Abstract
The title compound, C15H11N3, crystallizes with two independent molecules in the asymmetric unit. The dihedral angles between the phenyl and pyridine rings in each molecule are 53.48 (5) and 50.80 (5)°. In the crystal structure, weak intermolecular C—H⋯N hydrogen bonds connect molecules into one-dimensional chains. In addition, the crystal structure is stabilized by weak C—H⋯π(arene) interactions.
Related literature
For related literature, see: Fang et al. (2002 ▶, 2007 ▶); Medlycott & Hanan (2005 ▶, 2006 ▶); Spek (2003 ▶).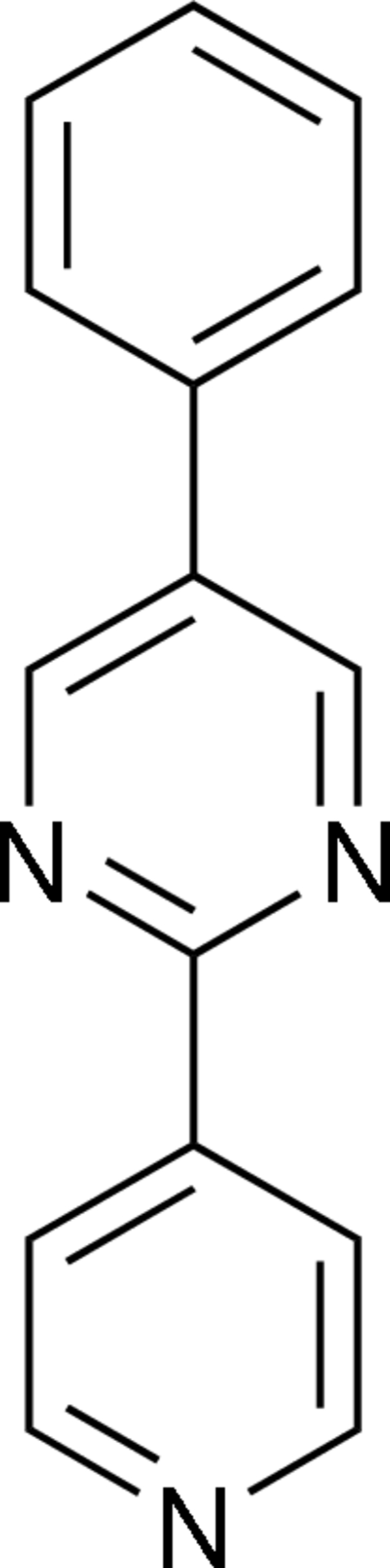
Experimental
Crystal data
C15H11N3
M r = 233.27
Triclinic,
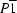
a = 9.2813 (5) Å
b = 9.3609 (5) Å
c = 13.9001 (7) Å
α = 71.462 (2)°
β = 86.957 (2)°
γ = 75.788 (3)°
V = 1109.54 (10) Å3
Z = 4
Cu Kα radiation
μ = 0.68 mm−1
T = 100 (2) K
0.40 × 0.38 × 0.08 mm
Data collection
Bruker SMART 6000 diffractometer
Absorption correction: multi-scan (SADABS; Sheldrick, 1996 ▶) T min = 0.734, T max = 0.947
15167 measured reflections
3967 independent reflections
3226 reflections with I > 2σ(I)
R int = 0.046
Refinement
R[F 2 > 2σ(F 2)] = 0.044
wR(F 2) = 0.139
S = 1.00
3967 reflections
325 parameters
H-atom parameters constrained
Δρmax = 0.27 e Å−3
Δρmin = −0.30 e Å−3
Data collection: SMART (Bruker, 2003 ▶); cell refinement: SAINT (Bruker, 1999 ▶); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: SHELXTL (Sheldrick, 2008 ▶); software used to prepare material for publication: UdMX (local program).
Supplementary Material
Crystal structure: contains datablocks I, global. DOI: 10.1107/S160053680800398X/lh2593sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S160053680800398X/lh2593Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| C7—H7⋯N5i | 0.95 | 2.55 | 3.3818 (18) | 147 |
| C9—H9⋯N6ii | 0.95 | 2.56 | 3.4027 (18) | 148 |
| C22—H22⋯N2i | 0.95 | 2.54 | 3.3784 (18) | 147 |
| C24—H24⋯N3ii | 0.95 | 2.56 | 3.4049 (19) | 149 |
| C1—H1⋯Cg3iii | 0.95 | 2.91 | 3.5925 (15) | 129 |
| C4—H4⋯Cg3 | 0.95 | 2.72 | 3.4163 (15) | 130 |
| C12—H12⋯Cg1i | 0.95 | 2.89 | 3.5844 (15) | 131 |
| C15—H15⋯Cg1iv | 0.95 | 2.92 | 3.5202 (15) | 122 |
| C17—H17⋯Cg6v | 0.95 | 2.84 | 3.5561 (16) | 133 |
| C20—H20⋯Cg6ii | 0.95 | 2.85 | 3.5251 (15) | 129 |
| C26—H26⋯Cg5 | 0.95 | 2.86 | 3.5242 (15) | 128 |
| C29—H29⋯Cg5vi | 0.95 | 2.77 | 3.4451 (15) | 129 |
Symmetry codes: (i) 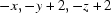 ; (ii)
; (ii) 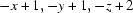 ; (iii)
; (iii) 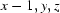 ; (iv)
; (iv) 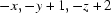 ; (v)
; (v) 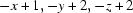 ; (vi)
; (vi) 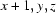 . Cg1 is the centroid of the N1/C1-C5 ring, Cg3 is the centroid of the N4/C16–C20 ring, Cg5 is the centroid of the C10–C15 ring and Cg6 is the centroid of the C25–C30 ring.
