Abstract
The hepatitis C virus (HCV) NS5b protein is an RNA-dependent RNA polymerase essential for replication of the viral RNA genome. In vitro and presumably in vivo, NS5b initiates RNA synthesis by a de novo mechanism. Different structural elements of NS5b have been reported to participate in RNA synthesis, especially a so-called “β-flap” and a C-terminal segment (designated “linker”) that connects the catalytic core of NS5b to a transmembrane anchor. High concentrations of GTP have also been shown to stimulate de novo RNA synthesis by HCV NS5b. Here we describe a combined structural and functional analysis of genotype 1 HCV-NS5b of strains H77 (subtype 1a), for which no structure has been previously reported, and J4 (subtype 1b). Our results highlight the linker as directly involved in lifting the first boundary to processive RNA synthesis, the formation of the first dinucleotide primer. The transition from this first dinucleotide primer state to processive RNA synthesis requires removal of the linker and of the β-flap with which it is shown to strongly interact in crystal structures of HCV NS5b. We find that GTP specifically stimulates this transition irrespective of its incorporation in neosynthesized RNA.
Keywords: Crystal Structure, Hepatitis Virus, Positive-strand RNA Viruses, RNA Polymerase, RNA Synthesis
Introduction
Hepatitis C virus (HCV)7 is a member of the Flaviviridae family that induces severe liver disease in humans (1). The viral genome is a single-stranded RNA of positive polarity containing a single open reading frame (ORF) flanked by two untranslated regions (UTRs), the 5′-UTR and 3′-UTR. The single ORF is translated into a large (∼3000 residues) polyprotein that is processed into some 10 mature proteins. Thus, the RNA-dependent RNA polymerase (RdRp) NS5b is cleaved from the C terminus of the polyprotein. In vivo the 591-residue NS5b is the central player in the synthesis of new genomic RNAs, in association with other viral and cellular proteins. This viral replication complex is associated with membranes (2) with the highly hydrophobic C-terminal 21 residues of NS5b forming a transmembrane helix (3). In vitro, NS5b has been shown to be capable of template-directed RNA synthesis on its own, requiring only divalent metals (magnesium or manganese) as cofactors. Indeed, NS5b can catalyze both de novo synthesis from a single-stranded template (4) and primer extension from the subsequent RNA duplex or from a preannealed template/primer duplex. The NS5b C-terminal transmembrane helix is dispensable for these activities, and C-terminal deletions of 21 residues (NS5b_Δ21) or more (NS5b_Δ47 to NS5b_Δ60) have been used in most activity and all crystallographic studies. The latter (5–7) has shown that the catalytic core of NS5b comprises residues 1–530 (Fig. 1E). They have also brought a puzzle to light; the 40-residue stretch (termed “linker” throughout this manuscript) between the catalytic core (fingers, palm, and thumb, Fig. 1) and the C-terminal membrane anchor occludes the catalytic cleft (5) in the crystal structures in which it is present (i.e. _Δ21 forms). The only reported exception is the case of the consensus subtype 2a NS5b_Δ21 (8), where two conformations of the same construct were crystallized, one with the linker in its usual, occluding position and one with the linker disordered. The arrangement of the linker in the active site of NS5b_Δ21 can be disrupted by mutations of critical linker residues interacting with the β-flap (Fig. 1B) of the thumb domain (9). The β-flap itself was proposed to participate in initiation of RNA synthesis by selective (10) and precise (11) positioning of a single-stranded RNA template along the finger RNA binding cleft, a prerequisite for proper de novo initiation.
FIGURE 1.
Structures of H77_NS5b in complex with GTP and of J4_NS5b_S556K. NS5b is in ribbon representation and colored according to domains; fingers are in red, palm is in yellow, thumb is in blue, and linker is in light brown. A–C, shown is H77_NS5b in complex with GTP. GTP molecules are in stick representation and are colored by element; Carbon, green, nitrogen, blue; oxygen, red; phosphorus, orange. A and B, two 90° rotation views of the complex are shown. The fingertips are labeled. Orientation in A is indicated by labeling of the thumb, palm, and fingers. The linker and β-flap (see Introduction for details) are labeled in B. C, details of the H77_NS5b GTP binding site are shown. Orientation is as in B, with the palm domain removed for clarity. Residues contacting the putative priming GTP (labeled P) are shown as sticks and colored by element. All residues thus highlighted are strictly conserved among HCV strains, except the two underlined. Position 556 is always a short hydrophilic residue (Gly, Ser, Asp, or Asn), and position 446 is always an acidic residue that is conserved within genotypes (Glu for 1a, 2, and 5; Asp for 3, 4, and 6), except for subtype 1b, where 446 is a conserved Gln. Pink dashes denote hydrogen bonds. D, the same view as C is shown of the J4_NS5b_S556K structure. 2Fo − Fc final electron density is displayed at a 1-δ level in a 1.6 Å radius around the linker and β-flap. Note that the well defined lysine 556 side chain points into the region occupied by the P GTP in C. E, shown is the primary structure of HVC_NS5b. The C-terminal transmembrane anchor, deleted from the constructs used in this study, is in dark gray.
In the present work we present a parallel structural and functional characterization of subtype 1a (H77 strain) and subtype 1b (J4 strain) NS5b enzymes with the aim of shedding new light on the puzzle of the linker involvement in the initiation of RNA synthesis. Our structural results include the first crystal structures of subtype 1a H77_NS5b_Δ21 alone and in complex with GTP and crystal structures of subtype 1b J4_NS5b_Δ21 in a new crystal form and of a mutant of the linker. A small-angle x-ray scattering (SAXS) characterization of J4_NS5b_Δ21 was also performed to assess the conformation of NS5b in solution. Functional analyses were done by in vitro RdRp assays of H77_NS5b_Δ21 and J4_NS5b_Δ21 (wild type and linker mutants). Taken together, these results clarify the early steps of RNA synthesis by HCV-NS5b; on the one hand, they point to the direct involvement of the linker in the very first steps of de novo initiation of RNA synthesis, leading to formation of the first dinucleotide primer. On the other hand, they show that GTP stimulates a later transition to processive elongation that is the true rate-limiting step in initiation.
EXPERIMENTAL PROCEDURES
Site-directed Mutagenesis
Site-directed mutagenesis of the serine 556 of H77 and J4 NS5b_Δ21 was performed by using the QuikChange site-directed mutagenesis kit (Stratagene) with oligonucleotides described in Table 1. Sequences of mutated fragments were verified by DNA sequencing using the ABI Prism Big Dye terminator sequencing kit at the Plateforme Genome transcriptome, Université de Bordeaux 2.
TABLE 1.
Oligonucleotides used in site-directed mutagenesis
Nucleotide changes are underlined.
| Oligonucleotides | Nucleotide sequences (5′→3′) |
|---|---|
| NS5b1b-S556Q | Sense, CGTCGCTGGTTACCAAGGGGGAGACATATATCACAGC |
| Anti-sense, ATATGTCTCCCCCTTGGTAACCAGCGACGAACCAGC | |
| NS5b1b-S556K | Sense, CGTCGCTGGTTACAAAGGGGGAGACATATATCACAGC |
| Anti-sense, ATATGTCTCCCCCTTTGTAACCAGCGACGAACCAGC | |
| NS5b1a-S556K | Sense, TTCACGGCTGGCTACAAAGGGGGAGACATTTATCACAG |
| Anti-sense, ATAAATGTCTCCCCCTTTGTAGCCAGCCGTGAACCAAC |
Protein Expression and Purification
The wild type or mutant NS5b_Δ21 of H77 and J4 were expressed in Escherichia coli and purified as previously described (12, 13). The H77_NS5b_Δ21 used in structural studies was the Q65H mutant described in Lou et al. (14) and was purified according to the protocol therein with 0.5% n-octylglucoside instead of 0.1% Nonidet P-40. The wild type J4_NS5b_Δ21 used in structural studies was expressed and purified according to the protocol of O'Farrell et al. (15), with minor modifications (i.e. 1% glucose was added to repress NS5b expression in all media, except the induction medium and carbenicillin was used as antibiotic instead of ampicillin). For all preparations used in structural work, the fractions containing the purified proteins were pooled, dialyzed against 5–10 mm Tris, pH 7.5, 0.2 m ammonium acetate, 1–4 mm DTT, 1 mm EDTA, flash-frozen in liquid nitrogen, and kept at −80 °C until use. Protein concentrations were determined by A280 with extinction coefficients calculated from the construct sequences.
