Abstract
Actin filaments are semiflexible polymers that display large-scale conformational twisting and bending motions. Modulation of filament bending and twisting dynamics has been linked to regulatory actin-binding protein function, filament assembly and fragmentation, and overall cell motility. The relationship between actin filament bending and twisting dynamics has not been evaluated. The numerical and analytical experiments presented here reveal that actin filaments have a strong intrinsic twist-bend coupling that obligates the reciprocal interconversion of bending energy and twisting stress. We developed a mesoscopic model of actin filaments that captures key documented features, including the subunit dimensions, interaction energies, helicity, and geometrical constraints coming from the double-stranded structure. The filament bending and torsional rigidities predicted by the model are comparable to experimental values, demonstrating the capacity of the model to assess the mechanical properties of actin filaments, including the coupling between twisting and bending motions. The predicted actin filament twist-bend coupling is strong, with a persistence length of 0.15–0.4 μm depending on the actin-bound nucleotide. Twist-bend coupling is an emergent property that introduces local asymmetry to actin filaments and contributes to their overall elasticity. Up to 60% of the filament subunit elastic free energy originates from twist-bend coupling, with the largest contributions resulting under relatively small deformations. A comparison of filaments with different architectures indicates that twist-bend coupling in actin filaments originates from their double protofilament and helical structure.
Introduction
Actin is an abundant and highly conserved cytoskeleton protein that powers a broad range of eukaryotic cell movements by self-associating into helical double-stranded filaments (1). Actin filaments also serve as tracks for contractile myosin molecular motors. Assembled filaments can reach tens of micrometers in length and are often arranged in networks and bundles that help determine cell shape and mechanical stability (2). Filament stability and assembly, network and bundle integrity and response, and overall force generation and cell motility are strongly influenced by the filament mechanical properties. Quantitative knowledge of actin filament elasticity is therefore of central importance for developing predictive molecular models of actin-based cell motility.
Actin filaments have complex mechanical properties and internal motions (bending, twisting, and subunit dynamics) occurring on different timescales and length scales. The thermal twisting and bending deformations of actin filaments occur on timescales of microseconds and milliseconds–seconds, respectively, and have been measured experimentally by various methods, including analysis of thermal fluctuations in shape (3–6), intrinsic curvature (7,8), rotational diffusion (9,10), single-molecule imaging (11) and nanometry (12,13), and all-atom simulations (14–16). Subunit dynamics are rapid (on a timescale of nanoseconds) and defined by small-amplitude fluctuations relative to filament dimensions (10).
Actin filament bending and twisting mechanics are typically described by continuum properties that assume an isotropic and homogeneous material (3,4,6,17,18). Filament bending and twisting rigidities are commonly associated with the bending persistence length (Lb) and torsional rigidity (C = LTkBT, where LT is the torsional persistence length), respectively. The total elastic free energy (Felastic) stored in the deformation (thermal or active) of an actin filament (described as a semiflexible continuum polymer) is determined by contributions from bending and twisting, as well any coupling that may exist between these motions (19), as defined by the elasticity Hamiltonian (see Table S1 in the Supporting Material for parameter and variable definitions and units, and Fig. S1 for illustration):
| (1) |
where the first product term on the rhs corresponds to bending, the second to twisting, and the third to twist-bend coupling; κ1, κ2, and κ3 are rotation strain vectors along the three filament axes, with κ3 representing that of the long axis (Fig. S1); and LTB is the twist-bend coupling persistence length. The elastic free-energy contributions arising from twist-bend coupling can be rather significant because it is proportional to 2LTB (20). Here, we assume that the coupling factors between twist (κ3) and the two bending rotation strains (κ1 and κ2) are identical, as predicted for homogeneous cylinders. Given the helical structure of actin filaments, anisotropies arising from local, noncylindrical fluctuations in shape will be averaged on length scales greater than the filament helical repeat. We also include an extra component, κ30, to account for the intrinsic twist in helical single- or double-stranded filaments (Fig. 1, C–E).
Figure 1.

Microscopic organization of model filaments. (A) Schematic of filament subunits modeled as ellipsoids connected by elastic bonds. The subunit centers of mass are given by G(k) or G(l) and their respective bond attachment coordinates by M(k)j and M(l)j). (B–E) Schematic of filament resting configurations: single-stranded nonhelical (B), single-stranded helical (C), aligned double-stranded helical (D), and staggered double-stranded helical (E). Longitudinal contacts are colored magenta, diagonal contacts are colored cyan, and lateral contacts are colored dark blue. (F) Schematic of subunit interface bond attachment dispersion areas. The interface is the convex envelope of the bond projections onto the plane normal to the line connecting the center of mass connecting two neighboring subunits.
