Abstract
Sensitive, high-throughput methods to detect malaria parasites in low-transmission settings are needed. PCR-based pooling strategies may offer a solution. We first used laboratory-prepared samples to compare 2 DNA extraction and 4 PCR detection methods across a range of pool sizes and parasite densities. Pooled Chelex extraction of DNA, followed by nested PCR of cytochrome b, was the optimal strategy, allowing reliable detection of a single low-parasitemic sample (100 parasites/μl) in pool sizes up to 50. This PCR-based pooling strategy was then compared with microscopy using 891 dried blood spots from a cohort of 77 Ugandan children followed for 2 years in an urban setting of low endemicity. Among 419 febrile episodes, 35 cases of malaria were detected using the PCR-based pooling strategy and 40 cases using microscopy. All five cases of malaria not detected by PCR were from samples stored for >2 years with parasitemia of <6,000/μl, highlighting the issue of possible DNA degradation with long-term storage of samples. Among 472 samples collected from asymptomatic children as part of routine surveillance, 15 (3.2%) were positive by PCR-based pooling compared to 4 (0.8%) by microscopy (P = 0.01). Thus, this PCR-based pooling strategy for detection of malaria parasites using dried blood spots offers a sensitive and efficient approach for malaria surveillance in low-transmission settings, enabling improved detection of asymptomatic submicroscopic infections and dramatic savings in labor and costs.
Due to large-scale implementation of effective control measures, many countries where malaria is endemic are experiencing dramatic declines in disease burden. With this success has come a shift in the end goal from control to elimination (9, 10). However, when the goal is elimination, accurate detection of persons infected with malaria parasites becomes essential (11). Standard surveillance systems depend on diagnosis by microscopy, a method that is technically challenging, labor-intensive, and often inaccurate in operational settings. More recently available rapid diagnostic tests (RDTs) provide convenience and ease of use, but they have limitations in specificity, sensitivity, species identification, and cost (22).
PCR-based methods for malaria parasite detection are relatively simple and provide improved sensitivity compared to microscopy and RDTs, especially in settings where infections have low parasitemia or contain mixed species (22). PCR can also be performed on dried blood spots, which are convenient for collection, storage, and transport. Nonetheless, PCR has a long turnaround time, making it an impractical tool for clinical care of individual patients. One exception would be real-time PCR, which has a short turnaround time but carries with it cost and capacity constraints that make it unavailable for rapid diagnosis in most settings. However, there is a potential role for PCR in situations where large numbers of samples are being screened: a high-throughput system could allow accurate, rapid, and cost-effective assessments to quantify prevalence and identify species.
The need for a high-throughput method to detect malaria parasites is especially important for surveillance in low-transmission and elimination settings, where low prevalence renders diagnostic testing at the individual sample level impractical and inefficient. Detection methods based on pooled nucleic acid could provide a simple solution. The pooling of sera or plasma specimens has long been used to improve the accuracy, cost-effectiveness, and throughput of screening for many infections (e.g., HIV-1, hepatitis B virus, hepatitis C virus, and West Nile virus) in low-prevalence settings, such as blood banks (4, 26). We recently reported methods for pooled analysis of dried blood spots for HIV-1 nucleic acids (1, 18).
The goal of this study was to develop and apply a novel PCR-based pooling strategy for the detection of malaria parasites using dried blood spots. We first established reference standards by using laboratory strains to create dried blood spots with known Plasmodium falciparum parasite densities. After extracting DNA from pools of dried blood spots, we compared the sensitivities and specificities of different nucleic acid amplification tests. The most robust pooling strategy was identified and then applied to 891 field samples collected using passive and active surveillance from a cohort of 77 children living in Kampala, Uganda—a city with relatively low malaria endemicity—who were followed for up to 2 years. The results were compared to those obtained by microscopy.
MATERIALS AND METHODS
Preparation of control dried blood spots with known parasite densities.
P. falciparum strain W2 parasites were cultured and synchronized at the ring stage using standard methods (14). Parasitemia was determined using flow cytometry. Positive controls were prepared by mixing infected red blood cells with uninfected whole blood to create parasite densities of 10 and 100 parasites/μl. Negative controls were prepared using blood from uninfected persons. To simulate collection of dried blood spots in the field, blood was spotted in 20-μl aliquots onto Whatman 3MM filter paper, air dried overnight, and stored at room temperature in plastic bags sealed with desiccant. Plasmodium vivax dried blood spots were obtained from the Centers for Disease Control and Prevention of the Republic of Korea. Punches 3 mm in diameter were removed from the dried blood spots using handheld hole punches.
