Abstract
Trehalose 6-phosphate (T6P), the precursor of trehalose, is a signaling molecule in plants with strong effects on metabolism, growth and development. We recently showed that in growing tissues T6P is an inhibitor of SnRK1 of the SNF1-related group of protein kinases.1 SnRK1 acts as transcriptional integrator in response to carbon and energy supply. In microarray experiments on seedlings of transgenic Arabidopsis with elevated T6P content we found that expression of SnRK1 marker genes was affected in a manner to be predicted by inhibition of SnRK1 by T6P in vivo.1 A large number of genes involved in reactions that utilize carbon, e.g., UDP-glucose dehydrogenase genes involved in cell wall synthesis, were upregulated. T6P was also found to affect developmental signaling pathways, probably in a SnRK1-independent manner. This includes upregulation of genes encoding UDP-glycosyltransferases that are involved in the glycosylation of hormones. In addition, genes involved in auxin response and light signaling were affected. Many of these genes belong to pathways that link the circadian clock to plant growth and development. The overall pattern of changes in gene expression supports a role for T6P in coordinating carbon supply with biosynthetic process involved in growth and development.
Key words: Arabidopsis, trehalose 6-phosphate, SnRK1, UDP-glucose, cell wall
With the exception of vertebrates trehalose synthesis occurs in all organisms. The chemistry of trehalose as a non-reducing disaccharide suits a role as a carbon source and protectant molecule, a function which trehalose performs in invertebrates, fungi, prokaryotes and archaea.2 Sucrose, the other naturally occurring non-reducing disaccharide, performs this function in plants. Nevertheless, plants still synthesize trehalose, but in low micromolar concentrations. A new distinct, previously unknown function of the pathway appears to have developed early in plant evolution.3 Clues to this function have been provided in mutant and transgenic plants, which show an indispensable role for the pathway in the utilization of carbon and in growth and development.4 Until recently a mechanistic basis to explain this function was not known.
Inhibition of SnRK1 by Trehalose 6-Phosphate Signals Carbon Availability
We recently showed that trehalose 6-phosphate (T6P) inhibits SnRK1 (SNF1-related protein kinase1) of the family of calcium-independent serine/threonine protein kinases that includes AMPK of mammals and SNF1 of yeast.1 These conserved kinases perform a fundamental role in transcriptional, metabolic and developmental regulation in response to energy limitation and starvation of carbon source.5 In Arabidopsis, SnRK1 has recently been found to affect the expression of a number of genes involved in metabolism, growth and development, in a role more extensive than previously appreciated.6 We recently demonstrated that in transgenic plants expressing an E. coli T6P synthase gene (TPS, otsA) to elevate T6P, 97% of the SnRK1 marker genes that showed a statistically significant change compared to wild type changed in a manner to be predicted by inhibition of SnRK1 in vivo. Analysis of microarray data revealed regulation of metabolic pathways and a general upregulation of biosynthetic processes by T6P.1 T6P levels have been shown to reflect sucrose content7 (Fig. 1) and to promote seedling growth on sucrose.8 These data fit a model where T6P content reflects sucrose supply and suggests that inhibition of SnRK1 by T6P enables the plant to activate processes that utilize carbon in growth and development.
Figure 1.
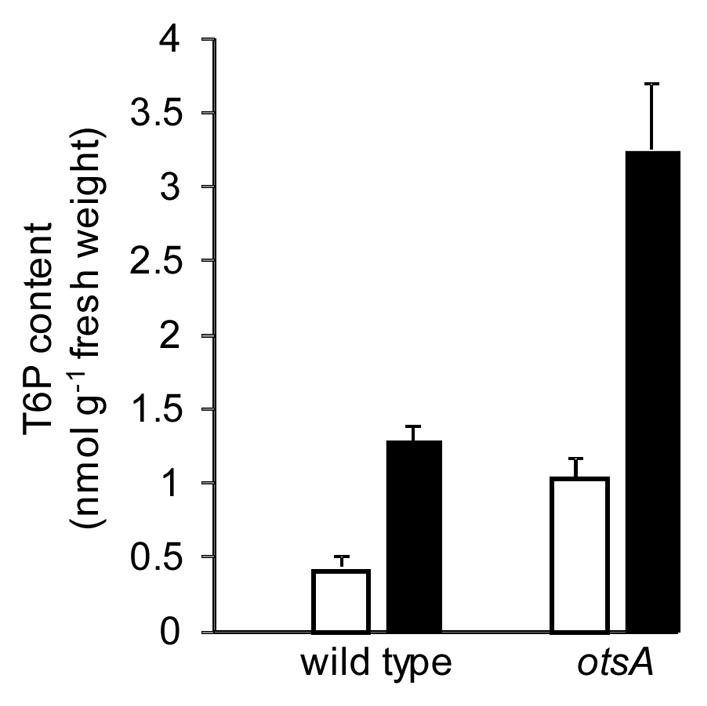
T6P content in 7-day-old wild-type and otsA-expressing seedlings grown in half-strength MS medium as in1 with 15 mM (□) or 100 mM (▪) sucrose. n = 4 with standard error of the mean.