Despite its potential contribution to the mechanical properties of filaments, the existence of twist-bend coupling and how it influences the conformational dynamics and energetics of actin filaments have not been evaluated. In this report, we demonstrate the existence and identify the origin of twist-bend coupling in actin filaments using a theoretical and geometric modeling approach that fills a gap between molecular-dynamics simulations and bulk rod-mechanics models of biopolymers. The derived mesoscopic models reveal the microscopic basis of actin filament mechanical properties and the origin of twist-bend coupling in complex linear polymers in general.
Materials and Methods
Simulation of Eqs. 2–4 supplemented with Eqs. S2 and S3 (Fig. 2) was carried out using the MATLAB (The MathWorks, Natick, MA) function ode15s (algorithm for stiff ordinary differential equations). The initial, resting filament configuration is linear. A constant inward load was applied to filament ends (Fig. S2), which caused the right- and leftmost filament subunit centers of mass to move toward the filament center. The new subunit positions correspond to points at a distance of three subunits' long-axis width (i.e., 3 × 6.7 = 21 nm) in the leftward direction for the rightmost filament subunit or in the rightward direction for the leftmost subunit. During simulations, the right- and leftmost subunit center positions were constrained at their new positions. However, in the absence of any torque applied to both filament ends, the two subunits were free to rotate around their center of mass. These particular initial conditions make the system of differential equations numerically stiff, hence the use of the MATLAB function ode15s.
Figure 2.

Equilibrium configurations of resting and strained filaments. Filaments of differing microscopic organization were loaded without twisting along the long filament axis. At equilibrium, configurations result from the balance between load-induced bending, responsible for large in-plane loops, and intrinsic twist-bend coupling arising from the microscopic subunit organization, responsible for out-of plane buckling. (A) Top and side views: single-stranded, nonhelical filaments do not present intrinsic twist-bend coupling. (B) Top and side views: helical filaments show minute out-of-plane buckling, indicating that the helicity present in the resting configuration is sufficient to drive the loop out of the bending plane. The equilibrium configuration for double-stranded with aligned (C) or staggered (D) subunits filaments presents a marked deviation from planar buckling alone, showing the presence of a strong coupling at the microscopic subunit arrangement level.
The addition of random forces in the pico-Newton range (comparable to forces of thermal origin) accelerates the convergence to equilibrium during the first initial calculation steps. Eqs. 5 and 6 obtained from Eqs. S4–S53, using appropriate relations (Eq. S24 or Eq. S52) were used to generate data presented in Figs. 3 and 4. In agreement with the use of the free-energy function normalized by the thermal energy factor kBT, the bond stiffness, usually given as J.m−2 (or equivalently as kcal/mole/Å2 (21)), is expressed in kBT.nm−2. The reported bond stiffness of 1 kcal/mole/Å2 for ATP-actin corresponds to ∼165 kBT.nm−2.
Figure 3.

Bending, twisting, and coupling persistence length landscapes. (A–D) Dependence of the bending, twisting, and coupling persistence lengths of single-stranded nonhelical (A), single-stranded helical (B), aligned double-stranded helical (C), and staggered double-stranded helical (D) filaments on the intersubunit bond stiffness and average subunit interface area. Note that the scales differ among panels and that both helicity and double-stranded structures contribute to the emergence of twist-bend coupling. The star and dot in panel D correspond to the average bond stiffness and interface areas of ATP-actin (14.6 nm2, 165 kBT.nm−2) and ADP-actin (14.6 nm2, 55 kBT.nm−2) filaments, respectively. The ATP- and ADP-actin filament bond stiffnesses range from 50 to 150 kcal mol−1 nm−2 and 20 to 50 kcal mol−1 nm−2, respectively (21). Normalization of the elastic free energy by kBT yields a corresponding range of 80–250 kBT nm−2 for ATP-actin and 30–80 kBT.nm−2 for ADP-actin filaments.
Figure 4.
