Abstract
Dihydropyrimidine dehydrogenase (DPD) is the initial enzyme acting in the catabolism of the widely used antineoplastic agent 5-fluorouracil (5FU). DPD deficiency is known to cause a potentially lethal toxicity following administration of 5FU. Here, we report novel genetic mechanisms underlying DPD deficiency in patients presenting with grade III/IV 5FU-associated toxicity. In one patient a genomic DPYD deletion of exons 21–23 was observed. In five patients a deep intronic mutation c.1129–5923C>G was identified creating a cryptic splice donor site. As a consequence, a 44 bp fragment corresponding to nucleotides c.1129–5967 to c.1129–5924 of intron 10 was inserted in the mature DPD mRNA. The deleterious c.1129–5923C>G mutation proved to be in cis with three intronic polymorphisms (c.483 + 18G>A, c.959–51T>G, c.680 + 139G>A) and the synonymous mutation c.1236G>A of a previously identified haplotype. Retrospective analysis of 203 cancer patients showed that the c.1129–5923C>G mutation was significantly enriched in patients with severe 5FU-associated toxicity (9.1%) compared to patients without toxicity (2.2%). In addition, a high prevalence was observed for the c.1129–5923C>G mutation in the normal Dutch (2.6%) and German (3.3%) population. Our study demonstrates that a genomic deletion affecting DPYD and a deep intronic mutation affecting pre-mRNA splicing can cause severe 5FU-associated toxicity. We conclude that screening for DPD deficiency should include a search for genomic rearrangements and aberrant splicing.
Electronic supplementary material
The online version of this article (doi:10.1007/s00439-010-0879-3) contains supplementary material, which is available to authorized users.
Introduction
Dihydropyrimidine dehydrogenase (DPD, EC 1.3.1.2) is the initial and rate-limiting enzyme in the catabolism of the pyrimidine bases uracil and thymine but also of the antineoplastic agent 5-fluorouracil. The pivotal role of DPD in 5FU-based chemotherapy has been shown in cancer patients with a complete or partial deficiency of this enzyme. These patients suffered from severe, possibly fatal multi-organ toxicity following the administration of 5FU (Johnson and Diasio 2001; Milano et al. 1999; van Kuilenburg et al. 2000).
The presence of mutated DPYD has been reported in patients experiencing severe 5FU toxicity (Morel et al. 2006; van Kuilenburg et al. 2000). For that reason, various strategies have been proposed to screen for patients with a DPD deficiency, including genotyping (Amstutz et al. 2009; Kleibl et al. 2009). However, in a significant number of patients with a reduced DPD activity, no mutations could be identified in the coding part of DPYD (Collie-Duguid et al. 2000; Mattison et al. 2004; van Kuilenburg et al. 2000). The recent observation that a DPYD haplotype not containing any nonsynonymous or splice-site mutations was associated with 5FU toxicity, suggested the presence of additional genetic variations in the noncoding region of DPYD (Amstutz et al. 2009). So far, no deep intronic mutations in DPYD have been described affecting the splicing of DPD pre-mRNA.
DPYD is present as a single copy gene on chromosome 1p21.3 and consists of 23 exons (Wei et al. 1998). Recently, we showed that genomic deletions affecting DPYD occurred in 7% of paediatric patients with a complete DPD deficiency (van Kuilenburg et al. 2009). Therefore, it is conceivable that genomic deletions encompassing a part or the entire DPYD gene might also provide a molecular basis for cancer patients with a phenotypically established DPD deficiency. In this study, we investigated the presence or absence of genomic rearrangements in DPYD, using multiplex ligation-dependent probe amplification (MLPA), in 92 patients with a reduced DPD activity and/or grade III/IV toxicity. Furthermore, we characterized a novel missense mutation K874R based on the crystal structure of DPD and identified the first deep intronic mutation in DPYD affecting the regulation of pre-mRNA splicing.
Materials and methods
Patients
Ninety-two cancer patients with reduced DPD activity and/or grade III/IV toxicity were examined for DPYD rearrangements. The study cohort consisted of 64 Dutch patients who had been referred for diagnostic purposes between 2007 and 2009 and proved to have a reduced DPD activity in peripheral blood mononuclear cells (Milano et al. 1999; van Kuilenburg et al. 2002b), ranging from 0.7–5.9 nmol/mg/h (local controls 9.6 ± 2.6 nmol/mg/h). The DPD activity in 11 French cancer patients ranged from 0.7 to 10.6 nmol/mg/h (local controls 13.3 ± 5.0 nmol/mg/min). In addition, 17 German patients presenting with grade IV 5FU-associated toxicity were analysed, including one previously reported patient (patient “2” from Gross et al. 2003). Toxicity grading was based on the common toxicity criteria and adverse event reporting guidelines from the National Cancer Institute (NCI-CTC AE, version 3.0; http://ctep.cancer.gov/reporting/ctc.html). Four additional Dutch subjects of whom exonic sequence data were available, including two patients with severe 5FU toxicity, a son of a patient with lethal toxicity and a healthy volunteer who was coincidently found to have a DPD deficiency during another study were further investigated for intronic mutations and alternative splicing. Subsequently, the presence of a deep intronic mutation was retrospectively analysed in 66 German cancer patients presenting with grade III/IV 5FU-associated toxicity and 137 patients with no or mild toxicity (Supplemental Table 1). In addition, the prevalence of the mutation was investigated in 453 and 191 healthy individuals from Germany and the Netherlands, respectively.
MLPA and array-based comparative genomic hybridization analysis
Genomic DNA was isolated from EDTA-blood using standard techniques. The MLPA test for DPYD (P103, MRC-Holland, Amsterdam, The Netherlands) contains 38 probes for DPYD, including one probe to detect the c.1905 + 1G>A mutation, and nine control probes specific for DNA sequences outside the DPYD gene. MLPA was performed as described before (Schouten et al. 2002). Data analysis was performed using Genescan and Genotyper software (Applied Biosystems, Nieuwekerk a/d IJssel, The Netherlands). The relative peak area of each probe was divided by the average relative peak area of this probe in control samples. In unaffected individuals this will result in a value of 1 (100%) representing two copies of the target sequence in the sample.