Stimulation of UDP-Glucose Metabolism Links Trehalose 6-Phosphate to Cell Wall Biosynthesis and Growth
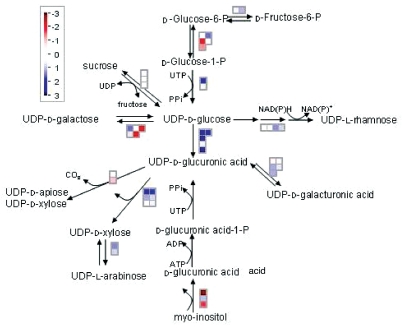
Statistical analysis using the Wilcoxon test in MapMan9 demonstrated differential regulation of cell wall biosynthesis in otsA-expressing seedlings. There were strong effects on genes encoding enzymes that utilize UDP-glucose (UDPG). UDPG is an important intermediate in plants as a direct substrate for cell wall and sucrose synthesis. In growing regions of plants that are receiving sucrose from leaves a major sink for UDPG is cell wall synthesis. The cell wall of Arabidopsis consists of large amounts of hemicelluloses which contain xylose and galacturonic acid in particular, and pectin which consists of α-1,4-linked galacturonic acid units.10 These are predominantly synthesized from UDP-glucuronic acid, from which it is estimated that in Arabidopsis 50% of cell wall biomass is derived.10 We found that four UDPG dehydrogenases (UDPG-DH) genes which form UDP-glucuronic acid from UDPG were upregulated by T6P (Table 1, Fig. 2). These enzymes are tightly correlated with cell division and growth.10 In trees they are used as a marker enzyme for cambial meristem. Three UDP-glucuronic acid decarboxylases which form xylose from UDP-glucuronic acid and a UDP-xylose epimerase which forms xylose from arabinose were also upregulated, together with two glucuronic acid epimerases which catalyze formation of UDP-galacturonic acid and a rhamnose synthase. In contrast, genes encoding enzymes that catalyze flux into mannose sugars were downregulated. There were small changes in cellulose synthases. A large number of changes were found for the cell-wall-modifying xyloglucan xyloglucosyl transferases (XET) which affect cell wall strength and flexibility. These changes are consistent with those observed in tps1 mutant embryos deficient in T6P where there was strong downregulation of genes encoding enzymes of cell wall biosynthesis.11 Interestingly, transcripts of ten pectin methylesterase/invertase inhibitor family protein genes were downregulated. The regulation of pectin methylesterases by these inhibitor proteins would potentially affect the degree of pectin demethylesterification as these enzymes catalyze the demethylesterification of the homogalacturonan component of pectins in cell walls. The degree of pectin demethylesterification affects the solidity of the cell wall which is important in growth and development; particularly physiological processes that require rearrangement of cell wall architecture such as root development and stem elongation.12
Table 1.