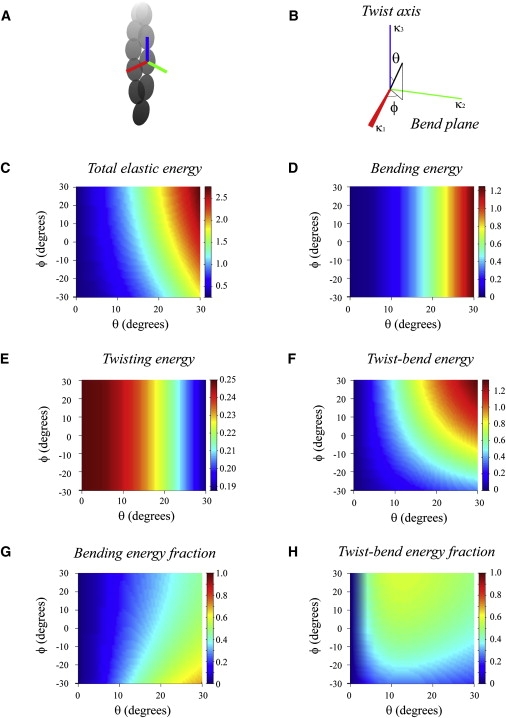
Contributions of bending, twisting, and coupling to the total elastic free energy. (A and B) Geometric coordinates associated with orientation of an individual filament subunit (see also Fig. S5) used to quantitate the elastic free-energy terms originating from bending, twisting, and coupling. The vector of strain rotation, κ, is determined by angles θ and ϕ, respectively. The angle between κ and the vertical axis is denoted by θ; the angle between the direction Ox and the projection of κ on the Oxy plane is ϕ. Note that a twist along the long subunit axis, parallel to the filament long axis, corresponds to a vector κ aligned with the vertical axis (i.e., θ = 0 or 180°); conversely, when θ = 90°, the rotation imposed to the subunit corresponds to bending. The angle ϕ controls the degree of coupling between bending and twisting strains. The rotation imposed to the subunit corresponds to an angle of 1 rad about the axis along the unit vector κ. (C–F) The dependence of total (C), bending (D), twisting (E), and coupling (F) elastic free energies on the angular strain rotation. (G and H) The fractional contributions from bending and coupling.
The average interaction area between subunits in an actin filament was determined by calculating the buried solvent-accessible surface area (SASA), defined as the difference in SASAs between the dimer and two free subunits. The SASA of an actin subunit, and the lateral and longitudinal dimers of the Oda filament model (22) were calculated using the calc-surface program (23) (accessed using the National Institutes of Health scientific supercomputing resource at http://helixweb.nih.gov/structbio/basic.html) for all atoms except waters in the PDB using a probe size of 1.4 Å.
Results and Discussion
Model filament geometries, subunit organization, and interfaces
We consider filaments as a series of nondeformable subunits with the dimensions of actin monomers (6.7 × 4.0 × 3.7 nm) (1,22) connected via elastic bonds dispersed uniformly over an interface of a defined area (Fig. 1 A). Each subunit is characterized by its center of mass position, G, and its orientation in three-dimensional space. The latter is defined by a set of three space vectors positioned along the three principal subunit axes (d1, d2, and d3, where d1 and d2 are unit vectors spanning the plane orthogonal to d3, the long filament axis (Fig. 1 A)). The bonds dispersed over the subunit interfaces are treated as extensible, freely rotating, and nondiffusible springs with constant stiffness (Fig. 1 A). We assume that the bonds adjoining a pair of connected subunits are parallel to the line connecting the subunit's center of mass. Therefore, the convex envelope of the bond projections onto the plane orthogonal to the line connecting the subunit defines the interface between these subunits. Below, we evaluate the effects of bond stiffness and interface area.
To determine the geometric factors that contribute to the filament's mechanical properties, we consider four different linear polymer architectures: 1), single-stranded nonhelical (Fig. 1 B); 2), single-stranded helical, equivalent to that of an individual actin filament strand (Fig. 1 C); 3), double-stranded helical with strands aligned (Fig. 1 D); and 4), double-stranded helical with strands staggered (Fig. 1 E). The latter mimics the structural organization of actin filaments. We do not consider double-stranded nonhelical filaments, because they are anisotropic in bending rigidity; the unit vectors spanning the plane orthogonal to the long filament axes (d1 and d2) are not equivalent, and therefore large-scale bending about d1 and d2 depends on the direction of the applied load.
The numbers of neighboring subunits in single- and double-stranded aligned and double-stranded staggered filaments differ (Fig. 1). Consequently, the relative position and total number of interfaces per subunit varies with the filament organization. Single-stranded linear and helical filament interfaces are restricted to longitudinal contacts (one on each end of a subunit, for a total of two per subunit; Fig. 1, B and C). Double-stranded aligned filaments are defined by three interface types (longitudinal, lateral and diagonal), with each subunit possessing five interfaces (two longitudinal, one lateral, and two diagonal; Fig. 1 D). Double-stranded staggered filaments (e.g., actin filaments) are defined by two interfaces (longitudinal and diagonal), with each subunit possessing two of each interface, for a total of four (Fig. 1 E). Note that what we define as diagonal contacts are commonly referred to as lateral actin filament contacts.