Optimizing the pooling methodology.
We used the laboratory-prepared P. falciparum blood samples to compare several DNA extraction and amplification methods across a range of pool sizes (10, 50, and 100) and parasite densities (0, 10, and 100 parasites/μl). Once the optimal strategy was identified, additional tests were performed in pool sizes of 5, 10, 25, and 50 to generate estimates of sensitivity. Pool sizes of 100 did not easily fit into the 1.5-ml tubes used for extraction and were therefore not pursued further. The optimal strategy was also used to test the P. vivax clinical samples in pools of 25.
Test pools contained one punch from a positive dried blood spot, the remainder being punches from malaria-negative dried blood spots. A single DNA extraction was performed on each pool of punches, and the extraction product was used as a template for amplification.
Two different DNA extraction methods were evaluated: the saponin/Chelex method (24) and the QIAamp DNA minikit (Qiagen, CA). The sensitivities of four detection methods were compared: two nested techniques, 18S ribosomal DNA (rDNA) nested PCR (29) and cytochrome b nested PCR (30), and two single-round techniques, 18S rDNA PCR (21) and 18S rDNA probe-based real-time PCR (28). Real-time amplification was performed using a 7500 Real-Time PCR System (Applied Biosystems, CA). Other PCR amplifications were performed on a Bio-Rad Thermocycler C1000 or S1000 (Bio-Rad Laboratories, CA). Amplification products were detected by agarose gel electrophoresis or via real-time PCR, as appropriate, by persons blinded to the status of the specimens.
Field samples from the study cohort.
Once Chelex extraction followed by cytochrome b nested PCR was identified as the optimal strategy, it was used to test dried blood spots collected using passive and active surveillance of a pediatric cohort randomly selected from the Mulago III parish of Kampala, Uganda (T. D. Clark, D. Njama-Meya, D. B. Nzarubara, C. Maiteki-Sebuguzi, B. Greenhouse, S. G. Staedke, M. R. Kamya, G. Dorsey, and P. J. Rosenthal, unpublished data). Children were enrolled from February to May 2007 and followed until December 2008 (7). As part of passive surveillance, blood smears and dried blood spots were collected whenever the children presented to the clinic with a new episode of fever (subjective or based on a temperature of ≥38°C in the previous 24 h). As part of active surveillance, blood smears and dried blood spots were collected from asymptomatic children presenting for routine visits approximately every 90 days. Final microscopy results were based on a rigorous quality control system that included rereading of all blood smears by a second microscopist and resolution of any discrepancies between the first and second readings by a third microscopist. To prepare dried blood spots, blood was blotted on Whatman 3MM filter paper, air dried, and stored at room temperature in plastic bags sealed with desiccant.
Pooling strategy applied to field samples.
We used a three-stage hierarchical pooling strategy (23). Samples were first divided into master pools with the optimal size dependent on the estimated prevalence, as well as the sensitivity and specificity of the pooling assay (33, 34). Positive master pools were divided into subpools, with the optimal subpool size being the square root of the master pool size. Positive subpools were tested as individual specimens. Based on the known prevalence by microscopy in this study (∼5%) and the expected high sensitivity and specificity of the assay (∼100% for parasite densities of 100 parasites/μl), the most efficient master and subpool sizes were determined to be nine and three, respectively (34).
In the final step, we performed a restriction digestion, as the cytochrome b PCR assay begins with genus-specific detection, followed by species determination based on AluI restriction digestion (31). Species were identified by comparing digestion patterns to those of known controls. (See the supplemental material for a detailed protocol of the pooling strategy.)
Statistical analysis.
Data were analyzed using Stata 10.0 (StataCorp LP, TX). P values were generated using a chi-square or Fisher's exact test as appropriate.
RESULTS
Optimizing the pooling strategy.