Array CGH was performed using commercially available 105 K Agilent slides, AMADID 019015 with a 21.7 kB overall median probe spacing (Oxford design, Agilent, Santa Clara, CA, USA) according to the manufacturer’s protocol (Oligonucleotide Array-Based CGH for Genomic DNA Analysis V5.0 June 2007). Fluorescent labelling of DNA was performed using the CGH labelling kit for Oligo Arrays (Enzo Life Sciences, Inc., Farmingdale, NY, USA). Slides were scanned using a 2 μm Agilent scanner, image analysis was performed with Agilent Feature extraction V9.5.3 and data analysis was performed using Agilent DNA Analytics (V4.0.76).
Sequence analysis of cDNA and gDNA coding for DPD
DNA was isolated from blood using standard techniques. PCR amplification of all 23 coding exons and flanking intronic regions of DPYD was carried out using intronic primer sets, essentially as described before (van Kuilenburg et al. 2000). Exon 6 was amplified using the forward primer DPYD6f (5′-CTGAAAATGTACTGCTCATTGCT-3′) and reverse primer DPYD6r (5′-GCTTCAAGCCAACTGCAAAT-3′). Part of intron 10 of DPYD was amplified using the forward primer DPYD10f (5′-TCAGACCAAATCATCGCATT-3′) and reversed primer DPYD10r (5′-TTCTCCTCATGGCACCCATA-3′).
Total RNA was isolated from 20 ml of heparinized blood and cultured fibroblasts using Trizol extraction (Invitrogen, Carlsbad, CA, USA). cDNA was prepared using a first strand cDNA synthesis kit for RT-PCR (Roche, Mannheim, Germany). PCR amplification of a part of the cDNA sequence of exons 8–11 was performed using the forward primer (5′-GCTACAAAGCTGCTTTCATTG-3′) and reverse primer (5′-CACTGATGACCACATCGGC-3′), corresponding to the nucleotides c.824_844 (exon 8) and c.1291_1309 (exon 11), respectively. cDNA analysis of exon 22 was performed using the forward primer (5′-TGGACCTTATCTGGAACAGC-3′) and reverse primer (5′-TCAGCATATGTAGGTGACATG-3′) corresponding to the nucleotides c.2637_2656 (exon 21) and c.3110_3130 (exon 23), respectively.
Amplification of cDNA and part of intron 10 was carried out in 25 μl reaction mixtures containing 20 mM Tris/HCl pH 8.4, 50 mM KCl, 1.5 mM MgCl2, 0.4 μM of each primer, 0.2 mM dNTPs and 0.02 U of Platinum Taq polymerase. After initial denaturation for 5 min at 95°C, amplification was carried out for 30 cycles (30 s 95°C, 30 s 55°C, 1 min 72°C) with a final extension step of 10 min at 72°C. PCR products were separated on 1.5% agarose gels, visualized with ethidium bromide or used for direct sequencing.
Population screening for genomic DPYD mutations was performed by direct sequence analysis and DHPLC, as previously described (Seck et al. 2005). Sequence analysis of cDNA and genomic fragments amplified by PCR was carried out on an Applied Biosystems model 3100 automated DNA sequencer using the dye-terminator method (Applied Biosystems, Nieuwekerk a/d IJssel, The Netherlands). The DPYD sequence of patients was compared to that observed in controls and in the reference sequence of DPYD (Ref Seq NM_000110.3; Ensembl ENST00000370192). Analysis of sequences for cryptic splice sites was performed with Alamut 1.5 software (Interactive Biosoftware, Rouen, France).
Quantitative PCR analysis
Quantitative PCR analysis of the wild-type DPD cDNA and the cDNA containing the 44 bp insertion of intron 10 was performed using the forward primers 5′-GAGGAGATGGAGCTTGCTAA-3′ and 5′-AGGAGCATCAGCCACATATC-3′, respectively, and the reverse primer 5′-CTGAACCAAAGGCACTGATG-3′. Quantitative PCR analysis of the wild-type DPD cDNA and the cDNA containing the exon 22 deletion was performed using the forward primers 5′-CCTACCATCAAGGATGTAAT-3′ and 5′-CCTACCATCAAGGCTATACA-3′, respectively, and the reverse primer 5′-AGATAAGGGTACGCCTCTCT-3′. The analysis was performed using the LightCycler® 480 SYBR Green I Master kit (Roche Diagnostics Nederland B.V, Almere, The Netherlands).
Crystal structure analysis
The homology model of human DPD was generated by manual adjustment of the sequence of the deposited crystal structure of recombinant pig DPD (PDB-ID: 1h7w; Dobritzsch et al. 2001) to that of the human enzyme using WinCoot (Emsley et al. 2010). An energetically preferred side chain conformation was chosen that causes no steric clashes with surrounding residues. The structural analysis was performed using WinCoot. Figure 5 was generated with the program PyMol (DeLano, W.L. The PyMOL Molecular Graphics System (2002). http://www.pymol.org).
Fig. 5.