Crystallization, Soaking, and Data Collection for H77_NS5b_Δ21
Monoclinic crystals (H77_X1) of H77_NS5b_Δ21 were initially obtained by the hanging drop vapor diffusion method. Equal volumes of protein solution (3.5 mg/ml in HEPES 8 mm, pH 7.5, 1.5 m ammonium acetate, 2.5 mm MgCl2, 1 mm EDTA, 4 mm DTT, also containing 55 μm concentrations of a short 5-base RNA and 0.5 mm GTP) and well solution (0.1 m MES, pH 6.5, 0.2 m ammonium sulfate, and PEG 5000 monomethyl ether) were mixed. To improve reproducibility and quality of the crystals, streak seeding and microseeding techniques were used. The final refined protocol uses the sitting drop vapor diffusion method and produces a triclinic crystal form (H77_X2). Equal volumes (2 μl each) of protein solution (3.5 mg/ml in 10 mm HEPES, pH 7.0, 0.2 m ammonium acetate, 1 mm EDTA, 5 mm DTT) and well solution (50 mm MES, pH 6.5, 0.2 m ammonium sulfate, 0.25 m ammonium acetate, 25–30% PEG 1000) were mixed and left to equilibrate overnight. Microseeding was then carried out with a dilute stock solution of crushed crystals, 0.2 μl of which was pipeted into each well. Crystals were transferred for a few seconds in a cryosolution containing 20% glycerol plus all the other components of the crystallization drop and flash-cooled in liquid ethane. In the case of soaks with substrates, these were added to a final concentration of 5–10 mm (nucleotides) and/or 0.1–1 mm (RNAs), MgCl2 was added to 10 mm, no ammonium sulfate was included, and soaking time was 20–40 min. High resolution synchrotron data (Table 2) were collected at beamline X06SA of the Swiss Light Source (Villigen, Switzerland).
TABLE 2.
Statistics of data collection and refinement
| Crystal form/PDB code | H77_X1/2XI2 | H77_X2/2XI3 | J4_02/2XHU | J4*_02/2XHV | J4_T/2XHW |
|---|---|---|---|---|---|
| Soaking conditions (immediately before cryocooling) | 80 μm CUGGC, 0.9 mm GTP, 0.14 m AmSO4, 25% PEG 5000 MME,d MES 0.1 m pH 6.5, 18% glycerol | 100 μm CUGGC, 10 mm GTP, 10 mm MgCl2, 25% PEG 5000 MME, MES 0.05 m pH 6.5, 20% glycerol | 0.2 m MgSO4, 25% PEG 2000 MME, 0.05 m HEPES pH 7.0, 15% glycerol | 0.2 m MgSO4, 25% PEG 2000 MME, 0.05 m HEPES pH 7.0, 15% glycerol | 0.2 m NaF, 8% PEG 3350, 0.05 m HEPES pH 7.0, 30% glycerol |
| Space group and cell parameters (Å, °) | P21 53.3 272.4 61.0 90 98.75 90 | P1 54.0 91.9 61.1 89.6 99.7 92.9 | P212121 106.5 107.8 133.8 90 90 90 | P212121 105.7 107.7 133.8 90 90 90 | P 32 2 1 125.4 125.4 140.1 90 90 120 |
| Number of molecules in the asymmetric unit | 3 | 2 | 2 | 2 | 1 |
| Resolution range (Å)a | 43.1-1.8 (1.85–1.8) | 30–1.7 (1.73-1.70) | 48.1-2.3 (2.37-2.29) | 34.7-1.9 (1.97-1.9) | 19.7–2.6 (2.72-2.66) |
| Rsymb | 0.082 (0.353) | 0.044 (0.348) | 0.15 (0.546) | 0.069 (0.359) | 0.20 (0.73) |
| I/σI | 13.2 (1.9) | 14.5 (2.0) | 12.90 (2.36) | 16.72 (3.21) | 8.59 (2.55) |
| Completeness (%) | 91.9 (81.6) | 95.5 (94.0) | 98.2 (97.1) | 98.5 (88.6) | 99.2 (92.4) |
| Unique reflections measured | 144,615 | 12,1329 | 69,005 | 118,899 | 36,792 |
| Average multiplicity | 2.9 (1.9) | 1.9 (1.8) | 5.5 (3.7) | 4.5 (2.9) | 9.1 (6.4) |
| Root mean square deviation bonds(Å)/angles(°) | 0.006/1.009 | 0.009/1.036 | 0.007/1.013 | 0.008/1.025 | 0.008/1.139 |
| R/Rfree (%)c | 16.6/20.7 (19.5/25.0) | 17.0/19.8 (27.3/32.0) | 16.5/21.6 (20.9/27.2) | 17.1/19.9 (22.3/27.2) | 17.2/22.0 (24.5/32.0) |
a In parentheses, statistics for the highest resolution shell.
b Rsym was determined by the equation R = ΣhklΣj|Ihkl,j − 〈Ihkl〉|/ΣhklΣjIhkl, where h, k, and l are the unique indices of all reflections measured more than once, and j the index for symmetry-redundant reflections.
c R and Rfree were determined by the equation R = Σhkl‖Fobs| − k |Fcalc‖/Σhkl|Fob‖, where h, k, and l are the indices of the reflections used in refinement (R) or of 5% of the reflections set aside and not used in refinement (Rfree). The same set of reflections was used for Rfree in all structures. Fobs is the structure factor deduced from the measured intensities, and Fcalc is the structure factor calculated from the model. k is a scale factor to put the Fcalc on the same scale as the Fobs.
d MME, monomethyl-ether.
Crystallization, Soaking, and Data Collection for J4_NS5b_Δ21 and J4_NS5b_S556K
The orthorhombic crystal form (J4_O1) reported by O'Farrell et al. (15) was reproduced by a different route and with significantly different cell parameters (our crystal form is hereafter called J4_O2). Our protocol is very reproducible and uses the hanging drop vapor diffusion method with microseeding; seeds were initially produced by crushing clusters of needles obtained by mixing equal 1-μl volumes of protein solution (6.3 mg/ml) and well solution (50 mm ammonium acetate, pH 5.0, 0.5 m NaCl, 10% glycerol, and 15–22% PEG 4000). Equal volumes (1 μl each) of protein solution and well solution (0.2 m magnesium sulfate, 15–25% PEG 2000 monomethyl-ether) were then mixed and left to equilibrate overnight. Microseeding was finally carried out with a 100–10,000× dilution of freshly crushed needles, 0.2 μl of which was pipeted into each drop.
A new, trigonal crystal form (J4_T) was obtained by lowering the salt concentration (initially 0.2 m ammonium acetate). Crystals were obtained by mixing 1 μl of protein solution (5 mg/ml) with 1 or 2 equal volumes of water and 1 or 2 equal volumes of reservoir solution (2–6% PEG 3350, 0.2 m NaF). The initial dilution promoted nucleation, and the subsequent drop shrinking allowed slow (∼1 week) growth of large (>0.3 × 0.3 × 0.3 mm3) trigonal crystals.
Crystals were transferred for a few seconds in a cryosolution containing all the components of the crystallization drop plus 15% (J4_O2) or 30% (J4_T) glycerol and flash-cooled in liquid nitrogen. In the case of soaks with substrates (J4_O2), magnesium sulfate was first substituted by 0.4 m ammonium acetate in 5 successive ∼2–5-fold dilutions of the drop. 10 mm MgCl2, 10 mm GTP, and/or 1 mm CUGGC RNA were then included, and soaking time was 30–40 min. High resolution synchrotron data (Table 2) were collected at beamline PROXIMA1 of the SOLEIL Synchrotron (Saclay, France) or beamline ID14–1 of the European Synchrotron Radiation Facility (Grenoble, France).
Structure Determination
Data were processed with the XDS package (16). Molecular replacement was performed with program Molrep (17). Models were rebuilt with COOT (18) and refined with phenix.refine (19). In the cases of H77_X1 and H77_X2, BK_NS5b_Δ21 was used as search model in molecular replacement (Protein Data Bank (PDB) code 1QUV), and the initial solution was first rebuilt automatically with arp_warp (20). In the cases of J4_O2 and J4_T, the previously published J4_NS5b_Δ21 (PDB code1NB4) was used as the search model in molecular replacement.
Objective Comparisons of Structures
Differences in main chain conformation between pairs of molecules were objectively assessed using program ESCET (21). We used significance values (ESCET “lolim” parameter) of 5.0 σ to identify broader conformational differences and 2.5 σ to take into account all significant conformational differences.
Small Angle X-ray Scattering
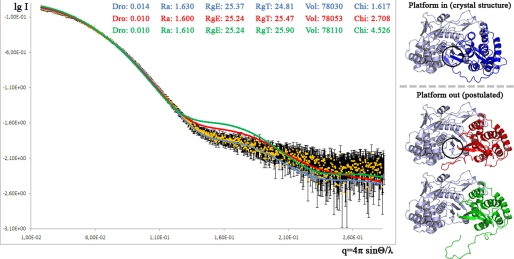
Protein samples of J4_NS5b_Δ21 were freshly dialyzed against Tris 10 mm, pH 7.5, ammonium acetate 200 mm, EDTA 1 mm, DTT 4 mm less than 48 h before and concentrated by ultrafiltration (Vivaspin, membrane cutoff, 30 kDa) to 4.0 mg/ml immediately before SAXS experiments. Data were collected at beamline ID14–3 of the European Synchrotron Radiation Facility (Grenoble, France). The beamline setup used was calibrated with a protein solution of known mass and concentration (bovine serum albumin at 4.5 mg/ml). Data were compared for radiation damage, but none was observed. Data were processed using standard procedures and extrapolated to infinite dilution by the program package PRIMUS (22). Program CRYSOL (23) was used to compare this experimental SAXS pattern to those calculated from the x-ray crystal structure and from more open models of J4_NS5b_Δ21. The latter models were constructed by opening the thumb and displacing the linker enough to accommodate a template/primer RNA duplex (see the legend to Fig. 7) taken from a Foot and Mouth Disease Virus RdRp complex (24).
