, and support a regulatory mechanism common to diverse AAA+ machines.
Supplementary Material
Acknowledgements
We thank Dr. S. Lee for advice on chimera design and helpful discussion, Dr. J. Lee for generating the Hsp70 construct, Mr. J.M. Choi for generating the ClpB construct, and Dr. T. Wensel for access to the plate reader. Work in the laboratory of F.T.F.T. is supported by grants from the National Institutes of Health (R01-AI076239), the Welch Foundation (Q-1530), the Department of Defense, and the American Cancer Society. B.S. was supported in part by a training fellowship from the Pharmacoinformatics Training Program of the Keck Center of the Gulf Coast Consortia (DK071505).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Bosl B, Grimminger V, Walter S. The molecular chaperone Hsp104--a molecular machine for protein disaggregation. J. Struct. Biol. 2006;156:139–148. doi: 10.1016/j.jsb.2006.02.004. [DOI] [PubMed] [Google Scholar]
- 2.Doyle SM, Wickner S. Hsp104 and ClpB: protein disaggregating machines. Trends Biochem. Sci. 2009;34:40–48. doi: 10.1016/j.tibs.2008.09.010. [DOI] [PubMed] [Google Scholar]
- 3.Haslberger T, Bukau B, Mogk A. Towards a unifying mechanism for ClpB/Hsp104-mediated protein disaggregation and prion propagation. Biochem. Cell Biol. 2010;88:63–75. doi: 10.1139/o09-118. [DOI] [PubMed] [Google Scholar]
- 4.Wickner RB, Edskes HK, Shewmaker F, Nakayashiki T. Prions of fungi: inherited structures and biological roles. Nat. Rev. Microbiol. 2007;5:611–618. doi: 10.1038/nrmicro1708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chernoff YO, Lindquist SL, Ono B, Inge-Vechtomov SG, Liebman SW. Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [PSI+] Science. 1995;268:880–884. doi: 10.1126/science.7754373. [DOI] [PubMed] [Google Scholar]
- 6.Shorter J, Lindquist S. Hsp104 catalyzes formation and elimination of self-replicating Sup35 prion conformers. Science. 2004;304:1793–1797. doi: 10.1126/science.1098007. [DOI] [PubMed] [Google Scholar]
- 7.Inoue Y, Taguchi H, Kishimoto A, Yoshida M. Hsp104 binds to yeast Sup35 prion fiber but needs other factor(s) to sever it. J. Biol. Chem. 2004;279:52319–52323. doi: 10.1074/jbc.M408159200. [DOI] [PubMed] [Google Scholar]
- 8.Wendler P, Shorter J, Plisson C, Cashikar AG, Lindquist S, Saibil HR. Atypical AAA+ subunit packing creates an expanded cavity for disaggregation by the protein-remodeling factor Hsp104. Cell. 2007;131:1366–1377. doi: 10.1016/j.cell.2007.10.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wendler P, Shorter J, Snead D, Plisson C, Clare DK, Lindquist S, Saibil HR. Motor mechanism for protein threading through Hsp104. Mol. Cell. 2009;34:81–92. doi: 10.1016/j.molcel.2009.02.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lee S, Sowa ME, Watanabe Y, Sigler PB, Chiu W, Yoshida M, Tsai FTF. The structure of ClpB: a molecular chaperone that rescues proteins from an aggregated state. Cell. 2003;115:229–240. doi: 10.1016/s0092-8674(03)00807-9. [DOI] [PubMed] [Google Scholar]
- 11.Lee S, Choi JM, Tsai FTF. Visualizing the ATPase cycle in a protein disaggregating machine: Structural basis for substrate binding by ClpB. Mol. Cell. 2007;25:261–271. doi: 10.1016/j.molcel.2007.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee S, Sielaff B, Lee J, Tsai FTF. CryoEM Structure of Hsp104 and Its Mechanistic Implication for Protein Disaggregation. Proc. Natl. Acad. Sci. USA. 2010;107:8135–8140. doi: 10.1073/pnas.1003572107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tipton KA, Verges KJ, Weissman JS. In vivo monitoring of the prion replication cycle reveals a critical role for Sis1 in delivering substrates to Hsp104. Mol. Cell. 2008;32:584–591. doi: 10.1016/j.molcel.2008.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tessarz P, Mogk A, Bukau B. Substrate threading through the central pore of the Hsp104 chaperone as a common mechanism for protein disaggregation and prion propagation. Mol. Microbiol. 2008;68:87–97. doi: 10.1111/j.1365-2958.2008.06135.x. [DOI] [PubMed] [Google Scholar]
- 15.Lum R, Tkach JM, Vierling E, Glover JR. Evidence for an unfolding/threading mechanism for protein disaggregation by Saccharomyces cerevisiae Hsp104. J. Biol. Chem. 2004;279:29139–29146. doi: 10.1074/jbc.M403777200. [DOI] [PubMed] [Google Scholar]