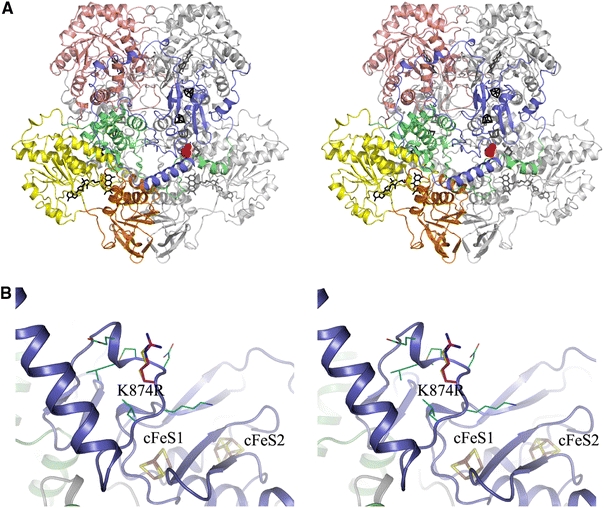
Location and environment of the K874R mutation site in the human DPD homology model. a Stereoview of the homodimeric DPD in cartoon representation. One subunit is shown entirely in grey, the domains of the other subunit are coloured as follows: FeS-cluster domain I (amino acids 27–172), green; FAD-binding domain II (173–286, 442–524), yellow; NADPH-binding domain III (287–441), orange; FMN- and pyrimidine-binding domain IV (525–847), salmon; FeS-cluster domain V (1–26, 848–1025), blue. Cofactors are depicted as stick models in black for the coloured subunit, and in grey for the other. The mutation site is indicated by a space-fill model of the native amino acid side chain, shown in red for one subunit only. b Close-up stereoview of the K874R mutation site and its environment. The structure of the human DPD is shown in cartoon representation and coloured according to a. Stick models of the iron sulphur clusters are labelled according to Dobritzsch et al. (2001) and shown with iron atoms in brown and sulphur atoms in yellow. The stick-model of the native K874 is shown with carbon atoms in cyan, that of the R874 side chain introduced by the mutation with carbon atoms in magenta and thicker sticks. Residues surrounding the mutation site (870–873, 875–876) are shown with thin sticks and carbon atoms in cyan (colour figure online)
Statistics
Differences in the distribution of a variant or haplotype between two groups were analysed by Pearson Chi-square and the Mann–Whitney U test for independent samples. Asymptotic significance (two-sided) is indicated. Statistical analysis was performed with the program PASW Statistics 18.
Results
Analysis of intragenic rearrangements
MLPA analysis showed the presence of the c.1905 + 1G>A (IVS14 + 1G>A) mutation in 28% (18/64) and 9% (1/11) of cancer patients with a reduced DPD activity from the Netherlands and France, respectively. One out of 17 (6%) German patients with grade IV toxicity proved to be a carrier for the c.1905 + 1G>A mutation. Figure 1a shows the MLPA analysis of the copy numbers of the 23 coding exons and 4 intronic sequences of DPYD in the 92 cancer patients. The mean variation observed for the various probes directed against the exon and intron sequences of DPYD was 8.1 ± 1.8%, ranging from 5.0% for the exon 11.2 probe to 14.4% for the exon 1.1 probe. A relative copy number below the cut-off value indicative for a deletion (relative copy number <0.7) was observed for exons 1, 11 and 21–23 (Fig. 1a). A relatively large variation was observed for the exon 1.1 probe and unfortunately, no DNA was available anymore from this patient for further studies.
Fig. 1.
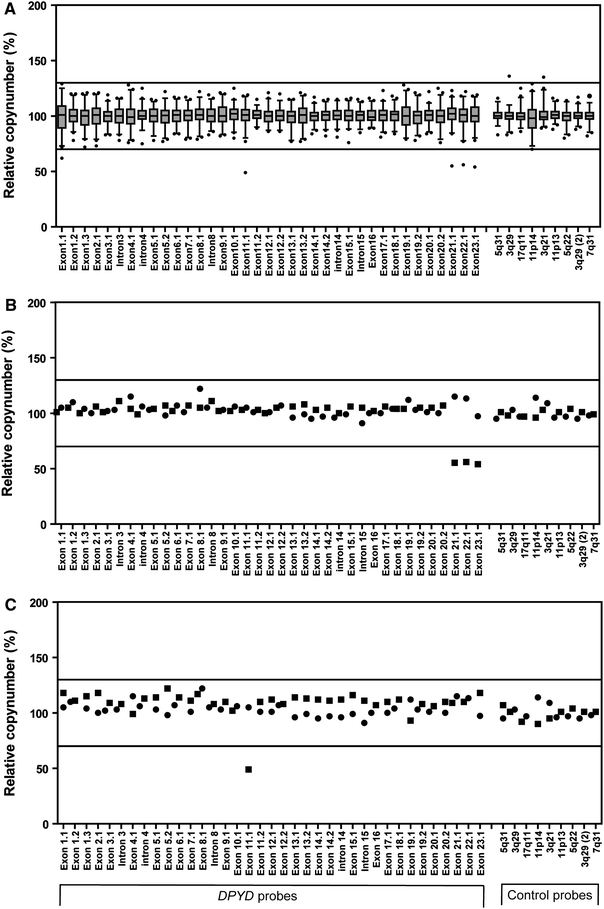
Analysis of copy number changes in DPYD using MLPA. a The box-plots of the quantitative analysis of the copy number of the 23 coding exons and four intronic sequences of DPYD and nine control probes specific for DNA sequences outside DPYD of 92 cancer patients. The top, bottom and line through the middle of a box correspond to the 75th, 25th and 50th percentile, respectively. The whiskers on the bottom extend from the 2.5th percentile and top 97.5th percentile. The circles represent outliers. b The MLPA analysis of patient 1 (filled square) and a control (filled circle). c The MLPA analysis of patient 2 (filled square) and a control (filled circle). The solid lines represent the cut-off values indicative for amplification (relative copy number > 1.3) or deletion (relative copy number < 0.7) of that particular sequence
In patient 1, in whom sequence analysis of DPYD did not provide unambiguous pathogenic mutations (patient 2 from Ref. Gross et al. 2003), MLPA analysis showed loss of heterozygosity for exons 21–23 (Fig. 1b). Array-based CGH was performed to delineate boundaries and size of the deletion. Detailed analysis of the chromosome 1p21.3 region encompassing DPYD showed a minimal deleted region of 61 kB ranging from 97320495 to 97381610, encompassing exons 22 and 23, and a maximal deletion of 99 kB ranging from 97302551 to 97401528, encompassing exons 21–23 of DPYD (Fig. 2).