. Cg1 is the centroid of the N1/C1-C5 ring, Cg3 is the centroid of the N4/C16–C20 ring, Cg5 is the centroid of the C10–C15 ring and Cg6 is the centroid of the C25–C30 ring.
Acknowledgments
The authors are grateful to the Natural Sciences and Engineering Research Council of Canada, the Ministère de l’Education du Québec and the Université de Montréal for financial support. The authors gratefully acknowledge Mme Françine Bélanger-Gariépy (Laboratoire de diffraction des rayons X, Université de Montréal, Canada) for the teaching of crystallography to MPS. Yuan-Qing Fang is acknowledged for help and guidance with the synthesis.
supplementary crystallographic information
Comment
Ruthenium polypyridyl complexes have long attracted attention due to their exceptional photophysical properties which makes them suitable as chromophores in light-harvesting devices (Medlycott & Hanan, 2005, 2006). Recently, we have reported new pyrimidine-substituted terpyridine ligands and their Ru(II) polypyridyl complexes (Fang et al., 2002, 2007). Introduction of the pyrimidine motif on a terpyridine unit leads to planarization of the system through hydrogen bonds, thus extending pi-delocalization in the acceptor ligand of the metal-to-ligand charge transfer (MLCT) emitting excited-states, which improves the photophysical properties of the complexes. The title compound C15H11N3 (3) [see Fig. 3] was designed for the enhanced π-acceptor character of pyridyl-type ligands for coordination and supramolecular chemistry.
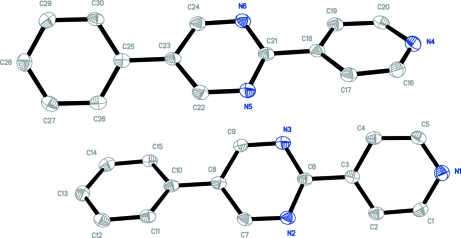
The title compound crystallizes with two molecules per asymmetric unit. The assignment of the nitrogen atoms was confirmed by comparing the observed and expected torsion angles and bond lengths. Ligand (3) is less planar than the terpyridyl analogue (Fang et al., 2007) despite weak intramolecular C—H···lone pair (N) interactions. All non-bonded N···H distances are shorter than 2.75 Å [N2···H2 = 2.57 Å, N3···H4 = 2.57 Å, N5···H17 = 2.62 Å, N6···H19 = 2.51 Å]. The dihedral angles between the phenyl and pyridine rings in each molecule are 53.48 (5)° and 50.80 (5)°. These deviations from planarity, in part, may be influenced by weak intermolecular C—H···N hydrogen bonds connecting molecules into one-dimensional chains and in addition, by the crystal structure being stabilized by weak C—H···π stacking interactions between "head-to-tail" molecules.
Experimental
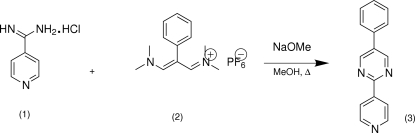
4-Pyridylamidine hydrochloride (1) (10,0 g, 63.5 mmol), 2-phenyl-1,3-bis(dimethylamino)trimethinium hexafluorophosphate (2) (1 eq, 22.1 g, 63.5 mmol) and NaOMe (1.2 eq, 4.13 g, 76.5 mmol) were dissolved in anhydrous MeOH (500 ml). The resulting yellow solution was refluxed for 15 h under N2. The white solid was isolated by filtration and dried under vacuum to give white shiny micro-crystals (11.0 g, 74%) of pure title compound (3). These crystals were suitable for X-ray diffraction measurements, m.p. 477.8–478.5 K. Anal Calcd for C15H11N3 (233.3): C, 77.23; H, 4.75; N, 18.01. Found: C, 77.04; H, 4.69; N, 17.86.
Refinement
H atoms were generated geometrically (C—H = 0.95 Å) and were included in the refinement in the riding model approximation; their temperature factors were set to 1.2 times those of the equivalent isotropic temperature factors of the parent site. A final verification of possible voids was performed using the VOID routine of the PLATON program (Spek, 2003).
