Abstract
Male fertility is declining and an underlying cause may be due to environment-epigenetic interactions in developing sperm, yet nothing is known of how the epigenome controls gene expression in sperm development. Histone methylation and acetylation are dynamically regulated in spermatogenesis and are sensitive to the environment. Our objectives were to determine how histone H3 methylation and acetylation contribute to the regulation of key genes in spermatogenesis. A germ cell line, GC-1, was exposed to either the control, or the chromatin modifying drugs tranylcypromine (T), an inhibitor of the histone H3 demethylase KDM1 (lysine specific demethylase 1), or trichostatin (TSA), an inhibitor of histone deacetylases, (HDAC). Quantitative PCR (qPCR) was used to identify genes that were sensitive to treatment. As a control for specificity the Myod1 (myogenic differentiation 1) gene was analyzed. Chromatin immunoprecipitation (ChIP) followed by qPCR was used to measure histone H3 methylation and acetylation at the promoters of target genes and the control, Myod1. Remarkably, the chromatin modifying treatment specifically induced the expression of spermatogonia expressed genes Pou5f1 and Gfra1. ChIP-qPCR revealed that induction of gene expression was associated with a gain in gene activating histone H3 methylation and acetylation in Pou5f1 and Gfra1 promoters, whereas CpG DNA methylation was not affected. Our data implicate a critical role for histone H3 methylation and acetylation in the regulation of genes expressed by spermatogonia – here, predominantly mediated by HDAC-containing protein complexes.
Introduction
The formation of spermatozoa from spermatogonial stem cells throughout adult life is dependent on a tightly orchestrated cell differentiation process governed by unique transcriptional programs and extensive chromatin remodeling [1]. While current information indicates that male germ cell differentiation is driven by tight transcriptional regulation [2], [3], [4], [5], [6], the epigenetic modifications and chromatin remodelers underlying the control of this gene expression are unknown. The epigenetic layer includes modification of histones such as methylation, acetylation, phosphorylation among others, and DNA methylation [7]. Recent large scale genome profiling experiments have revealed general roles for histone modifications in gene regulation whereby histone H3 methylation on lysine 4 (K4) is gene activating, and methylation on lysine 9 (K9) is gene silencing [8], [9]. Overall histone acetylation is associated to an open chromatin state and active gene transcription [10], [11], [12], [13], [14]. Advancements on the readout of these specific marks during cell differentiation are just now being made. For example as embryonic stem cells (ESC) progress from a pluripotent state to one of specialization, this process is characterized by the reconfiguration of developmental genes from a bivalent marking consisting of activating H3K4 trimethylation and repressive H3K27 trimethylation, to a loss of repressive marks and a gain in gene function [15]. Furthermore, there is evidence that epigenetic mechanisms such as DNA methylation underlie the control of expression of key genes such as Pou5f1 (POU domain class 5 transcription factor 1, also known as Oct4) and Nanog [16], [17].
In the testis, the production of haploid male gametes includes three phases: mitotic proliferation of spermatogonia, meiotic reshuffling of the genome to create genetic diversity and spermiogenesis, where unique morphological changes occur to transform a haploid, immotile round spermatid to an elongated spermatozoa, capable of fertilizing an oocyte [18], [19]. Spermatogonial stem cells either self-renew or differentiate and the lifetime production of mature spermatozoa is dependent on these stem cells [20], [21]. Inadequate establishment or depletion of the stem cell pool will lead to reduced sperm output and infertility [2], [3], [5]. Knockout and knockdown studies have revealed several transcription factors and proteins involved in signal transduction that are key regulators of stem cell biology including Pou5f1 and glial cell line derived neurotrophic factor family receptor alpha 1 (Gfra1, also known as GDNF-family receptor α1) [4], [22], [23], [24]. The Pou5f1 transcription factor is strongly expressed in spermatogonia and its ablation results in apoptosis of primordial germ cells. As spermatogonia differentiate Pou5f1 expression is downregulated [6], [25], [26]. GDNF-family receptor α1 localizes to type A spermatogonia including A single, A paired and A aligned and it is co-expressed with Pou5f1 [4]. Interestingly, knockdown of Gfra1 in the testicular stem cell population induced a phenotypic switch towards spermatogonial differentiation as indicated by a decrease in proliferation and expression of Pou5f1, as well as an increase in differentiation markers. Moreover GDNF cell-signaling is critical in spermatogonial stem cell renewal [4], [27]. The epigenetic mechanisms involved in regulating these key genes in spermatogenesis are unknown.
Accumulating evidence from gene knockouts has shown that chromatin modifiers such as histone demethylases, histone methyltransferases and histone deacetylases have a critical role in male germ cell development [28], [29], [30], [31], [32], [33], [34]. However information on gene-specific targets and how these chromatin modifiers regulate transcriptional processes and differentiation in spermatogenesis is lacking.
We and others have previously determined that in developing male germ cells histone methylation is highly dynamic and tightly controlled [35], [36]. It is well established that the histone H3 demethylase KDM1 (also known as LSD1/AOF2/BHC110) and the histone deacetylase 1 (HDAC1) form a repressor complex and that their histone modification activities are intimately linked [37], [38], [39]. Recently, we have shown that KDM1 is highly expressed in spermatogonia and acts in a repressor complex containing HDAC1 [35]. KDM1 is known to remove activating mono- and dimethylation marks from H3K4, which results in repression of gene transcription [40]. Conversely, in concert with the androgen receptor KDM1 targets repressive H3K9 mono- and dimethylation marks thereby activating target genes [41].
Our working hypothesis was that by inhibiting chromatin modifiers, KDM1 and HDAC, we would alter the epigenetic landscape of a cultured male germ cell line, GC-1 [42], and in turn change gene expression to reveal genes that are regulated by histone methylation and acetylation. Our specific aim was to identify key genes in spermatogenesis that may be targets of this complex and in turn regulated by histone methylation and acetylation. From a panel of genes known to be expressed in spermatogenesis we showed that the spermatogonia expressed genes Pou5f1 and Gfra1 were sensitive to the chromatin modifying treatment. We then examined how the expression of these genes was influenced by the levels of histone H3 methylation and acetylation, and DNA methylation in the gene promoters.
Materials and Methods
Cell Culture
The GC-1 cell line was obtained from ATCC (CRL-2053; Manassas, VA, USA). GC-1 cells were generated by Hofmann et al. and originally derived from postnatal day 10 mouse testes. They have been described as an intermediate spermatogenic cell type between type B spermatogonia and preleptotene spermatocytes due to morphological and gene expression characteristics [42]. The cells were cultured in DMEM (11965; Invitrogen, Carlsbad, CA, USA), supplemented with 10% FBS (12484-028; Invitrogen), 100 U of penicillin/100 µg of streptomycin (15140-122; Invitrogen) and 100 µM non-essential amino acids (11140-050; Invitrogen, Carlsbad, CA, USA), and grown in a humidified atmosphere containing 5% CO2 at 37°C. GC-1 cells were seeded at 20% of confluence and allowed to grow for 24 hours before treatment.
The monoamine oxidase inhibitor tranylcypromine (T) has been reported as a potent suppressor of the enzymatic activity of KDM1 [43], whereas trichostatin A (TSA) is a well characterized inhibitor of histone deacetylases [44]. In order to target KDM1 and HDAC1 with T and/or TSA, we examined the influence of these inhibitors on GC-1 cell viability, morphology and proliferation. Since trichostatin A is known to induce apoptosis and to affect cell cycle progression [45], [46], [47], GC-1 cells were exposed to varying TSA concentrations (25 nM, 50 nM, 100 nM, and 300 nM) to determine the optimal minimal dose which caused a reduction in histone H3 acetylation while having the least impact on cell viability. Treatment with 50 nM TSA was determined as most efficient – due to global changes in histone H3 acetylation levels without cytotoxicity (Figure S1). Cell viability was not altered by T concentrations ranging from 10 µM to 1 mM. A working concentration of 100 µM T was selected as this dose has been shown to inhibit KDM1 and cause epigenetic changes at the promoters of specific genes [43], [48]. As has been reported before [48], [49] no global changes in histone methylation occurred following treatment with T (Figures S1, S2). Medium was removed and cells were treated for 16 hours with either 100 µM T (13492-01-8, Biomol, Plymouth Meeting, PA, USA), or 50 nM TSA (T8552; Sigma-Aldrich, Saint Louis, MO, USA), or 0.000754% dimethylsulphoxide (DMSO) (D2650; Sigma-Aldrich), the solvent of TSA, or a cocktail of 100 µM T and 50 nM TSA. After 16 h of exposure, treated GC-1 cells were morphologically indistinguishable from control cells and cell counts after trypan blue staining revealed no differences in cell viability between treated and control cells.
Western Blotting
Total protein of control or treated GC-1 cells were extracted with Laemmli buffer (62.5 mM Tris-HCl, pH 6.8/2% SDS/25% Glycerol/0.01% Bromophenol blue) containing 5% of β-Mercaptoethanol. Proteins were resolved by standard SDS-PAGE and electroblotted onto nitrocellulose membranes as previously described [35]. The membranes were incubated with the primary antibody followed by the corresponding horseradish peroxidase-conjugated secondary antibody (all antibodies are listed in Table S1). Beta-Actin was used for normalization of KDM1 and HDAC1 levels, whereas the total amount of histone H3 was used for normalization of Western blots for histone H3 modifications. Antibody signals were detected using SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific, Rockford, IL, USA). Band intensities were measured using the AlphaDigiDoc analysis system (Alpha Innotech Corporation, San Leandro, CA, USA). If not otherwise indicated each experiment was replicated a minimum of three times.
Immunoprecipitation Assay
1×10∧6 GC-1 cells were harvested in 1 ml modified RIPA (50 mM Tris-HCl, pH 7.5/300 mM NaCl/1% NP-40/50 mM NaF (sodium fluoride)/1 mM PMSF/100 µM NaVO3/proteinase inhibitor cocktail (P8340; Sigma-Aldrich), sonicated (6×(5 sec on/5 sec off), amplitude: 25%) and kept on ice for 30 min. Soluble extracts were separated by centrifugation at 10000× g for 20 min. 250 µl of total cell extract were used for each IP or the corresponding control and each volume was made up to 500 µl with RIPA containing the full complement of inhibitors. Then, cell extracts were pre-cleared for 1 h on protein A-Agarose beads (Roche Diagnostics, Mannheim, Germany) and collected as supernatant after centrifugation. The supernatants were incubated over night at 4°C with either 5 µg of anti-HDAC1 (clone 2E10; Millipore, Billerica, MA, USA), or 5 µg of anti-KDM1 (Abcam, Cambridge, MA, USA) antibody, or the appropriate controls, which included mouse IgG and rabbit IgG (Jackson ImunoResearch Laboratories, West Grove, PA, USA). Protein A-Agarose beads were pre-adsorbed with 0.05% BSA prior to capturing the immune complex for 1 h at 4°C. The beads were then washed three times with RIPA buffer and captured proteins were eluted in SDS-PAGE sample buffer for Western blot analysis.