FIGURE 7.
Conformation of J4_NS5b in solution compared with the crystal structure and more open models. SAXS spectra measured in solution (yellow), calculated from the crystal structure J4*_O2 (blue, Platform in) or from opened models (red and green, Platform out) are shown. In a Δ55 structure (PDB code 1GX5), the thumb domain is slightly more open than in Δ21 structures (9). The red thumb model was obtained by applying this rotation to the thumb of J4*_O2 three times and moving the linker out of the catalytic core, which is the minimal deformation for the enzyme to accommodate a double-stranded RNA. The green thumb model was obtained by moving the linker further out starting from the red thumb model, which is more consistent with the elongation process occurring. The approximate expected position of the initiation platform buttressing the base of the priming nucleotide is circled in the blue thumb crystal structure and the red thumb minimally opened model.
In Vitro RdRp Assays
Three templates were used in these assays. The first two correspond to the 341 nt of the 3′ end of the minus strand RNA from HCV genotype 1 (G1-C) or 3 (G3-U). They were obtained by in vitro transcription as previously described (25). The third template (G3–4U) was obtained by in vitro transcription of a DNA template obtained by PCR amplification from the 5′-UTR of the geno 3 clone 4 HCV with the G3–4U forward primer AAAACCCTCTTACGAGGGGACACTC and the 5′-341-T7-2 reverse primer (25).
Single-round Replication Assay
HCV RdRp and RNA were preincubated for 30 min at 25 °C in a reaction mixture containing 20 mm Tris-HCl, pH 7.5, 1 mm DTT, 5 mm MgCl2, 40 mm NaCl, 17 units of RNasin (Promega), 0.5 mm concentrations of the two initiating nucleotides (GTP and CTP for RNA G1-C; ATP and CTP for RNA G3-U) or of GTP alone or no NTP, 86 nm RNA template, and 200 nm purified NS5b. Heparin (Mr 4000–6000, 200 μg/ml), then [32P]UTP (3000 Ci·mmol −1, PerkinElmer Life Sciences) and the NTP needed for elongation (0.5 mm each) were successively added. The reaction was run at 25 °C for 0, 5, 10, 20, or 60 min and stopped by the addition of 10% trichloroacetic acid (TCA). The radioactivity incorporated into neo-synthesized RNA was then determined by scintillation counting.
Gel-based Initiation and Elongation Assay
The assay was carried out in 20 μl of 20 mm Tris, pH 7.5, 50 mm NaCl, 5 mm MgCl2, 1 mm DTT, 0.4 units/μl RNasin, 1 μm RNA template, 1 μm NS5b. When the initiation phase alone was analyzed, 0.5 mm GTP or ATP and 10 μm CTP with 4 μCi of [α-32P] CTP (3000 Ci·mmol−1) were the only added nucleotides. At different time points, 4 μl were collected, the reaction was quenched by adding 1 μl of 100 mm SDS and diluted in 8 μl of 90% formamide, 10 mm EDTA, 0.025% SDS, 0.025% bromphenol blue, 0.025% xylene cyanol loading buffer. After denaturation at 70 °C for 5 min, samples were loaded onto a 22% polyacrylamide gel in Tris borate EDTA buffer containing 7 m urea. The migration was performed at 30 watts for 3 h, and the gel was submitted to electronic autoradiography using a Pharos apparatus and Quantity One software (Bio-Rad). The elongation phase was analyzed after a 2-h incubation in the same assay conditions except for the NTP used. GTP was added at different concentrations before or at the same time as 0.5 mm ATP, 3′-dUTP, and 10 or 100 μm CTP with 4 μCi of [α-32P]CTP (3000 Ci·mmol−1, PerkinElmer Life Sciences). When the elongation was stopped with 0.5 mm 3′-dGTP instead of 3′-dUTP, UTP (0.5 mm) was added in the elongation reaction mixture. The reaction was stopped by adding 5 μl of 100 mm SDS. Five microliters were diluted in 8 μl of formamide/dyes loading buffer, and the reaction products were analyzed on polyacrylamide gel as above. The molecular weight markers were synthesized as described in Dutartre et al. (26) or obtained from Dharmacon and 32P-labeled with [γ-32P]ATP (3000 Ci·mmol−1, PerkinElmer Life Sciences) and T4 polynucleotide kinase following the manufacturer instructions (Promega). To enhance RNA synthesis in gel-based assays with templates with a 3′-U (G3-U and G3–4U), DMSO (10%v/v) was added in the reaction mixtures using these templates as well as to those using G1-C in side-by-side comparisons (cf. the legends to Figs. 4 and 6). Control experiments with G1-C indicated that the addition of DMSO increased 2–3-fold RNA synthesis without modifying the pattern of initiation and elongation products.
FIGURE 4.
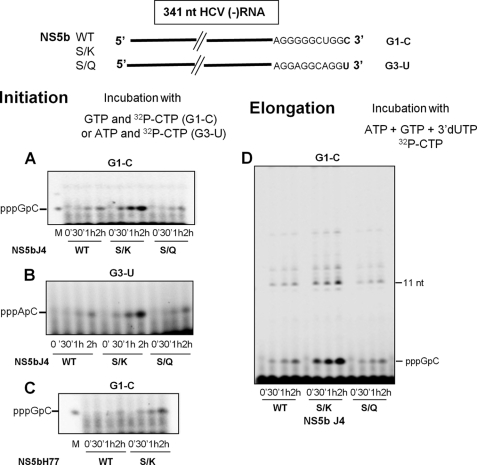
Effect of high GTP concentrations on the synthesis of initiation and elongation products. A, the nucleotide sequence of the 3′ end of template RNAs G1-C, G3-U, and G3-4U is shown. B, shown is the experimental scheme. C, RNA corresponding to the 341 nt from the 3′ end of the (-) strand RNA from H77 (G1-C) or genotype 3 clone 4 (G3-U) HCV were preincubated with J4 NS5b and 0, 50, 100, 200, and 500 μm GTP for 30 min at 25 °C. [α-32P]CTP and 3′-dUTP with and without ATP were then added, and incubation continued for 2 h in the presence of 10%(v/v) DMSO. The CTP concentration was 10 μm in synthesis with G3-U and 100 μm in synthesis with G1-C. The reaction was stopped by adding 5 μl of 100 mm SDS. Five microliters were diluted in 8 μl of formamide/dyes loading buffer, and the reaction products were analyzed on 22% polyacrylamide gel. The arrowhead indicates the position of the AC dinucleotide. M is for pppGpC molecular weight marker. D, G3–4U RNA was preincubated with J4 NS5b in the same conditions as above. [α-32P]CTP with and without ATP was then added with 3′-dUTP or 3′-dGTP plus UTP, and incubation continued for 2 h in the presence of 10%(v/v) DMSO. The reaction was stopped as in C, and the synthesized RNAs were analyzed on a 20% polyacrylamide gel. M is for RNA molecular weight markers of 4, 8, and 14 nt.
FIGURE 6.
Effect of S556 mutations on initiation and elongation of RNA synthesis. For quantification of initiation products, wild type or mutant NS5b and 341 nt (−) RNA were incubated in the RdRp reaction buffer with 0.5 mm GTP (G1-C) or ATP (G3-U) and 10 μm [α-32P]CTP as the only added nucleotides. 10% (v/v) DMSO was also added in the experiments performed with G3-U as template (panel B). At different time points, four microliters were collected and analyzed on 22% polyacrylamide gels as described under “Experimental Procedures.” The gels were submitted to electronic autoradiography, and the radioactive bands were quantified with the Quantity one software. A, shown is J4 NS5b with G1-C. B, shown is J4 NS5b with G3-U. C, shown is H77_NS5b with G1-C. D, for quantification of elongation products, wild type or mutant J4 NS5b and G1-C were incubated in a RdRp reaction buffer with 500 μm ATP, GTP, 3′-dUTP, and 100 μm [α-32P]CTP for 2 h at 25 °C. Four microliters of the reaction were analyzed on 20% polyacrylamide gels, and the products were quantified as above.
