Abstract
The NF-κB transcription factors control many physiological processes, including inflammation, immunity, and apoptosis. Its activity contributes to the development of various cell malignancies. NF-κB-inducing kinase (NIK) plays a pivotal role in NF-κB activation. However, the molecular mechanism to stabilize and activate NIK remains elusive, although it is known that cIAP1/2 (cellular inhibitor of apoptosis 1 and 2) ubiquitinate NIK for degradation. Here, we report a novel NF-κB-related zinc finger protein 91 (ZFP91) that stabilizes and activates NIK in a ubiquitination-dependent manner. We show that ZFP91 interacts with and promotes the Lys63-linked ubiquitination of NIK and subsequent processing of p100 to p52. The results of in vitro biochemical assays indicate that ZFP91 functions as an E3 ligase directly to NIK. Remarkably, the ubiquitination of NIK coincides with its Thr559 phosphorylation. Furthermore, knockdown of ZFP91 expression by RNA interference inhibits the CD40 ligation-induced activation of NIK and p100 processing as well as the expression of noncanonical NF-κB target genes. These data clearly indicate that ZFP91 is an important regulator of the noncanonical NF-κB pathway.
Keywords: E3 Ubiquitin Ligase, NF-κB Transcription Factor, RNA Interference (RNAi), TRAF, Ubiquitylation, CD40, Lys63-linked Ubiquitination, NIK, ZFP91
Introduction
The transcription factor NF-κB plays a central role in immune responses, development, and cell proliferation (1). It also links inflammation and immune responses to the development and progression of cancer (2). There are two major pathways leading to NF-κB activation. The canonical pathway requires IκB kinase (IKK)2 complex, consisting of IKKα/β/γ, whose activation results in the phosphorylation and ubiquitin-dependent degradation of IκB proteins and the nuclear translocation of p50-containing NF-κB complex (1). The noncanonical pathway requires NF-κB-inducing kinase (NIK), which cooperates with IKKα to induce NF-κB2 (p100) processing to p52, thereby resulting in translocation of the p52-RelB complex to the nucleus (1, 3). NIK is activated in a Thr559 phosphorylation-dependent manner (4) and then activates IKKα, which in turn phosphorylates p100 for proteasome-dependent processing to p52 (5). Although NIK mRNA is relatively abundant, NIK protein is undetectable in most cell types of unstimulated cells (6, 7). Tumor necrosis factor (TNF) receptor superfamily ligand-induced activation of the noncanonical NF-κB signaling pathway appears to prevent basally translated NIK proteins from undergoing proteasome-mediated degradation (6–8).
Protein ubiquitination is an important post-translational modification that regulates various biological functions (9). Although ubiquitination often results in protein degradation, a certain type of ubiquitination is important for signaling activation (10). Ubiquitination through Lys48 of the ubiquitin chain generally targets proteins for degradation, whereas ubiquitination through Lys63 plays a critical role in cellular signaling, DNA repair, protein localization, endocytosis, and protein kinase activation (11–15).
In this study, a cDNA microarray analysis with a diterpenoid kamebakaurin, which specifically inhibits DNA binding by the p50 subunit of NF-κB complex (16), revealed a novel gene, ZFP91 (zinc finger protein 91), which has consecutive zinc finger (ZnF) domains and is known to be highly expressed in most human acute myelogenous leukemia cases (17). We here demonstrate that ZFP91 is an atypical E3 ligase activating NIK via Lys63-linked ubiquitination in the noncanonical NF-κB signaling pathway.
EXPERIMENTAL PROCEDURES
Cell Culture, Transfection, and Luciferase Reporter Assay
HEK293 and MDA-MB231 cells were grown in Dulbecco's modified Eagle's medium with penicillin (100 units/ml) plus streptomycin (100 units/ml) (Invitrogen) and 10% heat-inactivated fetal bovine serum (Hyclone). Ramos and Jurkat cells were maintained in RPMI medium supplemented as above. Transfections were performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions. NF-κB-dependent luciferase activity was measured using the dual luciferase reporter assay system.
Plasmids and Reagents
The complete coding region of human ZFP91 cDNA was amplified from a human gastric cancer cell line SNU-638 cDNA library by PCR. We generated pFLAG-ZFP91 and pEntr-ZFP91 by PCR subcloning of ZFP91 into pCMV-Tag2B (Stratagene) and pEntr-BHRNX (Newgex, Seoul, Korea), respectively. ZFP91 and NIK deletion mutants were generated by standard PCR methods. ZFP91 point mutants were generated using a QuikChange kit (Stratagene) as directed by the manufacturer. Mammalian expression constructs were obtained as follows: Myc-NIK, Myc-NIK (KK429/430AA) (M. Jung, Georgetown University) and FLAG-cIAP1 (cellular inhibitor of apoptosis 1) (M. Naito, Tokyo University), HA-Ub, HA-UbK48R, HA-UbK63R, HA-Ub Lys48-only, and HA-Ub Lys63-only (C. H. Chung, Seoul National University). FLAG-TRAF2 (TNF receptor-associated factor 2) and Myc-NIK (T559A) were developed in our laboratory. Antibody reagents were purchased from the indicated vendors: anti-FLAG (catalog no. F3165, Sigma) and FLAG affinity gel (catalog no. A2220, Sigma), anti-α-tubulin (catalog no. T5168, Sigma), anti-HA (catalog no. 12CA5, Roche Applied Science), anti-Myc (catalog no. 9E10, Roche Applied Science), anti-NIK (catalog no. sc-7211 (Santa Cruz Biotechnology, Inc., Santa Cruz, CA) or 4994 (Cell Signaling)), anti-CD40 (catalog no. sc-975, Santa Cruz Biotechnology, Inc.), anti-phosphorylated NIK (catalog no. sc-12957, Santa Cruz Biotechnology, Inc.), anti-Ub (catalog no. sc-8017 (Santa Cruz Biotechnology, Inc.) or MMS-264R (Covance)), polyubiquitin Lys63 linkage-specific monoclonal antibody (clone HWA4C4, BIOMOL), anti-p52 (catalog no. 4882, Cell Signaling), anti-phosphorylated IKKα/β (catalog no. 2681, Cell Signaling), anti-IκB-α (catalog no. 9242, Cell Signaling), and anti-GST (catalog no. sc-138, Santa Cruz Biotechnology, Inc.). Recombinant human CD40 ligand CD154 and GST-NIK (aa 381–947) were purchased from R&D Systems and BPS Bioscience, respectively. Ubiquitin and Lys48-only and Lys63-only ubiquitin were obtained from Boston Biochem. We generated the anti-ZFP91 polyclonal antibody by immunizing mice with full-length ZFP91, which was purified by a Ni2+-NTA chelating agarose column (Peptron, Daejeon, Korea) from lysates of Sf21 insect cells transfected with recombinant baculovirus expressing full-length ZFP91.
