Abstract
Peroxisome proliferator-activated receptor-γ (PPARγ) agonists, a new class of antidiabetic agents, have been shown to possess antiinflammatory activity. In this study, we investigated the molecular mechanism by which PPARγ agonists inhibit proinflammatory cytokine expression in rat glomerular mesangial cells. Both natural and synthetic PPARγ agonists potently inhibited RANTES (regulated upon activation, normal T cell expressed and secreted) and monocyte chemoattractant protein-1 expression induced by TNF-α in mesangial cells, which was dependent on NF-κB signaling. However, PPARγ agonists had little effect on TNF-α-triggered IκBα phosphorylation and its subsequent degradation, p65 phosphorylation, and nuclear translocation. In the absence of PPARγ ligand, TNF-α induced a physical interaction between nuclear p65 and PPARγ, as demonstrated by co-immunoprecipitation. Such an interaction was mediated by the C-terminal region of p65. Activation of PPARγ by its agonist prevented PPARγ·p65 complex formation. Chromatin immunoprecipitation assay revealed that TNF-α induced p65 binding to the cis-acting κB elements in rat RANTES promoter, whereas disruption of PPARγ·p65 by its agonist blocked p65 interaction with its cognate κB sites. Knockdown of PPARγ via siRNA strategy completely abolished TNF-α-mediated p65 binding to κB sites and negated RANTES induction, suggesting that unliganded PPARγ is obligatory for NF-κB signaling. Consistently, overexpression of PPARγ in the absence of its ligand sensitized mesangial cells to TNF-α stimulation. These results uncover a paradoxical action of the unliganded and ligand-activated PPARγ in regulating NF-κB signaling and demonstrate PPARγ ligand as a molecular switch that controls its ability to modulate inflammatory responses in opposite directions.
Keywords: Chemokines, Gene Regulation, Inflammation, Kidney, NF-κB, PPAR, Tumor Necrosis Factor
Introduction
Peroxisome proliferator-activated receptor-γ (PPARγ),2 a ligand-dependent transcription factor that belongs to a subclass of the nuclear hormone receptor superfamily, plays a pivotal role in regulating a wide variety of biological processes such as insulin sensitivity, immune response, adipogenesis, and glucose homeostasis (1–3). PPARγ is also the molecular target of thiazolidinediones, which are insulin-sensitizing drugs that are used clinically in the treatment of type 2 diabetes (3–5). Although the mechanism underlying its insulin sensitization action remains to be fully elucidated, PPARγ activation by its agonists is known to negatively regulate the stimulus-dependent production of numerous inflammatory mediators that promote an insulin-resistant state (6–8). Evidence shows that PPARγ agonists are able to inhibit the expression and/or biological effects of tumor necrosis factor-α (TNF-α), interleukin-6 (IL-6), monocyte chemoattractant protein-1 (MCP-1), plasminogen activator inhibitor-1, and angiotensinogen (9). These studies underscore that the antiinflammatory potential of PPARγ agonists may play a crucial role in mediating their beneficial actions.
A large body of evidence demonstrates that PPARγ agonists also ameliorate renal fibrotic lesions and kidney dysfunction in both diabetic nephropathy and a variety of nondiabetic chronic kidney diseases (10–15). In rat remnant kidney model, PPARγ agonist troglitazone reduces proteinuria and serum creatinine, attenuates glomerulosclerosis, and suppresses plasminogen activator inhibitor-1 and transforming growth factor-β (TGF-β) expression (14). Similarly, in antiglomerular basement membrane nephritis rats, PPARγ agonist reduces urinary protein excretion and crescent formation and inhibits glomerular infiltration of monocytes/macrophages (15). Given that renal infiltration of inflammatory cells is an imperative pathomechanism in these disorders (16, 17), it is conceivable that PPARγ agonists may ameliorate renal injury primarily by inhibiting inflammation rather than by regulating the insulin sensitivity or glucose metabolism. However, the molecular mechanism by which PPARγ activation leads to inhibition of renal inflammation remains poorly understood.
The initiation of inflammatory gene expression is predominantly driven by nuclear factor-κB (NF-κB), a master transcription factor that trans-activates the expression of its target genes via binding to its cognate sequence-specific κB site (18, 19). Not surprisingly, NF-κB signaling is tightly controlled at multiple levels. In the basal, unstimulated state, p65 NF-κB is localized in the cytoplasm and associated with and inhibited by cytoplasmic inhibitory protein IκB. Upon stimulation by various cues such as TNF-α, IκB is phosphorylated and subsequently subjected to ubiquitin-mediated degradation, thereby liberating p65 and allowing it to undergo nuclear translocation (20). Earlier studies indicate that PPARγ agonists negatively influence NF-κB signaling by several possible mechanisms (21, 22), ranging from the up-regulation of IκBα expression to the reduction of p65 nuclear translocation. Whether PPARγ interacts directly with p65 in the nuclei in the postnuclear stage of NF-κB signaling, however, is unknown.
In this study, we have investigated the effect of PPARγ agonists on proinflammatory cytokine expression and dissected the underlying mechanism in rat glomerular mesangial cells. We found that PPARγ physically interacts with activated p65 in the nuclei; and unexpectedly, such an interaction is obligatory for NF-κB signaling. Our studies uncover a novel mechanism by which unliganded and ligand-activated PPARγ exert an opposite action in regulating NF-κB activity and inflammatory cytokine expression in mesangial cells.
EXPERIMENTAL PROCEDURES
Cell Culture and Treatment
Rat glomerular mesangial cells were provided by Dr. Cary Wu (University of Pittsburgh, Pittsburgh, PA) and cultured in DMEM-Ham's F12 medium supplemented with 10% fetal bovine serum (Invitrogen), as previously described (23, 24). Cells were seeded in culture plates to 60–70% confluence in the complete medium and incubated for 16 h prior to serum starvation. Serum-starved cells were treated with TNF-α (R&D Systems) at a final concentration of 5 ng/ml, or 15-deoxy-Δ12,14-prostaglandin J2 (15d-PGJ2) (Cayman Chemical, Ann Arbor, MI), troglitazone and ciglitazone (BIOMOL Research Laboratories, Plymouth Meeting, PA) at various concentrations as indicated. For some experiments such as assessing NF-κB activation, mesangial cells were pretreated with PPARγ agonists for 0.5 h followed by incubation with TNF-α for different periods of time as indicated. For blocking NF-κB signaling, mesangial cells were pretreated with a cell-permeable inhibitor peptide NF-κB SN50 (Calbiochem) (45 μm) for 1 h and then incubated with TNF-α. For knocking down PPARγ expression, mesangial cells were transfected with either control siRNA or PPARγ-specific siRNA (Ambion, Austin, TX) by using Lipofectamine 2000 reagent (Invitrogen).