Pooled Chelex extraction of DNA followed by cytochrome b nested PCR was identified as the most robust strategy. In pools containing one very low-P. falciparum-parasitemic sample (10 parasites/μl), the sensitivity ranged from 40% with pools of 100 to 93% with pools of 10. For all pool sizes that included one low-P. falciparum-parasitemic sample (100 parasites/μl), 100% sensitivity was achieved. The specificity was 100% throughout (Table 1). Sensitivity results from other DNA extraction/PCR combinations are shown in the supplemental material.
TABLE 1.
Sensitivity and specificity of pooled Chelex DNA extraction followed by cytochromeb nested PCR
| Pool size | % Sensitivity (n = 30)a |
% Specificity (n = 15)a |
|
|---|---|---|---|
| 100 p/μl | 10 p/μl | 0 p/μl | |
| 50 | 100 | 70 | 100 |
| 25 | 100 | 65 | 100 |
| 10 | 100 | 93 | 100 |
| 5 | 100 | 80 | 100 |
n refers to independent pooled extractions followed by PCR; p/μl, parasites/μl.
We also evaluated field samples from three patients diagnosed with P. vivax infection in a similar manner. The sensitivities for the detection of one positive sample in pools of 25 were 100% (10/10) when the sample had 640 parasites/μl, 90% (9/10) at 2,000 parasites/μl, and 100% at 8,118 parasites/μl (10/10).
Cohort characteristics.
A total of 419 samples were collected through passive surveillance in patients who presented with fever, and 472 samples were collected through active surveillance in asymptomatic subjects. The cohort included 77 children; the mean age at the start of the study period was 3 years (range, 1 to 10 years), and the median time of follow-up was 663 days (range, 616 to 730 days).
Application of pooling strategy to dried blood spots collected from the cohort.
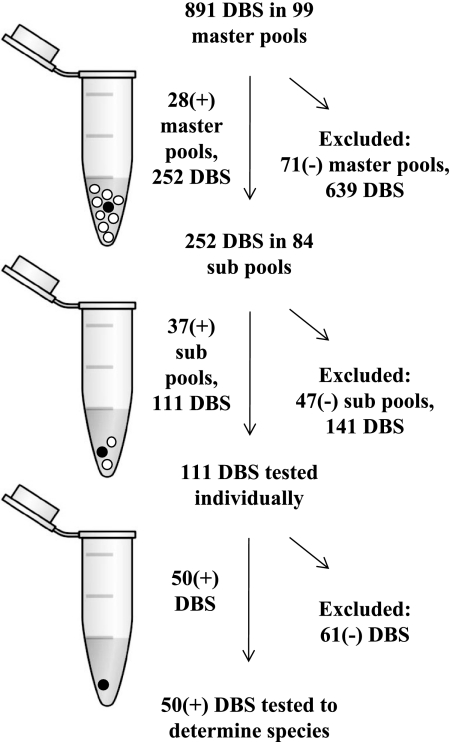
We used a three-stage pooling strategy, excluding samples that tested negative at each stage (Fig. 1). Ultimately, 50 of 891 dried blood spot samples were identified as positive, resulting in an overall prevalence of 5.6%. The efficiency of pooling, calculated as the number of tests performed per total number of individual samples evaluated, was 0.33. In other words, our pooling strategy, compared to performing individual PCRs on all samples, reduced labor and supply costs by about two-thirds.
FIG. 1.
Schematic of pooling strategy applied to cohort samples. The white circles represent Plasmodium-negative dried blood spots (DBS); the solid black circles represent Plasmodium-positive dried blood spots. At each stage, the dried blood spots were extracted together. The DNA extraction product was then used as a template for cytochrome b nested PCR.
Passive surveillance.
Of the 50 samples that were positive by pooling, 35 were among the 419 samples collected from febrile patients. All 35 of these samples were also positive by microscopy. Five samples that were positive by microscopy were negative by pooling (Table 2). The incidence of symptomatic infections was 0.32 case per person-year by microscopy versus 0.28 case per person-year by our pooling strategy.
TABLE 2.
Microscopy versus PCR-based pooling in passive surveillance of symptomatic subjects and active surveillance of asymptomatic subjects
| Surveillance | Poolinga | Microscopya |
Total | |
|---|---|---|---|---|
| + | − | |||
| Passive | + | 35 | 0 | 35 |
| − | 5 | 379 | 384 | |
| Total | 40 | 379 | 419 | |
| Active | + | 4 | 11 | 15 |
| − | 0 | 457 | 457 | |
| Total | 4 | 468 | 472 | |
+, positive; −, negative.