Immunoprecipitation and Immunoblotting
Cell lysates in lysis buffer (50 mm Tris, pH 7.4, 150 mm NaCl, 1 mm EDTA, 1% Triton X-100, and protease inhibitor mixture) were centrifuged at 15,000 rpm for 30 min at 4 °C, and 1 mg of protein of cleared lysates was used for each immunoprecipitation. The lysates were incubated overnight at 4 °C with primary antibodies with gentle rotation, and then 30 μl of protein A/G PLUS-agarose beads (Santa Cruz Biotechnology, Inc.) were added to the mixtures and rotated for an additional 1 h at 4 °C. Beads were washed three times with cold lysis buffer. The proteins were recovered by boiling in SDS-PAGE sample buffer. The eluted proteins were separated on SDS-PAGE and transferred to a PVDF membrane (Amersham Biosciences). To detect specific ubiquitination, a two-step immunoprecipitation was performed. After the first immunoprecipitation as described, pellets were washed three times with lysis buffer and incubated with 1% SDS/PBS for 1 h to disrupt non-covalent interactions. Samples were centrifuged, and supernatants were diluted with lysis buffer, followed by a second immunoprecipitation. The resultant precipitates were incubated in sample buffer and separated on SDS-PAGE as described.
Preparation of Recombinant ZFP91
To obtain baculovirus-expressed ZFP91 pEntr-ZFP91 and baculovirus, genomic DNA were incubated with LR clonase (Invitrogen) to obtain a recombinant baculovirus. These LR clonase reaction mixtures were transfected into Sf21 cells using Lipofectin (Invitrogen). Baculovirus-expressed ZFP91 was purified from Sf21 insect cells by a Ni2+-NTA chelating agarose column (Peptron).
In Vitro Binding Assay
ZFP91 and NIK purified from Sf21 cells were incubated with 1 ml of 1 mg/ml bovine serum albumin (BSA) buffer (1 mg/ml BSA, 0.25% Nonidet P-40, 2 mm EDTA, 0.2 mm PMSF, 1 mm DTT in phosphate-buffered saline) for 2–4 h at 4 °C. The resulting mixture was then immunoprecipitated with anti-NIK antibody and protein A/G PLUS-agarose beads, and the resultant precipitates were incubated in sample buffer and separated on SDS-PAGE as described.
Northern Blot
RNA was isolated from cells using RNeasy minikits according to the manufacturer's instructions (Qiagen). Ten μg of total RNA were resolved on 1% agarose-formaldehyde gel and transferred to a nylon membrane. Membranes were probed and washed according to the instructions of the manufacturer (Roche Applied Science). 32P-Labeled probes were generated by the random priming method using Rediprime II (Amersham Biosciences) and 50 μCi of [α-32P]dCTP (Amersham Biosciences). Unincorporated nucleotides were removed by purification through a G-25 spin column. The results were visualized by autoradiography.
Real-time PCR
RNA was isolated from cells using RNeasy minikits according to the manufacturer's instructions (Qiagen). Complementary DNA was synthesized from 1 μg of total RNA in a 20-μl reverse transcription reaction mixture according to the manufacturer's protocol (TaKaRa Bio, Kyoto, Japan). The following primer pairs were used for real-time PCR amplification: human interleukin (IL)-6, 5′-GAACTCCTTCTCCACAAGCGCCTT-3′ and 5′-CAAAAGACCAGTGATGATTTTCACCAGG-3′; IL-8, 5′-TCTGCAGCTCTGTGTGAAGG-3′ and 5′-ACTTCTCCACAACCCTCTG-3′; human MCP1 (monocyte chemotactic protein 1), 5′-CCCCAGTCACCTGCTGTTAT-3′ and 5′-AGATCTCCTTGGCCACAATG-3′; human TNF-α, 5′-CTGCCCCAATCCCTTTATT-3′ and 5′-CCCAATTCTCTTTTTGAGCC-3′; human BAFF (B cell activation factor), 5′-AATATCACTGGATGGAGATGT-3′ and 5′-GTTAGATTCTTTCTTCTTAGAGGTA-3′; human CCL19 (EBI-1-ligand chemokine), 5′-GCCTGCTGGTTCTCTGGAC-3′ and 5′-GGATGGGTTTCTGGGTCAC-3′. The quantitative real-time PCR was carried out using power SYBR Green (Bio-Rad). Reactions were performed in triplicate according to the manufacturer's protocol.
In Vitro Ubiquitination Assay
In vitro ubiquitination assays were performed in 50-μl reaction volumes containing 1 μg of ubiquitin, Ub Lys48-only, or Ub Lys63-only (Boston Biochem), 50 nm E1 (Biomol), 1.0 μg of E2 (Biomol), 100 ng of E3 ZFP91, and 5 μl of 10× reaction buffer (500 mm Tris, pH 7.5, 20 mm ATP, 50 mm MgCl2, 2 mm DTT). Reactions were incubated at 37 °C for 1 h and subsequently prepared for immunoblot analysis. For in vitro NIK ubiquitination by ZFP91, reactions were performed as described above, except adding 1 μg of recombinant NIK. Reactions were performed in 200-μl reaction volumes, and after incubation, 50 μl was reserved for immunoblotting, and the remaining 150 μl was diluted to a final volume of 1 ml with washing buffer (50 mm Tris, pH 7.5, 150 mm NaCl, pH 7.4, and protease inhibitor mixture), and then 3 μg of anti-NIK antibody was added. Samples were rotated for 12 h at 4 °C, and then 50 μl of protein A/G Plus-agarose beads (Santa Cruz Biotechnology, Inc.) were added. After 1 h of additional rotation, immunoprecipitates were washed three times with the washing buffer, and then samples were prepared for immunoblot analysis.