Equations for filament subunit displacement
In dynamical regimes characterized by a low Reynolds number (defined as the ratio of inertial forces to viscous forces) experienced by nanometer- to micrometer-sized biological macromolecules, inertial forces are insignificant and viscous forces dominate. Consequently, the balance of force and moment originating from elastic and viscous drags governs the dynamics of large-scale filament shape fluctuations and motion. In the case of slender bodies (e.g., filaments or elongated ellipsoids), the drag force exerted on the particle (filament subunit or segment) is proportional to the center of mass velocity. Hence, the viscous (CTdG(k)/dt) and elastic forces (F) experienced by the kth subunit (with nearest neighbor l) are balanced:
| (2) |
where CT is the translational drag coefficient of the entire filament, dG(k)/dt is the translational velocity, and F(k,l)j(k,l) is the elastic force arising from the extension of the jth bond between subunits k and l (Fig. 1 A). In the case of single-stranded filaments (Fig. 1, A and B), summation (Eq. 2) is limited to the two subunits flanking subunit k (i.e., l = k − 1 and k + 1). In the case of double-stranded filaments, the summation terms (Eq. 2) include two (Fig. 1 C) or one (Fig. 1 D) additional subunits on the opposite strand with a common interface contacting subunit k.
The moment of the elastic force (F) experienced by subunit k is determined by the net sum of all forces acting on subunit k times the change in center of mass position, and must balance the drag angular momentum, which is proportional to the product of the angular velocity (Ω(k)) and the rotational drag coefficient (CR):
| (3) |
The net rotation of subunit k is then given by the subunit orientation space vectors:
| (4) |
for i = 1, 2, 3 subunit axes. The global configuration of filaments in time and space are described by the geometry of filament subunit organization (Fig. 1) and the equations for filament subunit orientation and displacement (Eqs. 2–4).
Dependence of filament geometry on twist-bend coupling
To determine how the existence and magnitude of twist-bend coupling depends on the filament geometry, we computed the configuration of filaments under an external mechanical load applied at filament ends and evaluate the in- and out-of-plane deformations from the dispersion of subunit position in space (i.e., the fraction of total in plane subunits). The external load was applied without torque (i.e., no imposed twisting) toward the filament center. Under these conditions, out-of-plane buckling occurs only when twist-bend coupling exists; in the absence of twist-bend coupling, bending is confined to a plane (Fig. S2).
Single-stranded nonhelical filaments buckle in plane (Fig. 2 A). Single-stranded helical filaments buckle slightly out of plane, indicating that twist-bend coupling exists and that it arises from the helical organization of subunits since it is absent in nonhelical, single-stranded filaments (Fig. 2 B). Both aligned and staggered double-stranded filaments display more prominent twist-bend coupling than single-stranded helical filaments (Fig. 2, C and D).
These data reveal that three geometric factors govern the emergence of twist-bend coupling in linear polymers: 1), helicity; 2), a multistranded configuration; and 3), the relative alignment of subunits in the strands, with coupling in the staggered double-stranded alignment being the strongest of the configurations we considered in this study. Actin filaments possess all of these structural features and must therefore possess twist-bend coupling.
Elastic free energies of filaments
In the Supporting Material we provide derivations for the total free elastic energy stored in configurations of the various filament architectures evaluated in this study. The general expression for elastic free energy is:
| (5) |
where κ is a strain vector associated with local deformation, and the coefficients of matrix Q explicitly account for the different filament geometries and directly provide the filament mechanical parameters of interest, namely, the filament bending and torsional and twist-bend coupling persistence lengths. The bond stiffness and interface area dependence of the filament persistence length, torsional constant, and twist-bend coupling constant for the four filament types investigated in this study were computed using Eq. 5 (Fig. 3).
The elastic free-energy function derived in this work (Eq. 5 and Supporting Material) is formally equivalent to the Kirchhoff equation used in continuum model approaches (19). The advantage of this expression over the more familiar Kirchhoff equation is that all parameters are derived from the microscopic filament structure, and therefore explicitly account for the subunit dimensions, filament geometry (e.g.. helicity, number of strands, and staggered versus aligned double strands), the number and type (e.g.. lateral, diagonal, and longitudinal) of subunit interface bonds, and the intersubunit bond dispersion area.
Dependence of subunit interface areas and bond stiffness on filament mechanics
The filament mechanical properties (bending, twisting, and coupling; we assume that filaments are essentially inextensible—a reasonable approximation given the ∼2.3 GPa longitudinal elastic modulus (24,25) obtained from muscle fiber stiffness measurements and the <1% extensibility observed in contracting muscle (26–28)) depend on the bond stiffness and interface area (Fig. 3; note that the scales vary), but the extent to which they contribute is a function of the geometry, specifically the number of strands comprising the filament. Single-stranded filaments, helical and nonhelical, readily bend and twist unless the interface areas and/or bond stiffness are large (Fig. 3 A). Single-stranded helical filaments are marginally harder to bend and twist than single-stranded, nonhelical filaments due to their cylindrical width and the existence (albeit weak) of twist-bend coupling, respectively.