Reverse transcription and real-time PCR
Total RNA was extracted using Purezol (BioRad, Hercules, CA, USA) according to the manufacturer's instructions. Then, 2 µg of total RNA was reverse transcribed using the High Capacity cDNA Reverse Transcription kit, which includes random primers (Applied Biosystems, Foster City, CA, USA). Primers were designed using Primer Express software (Applied Biosystems). Primer information is given in Table S2. PCR reactions were performed following the Power SYBR Green PCR Master Mix protocol (Applied Biosystems) with an ABI Prism 7500 apparatus (Applied Biosystems). The measured amount of each cDNA was normalized to Gapdh. In order to avoid pseudogene amplification, Pou5f1 expression was additionally analyzed using the primer Mm00658129_gH Pou5f1 TaqMan gene expression assay (Applied Biosystems) and the TaqMan Fast Universal PCR Master Mix (Applied Biosystems), following manufacturer's instructions. The 18S rRNA TaqMan gene expression assay (primer Hs99999901_s1, Applied Biosystems) was used for normalization. If not otherwise indicated, each experiment was repeated independently a minimum of three times, and each sample and the corresponding negative controls were run in triplicates.
In silico promoter analysis
We performed an in silico analysis using the CSH TRED database [50], [51] (http://rulai.cshl.edu/cgibin/TRED/tred.cgi?process=home) to localize the transcriptional start sites (TSS) of Gfra1 (Gfra1 promoter ID 63194) and Myod1 (Myod1 promoter ID 76703), which were concordant with start sites of Gfra1 and Myod1 transcripts as given by ensemble genome browser (http://www.ensembl.org/index.html) and the UCSC genome browser (http://genome.ucsc.edu/index.html?org=Mouse&db=mm9&hgsid=125767812).
Chromatin Immunoprecipitation (ChIP) assay followed by real-time PCR
ChIPs were performed following the optimized instructions to measure the levels of histone methylation and acetylation at targeted gene enhancer and promoter regions in GC-1 cells following treatment. Moreover, ChIP assays were used to confirm the presence of KDM1- and/or HDAC1- at Pou5f1 and Gfra1 gene promoters in GC-1 cells. Cells were rinsed with ice-cold PBS once and fixed in 1% formaldehyde for 10 min at room temperature to cross-link DNA and proteins. Glycine (0.125 M) was then added to quench the formaldehyde. Subsequently, the cells were rinsed twice in ice-cold PBS, scraped and collected by centrifugation at 4°C. Cells were then resuspended in 2 mL of FA lysis buffer (50 mM HEPES pH 7.5, 140 mM NaCl, 1 mM EDTA, 1% triton X-100, 0.1% sodium deoxycholate 10%, 0.1% SDS and protease inhibitor cocktail (P8340; Sigma-Aldrich). The chromatin was sheared by sonication to an average fragment size of 500 to 1000 bp. 50 µL of sample was removed to serve as input control, whereas 50 µg of protein was used for each immunoprecipitation, including IgG and serum controls. After a dilution 1∶10 of each sample with RIPA buffer (50 mM Tris-HCl pH 8, 150 mM NaCl, 2 mM EDTA ph8, 1% NP-40, 0.5% sodium deoxycholate, 0.1%SDS and protease inhibitors), antibodies were added to the corresponding sample (find detailed antibody information in Table S1). 20 µL of protein A-Agarose beads (pre-absorbed with BSA (NEB, Pickering, Ontario, Canada) and sonicated single stranded herring sperm DNA (ab46666, Abcam) was added to all samples that were then rotated overnight at 4°C. After three washings with a buffer containing 0.1% SDS/1% Triton X-100/2 mM EDTA pH 8/150 mM NaCl/20 mM Tris-HCl pH 8 and one with a final wash buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA pH 8, 500 mM NaCl, 20 mM Tris-HCl pH 8) the chromatin-antibody complexes were eluted and the DNA-proteins cross-links were reversed for all the samples (including the input) at 65°C overnight in presence of 1 µg of RNase A (Sigma-Aldrich). Genomic DNA was recovered and purified with the QIAquick PCR purification kit (Qiagen, Hilden, Germany). Quantitative real-time PCR amplifications were performed using primers specific for Pou5f1 and Gfra1 promoters, or Gapdh control primers (Table S2), which were designed using Primer Express Software (Applied Biosystems). Primers spanned regions −2281 to −2212 and −854 to −781 relative to the Pou5f1 transcription start site, and −1071 to −997 and +104 to +179 relative to the putative Gfra1 transcription start site. In order to analyze a putative promoter occupancy of KDM1- and/or HDAC1-containig protein complexes approximately 3000 bp of Pou5f1 and Gfra1 promoter regions were screened by ChIP-qPCR for KDM1- and/or HDAC1 abundance. Power SYBR Green PCR Master Mix (Applied Biosystems) was used with an ABI Prism 7500 apparatus (Applied Biosystems) following manufacturer's instructions. Relative quantification of the genomic DNA received by ChIP was performed with the comparative CT method. Briefly, after confirmation of the PCR efficiency for each primer pair used in this study (Table S2), the ΔCT values were determined by subtracting the average CT value of the normalized input from the average CT value of the corresponding IP samples or the IgG and normal serum controls, respectively. In order to determine the fold change of target DNA in treated GC-1 cells over target DNA in DMSO controls, the ΔΔCT was calculated by subtracting the ΔCT of the DMSO from the ΔCT of TSA or TSA and T treated cells. The amount of target DNA is finally given by the formula 2∧-(ΔΔCT).
One experiment comprised the following treatments: cells exposed to DMSO, or TSA, or a cocktail of T and TSA. For one experiment two independent plates per treatment have been pooled. Each ChIP was performed on three independent experiments.
Pyrosequencing
CpG islands in Pou5f1 or Gfra1 promoter regions were identified with ”The sequence manipulation suite” database (http://www.bioinformatics.org/sms/index.html) [52] using the method described by Gardiner-Garden and Frommer [53]. Promoter methylation was determined by pyrosequencing for allele quantification (PSQ H96A, Biotage, Sweden), which is a real-time sequencing-based DNA analysis that quantifies multiple and consecutive CpG sites individually. Briefly 1000 ng of sample DNA was bisulfite treated using the Zymo DNA Methylation Kit (Zymo research, Orange, CA, USA). Bisulfite treated DNA was eluted in 10 µl volume and 1 µl of it is used for each PCR. PCR was performed using 10X PCR buffer, 3.0 mM MgCl2, 200 µM of each dNTP, 0.2 µM each of forward and reverse primers, HotStar DNA polymerase (Qiagen) 1.25 U, and ∼10 ng of bisulfite converted DNA per 50 µl reaction. PCR cycling conditions were: 94°C 15 min; 45 cycles of 94°C 30 s; 53°C 30 s; 72°C 30 s; 72°C 5 min; and then products were held at 4°C. The PCR was performed with one of the PCR primers biotinylated to convert the PCR product to single-stranded DNA templates. The PCR products (each 10 µl) were sequenced by Pyrosequencing PSQ96 HS System (Biotage AB) following the manufacturer's instructions (Biotage, Kungsgatan, Sweden). The methylation status of each locus was analyzed individually as a T/C SNP using QCpG software (Biotage, Kungsgatan, Sweden). Assay design and pyrosequencing sequencing was performed by EpigenDX (Worcester, MA, USA). The methylation assay ASY585 covered the region -190 to +26 relative to the Pou5f1 CpG transcriptional start site and the ADS932 assay comprised the region +26 to +143 relative to the putative Gfra1 transcriptional start site.
Statistical analyses
If not otherwise indicated, all experiments were repeated at least three times. All values are expressed as means ± SEM. To detect significant effects of treatments a one-way analysis of variance (ANOVA) was performed and to identify differences between means the Student-Newman-Keuls test was applied.
Results
The KDM1/HDAC1 complex is present in cultured GC-1
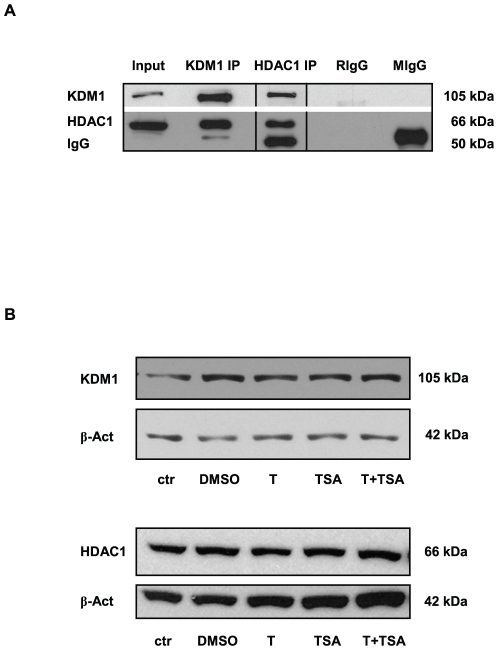
In vivo co-immunoprecipitation experiments on total GC-1 protein extracts demonstrated that in this cell line as in the testis KDM1 is associated with HDAC1 (Figure 1A). To determine whether targeting of the KDM1/HDAC1 repressor complex by T and/or TSA altered KDM1 or HDAC1 expression or protein stability, Western blot analysis was performed on total protein extracts isolated from GC-1 cells grown in regular culture medium, or culture medium supplemented with either T, TSA, DMSO (the solvent of TSA), or T and TSA. Treatment did not alter KDM1 or HDAC1 protein levels indicating that targeting the enzymatic activities of KDM1 and HDAC1, either individually or combined, does not influence their expression (Figure 1B).
Figure 1. KDM1 and HDAC1 form heteromeric complexes in GC-1 cells.
(A) Endogenous KDM1 and HDAC1 were co-immunoprecipitated with specific antibodies against KDM1 and HDAC1. Total GC-1 protein extract was used as input control. Immunoglobulin G (IgG), Rabbit IgG (RIgG), mouse IgG (MIgG). (B) Western blot analyses of HDAC1 and KDM1 in protein extracts isolated from GC-1 cells grown in culture medium (control, ctr), or in culture medium containing either dimethylsulphoxide (DMSO; TSA solvent), or tranylcypromine (T), or trichostatin A (TSA), or tranylcypromine and TSA (T+TSA). Western blots were reprobed with an antibody against beta-Actin as a loading control. Treatment of GC-1 cells with tranylcypromine and TSA does not alter protein levels of KDM1 and HDAC1.
