Abstract
Epidemiological and animal studies indicate that selenium supplementation suppresses risk of colorectal and other cancers. The majority of colorectal cancers are characterized by a defective DNA mismatch repair (MMR). Here, we have employed the MMR-deficient HCT 116 colorectal cancer cells and the MMR-proficient HCT 116 cells with hMLH1 complementation to investigate the role of hMLH1 in selenium-induced DNA damage response, a tumorigenesis barrier. The ATM (ataxia telangiectasia mutated) protein responds to clastogens and initiates DNA damage response. We show that hMLH1 complementation sensitizes HCT 116 cells to methylseleninic acid, methylselenocysteine, and sodium selenite via reactive oxygen species and facilitates the selenium-induced oxidative 8-oxoguanine damage, DNA breaks, G2/M checkpoint response, and ATM pathway activation. Pretreatment of the hMLH1-complemented HCT 116 cells with the antioxidant N-acetylcysteine or 2,2,6,6-tetramethylpiperidine-1-oxyl or the ATM kinase inhibitor KU55933 suppresses hMLH1-dependent DNA damage response to selenium exposure. Selenium treatment stimulates the association between hMLH1 and hPMS2 proteins, a heterodimer critical for functional MMR, in a manner dependent on ATM and reactive oxygen species. Taken together, the results suggest a new role of selenium in mitigating tumorigenesis by targeting the MMR pathway, whereby the lack of hMLH1 renders the HCT 116 colorectal cancer cells resistant to selenium-induced DNA damage response.
Keywords: Carcinogenesis, Checkpoint Control, DNA Repair, Reactive Oxygen Species (ROS), Selenium
Introduction
Selenium is an essential nutrient widely distributed in organic forms in certain foods and inorganic forms in soil. Selenium exerts its anticarcinogenic effects mainly through selenoproteins at nutritional levels and through selenium metabolites, including reactive oxygen species (ROS),3 at supranutritional levels (1, 2). Animal and epidemiological studies strongly implicate selenium as an effective chemoprevention agent against colon cancer (3–6); however, the Nutritional Prevention of Cancer Trial and the recent Selenium and Vitamin E Cancer Prevention Trial reported mixed results on the efficacy of selenium in suppressing prostate cancer (7–9). Whatever the reason, the molecular mechanism by which selenium mitigates colorectal tumorigenesis is largely unknown.
The majority of colorectal cancers are characterized by microsatellite instability due to a defective MMR system (10, 11). The MMR system senses DNA base mismatch after DNA replication and provokes repair, checkpoint and apoptotic responses (12). Among the many human MMR proteins, hMLH1 (human MutL homologue-1) and hPMS2 (human post-meiotic segregation protein-2) form a complex that recognizes and stabilizes mismatched DNA at an early stage and facilitates the DNA damage response (13). Somatic mutations in MMR genes and epigenetic silencing of hMLH1 expression are observed in a significant portion of sporadic colorectal cancers (8, 14), whereas germ line mutations in hMLH1 account for 60% of the autosomal dominant nonpolyposis colon cancer. Loss of both hMLH1 and hMSH2 (human MutS homologue-2) is associated with complete inactivation of MMR, whereas defects in other MMR proteins result in only partial MMR deficiency (15).
The G2/M checkpoint prevents damaged cells from entering mitosis by coordinating DNA repair and/or apoptosis pathways. In response to DNA damage, hMLH1 can regulate G2/M transition through Cdc2 protein (16), and ATM kinase is implicated in G2/M checkpoint response (17–19). Moreover, a linkage between hMLH1 and ATM after DNA damage has been demonstrated (20, 21). We have shown recently that selenium compounds at doses ≤ LD50 can activate DNA damage response through ATM and ROS in noncancerous but not in cancerous cells (22). In the current study, we hypothesized that lack of a G2/M checkpoint response renders the hMLH1-deficient HCT 116 cells resistant to selenium-induced DNA damage response. By employing isogenic cell lines with or without hMLH1 expression, we show herein that hMLH1 is required for selenium-induced DNA damage and G2/M checkpoint response in a manner dependent on ATM kinase and ROS in HCT 116 colorectal cancer cells.
EXPERIMENTAL PROCEDURES
Cell Cultures and Chemicals
HCT 116 human colorectal adenocarcinoma cells complemented with an empty vector or a hMLH1-expressing vector (HCT 116+hMLH1) (23, 24), HeLa cervical cancer cells (ATCC, Manassas, VA), and Caco-2 epithelial colorectal adenocarcinoma cells (ATCC) were cultured in Dulbecco's modified Eagle's medium (Mediatech, Inc., Herndon, VA) supplemented with 10% fetal bovine serum (Atlanta Biologicals, Lawrenceville, GA), penicillin (50 international units/ml), and streptomycin (50 μg/ml). The engineered HCT 116 cells were maintained under the selection of hygromycin B (250 μg/ml, Invitrogen). MMR in the HCT 116 cells is defective due to a loss of chromosome 3 where hMLH1 resides; hMLH1 complementation restores MMR function and the hPMS2 protein level (25). CCD 841 CoN normal human colon cells (ATCC) were maintained in minimum Eagle's medium (Mediatech, Inc., Manassas, VA) supplemented with 15% fetal calf serum (Atlanta Biologicals), essential amino acids (1 ng/ml), nonessential amino acids (1 ng/ml), vitamins (1 ng/ml), penicillin (50 international units/ml), and streptomycin (50 μg/ml) at 37 °C in a 5% CO2 incubator. Sodium selenite (Na2SeO3), methylseleninic acid (MSeA), Se-methylseleno-l-cysteine (MSeC), nocodazole (Noc), N-acetylcysteine (NAC), and Tempo (2,2,6,6-tetramethylpiperidine-1-oxyl) were obtained from Sigma-Aldrich, and all were dissolved in water except for Noc (dissolved in DMSO). 3-[4,5-Dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide was from Calbiochem (La Jolla, CA) and was dissolved in ethanol. The ATM kinase inhibitor KU55933 was obtained from Tocris (Ellisville, MO) and dissolved in DMSO.