Genes with altered transcript abundance compared to wild type in transgenic arabidopsis seedlings expressing the E. coli trehalose phosphate synthase gene otsA and containing elevated trehalose 6-phosphate
| Gene code | Up | Gene code | Down | |
| Utilisation of UDPG for cell wall synthesis | ||||
| UDP-glucose 6-dehydrogenase | At3g29360 | 6.16 | ||
| At5g15490 | 6.11 | |||
| At3g01010 | 5.07 | |||
| At5g39320 | 3.77 | |||
| UDP-glucuronic acid decarboxylase (xylose synthase) | At2g28760 | 4.16 | ||
| At3g62830 | 3.44 | |||
| At3g46440 | 2.26 | |||
| UDP-D-xylose 4-epimerase | At1g30620 | 2.64 | ||
| UDP-glucuronic acid 4-epimerase | At4g12250 | 2.24 | ||
| At4g00110 | 2.22 | |||
| Rhamnose synthase | At1g53500 | 2.52 | ||
| UDP-glucose 4-epimerase | At4g10960 | 2.92 | At1g12780 | 0.19 |
| At1g63180 | 0.24 | |||
| Phosphomannose isomerase | At1g67070 | 0.14 | ||
| GDP-D-mannose 4,6-dehydratase | At5g66280 | 0.23 | ||
| GDP-D-mannose 3,5-epimerase | At5g28840 | 0.49 | ||
| Cellulose synthase | At1g23480 | 0.37 | ||
| At1g24070 | 2.29 | At2g32540 | 0.39 | |
| At2g25540 | 2.26 | At2g35650 | 0.47 | |
| At2g32530 | 0.49 | |||
| Other cell wall | ||||
| Xyloglucan:xyloglucosyl transferase | At4g25820 | 10.7 | ||
| At4g30280 | 8.15 | |||
| At4g30290 | 7.22 | At5g65730 | 0.07 | |
| At1g65310 | 6.72 | At5g57550 | 0.17 | |
| At4g25810 | 4.96 | At4g03210 | 0.40 | |
| At4g13090 | 4.30 | At4g14130 | 0.44 | |
| At5g57560 | 3.8* | |||
| At5g57540 | 2.18 | |||
| Invertase/pectin methylesterase inhibitor family protein | At5g20740 | 2.41 | At1g14890 | 0.30 |
| At3g47380 | 0.31 | |||
| At1g47960 | 0.34 | |||
| At5g64620 | 0.37 | |||
| At5g62350 | 0.38 | |||
| At5g46980 | 0.43 | |||
| At4g25260 | 0.43 | |||
| At1g70720 | 0.47 | |||
| At4g12390 | 0.48 | |||
| At5g24370 | 0.49 | |||
| Hormone glycosylation | ||||
| IAA/ABA | At1g05680 | 10.4 3.13 | ||
| At2g23260 | ||||
| Flavonol/flavonoid/anthocyanin | At2g36790 | 14.7 | At5g65550 | 0.16 |
| At1g06000 | 3.90 | At4g15260 | 0.25 | |
| At1g30530 | 0.43 | |||
| Zeatin (cytokinin) | At2g36750 | 5.24 | At5g05860 | 0.44 |
| At1g22400 | 2.10 | |||
| Zeatin (brassinosteroid) | At2g36800 | 3.07 | ||
| ABA | At2g31750 | 0.31 | ||
| Sterol | At3g07020 | 2.98 | ||
| Glucosinolate | At1g24100 | 2.60 | ||
| Light signaling | At1g49130 | 0.08 | ||
| At1g75540 STH2 | 0.10 | |||
| At3g19850 | 0.10 | |||
| At5g52250 | 0.12 | |||
| At3g26740 CCL | 0.12 | |||
| At3g02380 COL2 | 0.12 | |||
| At1g18330 EPR1 | 0.12 | |||
| At5g37260 CIR1 | 0.13 | |||
| At3g45780 NPH1 | 0.14 | |||
| At5g17300 | 0.15 | |||
| At3g15570 | 0.16 | |||
| At1g14280 | 0.17 | |||
| At3g09150 | 0.21 | |||
| At5g66560 | 0.22 | |||
| At1g67900 | 0.25 | |||
| At5g67385 | 0.26 | |||
| At2g43010 PIF4 | 0.27 | |||
| At3g50840 | 0.27 | |||
| At3g22104 | 0.27 | |||
| At2g02950 | 0.28 | |||
| At4g31820 ENP | 0.29 | |||
| At1g18810 | 0.37 | |||
| At4g37590 NPY5 | 0.42 | |||
| Auxin response | At4g27450 | 0.07 | ||
| At5g18020 | 0.10 | |||
| At5g27780 | 0.11 | |||
| At4g34800 | 0.11 | |||
| At4g38860 | 0.11 | |||
| At2g21210 | 0.14 | |||
| At3g15450 | 0.14 | |||
| At1g29440 | 0.16 | |||
| At1g29490 | 0.16 | |||
| At3g25290 | 4.44 | At1g29500 | 0.16 | |
| At1g59500 GH3.4 | 3.93 | At5g18080 | 0.16 | |
| At5g13320 GDG1 | 3.04 | At1g52830 IAA6 | 0.18 | |
| At5g53590 | 2.95* | At1g43040 | 0.23 | |
| At1g17345 | 2.58 | At2g22670 IAA8 | 0.25 | |
| At5g50760 | 2.40 | At4g12980 | 0.25 | |