Figures
Fig. 1.
The asymmetric unit with thermal ellipsoids shown at 50% probability levels. H atoms have been omitted.
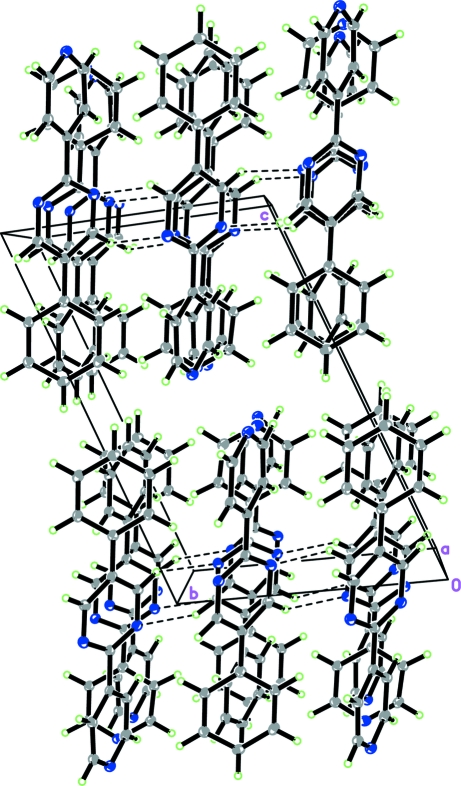
Fig. 2.
Part of the crystal structure of (3). Hydrogen bonds are shown as dashed lines. Ellipsoids are shown at the 30% probabilty level.
Fig. 3.
The reaction scheme for the title compound.
Crystal data
| C15H11N3 | Z = 4 |
| Mr = 233.27 | F(000) = 488 |
| Triclinic, P1 | Dx = 1.396 Mg m−3 |
| Hall symbol: -P 1 | Cu Kα radiation, λ = 1.54178 Å |
| a = 9.2813 (5) Å | Cell parameters from 5655 reflections |
| b = 9.3609 (5) Å | θ = 3.4–68.9° |
| c = 13.9001 (7) Å | µ = 0.68 mm−1 |
| α = 71.462 (2)° | T = 100 K |
| β = 86.957 (2)° | Block, colourless |
| γ = 75.788 (3)° | 0.40 × 0.38 × 0.08 mm |
| V = 1109.54 (10) Å3 |
Data collection
| Bruker SMART 6000 diffractometer | 3967 independent reflections |
| Radiation source: Rotating anode | 3226 reflections with I > 2σ(I) |
| Montel 200 optics | Rint = 0.046 |
| Detector resolution: 5.5 pixels mm-1 | θmax = 68.9°, θmin = 3.4° |
| ω scans | h = −11→11 |
| Absorption correction: multi-scan (SADABS; Sheldrick, 1996) | k = −11→11 |
| Tmin = 0.734, Tmax = 0.947 | l = −16→16 |
| 15167 measured reflections |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.044 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.139 | H-atom parameters constrained |
| S = 1.00 | w = 1/[σ2(Fo2) + (0.0925P)2 + 0.1892P] where P = (Fo2 + 2Fc2)/3 |
| 3967 reflections | (Δ/σ)max = 0.001 |
| 325 parameters | Δρmax = 0.27 e Å−3 |
| 0 restraints | Δρmin = −0.30 e Å−3 |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| N1 | −0.09769 (12) | 0.44347 (13) | 1.42209 (9) | 0.0235 (3) | |
| N2 | −0.12326 (12) | 0.79515 (13) | 1.05916 (8) | 0.0215 (3) | |
| N3 | 0.10382 (12) | 0.60327 (13) | 1.06789 (9) | 0.0230 (3) | |
| C28 | 0.60351 (14) | 1.04795 (15) | 0.59991 (10) | 0.0230 (3) | |
| H28 | 0.6235 | 1.0980 | 0.5313 | 0.028* | |
| C24 | 0.59612 (14) | 0.67217 (15) | 0.96148 (10) | 0.0211 (3) | |
| H24 | 0.6697 | 0.6196 | 0.9262 | 0.025* | |
| C22 | 0.40977 (14) | 0.89133 (15) | 0.96659 (10) | 0.0210 (3) | |
| H22 | 0.3519 | 0.9935 | 0.9349 | 0.025* | |
| C1 | −0.20867 (14) | 0.54005 (15) | 1.35782 (10) | 0.0222 (3) | |
| H1 | −0.3061 | 0.5581 | 1.3835 | 0.027* | |
| C2 | −0.18962 (14) | 0.61497 (15) | 1.25624 (10) | 0.0206 (3) | |
| H2 | −0.2717 | 0.6841 | 1.2144 | 0.025* | |
| C3 | −0.04783 (14) | 0.58719 (14) | 1.21624 (10) | 0.0188 (3) | |
| C4 | 0.06828 (14) | 0.48702 (15) | 1.28198 (10) | 0.0204 (3) | |
| H4 | 0.1664 | 0.4647 | 1.2580 | 0.025* | |
| C5 | 0.03846 (14) | 0.42042 (15) | 1.38280 (10) | 0.0218 (3) | |
| H5 | 0.1193 | 0.3543 | 1.4270 | 0.026* | |