Identification of spermatogenic genes sensitive to regulation by histone methylation and acetylation by treatment with trichostatin A and/or tranylcypromine
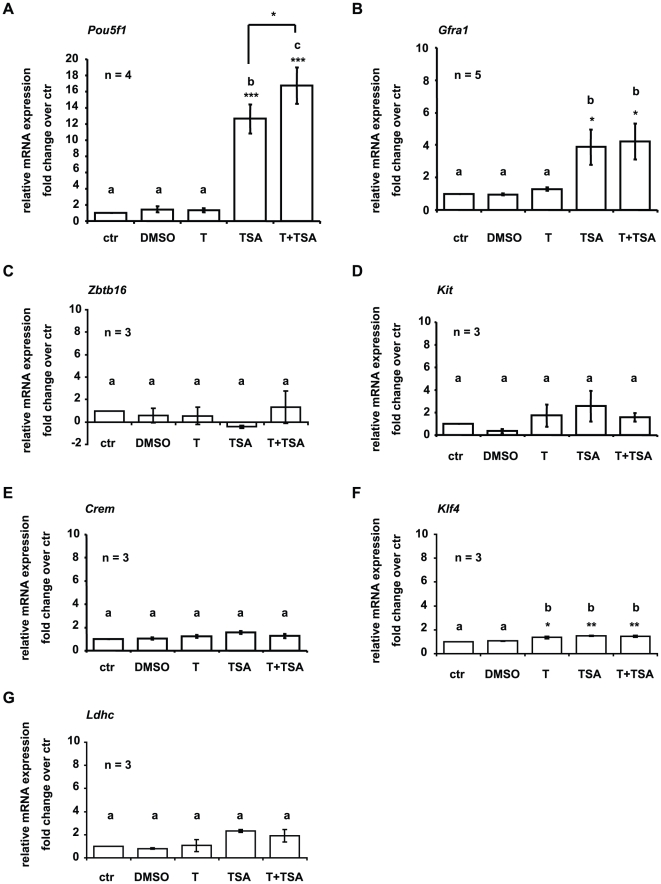
In order to identify target genes sensitive to the drug treatment we selected seven genes that have been shown to be expressed at defined stages of spermatogenesis. The Myod1 gene, which is not expressed in germ cells, was used as control for specificity of treatment to ensure that the drug treatments were not creating an overall more open chromatin state. Under control conditions we detected very low to no expression of the following spermatogonia markers (Table S3): Pou5f1 [6], [54], [55], zinc finger and BTB domain containing 16 (Zbtb16) [3], [54], or kit oncogene (Kit), [56], [57], [58], [59], [60]. However, Gfra1 [4], [61], [62], [63], [64] was weakly expressed in untreated GC-1 cells (Table S3). On the other hand transcripts of the following genes known to be expressed in spermatocytes were found under control conditions (Table S3): cAMP responsive element modulator (Crem) [65], [66] and in agreement with Hofmann and co-workers (1992) [42] low levels of testicular lactate dehydrogenase-C4 isozyme (Ldhc) [67]. Interestingly, Krueppel-like factor 4 (Klf4) is also expressed in untreated GC-1 cells (Table S3). In the mouse testis, Klf4 is strongly abundant in round spermatids [68], [69], but it is also found at low levels in cultured spermatogonial stem cells [70]. Remarkably, treatment of GC-1 cells with TSA or TSA in combination with T induced the expression of spermatogonia cell markers Pou5f1 (TSA induced a 12.6-fold increase, Figure 2A) and Gfra1 (TSA, or TSA and T combined induced a 4-fold increase, Figure 2B) and the maximal expression of Pou5f1 was induced by a combined treatment of TSA and T (16.8-fold increase, Figure 2A). There was no induction of Pou5f1 or Gfra1 expression when GC-1 cells were exposed to DMSO or T alone (Figure 2A,B). Treatment with TSA significantly increased expression of Gfra1 (4-fold increase) but combined treatment with TSA and T did not differ from TSA alone (Figure 2B). No induction or alterations in gene expression of the other marker genes were observed (Figure 2C–E,G) with the exception of Klf4 where a small but significant increase was detected following treatment with T (1.4-fold increase) or TSA alone (1.5-fold increase) and T and TSA combined (1.5-fold increase) (Figure 2F). (Ct values of genes analyzed in untreated GC-1 cells are summarized in Table S3. Changes in gene expression due to treatment are depicted as fold change over control conditions (untreated GC-1 cells) in Figure 2.)
Figure 2. Treatment induces Pou5f1 and Gfra1 gene expression in GC-1 cells.
QPCR analyses of spermatogenic gene expression in GC-1 cells, either grown in culture medium (control, ctr), or treated with tranylcypromine (T), or trichostatin A (TSA), or dimethylsulphoxid (DMSO; TSA solvent), or tranylcypromine and TSA (T+TSA). To avoid detection of expressed pseudogenes a TaqMan assay was performed to analyze Pou5f1 gene expression. Pou5f1 signals were normalized to 18S rRNA. All other genes were analyzed using SYBR green and signals were normalized to Gapdh. Values shown on the graphs are based on the change in expression in treated cells compared to the level of expression in control cells whereby the level of expression in control cells was defined as one. Thereby the fold change in expression indicates the ratio of normalized target gene expression in treated cells over normalized target gene expression in control cells. Data are expressed as mean ± SEM; *p<0.05, **p<0.01, ***p<0.001. Means with different letters are significantly different. N indicates the number of independent experiments, and per experiment each sample and the corresponding negative controls were run in triplicates. Trichostatin A treatment triggers Pou5f1 (A) and Gfra1 (B) gene expression in GC-1 cells. Treatment causes a slight but significant increase in Krueppel-like factor 4 (Klf4) (F) expression. Zinc finger and BTB domain containing 16 (Zbtb16) (C), kit oncogene (Kit) (D), cAMP responsive element modulator (Crem) (E), and lactate dehydrogenase C (Ldhc) (G) are not affected by treatment.
Trichostatin A treatment alters the epigenetic landscape at Pou5f1 and Gfra1 promoters
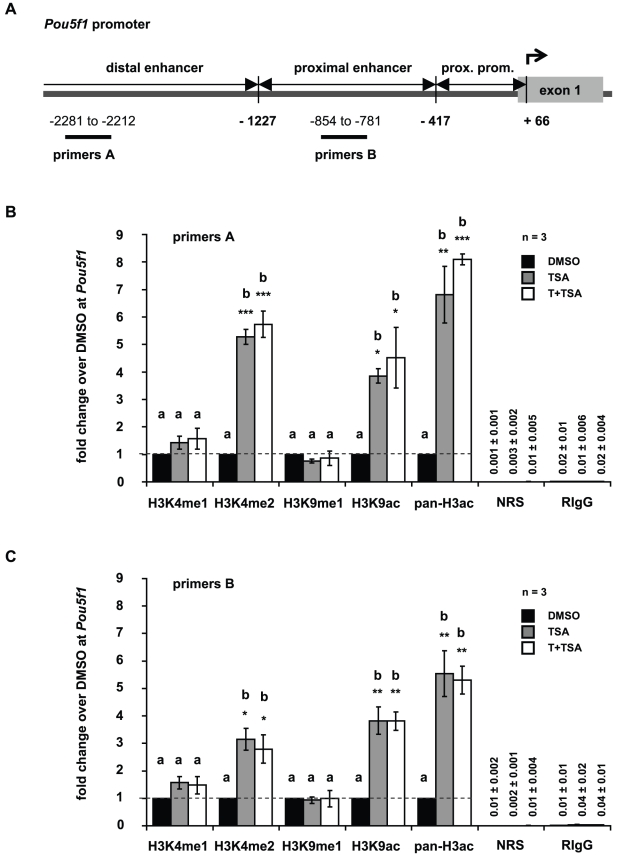
In all experiments performed in this study no differences were observed between GC-1 cells grown in culture medium and GC-1 cells exposed to DMSO. As the treatment with T alone did not induce changes in gene expression (Figure 2) or alter global histone methylation or acetylation levels (Figures S1, S2), we focused on TSA and the combined treatment of TSA and T in subsequent experiments to determine gene-specific alterations in histone methylation, histone acetylation, and DNA methylation at Pou5f1 and Gfra1 gene promoters. GC-1 cells grown in culture medium supplemented with DMSO were used as the control. To correlate epigenetic changes at these promoters with changes in gene expression, we performed chromatin immunoprecipitation (ChIP) followed by quantitative real-time PCR. The mouse Pou5f1 promoter has been well characterized and comprises defined regulatory regions (distal enhancer, proximal enhancer, and proximal promoter) that are critical for its activation [71], [72]. To cover the regulatory regions of the Pou5f1 promoter and a broad region of the Gfra1 promoter, ChIP-qPCR primers were positioned at intervals of approximately 1000 bp as depicted in Figures 3A and 4A, respectively. Since DNA fragments of 500–1000 bp were generated with sonication, epigenetic changes in the distal enhancer region of the Pou5f1 promoter were detected with primer pair A (−2281 to −2212 relative to TSS), and changes in the proximal enhancer/proximal promoter region with primer pair B (−854 to −781 relative to TSS) (Figure 3A). In order to identify epigenetic changes in the Gfra1 promoter, primer pair C (−1071 to −997 relative to TSS) and D (located in the 5′UTR of Gfra1 at +104 to +1011 relative to TSS) were used in qPCR analyses (Figure 4A).
Figure 3. Tranylcypromine and trichostatin A treatment influences posttranslational modifications of histone H3 at Pou5f1 promoter.
(A) Schematic representation of the Pou5f1 promoter. Numbers depict the positions of primer pairs used for ChIP-qPCR relative to the corresponding transcriptional start site. (B,C) Analyses of histone H3 methylation and acetylation levels in the Pou5f1 promoter by ChIP followed by qPCR in GC-1 cells that have been cultured in the presence of dimethylsulphoxide (DMSO, control, black bars), or trichostatin A (TSA, grey bars), or both inhibitors, tranylcypromine and TSA (T+TSA, white bars). Signals of target DNA received from DMSO control cells served as base and were defined as one. Fold change indicates 2∧-(ΔΔCT) of target DNA in treated cells over 2∧-(ΔΔCT) of target DNA in DMSO controls (Y-axis). Epigenetic modifications are depicted on the X-axis. Data are expressed as mean ± SEM; *p<0.05, **p<0.01, ***p<0.001. Means with different letters are significantly different. N indicates the number of independent experiments, and per experiment each sample and the corresponding negative controls were run in triplicates.
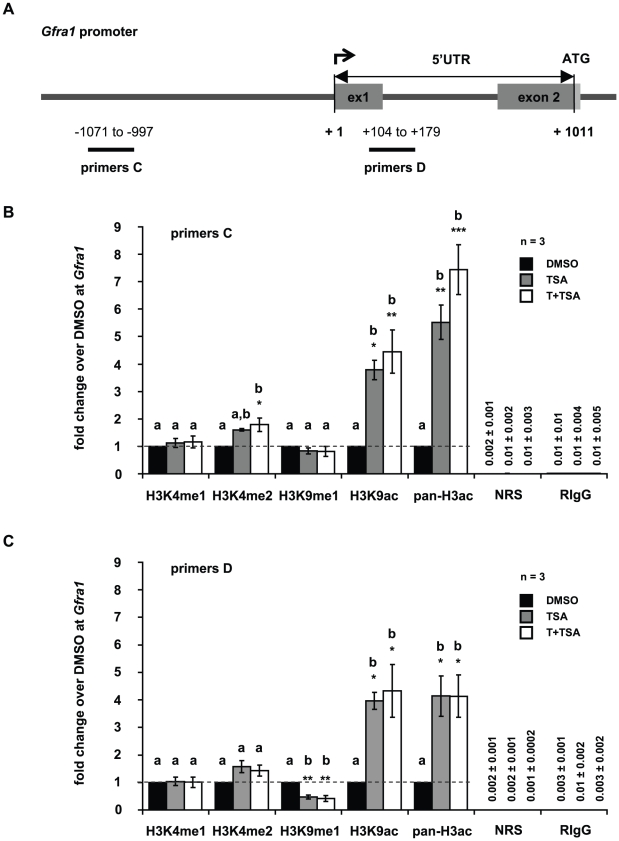
Figure 4. Tranylcypromine and trichostatin A treatment influences posttranslational modifications of histone H3 at Gfra1 promoter.