The bending, twisting, and coupling of aligned and staggered double-stranded filaments depend on the subunit interface area and intersubunit bond stiffness (Fig. 3, C and D). Double-stranded filaments have considerably larger bending persistence lengths than filaments of single-stranded geometries (Fig. 3, A and B), as predicted from contributions to the geometric moment and radial mass distribution. The twisting and coupling persistence lengths of double-stranded filaments are also larger by many orders of magnitude. The major determinant of twist-bend coupling in filaments is the double-stranded structure (geometry), although the integrity (bond stiffness) of the subunit contacts also contributes.
Mechanical properties of actin filaments
Knowledge of the actin filament bond stiffness and interface areas permits evaluation of the mesoscopic (double-stranded, staggered) filament model and theory presented in this work, as well as the development of novel predictions regarding actin filament mechanical properties. The interface contacts adjoining adjacent subunits in an actin filament behave as harmonic bonds with an effective stiffness that depends on the actin-bound nucleotide, displaying a range of 50–150 kcal mol−1 nm−2 for ATP-actin and 20–50 kcal mol−1 nm−2 for ADP-actin (21). The subunit interface areas estimated from the atomic actin filament model are ∼20 nm2 for longitudinal and ∼10 nm2 for lateral (i.e., diagonal) interactions, respectively (22,29), yielding a total interface area of ∼60 nm2 per subunit and an average of ∼15 nm2 per interface because each actin filament subunit contacts four neighboring subunits (Fig. 3 D).
The bending persistence lengths of ATP- and ADP-actin filaments predicted for double-stranded, staggered filaments agree with experimental values within a factor of ∼≤2 (4,5) (Table 1). The predicted torsional persistence length of ADP-actin filaments also agrees well with the experimentally determined value (Table 1). This small discrepancy may arise from uncertainties in the estimated bond stiffness (we use the average in our calculations) and/or subunit interface areas, our use of the average (lateral and longitudinal) interface area, and/or the existence of some subunit compliance (22).
Table 1.
Summary of actin filament mechanical properties
| Nucleotide | LB (predicted) | LB (measured) | LT (predicted) | LT (measured)† | LTB (predicted) |
|---|---|---|---|---|---|
| MgATP | 12.4 μm | 13 μm∗ | 1.4 μm | — | 0.4 μm |
| MgADP | 4.2 μm | 9.1 (±0.3) μm‡ | 0.45 μm | 0.5 (±0.2) μm | 0.15 μm |
The value of the MgATP actin filament bending persistence length (LP) is estimated from that of BeFx actin filaments (4).
The values of the filament torsional persistence length (LT) are calculated from the torsional rigidity (C = 2.3 ±1.0 × 10−27 N m−2 rad−1 (10);) using LT = C/(kBT) with kBT = 4.1 × 10−21 J.
The MgADP actin filament bending persistence length (LP) value represents an average of previously reported values (4,5,18).
Although actin filaments in solution are dynamic and adopt multiple structural states (30), the close agreement between the model predictions and the experimentally observed parameters indicates that the (double-stranded, staggered) geometric model reliably captures not only the predominant structural features of actin filaments but also their bending and twisting mechanics. Therefore, our approach permits prediction of twist-bend coupling in actin filaments, which we estimate to be 0.4 μm for ATP-actin and 0.15 μm for ADP-actin filaments (Fig. 3 D, Table 1). The nucleotide dependence of the coupling constant arises from the different greater subunit interface bond stiffness of ATP-actin. We note that the flexural rigidity of actin filaments calculated from the persistence length includes twist-bend coupling contributions and therefore underestimates the true intrinsic filament flexural rigidity.
Distribution of bending, twisting, and coupling elastic free energies under deformation
The fractional contributions of bending, twisting, and coupling to the total filament elastic free energy depend on the type (bend and/or twist) of deformation. The normalized total elastic free-energy density stored in a given filament configuration is a function of the external strain (defined by variables κ1, κ2, and κ3). The condition ensures energy density positivity under all strains and is fulfilled with the predicted filament persistence lengths of all filament types evaluated (Table 1), indicating the thermodynamic validity of the models and theory presented in this study.
The total elastic free-energy fraction originating from twist-bend coupling ,
| (6) |
is independent of the strain vector amplitude (i.e., the extent of imposed deformation). The fraction can be computed for different strain vector orientations in three-dimensional space using a spherical angle representation (Fig. 4, A and B). The total elastic free energy of a filament subunit and the fractional bending and twisting contributions can also be obtained (Fig. 4, C–F).