Cell Survival Assays
Four hours after being seeded into six-well plates (300 cells/well), the cells were treated with Na2SeO3 (0–20 μm), MSeA (0–10 μm), and MSeC (0–500 μm). Following 1 week of incubation, the colonies were fixed in 60% methanol, stained with 0.5% crystal violet, and then counted under a light microscope. A colony is defined as one containing >50 cells. Alternatively, cell survival was assessed by a 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide assay as described previously (23, 24).
Immunoprecipitation and Immunoblotting Analysis
Immunoprecipitation and immunoblotting analyses were performed as described previously (26, 27). Briefly, protein G-agarose-precleared lysates were immunoprecipitated with rabbit anti-PMS2 polyclonal antibodies (1:500, C-20, Santa Cruz Biotechnology, Santa Cruz, CA) for 16 h at 4 °C. After washing three times in lysis buffer and after heat denaturation, proteins in the anti-PMS2 immunoprecipitates were separated by SDS-PAGE and transferred to a PVDF membrane, followed by sequential incubation with rabbit anti-MLH1 polyclonal antibodies (1:1000, C-20, Santa Cruz Biotechnology), HRP-conjugated secondary antibodies, and the chemiluminescent reagents (SuperSignal, Pierce). Membranes were stripped using the Restore Western blot stripping buffer (Pierce) when necessary. ImageJ 1.43i (National Institutes of Health) was used to quantify the immunoblot signals on exposed films.
Flow Cytometric Assay
Cell cycle analysis was performed as described previously (22) with modifications. Briefly, HCT 116 and HCT 116+hMLH1 cells were seeded into 10-cm dishes and treated with selenium compounds for 0–48 h, with or without pretreatment with KU55933 (10 μm), NAC (10 mm), or Tempo (2 mm) for 24 h. Because G2/M cells tend to detach from the culture dishes, cell culture medium from selenium-treated cells were collected in an attempt to recover the loosely attached cells. Phases of the cell cycle were determined by flow cytometry (FACSCalibur cytometer with CELLQuest Pro, Becton Dickson, San Jose, CA) along with the ModFit LT software (version 3.0, Verity Software House, Topsham, ME). We used calibration standards (Molecular Probes, Carlsbad, CA) and a DNA QC particle kit (Becton Dickinson) to verify instrument performance.
Immunofluorescence
Immunofluorescence analysis was performed as described previously (22). Briefly, HCT 116 and HCT 116+hMLH1 cells were grown on coverslips and incubated with selenium compounds for 0–24 h. Alternatively, cells were pretreated with Noc (300 ng/ml) to capture cells in the G2/M phase. The cells were washed in PBS, fixed in 4% paraformaldehyde (in PBS) for 15 min, permeabilized in 0.3% Triton X-100 at room temperature for 10 min, followed by overnight incubation of the following antibodies at 4 °C: ATM (1:500, clone Y170, Epitomics, Burlingame, CA), phospho-ATM on Ser-1981 (pATM Ser-1981, 1:500, clone 10H11.E12, Rockland, Gilbertsville, PA), DNA-PKcs (DNA-dependent protein kinase catalytic subunit; 1:500, clone 18–2, Abcam, Cambridge, MA), phospho-DNA-PKcs on Thr-2647 (pDNA-PKcs Thr-2674, 1:500, Abcam), 8-oxoguanine (8-oxoG, 1:500, Abcam), H2AX (1:500, Abcam), and phospho-H2AX on Ser-139 (γH2AX, 1:500, clone 3F2, Abcam). Immunostaining was visualized by a fluorescence microscope (AxioObserver 100, Carl Zeiss, Oberkochen, Germany), and images were processed by using deconvolution with the software AxioVision Release 4.7.2.0.
Statistics
The data were analyzed using the GraphPad Prism 5.0 software (GraphPad Software, Inc., La Jolla, CA). Student's t test was applied to determine statistical significance between the treatments. Unless indicated otherwise, the level of statistical significance was set at p < 0.05.