(A) Schematic representation of the Gfra1 promoter. Numbers depict the positions of primer pairs used for ChIP-qPCR relative to the corresponding transcriptional start site. (B,C) Analyses of histone H3 methylation and acetylation levels in the Gfra1 promoter by ChIP and qPCR in GC-1 cells that have been cultured in the presence of dimethylsulphoxide (DMSO, control, black bars), or trichostatin A (TSA, grey bars), or both inhibitors, tranylcypromine and TSA (T+TSA, white bars). Signals of target DNA received from DMSO control cells served as base and were defined as one. Fold change indicates 2∧-(ΔΔCT) of target DNA in treated cells over 2∧-(ΔΔCT) of target DNA in DMSO controls (Y-axis). Epigenetic modifications are depicted on the X-axis. Data are expressed as mean ± SEM; *p<0.05, **p<0.01, ***p<0.001. Means with different letters are significantly different. N indicates the number of independent experiments, and per experiment each sample and the corresponding negative controls were run in triplicates.
Using ChIP-qPCR we compared the levels of H3K4 mono- and dimethylation, H3K9 monomethylation, H3K9 acetylation, and pan-H3 acetylation at the promoters of Pou5f1 and Gfra1 between cells treated with either TSA, or TSA and T, or the DMSO control. Treatment of GC-1 cells with TSA, or TSA in combination with T significantly increased gene activating histone H3K4 dimethylation, H3K9 acetylation, and pan-H3 acetylation marks in the distal enhancer and the proximal enhancer/proximal promoter region of Pou5f1, whereas levels of H3K4 monomethylation and H3K9 monomethylation were indistinguishable from DMSO controls (Figure 3B,C). No enhanced effect on histone H3K4 methylation was found by treating with TSA in combination with T.
Analyzing the epigenetic signature of the Gfra1 promoter region after TSA exposure or the combined treatment of TSA and T by ChIP-qPCR, we identified a significant increase in activating H3K9 acetylation and pan-H3 acetylation marks in the putative promoter region upstream of the TSS and downstream in the 5′UTR of Gfra1 (Figure 4B,C). A slight (approximately twofold) but significant increase in H3K4 dimethylation was observed upstream of the Gfra1 TSS using the combined treatment of TSA and T. There were no alterations in either H3K4 monomethylation or H3K9 monomethyaltion in this region (Figure 4B). On the contrary, we identified a slight but significant decrease in the repressive H3K9 monomethylation mark in the 5′UTR of Gfra1 using TSA alone or in combination with T, but H3K4 mono- and dimethylation levels were indistinguishable from the DMSO controls (Figure 4C). Here, no enhanced effect on histone H3K4- or H3K9 methylation was found by treating the cells with TSA in combination with T (Figure 4C).
Effect of tranylcypromine and trichostatin A on gene expression and the epigenetic landscape of Myod1
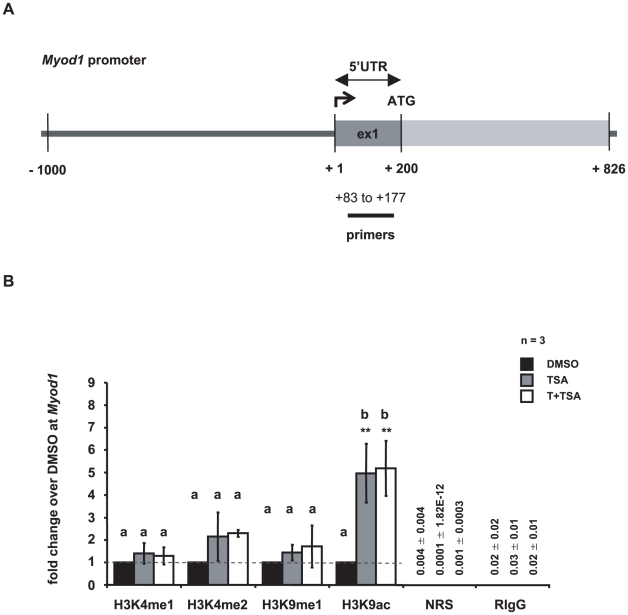
To demonstrate specificity of the effects on the expression of Pou5f1 and Gfra1 following treatment we examined the expression and epigenetic modifications of myogenic differentiation 1 (Myod1), which is important for muscle development (for review see [73]) and not expressed in GC-1 cells under control conditions (Table S4). ChIP analysis of the Myod1 promoter showed that treatment did not alter H3K4 mono- and dimethylation or H3K9 methylation. Treatment with TSA or T and TSA increased H3K9 acetylation (Figure 5). However these changes in acetylation alone did not induce Myod1 expression (Table S4).
Figure 5. Histone H3 methylation at Myod1 promoter is not affected by tranylcypromine and trichostatin A treatment.
(A) Schematic representation of the Myod1 promoter. Numbers depict the positions of primers used for ChIP-qPCR relative to the corresponding transcriptional start site. (B) ChIP analyses followed by qPCR reveal changes in the epigenetic landscape of the Myod1 promoter upon treatment. Despite the increase in H3K9 acetylation, Myod1 is not expressed in untreated GC-1 cells nor induced as a consequence of treatment, as shown in Table S4. Depicted are histone H3 methylation and acetylation levels in GC-1 cells that have been cultured in the presence of dimethylsulphoxide (DMSO, control, black bars), or trichostatin A (TSA, grey bars), or both inhibitors, tranylcypromine and TSA (T+TSA, white bars). Signals of target DNA received from DMSO control cells served as base and were defined as one. Fold change indicates 2∧-(ΔΔCT) of target DNA in treated cells over 2∧-(ΔΔCT) of target DNA in DMSO controls (Y-axis). Epigenetic modifications are depicted on the X-axis. Data are expressed as mean ± Stdev; *p<0.05, **p<0.01. Means with different letters are significantly different. N indicates the number of independent experiments, and per experiment each sample and the corresponding negative controls were run in triplicates.
Effect of tranylcypromine and trichostatin A on DNA methylation of CpG islands in Pou5f1 and Gfra1 promoters
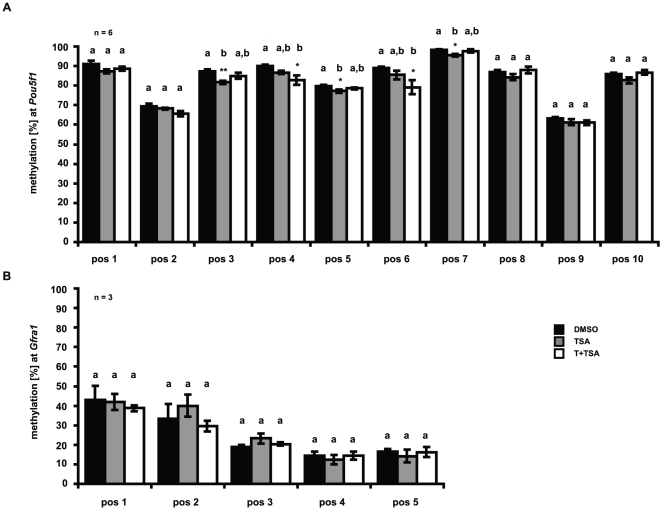
It is well known that histone methylation and DNA CpG methylation are functionally linked [49], [74], [75], [76], [77], [78]. We performed bisulfite treatment followed by pyrosequencing to determine the effect of TSA and T on the DNA methylation status of Pou5f1 and Gfra1 promoters. Primers used in these assays were located in the proximal promoter of Pou5f1 (−190 to +26 relative to TSS) and Gfra1 (+26 to +143 relative to TSS), overlapping or in closed vicinity with regions analyzed by ChIP-qPCR, which allowed us to relate the DNA methylation status of the promoter to the histone methylation and acetylation status and thus changes in gene expression between treated GC-1 cells and DMSO controls. Ten CpG dinucleotide positions were assayed for methylation in the defined region of the Pou5f1 promoter and identified as hypermethylated (>70% methylation) in all cells, comprising DMSO controls, or TSA-, or TSA and T treated GC-1 cells (Figure 6A). Interestingly, depending on the position of the dinucleotide and the treatment, slight but significant reductions in CpG methylation could be observed in the assayed Pou5f1 promoter region (Figure 6A). On the contrary, CpG dinucleotides of the assayed Gfra1 promoter region are slightly methylated as indicated in Figure 6B. Here, we were not able to detect any significant changes occurred between treated cells and controls.
Figure 6. Effect of tranylcypromine and trichostatin A treatment on CpG methylation at Pou5f1 and Gfra1 promoters.
DNA was extracted, treated with bisulfite, amplified by PCR and pyrosequencing was used to determine the bisulfite-converted sequence of CpG sites in (A) Pou5f1 (positions −190 to +26 relative to the transcriptional start site) and (B) Gfra1 promoters (positions +26 to +143 relative to the transcriptional start site). The methylation status of each position is depicted as the mean percentage of “n” independent experiments ± SEM; *p<0.05, **p<0.01. Means with different letters are significantly different. Black bars indicate DMSO controls, grey bars depict TSA treated samples and white bars represent samples that have been exposed to both inhibitors, tranylcypromine and TSA (T+TSA).