RESULTS
Crystal Structure of H77_NS5b and Comparison with J4_NS5b
We report here to high resolution the first structure of an NS5b of subtype 1a. The construct we used in the structural studies is a Q65H point mutant of strain H77_NS5b_Δ21. The Q65H mutation was isolated from a chronically infected chimpanzee, where it became fixed in the dominant virus present in the blood of the infected animal. The H77_NS5bQ65H_Δ21 construct was previously found to be even more active than the wild type in in vitro assays (14). We obtained two related crystal forms (X1 and X2) of H77_NS5bQ65H_Δ21 with, respectively, three and two molecules in the asymmetric units, yielding five high resolution views of the subtype 1a NS5b (Table 2). We subjected these five views to an objective conformational pairwise comparison with the program ESCET (21). Table 3 shows that we may ascribe the five molecules to two groups; X1_A, X1_B, and X2_A on the one hand and X1_C and X2_B on the other hand. The molecules are conformationally identical within each group but display small conformational differences between groups. These differences are ascribable to different crystal packing environments and are significant with a tolerance of 2.5 σ but not of 5 σ (Table 3, numbers in parentheses). In contrast, a structure of subtype 1b NS5b (strain J4, with nearly 90% sequence identity to H77_NS5b) is significantly different from all five views of H77_NS5b even with a tolerance of 5 σ (Table 3, last column). We may, therefore, break up the NS5b structure in subdomains according to the conformational differences between 1a and 1b enzymes (27). This analysis is pictured in Fig. 2. It shows that the main conformational difference lies in the shift of a block made of the structural elements taking part in the fingertips/thumb interaction, including helix A of the fingertips and helix S of the thumb. This difference leads to a maximum shift in this region in C-α position of 3.5 Å between J4_ and H77_NS5b (0.6 Å overall). In contrast, the part of the thumb (including the linker and β-flap) not directly involved in interactions with the fingertips is in the same rigid block as the palm and fingers.
TABLE 3.
Pairwise conformational comparisons between the five H77_NS5b molecules in two crystal forms (X1 and X2) and the best-defined J4_NS5b molecule in crystal form O (J4*_O2_B; see the legend to Table 4)
The letters A, B, and C indicate the protein chain used for the analysis; several protein molecules present in each asymmetric unit (Table 2). 〈esd〉, average coordinate error for α carbons in Å. The values are the percentages of residues whose α carbons are not in significantly different positions given tolerance (ESCET lolim parameter) of 2.5 σ (5 σ; for the values in parentheses). Values larger than 98% indicate identical molecules within error (49). With a higher lolim, significant global differences (values smaller than 98%, in bold) are still observed between H77_NS5b and J4_NS5b, whereas no differences remain between H77_NS5b chains and crystal forms. The underlined value refers to the molecules and lolim parameter used for Fig. 2.
| H77_X1_A | H77_X1_B | H77_X1_C | H77_X2_A | H77_X2_B | J4*_O2_B | |
|---|---|---|---|---|---|---|
| 〈esd〉 (Å) | 0.08 | 0.12 | 0.10 | 0.08 | 0.08 | 0.08 |
| H77_X1_A | 97.9 (99.9) | 95.3 (99.9) | 98.2 (100.0) | 93.0 (99.7) | 72.4 (94.9) | |
| H77_X1_B | 95.5 (99.7) | 98.0 (99.9) | 93.2 (99.4) | 80.9 (97.4) | ||
| H77_X1_C | 93.4 (99.8) | 99.4 (100.0) | 81.1 (97.6) | |||
| H77_X2_A | 89.9 (99.2) | 70.5 (93.3) | ||||
| H77_X2_B | 77.7 (96.5) | |||||
| J4*_O2_B |
FIGURE 2.
Objective comparison between H77_NS5b and J4_NS5b. Shown is a ribbon view of the H77_NS5b structure in the orientation of Fig. 1A and colored after ESCET comparison with J4_NS5b using a significance level of 5 σ (see the legend to Table 3). J4_NS5b_S556K was chosen for the comparison, as it is the best defined J4_NS5b structure available (Table 4, 〈esd〉 line) and conformationally identical to other J4_NS5b structures at this significance level (Table 4, numbers in parentheses). The conformational differences between H77_NS5b and J4_NS5b indicate two major blocks in the molecule; the first (434 residues) is colored blue, and the second (89 residues) is in green, with two smaller regions (cyan and magenta); red zones indicate flexible regions. Helix A of the fingertips and helix S of the thumb are labeled.
Noteworthy is the absence of ligands in the H77 X1 crystal form despite the presence in the crystallization drops of both a short RNA spanning the 3′-most 5 residues of the genotype 1 negative strand and GTP (see “Experimental Procedures”). Several sulfate ions were visible, however, one occupying the position of the triphosphate moiety of the “priming” nucleotide previously identified in our study of NS5b_1b_Δ55 in complex with nucleotides (28).
Crystal Structure of H77_NS5b in Complex with GTP
We reasoned that the absence of substrates in the H77_NS5b structure might be due to the presence of sulfate ions in the buffer, competing with the phosphate backbone of RNA and/or the triphosphate moieties of nucleotides. Therefore, we soaked crystals with solutions containing concentrated RNAs and/or nucleotides and no sulfates (see “Experimental Procedures”). These crystals belonged to the different space group X2 (Table 2), although whether from the soaking step or the seeding procedure with which they were obtained (see “Experimental Procedures”) is unknown. In the case of all soaks containing GTP (but with no other nucleotide), electron density for several nucleotides was apparent (Fig. 1, A, B, and C). Particularly interesting is the better ordered GTP (denoted P in Fig. 1C) present in the active site region. Its triphosphate moiety is stabilized by residues Arg-158, Arg-386, and Ser-367, the conserved triad identified as part of the priming site for a large class of de novo initiating RNA-dependent RNA polymerases (28). The base is sandwiched between residues 556 and 557 of the linker (depicted in light brown in Fig. 1) and the base of another, less well ordered nucleotide. Hydrogen bonds of the P GTP through its N1 and N2 to the carbonyl of Ser-282 seem to be the specificity determinant of this site for GTP. Finally, the ribose is held firmly in place by a bifurcated hydrogen bond between its 2′- and 3′-OH and Glu-446 of the β-flap (Fig. 1C) and by a hydrogen bond between its 2′-OH and the hydroxyl of Ser-556. Rather weak density for several other GTP molecules is localized at interfaces between NS5b molecules.
Involvement of GTP in the First Steps of RNA Synthesis from Template RNAs with a 3′-C or a 3′-U
In view of the H77_NS5b complex with GTP, we next investigated the role of high concentrations of GTP and that of the first two incorporated NTPs in the first steps of RNA synthesis by NS5b. To that end, we used templates corresponding to the 341 nucleotides from the HCV minus RNA of genotype 1 (G1-C) or genotype 3 (G3-U). We have previously shown that this RNA fragment is a good template for RNA synthesis by purified HCV NS5b in vitro and that initiation occurs by a de novo mechanism on the 3′ cytosine in the case of the H77 isolate-derived G1-C (12). In experimental conditions allowing multiple rounds of replication and/or template switching, it was 8–10-fold less replicated overall when the 3′ cytosine of G1-C was replaced by a 3′ uridine or with the genotype 3 clone 4 G3-U RNA that harbors a wild type 3′ uridine (25). We performed single-round kinetic experiments with G1-C or G3-U RNA templates and H77 and J4 NS5b. The RNA templates were first preincubated with NS5b alone or in the presence of a high concentration of GTP or of the two first nucleotides incorporated. Heparin, a trapping agent able to bind all NS5b unengaged in an initiation complex, was then added as well as the NTPs needed for further RNA synthesis. As shown in Fig. 3A, only low levels of productive initiation complexes were formed when either enzyme was preincubated with G1-C in the absence of any NTP. An 8–18-fold increase was observed when preincubation was performed in the presence of GTP, and a higher level still of RNA synthesis was obtained after preincubation with GTP and CTP, the two nucleotides needed for the synthesis of the initiation di- and trinucleotides (a further 17–30-fold increase compared with preincubation with GTP alone). Surprisingly, when G3-U was used as template, the RNA synthesis was 5-fold higher when preincubation was performed in the presence of GTP alone than with ATP and CTP, the two nucleotides complementary to the 3′ end of this template (Fig. 3B).
FIGURE 3.
Single round RNA synthesis by H77 and J4 NS5b. HCV RdRp and template RNA were preincubated for 30 min at 25 °C in the reaction mixture without NTP or with GTP or with the 2 initiating oligonucleotides. Heparin (Mr 4000–6000, 200 μg/ml) was then added followed by [α-32P]CTP and NTP needed to start the elongation. The reaction mixture was further incubated at 25 °C for 0, 5, 10, 20, and 60 min. The 32P RNA products were quantified after TCA precipitation and counted in a Wallac Counter. A, reactions were performed with G1-C and H77_NS5b (upper panel) or J4_NS5b (lower panel: empty diamonds, preincubation without NTP; filled squares, preincubation with GTP; filled triangles, preincubation with CTP and GTP). B, reactions were performed with G3-U and H77_NS5b (upper panel) or J4_NS5b (lower panel: filled squares, preincubation with GTP; filled circles, preincubation with ATP and CTP). Data were the mean of 3–6 independent experiments ± S.D.