| C6 | −0.02146 (13) | 0.66572 (15) | 1.10832 (10) | 0.0190 (3) | |
| C7 | −0.09684 (14) | 0.86669 (15) | 0.96277 (10) | 0.0207 (3) | |
| H7 | −0.1671 | 0.9585 | 0.9265 | 0.025* | |
| C8 | 0.02937 (13) | 0.81303 (14) | 0.91253 (10) | 0.0191 (3) | |
| C9 | 0.12654 (14) | 0.67759 (15) | 0.97125 (10) | 0.0224 (3) | |
| H9 | 0.2138 | 0.6359 | 0.9407 | 0.027* | |
| C10 | 0.05403 (13) | 0.89220 (15) | 0.80502 (10) | 0.0192 (3) | |
| C11 | 0.00409 (13) | 1.05394 (15) | 0.76275 (10) | 0.0203 (3) | |
| H11 | −0.0429 | 1.1139 | 0.8045 | 0.024* | |
| C12 | 0.02259 (14) | 1.12719 (15) | 0.66058 (10) | 0.0217 (3) | |
| H12 | −0.0111 | 1.2368 | 0.6330 | 0.026* | |
| C13 | 0.09030 (14) | 1.04032 (16) | 0.59868 (10) | 0.0223 (3) | |
| H13 | 0.1021 | 1.0903 | 0.5287 | 0.027* | |
| C14 | 0.14063 (14) | 0.88000 (15) | 0.63960 (10) | 0.0227 (3) | |
| H14 | 0.1872 | 0.8206 | 0.5974 | 0.027* | |
| C15 | 0.12309 (13) | 0.80658 (15) | 0.74159 (10) | 0.0202 (3) | |
| H15 | 0.1582 | 0.6971 | 0.7689 | 0.024* | |
| C29 | 0.71673 (14) | 0.94102 (15) | 0.66453 (10) | 0.0221 (3) | |
| H29 | 0.8144 | 0.9178 | 0.6400 | 0.027* | |
| C30 | 0.68821 (13) | 0.86809 (15) | 0.76430 (10) | 0.0197 (3) | |
| H30 | 0.7664 | 0.7944 | 0.8077 | 0.024* | |
| C25 | 0.54460 (14) | 0.90173 (15) | 0.80227 (10) | 0.0186 (3) | |
| C26 | 0.43120 (14) | 1.00916 (15) | 0.73613 (10) | 0.0203 (3) | |
| H26 | 0.3332 | 1.0326 | 0.7601 | 0.024* | |
| C27 | 0.46033 (15) | 1.08163 (15) | 0.63604 (10) | 0.0230 (3) | |
| H27 | 0.3825 | 1.1545 | 0.5919 | 0.028* | |
| C23 | 0.51637 (13) | 0.82266 (15) | 0.90904 (10) | 0.0184 (3) | |
| N6 | 0.57446 (11) | 0.59835 (13) | 1.05838 (8) | 0.0210 (3) | |
| C21 | 0.46901 (13) | 0.67626 (15) | 1.10567 (10) | 0.0187 (3) | |
| N5 | 0.38525 (12) | 0.82082 (12) | 1.06340 (8) | 0.0211 (3) | |
| C18 | 0.44441 (13) | 0.59364 (15) | 1.21314 (10) | 0.0186 (3) | |
| C19 | 0.49348 (13) | 0.43229 (15) | 1.25172 (10) | 0.0203 (3) | |
| H19 | 0.5441 | 0.3745 | 1.2097 | 0.024* | |
| C20 | 0.46761 (14) | 0.35763 (15) | 1.35169 (10) | 0.0215 (3) | |
| H20 | 0.4998 | 0.2476 | 1.3760 | 0.026* | |
| N4 | 0.39970 (12) | 0.43088 (13) | 1.41673 (8) | 0.0238 (3) | |
| C16 | 0.35492 (14) | 0.58659 (16) | 1.37925 (10) | 0.0231 (3) | |
| H16 | 0.3079 | 0.6415 | 1.4238 | 0.028* | |
| C17 | 0.37312 (13) | 0.67179 (16) | 1.27954 (10) | 0.0207 (3) | |
| H17 | 0.3377 | 0.7816 | 1.2568 | 0.025* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| N1 | 0.0246 (6) | 0.0248 (6) | 0.0228 (6) | −0.0064 (5) | −0.0001 (5) | −0.0092 (5) |
| N2 | 0.0197 (5) | 0.0236 (6) | 0.0209 (6) | −0.0007 (5) | −0.0027 (4) | −0.0095 (5) |
| N3 | 0.0199 (5) | 0.0226 (6) | 0.0238 (6) | −0.0003 (5) | 0.0001 (5) | −0.0071 (5) |