RESULTS
hMLH1 Complementation Sensitizes HCT 116 Cells to Selenium Compounds and Potentiates the Selenium-induced DNA Oxidation and Breaks
Selenium-induced cell cycle arrest and apoptosis in colon cancer cells has been well studied (28–30); however, a direct role of hMLH1 in selenium-mediated DNA damage response has not been studied under an isogenic background. To determine whether hMLH1 plays a role in the response of the cancerous cells to selenium compounds, we first assessed sensitivity of HCT 116 and HCT 116+hMLH1 cells to Na2SeO3 (Fig. 1A), MSeA (Fig. 1B), and MSeC (Fig. 1C) by colony formation assays. We chose the selenium compounds because the organic MSeA and MSeC are known to be very efficacious in suppressing tumors in animal models (31), yet Na2SeO3 is as effective as the organic selenium compounds in activating early barriers of tumorigenesis in cell models (22). Here, we found that HCT 116+hMLH1 cells were significantly (p < 0.05) more sensitive than HCT 116 cells to the three selenium compounds in a dose-dependent manner. This observation was confirmed by 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide assays (supplemental Fig. S1) and sub G0/G1 cell profiles containing fragmented DNA (HCT 116+hMLH1 cells, 7–10%; HCT 116 cells, trace amount) after selenium exposure (supplemental Fig. S2). Consistent with the notion that selenium compounds can induce ROS formation (22, 32), addition of the antioxidant NAC and Tempo dramatically suppressed the selenium-induced cell death in HCT 116+hMLH1 (Fig. 1, D–F) and HCT 116 cells (supplemental Fig. S3) as evidenced by colony formation assays. The expression of hMLH1 in the HCT 116+hMLH1 cells was comparable to that in the hMLH1-proficient Caco-2 colorectal cancer cells but was substantially lower than the HeLa cervical cancer cells and higher than the noncancerous CCD841 colorectal cells (supplemental Fig. S4). This suggests that the hMLH1 expression in the HCT 116+hMLH1 cells is not hyperexpressed and is at a physiological level.
FIGURE 1.
Sensitivity of HCT 116 cells and hMLH1-complemented HCT 116 cells to the selenium compounds and antioxidants. A–C, HCT 116 cells and HCT 116+hMLH1 cells were cultured in six-well plates and treated with Na2SeO3, MSeA, or MSeC. D–F, HCT116+hMLH1 cells were pretreated with antioxidants NAC or Tempo, followed by treatment with the selenium compounds. Cell survival was assessed by colony formation assay. The number of colonies in the condition without selenium treatment was set as 100%. Values are mean ± S.E. (n = 3). *, p < 0.05 compared with HCT 116+hMLH1 cells (A–C) or with antioxidant treatments (D–F).
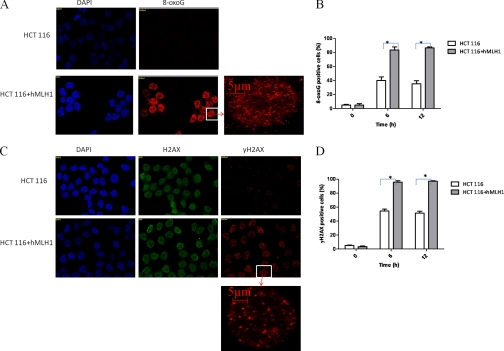
We next determined the nature of DNA damage in the selenium-treated colorectal cancer cells. The mutagenic 8-oxoG oxidative DNA lesions can lead to GC → TA mutations and DNA strand breaks if left unrepaired (33). H2AX is a phosphorylation substrate of ATM, resulting in the expression of γH2AX, a marker of DNA breaks. Analyses of immunofluorescence results indicated that 8-oxoG (Fig. 2, A and B) and γH2AX (Fig. 2, C and D) in the nuclei were induced after treatment with Na2SeO3 (2 μm), and the extent was much greater (p < 0.05) in HCT 116+hMLH1 than in HCT 116 cells. Pretreating HCT 116+hMLH1 cells with the antioxidant NAC or Tempo significantly (p < 0.05) reduced the Na2SeO3-induced 8-oxoG and γH2AX formation (supplemental Fig. S5). Altogether, hMLH1 sensitizes HCT 116 cells to selenium compounds in a manner dependent on ROS and promotes oxidative 8-oxoG damage and DNA breaks.
FIGURE 2.
Complementing HCT 116 cells with hMLH1 potentiates selenium-induced DNA oxidation and breaks. HCT 116 and HCT 116+hMLH1 cells were cultured on coverslips and treated with Na2SeO3 (2 μm) for 6 and 12 h, followed by immunofluorescence analyses of 8-oxoG and γH2AX. Ratios of nuclei (DAPI staining) showing 8-oxoG staining (A and B) and ratios of cells expressing γH2AX (C and D) to cells expressing total H2AX are presented with their respective S.E. values (n = 3). *, p < 0.05. A and C, representative pictures showing immunofluorescent signals of DAPI (blue), total H2AX (green), and γH2AX or 8-oxoG (red).
hMLH1 Complementation Facilitates G2/M arrest in Selenium-treated HCT 116 Cells
Previous results indicate a selenomethionine dose-dependent cell cycle arrest in HCT 116 cells (29, 30). Here, we determined a role of hMLH1 in selenium-induced DNA damage checkpoint response in colorectal cancer cells. Flow cytometric analyses indicated a time-dependent G2/M arrest 0–12 h after Na2SeO3 exposure (2 μm) (Fig. 3C) in HCT 116+hMLH1, but not in HCT 116 cells. The G2/M arrest in HCT 116+hMLH1 cells subsided at 24 and 48 h. Consequently, G0/G1 (Fig. 3A) and S (Fig. 3B) populations were less abundant in HCT 116+hMLH1 than in HCT 116 cells at 6 and 12 h. Similar patterns of cell cycle profiles exist 0–48 h after cellular exposure with MSeA (2 μm) or Na2SeO3 (5 μm) (supplemental Fig. S6). The above studies implicate hMLH1 in G2/M cell cycle arrest in selenium-treated HCT 116 cells.