| At3g09870 | 2.37 | At4g38840 | 0.27 | |
| At3g15540 IAA19 | 2.29 | At2g22670 | 0.29 | |
| At5g54510 DFL1 | 2.22 | At5g18050 | 0.32 | |
| At3g07390 AIR12 | 2.21 | At5g18060 | 0.35 | |
| At1g72430 | 2.06 | At5g66260 | 0.38 | |
| At1g29430 | 0.38 | |||
| At2g01200 | 0.40 | |||
| AT3G04730IAA16 | 0.41 | |||
| At3g23050 | 0.41 | |||
| At2g28350 | 0.42 | |||
| At4g34790 | 0.43 | |||
| At5g57420 | 0.48 | |||
| At2g16580 | 0.49 | |||
| At5g18010 | 0.49 | |||
| Auxin carrier | At2g17500 | 2.12 | At1g70940 | 0.19 |
| At5g57090 AGR1 | 2.11 | At1g76520 | 0.25 | |
| At2g01420 | 0.29 | |||
| Dormancy/auxin associated family protein | At2g33830 | 0.14 | ||
| At1g28330 | 0.14 | |||
| At1g56220 | 0.15 | |||
| At2g33830 | 0.21 | |||
| At1g56220 | 0.28 | |||
| At2g33830 | 0.30 | |||
denotes a SnRK1 marker gene that changes in a direction opposite to that that would be predicted from inhibition of SnRK1 by T6P. Numbers in bold are SnRK1 marker genes according to Baena-Gonzalez et al. 2007.1
Figure 2.
Synthesis of cell wall precursors from UD P-glucose. Changes in gene expression were analyzed using Mapman9 for Arabidopsis seedlings expressing the E. coli trehalose phosphate synthase gene otsA and containing elevated trehalose 6-phosphate. Blue denotes upregulation, red denotes downregulation.
Expression of UDP glycosyltransferase genes thought to be involved in the glycosylation of hormones, including auxin, was induced (Table 1). Since glycosylation modifies the action of hormones,13 this provides a possible means through which carbon availability can be linked to hormone signaling.
Light and Auxin Signaling are Involved in the Growth Response
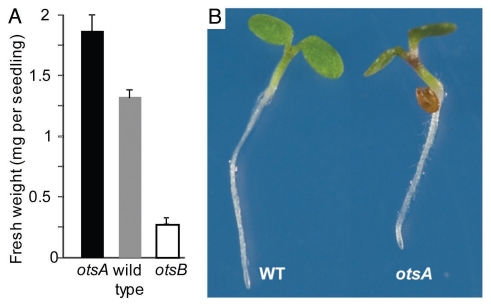
In agreement with enhanced cell wall biosynthesis, seedling weight was increased in otsA-expressing Arabidopsis with high T6P content, especially in response to sucrose supply8 (Fig. 3A). Hypocotyls were thickened, most strongly around the base, and often shorter. In addition, otsA-expressing seedlings had epinastic cotyledons with thicker petioles and accumulated anthocyanins (Fig. 3B). This phenotype is indicative of altered light signaling (Fig. 3B). It is thus not surprising that the expression of genes involved in light signaling and circadian rhythms was altered in seedlings with increased T6P (Table 1; Fig. 4). Statistical analysis using MapMan indicated that, in addition to genes with a role in light signaling, genes involved in auxin response were differentially expressed. For example, expression of the growth-promoting transcription factor gene Phytochrome Interacting Factor 4 (PIF4) was downregulated. PIF4 plays an important role in hypocotyl elongation in the shade avoidance response and during circadian growth.14 PIF gene expression is regulated by the circadian clock15 while PIF proteins are inactivated by 26S proteasome-mediated degradation in response to light. Gene expression of the 26S proteasome, in particular of genes for AAA-type ATPases, was upregulated, which is in agreement with the high-light phenotype of otsA. PIF4 may also be involved in auxin signaling as shown by the lack of induction of the auxin-inducible IAA29 transcript in response to high temperature in a pif4 mutant.16
Figure 3.
(A) Fresh weight of seedlings of otsA-expressing plants with increased T6P (▪), wild type (▪) and otsB-expressing plants with decreased T6P (□) grown on 100 mM sucrose. (B) Phenotype of otsA seedlings.