KDM1 and HDAC1 occupy Pou5f1 and Gfra1 promoters in GC-1 cells
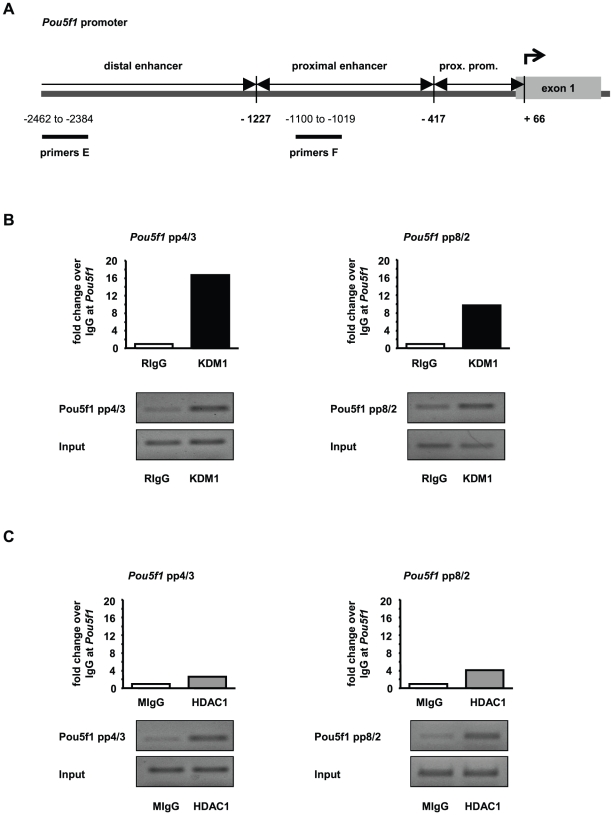
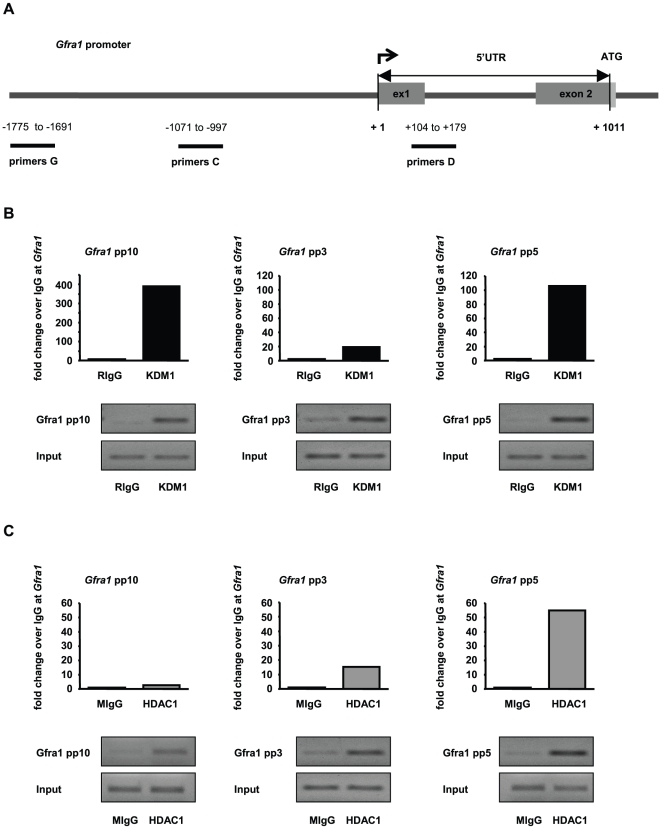
ChIP-QPCR was used to confirm that the protein targets of T and TSA, namely KDM1 and/or HDAC1 interact with Pou5f1 and Gfra1 gene promoters. These assays demonstrated a 17-fold enrichment of KDM1 in the distal enhancer and a 10-fold enrichment of KDM1 in the proximal enhancer of the Pou5f1 promoter in GC-1 cells (Figure 7B). HDAC1 was found to be enriched at the Pou5f1 promoter, namely 3-fold in the distal and 4-fold in the proximal enhancer region (Figure 7C). Analyzing KDM1 and HDAC1 occupancy of the Gfra1 promoter in GC-1 cells, we were able to detect a 390-fold enrichment of KDM1 in the distal (−1775 relative to TSS) and a 20-fold enrichment in the more proximal promoter region (−1071 relative to TSS). Moreover KDM1 was found to be 106-times enriched in the 5′UTR of the Gfra1 promoter (Figure 8B). HDAC1 was 3-times enriched in the distal Gfra1 promoter region, an over 15-fold enrichment of HDAC1 was detectable in the proximal promoter region and a 55-fold enrichment in the 5′UTR of the Gfra1 promoter (Figure 8C). According to a BLAST search at NCBI (http://blast.ncbi.nlm.nih.gov/) the antibody used to detect HDAC1 may have some cross-reactivity with two isoforms of HDAC1 (XP_001480415.1, XP_001480417.1) which would impact the results by diluting the signal at the target regions. These results strongly suggest that KDM1 and HDAC1 occupy Pou5f1 and Gfra1 promoters in GC-1 cells and function in the regulation of expression.
Figure 7. KDM1- and HDAC1-containing protein complexes are located at the Pou5f1 promoter in GC-1 cells.
(A) Schematic representation of the Pou5f1 promoter. Numbers depict the positions of primer pairs used for ChIP-qPCR relative to the corresponding transcriptional start site. (B,C) ChIP-qPCRs demonstrate occupancy of the Pou5f1 promoter by KDM1- (B) and HDAC1-containing protein complexes (C). ChIP assays were performed on GC-1 cells that have been cultured in the presence of dimethylsulphoxide using either a specific antibody (recognizing either KDM1 (black bars) or HDAC1 (grey bars)) or the corresponding IgG controls (white bars), (rabbit IgG = RIgG; mouse IgG = MIgG). Each bar represents the ChIP-qPCR results of two independent, pooled experiments. Signals of target DNA received from IgG controls served as base and were defined as one. Fold change represents the 2∧-(ΔΔCT) value, where the normalized IgG signal is subtracted from the normalized signal of the specific antibody. QPCR results were analyzed on an agarose gel and shown underneath the corresponding graphs.
Figure 8. KDM1- and HDAC1-containing protein complexes are located at the Gfra1 promoter in GC-1 cells.
(A) Schematic representation of the Gfra1 promoter. Numbers depict the positions of primer pairs used for ChIP-qPCR relative to the corresponding transcriptional start site. (B,C) ChIP-qPCRs demonstrate occupancy of the Gfra1 promoter by KDM1- (B) and HDAC1-containing protein complexes (C). ChIP assays were performed on GC-1 cells that have been cultured in the presence of dimethylsulphoxide using either a specific antibody (recognizing either KDM1 (black bars) or HDAC1 (grey bars)) or the corresponding IgG controls (white bars), (rabbit IgG = RIgG; mouse IgG = MIgG). Each bar represents the ChIP-qPCR results of two independent, pooled experiments. Signals of target DNA received from IgG controls served as base and were defined as one. Fold change represents the 2∧-(ΔΔCT) value, where the normalized IgG signal is subtracted from the normalized signal of the specific antibody. QPCR results were analyzed on an agarose gel and shown underneath the corresponding graph.
Discussion
In this study we revealed epigenetic mechanisms that are likely to be involved in the expression of key genes in spermatogonia, namely Pou5f1 and Gfra1. These experiments implicate histone H3 methylation and acetylation in the regulation of their expression and suggest that transcription is influenced by HDACs and KDM1. The underlying molecular mechanisms revealed by pharmacological treatments of the GC-1 cell line suggest that in spermatogonia expression of Pou5f1 and Gfra1 is maintained by an enrichment of histone H3 acetylation and dimethylation of histone H3 at lysine 4. Based on our results we suggest that as spermatogonia differentiate the loss of expression of Pou5f1 and Gfra1 may be concomitant with gene-specific activity of HDACs and KDM1 and the consequent removal of gene activating histone H3 acetylation and H3K4 dimethylation. These epigenetic changes are summarized in a model shown in Figure 9.
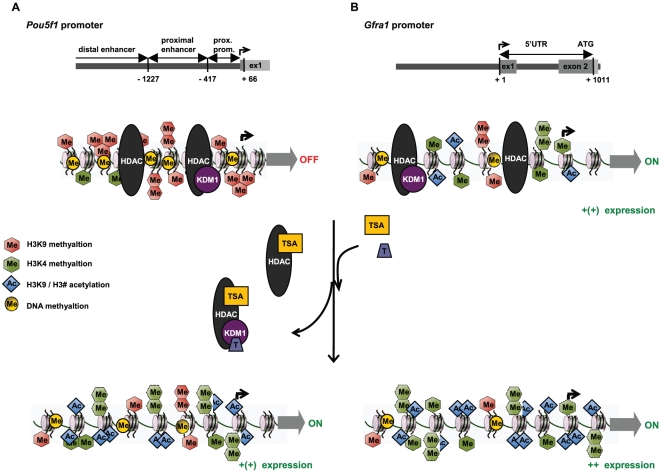
Figure 9. Epigenetic remodeling in Pouf5f1 and Gfra1 promoter regions - a proposed model to explain changes in gene expression in GC-1 cells after Tranylcypromine and TSA treatment.
(A) In untreated GC-1 cells (regular growth conditions), repressive epigenetic marks (e.g. H3K9 methylation and CpG methylation) keep the Pou5f1 promoter silent, by creating a tight chromatin structure and preventing the binding of the transcription machinery. Multi-protein complexes that contain writers of repressive epigenetic marks, e.g. HDACs and/or HDAC/KDM1, create a repressive chromatin state. Blocking these enzymes using TSA (targets HDACs) and/or tranylcypromine (represses KDM1) results in the accumulation of activating epigenetic marks (H3K4 methylation, H3 acetylation), thereby making it accessible for transcription factors and the transcription machinery, which eventually results in the induction of Pou5f1 expression. (B) Repressive and activating epigenetic marks at the promoter are responsible for the weak expression of Gfra1 in untreated GC-1 cells. The exposure of GC-1 cells to tranylcypromine and TSA blocks the gene silencing activities of HDACs and/or HDAC/KDM1 and allows for the accumulation of activating epigenetic marks. Thus the treatment shifts the epigenetic landscape at the Gfra1 promoter to a more active state and eventually triggers a stronger transcription of Gfra1. Histone deacetylase (HDAC), lysine-specific demethylase 1 (KDM1), tranylcypromine (T), trichostatin A (TSA).
Histone acetylation is intimately involved in the epigenetic regulation of transcription and at a single gene locus a fine balance of histone acetyltransferase and deacetylase activity controls gene expression [79], [80], [81]. Histone hyperacetylation is associated with an open chromatin state and general transcriptional activity [10], [11], [12], [13], [14]. Inhibition of histone deacetylase activity by TSA has many faces: it can for instance result in de-differentiation of primordial germ cells into embryonic germ cells [82], accelerate murine stem cell differentiation [83], force cancer cells into a differentiated state [84], or in concert with a DNA methyltransferase inhibitor convert neurosphere cells into neurosphere-derived haematopoietic cells [85]. While we did not observe changes in cell morphology or behavior to indicate the GC-1 cell line altered its differentiation state, we did observe specific changes in gene expression that were associated to gene-specific changes in the epigenetic profile. In the brief time-scale of these experiments changes in differentiation would not be expected. For example in the case of induced pluripotent stem cells, virally transduced pluripotency factors (e.g. Pou5f1, Klf4, Sox2) need to be expressed for approximately twelve to eighteen days to initiate de-differentiation and reprogramming of mouse embryonic fibroblast cells [86].
Interestingly TSA treatment of the GC-1 cell line turned on specific genes expressed in spermatogonia as we saw an activation of spermatogonia cell markers Gfra1 and Pou5f1. The lack of change in expression in the panel of genes examined suggests that expression of Pou5f1 and Gfra1 is a consequence of gene-specific epigenetic effects of the HDAC and KDM1 inhibitors and not due to global changes to the epigenome. In the present study, induced Pou5f1 expression was associated with increased H3K4 dimethylation, H3K9 acetylation and pan-H3 acetylation at regulatory regions of the mouse Pou5f1 promoter. Although we demonstrated that Pou5f1 expression was maximal under the co-suppression of KDM1 and HDAC1, a closer look at the epigenetic marks on the promoter revealed that co-inhibition of HDAC and KDM1 did not further enhance gene activating H3K4 methylation marks. Therefore the enhanced gene activation under KDM1/HDAC1 inhibition may be a consequence of indirect KDM1/HDAC1 signaling. We postulate that the inhibition by TSA on HDACs and the consequent increased H3K4 dimethylation was due to an indirect effect on KDM1 since TSA does not target demethylases, but specifically inhibits histone deacetylases by binding to the catalytic core [87]. Evidence supporting this potential molecular mechanism comes from experiments showing that hyperacetylation of lysine residues on histone H3 abolished or attenuated KDM1 function, and that acetyl groups had to be removed before H3K4 demethylation by KDM1 occurred [88], [89]. Moreover we detected KDM1 and HDAC at our target genes promoters suggesting that effects are a consequence of drug action on the KDM1/HDAC complex.