FIGURE 3.
Complementing HCT 116 cells with hMLH1 restores G2/M arrest after selenium exposure. HCT 116 and HCT 116+hMLH1 cells were treated with Na2SeO3 (2 μm), harvested at the indicated time points, and analyzed for cell cycle profiles by flow cytometry (n = 3). Cells in G0/G1 (A), S (B), and G2/M (C) phases were quantified and presented. Values are mean ± S.E. (n = 3). *, p < 0.05.
Selenium-induced G2/M Arrest in HCT 116+hMLH1 Cells Depends on ATM Kinase Activity and ROS
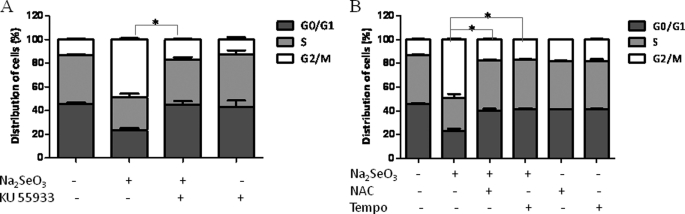
We have shown previously that selenium-induced DNA damage response involves ROS and ATM (22). ATM activation initiates a cascade of DNA damage responses, including G2/M arrest, to various types of DNA damage (17–19). Therefore, we treated HCT 116+hMLH1 cells sequentially with the ATM kinase inhibitor KU55933, the antioxidant NAC, or the superoxide dismutase mimic Tempo and Na2SeO3 (2 μm). The results showed that pretreatment of the cells with KU55933 (Fig. 4A), NAC, or Tempo (Fig. 4B) attenuated the selenium-induced G2/M arrest. KU55933, NAC, or Tempo alone did not impact on cell cycle profiles in HCT 116+hMLH1 cells.
FIGURE 4.
Effects of ATM kinase activity and ROS on the hMLH1-dependent G2/M arrest after selenium exposure. HCT 116+hMLH1 cells were pretreated with 10 μm KU55933 (A) and 10 mm NAC or 2 mm Tempo (B) for 24 h, followed by Na2SeO3 (2 μm) exposure for 12 h. Cell cycle profiles were analyzed as described in the legend to Fig. 3.
Selenium-induced ATM Pathway Activation Requires hMLH1 and Occurs Mainly in G2/M Phase
The above results suggest that hMLH1 may mediate selenium-induced ATM pathway activation in the G2/M phase. To test this hypothesis, we determined the expression of G2/M phase pATM Ser-1981, a general marker of ATM pathway activation (34), and pDNA-PKcs Thr-2647 (phospho-DNA-PKcs on Thr-2647), a phosphorylation event downstream of ATM kinase activation (35), in HCT 116 and HCT 116+hMLH1 cells treated with Na2SeO3 (2 μm) with or without pretreatment of the microtubule blocker Noc (300 ng/ml) for 16 h. This condition traps about 90% cells in G2/M phase (36, 37). Immunofluorescence analysis showed that pATM Ser-1981 (Fig. 5, A and B) and pDNA-PKcs Thr-2647 (Fig. 5, C and D) levels increased marginally in HCT 116 cells after selenium exposure. In stark contrast, both the phosphorylation events are substantially induced in HCT 116+hMLH1 6 and 12 h after the Na2SeO3 exposure. Furthermore, the Na2SeO3-induced expression of pATM Ser-1981 and pDNA-PKcs Thr-2647 were significantly enhanced by Noc treatment in HCT 116+hMLH1 but not in HCT 116 cells. Noc treatment alone did not induce expression of pATM Ser-1981 or pDNA-PKcs Thr-2647. Pretreatment of HCT 116+hMLH1 cells with NAC, Tempo, or KU55933 significantly (p < 0.05) attenuated Na2SeO3-induced pATM Ser-1981 formation (supplemental Fig. S7). Collectively, results from Figs. 4 and 5 show that the hMLH1-dependent, selenium-induced G2/M arrest requires ATM and ROS, and the ATM pathway activation occurs mainly in G2/M phase.
FIGURE 5.
hMLH1 mediates Na2SeO3-induced ATM pathway activation in G2/M. Undisturbed or G2/M-arrested (300 ng/ml Noc, 16 h) cells were treated with Na2SeO3 (2 μm) for 0–12 h prior to immunofluorescence analyses. Ratios of cells expressing pATM Ser-1981 (A and B) or pDNA-PKcs Thr-2647 (C and D) to cells expressing total ATM or total DNA-PKcs are presented with their respective S.E. values. *, p < 0.05.
The Association between hMLH1 and hPMS2 Is Stimulated by Selenium Treatment and Depends on ATM Kinase Activity and ROS
At the early stage of MMR, hMLH1 and hPMS2 form a heterodimer to stabilize mismatched DNA and facilitate DNA damage response (13). Therefore, we performed co-immunoprecipitation experiment to determine whether the association between hMLH1 and hPMS2 was altered after selenium treatment. We found that treatment of HCT 116+hMLH1 cells with Na2SeO3 (Fig. 6A) and MSeA (supplemental Fig. S8) increases the amount of hMLH1 in the anti-hPMS2 immunoprecipitates at 24 h in a dose-dependent manner. Reciprocal co-immunoprecipitation was not achievable in our hands using available anti-hMLH1 antibodies. Incubation of the immunoprecipitates with ethidium bromide did not affect the protein associations (supplemental Fig. S9), suggesting that the protein-protein interaction was not due to DNA bridging. Next, we determined whether ROS and ATM kinase were involved in the increased hMLH1-hPMS2 interaction. Pretreatment with NAC (Fig. 6B), Tempo (Fig. 6C), or KU55933 (Fig. 6D) prevented the hMLH1-hPMS2 association in HCT 116+hMLH1 cells with or without selenium treatment. Similar to a previous report (25), the selenium compounds, antioxidants, and KU55933 treatment had no effect on protein levels of hMLH1 or hPMS2 (Fig. 6, A–D and supplemental Fig. S8). Taken together, selenium treatment stimulates hMLH1-hPMS2 association in a manner dependent on ATM kinase and ROS.