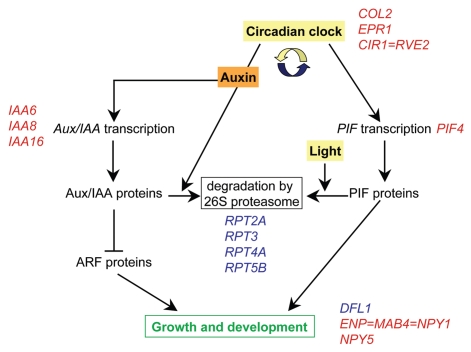
Figure 4.
Model for the interactions of light signaling, circadian rhythms and auxin-regulated growth in response to T6P. The circadian clock regulates transcript levels of the growth regulator PIF4, while PIF4 protein is degraded by the 26S proteasome in response to light. Auxin induces the expression of Aux/IAA proteins as well as promoting their degradation by the 26S proteasome in a process gated by the circadian clock. Aux/IAA proteins regulate gene expression by heterodimerizing with auxin response factor (ARF) proteins. DFL1, ENP = MAB4 = NPY1 (At4g31820) and NPY5 are possible downstream targets that are involved in plant development in response to auxin as well as to light. Examples of genes whose expression was affected in otsA-expressing seedlings with increased T6P are listed in blue (upregulated) or red (downregulated). See Table 1 for list of genes.
In addition to PIF proteins, auxin is considered to provide a link between the circadian clock and growth.17,18 Auxin effects on growth are complex, and, in addition to auxin-dependent growth being gated by the circadian clock, it depends on the developmental stage of a seedling whether auxin inhibits or promotes hypocotyl elongation.18 Expression of Aux/IAA genes was downregulated in seedlings with increased T6P (Fig. 3). Aux/IAA proteins are transcriptional regulators that heterodimerize with auxin response factor (ARF) proteins, which, in turn, function as transcriptional activators or repressors. Auxin promotes the degradation of Aux/IAA proteins by enhancing binding to TIR1, ubiquitination and degradation by the 26S proteasome19 in a process gated by the circadian clock.18 While expression of genes belonging to the 26S proteasome was induced, TIR1 expression was downregulated in otsA-expressing plants. Although the expression of central clock genes was not altered, the expression of transcription factors genes that are involved in circadian clock responses (COL2, EPR1 and CIR1 = RVE2) was downregulated. Interestingly, the expression of genes involved in developmental processes in response to auxin as well as light (DFL1, ENP = MAB4 = NPY1 and NPY5) was also affected in otsA-expressing plants. Such genes could thus provide a link between T6P and plant development in a pathway mediated by auxin and light. Overall, these auxin and light-related changes in gene expression appear to be independent of SnRK1.
Conclusion
Our data show extensive effects of T6P on gene expression in Arabidopsis seedlings. Many of these effects can be explained through a direct impact on SnRK1, while others are probably secondary effects of altered growth or SnRK1-independent. Our data support a role for T6P in inhibiting the starvation response mediated by SnRK1 and in the upregulation of genes involved in biosynthetic processes. Particularly significant for the regulation of growth in Arabidopsis seedlings are effects on genes encoding enzymes of cell wall synthesis, which accounts for a major portion of carbon utilization. There were also effects on the expression of genes involved in auxin and light signaling pathways that are implicated in plant growth and development. Overall these data suggest that T6P regulates growth in relation to sucrose supply by regulating biosynthetic reactions and through regulating hormone signaling either directly or indirectly.
Acknowledgements
Rothamsted Research receives grant-aided support from the Biotechnological and Biological Sciences Research Council (BBSRC) of the United Kingdom. Research was supported by BBSRC grants BB/C51257X/1, BB/D006112/1 and BB/C512645/1.