| C28 | 0.0266 (7) | 0.0257 (7) | 0.0181 (7) | −0.0088 (6) | 0.0001 (5) | −0.0070 (6) |
| C24 | 0.0192 (6) | 0.0235 (7) | 0.0211 (7) | −0.0017 (5) | −0.0004 (5) | −0.0104 (6) |
| C22 | 0.0198 (6) | 0.0187 (6) | 0.0229 (7) | −0.0011 (5) | −0.0013 (5) | −0.0068 (5) |
| C1 | 0.0192 (6) | 0.0252 (7) | 0.0250 (7) | −0.0050 (5) | 0.0014 (5) | −0.0119 (6) |
| C2 | 0.0174 (6) | 0.0218 (7) | 0.0243 (7) | −0.0032 (5) | −0.0030 (5) | −0.0102 (6) |
| C3 | 0.0191 (6) | 0.0176 (6) | 0.0215 (7) | −0.0033 (5) | −0.0019 (5) | −0.0090 (5) |
| C4 | 0.0173 (6) | 0.0208 (6) | 0.0254 (7) | −0.0033 (5) | −0.0016 (5) | −0.0108 (6) |
| C5 | 0.0205 (6) | 0.0209 (6) | 0.0234 (7) | −0.0029 (5) | −0.0046 (5) | −0.0070 (6) |
| C6 | 0.0165 (6) | 0.0191 (6) | 0.0225 (7) | −0.0019 (5) | −0.0035 (5) | −0.0094 (6) |
| C7 | 0.0199 (6) | 0.0213 (7) | 0.0197 (7) | −0.0001 (5) | −0.0047 (5) | −0.0078 (5) |
| C8 | 0.0182 (6) | 0.0194 (7) | 0.0212 (7) | −0.0025 (5) | −0.0035 (5) | −0.0094 (6) |
| C9 | 0.0180 (6) | 0.0250 (7) | 0.0225 (7) | −0.0012 (5) | 0.0010 (5) | −0.0080 (6) |
| C10 | 0.0135 (6) | 0.0229 (7) | 0.0225 (7) | −0.0037 (5) | −0.0025 (5) | −0.0090 (6) |
| C11 | 0.0171 (6) | 0.0225 (7) | 0.0230 (7) | −0.0028 (5) | −0.0029 (5) | −0.0106 (6) |
| C12 | 0.0194 (6) | 0.0208 (6) | 0.0247 (7) | −0.0041 (5) | −0.0049 (5) | −0.0064 (6) |
| C13 | 0.0211 (6) | 0.0283 (7) | 0.0191 (7) | −0.0085 (6) | −0.0010 (5) | −0.0075 (6) |
| C14 | 0.0187 (6) | 0.0285 (7) | 0.0257 (7) | −0.0066 (5) | 0.0013 (5) | −0.0145 (6) |
| C15 | 0.0160 (6) | 0.0193 (6) | 0.0256 (7) | −0.0026 (5) | −0.0017 (5) | −0.0085 (6) |
| C29 | 0.0200 (6) | 0.0279 (7) | 0.0228 (7) | −0.0077 (6) | 0.0017 (5) | −0.0128 (6) |
| C30 | 0.0167 (6) | 0.0221 (7) | 0.0206 (7) | −0.0016 (5) | −0.0044 (5) | −0.0089 (5) |
| C25 | 0.0182 (6) | 0.0188 (6) | 0.0213 (7) | −0.0040 (5) | −0.0016 (5) | −0.0096 (5) |
| C26 | 0.0166 (6) | 0.0211 (6) | 0.0242 (7) | −0.0030 (5) | −0.0008 (5) | −0.0093 (6) |
| C27 | 0.0224 (7) | 0.0215 (7) | 0.0239 (7) | −0.0032 (5) | −0.0061 (5) | −0.0059 (6) |
| C23 | 0.0143 (6) | 0.0211 (7) | 0.0211 (7) | −0.0030 (5) | −0.0025 (5) | −0.0092 (6) |
| N6 | 0.0197 (5) | 0.0231 (6) | 0.0197 (6) | −0.0008 (5) | −0.0017 (4) | −0.0090 (5) |
| C21 | 0.0154 (6) | 0.0209 (7) | 0.0214 (7) | −0.0029 (5) | −0.0031 (5) | −0.0094 (6) |
| N5 | 0.0195 (5) | 0.0203 (6) | 0.0214 (6) | −0.0014 (5) | 0.0000 (4) | −0.0063 (5) |
| C18 | 0.0135 (6) | 0.0232 (7) | 0.0202 (7) | −0.0038 (5) | −0.0024 (5) | −0.0081 (6) |
| C19 | 0.0170 (6) | 0.0229 (7) | 0.0230 (7) | −0.0031 (5) | −0.0034 (5) | −0.0104 (6) |
| C20 | 0.0181 (6) | 0.0217 (7) | 0.0231 (7) | −0.0030 (5) | −0.0051 (5) | −0.0054 (6) |
| N4 | 0.0208 (6) | 0.0288 (6) | 0.0214 (6) | −0.0051 (5) | −0.0024 (5) | −0.0076 (5) |