This prompted us to perform gel-based analyses of initiation and elongation products synthesized from G1-C or G3-U after preincubation in the absence or with increasing concentrations of GTP (Fig. 4B). As expected, a GC dinucleotide was only produced from G1-C in the presence of GTP, and its amount increased with GTP concentration (Fig. 4C). The expected 11-nt elongation product generated after incorporation of 3′-dUTP was also only visible in the presence of GTP, confirming that initiation occurred at the 3′ end of the template. The band of higher molecular weight visible on the gel was probably due to enzyme stuttering at the CTP concentration used (100 μm). Alternatively, it could be produced from another initiation site. The first hypothesis is more likely as this higher band was the major elongation product formed at lower CTP concentration (10 μm), and it disappeared when the synthesis was performed with 500 μm CTP and ATP at 40 μm as 32P-labeled nucleotide (data not shown). Conversely, when G3-U was used as template, the same amount of AC initiation product was synthesized independently of preincubation with GTP (Fig. 4C). Longer products were only visible at high GTP concentration, but their profile was complex with several labeled bands, one of which migrating slightly slower than the pACCU marker. As mentioned above, this heterogeneity could be due to enzyme stuttering at the low CTP concentration used with this template (10 μm) or initiations at other sites in the template. To verify this point, we designed a new template (G3–4U RNA, Fig. 4A) by modifying the 3′ end of G3-U. In the new construct, the second and the third G residues were changed to U, and the C residue at the fifth position was changed to G. Finally, the G residue at 18 nt from the 3′ end was changed to C to restore the base pair at the bottom of the SL-A1 stem present in the wild type RNA. RNA synthesis was performed with G3–4U with the same method as in Fig. 4C. In these conditions, the concentration of the first 3 incorporated nt was not limiting (0.5 mm ATP), and the polymerization reaction should lead to an 8-nt elongation product in the presence of 3′-dUTP or a 14-nt elongation product in the presence of 3′-dGTP. As shown in Fig. 4D, left panel, an 8-nt RNA product migrating at the same position or very close to the 8-nt RNA marker was the major RNA synthesized in the presence of 3′-dUTP. Its amount increased with the concentration of GTP during preincubation, and it was undetectable in the absence of ATP or GTP. When the RNA synthesis was performed in the presence of 3′-dGTP, the expected 14-nt product was visible only when preincubation with GTP was performed before the addition of the three other nucleotides (compare lane 2 with lanes 3–6, Fig. 4D, right panel). Higher molecular weight products were also synthesized at the higher GTP concentrations due to GTP competing for incorporation with 3′-dGTP. Accordingly, the dominant product at the highest GTP concentration is the highest band visible on our gel. It might correspond to a complete copy of the template, poorly resolved on the 20% acrylamide gel. RNA products longer than 14 nt are detected at high GTP concentration even in the absence of ATP (Fig. 4D, right panel, first lane) and might be the results of heterogeneous initiations.
Altogether, these results show that preincubation of NS5b with template and high GTP concentrations can enhance RNA synthesis at two steps. The first step is the formation of the initial dinucleotide when GTP is the priming nucleotide. The second step, which can be singled out with the 3′-U RNA templates, is a later and rate-limiting transition from the initiation to an elongation phase.
S556K Mutation Enhances Initiation of RNA Synthesis in Both H77_ and J4_NS5b
In light of our crystal structure of H77_NS5b in complex with GTP, we decided to reinvestigate the role of the linker in GTP binding in the first steps of RNA synthesis after changes of the serine 556 to lysine or glutamine. Kinetic experiments in the presence of heparin were performed after preincubation of wild type or mutant J4_NS5b with the G3-U template and ATP and CTP or GTP alone. The results are presented in Fig. 5A. The curves obtained with WT and S556Q NS5b displayed very similar patterns, and in both cases RNA synthesis was higher after preincubation with GTP alone than with ATP and CTP (5.5 ± 0.7- and 5.6 ± 0.6-fold at the different time points, respectively). With the S556K NS5b, a higher amount of RNA was also synthesized after preincubation with GTP than with ATP and CTP (6.4 ± 0.8-fold), but the level of RNA synthesis was about 2-fold that of WT or S556Q enzyme (Fig. 5A, compare levels at the plateau for wild type and mutant enzymes). The same observations were made when the experiment was conducted with WT and S556K H77_NS5b (Fig. 5B).
FIGURE 5.
Single-round RNA synthesis by wild type or S556 mutant NS5b. HCV RdRp and G3-U RNA were preincubated for 30 min at 25 °C in the RdRp buffer with GTP or with ATP and CTP. Heparin was then added followed by [α-32P]CTP and NTP needed to start the elongation. The reaction mixture was further incubated at 25 °C for 0, 5, 10, 20, and 60 min. The 32P RNA products were quantified after TCA precipitation and counted in a Wallac Counter. A, reactions were performed with wild type, S556K, or S556Q J4 NS5b (filled squares, preincubation with GTP; filled circles, preincubation with ATP and CTP). B, the same experiments were performed with wild type or S556K H77_NS5b. Data are the mean of six independent experiments ± S.D.
With the aim of identifying the step of RNA synthesis, which was enhanced by the serine to lysine mutation in the NS5b linker, we performed gel-based initiation and elongation assays with wild type and mutant enzymes. To quantify the initiation phase, we performed kinetic experiments with the first two initiating nucleotides as only added NTPs, i.e. GTP and CTP for G1-C and ATP and CTP for G3-U. With both templates, the J4_NS5b with the S556K mutation synthesized 4.8 ± 0.8-fold more of dinucleotide than the WT or S556Q enzyme (Fig. 6, A and B). The same analysis was done with the WT and S556K H77_NS5b with G1-C only, as the level of activity of the WT H77_NS5b with G3-U was too low to allow accurate quantification of the initiation dinucleotide. In this case, a higher amount of GC dinucleotide was also observed with the S556K H77_NS5b compared with the WT enzyme (Fig. 6C). The effect of the amino acid changes on the elongation phase was then determined by using G1-C as template and J4_NS5b. Data illustrated in Fig. 6D indicated that the S556K mutation led to an overall 2-fold increase in the synthesis of the 11-nt elongation product compared with WT or S556Q mutant NS5b. This relatively low value compared with the 5-fold enhancement of initiation suggests that the S556K mutation does not interfere with the transition or the elongation phases. The 2-fold increase in elongation is likely simply due to the higher quantity of the dinucleotide primer synthesized during the initiation phase.
Altogether, these data indicate that the modifications introduced in the linker domain of H77 or J4 NS5b do not interfere with the increase of RNA synthesis observed after preincubation of the viral polymerase, the G3-U template, and GTP. However, the introduction of a lysine in place of the wild type serine greatly enhances de novo initiation up to formation of the first dinucleotide primer, seemingly with no further effects on subsequent polymerization steps.
Compared Structural Analysis of J4_NS5b, J4_NS5b_S556K, and H77_NS5b
To investigate the molecular basis of this enhancement of initiation, we crystallized a S556K mutant of J4_NS5b (J4*_NS5b) and refined its structure to high resolution (Table 2). When compared with the wild type J4_NS5b, the results are identical with those found for H77_NS5b with or without GTP; the conformational differences are significant only at the lower significance level (ESCET lolim parameter of 2.5 σ) and in different crystal environments (Table 4). In the same crystal environment, there are no discernible conformational differences between J4*_NS5b and J4_NS5b (Table 4, J4*_O2_B versus J4_O2_B). The only difference is in the side chain of the mutated residue; the substituted lysine side chain is well defined in the electron density map (Fig. 1D) and points into the space occupied by the P nucleotide in the H77_NS5b complex with GTP (Fig. 1C). We next tried to soak substrates into the J4_NS5b and J4*_NS5b crystals by a protocol similar to that used to obtain the H77_NS5b/GTP complex (i.e. by removing sulfate ions and introducing high concentrations of CUGGC RNA and GTP; see “Experimental Procedures”). We detected no binding of substrates, however, beyond a replacement of sulfates by triphosphates next to the Arg-158—Arg-386—Ser-367 (not shown). This phenomenon we routinely observe when soaking nucleotides into NS5b crystals and interpret as binding of nucleotides with disordered ribose and base at the priming site (28). Of possible note is the fact that part of the priming site (including Arg-386 and Arg-394) is slightly shifted in J4_NS5b with respect to H77_NS5b, being part of the fingertips-thumb block (in green on Fig. 2) that moves from one structure to the other (see above, “Crystal Structure of H77_NS5b and Comparison with J4_NS5b”).
TABLE 4.