FIGURE 6.
Selenium treatment increases hMLH1-hPMS2 association in a manner dependent on ATM and ROS. A, HCT 116 and HCT 116+hMLH1 cells were treated with Na2SeO3 (2 μm) for 24 h. B–D, HCT 116+hMLH1 cells were pretreated with NAC (10 mm), Tempo (2 mm), or KU55933 (10 μm) for 24 h, followed by Na2SeO3 (2 μm) exposure for 24 h. The lysates were immunoprecipitated (IP) using anti-IgG or anti-PMS2 antibodies, followed by immunoblotting using the indicated antibodies. The amount of hMLH1 associated with anti-hPMS2 immunoprecipitates was estimated based on the band intensity of the inputs. The related ratios in cells without selenium treatment were set as 1 and were presented as S.E. values. *, p < 0.05; **, p < 0.01 compared with no Na2SeO3 treatment. Inputs contained 5% of the amount of lysates used for immunoprecipitation.
DISCUSSION
In this study, we have identified hMLH1 as a selenium target for ATM-dependent G2/M arrest in colorectal cancer cells and provided insight into the resistance of the cancer cells to selenium treatment. Selenium may mitigate colorectal tumorigenesis at various stages by different mechanisms. Our recent study provides evidence that selenium compounds at low doses can activate an ATM- and ROS-dependent senescence response, an early barrier of tumorigenesis, in the noncancerous CCD841 but not the cancerous HCT 116 colorectal cells (22). The current study describes an essential role of hMLH1 in the ATM-dependent DNA damage response in the G2/M phase of HCT 116 cells exposed to relatively high doses of selenium. We show that hMLH1 complementation sensitizes HCT 116 cells to selenium-induced oxidative and broken DNA damage and stimulates the ATM-dependent G2/M checkpoint response in HCT 116 cells. After selenium exposure, the ATM-dependent DNA damage response promotes the hMLH1-hPMS2 association that is critical to initiate MMR. Without the hMLH1-depenedent ATM pathway activation, the cancerous HCT 116 cells fail to effectively transmit the selenium-induced oxidative DNA damage signal to checkpoint and cell death pathways. Consequently, the colorectal cancer cell may proliferate with unrepaired DNA damage and exacerbate the intrinsic genome instability.
How does hMLH1 complementation sensitize HCT 116 cells to selenium-induced cell killing effect? Although the primary function of MMR is to correct errors arising during DNA replication, MMR proteins are known to strongly enhance cellular and genetic toxicity of several chemotherapeutic agents (38, 39). Metabolism of selenium compounds at chemopreventive or therapeutic doses are known to induce ROS formation and the subsequent oxidative DNA base modifications and breaks in cancerous cells, including HCT 116 cells (22, 32, 40–45). MMR may mediate selenium-induced DNA breaks by two mechanisms. Firstly, the model of futile MMR cycle depicts that the generation of single-stranded breaks and gaps after the first S-phase can be converted into DNA DSBs by replication fork collapse when the unrepaired DNA lesions enter the next cycle of S-phase (38, 39). Secondly, recognition and processing of DNA damage by MMR proteins may directly induce DNA DSBs (46, 47). Because the selenium-induced γH2AX formation appears as fast as 8-oxoG formation, our study favors the latter scenario in which hMLH1 plays a signaling but not repair role in recognizing and processing the selenium-induced oxidative DNA damage in G2/M phase. The hMLH1-hPMS2 complex may be recruited to the region containing 8-oxoG, and its intrinsic nuclease activity may process the oxidatively modified DNA into breaks (48). Similarly, in the cellular response to the carcinogenic chromium, the MMR signaling complexes MSH2-MSH3 and MSH2-MSH6 contribute to a rapid formation of chromium-induced DNA DSBs (47). Consequently, the selenium-induced HCT 116+hMLH1 cells accumulate the unrepaired and highly toxic DNA breaks, arrest in G2/M phase, and enter the cell death pathway.
The ATM kinase is crucial in the response to DNA breaks and then transmits the signal to downstream substrates for S and G2/M arrest and/or cell death. An hMLH1-dependent ATM pathway activation has been demonstrated (20, 21), and cells deficient in MMR have defective G2/M arrest after DNA DSBs (16). As discussed above, the hMLH1-hPMS2 complex may rapidly process oxidative DNA damage into DNA breaks, which in turn activates the ATM kinase. Similar to chromium-induced DNA DSBs (46), the selenium-induced lesions are found primarily in G2/M phase. The nature of the DNA DSB induction by chromium and selenium is not totally understood, but this may involve cell cycle-specific assembly of DNA DSB-sensing protein complex such as the endonuclease-containing Mre11/Rad50/Nbs1 complex that can be recruited to damaged site in the S and G2 phase (49). In particular, the Mre11/Rad50/Nbs1 complex interacts with hMLH1 in G2 phase after the formation of O(6)-methylguanine induced by the chemotherapeutic agent temozolomide (50). Whatever the attribution, the hMLH1-dependent DNA DSB induction by selenium may activate ATM kinase in G2 phase, which in turn signals apoptosis pathway through key players such as p53, a prominent phosphorylation substrate of ATM (51). Interestingly, immunohistochemistry analysis of colorectal cancer samples reveals that expression of ATM is associated with cancer survival (52).