Of the 5 microscopy-positive samples that were missed by pooling, 3 were excluded at the master pool level and 2 at the subpool level. These samples had two characteristics: they had low parasite densities (64, 400, 400, 880, and 5,680 parasites/μl) and were tested more than 2 years after collection. Among all 40 microscopy-positive samples from symptomatic subjects, we compared those that had both characteristics (parasite density of less than 6,000 parasites/μl and storage for more than 2 years) with those that did not. The sensitivity of pooling for detection of parasites was 29% (2/7) among samples with both characteristics versus 100% (33/33) among samples without both characteristics (P < 0.001).
Active surveillance.
Of the 50 samples positive by pooling, 15 were among the 472 samples collected from asymptomatic patients. Four were detected by microscopy, and 11 were not. Pooling successfully identified all the microscopy-positive samples (Table 2). The prevalence of asymptomatic infections was 0.8% by microscopy versus 3.2% by pooling (P = 0.01).
For the 11 samples that were negative by microscopy but positive by pooling, reviews of the medical histories suggested that many represented asymptomatic parasitemia. Two samples were from patients who presented symptomatic with positive blood smears 5 to 11 days later. Five samples represented consecutive collections from two patients. One sample was obtained from a patient who had gametocytes on smear 3 days later. Two samples were obtained at enrollment, when the prevalence of asymptomatic parasitemia was high in the cohort (8). The last sample was from a patient who did not have any follow-up smears; we were therefore unable to assess the possibility of persistent infection.
Species determination.
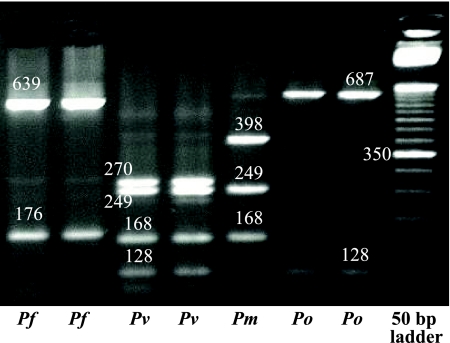
Species determination was based on AluI restriction digestion of the amplified cytochrome b gene (Fig. 2). Among the 35 samples that were positive by both microscopy and pooling, there was agreement in the detection of 31 P. falciparum infections and 1 Plasmodium ovale infection. Three samples identified as P. falciparum by microscopy were identified by pooling as 2 P. ovale infections and 1 mixed P. falciparum-P. vivax infection. Of the 11 asymptomatic cases positive by PCR but not microscopy, there were 7 P. falciparum infections, 1 Plasmodium malariae infection, 1 P. ovale infection, and 2 mixed P. falciparum-P. ovale infections.
FIG. 2.
AluI digestion for species determination. P. ovale digestion patterns may vary depending on the strain type. Our samples showed a pattern that correlates with a published sequence (GenBank accession number AB182497). However, another sequence (GenBank accession number AB182496) would produce a different AluI restriction pattern with expected band sizes of 205 and 610 bp. Pf, P. falciparum; Pm, P. malariae; Po, P. ovale; Pv, P. vivax.
DISCUSSION
We developed and applied a novel strategy for PCR-based pooled detection of malaria parasites using dried blood spots. After evaluating numerous methods on laboratory-prepared samples, we found the optimal strategy to be Chelex extraction of pooled 3-mm punches from dried blood spots, followed by cytochrome b nested PCR for Plasmodium detection and AluI restriction digestion for species determination. Because of the sensitive detection associated with this method, we were able to reliably identify a single 100-parasite/μl sample in pool sizes up to 50. We then applied the optimized strategy to 891 field samples collected through both passive and active surveillance in a well-characterized pediatric cohort. Compared to microscopy, we detected fewer cases among symptomatic patients (35/419 versus 40/419), missing 5 microscopy-positive samples that were all stored for longer than 2 years and that tended to have lower parasitemia. Among asymptomatic patients, we detected 3 times as many infections as by microscopy (15/472 versus 4/472). The pooling strategy also allowed improved detection of non-falciparum and mixed infections.