As predicted from the theory of elastic rods, the total elastic free energy of an actin filament subunit (note that this does not refer to the elastic energy of the entire filament, but to that of an individual subunit) depends on the magnitude and type of deformation (Fig. 4 C). The existence of twist-bend coupling introduces asymmetry (with respect to bend and twist deformation) to the total free-energy landscape (Fig. 4 C) despite the perfectly symmetrical and isotropic (i.e., they do not depend on the angle ϕ) bending and twisting energy landscapes (Fig. 4, D and E). Twist-bend coupling contributes a substantial fraction of the total filament elastic free energy—up to ∼60%—under small deformations (Fig. 4 H), such as those experienced by thermally fluctuating actin filaments (3–6,10). This elastic behavior of actin filaments differs from that of DNA. In the case of DNA, twist-bend coupling is relatively insignificant in most situations (31), particularly when the polymer length exceeds that of a single helical turn (32–34), because DNA stretches under tension (20). The relative contribution from pure twisting is relatively small (≤3% of total elastic energy) at all deformations (Fig. 4 E) because filaments twist more easily than they bend (LT < LP; Table 1).
Conclusions
The models and theory developed in this study demonstrate the existence of twist-bend coupling as an emergent property of actin filaments. Twist-bend coupling can dramatically influence the energetic and mechanical properties of filaments, such that conformational (elastic) energy is either stored or dissipated depending on the combination of bending and twisting deformation. This behavior arises because of the helical and double-stranded organization of actin filament subunits, which allows for interconversion of bending and twisting elastic energies.
Twist-bend coupling is an intrinsic property of helical multistranded helical polymers with stable interstrand interactions. Coupling between bending and twisting likely reflects a general functional adaptation of multistranded polymers, in similarity to a critical concentration and rate-limiting nucleation of assembly, terminal subunit addition and dissociation, and long length distribution (25). The emergence of twist-bend coupling introduces important physicochemical properties that can modulate energy transduction and work production, which may reflect a general evolutionary adaptation for driving cell global organization.
The reciprocal conversion of bending and twisting energies enables actin filaments and filament assemblies (i.e., bundles and networks) to respond adaptively to external stresses by allowing elastic energies associated with external bending or twisting loads to dissipate through random fluctuations if the filaments are free to translate or rotate, thereby modulating mechanical properties and preventing large-scale deformation or possible failure. Alternatively, elastic energy can be stored and subsequently released in the form of work production when actin filament motions are restricted, as occurs in assemblies or near surfaces (e.g., cell membranes and organelles). Such dissipation of stored elastic free energy can drive motility (35).
The existence of twist-bend coupling in actin filaments could therefore have important implications for cell physiology and function. Coupling likely contributes to actin-based cell motility and mechanics because it is most prominent at small filament deformations such as those associated with subunit incorporation (i.e., filament growth) at the leading edge of migrating cells (35), interaction with regulatory proteins (36–38), displacement of filament bundles (39), and interaction with contractile motor proteins (40). Modulation of filament bending (5) and twisting (10) dynamics by regulatory proteins could contribute to stress accumulation, which increases the probability of failure (i.e., filament severing (5,36,37)).
Twist-bend coupling may also play a role in higher-order assemblies, such as filament bundles and networks, in which filament motions are restricted. The contributions from twist-bend coupling to the elastic properties of such assemblies have yet to be evaluated. However, given that coupling can dominate contributions to the filament elastic energy, it may be critical for driving the assembly dynamics of lamellipodia, filopodia, and other higher-order actin filament structures.
Acknowledgments
Author contributions: E.M.D.L.C., L.B., and J.L.M. designed the research; E.M.D.L.C., J.R., B.M., and J.L.M. performed research; and E.M.D.L.C. and J.L.M. contributed new reagents or analytic tools, analyzed data, and wrote the article.
This work was supported by grants from the National Institutes of Health (GM071688 and GM071688-03S1 to E.M.D.L.C.), the American Heart Association (0940075N to E.M.D.L.C.), the Agence Nationale de la Recherche (ANR-06-PCV-135054 and ANR-08-SYSC-013 to L.B. and J.L.M.), and the Institute of Complex Systems IXXI, Rhône-Alpes (to J.L.M.). E.M.D.L.C. is an American Heart Association established investigator, an NSF-CAREER Award recipient (MCB-0546353), and a Hellman Family fellow.
Contributor Information
Enrique M. De La Cruz, Email: enrique.delacruz@yale.edu.
Jean-Louis Martiel, Email: jean-louis.martiel@imag.fr.