Western Blot Analysis
Cell lysates were prepared as described previously (23). Samples were heated at 100 °C for ∼5–10 min before loading and separated on 10% SDS-polyacrylamide gels. Western blot analysis of protein expression was carried out by using routine procedures as described elsewhere (25). The primary antibodies were obtained from following sources: anti-RANTES (sc-1410), anti-MCP-1 (sc-1304), anti-PPARγ (sc-7273) (Santa Cruz Biotechnology, Santa Cruz, CA), anti-p65 NF-κB, anti-phospho-p65 NF-κB (Ser536), anti-IκBα and anti-phospho-IκBα (Ser32/Ser36) (Cell Signaling Technology, Danvers, MA), and anti-α-tubulin (Sigma).
Nuclear Protein Preparation
For preparation of nuclear protein, mesangial cells after various treatments as indicated were washed twice with cold phosphate-buffered saline (PBS) and scraped off the plate with a rubber policeman. After centrifugation, cell pellets were resuspended in buffer A (10 mm HEPES, pH 7.9, 1.5 mm MgCl2, 10 nm KCl, 0.5% Nonidet P-40, and 1% protease inhibitor mixture (Sigma)) and lysed with homogenizer. Cell nuclei were collected by centrifugation at 5,000 rpm for 15 min. After washing with buffer B (10 mm HEPES, pH 7.9, 1.5 mm MgCl2, 10 nm KCl, and 1% protease inhibitor mixture), nuclei were lysed in SDS sample buffer and subjected to Western blot analysis as described previously (26).
Immunofluorescence Staining
Indirect immunofluorescence staining was carried out according to the procedures described previously (20). Briefly, mesangial cells cultured on coverslips were washed with cold PBS twice and fixed with cold methanol for 10 min at −20 °C. After extensive washing with PBS containing 0.5% BSA, cells were blocked with 20% normal donkey serum in PBS and then incubated with specific primary antibodies against p65 NF-κB and PPARγ. For visualizing primary antibodies, cells were stained with cyanine Cy3- or Cy5-conjugated secondary antibodies (Jackson ImmunoResearch Laboratories, West Grove, PA). Cells were double stained with DAPI (4′,6-diamidino-2-phenylindole, HCl) to visualize the nuclei. Nonimmune normal control IgG was used to replace the primary antibody as negative control, and no staining occurred. Slides were viewed with a Nikon Eclipse E600 microscope equipped with a digital camera (Melville, NY).
Plasmid Constructs and Transfection
The green fluorescence protein (GFP)-tagged p65 NF-κB expression vector (pGFP-p65) was kindly provided by Dr. Johannes A. Schmid (University of Vienna, Vienna, Austria) (27). The p65 deletion mutants, p65Δ(18–167) and p65Δ(308–551), cloned in the pEF/myc/nuc expression vector (Invitrogen), were described elsewhere (28) and provided by Dr. Mark S. Nanes (Emory University, Atlanta, GA). Mouse PPARγ expression vector was described previously (29) Rat mesangial cells were transiently transfected with various expression vectors by using Lipofectamine 2000 reagent (Invitrogen). Cell lysates were subjected to subsequent immunoblotting and immunoprecipitation, respectively.
Immunoprecipitation
Immunoprecipitation was carried out by using an established method (20). Briefly, mesangial cells were transfected with GFP-tagged p65 NF-κB expression vector (pGFP-p65), or Myc-tagged p65 deletion mutants by using Lipofectamine 2000 reagent, and then incubated with or without TNF-α and/or 15d-PGJ2 for 30 min as indicated. Cells were lysed on ice in 1 ml of nondenaturing lysis buffer that contained 1% Triton X-100, 0.01 m Tris-HCl, pH 8.0, 0.14 m NaCl, 0.025% NaN3, 1% protease inhibitors mixture, and 1% phosphatase inhibitors mixture I and II (Sigma). After preclearing with normal IgG, cell lysates (1 mg of protein) were incubated overnight at 4 °C with 4 μg of anti-PPARγ (Santa Cruz Biotechnology), followed by precipitation with 30 μl of protein A/G plus-agarose for 1 h at 4 °C. The precipitated complexes were separated on SDS-polyacrylamide gels and immunoblotted with anti-GFP (ab-6556; Abcam, Cambridge, MA) or anti-Myc (catalog no. 9402; Cell Signaling Technology) antibodies.
Chromatin Immunoprecipitation (ChIP) Assay
ChIP assay was performed to analyze in vivo interactions of NF-κB and its cognate cis-acting element in rat RANTES promoter (20, 30). This assay was carried out essentially according to the protocols specified by the manufacturer (ChIP assay kit; Upstate, Charlottesville, VA). Briefly, mesangial cells after various treatments as indicated were cross-linked with 1% formaldehyde and then resuspended in SDS lysis buffer containing protease inhibitors. The chromatin solution was sonicated, and the supernatant was diluted 10-fold. An aliquot of total diluted lysate was used for total genomic DNA as input DNA control. The anti-p65 NF-κB antibody (sc-109; Santa Cruz Biotechnology) was added and incubated at 4 °C overnight, followed by incubation with protein A-agarose for 1 h. The precipitates were washed, and chromatin complexes were eluted. After reversal of the cross-linking at 65 °C for 4 h, the DNA was purified, and ChIP samples were used as a template for PCR using the primer sets for rat RANTES promoter regions (from −114 to +36) containing two putative κB binding sites (31). The sequences of primers used for ChIP assay were as follows: forward, 5′-ctgacagcagccagggtttg-3′; and reverse, 5′-agatgcatgcgttgtctcag-3′, which generated a PCR product of 150 bp. PCR samples were analyzed by electrophoresis on a 2.0% agarose gel.
Statistical Analyses
All data examined were expressed as mean ± S.E. Statistical analyses of the data were performed using SigmaStat software (Jandel Scientific, San Rafael, CA). Comparison between groups was made using one-way ANOVA, followed by Student-Newman-Keuls test. p < 0.05 was considered significant.