Pairwise conformational comparisons between four different views of J4_NS5b
Table definitions are as in Table 3. In crystal form O (variants O1 and O2), the two molecules in the asymmetric unit are conformationally identical (not shown). The better-defined molecule B (lower 〈esd〉) is presented. Note that there are significant differences at 2.5 σ between molecules in crystal forms O1 (previously published, PDB code 1NB4), O2, and T (this report) but not between wild type J4_NS5b and J4_NS5b_S556K (denoted J4*) in the same crystal packing environment (O2, this report). Note also that with a tolerance of 5 σ, there are no significant conformational differences between any of the J4_NS5b molecules.
| J4_O1_B | J4*_O2_B | J4_O2_B | J4_T_A | |
|---|---|---|---|---|
| 〈esd〉 (Å) | 0.12 | 0.08 | 0.13 | 0.14 |
| J4_O1_B | 78.0 (99.1) | 88.1 (99.7) | 88.5 (99.3) | |
| J4*_O2_B | 100.0 (100.0) | 90.5 (98.9) | ||
| J4_O2_B | 94.2 (99.3) | |||
| J4_T_A |
Oligomerization State and Conformation of J4_NS5b_Δ21 in Solution
To analyze the oligomerization state and conformation of NS5b_Δ21 in solution and further probe the role of the linker, we performed small angle x-ray scattering analyses of J4_NS5b_Δ21. For a homogeneous sample, SAXS allows a measure of the molecular mass from equation I0 = KMC where I0 is the forward scatter, K is a constant for a given experimental setup, M the molecular mass, and C is the concentration of the object in solution. Results showed that protein samples were monodisperse in the buffer conditions (including 200 mm monovalent salt, cf. “Experimental Procedures”) with slight attractive interactions between molecules leading to an overestimation of the radius of gyration and molecular mass at higher protein concentrations (Table 5). Extrapolation to infinite dilution yielded an estimate of the molecular mass of the construct of 56 kDa. This estimate is the same for other HCV strains (not shown) and falls within 15% of the 64.4-kDa mass calculated from the sequence. This systematic error is, thus, of the same order of magnitude as the expected error on concentration of NS5b measured by A280. The same result was also obtained with 20 mm monovalent salt, albeit the samples then displayed stronger attractive interactions (not shown). Thus, J4_NS5b_Δ21 is a monomer in solution in the buffer conditions used, with no trace of oligomers or higher order aggregates. The curve calculated from the crystal structures of J4_NS5b_ Δ21 can be readily fitted (Fig. 7, blue model, curve and values) to the experimental SAXS data (Fig. 7, yellow points with error bars). This is not the case, however, of more open models constructed by the minimal opening of the thumb required to accommodate a double-stranded RNA, whether this is accompanied by a small (Fig. 7, red model, curve and values) or large (Fig. 7, green model, curve and values) displacement of the linker. More exotic models may be constructed where a combination of opening of the thumb and judicious placement of the linker yields a calculated curve closer to the SAXS data (not shown), so our data do not completely rule out that NS5b may be more open in solution than in the crystal state. Still, our results are strong evidence that the basal conformation of NS5b_Δ21 in solution is very close to that seen in crystal structures.
TABLE 5.
SAXS analysis of J4_NS5b_Δ21; concentration dependence of radius of gyration (Rg) and estimated molecular weight (Mr) and extrapolation to infinite dilution
| Sample | Concentration | Rg | Mr = I(0)/kc |
|---|---|---|---|
| mg/ml | nm | kDa | |
| J4Δ21 | 4.04 | 2.94 | 68.7 |
| 2.14 | 2.77 | 62.3 | |
| 0.98 | 2.64 | 61.2 | |
| Extrapolation | 0 | 2.49 | 56.4 |
DISCUSSION
Genotype 1 is both the most prevalent HCV genotype in developed countries and the genotype for which the current standard of care is least satisfactory (29). As strains H77 (subtype 1a) and J4 (subtype 1b) are known to be infectious in the chimpanzee model (30, 31), the polymerases from these strains have been extensively studied biochemically (15, 32–34). This has included extensive characterization of the modes of action of nonnucleoside NS5b inhibitors, including numerous structure-activity relationship studies (e.g. Ref. 33). These studies have taken advantage of the known structure and crystallization conditions of 1b_NS5b from the BK strain (that is also the alternative 1b strain NS5b used in activity and structure-activity relationship studies (32)) and of late the structure of a complex between a palm nonnucleoside inhibitor and NS5b_Δ21 of the important 1b Con1 strain (used in most genotype 1 replicon systems) (35). The structure of J4_NS5b_Δ21 is also available as well as its complex with a poorly ordered U5 RNA (15). Altogether, there are now more than 70 crystal structures of 1b NS5b available in the PDB, most of them complexes with nonnucleoside inhibitors.8 In contrast, no structure of a subtype 1a NS5b has been reported to date.
Here we report the crystal structure of an H77_NS5b_Δ21 point mutant (Q65H) that is more soluble than the wild type and, hence, more amenable to crystallization (14). Objective comparison (21) with the J4_NS5b_Δ21 structure shows that 1a and 1b enzymes have quite significantly different conformations in the region of interaction between the back of the thumb and the fingertips (Fig. 2). Our previous structure of the subtype 2a JFH1 NS5b had also pinpointed this region (including helix A from the fingertips and the thumb helix S, the top of which provides part of the resting place of helix A) as harboring significant conformational variation (36). This variation was linked to the special properties of JFH1_NS5b and specifically to a de novo RNA synthesis efficiency much higher than that of the NS5bs from other strains. We may, therefore, hypothesize that the lower specific activity reported by several groups for H77_NS5b compared with 1b enzymes such as J4_NS5b (Ref. 14; cf. also Fig. 3) may be linked to the difference in this region. Our finding that GTP binds at the priming site of H77_NS5b (Fig. 1, A–C) more easily than it does for J4_NS5b (in the same soaking conditions, we could detect only the triphosphate moiety) would seem at odds with H77_NS5b lesser de novo synthesis efficiency (e.g. Fig. 3A). However, during de novo initiation by viral RdRps such as HCV-NS5b, several steps may be rate-limiting. Clearly, in the case of NS5b there are at least two early steps that are disfavored (Fig. 8).The first step is formation of the first dinucleotide; hence, the requirements for high concentrations of the first two nucleotides (i.e. the priming and second nucleotides) (37). The second, and rate-limiting step is subsequent use of this dinucleotide as a primer; hence, there is a marked accumulation of dinucleotide in de novo initiation assays (cf. e.g. Fig. 4C) and requirement of a high concentration of the third nucleotide for further RNA synthesis (26) (37). HCV-NS5b then switches to elongation mode, a regime in which it is known to processively copy RNAs as long as the viral genome (38). A third slow step has been reported by some groups in this transition to elongation, leading to accumulation of a five-nucleotide product (26). In our hands, this behavior is only detected when manganese is used instead of magnesium as the catalytic ion (not shown). With the (presumably physiological) magnesium then, the critical steps are those leading to de novo formation of a trinucleotide primer. The requirement of high NTP concentrations in these early steps of RNA synthesis by HCV NS5b has been extensively analyzed (4, 37). It has also been reported that high GTP concentrations selectively stimulate RNA synthesis (38) and that high levels of initiating GTP favor de novo RNA synthesis indirectly by repressing primer extension (10). In these two latter reports the authors used either a full-length HCV genome or a short RNA with a C residue at the 3′ end, respectively. Thus, it was difficult to identify precisely the initiation site or to discriminate between various implications of GTP in early steps of RNA synthesis as it was the initiating nucleotide. We reinvestigated the possible involvement of GTP during initiation with an A residue on RNA templates with a 3′-U. In vitro the recombinant NS5b preferentially initiates RNA synthesis on a C residue. However, the genomic RNA of all HCV genotypes harbors a U residue at its 3′ end, and the replication intermediate RNA (−) has a 3′-U residue in most of the genotype 2, 3, and 4 HCV strains. Furthermore, Cai et al. (39) have reported that in the context of a cell-based replicon system (genotype 1), the replicated genomic RNA has a 5′-A residue (hence, the replication intermediate RNA (−) has a 3′-U residue) even when the transfected genomic RNA initially has a 5′-G. In the present report we demonstrate that GTP also stimulates initiation of RNA synthesis independently of its incorporation in the initiation and in the elongation products (Fig. 4, C and D). This contribution to GTP stimulation does not stem from increased dinucleotide synthesis (Fig. 4C) but from stimulation of the transition to elongation (Fig. 4D).
FIGURE 8.
Schematic of the early steps in RNA synthesis by HCV-NS5b from the HCV minus strand in vitro. The methods used in deriving this model are indicated in boxed text on the left. Increasing details are given from top to bottom, starting with the most macroscopic level of overall de novo RNA synthesis and ending with the structural elements likely involved in the salient steps of dinucleotide formation and transition to elongation. The sequence of genotype 1 (−) strand is used as an example, but the activation by GTP of the transition to elongation is independent of its incorporation in neosynthesized RNA (as shown with the G3-U and G3–4U templates under “Results”). The underlined sequence in the RNA template is paired to a complementary sequence in the conserved, stable stem-loop SL-A1 at the 3′ end of the (−) strand.