Supporting Material
References
- 1.Holmes K.C., Popp D., Kabsch W. Atomic model of the actin filament. Nature. 1990;347:44–49. doi: 10.1038/347044a0. [DOI] [PubMed] [Google Scholar]
- 2.Fletcher D.A., Mullins R.D. Cell mechanics and the cytoskeleton. Nature. 2010;463:485–492. doi: 10.1038/nature08908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gittes F., Mickey B., Howard J. Flexural rigidity of microtubules and actin filaments measured from thermal fluctuations in shape. J. Cell Biol. 1993;120:923–934. doi: 10.1083/jcb.120.4.923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Isambert H., Venier P., Carlier M.F. Flexibility of actin filaments derived from thermal fluctuations. Effect of bound nucleotide, phalloidin, and muscle regulatory proteins. J. Biol. Chem. 1995;270:11437–11444. doi: 10.1074/jbc.270.19.11437. [DOI] [PubMed] [Google Scholar]
- 5.McCullough B.R., Blanchoin L., De la Cruz E.M. Cofilin increases the bending flexibility of actin filaments: implications for severing and cell mechanics. J. Mol. Biol. 2008;381:550–558. doi: 10.1016/j.jmb.2008.05.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Le Goff L., Hallatschek O., Amblard F. Tracer studies on f-actin fluctuations. Phys. Rev. Lett. 2002;89:258101. doi: 10.1103/PhysRevLett.89.258101. [DOI] [PubMed] [Google Scholar]
- 7.Orlova A., Egelman E.H. A conformational change in the actin subunit can change the flexibility of the actin filament. J. Mol. Biol. 1993;232:334–341. doi: 10.1006/jmbi.1993.1393. [DOI] [PubMed] [Google Scholar]
- 8.De La Cruz E.M., Mandinova A., Pollard T.D. Polymerization and structure of nucleotide-free actin filaments. J. Mol. Biol. 2000;295:517–526. doi: 10.1006/jmbi.1999.3390. [DOI] [PubMed] [Google Scholar]
- 9.Prochniewicz E., Zhang Q., Thomas D.D. Microsecond rotational dynamics of actin: spectroscopic detection and theoretical simulation. J. Mol. Biol. 1996;255:446–457. doi: 10.1006/jmbi.1996.0037. [DOI] [PubMed] [Google Scholar]
- 10.Prochniewicz E., Janson N., De la Cruz E.M. Cofilin increases the torsional flexibility and dynamics of actin filaments. J. Mol. Biol. 2005;353:990–1000. doi: 10.1016/j.jmb.2005.09.021. [DOI] [PubMed] [Google Scholar]
- 11.Forkey J.N., Quinlan M.E., Goldman Y.E. Measurement of single macromolecule orientation by total internal reflection fluorescence polarization microscopy. Biophys. J. 2005;89:1261–1271. doi: 10.1529/biophysj.104.053470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Tsuda Y., Yasutake H., Yanagida T. Torsional rigidity of single actin filaments and actin-actin bond breaking force under torsion measured directly by in vitro micromanipulation. Proc. Natl. Acad. Sci. USA. 1996;93:12937–12942. doi: 10.1073/pnas.93.23.12937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yasuda R., Miyata H., Kinosita K., Jr. Direct measurement of the torsional rigidity of single actin filaments. J. Mol. Biol. 1996;263:227–236. doi: 10.1006/jmbi.1996.0571. [DOI] [PubMed] [Google Scholar]
- 14.Chu J.W., Voth G.A. Allostery of actin filaments: molecular dynamics simulations and coarse-grained analysis. Proc. Natl. Acad. Sci. USA. 2005;102:13111–13116. doi: 10.1073/pnas.0503732102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pfaendtner J., Branduardi D., Voth G.A. Nucleotide-dependent conformational states of actin. Proc. Natl. Acad. Sci. USA. 2009;106:12723–12728. doi: 10.1073/pnas.0902092106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pfaendtner J., De La Cruz E.M., Voth G.A. Actin filament remodeling by actin depolymerization factor/cofilin. Proc. Natl. Acad. Sci. USA. 2010;107:7299–7304. doi: 10.1073/pnas.0911675107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Käs J., Strey H., Janmey P.A. F-actin, a model polymer for semiflexible chains in dilute, semidilute, and liquid crystalline solutions. Biophys. J. 1996;70:609–625. doi: 10.1016/S0006-3495(96)79630-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Greenberg M.J., Wang C.L., Moore J.R. Modulation of actin mechanics by caldesmon and tropomyosin. Cell Motil. Cytoskeleton. 2008;65:156–164. doi: 10.1002/cm.20251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Landau L.D., Lifshitz E.M. Elsevier; Amsterdam: 2002. Theory of Elasticity. [Google Scholar]