RESULTS
PPARγ Agonists Inhibit RANTES and MCP-1 Expression in Mesangial Cells
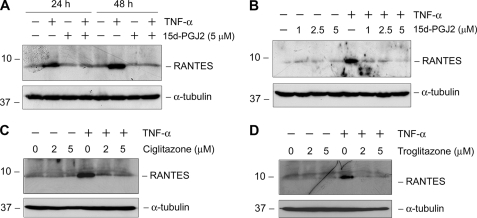
We first examined the effect of PPARγ agonists on the expression of proinflammatory cytokines in mesangial cells. As shown in Fig. 1A, expression of RANTES, also known as CC-chemokine ligand 5 (CCL5), was markedly induced in mesangial cells after incubation with TNF-α. Interestingly, PPARγ agonist 15d-PGJ2 completely abolished RANTES induction. The inhibition of RANTES induction by 15d-PGJ2 was quite effective, as it fully negated the TNF-α-mediated RANTES induction at the concentration as low as 1 μm (Fig. 1B). Similarly, ciglitazone and troglitazone, two synthetic PPARγ agonists, also completely blocked RANTES induction following TNF-α stimulation (Fig. 1, C and D).
FIGURE 1.
PPARγ agonists inhibit RANTES expression induced by TNF-α in rat glomerular mesangial cells. Mesangial cells were incubated with TNF-α (5 ng/ml) in the absence or presence of either endogenous (A and B) or synthetic PPARγ agonists (C and D) as indicated. Cells were treated either with 15d-PGJ2 (5 μm) for different periods of time (A) or various concentrations (B) for 48 h or with different doses of ciglitazone (C) and troglitazone (D) for 48 h. Cell lysates were immunoblotted with antibodies against RANTES or α-tubulin, respectively.
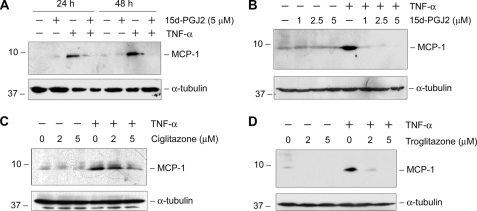
We also assessed the effects of PPARγ agonists on the expression of MCP-1, also known as CCL2, another major chemokine that is involved in regulating renal inflammation (32). As presented in Fig. 2, TNF-α also markedly induced MCP-1 expression in mesangial cells, and activation of PPARγ by 15d-PGJ2 completely abolished MCP-1 induction. Likewise, ciglitazone and troglitazone also inhibited MCP-1 expression induced by TNF-α (Fig. 2, C and D). Together, these results indicate that activation of PPARγ by its agonists effectively inhibits inflammatory response by preventing RANTES and MCP-1 expression in mesangial cells.
FIGURE 2.
PPARγ agonist also inhibits MCP-1 expression in rat mesangial cells. Rat mesangial cells were treated with TNF-α (5 ng/ml) in the absence or presence of either endogenous (A and B) or synthetic PPARγ agonists (C and D) as indicated. Cell lysates were prepared after incubation with 15d-PGJ2 (5 μm) for different periods of time (A) or various concentrations (B) for 48 h or with different doses of ciglitazone (C) and troglitazone (D) for 48 h, followed by immunoblotting with antibodies against MCP-1 or α-tubulin, respectively.
PPARγ Agonist Displays Little Effect on the Early Events of NF-κB Activation
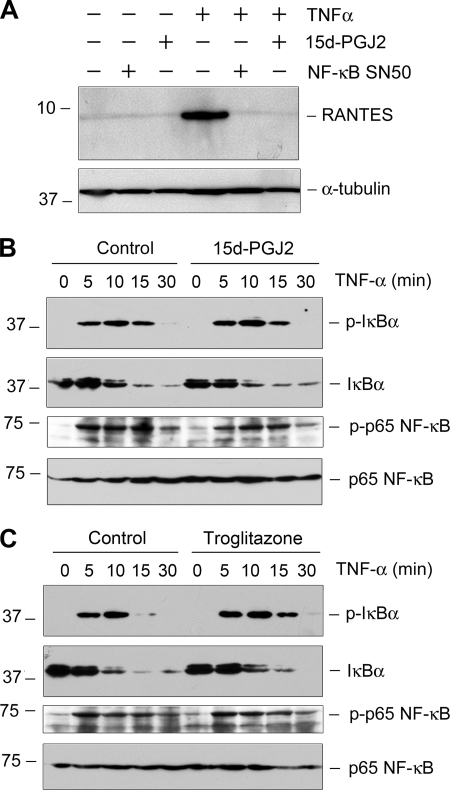
To explore the mechanism by which PPARγ activation inhibits proinflammatory cytokine expression, we first investigated the potential pathway leading to RANTES expression in mesangial cells. As shown in Fig. 3A, blockade of NF-κB signaling by a cell-permeable inhibitor peptide (NF-κB SN50) completely abolished RANTES induction by TNF-α, suggesting that NF-κB signaling is essential for mediating the TNF-α-induced RANTES expression in mesangial cells.
FIGURE 3.
PPARγ agonist displays little effect on the early events of NF-κB activation. A, TNF-α-mediated RANTES induction is dependent on NF-κB signaling. Mesangial cells were pretreated with NF-κB inhibitor SN50 (45 μm) or 15d-PGJ2 (5 μm) for 0.5 h followed by incubation with TNF-α (5 ng/ml) for 48 h. NF-κB inhibitor SN50 completely blocked TNF-α-mediated RANTES induction. B and C, both 15d-PGJ2 and troglitazone had little effect on the early events of NF-κB activation (IκBα phosphorylation, its degradation, and p65 NF-κB phosphorylation) induced by TNF-α. Mesangial cells were pretreated with either 15d-PGJ2 (5 μm) or troglitazone (5 μm) for 0.5 h, followed by incubation with TNF-α (5 ng/ml) for different periods of time as indicated. Cell lysates were immunoblotted with specific antibodies against phospho-IκBα, total IκBα, phospho-p65, or total p65, respectively.