| C16 | 0.0203 (6) | 0.0288 (7) | 0.0223 (7) | −0.0043 (6) | −0.0002 (5) | −0.0119 (6) |
| C17 | 0.0173 (6) | 0.0228 (7) | 0.0229 (7) | −0.0026 (5) | −0.0022 (5) | −0.0097 (6) |
Geometric parameters (Å, °)
| N1—C1 | 1.3429 (18) | C11—C12 | 1.3890 (18) |
| N1—C5 | 1.3451 (17) | C11—H11 | 0.95 |
| N2—C7 | 1.3338 (17) | C12—C13 | 1.3904 (18) |
| N2—C6 | 1.3445 (17) | C12—H12 | 0.95 |
| N3—C9 | 1.3344 (17) | C13—C14 | 1.3911 (19) |
| N3—C6 | 1.3474 (16) | C13—H13 | 0.95 |
| C28—C29 | 1.3874 (19) | C14—C15 | 1.3854 (18) |
| C28—C27 | 1.3935 (19) | C14—H14 | 0.95 |
| C28—H28 | 0.95 | C15—H15 | 0.95 |
| C24—N6 | 1.3341 (17) | C29—C30 | 1.3810 (18) |
| C24—C23 | 1.3987 (19) | C29—H29 | 0.95 |
| C24—H24 | 0.95 | C30—C25 | 1.4068 (18) |
| C22—N5 | 1.3321 (17) | C30—H30 | 0.95 |
| C22—C23 | 1.4010 (18) | C25—C26 | 1.3998 (19) |
| C22—H22 | 0.95 | C25—C23 | 1.4746 (18) |
| C1—C2 | 1.3861 (18) | C26—C27 | 1.3853 (18) |
| C1—H1 | 0.95 | C26—H26 | 0.95 |
| C2—C3 | 1.3982 (18) | C27—H27 | 0.95 |
| C2—H2 | 0.95 | N6—C21 | 1.3446 (16) |
| C3—C4 | 1.3924 (19) | C21—N5 | 1.3442 (17) |
| C3—C6 | 1.4829 (18) | C21—C18 | 1.4825 (18) |
| C4—C5 | 1.3847 (18) | C18—C19 | 1.3962 (19) |
| C4—H4 | 0.95 | C18—C17 | 1.3970 (18) |
| C5—H5 | 0.95 | C19—C20 | 1.3808 (18) |
| C7—C8 | 1.3998 (18) | C19—H19 | 0.95 |
| C7—H7 | 0.95 | C20—N4 | 1.3440 (17) |
| C8—C9 | 1.3949 (19) | C20—H20 | 0.95 |
| C8—C10 | 1.4757 (18) | N4—C16 | 1.3458 (18) |
| C9—H9 | 0.95 | C16—C17 | 1.3871 (19) |
| C10—C15 | 1.4027 (18) | C16—H16 | 0.95 |
| C10—C11 | 1.4039 (19) | C17—H17 | 0.95 |
| C1—N1—C5 | 116.27 (12) | C12—C13—C14 | 119.78 (13) |
| C7—N2—C6 | 116.84 (11) | C12—C13—H13 | 120.1 |
| C9—N3—C6 | 116.34 (11) | C14—C13—H13 | 120.1 |
| C29—C28—C27 | 119.71 (12) | C15—C14—C13 | 120.31 (12) |
| C29—C28—H28 | 120.1 | C15—C14—H14 | 119.8 |
| C27—C28—H28 | 120.1 | C13—C14—H14 | 119.8 |
| N6—C24—C23 | 123.58 (11) | C14—C15—C10 | 120.70 (12) |
| N6—C24—H24 | 118.2 | C14—C15—H15 | 119.6 |
| C23—C24—H24 | 118.2 | C10—C15—H15 | 119.6 |
| N5—C22—C23 | 123.44 (12) | C30—C29—C28 | 120.36 (12) |
| N5—C22—H22 | 118.3 | C30—C29—H29 | 119.8 |
| C23—C22—H22 | 118.3 | C28—C29—H29 | 119.8 |
| N1—C1—C2 | 123.93 (12) | C29—C30—C25 | 120.69 (12) |
| N1—C1—H1 | 118 | C29—C30—H30 | 119.7 |
| C2—C1—H1 | 118 | C25—C30—H30 | 119.7 |
| C1—C2—C3 | 119.00 (12) | C26—C25—C30 | 118.38 (12) |
| C1—C2—H2 | 120.5 | C26—C25—C23 | 121.73 (11) |
| C3—C2—H2 | 120.5 | C30—C25—C23 | 119.88 (11) |
| C4—C3—C2 | 117.68 (12) | C27—C26—C25 | 120.69 (12) |
| C4—C3—C6 | 121.26 (11) | C27—C26—H26 | 119.7 |
| C2—C3—C6 | 121.03 (12) | C25—C26—H26 | 119.7 |
| C5—C4—C3 | 118.96 (12) | C26—C27—C28 | 120.18 (12) |