RNA Interference
RNA interference was carried out using specific siRNA to silence ZFP91 in HEK293 or MDA-MB231 cells. ZFP91-specific double-stranded siRNA oligonucleotides were obtained from Qiagen Inc. The siRNA duplexes prepared in diethyl pyrocarbonate-treated water at a 20 μm concentration were transfected in subconfluent cells with Lipofectamine RNAiMAX (Invitrogen) following the manufacturer's instructions and then incubated for 24–48 h before analysis.
RESULTS
ZFP91 Activates Noncanonical NF-κB Signaling
To identify a novel NF-κB-related gene, a cDNA microarray analysis was performed using total RNA from MDA-MB231 cells treated with or without 10 μg/ml kamebakaurin for 5 h. Among 50 candidates, which were down-regulated more than 2-fold after kamebakaurin treatment, TNFAIP3 and ZFP91 were confirmed by Northern blot analyses (Fig. 1A). TNFAIP3, also called A20, is a well known regulator of NF-κB signaling (18), showing that the microarray analysis was successful.
FIGURE 1.
ZFP91 activates the noncanonical NF-κB signaling pathway. A, MDA-MB231 cells were treated with kamebakaurin (KA) (10 μg/ml) for 5 h, and the extracted mRNAs were subject to Northern blot analysis using 28 S RNA as a control. B, 24 h after the transfection with titration of FLAG-ZFP91 and NF-κB reporter construct pNF-κB-Luc, the lysates of HEK293 cells were subject to the measurement of dual luciferase activity. Data represented as mean ± S.D. of three independent experiments. C, HEK293 cells transfected with titration of FLAG-ZFP91 were lysed, and the lysates were analyzed by immunoblot using the indicated antibodies for NIK, p52, p50, FLAG, and tubulin. D, HEK293 cells were co-transfected with FLAG-ZFP91 and Myc-NIK as indicated. The cell lysates were analyzed by immunoblot using the indicated antibodies for p52, p-IKKβ/α, IκB-α, Myc, FLAG, and tubulin.
To gain insight into ZFP91 function, we tested whether ZFP91 affects NF-κB signaling. Intriguingly, ZFP91 induced NF-κB-dependent reporter gene expression dose-dependently (Fig. 1B). Furthermore, ZFP91 increased the NIK level and p100 processing to p52 dose-dependently (Fig. 1C, top two panels), whereas it did not affect the expression level of p50 (Fig. 1C, third panel), suggesting that ZFP91 may be associated with the noncanonical NF-κB pathway.
To address the effect of ZFP91 on the noncanonical NF-κB signaling, we investigated whether ZFP91 regulates NIK-induced p100 processing to p52 and IKKα phosphorylation. Co-expression of ZFP91 and NIK increased NIK level (Fig. 1D, fourth panel) and further increased downstream p52 and IKKα phosphorylation levels (Fig. 1D, top two panels). However, the IκBα level was not changed significantly by expression of ZFP91 or NIK (Fig. 1D, third panel), indicating that ZFP91 may act in conjunction with NIK.
ZFP91 Interacts with NIK
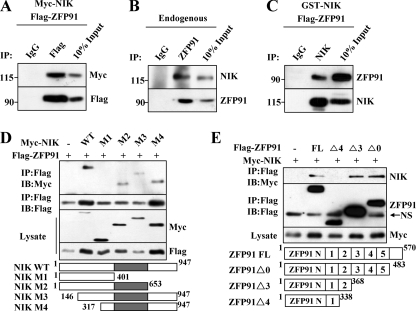
Next, we examined whether ZFP91 interacts with NIK. Co-immunoprecipitation assays revealed that ZFP91 physically interacted with NIK upon co-expression of ZFP91 and NIK (Fig. 2A), and endogenous interaction was observed under the treatment of proteasome inhibitor MG-132, which prevents NIK degradation (Fig. 2B). This interaction was further confirmed by an in vitro binding assay (Fig. 2C). To characterize the region of these two proteins responsible for their interaction, we constructed various truncation mutants of ZFP91 and NIK. We showed that this interaction was mediated by the kinase domain of NIK (aa 401–653) and the second ZnF domain of ZFP91 (aa 338–368) (Fig. 2, D and E).
FIGURE 2.
ZFP91 interacts with NIK. A, the lysates of HEK293 cells co-transfected with FLAG-ZFP91 and Myc-NIK were incubated with IgG and anti-FLAG antibody, and the immune complexes were analyzed by immunoblots using appropriate antibodies. B, the lysates of HEK293 cells treated with proteasomal inhibitor MG132 (10 μm) for the final 4 h were incubated with IgG and anti-ZFP91 antibody, and the immune complexes were subject to immunoblots using the indicated antibodies. C, FLAG-ZFP91 and GST-NIK purified from Sf21 cells were incubated in 0.1% BSA binding buffer. The associated complexes were incubated with IgG and anti-NIK antibody and then analyzed by immunoblots using appropriate antibodies. D and E, lysates of HEK293 cells co-transfected with the indicated FLAG-ZFP91 and Myc-NIK constructs were incubated with anti-FLAG antibody, and the immune complexes were analyzed by immunoblots (top two panels). FLAG and Myc immunoblots were completed from the same cell lysates (bottom panels). NS, nonspecific bands. IP, immunoprecipitation; IB, immunoblot.
ZFP91 Stabilizes, Activates, and Ubiquitinates NIK
Because NF-κB signaling has been known to be tightly regulated by ubiquitin in both positive and negative manners (19), we tested whether ubiquitin affects the functional regulation of NIK by ZFP91. Surprisingly, co-expression of NIK with ubiquitin or ZFP91 activated NF-κB-dependent reporter gene expression, and this effect was further potentiated by both (Fig. 3A). Furthermore, co-expression of ZFP91 and ubiquitin resulted in a strong increase in endogenous level of NIK and the processing of p100 to p52 as compared with those under the expression of ZFP91 or ubiquitin alone (Fig. 3B), indicating that ZFP91 stabilizes NIK and further activates it in a ubiquitin-dependent manner.