We next investigated whether PPARγ activation leads to inhibition of NF-κB signaling. To this end, we examined the effects of PPARγ agonists on the early events of NF-κB activation, including IκBα phosphorylation and its subsequent degradation, as well as p65 NF-κB phosphorylation. As shown in Fig. 3B, IκBα was phosphorylated and underwent rapid degradation at 5–15 min after TNF-α treatment; and at the same time, p65 NF-κB was phosphorylated and activated. However, preincubation with 15d-PGJ2 did not significantly affect the TNF-α-induced IκBα phosphorylation, its degradation, and p65 phosphorylation (Fig. 3B). Similarly, synthetic PPARγ agonist troglitazone also failed to disrupt IκBα degradation and p65 activation (Fig. 3C). These results suggest that PPARγ activation by its agonists does not affect the early events of NF-κB signaling, such as IκBα phosphorylation, its subsequent degradation, and p65 phosphorylation.
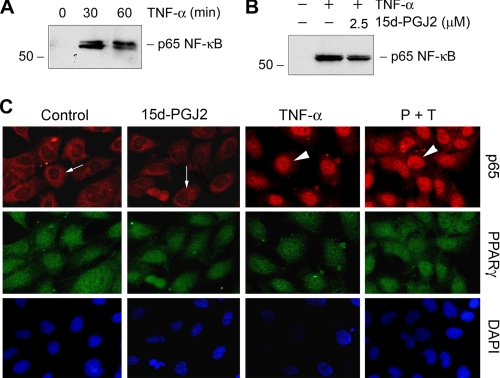
As one key event during NF-κB signaling, p65 is liberated after IκBα degradation and translocated into the nucleus. We further examined whether PPARγ activation affects this episode of NF-κB signaling. Cellular fractionation revealed that TNF-α induced a marked accumulation of p65 in the nuclei of mesangial cells (Fig. 4A). Preincubation with 15d-PGJ2, however, did not significantly affect p65 nuclear translocation (Fig. 4B). Similar results were obtained when examining the cellular localization of p65 by immunofluorescence staining. p65 protein was largely localized in the cytoplasm in the basal condition (Fig. 4C, arrows), and its nuclear accumulation was evident after TNF-α stimulation (Fig. 4C, arrowheads). Once again, 15d-PGJ2 did not block p65 nuclear translocation after TNF-α treatment. Notably, PPARγ was localized primarily and constitutively in the nuclei of mesangial cells under various conditions.
FIGURE 4.
PPARγ agonist does not block p65 NF-κB nuclear translocation induced by TNF-α. A and B, rat mesangial cells were treated with TNF-α for various periods of time as indicated (A), or pretreated with 2.5 μm 15d-PGJ2 for 0.5 h, followed by incubation in the absence or presence of TNF-α for an additional 0.5 h (B). Nuclear proteins were prepared and immunoblotted for p65 NF-κB. C, immunofluorescence staining showed the inability of PPARγ agonist to block p65 nuclear translocation. Cells were pretreated with 2.5 μm 15d-PGJ2 for 0.5 h, followed by incubation in the absence or presence of TNF-α for an additional 0.5 h. TNF-α induced p65 nuclear translocation; however, 15d-PGJ2 had little effect on p65 nuclear translocation. Arrows indicate cytoplasmic localization of p65. Arrowheads denote nuclear staining of p65.
Inhibition of RANTES by PPARγ Agonist Is Not Dependent on Hepatocyte Growth Factor (HGF)/c-met Pathway
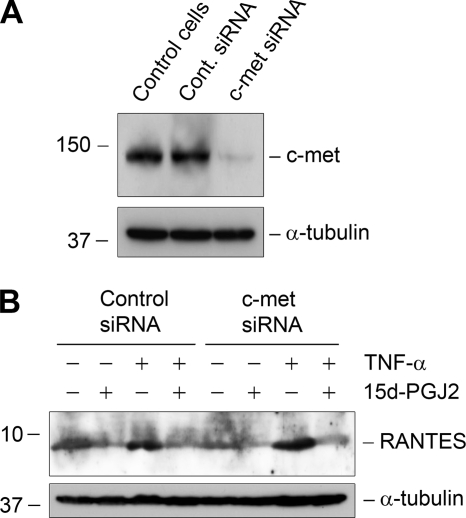
Given the inability of PPARγ agonists to inhibit the early events of NF-κB signaling, we sought to explore whether PPARγ activation leads to inhibition of inflammation by an indirect mechanism. In that regard, it has been shown that PPARγ activation induces HGF expression in mesangial cells (33) and that HGF is able to inhibit NF-κB signaling and RANTES expression (30). To test this possibility, we knocked down the expression of c-met, the sole receptor for HGF, by siRNA strategy in mesangial cells. As shown in Fig. 5A, effective knockdown of c-met expression was evident. However, knockdown of c-met did not negate the inhibition of RANTES expression by 15d-PGJ2 (Fig. 5B). Therefore, it appears that inhibition of RANTES by PPARγ agonist is independent of HGF/c-met signaling.
FIGURE 5.
Inhibition of RANTES by PPARγ agonist is independent of HGF/c-met signaling. A, knockdown of c-met receptor expression by siRNA strategy is shown. Mesangial cells were transfected with either control or c-met-specific siRNA followed by Western blot analysis for c-met expression. Mesangial cells without transfection served as controls (control cells). B, knockdown of c-met did not negate 15d-PGJ2-mediated suppression of RANTES expression. Mesangial cells were transfected with either control or c-met-specific siRNA followed by incubation with TNF-α and/or 15d-PGJ2 as indicated.
Activation of PPARγ Blocks Its Interaction with p65 NF-κB
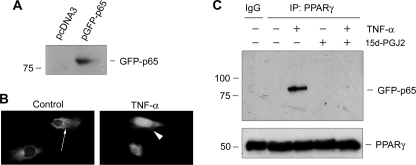
We next investigated the effect of PPARγ activation on the postnuclear events of NF-κB signaling. Because both PPARγ and p65 are localized in the nucleus after TNF-α stimulation (Fig. 4C), we sought to explore any potential, physical interaction between them. As shown in Fig. 6, A and B, GFP-tagged p65 also underwent nuclear translocation following TNF-α treatment in mesangial cells. Co-immunoprecipitation assay demonstrated that PPARγ could bind to p65 to form a physical complex after TNF-α stimulation, as p65 was detectable in the complexes precipitated by anti-PPARγ antibody (Fig. 6C). This interaction between PPARγ and p65 was apparently induced by TNF-α, as no complex formation was observed in mesangial cells under basal, unstimulated conditions. Interestingly, incubation with 15d-PGJ2 completely prevented PPARγ from binding to p65 (Fig. 6C). These results indicate that only unliganded PPARγ binds to p65, and activation of PPARγ after ligand binding prevents their interaction.