Fig. 2.
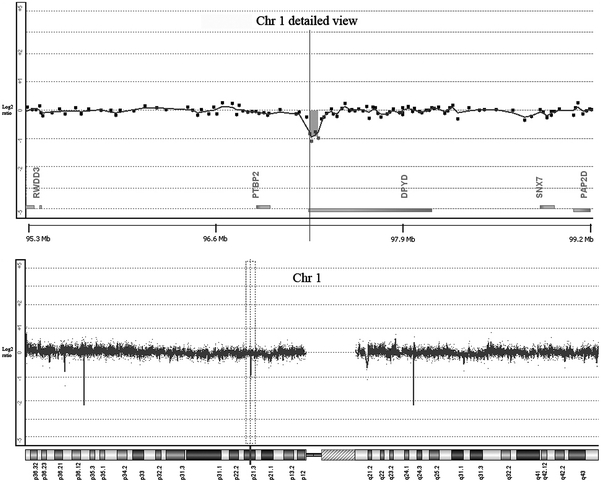
Detection of copy number changes by oligo array-based genomic hybridization for patient 1. The Y axis represents the log2 ratio of the intensities of patient and reference DNA. On the X axis oligos are ordered by Mb position. The lower panel shows the observed log2 ratios on chromosome (Chr) 1. The spikes outside the selected area are known copy number variants in the human DNA. The upper panel shows a magnified view of the selected area in the lower panel for the probes located in DPYD and flanking regions. The genes located in the flanking region of DPYD are indicated on the X axis. The grey box represents the minimal deleted region for patient 1
In patient 2, MLPA analysis suggested a deletion of part of exon 11 (Fig. 1c). However, sequence analysis of exon 11 and flanking regions showed that the patient was apparent homozygous for the c.1236G>A (p.E412E) mutation which is located near the ligation site of the MLPA probe 11.1. Therefore, the decreased signal is, most likely, caused by a decreased hybridization or ligation of the MLPA probe. Indeed, hybridization and ligation at a lower temperature resulted in normalization of the signal for the MLPA probe 11.1, thus excluding the presence of a deletion (results not shown).
Identification of a deep intronic mutation affecting pre-mRNA splicing
Total RNA was isolated from EDTA-blood of patient 2 and a control and subjected to RT-PCR amplification. A part of the coding sequence of the DPD cDNA, corresponding to the nucleotides c.824_c.1309, was amplified and the PCR fragments were analysed by gel electrophoresis. A normal sized fragment of 486 bp was detected in a control subject whereas the 486 bp fragment was found together with a larger sized fragment of 530 bp in patient 2 (Fig. 3a). Sequence analysis of the PCR fragments showed that the 530 bp fragment originated from the 486 bp fragment by an insertion of 44 bp, corresponding to the nucleotides c.1129–5967_c.1129–5924 of intron 10. Subsequent analysis of the corresponding sequence of intron 10 showed that the patient was homozygous for a c.1129–5923C>G mutation, creating a cryptic splice donor site (Fig. 3b). As a consequence, a 44 bp fragment of intron 10 was inserted in the mature DPD mRNA (Fig. 3b). The insertion leads to a shifted reading frame, resulting in a premature stop codon in exon 11.
Fig. 3.
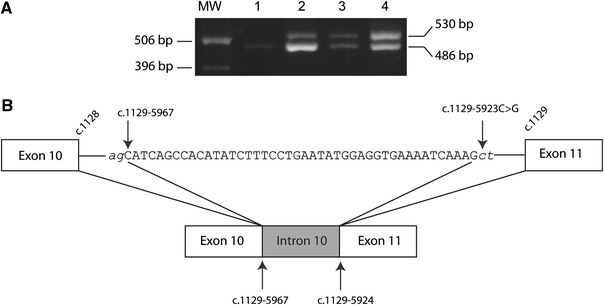
PCR and sequence analysis of exon 11 and flanking regions of DPYD. a The PCR amplification of a cDNA fragment using forward and reverse primers located in exons 8 and 11, respectively. The 486 and 530 bp fragments correspond to the wild-type and mutated cDNA fragments, respectively. MW marker: molecular weight marker; 1 control, 2 patient 3, 3 patient 2, 4 patient 4. b A schematic representation of the region of intron 10 and the effect of the mutation on splicing of the DPD pre-mRNA. The intronic sequence inserted into the DPD mRNA is indicated in capital letters. The cryptic splice acceptor and splice donor sites are indicated in italic
Surprisingly, cDNA analysis of patient 2 also revealed the presence of an exon 22 deletion (c.[2767_2907del]) encoding the amino acids 923–969 (Fig. 4). Quantitative PCR showed that the ratio of the mutant DPD mRNA lacking exon 22 and wild-type DPD mRNA was 25 ± 3%. Sequence analysis of exon 22 and flanking intronic sequences of DPYD of patient 2 did not reveal any mutations.
Fig. 4.
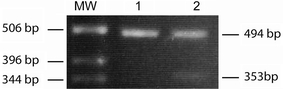
cDNA analysis of exon 22 and flanking regions of DPYD of patient 2. PCR amplification of a cDNA fragment using forward and reverse primers located in exons 21 and 23, respectively. The 494 and 353 bp fragments correspond to the wild-type and mutated cDNA fragments, respectively
Retrospective analysis of our database with previously analysed patients and controls revealed four additional subjects (subjects 3–6) who were heterozygous for the c.1236G>A (p.E412E) mutation (Table 1). Analysis of intron 10 showed that these four subjects were heterozygous for the c.1129–5923C>G mutation with the concomitant insertion of nucleotides c.1129–5967_c.1129–5924 into the DPD mRNA (Fig. 3). Thus, the c.1129–5923C>G mutation and c.1236G>A (p.E412E) mutation are likely to be in cis. Quantitative PCR showed that the ratio of the mutant DPD mRNA, containing the 44 bp fragment of intron 10, and wild-type DPD mRNA was 144 ± 6 and 28 ± 13% in patient 2 and subject 3, respectively.