Monitoring, evaluation, and appropriate targeting of interventions depend on accurate surveillance, particularly in low-transmission and elimination settings. Compared to microscopy and RDTs, PCR is well known to provide improved sensitivity and specificity (22). However, the long turnaround time for PCR makes it impractical for rapid diagnosis of individual patients, and in surveillance settings, it is labor and cost prohibitive to perform individual PCRs on large numbers of samples. Pooling is a simple strategy that has recently been applied to the detection of malaria parasites in surveillance settings. In an exploratory study using 18S rDNA nested PCR and pool sizes of 10, there was successful detection of 6 positives among 200 serum samples (2). In a larger study that utilized dried blood spot samples collected from a cohort of pregnant women, individual DNA extractions were performed, and then the extraction products from 4 samples were pooled in a single 18S rDNA real-time PCR. Among 1,092 microscopy-negative samples that were tested by pooling, 35 positive samples were identified. Among 176 microscopy-positive samples that were tested in individual PCRs, 74 were positive (32). Our study offers advances over prior studies. Importantly, we used easily collected dried blood spots, optimized strategies after comparison of multiple related methods, validated the sensitivity and specificity of our strategy by comparison with laboratory controls, and were able to pool large numbers of samples.
The inexpensive Chelex method performed better than the commonly used spin column system, and at lower parasite densities and higher pool sizes, cytochrome b nested PCR provided the highest sensitivity. Others have also found the cytochrome b method to be more sensitive than 18S rDNA nested PCR (13, 30), likely because there are more copies of the gene in each parasite (15, 25). It also performed better than two other single-round PCR methods, including one real-time method. The cytochrome b PCR method is also advantageous in that amplification targets a DNA sequence common to all species (30). For this reason, our pooling strategy requires tests of speciation only at the final stage of pooling, further simplifying the assay.
Unlike published strategies in which pooling occurred at the time of PCR detection (32, 34), we pooled at the time of DNA extraction. By using this approach and by using optimal pool sizes based on the anticipated prevalence, our strategy allowed pooled detection of large numbers of samples and reduced much of the labor associated with DNA extraction. The findings that sensitivity was maintained and that experiments were easy to perform at pool sizes up to 50 are particularly important, since the ability to pool many samples dramatically increases efficiency, especially in low-prevalence settings. For example, if our pooling strategy were used in a setting where the prevalence of parasitemia was 0.01%, using pool sizes of 49 would result in an efficiency of 0.05, saving approximately 95% of the labor and supply costs (34). Also, of particular relevance to low-prevalence settings, our proposed strategy can potentially provide improved specificity, given that an individual sample must be amplified by PCR three times to be identified as positive. Specificity was not a problem in our study, but false positives could occur in situations where there is cross-contamination. The fact that our strategy involves pooled DNA extractions with resampling from the original dried blood spot at each stage is a further advance over methods that require resampling from a potentially contaminated DNA extraction product.
When applied to field samples, we found that the sensitivity of low-parasitemic samples stored for at least 2 years was compromised, highlighting the possibility of DNA degradation with long-term storage of samples. No microscopy-positive samples stored for less than 2 years were missed by pooling, suggesting that pooled testing of samples within 2 years of collection is reliable. In addition, extremely low parasite densities, around 10 parasites/μl, may not be detected reliably, likely due to the diluting effect of pooling. However, increased sensitivity was demonstrated with smaller pool sizes (see Table S1 in the supplemental material). Also, this strategy enabled the detection of 11 submicroscopic infections, 6 of which were detected in samples that had been stored for more than 2 years.
There are several potential applications for our pooling strategy. First, it is a sensitive and high-throughput method that can improve the detection of malaria infections in community surveys of parasite prevalence (12). Second, there is potential for our pooling strategy to be used for quality assurance of standard diagnostics in research or clinical settings (17). Third, it allows the identification of asymptomatic submicroscopic infections. Similar to the results of a recently performed systematic review of submicroscopic infections, we detected about three times as many parasitemic samples as with microscopy (20). Submicroscopic asymptomatic infections are known to be a major driver of malaria transmission (3, 6, 29), but how and to what extent remains unclear. Using samples from easily collected dried blood spots, our pooling strategy can facilitate further studies on this topic.