FIGURE 3.
ZFP91 stabilizes NIK and induces Lys63-linked NIK ubiquitination. A, HEK293 cells were co-transfected with pNF-κB-Luc and Myc-NIK in combination with HA-Ub or FLAG-ZFP91 or together with HA-Ub and FLAG-ZFP91. Dual luciferase activity was measured 24 h after transfection. Data are represented as mean ± S.D. of three independent experiments. B, the lysates from HEK293 cells co-transfected with HA-Ub, FLAG-ZFP91, or both were subject to endogenous NIK and p52 immunoblots (top two panels). FLAG and tubulin immunoblots were also completed from the same cell lysates (bottom two panels). C, the lysates of HEK293 cells co-transfected with Myc-NIK, FLAG-ZFP91, and HA-Ub constructs as indicated were subject to immunoprecipitation (IP) using anti-Myc antibody, and the immune complexes were analyzed by immunoblots (IB; top two panels). FLAG, Myc, and tubulin immunoblot was completed from the same cell lysates (bottom panel). D, the lysates of HEK293 cells co-transfected with Myc-NIK and HA-UbK48R along with a control siRNA, ZFP91 siRNA, or FLAG-ZFP91 were incubated with anti-Myc antibody, and the immune complexes were analyzed by immunoblots (top two panels). The immunoblots using anti-ZFP91, anti-Myc, and anti-tubulin antibodies were completed from the same cell lysates (bottom two panels). E, the lysates of HEK293 cells co-transfected with Myc-NIK, FLAG-ZFP91, and HA-Ub constructs as indicated were subject to immunoprecipitation using anti-Myc antibody, a portion of the precipitated pellets were saved for immunoblots, and the remainder of the precipitated pellets were treated with 1% SDS, PBS to disrupt non-covalent interactions, and supernatants were diluted with lysis buffer, followed by a second immunoprecipitation using anti-Myc antibody. The immune complexes were analyzed by immunoblots (top two panels). FLAG immunoblot was completed from the same cell lysates.
The ZFP91- and ubiquitin-dependent activation of NIK prompted us to investigate whether ZFP91 is related with NIK ubiquitination. Indeed, ZFP91 induced NIK ubiquitination with the ubiquitin K48R mutant, whereas the ubiquitin K63R mutant was not efficiently ligated to NIK (Fig. 3C). This result was further confirmed by the data indicating that ZFP91 siRNA reduced ubiquitin K48R-linked NIK (Fig. 3D). To determine whether co-eluting protein complexes were involved in the NIK ubiquitination, a two-step, denaturing protocol was used to immunoprecipitate Myc-NIK. Ubiquitination of NIK observed in both the first and second immunoprecipitation appeared to be comparable except intensity of ubiquitination (Fig. 3E).
ZnF Domain 2 of ZFP91 Is Critical for Lys63-linked Ubiquitination of NIK
To identify the domain(s) responsible for mediating the ubiquitination, we generated various truncation mutations of ZFP91 and analyzed their activities in HEK293 cells. Ubiquitin-conjugated NIK was readily detectable when NIK was co-expressed with full-length ZFP91 or ZFP91-C (aa 242–570) but was not detectable with ZFP91-N (aa 1–242) (Fig. 4A). These results suggest that the ZnF domains of ZFP91 are required for NIK ubiquitination. Further deletion analyses demonstrated that any mutants retaining ZnF2 strongly ubiquitinated NIK (Fig. 4B), indicating that ZnF2 is critical for NIK ubiquitination. This conclusion was confirmed with point mutants of conserved cysteines in ZnF1, -2, -3, -4, or -5. When the ZnF2 (C344A/C349A) mutant of ZFP91 was expressed with NIK, NIK ubiquitination was almost lost (Fig. 4C, lane 4). All of the results consistently support the idea that ZFP91 mediates the Lys63-linked ubiquitination of NIK.
FIGURE 4.
ZnF domain 2 of ZFP91 is critical for Lys63-linked ubiquitination of NIK. A, the lysates of HEK293 cells co-transfected with Myc-NIK and HA-Ub along with FLAG-ZFP91FL (aa 1–570), FLAG-ZFP91N (aa 1–242), or FLAG-ZFP91C (aa 242–570) were incubated with anti-Myc antibody, and the immune complexes were analyzed by immunoblots (top two panels). A FLAG, Myc, and tubulin immunoblot was completed from the same cell lysates (bottom panel). NS, nonspecific bands. B, the lysates of HEK293 cells co-transfected with Myc-NIK and HA-Ub along with various FLAG-ZFP91 constructs as indicated were incubated with anti-Myc antibody, and the immune complexes were analyzed by immunoblots (top two panels). A FLAG, Myc, and tubulin immunoblot was completed from the same cell lysates (bottom panel). C, the lysates of HEK293 cells co-transfected with Myc-NIK and HA-Ub along with FLAG-ZFP91 or point mutants of conserved cysteines in ZnF1 MT (C313A/C318A), ZnF2 MT (C344A/C349A), ZnF3 MT (C374A/C377A), ZnF4 MT (C402A/C405A), or ZnF5 MT (C432A/C435A) were incubated with anti-Myc antibody, and the immune complexes were analyzed by immunoblots (top two panels). A FLAG, Myc, and tubulin immunoblot was completed from the same cell lysates (bottom). D, the lysates of HEK293 cells co-transfected with Myc-NIK and HA-Ub48K (treated with proteasomal inhibitor MG132 for the final 4 h) or HA-Ub63K alone or together with FLAG-tagged TRAF2, cIAP1, and ZFP91, as indicated, were incubated with anti-Myc antibody, and the immune complexes were analyzed by immunoblots (top two panels). A FLAG, Myc, and tubulin immunoblot was completed from the same cell lysates (bottom). IP, immunoprecipitation; IB, immunoblot.