| C5—C4—H4 | 120.5 | C26—C27—H27 | 119.9 |
| C3—C4—H4 | 120.5 | C28—C27—H27 | 119.9 |
| N1—C5—C4 | 124.14 (12) | C24—C23—C22 | 114.48 (12) |
| N1—C5—H5 | 117.9 | C24—C23—C25 | 122.19 (11) |
| C4—C5—H5 | 117.9 | C22—C23—C25 | 123.33 (12) |
| N2—C6—N3 | 125.23 (12) | C24—N6—C21 | 116.46 (11) |
| N2—C6—C3 | 117.34 (11) | N5—C21—N6 | 125.40 (12) |
| N3—C6—C3 | 117.42 (11) | N5—C21—C18 | 118.22 (11) |
| N2—C7—C8 | 123.18 (12) | N6—C21—C18 | 116.37 (11) |
| N2—C7—H7 | 118.4 | C22—N5—C21 | 116.63 (11) |
| C8—C7—H7 | 118.4 | C19—C18—C17 | 117.45 (12) |
| C9—C8—C7 | 114.70 (12) | C19—C18—C21 | 120.29 (11) |
| C9—C8—C10 | 123.20 (11) | C17—C18—C21 | 122.26 (12) |
| C7—C8—C10 | 122.08 (12) | C20—C19—C18 | 119.28 (12) |
| N3—C9—C8 | 123.71 (11) | C20—C19—H19 | 120.4 |
| N3—C9—H9 | 118.1 | C18—C19—H19 | 120.4 |
| C8—C9—H9 | 118.1 | N4—C20—C19 | 124.09 (12) |
| C15—C10—C11 | 118.38 (12) | N4—C20—H20 | 118 |
| C15—C10—C8 | 120.59 (12) | C19—C20—H20 | 118 |
| C11—C10—C8 | 120.99 (11) | C20—N4—C16 | 116.18 (12) |
| C12—C11—C10 | 120.74 (12) | N4—C16—C17 | 124.03 (12) |
| C12—C11—H11 | 119.6 | N4—C16—H16 | 118 |
| C10—C11—H11 | 119.6 | C17—C16—H16 | 118 |
| C11—C12—C13 | 120.08 (12) | C16—C17—C18 | 118.96 (12) |
| C11—C12—H12 | 120 | C16—C17—H17 | 120.5 |
| C13—C12—H12 | 120 | C18—C17—H17 | 120.5 |
| C5—N1—C1—C2 | −0.23 (18) | C27—C28—C29—C30 | −0.06 (19) |
| N1—C1—C2—C3 | 1.52 (19) | C28—C29—C30—C25 | −0.50 (18) |
| C1—C2—C3—C4 | −1.28 (18) | C29—C30—C25—C26 | 0.86 (18) |
| C1—C2—C3—C6 | −179.38 (11) | C29—C30—C25—C23 | −179.99 (11) |
| C2—C3—C4—C5 | −0.09 (18) | C30—C25—C26—C27 | −0.67 (18) |
| C6—C3—C4—C5 | 178.00 (11) | C23—C25—C26—C27 | −179.80 (11) |
| C1—N1—C5—C4 | −1.28 (19) | C25—C26—C27—C28 | 0.13 (18) |
| C3—C4—C5—N1 | 1.46 (19) | C29—C28—C27—C26 | 0.25 (19) |
| C7—N2—C6—N3 | −0.15 (18) | N6—C24—C23—C22 | −0.22 (18) |
| C7—N2—C6—C3 | 178.6 (1) | N6—C24—C23—C25 | 179.12 (11) |
| C9—N3—C6—N2 | −0.02 (19) | N5—C22—C23—C24 | 0.14 (18) |
| C9—N3—C6—C3 | −178.77 (11) | N5—C22—C23—C25 | −179.20 (11) |
| C4—C3—C6—N2 | −158.60 (12) | C26—C25—C23—C24 | 148.58 (13) |
| C2—C3—C6—N2 | 19.42 (17) | C30—C25—C23—C24 | −30.54 (17) |
| C4—C3—C6—N3 | 20.25 (17) | C26—C25—C23—C22 | −32.13 (18) |
| C2—C3—C6—N3 | −161.73 (12) | C30—C25—C23—C22 | 148.75 (13) |
| C6—N2—C7—C8 | 0.11 (18) | C23—C24—N6—C21 | 0.22 (18) |
| N2—C7—C8—C9 | 0.08 (18) | C24—N6—C21—N5 | −0.14 (18) |
| N2—C7—C8—C10 | 178.41 (11) | C24—N6—C21—C18 | 179.6 (1) |
| C6—N3—C9—C8 | 0.23 (19) | C23—C22—N5—C21 | −0.07 (18) |
| C7—C8—C9—N3 | −0.26 (19) | N6—C21—N5—C22 | 0.07 (19) |
| C10—C8—C9—N3 | −178.57 (11) | C18—C21—N5—C22 | −179.67 (11) |
| C9—C8—C10—C15 | 33.47 (18) | N5—C21—C18—C19 | 160.88 (12) |