FIGURE 6.
PPARγ activation disrupts the interaction between PPARγ and p65 NF-κB upon TNF-α stimulation. A, Western blot shows ectopic expression of GFP-tagged p65 in mesangial cells. Cells were transfected with either empty vector (pcDNA3) or expression vector containing GFP-tagged p65, and cell lysates were immunoblotted with anti-GFP antibody. B, GFP fluorescence illustrates the nuclear translocation GFP-p65 NF-κB after treatment with TNF-α (5 ng/ml) for 30 min. C, co-immunoprecipitation assay demonstrates the interaction of p65 and unliganded PPARγ in the presence of TNF-α. Mesangial cells were transfected with GFP-p65 expression vector and treated with TNF-α (5 ng/ml) and/or PGJ2 (5 μm) as indicated. Cell lysates were immunoprecipitated with anti-PPARγ antibody, followed by immunoblotting with anti-GFP.
C-terminal p65 NF-κB Mediates Its Interaction with PPARγ
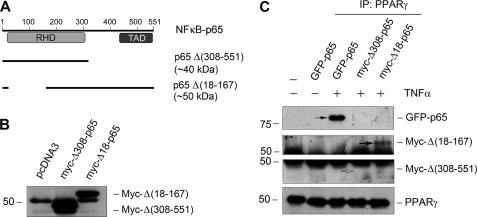
To delineate the structural domain of p65 that mediates its interaction, we employed p65 deletion mutants to examine their ability to bind to PPARγ. Fig. 7A shows the construction of GFP-tagged full-length p65 as well as Myc-tagged p65 deletion mutants. p65Δ(308–551) contained the N-terminal region of p65, but lacked the C-terminal region (308–551). p65Δ(18–167) contained the C-terminal region of p65, but a region from amino acids 18 to 167 was deleted. Transfection of mesangial cells with Δ(308–551) and Δ(18–167) truncated p65 vectors gave rise to a 46-kDa and 57-kDa fusion protein, respectively (Fig. 7B). When these constructs were transfected into mesangial cells, full-length p65 readily interacted with PPARγ after TNF-α stimulation (Fig. 7C). However, only p65Δ(18–167), but not p65 Δ(308–551), was detected in the immunocomplexes precipitated by anti-PPARγ antibody (Fig. 7C). These results indicate that the C-terminal region of p65, but not the N-terminal Rel homology domain, is essential for mediating its interaction with PPARγ.
FIGURE 7.
C-terminal p65 NF-κB mediates its interaction with PPARγ. A, diagram shows the construction of various p65 NF-κB expression vectors. Plasmids containing GFP-tagged full-length p65 as well as Myc-tagged p65 deletion mutants are illustrated. The calculated molecular masses of these mutant proteins are shown. RHD, Rel homology domain; TAD, trans-activation domain. B, mesangial cells were transfected with Δ(308–551) and Δ(18–167) truncated vectors and pcDNA3 control, followed by Western blotting for Myc. Myc-tagged Δ(308–551) (44 kDa) and Δ(18–167) (56 kDa) are indicated. The discrepancy between the predicted and actual molecular masses of the mutant proteins presumably resulted from the posttranslational modifications. C, mesangial cells were transfected to express GFP-tagged full-length p65 as well as Myc-tagged truncated p65 mutants Δ(308–551) and Δ(18–167). After treatment with TNF-α (5 ng/ml), cell lysates were immunoprecipitated (IP) with anti-PPARγ, followed by immunoblotting with anti-GFP and anti-Myc, respectively. Interaction of PPARγ with full-length p65 and Δ(18–167) (solid arrows), but not Δ(308–551) (open arrow), was evident.
PPARγ Activation Blocks p65 Binding to Its Cognate DNA Elements
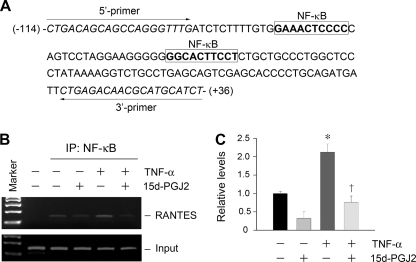
NF-κB regulates gene transcription primarily through binding to its cis-acting κB element. To explore whether PPARγ activation affect p65 binding to its cognate DNA element, we utilized an in situ ChIP assay. As shown in Fig. 8A, there were two putative NF-κB binding sites in the proximal promoter region of rat RANTES gene (31). Stimulation of mesangial cells with TNF-α increased p65 binding to κB elements in rat RANTES promoter, as demonstrated in Fig. 8, B and C. However, activation of PPARγ by 15d-PGJ2 prevented p65 binding to κB elements induced by TNF-α (Fig. 8, B and C). Because 15d-PGJ2 abolished the interaction between p65 and PPARγ (Fig. 6C), these findings imply that the recruitment of PPARγ is likely a prerequisite for p65 to bind to its cognate DNA elements. Notably, 15d-PGJ2 had the tendency to inhibit p65 binding to κB elements in the absence of TNF-α, suggesting that PPARγ activation also inhibits the basal endogenous p65/κB element interaction in the unstimulated conditions.
FIGURE 8.
PPARγ activation blocks p65 NF-κB binding to its cognate elements in rat RANTES promoter. A, partial sequence of rat RANTES gene promoter region. Boxed sequences are the putative κB sites in this region. B and C, ChIP assay revealed that TNF-α induced p65 binding to the RANTES promoter, which was blocked by PPARγ agonist 15d-PGJ2. Representative ChIP assay (B) and quantitative ChIP data (C) are presented. *, p < 0.05 versus controls; †, p < 0.05 versus TNF-α alone (n = 3). Error bars, S.E.
Unliganded PPARγ Is Essential for Mediating RANTES Expression and NF-κB Signaling
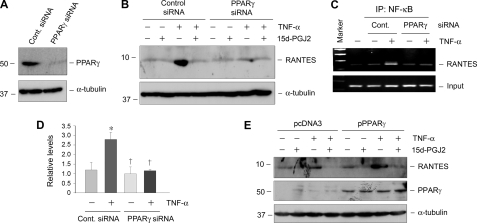
To examine the importance of p65/PPARγ interaction in NF-κB signaling, we assessed the effect of PPARγ depletion on TNF-α-mediated RANTES expression. Mesangial cells were transfected with either control or PPARγ-specific siRNA, followed by incubation with TNF-α or/and 15d-PGJ2 as indicated. Substantial knockdown of PPARγ was confirmed by Western blot analysis (Fig. 9A). Interestingly, knockdown of PPARγ completely abolished RANTES induction by TNF-α (Fig. 9B), suggesting that unliganded PPARγ is essential for mediating TNF-α-induced RANTES expression.