Table 1.
DPD activity and DPD genotype in patients
| gDNA | cDNA | Effect | Patient 1 (F, 49)a | Patient 2 (F, 50) | Subject 3 (F, 38) | Subject 4 (M, 50)b | Subject 5 (M, 34)c | Subject 6 (F, 62) |
|---|---|---|---|---|---|---|---|---|
| c.85T>C | c.85T>C | p.C29R | + | ++ | + | + | + | |
| c.496A>G | c.496A>G | p.M166 V | + | + | + | |||
| c.483 + 18G > A | − | ++ | + | + | + | + | ||
| c.680 + 139G > A | − | ++ | ++ | ++ | + | + | ||
| c.959–51T > G | − | ++ | + | + | + | + | ||
| c.1129–5923C>G | c.1128_1129ins1129–5967_1129–5924 | Frameshift | ++ | + | + | + | + | |
| c.1138–15T>C | – | + | + | |||||
| c.1236G > A | c.1236G>A | p.E412E | ++ | + | + | + | + | |
| c.1601G>A | c.1601G>A | p.S534N | + | + | ||||
| c.1627A>G | c.1627A>G | p.I534 V | + | + | ||||
| c.1740 + 39C>T | − | + | + | + | ||||
| c.1740 + 40A>G | − | + | ++ | ++ | + | ++ | + | |
| c.1905 + 1G>A | c.1741_1905del | p.581_635del | + | |||||
| c.1974 + 75A>G | − | + | ||||||
| c.2194G>A | c.2194G>A | p.V732I | + | |||||
| c.2300–39G>A | − | + | + | |||||
| c.2621A>G | c.2621A>G | p.K874R | + | |||||
| Unknown | c.2767_2907del | p.923_969del | + | |||||
| g.97302551_97401528deld | c.875_1025del | Truncated protein | + | |||||
| DPD activity (nmol/mg/h)e | 1.2 | 2.9 | 4.2 | 6.2 | n.a. | 1.6 | ||
| Toxicity | IV | III | III | III |
The mutations indicated in bold were part of a haplotype associated with severe 5FU toxicity (Amstutz et al. 2009)
n.a. not available
+, heterozygous; ++, homozygous
aData taken from Gross et al. (2003)
bHealthy volunteer
cSon of a patient experiencing lethal toxicity
dMaximal deleted region
eDPD activity in peripheral blood mononuclear cells (control DPD activity was 9.6 ± 2.6 nmol/mg/h; van Kuilenburg et al. 2007)
Analysis of the c.1236G>A and c.1129–5923C>G mutation in 191 Dutch and 453 German healthy individuals revealed a heterozygote frequency of 2.6 and 3.3%, respectively. In 203 German patients who had been diagnosed for cancer between 2002 and 2009 and assessed for 5FU-associated side effects (Supplemental Table 1), we retrospectively found nine cases carrying the c.1236G>A and c.1129–5923C>G mutations. Six of them had experienced severe grade III/IV 5FU-associated toxicity, mostly neutropenia, leucopenia and diarrhoea. Overall, the deleterious allele was associated with 9.1% (6 of 66) of the individuals with severe toxic effects compared to 2.2% (3 of 137) of the patients who tolerated the chemotherapy regimen (P = 0.033).
Crystal structure analysis
Sequence analysis of the DPD gene of subject 3 also revealed the presence of a novel missense mutation c.2621A>G (p.K874R). Analysis of a three-dimensional model of the human enzyme, based on the crystal structure of the recombinant pig DPD, showed that the K874R site belongs to a stretch of about 85 amino acids (856–941) that connects the barrel domain IV with the ferredoxin-like core of the C-terminal FeS-cluster domain V (Fig. 5). This stretch is least conserved between DPDs from different organisms and K874 itself is in mouse and rat DPD replaced by a glutamine. The point mutation replaces the long, flexible and basic lysine side chain by a longer one of equal charge (Fig. 5). Due to the location of the mutation site on the protein surface, the orientation of the side chain towards the solvent and the limited interaction with surrounding residues, the effect of the amino acid exchange is probably negligible.
Discussion
5-Fluorouracil (5FU) and its oral prodrug capecitabine (Xeloda®) are two of the most frequently prescribed chemotherapeutic drugs for the adjuvant and palliative treatment of patients with cancers of the gastrointestinal tract, breast and head and neck (Meyerhardt and Mayer 2005; Twelves et al. 2005). Although the beneficial effects of 5FU and capecitabine are well established, the toxicity associated with the treatment can be severe (Meyerhardt and Mayer 2005). In this respect, it has been shown that a DPD deficiency is an important pharmacogenetic syndrome and the identification of predictive genetic markers of toxicity has become a challenging topic in the field of oncology (Amstutz et al. 2009; Gross et al. 2008; Morel et al. 2006).
Analysis of the prevalence of the various DPYD mutations, in cancer patients experiencing severe toxicity, showed that the splice-site mutation c.1905 + 1G>A (IVS14 + 1G>A) and the c.2846A>T (D949V) were the most common pathological mutations (Loganayagam et al. 2010; Morel et al. 2006; Schwab et al. 2008; van Kuilenburg 2004). The high prevalence of c.1905 + 1G>A mutations in cancer patients with a reduced DPD activity from the Netherlands is in line with a previous study in tumour patients with severe 5FU toxicity (van Kuilenburg et al. 2002a). Moreover, a relatively high incidence of the splice-site mutation with 1.8% heterozygotes in the normal population has been described (van Kuilenburg et al. 2001).