Structural analyses of RdRps homologous to HCV-NS5b have shed much light on the molecular basis of these processes (40–42). They have highlighted the requirement for the first dinucleotide formation in de novo initiation of a platform to buttress the base of the priming nucleotide. Hence, this platform must be “in” for formation of the first dinucleotide primer, then move “out” before the addition of the third nucleotide to allow translocation of the nascent primer-template complex (Fig. 8). Structures of de novo initiation complexes unequivocally identifying the initiation platforms are available for only two RdRps; the phage φ6 polymerase (40) and the Reovirus polymerase (41). Strikingly, these two RdRps use different structural elements for the initiation platform. In the case of the φ6 polymerase, it is a tyrosine provided by a C-terminal domain homologous to the NS5b linker, whereas in the case of the Reovirus polymerase, the base of the priming GTP rests against a Gly-Ser motif inserted in the palm domain that retracts upon the transition from initiation to elongation. This double similarity with the H77_NS5b complex with GTP reported here (Fig. 1, B and C) has led us to investigate whether the linker might provide the initiation platform in the case of HCV-NS5b. Our functional data (Figs. 5 and 6) do argue for a direct involvement of the linker in formation of the first dinucleotide. The structural data, however, remain unconclusive (Fig. 1D). In the absence of the structure of a bona fide initiation complex for HCV-NS5b, the higher dinucleotide formation activity of the S556K mutants could in principle be ascribed to several alternative reasons. Chief among them would be the linker involvement in correct positioning of the template during initiation, a function to which the homologous C-terminal domain of the φ6 polymerase also participates. We have been unable, however, to bind a short single-stranded RNA in our crystals to either the H77 or J4 polymerase (wild type or S556K) despite a previous report of success with J4_NS5b (15). A conserved aromatic residue is found in Flavivirus and HCV polymerases at a position spatially very similar to the φ6 polymerase initiation platform (42). In those enzymes though, it is carried by the β-flap (Fig. 1B), a special feature of the Flaviviridae RdRp thumb. In the case of pestiviruses (the closest relative to HCV for which an RdRp structure is known) the β-flap is too short to reach deeply enough into the active site. On the basis of a complex with a GTP partially overlapping the priming site but with the ribose and base in a distinctly different position (as in our HCV-H77_NS5b complex with GTP), Choi et al. (43) have proposed that this specific GTP site might serve as stabilizer of the initiation complex, i.e. that the initiation platform might be provided by this GTP rather than directly by the Pestivirus polymerase. This hypothesis has been confirmed to some extent by D'Abramo et al. (44), who found that this GTP seemingly stabilizes the interaction between the 3′ end of the template and the priming nucleotide. A similar phenomenon could also explain our observations with the HCV-NS5b S556K mutants. Still quite elusive, however, is the structural basis of the selective stimulation by GTP of the transition to elongation reported here for HCV-NS5b. We previously described a surface GTP binding site in the HCV-BK_NS5b_Δ55 (28), and a role for this site in the transition to elongation has been suggested by Dutartre et al. (26) using the same 1b NS5b construct. However, amino acid 499, an integral part of this surface GTP binding site, is variable among HCV isolates. Thus, the BK isolate, like most 1b strains, harbors a valine at this position, whereas it is an alanine in H77 (as in most HCV strains of subtype other than 1b) and a threonine in J4. This and unfavorable crystal packing in the case of H77_NS5b may explain in part why we could not detect GTP binding at this site in our structures of J4_NS5b and H77_NS5b. Further studies are needed to identify the interaction between NS5b and GTP favoring the transition from initiation to elongation.
A feature almost universally found in de novo initiating RdRps is a C-terminal extension to the polymerase domain (the linker in HCV-NS5b) that occludes the exit for the double-stranded product from the catalytic site. The only known exceptions so far are the Flavivirus polymerase, which has no such extension (45, 46), and the Reovirus polymerase (41), for which the C-terminal domain (called “bracelet” domain) is not involved in de novo initiation and is wrought in such a way that it harbors a large channel, allowing exit of a double-stranded RNA. Remarkably, the C-terminal occlusion is even found in the few Picornaviridae-like RdRps that synthesize one strand de novo, whereas synthesis of the other strand follows the picornavirus RdRp “protein-primed” mechanism (42). This is the case among Caliciviridae of noroviruses and sapoviruses (47). The switch from de novo initiation to elongation requires that the C-terminal part move away to allow exit of the template-primer duplex, as long postulated for the φ6 polymerase and actually observed in the norovirus polymerase elongation complex (47). Indeed, HCV-NS5b is an extreme case in this respect. first, the C-terminal linker, although less bulky than the corresponding φ6 RdRp region, is directly connected to a transmembrane helix (Fig. 1E). This implies a considerable movement of the polymerase relative to the membrane and probably a complete rearrangement of the replication complex when the linker moves away from the catalytic cleft. Second, even with the linker removed, the thumb still has to rotate away to allow elongation, as proposed with the first crystal structure of NS5b_Δ55 (5), due in part but not only to the protrusion of the β-flap. Our SAXS data are strong evidence that the basal conformation of NS5b_Δ21 in solution is very close to that seen in crystal structures (Fig. 7). Consequently, the two slow steps detected in de novo initiation in solution can be interpreted in terms of conformational changes starting from these structures. The rate-limiting formation of the trinucleotide and subsequent processive elongation indicate that removal of the initiation platform is coincident with removal of the linker and opening of the thumb. More puzzling is the inefficiency of formation of the first dinucleotide. It indicates that simultaneous and productive binding of template and the first two nucleotides does not occur spontaneously in recombinant, isolated HCV-NS5b. This suggests that other cofactors in the replication complex in vivo are necessary for RNA synthesis initiation (though not for subsequent elongation) and that these requirements are only crudely substituted for by our in vitro assay conditions (particularly high nucleotide concentrations). Thus, not only is the basal state of HCV-NS5b not competent for primer elongation, it is not quite competent for initiation either. There are two roadblocks on the way to productive RNA synthesis, and both have to be lifted. This 2-fold negative regulation is to be related to the fact that HCV, like all Flaviviridae, produces a vast excess of RdRp and that only a small fraction of these must actually engage in RNA synthesis in cells (48).
Acknowledgments
We thank Dr. Jens Bukh for providing the pCV-H77 and the pCV-J4L6 molecular clones and Dr. Javier Perez, Dr. François Ternois, John E. LaVoy, and Nathalie Scrima for participation in early stages of this work. We are grateful to Drs. Patrice Vachette, Michel Castroviejo, Christian Cazenave, and Michel Ventura for helpful discussions and Drs. Célia Caillet-Saguy and Volker Lohmann for critical reading of the manuscript. We thank the Structural biology and Proteomics pole of the IMAGIF integrated platform for access to Crystallization and Mass Spectrometry services and Synchrotrons SOLEIL (beamline PROXIMA 1) and the European Synchrotron Radiation Facility for generous allocation of beamtime.
This work was supported, in whole or in part, by National Institutes of Health Grant RO1 CA63640 (to C. H. H.). This work was also supported in part by Agence Nationale de Recherche sur le Sida et les Hépatites Virales Grants AO 2008, CSS4 (to S. B.) and AO 2008, CSS4 (T. A.-G.) and predoctoral (to D. H. and K. M.) and postdoctoral (to P. C. S.) fellowships.
The atomic coordinates and structure factors (codes 2XI2, 2XI3, 2XHU, 2XHV, and 2XHW) have been deposited in the Protein Data Bank, Research Collaboratory for Structural Bioinformatics, Rutgers University, New Brunswick, NJ (http://www.rcsb.org/).
P. C. Simister and S. Bressanelli, unpublished information.
- HCV
- hepatitis C virus
- RdRp
- RNA-dependent RNA polymerase
- SAXS
- small-angle X-ray scattering
- nt
- nucleotide(s)
- 〈esd〉
- average estimated standard deviation.