| C7—C8—C10—C15 | −144.72 (13) | N6—C21—C18—C19 | −18.89 (17) |
| C9—C8—C10—C11 | −148.99 (13) | N5—C21—C18—C17 | −19.43 (18) |
| C7—C8—C10—C11 | 32.82 (18) | N6—C21—C18—C17 | 160.81 (12) |
| C15—C10—C11—C12 | 0.05 (18) | C17—C18—C19—C20 | 1.14 (18) |
| C8—C10—C11—C12 | −177.54 (11) | C21—C18—C19—C20 | −179.15 (10) |
| C10—C11—C12—C13 | 0.43 (18) | C18—C19—C20—N4 | −1.45 (19) |
| C11—C12—C13—C14 | −0.57 (19) | C19—C20—N4—C16 | 0.40 (18) |
| C12—C13—C14—C15 | 0.24 (19) | C20—N4—C16—C17 | 0.95 (19) |
| C13—C14—C15—C10 | 0.25 (18) | N4—C16—C17—C18 | −1.19 (19) |
| C11—C10—C15—C14 | −0.39 (18) | C19—C18—C17—C16 | 0.08 (18) |
| C8—C10—C15—C14 | 177.20 (11) | C21—C18—C17—C16 | −179.62 (11) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| C7—H7···N5i | 0.95 | 2.55 | 3.3818 (18) | 147 |
| C9—H9···N6ii | 0.95 | 2.56 | 3.4027 (18) | 148 |
| C22—H22···N2i | 0.95 | 2.54 | 3.3784 (18) | 147 |
| C24—H24···N3ii | 0.95 | 2.56 | 3.4049 (19) | 149 |
| C1—H1···Cg3iii | 0.95 | 2.91 | 3.5925 (15) | 129. |
| C4—H4···Cg3 | 0.95 | 2.72 | 3.4163 (15) | 130. |
| C12—H12···Cg1i | 0.95 | 2.89 | 3.5844 (15) | 131. |
| C15—H15···Cg1iv | 0.95 | 2.92 | 3.5202 (15) | 122. |
| C17—H17···Cg6v | 0.95 | 2.84 | 3.5561 (16) | 133. |
| C20—H20···Cg6ii | 0.95 | 2.85 | 3.5251 (15) | 129. |
| C26—H26···Cg5 | 0.95 | 2.86 | 3.5242 (15) | 128. |
| C29—H29···Cg5vi | 0.95 | 2.77 | 3.4451 (15) | 129. |
Symmetry codes: (i) −x, −y+2, −z+2; (ii) −x+1, −y+1, −z+2; (iii) x−1, y, z; (iv) −x, −y+1, −z+2; (v) −x+1, −y+2, −z+2; (vi) x+1, y, z.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: LH2593).
References
- Bruker (1999). SAINT Release 7.06A. Bruker AXS Inc., Madison, Wisconsin, USA.
- Bruker (2003). SMART Release 5.625. Bruker AXS Inc., Madison, Wisconsin, USA.
- Fang, Y.-Q., Taylor, N. J., Hanan, G. S., Loiseau, F., Passalacqua, R., Campagna, S., Nierengarten, H. & Van Dorsselaer, A. (2002). J. Am. Chem. Soc.124, 7912–7913. [DOI] [PubMed]
- Fang, Y.-Q., Taylor, N. J., Laverdière, F., Hanan, G. S., Loiseau, F., Nastasi, F., Campagna, S., Nierengarten, H., Leize-Wagner, E. & Van Dorsselaer, A. (2007). Inorg. Chem.46, 2854–2863. [DOI] [PubMed]
- Medlycott, E. A. & Hanan, G. S. (2005). Chem. Soc. Rev.34, 133–142. [DOI] [PubMed]
- Medlycott, E. A. & Hanan, G. S. (2006). Coord. Chem. Rev.250, 1763–1782.
- Sheldrick, G. M. (1996). SADABS University of Göttingen, Germany.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Spek, A. L. (2003). J. Appl. Cryst.36, 7–13.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks I, global. DOI: 10.1107/S160053680800398X/lh2593sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S160053680800398X/lh2593Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report