Previously, the analysis of 68 patients with high-grade 5FU-associated toxicity did not reveal large intragenic rearrangements of DPYD (Ticha et al. 2009). In the present study, unambiguous evidence for such a large genomic deletion within DPYD was obtained in a patient with reduced DPD activity and severe 5FU toxicity. The deleted region, encompassing exons 21–23, was located outside the common fragile site FRA1E which extends over 370 kb within DPYD (Hormozian et al. 2007; van Kuilenburg et al. 2009). Common fragile sites represent chromosome structures that are particular prone to breakage under replication stress and the genomic instability can give rise to deletions, translocations and amplifications (Hormozian et al. 2007). Recently, we showed the presence of large intragenic rearrangements of DPYD and a de novo interstitial deletion del(1)(p13.3p21.3) in five patients suffering from a complete DPD deficiency accompanied by a severe neurological disorder (van Kuilenburg et al. 2009). The deletions involving exons 12 and 14–16 were located within the common fragile site FRA1E (Hormozian et al. 2007; van Kuilenburg et al. 2009). The fact that a genomic deletion affecting DPYD was observed in only 1 out of 92 patients with reduced DPD activity and/or severe 5FU toxicity indicates that rearrangements in DPYD are not a major genetic cause underlying a partial DPD deficiency.
Recent advances in our understanding of pre-mRNA splicing has led to an increased awareness that mutations outside the splice sites may affect pre-mRNA splicing and thereby causing disease (Pagani and Baralle 2004; Wang and Cooper 2007). To date, alternative splicing of DPD pre-mRNA has only been described for mutations in the invariant splice donor sites of exons 11 and 14 (van Kuilenburg et al. 2005; Vreken et al. 1996; Wei et al. 1996). Therefore, a conspicuous finding was a deep intronic mutation affecting DPD pre-mRNA splicing in five individuals. The c.1129–5923C>G mutation in intron 10 created a cryptic splice donor site and as a consequence, a 44 bp fragment of intron 10 was inserted in the mature DPD mRNA. A systematic analysis of the sequences present in the vicinity of the splice junctions has led to consensus sequences for both the splice acceptor and splice donor sites (Shapiro and Senapathy 1987). A scoring system developed by Shapiro and Senapathy showed that most naturally occurring splice acceptor and splice donor sites yielded scores above 0.70. Analysis of the cryptic splice acceptor and splice donor sites, created by the c.1129–5923C>G mutation, showed consensus values of 0.87 and 0.90, respectively. However, the c.1129–5923C>G mutation did not result in the exclusive generation of a mutant transcript as the wild-type mRNA was found still to be present in a patient apparent homozygous for the c.1129–5923C>G mutation.
The c.1129–5923C>G mutation was presently detected in all patients who were also a carrier of the c.1236G>A (p.E412E) mutation and these two genetic mutations are likely to be in cis. Genotype analysis of healthy individuals showed that the frequency of heterozygosity for the c.1236G>A mutation was 2.6, 3.3 and 1.9% in the Dutch, German and Tunisian populations, respectively (Ben et al. 2007). In contrast, the mutation was not reported in 121 Korean and 341 Japanese individuals (Cho et al. 2007; Maekawa et al. 2007). Conflicting data exist as to weather the c.1236G>A mutation is associated with an increased risk of development of severe 5FU-associated toxicity (Kleibl et al. 2009; Schwab et al. 2008). However, a haplotype containing three intronic polymorphisms (c.483 + 18G>A, c.959–51T>G, c.680 + 139G>A) and the c.1236G>A mutation was demonstrated to be associated with severe 5FU toxicity (Amstutz et al. 2009). In our study, all patients who were a carrier of the c.1129–5923C>G and c.1236G>A mutation, also possessed the three intronic mutations c.483 + 18G>A, c.959–51T>G and c.680 + 139G>A. Thus, the increased susceptibility for 5FU toxicity of carriers of this haplotype is most likely due to the presence of the c.1129–5923C>G mutation. This would be in line with our results which showed that the c.1129–5923C>G mutation was significantly enriched in patients suffering from severe 5FU toxicity.
Studies at the mRNA level are rarely performed and are usually considered only when the effect of the mutation on the encoded protein clearly does not explain the phenotype, as in synonymous variations. The present study showed the presence of a genomic deletion affecting DPYD and for the first time, a deep intronic mutation affecting DPD pre-mRNA splicing. Therefore, screening of DPD deficient patients for genomic deletions and aberrant splicing should be considered in case initial sequence analysis does not reveal pathogenic mutations. In particular, the apparently high prevalence of the c.1129–5923C>G mutation in the normal population warrants genetic screening for this mutation in cancer patients prior to the start of 5FU-containing chemotherapy.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
References
- Amstutz U, Farese S, Aebi S, Largiader CR. Dihydropyrimidine dehydrogenase gene variation and severe 5-fluorouracil toxicity: a haplotype assessment. Pharmacogenomics. 2009;10:931–944. doi: 10.2217/pgs.09.28. [DOI] [PubMed] [Google Scholar]