- 16.Lum R, Niggemann M, Glover JR. Peptide and protein binding in the axial channel of Hsp104: insights into the mechanism of protein unfolding. J. Biol. Chem. 2008;283:30139–30150. doi: 10.1074/jbc.M804849200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Scott A, Chung HY, Gonciarz-Swiatek M, Hill GC, Whitby FG, Gaspar J, Holton JM, Viswanathan R, Ghaffarian S, Hill CP, Sundquist WI. Structural and mechanistic studies of VPS4 proteins. EMBO J. 2005;24:3658–3669. doi: 10.1038/sj.emboj.7600818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Glover JR, Lindquist S. Hsp104, Hsp70, and Hsp40: a novel chaperone system that rescues previously aggregated proteins. Cell. 1998;94:73–82. doi: 10.1016/s0092-8674(00)81223-4. [DOI] [PubMed] [Google Scholar]
- 19.Krzewska J, Langer T, Liberek K. Mitochondrial Hsp78, a member of the Clp/Hsp100 family in Saccharomyces cerevisiae, cooperates with Hsp70 in protein refolding. FEBS Lett. 2001;489:92–96. doi: 10.1016/s0014-5793(00)02423-6. [DOI] [PubMed] [Google Scholar]
- 20.Schlee S, Beinker P, Akhrymuk A, Reinstein J. A chaperone network for the resolubilization of protein aggregates: direct interaction of ClpB and DnaK. J. Mol. Biol. 2004;336:275–285. doi: 10.1016/j.jmb.2003.12.013. [DOI] [PubMed] [Google Scholar]
- 21.Kedzierska S, Chesnokova LS, Witt SN, Zolkiewski M. Interactions within the ClpB/DnaK bi-chaperone system from Escherichia coli. Arch. Biochem. Biophys. 2005;444:61–65. doi: 10.1016/j.abb.2005.10.005. [DOI] [PubMed] [Google Scholar]
- 22.Doyle SM, Hoskins JR, Wickner S. Collaboration between the ClpB AAA+ remodeling protein and the DnaK chaperone system. Proc. Natl. Acad. Sci. USA. 2007;104:11138–11144. doi: 10.1073/pnas.0703980104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hung GC, Masison DC. N-terminal domain of yeast Hsp104 chaperone is dispensable for thermotolerance and prion propagation but necessary for curing prions by Hsp104 overexpression. Genetics. 2006;173:611–620. doi: 10.1534/genetics.106.056820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cashikar AG, Schirmer EC, Hattendorf DA, Glover JR, Ramakrishnan MS, Ware DM, Lindquist SL. Defining a pathway of communication from the C-terminal peptide binding domain to the N-terminal ATPase domain in a AAA protein. Mol. Cell. 2002;9:751–760. doi: 10.1016/s1097-2765(02)00499-9. [DOI] [PubMed] [Google Scholar]
- 25.Watanabe Y, Takano M, Yoshida M. ATP binding to nucleotide binding domain (NBD)1 of the ClpB chaperone induces motion of the long coiled-coil, stabilizes the hexamer, and activates NBD2. J. Biol. Chem. 2005;280:24562–24567. doi: 10.1074/jbc.M414623200. [DOI] [PubMed] [Google Scholar]
- 26.Haslberger T, Weibezahn J, Zahn R, Lee S, Tsai FTF, Bukau B, Mogk A. M domains couple the ClpB threading motor with the DnaK chaperone activity. Mol. Cell. 2007;25:247–260. doi: 10.1016/j.molcel.2006.11.008. [DOI] [PubMed] [Google Scholar]
- 27.Watanabe Y-H, Nakazaki Y, Suno R, Yoshida M. Stability of the two wings of the coiled-coil domain of ClpB chaperone is critical for its disaggregation activity. Biochem J. 2009;421:71–77. doi: 10.1042/BJ20082238. [DOI] [PubMed] [Google Scholar]
- 28.Zolkiewski M. ClpB cooperates with DnaK, DnaJ, and GrpE in suppressing protein aggregation. J. Biol. Chem. 1999;274:28083–28086. doi: 10.1074/jbc.274.40.28083. [DOI] [PubMed] [Google Scholar]
- 29.Goloubinoff P, Mogk A, Zvi AP, Tomoyasu T, Bukau B. Sequential mechanism of solubilization and refolding of stable protein aggregates by a bichaperone network. Proc. Natl. Acad. Sci. USA. 1999;96:13732–13737. doi: 10.1073/pnas.96.24.13732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Higurashi T, Hines JK, Sahi C, Aron R, Craig EA. Specificity of the J-protein Sis1 in the propagation of 3 yeast. Proc Natl Acad Sci U S A. 2008;105:16596–16601. doi: 10.1073/pnas.0808934105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Doyle SM, Shorter J, Zolkiewski M, Hoskins JR, Lindquist S, Wickner S. Asymmetric deceleration of ClpB or Hsp104 ATPase activity unleashes protein-remodeling activity. Nat. Struct. Mol. Biol. 2007;14:114–122. doi: 10.1038/nsmb1198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Schaupp A, M. M, Grimminger V, Bösl B, Walter S. Processing of proteins by the molecular chaperone Hsp104. J. Mol. Biol. 2007;370:674–686. doi: 10.1016/j.jmb.2007.04.070. [DOI] [PubMed] [Google Scholar]
- 33.Schlothauer T, Mogk A, Dougan DA, Bukau B, Turgay K. MecA, an adaptor protein necessary for ClpC chaperone activity. Proc. Natl. Acad. Sci. USA. 2003;100:2306–2311. doi: 10.1073/pnas.0535717100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bosl B, Grimminger V, Walter S. Substrate binding to the molecular chaperone Hsp104 and its regulation by nucleotides. J. Biol. Chem. 2005;280:38170–38176. doi: 10.1074/jbc.M506149200. [DOI] [PubMed] [Google Scholar]
- 35.Twining SS. Fluorescein isothiocyanate-labeled casein assay for proteolytic enzymes. Anal. Biochem. 1984;143:30–34. doi: 10.1016/0003-2697(84)90553-0. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.