Recent studies suggest that NIK is suppressed through constitutive proteasome-mediated degradation regulated by cIAP1/2 that were activated by TRAF2 in a ubiquitination-dependent manner (8, 20–22). Therefore, we examined whether cIAP1 and TRAF2 affect ZFP91-mediated Lys63-linked ubiquitination of NIK. Consistent with the previous reports (21, 22), TRAF2 and cIAP1 promoted Lys48-linked ubiquitination of NIK. ZFP91, however, did not induce ubiquitination of NIK with Lys48-only ubiquitin (Fig. 4D, lane 4) and did not alter the ubiquitination of NIK induced by cIAP1 and TRAF2 (Fig. 4D, lane 3). On the other hand, ZFP91 promoted Lys63-linked ubiquitination of NIK (Fig. 4D, lane 8), whereas cIAP1 and TRAF2 did not (Fig. 4D, lane 6). Furthermore, ZFP91-mediated Lys63-linked ubiquitination of NIK was not affected by cIAP1 and TRAF2 (Fig. 4D, lane 7), indicating that the Lys63-linked ubiquitination of NIK is independent of cIAP1 and TRAF2.
ZFP91 Is an E3 Ligase for Lys63-linked Ubiquitination of NIK
The fact that ZFP91 induces Lys63-linked ubiquitination of NIK in vivo raised the possibility that ZFP91 has an intrinsic E3 ligase activity because the ZnF domain of A20/TNFAIP3, another NF-κB pathway regulator, has been known as an atypical E3 ligase (18). Thus, we investigated in vitro E3 ligase activity of ZFP91. ZFP91 purified from insect cells was subject to in vitro ubiquitination assays. Remarkably, despite the absence of a known E3 ligase domain, such as the RING (really interesting new gene) finger or HECT (homologous to the E6-associated protein C terminus) motif (23, 24), ZFP91 catalyzed in vitro autoubiquitination when a specific ubiquitin-conjugating enzyme, UbcH13/Mms2, and all reaction components were included (Fig. 5, A and B). Moreover, we showed that ZFP91 catalyzed Lys63-linked ubiquitination preferentially over Lys48-linked ubiquitination (Fig. 5C), as predicted from the in vivo assay (Figs. 3C and 4D). Furthermore, we showed that ZFP91 directly ubiquitinated NIK with Lys63-only ubiquitin (Fig. 5D). These results demonstrate that ZFP91 is a novel E3 ligase that conjugates Lys63-linked ubiquitin chains to NIK.
FIGURE 5.
ZFP91 is an E3 ligase for Lys63-linked ubiquitination of NIK. A, in vitro ubiquitination assays were performed in reaction mixtures containing purified ZFP91, ubiquitin, E1 enzyme, and ATP in the presence of a panel of E2 ubiquitin-conjugating enzymes. Reactions were incubated at 37 °C for 1 h, and the reaction mixtures were detected by anti-HA immunoblot (upper panel). The amount of ZFP91 proteins used in the assay is shown in a FLAG immunoblot (lower panel). B, in vitro ubiquitination assays were performed in reaction mixtures containing purified ZFP91, ubiquitin, E1 enzyme, and ATP in the presence of an E2 ubiquitin-conjugating enzyme, UbcH13/Mms2. Reactions were incubated at 37 °C for 1 h, and the ubiquitinated ZFP91 was detected by an anti-HA and anti-FLAG immunoblot. C, in vitro ubiquitination assays were performed as described above using purified ZFP91 with Lys48-only or Lys63-only ubiquitin, and the ubiquitinated ZFP91 was detected by anti-Ub immunoblot (upper panel). The amount of ZFP91 proteins used in the assay is shown in the FLAG immunoblot (lower panel). D, in vitro ubiquitination assays were performed using ZFP91 and GST-NIK (aa 381–947) as described above and under “Experimental Procedures.” The reaction mixtures were immunoprecipitated with anti-GST antibody and analyzed by immunoblots (top two panels). Anti-Ub and -FLAG immunoblots were completed using the same samples (bottom two panels). IB, immunoblot; IP, immunoprecipitation.
ZFP91 Mediates the Lys63-linked Ubiquitination and Phosphorylation of NIK in CD40 Signaling
Next, we investigated the relationship of the Lys63-linked ubiquitination and phosphorylation of NIK in the processing of p100 to p52. Interestingly, ZFP91-induced ubiquitination was mostly abrogated in NIK mutants K429A/K430A (KK429/430AA) and T559A (Fig. 6A). Furthermore, Lys63-linked ubiquitination of NIK by ZFP91 coincided with Thr559 phosphorylation of NIK, enhanced processing of p100 to p52, and activation of NF-κB reporter gene (Fig. 6, A and B), suggesting the strong correlation between Lys63-linked ubiquitination of NIK by ZFP91 and its kinase function.
FIGURE 6.
ZFP91 mediates the Lys63-linked ubiquitination and phosphorylation of NIK in CD40 signaling. A, the lysates of HEK293 cells co-transfected with FLAG-ZFP91 and HA-Ub together with Myc-NIK wild type, Myc-NIK K429A/K430A (KK429/430AA), or Myc-NIK T559A were incubated with anti-Myc antibody. Immunoblot analyses for immunoprecipitated proteins were completed by using anti-HA and Myc antibodies (top two panels) from the immune complexes and anti-phospho-NIK, p52, FLAG, Myc, and tubulin antibodies (bottom four panels) from the same cell lysates. B, lysates from HEK293 cells co-transfected with NF-κB luciferase reporter construct pNF-κB-Luc, Myc-NIK, and FLAG-ZFP91 along with HA-Ub, HA-Ub Lys48-only (HA-UbK48only), or HA-Ub Lys63-only (HA-UbK63only), as indicated, were analyzed by luciferase activity and immunoblot. C, MDA-MB231 cells transfected with a control or ZFP91 siRNA stimulated with CD154 for the indicated time. Cell lysates were subject to immunoblots using anti-p52, NIK, ZFP91, and tubulin antibodies. D, MDA-MB231 cells co-transfected with Myc-NIK and a control or ZFP91 siRNA stimulated with CD154 for 6 h. Cell lysates were immunoprecipitated with anti-Myc antibody, and the immune complexes were analyzed with an antibody specific for Lys63-linked ubiquitin. The immunoblots (IB) using anti-ZFP91, anti-Myc, and anti-tubulin antibodies were completed from the same cell lysates. E, MDA-MB231 cells transfected with a control or ZFP91 siRNA stimulated with CD154 for the indicated time. RNAs were isolated from cells, reverse-transcribed, and analyzed by real-time PCR for IL-6, IL-8, MCP1, TNF-α, BAFF, and CCL19. Data are represented as mean ± S.D. (error bars) of three independent experiments. IB, immunoblot; IP, immunoprecipitation.