Lastly, there are several possible applications of our pooling strategy for malaria elimination programs. It can be applied in mass screening and treatment of asymptomatic populations, which is emerging as a potentially important strategy to facilitate malaria elimination (19). For example, mass screening and treatment are being applied as part of recent efforts to eliminate artemisinin-resistant parasites that have emerged at the Thai-Cambodian border (World Health Organization, presented at the Strategy for the Containment of Artemisinin Tolerant Malaria Parasites in South-East Asia Project Meeting, Pailin, Cambodia, 2009). Areas that have already achieved elimination have instituted high-volume border-screening programs in order to prevent reintroduction of malaria. Mauritius (5), Oman (27), and Singapore (16) are examples of countries that perform individual screening of asymptomatic visitors and/or immigrants using microscopy or RDTs. While such mass screening strategies are feasible using existing techniques in resource-rich countries, use of the proposed pooling strategy may put mass screening within the reach of resource-poor sites (5, 35).
In summary, we have developed and applied a robust and novel strategy for high-throughput PCR-based detection of malaria parasites using dried blood spots. Compared to microscopy, our PCR-based pooling strategy allows improved detection of species and, most importantly, asymptomatic infections. There is a need for improved surveillance in low-transmission settings, and our proposed strategy holds promise as a simple new tool to facilitate progress toward malaria elimination.
Supplementary Material
Acknowledgments
This work was supported by grants from the Pediatric Infectious Diseases Society, National Institutes of Health (grants U01 AI52142, R01 MH068686, and P01 AI071713), the Bill and Melinda Gates Foundation (grant 48820), and the University of California, San Francisco, Dean's Research Fellowship Program.
We thank Nicolas Steenkeste for sharing details of the cytochrome b nested PCR and AluI digestion for species detection, Ric Price for sharing standard operating procedures of other tested PCR methods, and Jung-Yeon Kim for sharing of clinical P. vivax samples. We thank Phil Rosenthal and Jay Tureen for their guidance and helpful review of the manuscript and Richard Feachem for his support of the project.
We declare that we have no conflicts of interest for this article.
Footnotes
Published ahead of print on 4 August 2010.
Supplemental material for this article may be found at http://jcm.asm.org/.
REFERENCES
- 1.Bebell, L. M., C. D. Pilcher, G. Dorsey, D. Havlir, M. R. Kamya, M. P. Busch, J. D. Williams, C. T. Nugent, C. Bentsen, P. J. Rosenthal, and E. D. Charlebois. 2010. Acute HIV-1 infection is highly prevalent in Ugandan adults with suspected malaria. AIDS 24:1945-1952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bharti, A. R., S. L. Letendre, K. P. Patra, J. M. Vinetz, and D. M. Smith. 2009. Malaria diagnosis by a polymerase chain reaction-based assay using a pooling strategy. Am. J. Trop. Med. Hyg. 81:754-757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Boudin, C., M. Olivier, J. F. Molez, J. P. Chiron, and P. Ambroise-Thomas. 1993. High human malarial infectivity to laboratory-bred Anopheles gambiae in a village in Burkina Faso. Am. J. Trop. Med. Hyg. 48:700-706. [DOI] [PubMed] [Google Scholar]
- 4.Busch, M. P., D. J. Wright, B. Custer, L. H. Tobler, S. L. Stramer, S. H. Kleinman, H. E. Prince, C. Bianco, G. Foster, L. R. Petersen, G. Nemo, and S. A. Glynn. 2006. West Nile virus infections projected from blood donor screening data, United States, 2003. Emerg. Infect. Dis. 12:395-402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4a.Clark, T. D., D. Njama-Meya, B. Nzarubara, C. Maiteki-Sebuguzi, B. Greenhouse, S. G. Staedke, M. R. Kamya, G. Dorsey, and P. J. Rosenthal. 2010. Incidence of malaria and efficacy of combination antimalarial therapies over 4 years in an urban cohort of Ugandan children. PLoS One 5:e11759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cohen, J. M., D. L. Smith, A. Vallely, G. Taleo, G. Malefoasi, and O. Sabot. 2009. Holding the line, p. 40-60. In R. G. A. Feachem, A. Phillips, and G. A. Targett (ed.), A prospectus on malaria elimination. Global Health Group, San Francisco, CA.