- Ben FR, Gross E, Chouchen L, B’Chir F, Ben AS, Neubauer S, Kiechle M, Saguem S. Mutational spectrum of dihydropyrimidine dehydrogenase gene (DPYD) in the Tunisian population. C R Biol. 2007;330:764–769. doi: 10.1016/j.crvi.2007.08.003. [DOI] [PubMed] [Google Scholar]
- Cho HJ, Park YS, Kang WK, Kim JW, Lee SY. Thymidylate synthase (TYMS) and dihydropyrimidine dehydrogenase (DPYD) polymorphisms in the Korean population for prediction of 5-fluorouracil-associated toxicity. Ther Drug Monit. 2007;29:190–196. doi: 10.1097/FTD.0b013e318040b1fe. [DOI] [PubMed] [Google Scholar]
- Collie-Duguid ESR, Etienne MC, Milano GA, McLeod HL. Known variant DPYD alleles do not explain DPD deficiency in cancer patients. Pharmacogenetics. 2000;10:217–223. doi: 10.1097/00008571-200004000-00002. [DOI] [PubMed] [Google Scholar]
- Dobritzsch D, Schneider G, Schnackerz KD, Lindqvist Y. Crystal structure of dihydropyrimidine dehydrogenase, a major determinant of the pharmacokinetics of the anti-cancer drug 5-fluorouracil. EMBO J. 2001;20:650–660. doi: 10.1093/emboj/20.4.650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gross E, Ullrich T, Seck K, Mueller V, de Wit M, von Schilling C, Meindl A, Schmitt M, Kiechle M. Detailed analysis of five mutations in dihydropyrimidine dehydrogenase detected in cancer patients with 5-fluorouracil-related side effects. Hum Mutat. 2003;22:498. doi: 10.1002/humu.9201. [DOI] [PubMed] [Google Scholar]
- Gross E, Busse B, Riemenschneider M, Neubauer S, Seck K, Klein HG, Kiechle M, Lordick F, Meindl A. Strong association of a common dihydropyrimidine dehydrogenase gene polymorphism with fluoropyrimidine-related toxicity in cancer patients. PLoS One. 2008;3:e4003. doi: 10.1371/journal.pone.0004003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hormozian F, Schmitt JG, Sagulenko E, Schwab M, Savelyeva L. FRA1E common fragile site breaks map within a 370kilobase pair region and disrupt the dihydropyrimidine dehydrogenase gene (DPYD) Cancer Lett. 2007;246:82–91. doi: 10.1016/j.canlet.2006.02.004. [DOI] [PubMed] [Google Scholar]
- Johnson MR, Diasio RB. Importance of dihydropyrimidine dehydrogenase (DPD) deficiency in patients exhibiting toxicity following treatment with 5-fluorouracil. Adv Enzyme Regul. 2001;41:151–157. doi: 10.1016/S0065-2571(00)00011-X. [DOI] [PubMed] [Google Scholar]
- Kleibl Z, Fidlerova J, Kleiblova P, Kormunda S, Bilek M, Bouskova K, Sevcik J, Novotny J. Influence of dihydropyrimidine dehydrogenase gene (DPYD) coding sequence variants on the development of fluoropyrimidine-related toxicity in patients with high-grade toxicity and patients with excellent tolerance of fluoropyrimidine-based chemotherapy. Neoplasma. 2009;56:303–316. doi: 10.4149/neo_2009_04_303. [DOI] [PubMed] [Google Scholar]
- Loganayagam A, renas-Hernandez M, Fairbanks L, Ross P, Sanderson JD, Marinaki AM. The contribution of deleterious DPYD gene sequence variants to fluoropyrimidine toxicity in British cancer patients. Cancer Chemother Pharmacol. 2010;65:403–406. doi: 10.1007/s00280-009-1147-x. [DOI] [PubMed] [Google Scholar]
- Maekawa K, Saeki M, Saito Y, Ozawa S, Kurose K, Kaniwa N, Kawamoto M, Kamatani N, Kato K, Hamaguchi T, Yamada Y, Shirao K, Shimada Y, Muto M, Doi T, Ohtsu A, Yoshida T, Matsumura Y, Saijo N, Sawada J. Genetic variations and haplotype structures of the DPYD gene encoding dihydropyrimidine dehydrogenase in Japanese and their ethnic differences. J Hum Genet. 2007;52:804–819. doi: 10.1007/s10038-007-0186-6. [DOI] [PubMed] [Google Scholar]
- Mattison LK, Ezzeldin H, Carpenter M, Modak A, Johnson MR, Diasio RB. Rapid identification of dihydropyrimidine dehydrogenase deficiency by using a novel 2–13C-uracil breath test. Clin Cancer Res. 2004;10:2652–2658. doi: 10.1158/1078-0432.CCR-03-0374. [DOI] [PubMed] [Google Scholar]
- Meyerhardt JA, Mayer RJ. Systemic therapy for colorectal cancer. N Engl J Med. 2005;352:476–487. doi: 10.1056/NEJMra040958. [DOI] [PubMed] [Google Scholar]
- Milano GA, Etienne MC, Pierrefite V, Barberi-Heyob M, Deporte-Fety R, Renée N. Dihydropyrimidine dehydrogenase deficiency and fluorouracil-related toxicity. Br J Cancer. 1999;79:627–630. doi: 10.1038/sj.bjc.6690098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morel A, Boisdron-Celle M, Fey L, Soulie P, Craipeau MC, Traore S, Gamelin E. Clinical relevance of different dihydropyrimidine dehydrogenase gene single nucleotide polymorphisms on 5-fluorouracil tolerance. Mol Cancer Ther. 2006;5:2895–2904. doi: 10.1158/1535-7163.MCT-06-0327. [DOI] [PubMed] [Google Scholar]
- Pagani F, Baralle FE. Genomic variants in exons and introns: identifying the splicing spoilers. Nat Rev Genet. 2004;5:389–396. doi: 10.1038/nrg1327. [DOI] [PubMed] [Google Scholar]
- Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res. 2002;30:e57. doi: 10.1093/nar/gnf056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwab M, Zanger UM, Marx C, Schaeffeler E, Klein K, Dippon J, Kerb R, Blievernicht J, Fischer J, Hofmann U, Bokemeyer C, Eichelbaum M. Role of genetic and nongenetic factors for fluorouracil treatment-related severe toxicity: a prospective clinical trial by the German 5-FU Toxicity Study Group. J Clin Oncol. 2008;26:2131–2138. doi: 10.1200/JCO.2006.10.4182. [DOI] [PubMed] [Google Scholar]