FIGURE 9.
Unliganded PPARγ is required for mediating TNF-α-induced RANTES expression in mesangial cells. A and B, knockdown of PPARγ abolishes RANTES induction by TNF-α. Mesangial cells were transfected with either control or PPARγ-specific siRNA, followed by incubation with TNF-α (5 ng/ml) or/and PGJ2 (5 μm) as indicated. Knockdown of PPARγ was confirmed by Western blot analysis (A), and knockdown of PPARγ completely abolished RANTES induction by TNF-α (B). C and D, the binding of p65 to RANTES promoter upon TNF-α stimulation was dependent on the presence of unliganded PPARγ. Representative ChIP assay (C) and quantitative ChIP data (D) are presented. *, p < 0.05 (n = 3). Error bars, S.E. E, overexpression of PPARγ in mesangial cells sensitized to TNF-α stimulation. Mesangial cells were transfected with PPARγ expression vector (pPPARγ) or empty vector (pcDNA3) followed by incubation with TNF-α (5 ng/ml) and/or 15d-PGJ2 (5 μm) as indicated. Cell lysates were immunoblotted with specific antibodies against RANTES, PPARγ, and α-tubulin, respectively.
We also investigated the effect of PPARγ depletion on p65 binding to its cognate κB element. As shown in Fig. 9, C and D, TNF-α induced p65 binding to its DNA element; however, PPARγ depletion by siRNA strategy completely prevented the TNF-α-induced binding of p65 to the κB sites in RANTES promoter, suggesting that the binding of p65 to RANTES promoter upon TNF-α stimulation is also dependent on the presence of unliganded PPARγ.
Using an opposite strategy, we found that overexpression of PPARγ in the absence of its ligand sensitized mesangial cells to TNF-α stimulation. As shown in Fig. 9E, more robust induction of RANTES was observed in the mesangial cells tranfected with exogenous PPARγ expression vector, compared with that with pcDNA3 empty vector controls. Taken together, although activation of PPARγ inhibits NF-κB signaling, unliganded PPARγ is obligatory for the TNF-α-mediated NF-κB signaling and RANTES induction in mesangial cells.
DISCUSSION
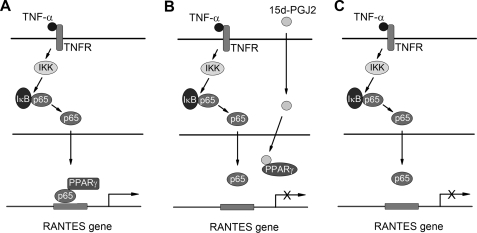
PPARγ activation by its agonists is known to lead to inhibition of proinflammatory cytokine expression (1, 5, 22); however, the underlying mechanism remains ambiguous. In the present study, we demonstrate that activation of PPARγ by its agonists prevents PPARγ·p65 complex formation, which is required for proper NF-κB signaling in mesangial cells. As summarized in Fig. 10, in the absence of PPARγ agonist, TNF-α triggers the phosphorylation and subsequent degradation of IκBα and induces p65 phosphorylation and nuclear translocation. Nuclear p65 recruits unliganded PPARγ to form a complex and then binds to the cognate κB sites in the RANTES promoter and directs its gene transcription (Fig. 10A). However, the binding of PPARγ agonists prevents or/and disrupts the interaction between p65 and PPARγ, presumably by triggering a conformational change of PPARγ. This disables p65 to bind to the RANTES promoter, thereby blocking RANTES expression (Fig. 10B). Interestingly, it appears that unliganded PPARγ is absolutely obligatory for p65 binding to the κB sites of RANTES promoter and for RANTES induction, as knockdown of PPARγ completely prevents TNF-α-mediated RANTES induction in mesangial cells (Fig. 10C). These studies provide significant insights into understanding the molecular mechanism by which PPARγ agonists inhibits inflammatory responses. Our studies also uncover an opposite effect of unliganded and ligand-activated PPARγ in regulating NF-κB signaling and proinflammatory cytokine expression.
FIGURE 10.
Schematic model indicates the opposite effects of unliganded and ligand-activated PPARγ on RANTES expression. A, in the absence of PPARγ agonist, TNF-α triggers the phosphorylation and degradation of IκB and induces p65 phosphorylation and nuclear translocation. Nuclear p65 recruits unliganded PPARγ to form a complex and then binds to the cognate sequence in the RANTES promoter and directs gene transcription. B, in the presence of PPARγ agonists, the binding of agonist leads to conformational changes of PPARγ, which prevents or/and disrupts the interaction of p65 and PPARγ, upon TNF-α stimulation. This disables p65 to bind to the RANTES promoter, thereby blocking RANTES expression. C, unliganded PPARγ is obligatory for p65 binding to RANTES promoter and for RANTES induction. Knockdown of PPARγ completely prevents TNF-α-mediated RANTES expression in mesangial cells.
The results presented in the current study show that both natural (15d-PGJ2) and synthetic agonists (ciglitazone and troglitazone) inhibit RANTES and MCP-1 expression in rat mesangial cells (Figs. 1 and 2). Although not tested, it would not be surprising that other thiazolidinediones such as rosglitazone and pioglitazone might also be able to inhibit the expression of these chemokines. In that regard, earlier studies have shown that rosglitazone and pioglitazone repress RANTES expression in human endometrial stromal cells and human monocyte-derived dendritic cells (34, 35), respectively. Several previous studies have examined the mechanism behind the antiinflammatory action of PPARγ agonists in different contexts by focusing on the early events of NF-κB signaling, such as IκBα expression, phosphorylation, and its subsequent degradation, as well as p65 nuclear translocation (21, 36–38). For example, PPARγ activation has been shown to induce IκBα mRNA and protein expression in mouse cardiac tissues (21) and reduces p65 expression in human peripheral monocytes (37). It is also revealed that PPARγ activation by its agonists suppresses IκB kinase activity in pancreatic acinar AR42J cells (36), whereas it inhibits IL-1β and IFNγ-stimulated p65 nuclear translocation and its DNA binding activity in rat pancreatic β-cells (38). Therefore, it appears that PPARγ activation can influence distinct events in the processes of NF-κB signaling in a cell context-specific fashion. In mesangial cells, however, PPARγ agonists apparently fail to affect the early events of NF-κB signaling, such as IκBα phosphorylation and its subsequent degradation, as well as p65 phosphorylation and its nuclear translocation (Figs. 3 and 4). We have also ruled out the possibility that PPARγ agonists may inhibit renal inflammation by an indirect mechanism through inducing HGF (33), a well characterized cytokine with potent antiinflammatory potential (30, 39), as depletion of HGF receptor (c-met) does not abolish the inhibitory effect of PPARγ agonists on RANTES expression (Fig. 5). Taken together, it is becoming clear that PPARγ agonists most likely inhibit inflammatory responses in mesangial cells by targeting the postnuclear events during NF-κB signaling.