What does the increased association between hMLH1 and hPMS2 in HCT116+hMLH1 cells after selenium exposure implicate? To the best of our knowledge, this is the first report showing an increased association between hMLH1 and hPMS2 after DNA damage. hPMS2 is not expressed in the hMLH1-deficient HCT116 cells, suggesting the requirement of hMLH1-hPMS2 complex for functional MMR in colorectal cancer cells. Interestingly, ATM has been shown to interact with and possibly phosphorylates hMLH1 (53). Consistent with this notion, we found the increased hMLH1-hPMS2 association by selenium treatment is dependent on ATM kinase activity. ATM activation after DNA damage may stimulate hMLH1-hPMS2 association as observed in the case of an increased association between hPMS2 and p73 protein in the apoptotic response to cisplatin-induced DNA damage (25). Moreover, hMLH1 and hPMS2 both exhibit nuclear import and export signal domains, but hMLH1-hPMS2 dimerization masks the export peptides located at the C terminus (54). It is conceivable that ATM kinase activity facilitates the association between hMLH1 and hPMS2 and also confines the complex in the nucleus for efficiently sensing and processing the selenium-induced oxidative DNA damage.
In animal tumor models, MSeA and MSeC are more efficacious than Na2SeO3 in suppressing chemical-induced tumorigenesis (31). Nonetheless, LD50 of HCT 116+hMLH1 cells by MSeC is 100-fold higher than that by MSeA or Na2SeO3. Cultured cells lack a lyase and other enzymes necessary for conversion of MSeC to more effective downstream selenium metabolites (55, 56). Therefore, future studies employing selenium compounds in cultured cells for chemoprevention perspectives should avoid using MSeC.
Altogether, we propose that the hMLH1-hPMS2 complex senses and processes selenium-induced oxidative DNA damage and transmits the signal to ATM kinase, leading to the activation of G2/M checkpoint and death pathways. Nonetheless, other kinases such as DNA-PKcs also can be involved in the hMLH1-dependent sensitivity to selenium compounds. Because pDNA-PKcs Thr-2647 induction by DNA DSBs requires ATM (35), hMLH1 may mediate the selenium-induced formation of pDNA-PKcs on Thr-2647 via ATM. In HCT 116 cells, loss of a functional MMR compromises the ATM-dependent G2/M checkpoint response and contributes to resistance to selenium compounds and tolerance to aberrant DNA adducts. Consequently, cells may avoid apoptosis and enter mitosis with persistent DNA damage, exacerbating genome instability in the cancer cell. Although the majority of colorectal cancer progressively loses MMR function, activation of the hMLH1-dependent G2/M checkpoint response by selenium is a promising therapeutic target in precancerous or hMLH1-expressing colorectal cancer cells. Along with our recent publication (22), we propose that selenium compounds can differentially activate DNA damage response at various stages of colorectal tumorigenesis. In the initiation stage, DNA damage induced by selenium compounds at sublethal doses activates a senescence response (22), thus stifling cancer progression at a very early stage. During the promotion and progression stages of tumorigenesis, selenium compounds at lethal doses halt cells in the G2/M phase and promote cell death in a hMLH1-depdenent manner. The identification of the hMLH1-dependent activation of ATM DNA damage response provides mechanistic insight into selenium chemoprevention on colorectal cancer and suggests a promising molecular pathway for selectively killing of cancer cells by selenium.
Acknowledgments
We thank Dr. Françoise Praz and Dr. Thomas Kunkel for donating HCT 116 cells; Dr. Liangli Yu for Caco-2 cells; and Dr. Huawei Zeng, Dr. Benjamin Chen, and Ryan T. Y. Wu for comments.
This work was partially supported by the General Research Board, the University of Maryland at College Park (to W.-H. C.), the Chinese Top University Graduate Student Studying Abroad Program, China Scholarship Council (to Y. Q.), and the U. S. Department of Agriculture Agricultural Research Service Project 1235-51530-052–00D (to N. W. S).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S9.
- ROS
- reactive oxygen species
- MMR
- mismatch repair
- DSB
- double-strand break
- Na2SeO3
- sodium selenite
- MSeA
- methylseleninic acid
- MSeC
- Se-methylseleno-l-cysteine
- NAC
- N-acetylcysteine
- Noc
- nocodazole
- 8-oxoG
- 8-oxoguanine.