- Seck K, Riemer S, Kates R, Ullrich T, Lutz V, Harbeck N, Schmitt M, Kiechle M, Diasio R, Gross E. Analysis of the DPYD gene implicated in 5-fluorouracil catabolism in a cohort of Caucasian individuals. Clin Cancer Res. 2005;11:5886–5892. doi: 10.1158/1078-0432.CCR-04-1784. [DOI] [PubMed] [Google Scholar]
- Shapiro MB, Senapathy P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 1987;15:7155–7174. doi: 10.1093/nar/15.17.7155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ticha I, Kleiblova P, Fidlerova J, Novotny J, Pohlreich P, Kleibl Z. Lack of large intragenic rearrangements in dihydropyrimidine dehydrogenase (DPYD) gene in fluoropyrimidine-treated patients with high-grade toxicity. Cancer Chemother Pharmacol. 2009;64:615–618. doi: 10.1007/s00280-009-0970-4. [DOI] [PubMed] [Google Scholar]
- Twelves C, Wong A, Nowacki MP, Abt M, Burris H, III, Carrato A, Cassidy J, Cervantes A, Fagerberg J, Georgoulias V, Husseini F, Jodrell D, Koralewski P, Kroning H, Maroun J, Marschner N, McKendrick J, Pawlicki M, Rosso R, Schuller J, Seitz JF, Stabuc B, Tujakowski J, Van HG, Zaluski J, Scheithauer W. Capecitabine as adjuvant treatment for stage III colon cancer. N Engl J Med. 2005;352:2696–2704. doi: 10.1056/NEJMoa043116. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP. Dihydropyrimidine dehydrogenase and the efficacy and toxicity of 5-fluorouracil. Eur J Cancer. 2004;40:939–950. doi: 10.1016/j.ejca.2003.12.004. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP, Haasjes J, Richel DJ, Zoetekouw L, van Lenthe H, De Abreu RA, Maring JG, Vreken P, van Gennip AH. Clinical implications of dihydropyrimidine dehydrogenase (DPD) deficiency in patients with severe 5-fluorouracil-associated toxicity: identification of new mutations in the DPD gene. Clin Cancer Res. 2000;6:4705–4712. [PubMed] [Google Scholar]
- van Kuilenburg ABP, Muller EW, Haasjes J, Meinsma JR, Zoetekouw L, Waterham HR, Baas F, Richel DJ, van Gennip AH. Lethal outcome of a patient with a complete dihydropyrimidine dehydrogenase (DPD) deficiency after administration of 5-fluorouracil: frequency of the common IVS14 + 1G > A mutation causing DPD deficiency. Clin Cancer Res. 2001;7:1149–1153. [PubMed] [Google Scholar]
- van Kuilenburg ABP, Meinsma JR, Zoetekouw L, van Gennip AH. High prevalence of the IVS14 + 1G > A mutation in the dihydropyrimidine dehydrogenase gene of patients with severe 5-fluorouracil-associated toxicity. Pharmacogenetics. 2002;12:555–558. doi: 10.1097/00008571-200210000-00007. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP, Meinsma JR, Zoetekouw L, van Gennip AH. Increased risk of grade IV neutropenia after administration of 5-fluorouracil due to a dihydropyrimidine dehydrogenase deficiency: high prevalence of the IVS14 + 1 g > a mutation. Int J Cancer. 2002;101:253–258. doi: 10.1002/ijc.10599. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP, Meinsma R, Beke E, Bobba B, Boffi P, Enns GM, Witt DR, Dobritzsch D. Identification of three novel mutations in the dihydropyrimidine dehydrogenase gene associated with altered pre-mRNA splicing or protein function. Biol Chem. 2005;386:319–324. doi: 10.1515/BC.2005.038. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP, Klumpen HJ, Westermann AM, Zoetekouw L, van LH, Bakker PJ, Richel DJ, Guchelaar HJ. Increased dihydropyrimidine dehydrogenase activity associated with mild toxicity in patients treated with 5-fluorouracil and leucovorin. Eur J Cancer. 2007;43:459–465. doi: 10.1016/j.ejca.2006.09.017. [DOI] [PubMed] [Google Scholar]
- van Kuilenburg ABP, Meijer J, Mul ANPM, Hennekam RCM, Hoovers JMN, de Die-Smulders CEM, Weber P, Mori AC, Bierau J, Fowler B, Macke K, Sass JO, Meinsma R, Hennermann JB, Miny P, Zoetekouw L, Vijzelaar R, Nicolai J, Ylstra B, Rubio-Gozalbo ME. Analysis of severely affected patients with dihydropyrimidine dehydrogenase deficiency reveals large intragenic rearrangements of DPYD and a de novo interstitial deletion del(1)(p13.3p21.3) Hum Genet. 2009;125:581–590. doi: 10.1007/s00439-009-0653-6. [DOI] [PubMed] [Google Scholar]
- Vreken P, van Kuilenburg ABP, Meinsma JR, Smit GPA, Bakker HD, De Abreu RA, van Gennip AH. A point mutation in an invariant splice donor site leads to exon skipping in two unrelated Dutch patients with dihydropyrimidine dehydrogenase deficiency. J Inherit Metab Dis. 1996;19:645–654. doi: 10.1007/BF01799841. [DOI] [PubMed] [Google Scholar]
- Wang GS, Cooper TA. Splicing in disease: disruption of the splicing code and the decoding machinery. Nat Rev Genet. 2007;8:749–761. doi: 10.1038/nrg2164. [DOI] [PubMed] [Google Scholar]
- Wei X, McLeod HL, McMurrough J, Gonzalez FJ. Molecular basis of the human dihydropyrimidine dehydrogenase deficiency and 5-fluorouracil toxicity. J Clin Invest. 1996;98:610–615. doi: 10.1172/JCI118830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei X, Elizondo G, Sapone A, McLeod HL, Raunio H, Gonzalez FJ. Characterization of the human dihydropyrimidine dehydrogenase gene. Genomics. 1998;51:391–400. doi: 10.1006/geno.1998.5379. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