The gain in histone H3K4 dimethylation at the Pou5f1 promoter and associated gene activation we report here following HDAC inhibition and under the combined suppression of KDM1/HDAC is similar to what has been observed in embryonal carcinoma cells [43]. However, unlike the GC-1 cells, embryonal carcinoma cells increased Pou5f1 expression when KDM1 was inhibited by T alone [43]. The differences in cellular response to T between the embryonal carcinoma cells and GC-1 cell used here may be due to differences in sensitivity to inhibition of KDM1 by T. Unlike GC-1 cells, embryonal carcinoma cells express high levels of KDM1 and have detectable levels of Pou5f1 under control conditions [43].
Like Pou5f1, the expression of Gfra1 in GC-1 cells seemed to be directed principally by HDACs since treatment with TSA caused a strong increase in activating H3K9- and total H3 acetylation in the Gfra1 promoter region and consequent gene expression. In contrast to the major changes in acetylation only a slight increase in gene activating H3K4 dimethylation under the combined inhibition of KDM1/HDAC1 by TSA and T occurred, and this was not associated with enhanced expression in comparison to TSA alone. In this study there was no effect of inhibition of KDM1 by T alone on global histone modifications or changes in gene expression. Rather, treatment with T was effective only when used in combination with TSA. These effects are in line with previous studies where it has been reported that there was a lack of globally increased histone H3 methylation after genetic ablation of KDM1 in murine ES cells [49], and following the suppression of KDM1 by T, or its knockdown by siRNA in human cervical cancer cell lines [48]. Taken together these findings suggest that KDM1 activities are highly gene-specific and likely to involve its function in a protein complex with HDAC and other proteins.
At some gene promoters there is crosstalk between histone modifications and DNA methylation. In a previous study Ou and coworkers associated a TSA induced increase of histone acetylation, with a significant decrease in global DNA methylation [90]. It was thereby pertinent to assess the DNA methylation status at the promoter regions of Pou5f1 and Gfra1. Overall no significant changes in DNA methylation occurred in either the Pou5f1 or Gfra1 promoter indicating that histone methylation and acetylation are the principles means of regulating gene expression at these promoters in GC-1 cells. However, depending on the treatment and the position of the CpG dinucleotides we observed a slight but significant decrease in DNA methylation in the analyzed Pou5f1 promoter region, but none in the Gfra1 promoter. We cannot completely rule out that the changes in DNA methylation observed in the Pou5f1 promoter had an impact on the induction of its expression in treated GC-1 cells. But this seems to be unlikely as >60 to 70% methylation remained following treatment at the Pou5f1 promoter and low methylation was found on the Gfra1 promoter.
As the mechanisms and interpretation discussed here are based on a transformed spermatocyte-like cell line, this underlies the extrapolation to events in vivo. The use of an immortalized cell line such as the GC-1 holds certain limitations but the positive factors outweighed the negatives when choosing a model system and examining the alternatives. The GC-1 cells represent a relatively homogenous cell line that can be readily and robustly expanded without being co-cultured with somatic cells, while sustaining germ cell characteristics [42]. Moreover they are easy to manipulate and deliver an unlimited source of material, which is critical for ChIP experiments, which require millions of cells per treatment for analysis.
Importantly this study has revealed epigenetic mechanisms likely to be involved in the complex gene regulation in spermatogenesis. Worth considering is that environmental induced epigenetic mutations in developing germ cells may have consequences for lifetime fertility as there is likely to be plasticity in histone modifications.
Supporting Information
Antibodies used for Western blotting (WB) and chromatin-immunoprecipitation (ChIP).
(0.04 MB DOC)
Primers and TaqMan gene expression assays used for qPCR or ChIP-qPCR.
(0.07 MB DOC)
qPCR analysis of male germ cell marker gene expression in untreated GC-1 cells.
(0.04 MB DOC)
qPCR analysis of MyoD gene expression in untreated and treated GC-1 cells.
(0.03 MB DOC)
Effect of tranylcypromine and trichostatin A on global Histone H3K4 methylation and H3 acetylation levels. Western blot analyses of global histone (A) H3K9 acetylation, (B) pan-H3 acetylation, (C) H3K4 monomethylation, and (D) H3K4 dimethylation on protein extracts isolated from GC-1 cells either grown in culture medium (control, ctr), or treated with tranylcypromine (T), trichostatin A (TSA), or dimethylsulphoxid (DMSO; TSA solvent), or tranylcypromine and TSA (T+TSA). (E) Histone H3 visualized on a coomassie gel was used for normalization. Tranylcypromine alone does not affect global H3 acetylation (A,B) or H3K4 methylation (C,D) in GC-1 cells. TSA exposure causes a global increase in H3K9 (A) and total H3 acetylation (B) and, probably due to an indirect effect [91], H3K4 dimethylation (D). The combined treatment of T+TSA induces a global increase in H3K4 monomethylation (C), but does not further enhance the effect on H3 acetylation (A,B) or H3K4 dimethylation (C). Signals from proteins isolated from control cells served as the base and were defined as one. Fold change indicates the ratio of signal intensity in treated cells normalized to histone H3 over signal intensity in control cells normalized histone H3. Data are expressed as mean ± SEM; **p<0.01, ***p<0.001. Means with different letters are significantly different. N indicates the number of independent experiments.
(2.21 MB EPS)
Tranylcypromine and trichostatin A do not affect global histone H3K9 methylation or H3K4 trimethylation levels. Western blot analyses of global histone (A) H3K9 mono-, (B) H3K9 di-, (D) H3K9 tri-, and (E) H3K4 trimethylation on protein extracts isolated from GC-1 cells either grown in culture medium (control, ctr), or treated with tranylcypromine (T), trichostatin A (TSA), or dimethylsulphoxid (DMSO; TSA solvent), or tranylcypromine and TSA (T+TSA). (C) and (F) Histone H3 visualized on coomassie gels was used for normalization. Signals achieved with proteins isolated from control cells served as base and were defined as one. Fold change indicates the ratio of signal intensity in treated cells normalized to histone H3 over signal intensity in control cells normalized histone H3. Data are expressed as mean ± SEM. N indicates the number of independent experiments.
(3.02 MB EPS)
Acknowledgments
We are grateful to Dr. Romain Lambrot for helpful comments and discussions, Dr. Jose Moro for his statistical support, and all other Kimmins laboratory group members.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: This research was funded by the Canadian Institutes of Health Research (SK grant # 82820). MG is supported by a post-doctoral fellowship from the Canadian Institutes of Health Research. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Kimmins S, Sassone-Corsi P. Chromatin remodelling and epigenetic features of germ cells. Nature. 2005;434:583–589. doi: 10.1038/nature03368. [DOI] [PubMed] [Google Scholar]
- 2.Chen C, Ouyang W, Grigura V, Zhou Q, Carnes K, et al. ERM is required for transcriptional control of the spermatogonial stem cell niche. Nature. 2005;436:1030–1034. doi: 10.1038/nature03894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Costoya JA, Hobbs RM, Barna M, Cattoretti G, Manova K, et al. Essential role of Plzf in maintenance of spermatogonial stem cells. Nat Genet. 2004;36:653–659. doi: 10.1038/ng1367. [DOI] [PubMed] [Google Scholar]
- 4.He Z, Jiang J, Hofmann MC, Dym M. Gfra1 silencing in mouse spermatogonial stem cells results in their differentiation via the inactivation of RET tyrosine kinase. Biol Reprod. 2007;77:723–733. doi: 10.1095/biolreprod.107.062513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Oatley JM, Avarbock MR, Telaranta AI, Fearon DT, Brinster RL. Identifying genes important for spermatogonial stem cell self-renewal and survival. Proc Natl Acad Sci U S A. 2006;103:9524–9529. doi: 10.1073/pnas.0603332103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Pesce M, Wang X, Wolgemuth DJ, Scholer H. Differential expression of the Oct-4 transcription factor during mouse germ cell differentiation. Mech Dev. 1998;71:89–98. doi: 10.1016/s0925-4773(98)00002-1. [DOI] [PubMed] [Google Scholar]
- 7.Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–1080. doi: 10.1126/science.1063127. [DOI] [PubMed] [Google Scholar]
- 8.Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell. 2005;120:169–181. doi: 10.1016/j.cell.2005.01.001. [DOI] [PubMed] [Google Scholar]
- 9.Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. doi: 10.1016/j.cell.2007.02.005. [DOI] [PubMed] [Google Scholar]
- 10.Bulger M. Hyperacetylated chromatin domains: lessons from heterochromatin. J Biol Chem. 2005;280:21689–21692. doi: 10.1074/jbc.R500004200. [DOI] [PubMed] [Google Scholar]
- 11.Grimes SR, Jr, Henderson N. Hyperacetylation of histone H4 in rat testis spermatids. Exp Cell Res. 1984;152:91–97. doi: 10.1016/0014-4827(84)90232-5. [DOI] [PubMed] [Google Scholar]
- 12.Hazzouri M, Pivot-Pajot C, Faure AK, Usson Y, Pelletier R, et al. Regulated hyperacetylation of core histones during mouse spermatogenesis: involvement of histone deacetylases. Eur J Cell Biol. 2000;79:950–960. doi: 10.1078/0171-9335-00123. [DOI] [PubMed] [Google Scholar]
- 13.Ma J, Svoboda P, Schultz RM, Stein P. Regulation of zygotic gene activation in the preimplantation mouse embryo: global activation and repression of gene expression. Biol Reprod. 2001;64:1713–1721. doi: 10.1095/biolreprod64.6.1713. [DOI] [PubMed] [Google Scholar]
- 14.Millar CB, Grunstein M. Genome-wide patterns of histone modifications in yeast. Nat Rev Mol Cell Biol. 2006;7:657–666. doi: 10.1038/nrm1986. [DOI] [PubMed] [Google Scholar]