- 6.Coleman, R. E., C. Kumpitak, A. Ponlawat, N. Maneechai, V. Phunkitchar, N. Rachapaew, G. Zollner, and J. Sattabongkot. 2004. Infectivity of asymptomatic Plasmodium-infected human populations to Anopheles dirus mosquitoes in western Thailand. J. Med. Entomol. 41:201-208. [DOI] [PubMed] [Google Scholar]
- 7.Davis, J. C., T. D. Clark, S. K. Kemble, N. Talemwa, D. Njama-Meya, S. G. Staedke, and G. Dorsey. 2006. Longitudinal study of urban malaria in a cohort of Ugandan children: description of study site, census and recruitment. Malar. J. 5:18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dorsey, G., S. Staedke, T. D. Clark, D. Njama-Meya, B. Nzarubara, C. Maiteki-Sebuguzi, C. Dokomajilar, M. R. Kamya, and P. J. Rosenthal. 2007. Combination therapy for uncomplicated falciparum malaria in Ugandan children: a randomized trial. JAMA 297:2210-2219. [DOI] [PubMed] [Google Scholar]
- 9.Feachem, R., and O. Sabot. 2008. A new global malaria eradication strategy. Lancet 371:1633-1635. [DOI] [PubMed] [Google Scholar]
- 10.Feachem, R. G. A., and the Malaria Elimination Group. 2009. Shrinking the malaria map, a guide on malaria elimination for policy makers. Global Health Group, San Francisco, CA.
- 11.Greenwood, B. M. 2008. Control to elimination: implications for malaria research. Trends Parasitol. 24:449-454. [DOI] [PubMed] [Google Scholar]
- 12.Guerra, C. A., P. W. Gikandi, A. J. Tatem, A. M. Noor, D. L. Smith, S. I. Hay, and R. W. Snow. 2008. The limits and intensity of Plasmodium falciparum transmission: implications for malaria control and elimination worldwide. PLoS Med. 5:e38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hasan, A. U., S. Suguri, J. Sattabongkot, C. Fujimoto, M. Amakawa, M. Harada, and H. Ohmae. 2009. Implementation of a novel PCR based method for detecting malaria parasites from naturally infected mosquitoes in Papua New Guinea. Malar. J. 8:182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jensen, J. B. 2002. In vitro culture of Plasmodium parasites. Methods Mol. Med. 72:477-488. [DOI] [PubMed] [Google Scholar]
- 15.Langsley, G., J. E. Hyde, M. Goman, and J. G. Scaife. 1983. Cloning and characterisation of the rRNA genes from the human malaria parasite Plasmodium falciparum. Nucleic Acids Res. 11:8703-8717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lee, Y. C., C. S. Tang, L. W. Ang, H. K. Han, L. James, and K. T. Goh. 2009. Epidemiological characteristics of imported and locally-acquired malaria in Singapore. Ann. Acad. Med. Singapore 38:840-849. [PubMed] [Google Scholar]
- 17.McMorrow, M. L., M. I. Masanja, E. Kahigwa, S. M. Abdulla, and S. P. Kachur. 2010. Quality assurance of rapid diagnostic tests for malaria in routine patient care in rural Tanzania. Am. J. Trop. Med. Hyg. 82:151-155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nugent, C. T., J. Dockter, F. Bernardin, R. Hecht, D. Smith, E. Delwart, C. Pilcher, D. Richman, M. Busch, and C. Giachetti. 2009. Detection of HIV-1 in alternative specimen types using the APTIMA HIV-1 RNA Qualitative Assay. J. Virol. Methods 159:10-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ogutu, B., A. B. Tiono, M. Makanga, Z. Premji, A. D. Gbadoe, D. Ubben, A. C. Marrast, and O. Gaye. 2010. Treatment of asymptomatic carriers with artemether-lumefantrine: an opportunity to reduce the burden of malaria? Malar. J. 9:30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Okell, L. C., A. C. Ghani, E. Lyons, and C. J. Drakeley. 2009. Submicroscopic infection in Plasmodium falciparum-endemic populations: a systematic review and meta-analysis. J. Infect. Dis. 200:1509-1517. [DOI] [PubMed] [Google Scholar]
- 21.Padley, D., A. H. Moody, P. L. Chiodini, and J. Saldanha. 2003. Use of a rapid, single-round, multiplex PCR to detect malarial parasites and identify the species present. Ann. Trop. Med. Parasitol. 97:131-137. [DOI] [PubMed] [Google Scholar]