It is generally accepted that upon stimulation, p65 is liberated from the sequestration by IκBα in the cytoplasm and enters the nuclei, wherein it binds to the κB sites of its target genes. What is less understood, however, is the molecular details by which activated p65 binds to the cis-acting κB element. Interestingly, we have uncovered in this study that, prior to binding to κB element, p65 actually recruits PPARγ by forming a physical complex through its C-terminal region that harbors the trans-activation domain, and such a recruitment of PPARγ could be a prerequisite for a functional NF-κB signaling. This notion is supported by several lines of evidence. First, activated p65 forms a complex with PPARγ in mesangial cells after TNF-α stimulation (Fig. 6C). Second, activation of PPARγ by its agonists prevents PPARγ·p65 complex formation, which is associated with a disruptive NF-κB signaling and inhibition of RANTES induction. Third, depletion of PPARγ prevents p65 from binding to the κB sites and abolishes RANTES expression after TNF-α stimulation in mesangial cells (Fig. 9). In addition, overexpression of unliganded PPARγ sensitizes mesangial cells to TNF-α stimulation (Fig. 9E). Therefore, the p65/PPARγ interaction is absolutely obligatory for proper function of NF-κB signaling. These results unravel a previously unrecognized, physiologically important step in NF-κB signaling, in which PPARγ facilitates p65 binding to the κB elements by forming a PPARγ·p65 complex.
Our model, as presented in Fig. 10, illustrates that despite the importance of PPARγ in mediating NF-κB signaling, its activation by agonists paradoxically inhibits NF-κB signaling and proinflammatory cytokine expression in mesangial cells. This dramatic shift of PPARγ function from pro- to anti-NF-κB signaling is clearly dictated by its ligand, as PPARγ activation by its agonists prevents its interaction with p65 in mesangial cells. The exact reason behind the inability of ligand-bound PPARγ to interact with p65 remains to be elucidated, but it could be related to any perceived conformational change of PPARγ triggered by ligand binding. This notion is consistent with earlier observations that the ligand binding domain of PPARγ is subjected to ligand-dependent sumoylation (40–42). Whether this ligand-dependent sumoylation of PPARγ plays a role in preventing p65/PPARγ interaction in mesangial cells remains to be determined.
In summary, we have shown in this study that activation of PPARγ by its agonists in mesangial cells inhibits the stimulus-dependent induction of RANTES and MCP-1, two important chemokines that are instrumental in recruiting the inflammatory cells such as T cells and monocytes/macrophages in diseased kidney (43, 44). Our results also unravel a novel mechanism by which PPARγ agonists block NF-κB signaling through preventing p65/PPARγ interaction. These studies provide an explanation for how its ligand acts as a molecular switch that controls the ability of PPARγ to modulate NF-κB signaling and inflammatory responses in opposite directions.
Acknowledgments
We thank Drs. Johannes A. Schmid and Mark S. Nanes for providing GFP-p65 and truncated p65 mutant expression vectors.
This work was supported, in whole or in part, by National Institutes of Health Grants DK061408, DK064005, and DK071040.
- PPARγ
- peroxisome proliferator-activated receptor-γ
- 15d-PGJ2
- 15-deoxy-Δ12,14-prostaglandin J2
- HGF
- hepatocyte growth factor
- MCP-1
- monocyte chemoattractant protein-1
- NF-κB
- nuclear factor-κB
- RANTES
- regulated upon activation, normal T cell expressed and secreted.
REFERENCES
- 1.Fogo A. B. (2008) Curr. Opin. Nephrol. Hypertens. 17, 282–285 [DOI] [PubMed] [Google Scholar]
- 2.Guan Y., Breyer M. D. (2001) Kidney Int. 60, 14–30 [DOI] [PubMed] [Google Scholar]
- 3.Ruan X., Zheng F., Guan Y. (2008) Am. J. Physiol. Renal Physiol. 294, F1032–F1047 [DOI] [PubMed] [Google Scholar]
- 4.Lehmann J. M., Moore L. B., Smith-Oliver T. A., Wilkison W. O., Willson T. M., Kliewer S. A. (1995) J. Biol. Chem. 270, 12953–12956 [DOI] [PubMed] [Google Scholar]
- 5.Wu J., Chen L., Zhang D., Huo M., Zhang X., Pu D., Guan Y. (2009) Front. Biosci. 14, 995–1009 [DOI] [PubMed] [Google Scholar]
- 6.Ricote M., Li A. C., Willson T. M., Kelly C. J., Glass C. K. (1998) Nature 391, 79–82 [DOI] [PubMed] [Google Scholar]
- 7.Odegaard J. I., Ricardo-Gonzalez R. R., Goforth M. H., Morel C. R., Subramanian V., Mukundan L., Red Eagle A., Vats D., Brombacher F., Ferrante A. W., Chawla A. (2007) Nature 447, 1116–1120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hevener A. L., Olefsky J. M., Reichart D., Nguyen M. T., Bandyopadyhay G., Leung H. Y., Watt M. J., Benner C., Febbraio M. A., Nguyen A. K., Folian B., Subramaniam S., Gonzalez F. J., Glass C. K., Ricote M. (2007) J. Clin. Invest. 117, 1658–1669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sharma A. M., Staels B. (2007) J. Clin. Endocrinol. Metab. 92, 386–395 [DOI] [PubMed] [Google Scholar]