Because NIK is activated by TNF receptor superfamily ligands, such as CD154, BAFF, and TWEAK (7, 8, 25), we next examined whether ZFP91 mediates CD40 signaling to NIK in MDA-MB231 cells, which are known to highly express CD40 (26, 27) (supplemental Fig. 1, A and B). Consistent with the previous reports, CD154 led NIK to stabilization and activated subsequent p100 processing to p52 (Fig. 6C, left four lanes). siRNA-mediated knockdown of ZFP91, however, almost completely blocked these processes (Fig. 6C, right four lanes). Furthermore, CD40 ligation indeed elicited Lys63-linked NIK ubiquitination in vivo (Fig. 6D, lane 2, and supplemental Fig. 2, lane 2), whereas ZFP91 siRNA expression blocked the CD154-induced ubiquitination of NIK (Fig. 6D, lane 4, and supplemental Fig. 2, lane 4). These data clearly demonstrate that ZFP91 is essential for the Lys63-linked ubiquitination, stabilization, and activation of NIK in CD40 signaling.
Because ligation of CD40 is well established to activate NF-κB (28), we tested whether ZFP91 regulates the transcription of NF-κB target genes upon CD154 stimulation. Consistent with the previous reports (29, 30), IL-6, IL-8, MCP1, and TNF-α mRNA levels were rapidly increased after CD154 stimulation and decreased to basal level over time. siRNA-mediated knockdown of ZFP91, however, did not alter the IL-6, IL-8, MCP1, and TNF-α mRNA levels induced by CD154 (Fig. 6E). On the other hand, the expression of the BAFF and CCL19 mRNA levels increased persistently over time upon CD154 stimulation, whereas ZFP91 siRNA expression blocked the CD154-induced expression of BAFF and CCL19 mRNA levels (Fig. 6E). These results demonstrate that ZFP91 is an important regulator of the noncanonical NF-κB pathway via CD40.
DISCUSSION
Although mounting evidence suggests that Lys63-linked ubiquitination plays a key role in the activation of protein kinases in multiple pathways leading to NF-κB activation (14, 15), it has not been known whether Lys63-linked ubiquitination of NIK is important for NIK activation. This study may provide an answer to the question of how noncanonical NF-κB signaling is activated via Lys63-linked ubiquitination of NIK. We demonstrated for the first time the molecular function of a novel E3 ligase ZFP91: the stabilization and activation of NIK via conjugation of Lys63-linked ubiquitin chains. It is of note that, despite the absence of a known E3 ligase domain, ZFP91 efficiently catalyzed non-degradative Lys63-linked ubiquitination with a specific ubiquitin-conjugating enzyme, Ubc13.
The E3 ligases are critical components that are needed to determine the enzymatic specificity in the ubiquitin cascade as a result of direct interaction with substrates (31). E3 ligases can generally be classified into two families: RING-type E3 ligases and HECT-type E3 ligases (23, 24). Additionally, the A20-type ZnF represents a novel class of E3 ligases. A20 is a protein that contains an N-terminal ovarian tumor domain and novel ZnF domains in the C-terminal portion of the protein, which mediate the transfer of ubiquitin from E2 enzyme to the substrate (18). Comparable with A20 composed of seven consecutive ZnF domains, the ZFP91 is also a ZnF protein having consecutive five ZnF domains. We obtained substantive evidence that ZFP91 functions as an E3 ligase of NIK. The evidence is noted as follows: 1) ZFP91 and NIK form a stable complex, 2) ZFP91 catalyzes NIK ubiquitination in vivo and in vitro, and 3) the enzymatic activity is dependent on active site cysteines in the ZnF domain as demonstrated in the A20 (18). Therefore, NIK might be a physiological substrate for the ZFP91 E3 ligase.
Our findings suggest that Ubc13 is required for the activation of noncanonical NF-κB by ZFP91-mediated Lys63 ubiquitination of NIK. It has previously been shown that Ubc13 is dispensable for IL-1- and TNF-α-mediated NF-κB activation in Ubc13-deficient MEF cells, and processing of p100 to p52 was normal in Ubc13-deficient B cells stimulated with CD154 and BAFF (32). On the other hand, Ubc13 is essential for Tax-linked Lys63 ubiquitination and subsequent activation of the noncanonical NF-κB pathway in Ubc13-deficient MEF cells (33). Thus, the mechanisms used by ZFP91 to activate noncanonical NF-κB are similar with the Tax. In ZFP91-mediated noncanonical NF-κB activation, the role of Ubc13 is to facilitate Lys63-linked ubiquitination of ZFP91 (Fig. 5, A–C), which in turn mediates the ubiquitination of NIK (Fig. 5D) and activates the noncanonical NF-κB pathway.