Footnotes
Previously published online: www.landesbioscience.com/journals/psb/article/10792
References
- 1.Zhang Y, Primavesi LF, Jhurreea D, Andralojc PJ, Mitchell RAC, Powers SJ, et al. Inhibition of Snf1-related protein kinase (SnRK1) activity and regulation of metabolic pathways by trehalose 6-phosphate. Plant Physiol. 2009;149:1860–1871. doi: 10.1104/pp.108.133934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Paul MJ, Primavesi LF, Jhurreea D, Zhang Y. Trehalose metabolism and signalling. Annu Rev Plant Biol. 2008;59:417–441. doi: 10.1146/annurev.arplant.59.032607.092945. [DOI] [PubMed] [Google Scholar]
- 3.Avonce N, Wuyts J, Verschooten K, Vandesteene L, Van Dijck P. The Cytophaga hutchinsonii ChTPSP: first characterized bifunctional TPS-TPP protein as putative ancestor of all eukaryotic biosynthesis proteins. Mol Biol Evol. 2009:359–369. doi: 10.1093/molbev/msp241. [DOI] [PubMed] [Google Scholar]
- 4.Paul M. Trehalose 6-phosphate. Curr Opin Plant Biol. 2007;10:303–309. doi: 10.1016/j.pbi.2007.04.001. [DOI] [PubMed] [Google Scholar]
- 5.Hardie DG. AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat Rev Mol Cell Biol. 2007;8:774–785. doi: 10.1038/nrm2249. [DOI] [PubMed] [Google Scholar]
- 6.Baena-Gonzalez E, Rolland F, Thevelein JM, Sheen J. A central integrator of transcription networks in plant stress and energy signalling. Nature. 2007;448:938–943. doi: 10.1038/nature06069. [DOI] [PubMed] [Google Scholar]
- 7.Lunn JE, Feil R, Hendriks JHM, Gibon Y, Morcuende R, Osuna D, et al. Sugar-induced increases in trehalose 6-phosphate are correlated with redox activation of ADP-glucose pyrophosphorylase and higher rates of starch synthesis in Arabidospis thaliana. Biochem J. 2006;397:139–148. doi: 10.1042/BJ20060083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Schluepmann H, Pellny T, van Dijken A, Smeekens S, Paul M. Trehalose 6-phosphate is indispensable for carbohydrate utilisation and growth in Arabidopsis thaliana. Proc Natl Acad Sci USA. 2003;100:6849–6854. doi: 10.1073/pnas.1132018100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Usadel B, Nagel A, Thimm O, Redestig H, Blaesing OE, Palacios-Rojas N, et al. Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of co-responding genes and comparison with known responses. Plant Physiol. 2005;138:1195–1204. doi: 10.1104/pp.105.060459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Klinghammer M, Tenhaken R. Genome-wide analysis of the UDP-glucose dehydrogenase gene family in Arabidopsis, a key enzyme for matrix polysaccharides in cell walls. J Exp Bot. 2007;58:3609–3621. doi: 10.1093/jxb/erm209. [DOI] [PubMed] [Google Scholar]
- 11.Gomez LD, Baud S, Gilday A, Graham IA. Delayed embryo development in the Arabidopsis trehalose 6-phosphate synthase 1 mutant is associated with altered cell wall structure, decreased cell division and starch accumulation. Plant J. 2006;46:69–84. doi: 10.1111/j.1365-313X.2006.02662.x. [DOI] [PubMed] [Google Scholar]
- 12.Hothorn M, Wolf S, Aloy P, Greiner S, Scheffzek K. Structural insights into the target specificity of plant invertase and pectin methylesterase inhibitory proteins. Plant Cell. 2004;16:3437–3447. doi: 10.1105/tpc.104.025684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lim E-K, Bowles DJ. A class of plant glycosyltransferases involved in cellular homeostasis. EMBO J. 2004;23:2915–2922. doi: 10.1038/sj.emboj.7600295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Franklin KL. Shade avoidance. New Phytol. 2008;179:930–944. doi: 10.1111/j.1469-8137.2008.02507.x. [DOI] [PubMed] [Google Scholar]
- 15.Nozue K, Covington MF, Duek PD, Lorrain S, Fankhauser C, Harmer SL, Maloof JN. Rhythmic growth explained by coincidence between internal and external clues. Nature. 2007;448:358–363. doi: 10.1038/nature05946. [DOI] [PubMed] [Google Scholar]
- 16.Koini MA, Alvey L, Allen T, Tilley CA, Harberd NP, Whitelam GC, Franklin KA. High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Current Biol. 2009;19:408–413. doi: 10.1016/j.cub.2009.01.046. [DOI] [PubMed] [Google Scholar]
- 17.Albadí D, Bláquez M. Molecular interactions between light and hormone signaling to control plant growth. Plant Mol Biol. 2009;69:409–417. doi: 10.1007/s11103-008-9400-y. [DOI] [PubMed] [Google Scholar]
- 18.Covington MF, Harmer SL. The circadian clock regulates auxin signaling and responses in Arabidopsis. PLoS Biol. 2007;5:1773–1784. doi: 10.1371/journal.pbio.0050222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Dos Santos Maraschin F, Memelink J, Offringa R. Auxin-induced, SCFTIR1-mediated poly-ubiquitination marks of AUX/IAA proteins for degradation. Plant J. 2009;59:100–109. doi: 10.1111/j.1365-313X.2009.03854.x. [DOI] [PubMed] [Google Scholar]