- 15.Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–326. doi: 10.1016/j.cell.2006.02.041. [DOI] [PubMed] [Google Scholar]
- 16.Lagarkova MA, Volchkov PY, Lyakisheva AV, Philonenko ES, Kiselev SL. Diverse epigenetic profile of novel human embryonic stem cell lines. Cell Cycle. 2006;5:416–420. doi: 10.4161/cc.5.4.2440. [DOI] [PubMed] [Google Scholar]
- 17.Yeo S, Jeong S, Kim J, Han JS, Han YM, et al. Characterization of DNA methylation change in stem cell marker genes during differentiation of human embryonic stem cells. Biochem Biophys Res Commun. 2007;359:536–542. doi: 10.1016/j.bbrc.2007.05.120. [DOI] [PubMed] [Google Scholar]
- 18.Clermont Y. Kinetics of spermatogenesis in mammals: seminiferous epithelium cycle and spermatogonial renewal. Physiol Rev. 1972;52:198–236. doi: 10.1152/physrev.1972.52.1.198. [DOI] [PubMed] [Google Scholar]
- 19.Russell LD, Ettlin RA, Sinha Hikim AP, Clegg ED. Clearwater, Florida: Cache River Press; 1990. Histological and histopathological evaluation of the testis. [Google Scholar]
- 20.de Rooij DG. Proliferation and differentiation of spermatogonial stem cells. Reproduction. 2001;121:347–354. doi: 10.1530/rep.0.1210347. [DOI] [PubMed] [Google Scholar]
- 21.de Rooij DG, Russell LD. All you wanted to know about spermatogonia but were afraid to ask. J Androl. 2000;21:776–798. [PubMed] [Google Scholar]
- 22.Naughton CK, Jain S, Strickland AM, Gupta A, Milbrandt J. Glial cell-line derived neurotrophic factor-mediated RET signaling regulates spermatogonial stem cell fate. Biol Reprod. 2006;74:314–321. doi: 10.1095/biolreprod.105.047365. [DOI] [PubMed] [Google Scholar]
- 23.Nichols J, Zevnik B, Anastassiadis K, Niwa H, Klewe-Nebenius D, et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell. 1998;95:379–391. doi: 10.1016/s0092-8674(00)81769-9. [DOI] [PubMed] [Google Scholar]
- 24.Niwa H, Miyazaki J, Smith AG. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet. 2000;24:372–376. doi: 10.1038/74199. [DOI] [PubMed] [Google Scholar]
- 25.Ohbo K, Yoshida S, Ohmura M, Ohneda O, Ogawa T, et al. Identification and characterization of stem cells in prepubertal spermatogenesis in mice small star, filled. Dev Biol. 2003;258:209–225. doi: 10.1016/s0012-1606(03)00111-8. [DOI] [PubMed] [Google Scholar]
- 26.Ohmura M, Yoshida S, Ide Y, Nagamatsu G, Suda T, et al. Spatial analysis of germ stem cell development in Oct-4/EGFP transgenic mice. Arch Histol Cytol. 2004;67:285–296. doi: 10.1679/aohc.67.285. [DOI] [PubMed] [Google Scholar]
- 27.Kubota H, Avarbock MR, Brinster RL. Growth factors essential for self-renewal and expansion of mouse spermatogonial stem cells. Proc Natl Acad Sci U S A. 2004;101:16489–16494. doi: 10.1073/pnas.0407063101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fenic I, Sonnack V, Failing K, Bergmann M, Steger K. In vivo effects of histone-deacetylase inhibitor trichostatin-A on murine spermatogenesis. J Androl. 2004;25:811–818. doi: 10.1002/j.1939-4640.2004.tb02859.x. [DOI] [PubMed] [Google Scholar]
- 29.Godmann M, Lambrot R, Kimmins S. The dynamic epigenetic program in male germ cells: Its role in spermatogenesis, testis cancer, and its response to the environment. Microsc Res Tech. 2009 doi: 10.1002/jemt.20715. [DOI] [PubMed] [Google Scholar]
- 30.Hayashi K, Yoshida K, Matsui Y. A histone H3 methyltransferase controls epigenetic events required for meiotic prophase. Nature. 2005;438:374–378. doi: 10.1038/nature04112. [DOI] [PubMed] [Google Scholar]
- 31.McBurney MW, Yang X, Jardine K, Hixon M, Boekelheide K, et al. The mammalian SIR2alpha protein has a role in embryogenesis and gametogenesis. Mol Cell Biol. 2003;23:38–54. doi: 10.1128/MCB.23.1.38-54.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Okada Y, Scott G, Ray MK, Mishina Y, Zhang Y. Histone demethylase JHDM2A is critical for Tnp1 and Prm1 transcription and spermatogenesis. Nature. 2007;450:119–123. doi: 10.1038/nature06236. [DOI] [PubMed] [Google Scholar]
- 33.Peters AH, O'Carroll D, Scherthan H, Mechtler K, Sauer S, et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell. 2001;107:323–337. doi: 10.1016/s0092-8674(01)00542-6. [DOI] [PubMed] [Google Scholar]
- 34.Tachibana M, Nozaki M, Takeda N, Shinkai Y. Functional dynamics of H3K9 methylation during meiotic prophase progression. Embo J. 2007;26:3346–3359. doi: 10.1038/sj.emboj.7601767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Godmann M, Auger V, Ferraroni-Aguiar V, Di Sauro A, Sette C, et al. Dynamic regulation of histone H3 methylation at lysine 4 in mammalian spermatogenesis. Biol Reprod. 2007;77:754–764. doi: 10.1095/biolreprod.107.062265. [DOI] [PubMed] [Google Scholar]
- 36.Payne C, Braun RE. Histone lysine trimethylation exhibits a distinct perinuclear distribution in Plzf-expressing spermatogonia. Dev Biol. 2006;293:461–472. doi: 10.1016/j.ydbio.2006.02.013. [DOI] [PubMed] [Google Scholar]
- 37.Hakimi MA, Dong Y, Lane WS, Speicher DW, Shiekhattar R. A candidate X-linked mental retardation gene is a component of a new family of histone deacetylase-containing complexes. J Biol Chem. 2003;278:7234–7239. doi: 10.1074/jbc.M208992200. [DOI] [PubMed] [Google Scholar]
- 38.Lee MG, Wynder C, Norman J, Shiekhattar R. Isolation and characterization of histone H3 lysine 4 demethylase-containing complexes. Methods. 2006;40:327–330. doi: 10.1016/j.ymeth.2006.07.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Shi YJ, Matson C, Lan F, Iwase S, Baba T, et al. Regulation of LSD1 histone demethylase activity by its associated factors. Mol Cell. 2005;19:857–864. doi: 10.1016/j.molcel.2005.08.027. [DOI] [PubMed] [Google Scholar]
- 40.Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–953. doi: 10.1016/j.cell.2004.12.012. [DOI] [PubMed] [Google Scholar]
- 41.Metzger E, Wissmann M, Yin N, Muller JM, Schneider R, et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature. 2005;437:436–439. doi: 10.1038/nature04020. [DOI] [PubMed] [Google Scholar]
- 42.Hofmann MC, Narisawa S, Hess RA, Millan JL. Immortalization of germ cells and somatic testicular cells using the SV40 large T antigen. Exp Cell Res. 1992;201:417–435. doi: 10.1016/0014-4827(92)90291-f. [DOI] [PubMed] [Google Scholar]
- 43.Lee MG, Wynder C, Schmidt DM, McCafferty DG, Shiekhattar R. Histone H3 lysine 4 demethylation is a target of nonselective antidepressive medications. Chem Biol. 2006;13:563–567. doi: 10.1016/j.chembiol.2006.05.004. [DOI] [PubMed] [Google Scholar]
- 44.Yoshida M, Kijima M, Akita M, Beppu T. Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J Biol Chem. 1990;265:17174–17179. [PubMed] [Google Scholar]
- 45.Medina V, Edmonds B, Young GP, James R, Appleton S, et al. Induction of caspase-3 protease activity and apoptosis by butyrate and trichostatin A (inhibitors of histone deacetylase): dependence on protein synthesis and synergy with a mitochondrial/cytochrome c-dependent pathway. Cancer Res. 1997;57:3697–3707. [PubMed] [Google Scholar]
- 46.Minucci S, Horn V, Bhattacharyya N, Russanova V, Ogryzko VV, et al. A histone deacetylase inhibitor potentiates retinoid receptor action in embryonal carcinoma cells. Proc Natl Acad Sci U S A. 1997;94:11295–11300. doi: 10.1073/pnas.94.21.11295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Taylor SJ, Resnick RJ, Shalloway D. Sam68 exerts separable effects on cell cycle progression and apoptosis. BMC Cell Biol. 2004;5:5. doi: 10.1186/1471-2121-5-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhu Q, Liu C, Ge Z, Fang X, Zhang X, et al. Lysine-specific demethylase 1 (LSD1) Is required for the transcriptional repression of the telomerase reverse transcriptase (hTERT) gene. PLoS ONE. 2008;3:e1446. doi: 10.1371/journal.pone.0001446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wang J, Hevi S, Kurash JK, Lei H, Gay F, et al. The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat Genet. 2009;41:125–129. doi: 10.1038/ng.268. [DOI] [PubMed] [Google Scholar]
- 50.Jiang C, Xuan Z, Zhao F, Zhang MQ. TRED: a transcriptional regulatory element database, new entries and other development. Nucleic Acids Res. 2007;35:D137–140. doi: 10.1093/nar/gkl1041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zhao F, Xuan Z, Liu L, Zhang MQ. TRED: a Transcriptional Regulatory Element Database and a platform for in silico gene regulation studies. Nucleic Acids Res. 2005;33:D103–107. doi: 10.1093/nar/gki004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Stothard P. The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques. 2000;28:1102–1104. doi: 10.2144/00286ir01. [DOI] [PubMed] [Google Scholar]
- 53.Gardiner-Garden M, Frommer M. CpG islands in vertebrate genomes. J Mol Biol. 1987;196:261–282. doi: 10.1016/0022-2836(87)90689-9. [DOI] [PubMed] [Google Scholar]
- 54.Buaas FW, Kirsh AL, Sharma M, McLean DJ, Morris JL, et al. Plzf is required in adult male germ cells for stem cell self-renewal. Nat Genet. 2004;36:647–652. doi: 10.1038/ng1366. [DOI] [PubMed] [Google Scholar]
- 55.Yoshimizu T, Sugiyama N, De Felice M, Yeom YI, Ohbo K, et al. Germline-specific expression of the Oct-4/green fluorescent protein (GFP) transgene in mice. Dev Growth Differ. 1999;41:675–684. doi: 10.1046/j.1440-169x.1999.00474.x. [DOI] [PubMed] [Google Scholar]
- 56.Manova K, Nocka K, Besmer P, Bachvarova RF. Gonadal expression of c-kit encoded at the W locus of the mouse. Development. 1990;110:1057–1069. doi: 10.1242/dev.110.4.1057. [DOI] [PubMed] [Google Scholar]
- 57.Ohta H, Yomogida K, Dohmae K, Nishimune Y. Regulation of proliferation and differentiation in spermatogonial stem cells: the role of c-kit and its ligand SCF. Development. 2000;127:2125–2131. doi: 10.1242/dev.127.10.2125. [DOI] [PubMed] [Google Scholar]
- 58.Sorrentino V, Giorgi M, Geremia R, Besmer P, Rossi P. Expression of the c-kit proto-oncogene in the murine male germ cells. Oncogene. 1991;6:149–151. [PubMed] [Google Scholar]
- 59.Vincent S, Segretain D, Nishikawa S, Nishikawa SI, Sage J, et al. Stage-specific expression of the Kit receptor and its ligand (KL) during male gametogenesis in the mouse: a Kit-KL interaction critical for meiosis. Development. 1998;125:4585–4593. doi: 10.1242/dev.125.22.4585. [DOI] [PubMed] [Google Scholar]