REFERENCES
- 1.Hatfield D. L., Yoo M. H., Carlson B. A., Gladyshev V. N. (2009) Biochim. Biophys. Acta 1790, 1541–1545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zeng H., Combs G. F., Jr. (2008) J. Nutr. Biochem. 19, 1–7 [DOI] [PubMed] [Google Scholar]
- 3.Clark L. C., Combs G. F., Jr., Turnbull B. W., Slate E. H., Chalker D. K., Chow J., Davis L. S., Glover R. A., Graham G. F., Gross E. G., Krongrad A., Lesher J. L., Jr., Park H. K., Sanders B. B., Jr., Smith C. L., Taylor J. R. (1996) JAMA 276, 1957–1963 [PubMed] [Google Scholar]
- 4.Davis C. D., Uthus E. O. (2002) J. Nutr. 132, 292–297 [DOI] [PubMed] [Google Scholar]
- 5.Feng Y., Finley J. W., Davis C. D., Becker W. K., Fretland A. J., Hein D. W. (1999) Toxicol. Appl. Pharmacol. 157, 36–42 [DOI] [PubMed] [Google Scholar]
- 6.Finley J. W., Davis C. D., Feng Y. (2000) J. Nutr. 130, 2384–2389 [DOI] [PubMed] [Google Scholar]
- 7.Shamberger R. J., Frost D. V. (1969) Can. Med. Assoc. J. 100, 682. [PMC free article] [PubMed] [Google Scholar]
- 8.Waters R. (2006) EMBO Rep. 7, 377–381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lippman S. M., Klein E. A., Goodman P. J., Lucia M. S., Thompson I. M., Ford L. G., Parnes H. L., Minasian L. M., Gaziano J. M., Hartline J. A., Parsons J. K., Bearden J. D., 3rd, Crawford E. D., Goodman G. E., Claudio J., Winquist E., Cook E. D., Karp D. D., Walther P., Lieber M. M., Kristal A. R., Darke A. K., Arnold K. B., Ganz P. A., Santella R. M., Albanes D., Taylor P. R., Probstfield J. L., Jagpal T. J., Crowley J. J., Meyskens F. L., Jr., Baker L. H., Coltman C. A., Jr. (2009) JAMA 301, 39–51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Jiricny J. (2006) Nat. Rev. Mol. Cell Biol. 7, 335–346 [DOI] [PubMed] [Google Scholar]
- 11.Kolodner R. D., Marsischky G. T. (1999) Curr. Opin. Genet. Dev. 9, 89–96 [DOI] [PubMed] [Google Scholar]
- 12.Hardman R. A., Afshari C. A., Barrett J. C. (2001) Cancer Res. 61, 1392–1397 [PubMed] [Google Scholar]
- 13.Plotz G., Raedle J., Brieger A., Trojan J., Zeuzem S. (2002) Nucleic Acids Res. 30, 711–718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kunkel T. A., Erie D. A. (2005) Annu. Rev. Biochem. 74, 681–710 [DOI] [PubMed] [Google Scholar]
- 15.Brieger A., Plotz G., Zeuzem S., Trojan J. (2007) Biochem. Biophys. Res. Commun. 364, 731–736 [DOI] [PubMed] [Google Scholar]
- 16.Yan T., Schupp J. E., Hwang H. S., Wagner M. W., Berry S. E., Strickfaden S., Veigl M. L., Sedwick W. D., Boothman D. A., Kinsella T. J. (2001) Cancer Res. 61, 8290–8297 [PubMed] [Google Scholar]
- 17.Arlander S. J., Greene B. T., Innes C. L., Paules R. S. (2008) Cancer Res. 68, 89–97 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shi M., Vivian C. J., Lee K. J., Ge C., Morotomi-Yano K., Manzl C., Bock F., Sato S., Tomomori-Sato C., Zhu R., Haug J. S., Swanson S. K., Washburn M. P., Chen D. J., Chen B. P., Villunger A., Florens L., Du C. (2009) Cell 136, 508–520 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 19.Shiloh Y. (2006) Trends Biochem. Sci. 31, 402–410 [DOI] [PubMed] [Google Scholar]
- 20.Adamson A. W., Beardsley D. I., Kim W. J., Gao Y., Baskaran R., Brown K. D. (2005) Mol. Biol. Cell 16, 1513–1526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Stojic L., Mojas N., Cejka P., Di Pietro M., Ferrari S., Marra G., Jiricny J. (2004) Genes Dev. 18, 1331–1344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wu M., Kang M. M., Schoene N. W., Cheng W. H. (2010) J. Biol. Chem. 285, 12055–12062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yanamadala S., Ljungman M. (2003) Mol. Cancer Res. 1, 747–754 [PubMed] [Google Scholar]
- 24.Koi M., Umar A., Chauhan D. P., Cherian S. P., Carethers J. M., Kunkel T. A., Boland C. R. (1994) Cancer Res. 54, 4308–4312 [PubMed] [Google Scholar]
- 25.Shimodaira H., Yoshioka-Yamashita A., Kolodner R. D., Wang J. Y. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 2420–2425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cheng W. H., von Kobbe C., Opresko P. L., Arthur L. M., Komatsu K., Seidman M. M., Carney J. P., Bohr V. A. (2004) J. Biol. Chem. 279, 21169–21176 [DOI] [PubMed] [Google Scholar]