- 20.Marko J.F. DNA under high tension: overstretching, undertwisting, and relaxation dynamics. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 1998;57:2134–2149. [Google Scholar]
- 21.Chu J.W., Voth G.A. Coarse-grained modeling of the actin filament derived from atomistic-scale simulations. Biophys. J. 2006;90:1572–1582. doi: 10.1529/biophysj.105.073924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Oda T., Iwasa M., Narita A. The nature of the globular- to fibrous-actin transition. Nature. 2009;457:441–445. doi: 10.1038/nature07685. [DOI] [PubMed] [Google Scholar]
- 23.Gerstein M. A resolution-sensitive procedure for comparing protein surfaces and its application to the comparison of antigen-combining sites. Acta Crystallogr. A. 1992;48:271–276. [Google Scholar]
- 24.Higuchi H., Yanagida T., Goldman Y.E. Compliance of thin filaments in skinned fibers of rabbit skeletal muscle. Biophys. J. 1995;69:1000–1010. doi: 10.1016/S0006-3495(95)79975-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Howard J. Sinauer Associates; Sunderland, MA: 2001. Mechanics of Motor Proteins and the Cytoskeleton. [Google Scholar]
- 26.Huxley H.E., Stewart A., Irving T. X-ray diffraction measurements of the extensibility of actin and myosin filaments in contracting muscle. Biophys. J. 1994;67:2411–2421. doi: 10.1016/S0006-3495(94)80728-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wakabayashi K., Sugimoto Y., Amemiya Y. X-ray diffraction evidence for the extensibility of actin and myosin filaments during muscle contraction. Biophys. J. 1994;67:2422–2435. doi: 10.1016/S0006-3495(94)80729-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bordas J., Svensson A., Boesecke P. Extensibility and symmetry of actin filaments in contracting muscles. Biophys. J. 1999;77:3197–3207. doi: 10.1016/S0006-3495(99)77150-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sept D., McCammon J.A. Thermodynamics and kinetics of actin filament nucleation. Biophys. J. 2001;81:667–674. doi: 10.1016/S0006-3495(01)75731-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Reisler E., Egelman E.H. Actin structure and function: what we still do not understand. J. Biol. Chem. 2007;282:36133–36137. doi: 10.1074/jbc.R700030200. [DOI] [PubMed] [Google Scholar]
- 31.Marko J.F., Siggia E.D. Bending and twisting elasticity of DNA. Macromolecules. 1994;27:981–988. [Google Scholar]
- 32.Lankas F., Sponer J., Langowski J. Sequence-dependent elastic properties of DNA. J. Mol. Biol. 2000;299:695–709. doi: 10.1006/jmbi.2000.3781. [DOI] [PubMed] [Google Scholar]
- 33.Panyukov S., Rabin Y. Fluctuating filaments: statistical mechanics of helices. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics. 2000;62(5 Pt B):7135–7146. doi: 10.1103/physreve.62.7135. [DOI] [PubMed] [Google Scholar]
- 34.Panyukov S., Rabin Y. Thermal fluctuations of elastic filaments with spontaneous curvature and torsion. Phys. Rev. Lett. 2000;85:2404–2407. doi: 10.1103/PhysRevLett.85.2404. [DOI] [PubMed] [Google Scholar]
- 35.Mogilner A., Oster G. Cell motility driven by actin polymerization. Biophys. J. 1996;71:3030–3045. doi: 10.1016/S0006-3495(96)79496-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.De La Cruz E.M. Cofilin binding to muscle and non-muscle actin filaments: isoform-dependent cooperative interactions. J. Mol. Biol. 2005;346:557–564. doi: 10.1016/j.jmb.2004.11.065. [DOI] [PubMed] [Google Scholar]
- 37.De La Cruz E.M. How cofilin severs an actin filament. Biophys. Rev. 2009;1:51–59. doi: 10.1007/s12551-009-0008-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Prochniewicz E., Henderson D., Thomas D.D. Dystrophin and utrophin have distinct effects on the structural dynamics of actin. Proc. Natl. Acad. Sci. USA. 2009;106:7822–7827. doi: 10.1073/pnas.0812007106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bathe M. A finite element framework for computation of protein normal modes and mechanical response. Proteins. 2008;70:1595–1609. doi: 10.1002/prot.21708. [DOI] [PubMed] [Google Scholar]
- 40.Prochniewicz E., Chin H.F., De La Cruz E.M. Myosin isoform determines the conformational dynamics and cooperativity of actin filaments in the strongly bound actomyosin complex. J. Mol. Biol. 2010;396:501–509. doi: 10.1016/j.jmb.2009.11.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