NIK was originally identified as a mitogen-activated protein kinase kinase kinase interacting with TRAF2 (34). Genetic analysis of both aly/aly mice, which harbor a NIK-inactivating mutation, and Nik−/− mice revealed that NIK is mainly involved in activation of the noncanonical NF-κB signaling pathway (35, 36). Activation of this pathway through engagement of CD40, BAFF receptor, or TWEAK receptor results in NIK stabilization, leading to accumulation of NIK (6–8, 25). NIK stabilization and continuous activation of the noncanonical pathway was regulated by TRAF2, TRAF3, and cIAP1/2 (6, 21, 22, 25, 37, 38). It is well documented that TRAF2 and cIAP1/2 form an E3 ligase complex, and importantly, in this complex, cIAP1/2 act as Lys48-specific E3 ligases in the negative regulation of NIK and noncanonical NF-κB activation (21, 25). These results could raise the question of whether ZFP91 regulates cIAP1/2 and TRAF2 E3 ligase activity to influence Lys48-linked ubiquitination of NIK. We showed, however, that ZFP91 did not inhibit TRAF2 and cIAP1 E3 ligase activity (Fig. 4D, lanes 2 and 3). In addition, the fact that TRAF2 and cIAP1/2 catalyze Lys63-linked ubiquitination (21, 39, 40) could also raise the possibility that the TRAF2 and cIAP1/2 ubiquitin ligase complex may be responsible for the Lys63-linked ubiquitination of NIK. However, TRAF2 and cIAP1 did not induce Lys63-linked ubiquitination of NIK (Fig. 4D, lane 6) and did not alter the Lys63-linked ubiquitination of NIK by ZFP91 (Fig. 4D, lanes 7 and 8). These results indicated that NIK is regulated not only in its stabilization via dissociation of the NIK-TRAF3-TRAF2-cIAP1/2 complex but also in its activation via Lys63-linked ubiquitination.
This study also uncovered that Lys63-specific ubiquitination is linked to the phosphorylation at Thr559 in the activation loop of NIK. The association of ubiquitination with the phosphorylation of protein kinase is not unprecedented in protein kinase activation. For example, the activation of MEKK1 coincides with ubiquitination and phosphorylation (41). In addition, TAK1 is activated by unanchored Lys63-specific ubiquitin chains (42). It has also been suggested that Lys63-linked ubiquitination of IL-1β receptor-associated kinase 1 is critical for IKK activation (43). Our results also suggest that the dominant negative function of NIK mutants K429A/K430A and T559A (4, 34) could be at least in part associated with their incapability to be ubiquitinated by ZFP91. Further studies are required to elucidate the molecular relationship between Lys63-linked ubiquitination and activation of NIK.
Collectively, our current studies explore the mechanism by which ZFP91 induces noncanonical NF-κB activation through the ZFP91-mediated Lys63-ubiquitination of NIK. These studies also suggest that ZFP91 may have a role in the pathophysiology of the NIK-driven immune system in a variety of diseases.
Supplementary Material
Acknowledgments
We are grateful to many researchers, who provided valuable research materials, listed under “Experimental Procedures.” Our thanks extend to Drs. H. G. Lee and H. S. Son for help with the production of anti-ZFP91 antibody and ZFP91 protein in insect cells, respectively.
This work was supported by a grant from the 21C Frontier Genome Project from the Ministry of Education and Science and Technology, Republic of Korea.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. 1 and 2.
- IKK
- IκB kinase
- NIK
- NF-κB-inducing kinase
- ZnF
- zinc finger
- aa
- amino acids
- Ub
- ubiquitin.
REFERENCES
- 1.Vallabhapurapu S., Karin M. (2009) Annu. Rev. Immunol. 27, 693–733 [DOI] [PubMed] [Google Scholar]
- 2.Karin M., Greten F. R. (2005) Nat. Rev. Immunol. 5, 749–759 [DOI] [PubMed] [Google Scholar]
- 3.Senftleben U., Cao Y., Xiao G., Greten F. R., Krähn G., Bonizzi G., Chen Y., Hu Y., Fong A., Sun S. C., Karin M. (2001) Science 293, 1495–1499 [DOI] [PubMed] [Google Scholar]
- 4.Lin X., Mu Y., Cunningham E. T., Jr., Marcu K. B., Geleziunas R., Greene W. C. (1998) Mol. Cell. Biol. 18, 5899–5907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Xiao G., Rabson A. B., Young W., Qing G., Qu Z. (2006) Cytokine Growth Factor Rev. 17, 281–293 [DOI] [PubMed] [Google Scholar]
- 6.Liao G., Zhang M., Harhaj E. W., Sun S. C. (2004) J. Biol. Chem. 279, 26243–26250 [DOI] [PubMed] [Google Scholar]
- 7.Qing G., Qu Z., Xiao G. (2005) J. Biol. Chem. 280, 40578–40582 [DOI] [PubMed] [Google Scholar]
- 8.Varfolomeev E., Blankenship J. W., Wayson S. M., Fedorova A. V., Kayagaki N., Garg P., Zobel K., Dynek J. N., Elliott L. O., Wallweber H. J., Flygare J. A., Fairbrother W. J., Deshayes K., Dixit V. M., Vucic D. (2007) Cell 131, 669–681 [DOI] [PubMed] [Google Scholar]
- 9.Mukhopadhyay D., Riezman H. (2007) Science 315, 201–205 [DOI] [PubMed] [Google Scholar]
- 10.Pickart C. M. (2001) Annu. Rev. Biochem. 70, 503–533 [DOI] [PubMed] [Google Scholar]
- 11.Habelhah H., Takahashi S., Cho S. G., Kadoya T., Watanabe T., Ronai Z. (2004) EMBO J. 23, 322–332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Clague M. J., Urbé S. (2006) Trends Cell Biol. 16, 551–559 [DOI] [PubMed] [Google Scholar]