- 60.Yoshinaga K, Nishikawa S, Ogawa M, Hayashi S, Kunisada T, et al. Role of c-kit in mouse spermatogenesis: identification of spermatogonia as a specific site of c-kit expression and function. Development. 1991;113:689–699. doi: 10.1242/dev.113.2.689. [DOI] [PubMed] [Google Scholar]
- 61.Buageaw A, Sukhwani M, Ben-Yehudah A, Ehmcke J, Rawe VY, et al. GDNF family receptor alpha1 phenotype of spermatogonial stem cells in immature mouse testes. Biol Reprod. 2005;73:1011–1016. doi: 10.1095/biolreprod.105.043810. [DOI] [PubMed] [Google Scholar]
- 62.Dettin L, Ravindranath N, Hofmann MC, Dym M. Morphological characterization of the spermatogonial subtypes in the neonatal mouse testis. Biol Reprod. 2003;69:1565–1571. doi: 10.1095/biolreprod.103.016394. [DOI] [PubMed] [Google Scholar]
- 63.Meng X, Lindahl M, Hyvonen ME, Parvinen M, de Rooij DG, et al. Regulation of cell fate decision of undifferentiated spermatogonia by GDNF. Science. 2000;287:1489–1493. doi: 10.1126/science.287.5457.1489. [DOI] [PubMed] [Google Scholar]
- 64.Tadokoro Y, Yomogida K, Ohta H, Tohda A, Nishimune Y. Homeostatic regulation of germinal stem cell proliferation by the GDNF/FSH pathway. Mech Dev. 2002;113:29–39. doi: 10.1016/s0925-4773(02)00004-7. [DOI] [PubMed] [Google Scholar]
- 65.Foulkes NS, Mellstrom B, Benusiglio E, Sassone-Corsi P. Developmental switch of CREM function during spermatogenesis: from antagonist to activator. Nature. 1992;355:80–84. doi: 10.1038/355080a0. [DOI] [PubMed] [Google Scholar]
- 66.Nantel F, Monaco L, Foulkes NS, Masquilier D, LeMeur M, et al. Spermiogenesis deficiency and germ-cell apoptosis in CREM-mutant mice. Nature. 1996;380:159–162. doi: 10.1038/380159a0. [DOI] [PubMed] [Google Scholar]
- 67.Hintz M, Goldberg E. Immunohistochemical localization of LDH-x during spermatogenesis in mouse testes. Dev Biol. 1977;57:375–384. doi: 10.1016/0012-1606(77)90222-6. [DOI] [PubMed] [Google Scholar]
- 68.Behr R, Kaestner KH. Developmental and cell type-specific expression of the zinc finger transcription factor Kruppel-like factor 4 (Klf4) in postnatal mouse testis. Mech Dev. 2002;115:167–169. doi: 10.1016/s0925-4773(02)00127-2. [DOI] [PubMed] [Google Scholar]
- 69.Godmann M, Gashaw I, Katz JP, Nagy A, Kaestner KH, et al. Kruppel-like factor 4, a “pluripotency transcription factor” highly expressed in male postmeiotic germ cells, is dispensable for spermatogenesis in the mouse. Mech Dev. 2009;126:650–664. doi: 10.1016/j.mod.2009.06.1081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Kanatsu-Shinohara M, Lee J, Inoue K, Ogonuki N, Miki H, et al. Pluripotency of a single spermatogonial stem cell in mice. Biol Reprod. 2008;78:681–687. doi: 10.1095/biolreprod.107.066068. [DOI] [PubMed] [Google Scholar]
- 71.Hubner K, Fuhrmann G, Christenson LK, Kehler J, Reinbold R, et al. Derivation of oocytes from mouse embryonic stem cells. Science. 2003;300:1251–1256. doi: 10.1126/science.1083452. [DOI] [PubMed] [Google Scholar]
- 72.Yeom YI, Fuhrmann G, Ovitt CE, Brehm A, Ohbo K, et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development. 1996;122:881–894. doi: 10.1242/dev.122.3.881. [DOI] [PubMed] [Google Scholar]
- 73.Arnold HH, Braun T. Targeted inactivation of myogenic factor genes reveals their role during mouse myogenesis: a review. Int J Dev Biol. 1996;40:345–353. [PubMed] [Google Scholar]
- 74.Esteve PO, Chin HG, Smallwood A, Feehery GR, Gangisetty O, et al. Direct interaction between DNMT1 and G9a coordinates DNA and histone methylation during replication. Genes Dev. 2006;20:3089–3103. doi: 10.1101/gad.1463706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Jackson JP, Lindroth AM, Cao X, Jacobsen SE. Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature. 2002;416:556–560. doi: 10.1038/nature731. [DOI] [PubMed] [Google Scholar]
- 76.Lehnertz B, Ueda Y, Derijck AA, Braunschweig U, Perez-Burgos L, et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol. 2003;13:1192–1200. doi: 10.1016/s0960-9822(03)00432-9. [DOI] [PubMed] [Google Scholar]
- 77.Tamaru H, Selker EU. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature. 2001;414:277–283. doi: 10.1038/35104508. [DOI] [PubMed] [Google Scholar]
- 78.Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature. 2006;439:871–874. doi: 10.1038/nature04431. [DOI] [PubMed] [Google Scholar]
- 79.Liu R, Wang L, Chen G, Katoh H, Chen C, et al. FOXP3 up-regulates p21 expression by site-specific inhibition of histone deacetylase 2/histone deacetylase 4 association to the locus. Cancer Res. 2009;69:2252–2259. doi: 10.1158/0008-5472.CAN-08-3717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Vogelauer M, Wu J, Suka N, Grunstein M. Global histone acetylation and deacetylation in yeast. Nature. 2000;408:495–498. doi: 10.1038/35044127. [DOI] [PubMed] [Google Scholar]
- 81.Zhong X, Jin Y. Critical Roles of Coactivator p300 in Mouse Embryonic Stem Cell Differentiation and Nanog Expression. J Biol Chem. 2009;284:9168–9175. doi: 10.1074/jbc.M805562200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Durcova-Hills G, Tang F, Doody G, Tooze R, Surani MA. Reprogramming primordial germ cells into pluripotent stem cells. PLoS ONE. 2008;3:e3531. doi: 10.1371/journal.pone.0003531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Karantzal E, Schulz H, Hummel O, Hubner N, Hatzopoulos A, et al. Histone deacetylase inhibition accelerates the early events of stem cell differentiation: transcriptomic and epigenetic analysis. Genome Biol. 2008;9:R65. doi: 10.1186/gb-2008-9-4-r65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Svechnikova I, Almqvist PM, Ekstrom TJ. HDAC inhibitors effectively induce cell type-specific differentiation in human glioblastoma cell lines of different origin. Int J Oncol. 2008;32:821–827. [PubMed] [Google Scholar]
- 85.Schmittwolf C, Kirchhof N, Jauch A, Durr M, Harder F, et al. In vivo haematopoietic activity is induced in neurosphere cells by chromatin-modifying agents. Embo J. 2005;24:554–566. doi: 10.1038/sj.emboj.7600546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Brambrink T, Foreman R, Welstead GG, Lengner CJ, Wernig M, et al. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell. 2008;2:151–159. doi: 10.1016/j.stem.2008.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Finnin MS, Donigian JR, Cohen A, Richon VM, Rifkind RA, et al. Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature. 1999;401:188–193. doi: 10.1038/43710. [DOI] [PubMed] [Google Scholar]
- 88.Forneris F, Binda C, Dall'Aglio A, Fraaije MW, Battaglioli E, et al. A highly specific mechanism of histone H3-K4 recognition by histone demethylase LSD1. J Biol Chem. 2006;281:35289–35295. doi: 10.1074/jbc.M607411200. [DOI] [PubMed] [Google Scholar]
- 89.Forneris F, Binda C, Vanoni MA, Battaglioli E, Mattevi A. Human histone demethylase LSD1 reads the histone code. J Biol Chem. 2005;280:41360–41365. doi: 10.1074/jbc.M509549200. [DOI] [PubMed] [Google Scholar]
- 90.Ou JN, Torrisani J, Unterberger A, Provencal N, Shikimi K, et al. Histone deacetylase inhibitor Trichostatin A induces global and gene-specific DNA demethylation in human cancer cell lines. Biochem Pharmacol. 2007;73:1297–1307. doi: 10.1016/j.bcp.2006.12.032. [DOI] [PubMed] [Google Scholar]
- 91.Lee MG, Wynder C, Bochar DA, Hakimi MA, Cooch N, et al. Functional interplay between histone demethylase and deacetylase enzymes. Mol Cell Biol. 2006;26:6395–6402. doi: 10.1128/MCB.00723-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Antibodies used for Western blotting (WB) and chromatin-immunoprecipitation (ChIP).
(0.04 MB DOC)
Primers and TaqMan gene expression assays used for qPCR or ChIP-qPCR.
(0.07 MB DOC)
qPCR analysis of male germ cell marker gene expression in untreated GC-1 cells.
(0.04 MB DOC)
qPCR analysis of MyoD gene expression in untreated and treated GC-1 cells.
(0.03 MB DOC)
Effect of tranylcypromine and trichostatin A on global Histone H3K4 methylation and H3 acetylation levels. Western blot analyses of global histone (A) H3K9 acetylation, (B) pan-H3 acetylation, (C) H3K4 monomethylation, and (D) H3K4 dimethylation on protein extracts isolated from GC-1 cells either grown in culture medium (control, ctr), or treated with tranylcypromine (T), trichostatin A (TSA), or dimethylsulphoxid (DMSO; TSA solvent), or tranylcypromine and TSA (T+TSA). (E) Histone H3 visualized on a coomassie gel was used for normalization. Tranylcypromine alone does not affect global H3 acetylation (A,B) or H3K4 methylation (C,D) in GC-1 cells. TSA exposure causes a global increase in H3K9 (A) and total H3 acetylation (B) and, probably due to an indirect effect [91], H3K4 dimethylation (D). The combined treatment of T+TSA induces a global increase in H3K4 monomethylation (C), but does not further enhance the effect on H3 acetylation (A,B) or H3K4 dimethylation (C). Signals from proteins isolated from control cells served as the base and were defined as one. Fold change indicates the ratio of signal intensity in treated cells normalized to histone H3 over signal intensity in control cells normalized histone H3. Data are expressed as mean ± SEM; **p<0.01, ***p<0.001. Means with different letters are significantly different. N indicates the number of independent experiments.
(2.21 MB EPS)
Tranylcypromine and trichostatin A do not affect global histone H3K9 methylation or H3K4 trimethylation levels. Western blot analyses of global histone (A) H3K9 mono-, (B) H3K9 di-, (D) H3K9 tri-, and (E) H3K4 trimethylation on protein extracts isolated from GC-1 cells either grown in culture medium (control, ctr), or treated with tranylcypromine (T), trichostatin A (TSA), or dimethylsulphoxid (DMSO; TSA solvent), or tranylcypromine and TSA (T+TSA). (C) and (F) Histone H3 visualized on coomassie gels was used for normalization. Signals achieved with proteins isolated from control cells served as base and were defined as one. Fold change indicates the ratio of signal intensity in treated cells normalized to histone H3 over signal intensity in control cells normalized histone H3. Data are expressed as mean ± SEM. N indicates the number of independent experiments.
(3.02 MB EPS)