- 10.Yang H. C., Deleuze S., Zuo Y., Potthoff S. A., Ma L. J., Fogo A. B. (2009) J. Am. Soc. Nephrol. 20, 2380–2388 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Toblli J. E., Ferrini M. G., Cao G., Vernet D., Angerosa M., Gonzalez-Cadavid N. F. (2009) Nephrol. Dial. Transplant. 24, 2384–2391 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kawai T., Masaki T., Doi S., Arakawa T., Yokoyama Y., Doi T., Kohno N., Yorioka N. (2009) Lab. Invest. 89, 47–58 [DOI] [PubMed] [Google Scholar]
- 13.Calkin A. C., Giunti S., Jandeleit-Dahm K. A., Allen T. J., Cooper M. E., Thomas M. C. (2006) Nephrol. Dial. Transplant. 21, 2399–2405 [DOI] [PubMed] [Google Scholar]
- 14.Ma L. J., Marcantoni C., Linton M. F., Fazio S., Fogo A. B. (2001) Kidney Int. 59, 1899–1910 [DOI] [PubMed] [Google Scholar]
- 15.Haraguchi K., Shimura H., Onaya T. (2003) Clin. Exp. Nephrol. 7, 27–32 [DOI] [PubMed] [Google Scholar]
- 16.Ghosh S. S., Massey H. D., Krieg R., Fazelbhoy Z. A., Ghosh S., Sica D. A., Fakhry I., Gehr T. W. (2009) Am. J. Physiol. Renal Physiol. 296, F1146–F1157 [DOI] [PubMed] [Google Scholar]
- 17.Liu Y. (2010) J. Am. Soc. Nephrol. 21, 212–222 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hayden M. S., Ghosh S. (2008) Cell 132, 344–362 [DOI] [PubMed] [Google Scholar]
- 19.Rangan G., Wang Y., Harris D. (2009) Front. Biosci. 14, 3496–3522 [DOI] [PubMed] [Google Scholar]
- 20.Tan X., Wen X., Liu Y. (2008) J. Am. Soc. Nephrol. 19, 1741–1752 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Buroker N. E., Barboza J., Huang J. Y. (2009) FEBS J. 276, 3247–3255 [DOI] [PubMed] [Google Scholar]
- 22.Vanden Berghe W., Vermeulen L., Delerive P., De Bosscher K., Staels B., Haegeman G. (2003) Adv. Exp. Med. Biol. 544, 181–196 [DOI] [PubMed] [Google Scholar]
- 23.Wen X., Li Y., Hu K., Dai C., Liu Y. (2005) Am. J. Pathol. 167, 947–957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dai C., Liu Y. (2004) J. Am. Soc. Nephrol. 15, 1402–1412 [DOI] [PubMed] [Google Scholar]
- 25.Yang J., Liu Y. (2001) Am. J. Pathol. 159, 1465–1475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Yang J., Dai C., Liu Y. (2003) Am. J. Pathol. 163, 621–632 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Schmid J. A., Birbach A., Hofer-Warbinek R., Pengg M., Burner U., Furtmüller P. G., Binder B. R., de Martin R. (2000) J. Biol. Chem. 275, 17035–17042 [DOI] [PubMed] [Google Scholar]
- 28.Lu X., Farmer P., Rubin J., Nanes M. S. (2004) J. Cell. Biochem. 92, 833–848 [DOI] [PubMed] [Google Scholar]
- 29.Jiang J. G., Johnson C., Zarnegar R. (2001) J. Biol. Chem. 276, 25049–25056 [DOI] [PubMed] [Google Scholar]
- 30.Giannopoulou M., Dai C., Tan X., Wen X., Michalopoulos G. K., Liu Y. (2008) Am. J. Pathol. 173, 30–41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pocock J., Gómez-Guerrero C., Harendza S., Ayoub M., Hernández-Vargas P., Zahner G., Stahl R. A., Thaiss F. (2003) J. Immunol. 170, 6280–6291 [DOI] [PubMed] [Google Scholar]
- 32.Haberstroh U., Pocock J., Gómez-Guerrero C., Helmchen U., Hamann A., Gutierrez-Ramos J. C., Stahl R. A., Thaiss F. (2002) Kidney Int. 62, 1264–1276 [DOI] [PubMed] [Google Scholar]
- 33.Li Y., Wen X., Spataro B. C., Hu K., Dai C., Liu Y. (2006) J. Am. Soc. Nephrol. 17, 54–65 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pritts E. A., Zhao D., Ricke E., Waite L., Taylor R. N. (2002) J. Clin Endocrinol. Metab. 87, 1841–1844 [DOI] [PubMed] [Google Scholar]
- 35.Gosset P., Charbonnier A. S., Delerive P., Fontaine J., Staels B., Pestel J., Tonnel A. B., Trottein F. (2001) Eur. J. Immunol. 31, 2857–2865 [DOI] [PubMed] [Google Scholar]
- 36.Wan H., Yuan Y., Qian A., Sun Y., Qiao M. (2008) Biomed. Pharmacother. 62, 466–472 [DOI] [PubMed] [Google Scholar]
- 37.Hounoki H., Sugiyama E., Mohamed S. G., Shinoda K., Taki H., Abdel-Aziz H. O., Maruyama M., Kobayashi M., Miyahara T. (2008) Bone 42, 765–774 [DOI] [PubMed] [Google Scholar]
- 38.Kim E. K., Kwon K. B., Koo B. S., Han M. J., Song M. Y., Song E. K., Han M. K., Park J. W., Ryu D. G., Park B. H. (2007) Int. J. Biochem. Cell Biol. 39, 1260–1275 [DOI] [PubMed] [Google Scholar]
- 39.Gong R., Rifai A., Ge Y., Chen S., Dworkin L. D. (2008) J. Biol. Chem. 283, 7401–7410 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Jennewein C., Kuhn A. M., Schmidt M. V., Meilladec-Jullig V., von Knethen A., Gonzalez F. J., Brüne B. (2008) J. Immunol. 181, 5646–5652 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zelcer N., Tontonoz P. (2005) Cell Metab. 2, 273–275 [DOI] [PubMed] [Google Scholar]
- 42.Pascual G., Fong A. L., Ogawa S., Gamliel A., Li A. C., Perissi V., Rose D. W., Willson T. M., Rosenfeld M. G., Glass C. K. (2005) Nature 437, 759–763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ruster C., Wolf G. (2008) Front. Biosci. 13, 944–955 [DOI] [PubMed] [Google Scholar]
- 44.Krensky A. M., Ahn Y. T. (2007) Nat. Clin. Pract. Nephrol. 3, 164–170 [DOI] [PMC free article] [PubMed] [Google Scholar]