- 13.Wang C., Deng L., Hong M., Akkaraju G. R., Inoue J., Chen Z. J. (2001) Nature 412, 346–351 [DOI] [PubMed] [Google Scholar]
- 14.Chen Z. J. (2005) Nat. Cell Biol. 7, 758–765 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Krappmann D., Scheidereit C. (2005) EMBO Rep. 6, 321–326 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lee J. H., Koo T. H., Hwang B. Y., Lee J. J. (2002) J. Biol. Chem. 277, 18411–18420 [DOI] [PubMed] [Google Scholar]
- 17.Unoki M., Okutsu J., Nakamura Y. (2003) Int. J. Oncol. 22, 1217–1223 [PubMed] [Google Scholar]
- 18.Wertz I. E., O'Rourke K. M., Zhou H., Eby M., Aravind L., Seshagiri S., Wu P., Wiesmann C., Baker R., Boone D. L., Ma A., Koonin E. V., Dixit V. M. (2004) Nature 430, 694–699 [DOI] [PubMed] [Google Scholar]
- 19.Skaug B., Jiang X., Chen Z. J. (2009) Annu. Rev. Biochem. 78, 769–796 [DOI] [PubMed] [Google Scholar]
- 20.Vince J. E., Wong W. W., Khan N., Feltham R., Chau D., Ahmed A. U., Benetatos C. A., Chunduru S. K., Condon S. M., McKinlay M., Brink R., Leverkus M., Tergaonkar V., Schneider P., Callus B. A., Koentgen F., Vaux D. L., Silke J. (2007) Cell 131, 682–693 [DOI] [PubMed] [Google Scholar]
- 21.Vallabhapurapu S., Matsuzawa A., Zhang W., Tseng P. H., Keats J. J., Wang H., Vignali D. A., Bergsagel P. L., Karin M. (2008) Nat. Immunol. 9, 1364–1370 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zarnegar B. J., Wang Y., Mahoney D. J., Dempsey P. W., Cheung H. H., He J., Shiba T., Yang X., Yeh W. C., Mak T. W., Korneluk R. G., Cheng G. (2008) Nat. Immunol. 9, 1371–1378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Deshaies R. J., Joazeiro C. A. (2009) Annu. Rev. Biochem. 78, 399–434 [DOI] [PubMed] [Google Scholar]
- 24.Bernassola F., Karin M., Ciechanover A., Melino G. (2008) Cancer Cell 14, 10–21 [DOI] [PubMed] [Google Scholar]
- 25.Vince J. E., Chau D., Callus B., Wong W. W., Hawkins C. J., Schneider P., McKinlay M., Benetatos C. A., Condon S. M., Chunduru S. K., Yeoh G., Brink R., Vaux D. L., Silke J. (2008) J. Cell Biol. 182, 171–184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gomes E. M., Rodrigues M. S., Phadke A. P., Butcher L. D., Starling C., Chen S., Chang D., Hernandez-Alcoceba R., Newman J. T., Stone M. J., Tong A. W. (2009) Clin. Cancer Res. 15, 1317–1325 [DOI] [PubMed] [Google Scholar]
- 27.Hirano A., Longo D. L., Taub D. D., Ferris D. K., Young L. S., Eliopoulos A. G., Agathanggelou A., Cullen N., Macartney J., Fanslow W. C., Murphy W. J. (1999) Blood 93, 2999–3007 [PubMed] [Google Scholar]
- 28.Lapchak P. H., Melter M., Pal S., Flaxenburg J. A., Geehan C., Frank M. H., Mukhopadhyay D., Briscoe D. M. (2004) Am. J. Physiol. Renal Physiol. 287, F512–F520 [DOI] [PubMed] [Google Scholar]
- 29.Ait-Ghezala G., Mathura V. S., Laporte V., Quadros A., Paris D., Patel N., Volmar C. H., Kolippakkam D., Crawford F., Mullan M. (2005) Brain Res. Mol. Brain Res. 140, 73–85 [DOI] [PubMed] [Google Scholar]
- 30.Poggi M., Jager J., Paulmyer-Lacroix O., Peiretti F., Gremeaux T., Verdier M., Grino M., Stepanian A., Msika S., Burcelin R., de Prost D., Tanti J. F., Alessi M. C. (2009) Diabetologia 52, 1152–1163 [DOI] [PubMed] [Google Scholar]
- 31.Weissman A. M. (2001) Nat. Rev. Mol. Cell Biol. 2, 169–178 [DOI] [PubMed] [Google Scholar]
- 32.Yamamoto M., Okamoto T., Takeda K., Sato S., Sanjo H., Uematsu S., Saitoh T., Yamamoto N., Sakurai H., Ishii K. J., Yamaoka S., Kawai T., Matsuura Y., Takeuchi O., Akira S. (2006) Nat. Immunol. 7, 962–970 [DOI] [PubMed] [Google Scholar]
- 33.Shembade N., Harhaj N. S., Yamamoto M., Akira S., Harhaj E. W. (2007) J. Virol. 81, 13735–13742 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Malinin N. L., Boldin M. P., Kovalenko A. V., Wallach D. (1997) Nature 385, 540–544 [DOI] [PubMed] [Google Scholar]
- 35.Yin L., Wu L., Wesche H., Arthur C. D., White J. M., Goeddel D. V., Schreiber R. D. (2001) Science 291, 2162–2165 [DOI] [PubMed] [Google Scholar]
- 36.Xiao G., Harhaj E. W., Sun S. C. (2001) Mol. Cell 7, 401–409 [DOI] [PubMed] [Google Scholar]
- 37.He J. Q., Zarnegar B., Oganesyan G., Saha S. K., Yamazaki S., Doyle S. E., Dempsey P. W., Cheng G. (2006) J. Exp. Med. 203, 2413–2418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gardam S., Sierro F., Basten A., Mackay F., Brink R. (2008) Immunity 28, 391–401 [DOI] [PubMed] [Google Scholar]
- 39.Bertrand M. J., Milutinovic S., Dickson K. M., Ho W. C., Boudreault A., Durkin J., Gillard J. W., Jaquith J. B., Morris S. J., Barker P. A. (2008) Mol. Cell 30, 689–700 [DOI] [PubMed] [Google Scholar]
- 40.Lee T. H., Shank J., Cusson N., Kelliher M. A. (2004) J. Biol. Chem. 279, 33185–33191 [DOI] [PubMed] [Google Scholar]
- 41.Gallagher E., Enzler T., Matsuzawa A., Anzelon-Mills A., Otero D., Holzer R., Janssen E., Gao M., Karin M. (2007) Nat. Immunol. 8, 57–63 [DOI] [PubMed] [Google Scholar]
- 42.Xia Z. P., Sun L., Chen X., Pineda G., Jiang X., Adhikari A., Zeng W., Chen Z. J. (2009) Nature 461, 114–119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Windheim M., Stafford M., Peggie M., Cohen P. (2008) Mol. Cell. Biol. 28, 1783–1791 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.