- 22.Perkins, M. D., and D. Bell. 2008. Working without a blindfold: the critical role of diagnostics in malaria control. Malar. J. 7(Suppl. 1):S5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pilcher, C. D., J. J. Eron, Jr., S. Galvin, C. Gay, and M. S. Cohen. 2004. Acute HIV revisited: new opportunities for treatment and prevention. J. Clin. Invest. 113:937-945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Plowe, C. V., A. Djimde, M. Bouare, O. Doumbo, and T. E. Wellems. 1995. Pyrimethamine and proguanil resistance-conferring mutations in Plasmodium falciparum dihydrofolate reductase: polymerase chain reaction methods for surveillance in Africa. Am. J. Trop. Med. Hyg. 52:565-568. [DOI] [PubMed] [Google Scholar]
- 25.Preiser, P. R., R. J. Wilson, P. W. Moore, S. McCready, M. A. Hajibagheri, K. J. Blight, M. Strath, and D. H. Williamson. 1996. Recombination associated with replication of malarial mitochondrial DNA. EMBO J. 15:684-693. [PMC free article] [PubMed] [Google Scholar]
- 26.Quinn, T. C., R. Brookmeyer, R. Kline, M. Shepherd, R. Paranjape, S. Mehendale, D. A. Gadkari, and R. Bollinger. 2000. Feasibility of pooling sera for HIV-1 viral RNA to diagnose acute primary HIV-1 infection and estimate HIV incidence. AIDS 14:2751-2757. [DOI] [PubMed] [Google Scholar]
- 27.Roll Back Malaria Partnership. 2008. Elimination and eradication: achieving zero transmission, Global malaria action plan, part II.3. Roll Back Malaria, Geneva, Switzerland.
- 28.Shokoples, S. E., M. Ndao, K. Kowalewska-Grochowska, and S. K. Yanow. 2009. Multiplexed real-time PCR assay for discrimination of Plasmodium species with improved sensitivity for mixed infections. J. Clin. Microbiol. 47:975-980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Snounou, G., S. Viriyakosol, X. P. Zhu, W. Jarra, L. Pinheiro, V. E. do Rosario, S. Thaithong, and K. N. Brown. 1993. High sensitivity of detection of human malaria parasites by the use of nested polymerase chain reaction. Mol. Biochem. Parasitol. 61:315-320. [DOI] [PubMed] [Google Scholar]
- 30.Steenkeste, N., S. Incardona, S. Chy, L. Duval, M. T. Ekala, P. Lim, S. Hewitt, T. Sochantha, D. Socheat, C. Rogier, O. Mercereau-Puijalon, T. Fandeur, and F. Ariey. 2009. Towards high-throughput molecular detection of Plasmodium: new approaches and molecular markers. Malar. J. 8:86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Steenkeste, N., W. O. Rogers, L. Okell, I. Jeanne, S. Incardona, L. Duval, S. Chy, S. Hewitt, M. Chou, D. Socheat, F. X. Babin, F. Ariey, and C. Rogier. 2010. Sub-microscopic malaria cases and mixed malaria infection in a remote area of high malaria endemicity in Rattanakiri province, Cambodia: implication for malaria elimination. Malar. J. 9:108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Taylor, S. M., J. J. Juliano, P. A. Trottman, J. B. Griffin, S. H. Landis, P. Kitsa, A. K. Tshefu, and S. R. Meshnick. 2010. High-throughput pooling and real-time PCR-based strategy for malaria detection. J. Clin. Microbiol. 48:512-519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Westreich, D. J., M. G. Hudgens, S. A. Fiscus, and C. D. Pilcher. 1 March 2010, posting date. Optimal pooling strategies for acute HIV—web calculator v1.0. http://www.bios.unc.edu/∼mhudgens/optimal.pooling.htm.
- 34.Westreich, D. J., M. G. Hudgens, S. A. Fiscus, and C. D. Pilcher. 2008. Optimizing screening for acute human immunodeficiency virus infection with pooled nucleic acid amplification tests. J. Clin. Microbiol. 46:1785-1792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zanzibar Malaria Control Program. 2009. Malaria elimination in Zanzibar: a feasibility assessment. Ministry of Health and Social Welfare, Zanzibar City, Zanzibar.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.