- 27.Cheng W. H., von Kobbe C., Opresko P. L., Fields K. M., Ren J., Kufe D., Bohr V. A. (2003) Mol. Cell. Biol. 23, 6385–6395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Thant A. A., Wu Y., Lee J., Mishra D. K., Garcia H., Koeffler H. P., Vadgama J. V. (2008) Anticancer Res 28, 3579–3592 [PMC free article] [PubMed] [Google Scholar]
- 29.Goulet A. C., Chigbrow M., Frisk P., Nelson M. A. (2005) Carcinogenesis 26, 109–117 [DOI] [PubMed] [Google Scholar]
- 30.Goel A., Fuerst F., Hotchkiss E., Boland C. R. (2006) Cancer Biol. Ther. 5, 529–535 [DOI] [PubMed] [Google Scholar]
- 31.Ip C., Hayes C., Budnick R. M., Ganther H. E. (1991) Cancer Res. 51, 595–600 [PubMed] [Google Scholar]
- 32.Huang F., Nie C., Yang Y., Yue W., Ren Y., Shang Y., Wang X., Jin H., Xu C., Chen Q. (2009) Free Radic. Biol. Med. 46, 1186–1196 [DOI] [PubMed] [Google Scholar]
- 33.Grollman A. P., Moriya M. (1993) Trends Genet. 9, 246–249 [DOI] [PubMed] [Google Scholar]
- 34.Bakkenist C. J., Kastan M. B. (2003) Nature 421, 499–506 [DOI] [PubMed] [Google Scholar]
- 35.Chen B. P., Uematsu N., Kobayashi J., Lerenthal Y., Krempler A., Yajima H., Löbrich M., Shiloh Y., Chen D. J. (2007) J. Biol. Chem. 282, 6582–6587 [DOI] [PubMed] [Google Scholar]
- 36.Cheng W. H., Muftic D., Muftuoglu M., Dawut L., Morris C., Helleday T., Shiloh Y., Bohr V. A. (2008) Mol. Biol. Cell 19, 3923–3933 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ren J., Kan A., Leong S. H., Ooi L. L., Jeang K. T., Chong S. S., Kon O. L., Lee C. G. (2006) J. Biol. Chem. 281, 11413–11421 [DOI] [PubMed] [Google Scholar]
- 38.Fishel R., Lescoe M. K., Rao M. R., Copeland N. G., Jenkins N. A., Garber J., Kane M., Kolodner R. (1993) Cell 75, 1027–1038 [DOI] [PubMed] [Google Scholar]
- 39.Lin X., Howell S. B. (1999) Mol. Pharmacol. 56, 390–395 [DOI] [PubMed] [Google Scholar]
- 40.Zhou N., Xiao H., Li T. K., Nur-E-Kamal A., Liu L. F. (2003) J. Biol. Chem. 278, 29532–29537 [DOI] [PubMed] [Google Scholar]
- 41.Li G. X., Hu H., Jiang C., Schuster T., Lü J. (2007) Int. J. Cancer 120, 2034–2043 [DOI] [PubMed] [Google Scholar]
- 42.Rudolf E., Králová V., Cervinka M. (2008) Anticancer Agents Med. Chem. 8, 598–602 [PubMed] [Google Scholar]
- 43.Stewart M. S., Spallholz J. E., Neldner K. H., Pence B. C. (1999) Free Radic. Biol. Med. 26, 42–48 [DOI] [PubMed] [Google Scholar]
- 44.Wang H., Yang X., Zhang Z., Xu H. (2003) J. Inorg. Biochem. 97, 221–230 [DOI] [PubMed] [Google Scholar]
- 45.Hwang J. T., Kim Y. M., Surh Y. J., Baik H. W., Lee S. K., Ha J., Park O. J. (2006) Cancer Res. 66, 10057–10063 [DOI] [PubMed] [Google Scholar]
- 46.Peterson-Roth E., Reynolds M., Quievryn G., Zhitkovich A. (2005) Mol. Cell. Biol. 25, 3596–3607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Reynolds M. F., Peterson-Roth E. C., Bespalov I. A., Johnston T., Gurel V. M., Menard H. L., Zhitkovich A. (2009) Cancer Res. 69, 1071–1079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kadyrov F. A., Holmes S. F., Arana M. E., Lukianova O. A., O'Donnell M., Kunkel T. A., Modrich P. (2007) J. Biol. Chem. 282, 37181–37190 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Chen L., Nievera C. J., Lee A. Y., Wu X. (2008) J. Biol. Chem. 283, 7713–7720 [DOI] [PubMed] [Google Scholar]
- 50.Mirzoeva O. K., Kawaguchi T., Pieper R. O. (2006) Mol. Cancer Ther. 5, 2757–2766 [DOI] [PubMed] [Google Scholar]
- 51.Pandita T. K., Lieberman H. B., Lim D. S., Dhar S., Zheng W., Taya Y., Kastan M. B. (2000) Oncogene 19, 1386–1391 [DOI] [PubMed] [Google Scholar]
- 52.Grabsch H., Dattani M., Barker L., Maughan N., Maude K., Hansen O., Gabbert H. E., Quirke P., Mueller W. (2006) Clin. Cancer Res. 12, 1494–1500 [DOI] [PubMed] [Google Scholar]
- 53.Brown K. D., Rathi A., Kamath R., Beardsley D. I., Zhan Q., Mannino J. L., Baskaran R. (2003) Nat. Genet. 33, 80–84 [DOI] [PubMed] [Google Scholar]
- 54.Wu X., Platt J. L., Cascalho M. (2003) Mol. Cell. Biol. 23, 3320–3328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Drake E. N. (2006) Med. Hypotheses 67, 318–322 [DOI] [PubMed] [Google Scholar]
- 56.Zeng H. (2009) Molecules 14, 1263–1278 [DOI] [PMC free article] [PubMed] [Google Scholar]