REFERENCES
- 1.Moradpour D., Penin F., Rice C. M. (2007) Nat. Rev. Microbiol. 5, 453–463 [DOI] [PubMed] [Google Scholar]
- 2.Gosert R., Egger D., Lohmann V., Bartenschlager R., Blum H. E., Bienz K., Moradpour D. (2003) J. Virol. 77, 5487–5492 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ivashkina N., Wölk B., Lohmann V., Bartenschlager R., Blum H. E., Penin F., Moradpour D. (2002) J. Virol. 76, 13088–13093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Luo G., Hamatake R. K., Mathis D. M., Racela J., Rigat K. L., Lemm J., Colonno R. J. (2000) J. Virol. 74, 851–863 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ago H., Adachi T., Yoshida A., Yamamoto M., Habuka N., Yatsunami K., Miyano M. (1999) Structure 7, 1417–1426 [DOI] [PubMed] [Google Scholar]
- 6.Bressanelli S., Tomei L., Roussel A., Incitti I., Vitale R. L., Mathieu M., De Francesco R., Rey F. A. (1999) Proc. Natl. Acad. Sci. U.S.A. 96, 13034–13039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lesburg C. A., Cable M. B., Ferrari E., Hong Z., Mannarino A. F., Weber P. C. (1999) Nat. Struct. Biol. 6, 937–943 [DOI] [PubMed] [Google Scholar]
- 8.Biswal B. K., Cherney M. M., Wang M., Chan L., Yannopoulos C. G., Bilimoria D., Nicolas O., Bedard J., James M. N. G. (2005) J. Biol. Chem. 280, 18202–18210 [DOI] [PubMed] [Google Scholar]
- 9.Adachi T., Ago H., Habuka N., Okuda K., Komatsu M., Ikeda S., Yatsunami K. (2002) Biochim. Biophys. Acta 1601, 38–48 [DOI] [PubMed] [Google Scholar]
- 10.Ranjith-Kumar C. T., Gutshall L., Sarisky R. T., Kao C. C. (2003) J. Mol. Biol. 330, 675–685 [DOI] [PubMed] [Google Scholar]
- 11.Hong Z., Cameron C. E., Walker M. P., Castro C., Yao N., Lau J. Y., Zhong W. (2001) Virology 285, 6–11 [DOI] [PubMed] [Google Scholar]
- 12.Reigadas S., Ventura M., Sarih-Cottin L., Castroviejo M., Litvak S., Astier-Gin T. (2001) Eur. J. Biochem. 268, 5857–5867 [DOI] [PubMed] [Google Scholar]
- 13.Astier-Gin T., Bellecave P., Litvak S., Ventura M. (2005) FEBS J. 272, 3872–3886 [DOI] [PubMed] [Google Scholar]
- 14.Lou H., Choi Y. H., LaVoy J. E., Major M. E., Hagedorn C. H. (2003) Virology 317, 65–72 [DOI] [PubMed] [Google Scholar]
- 15.O'Farrell D., Trowbridge R., Rowlands D., Jäger J. (2003) J. Mol. Biol. 326, 1025–1035 [DOI] [PubMed] [Google Scholar]
- 16.Kabsch W. (2010) Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Vagin A., Teplyakov A. (2010) Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 [DOI] [PubMed] [Google Scholar]
- 18.Emsley P., Lohkamp B., Scott W. G., Cowtan K. (2010) Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Adams P. D., Afonine P. V., Bunkóczi G., Chen V. B., Davis I. W., Echols N., Headd J. J., Hung L. W., Kapral G. J., Grosse-Kunstleve R. W., McCoy A. J., Moriarty N. W., Oeffner R., Read R. J., Richardson D. C., Richardson J. S., Terwilliger T. C., Zwart P. H. (2010) Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Perrakis A., Harkiolaki M., Wilson K. S., Lamzin V. S. (2001) Acta Crystallogr. D Biol. Crystallogr. 57, 1445–1450 [DOI] [PubMed] [Google Scholar]
- 21.Schneider T. R. (2000) Acta Crystallogr. D Biol. Crystallogr. 56, 714–721 [DOI] [PubMed] [Google Scholar]
- 22.Konarev P. V., Volkov V. V., Sokolova A. V., Koch M. H., Svergun D. I. (2003) J. Appl. Crystallogr. 36, 1277–1282 [Google Scholar]
- 23.Svergun D. I., Barberato C., Koch M. H. (1995) J. Appl. Crystallogr. 28, 768–773 [Google Scholar]
- 24.Ferrer-Orta C., Arias A., Perez-Luque R., Escarmís C., Domingo E., Verdaguer N. (2004) J. Biol. Chem. 279, 47212–47221 [DOI] [PubMed] [Google Scholar]
- 25.Masante C., Mahias K., Lourenço S., Dumas E., Cahour A., Trimoulet P., Fleury H., Astier-Gin T., Ventura M. (2008) J. Gen. Virol 89, 212–221 [DOI] [PubMed] [Google Scholar]
- 26.Dutartre H., Boretto J., Guillemot J. C., Canard B. (2005) J. Biol. Chem. 280, 6359–6368 [DOI] [PubMed] [Google Scholar]
- 27.Schneider T. R. (2004) Acta Crystallogr. D Biol. Crystallogr. 60, 2269–2275 [DOI] [PubMed] [Google Scholar]
- 28.Bressanelli S., Tomei L., Rey F. A., De Francesco R. (2002) J. Virol. 76, 3482–3492 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Pawlotsky J. M., Chevaliez S., McHutchison J. G. (2007) Gastroenterology 132, 1979–1998 [DOI] [PubMed] [Google Scholar]
- 30.Yanagi M., Purcell R. H., Emerson S. U., Bukh J. (1997) Proc. Natl. Acad. Sci. U.S.A. 94, 8738–8743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yanagi M., St Claire M., Shapiro M., Emerson S. U., Purcell R. H., Bukh J. (1998) Virology 244, 161–172 [DOI] [PubMed] [Google Scholar]
- 32.Ellis D. A., Blazel J. K., Tran C. V., Ruebsam F., Murphy D. E., Li L. S., Zhao J., Zhou Y., McGuire H. M., Xiang A. X., Webber S. E., Zhao Q., Han Q., Kissinger C. R., Lardy M., Gobbi A., Showalter R. E., Shah A. M., Tsan M., Patel R. A., LeBrun L. A., Kamran R., Bartkowski D. M., Nolan T. G., Norris D. A., Sergeeva M. V., Kirkovsky L. (2009) Bioorg. Med. Chem. Lett. 19, 6047–6052 [DOI] [PubMed] [Google Scholar]
- 33.Rockway T. W., Zhang R., Liu D., Betebenner D. A., McDaniel K. F., Pratt J. K., Beno D., Montgomery D., Jiang W. W., Masse S., Kati W. M., Middleton T., Molla A., Maring C. J., Kempf D. J. (2006) Bioorg. Med. Chem. Lett. 16, 3833–3838 [DOI] [PubMed] [Google Scholar]
- 34.Cramer J., Jaeger J., Restle T. (2006) Biochemistry 45, 3610–3619 [DOI] [PubMed] [Google Scholar]
- 35.Hang J. Q., Yang Y., Harris S. F., Leveque V., Whittington H. J., Rajyaguru S., Ao-Ieong G., McCown M. F., Wong A., Giannetti A. M., Le Pogam S., Talamás F., Cammack N., Nájera I., Klumpp K. (2009) J. Biol. Chem. 284, 15517–15529 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Simister P., Schmitt M., Geitmann M., Wicht O., Danielson U. H., Klein R., Bressanelli S., Lohmann V. (2009) J. Virol. 83, 11926–11939 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ferrari E., He Z., Palermo R. E., Huang H. C. (2008) J. Biol. Chem. 283, 33893–33901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lohmann V., Overton H., Bartenschlager R. (1999) J. Biol. Chem. 274, 10807–10815 [DOI] [PubMed] [Google Scholar]
- 39.Cai Z., Liang T. J., Luo G. (2004) J. Virol. 78, 3633–3643 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Butcher S. J., Grimes J. M., Makeyev E. V., Bamford D. H., Stuart D. I. (2001) Nature 410, 235–240 [DOI] [PubMed] [Google Scholar]
- 41.Tao Y., Farsetta D. L., Nibert M. L., Harrison S. C. (2002) Cell 111, 733–745 [DOI] [PubMed] [Google Scholar]
- 42.Lescar J., Canard B. (2009) Curr. Opin. Struct. Biol. 19, 759–767 [DOI] [PubMed] [Google Scholar]
- 43.Choi K. H., Groarke J. M., Young D. C., Kuhn R. J., Smith J. L., Pevear D. C., Rossmann M. G. (2004) Proc. Natl. Acad. Sci. U.S.A. 101, 4425–4430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.D'Abramo C. M., Deval J., Cameron C. E., Cellai L., Götte M. (2006) J. Biol. Chem. 281, 24991–24998 [DOI] [PubMed] [Google Scholar]
- 45.Malet H., Egloff M. P., Selisko B., Butcher R. E., Wright P. J., Roberts M., Gruez A., Sulzenbacher G., Vonrhein C., Bricogne G., Mackenzie J. M., Khromykh A. A., Davidson A. D., Canard B. (2007) J. Biol. Chem. 282, 10678–10689 [DOI] [PubMed] [Google Scholar]
- 46.Yap T. L., Xu T., Chen Y. L., Malet H., Egloff M. P., Canard B., Vasudevan S. G., Lescar J. (2007) J. Virol. 81, 4753–4765 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zamyatkin D. F., Parra F., Alonso J. M., Harki D. A., Peterson B. R., Grochulski P., Ng K. K. (2008) J. Biol. Chem. 283, 7705–7712 [DOI] [PubMed] [Google Scholar]
- 48.Quinkert D., Bartenschlager R., Lohmann V. (2005) J. Virol. 79, 13594–13605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Schneider T. R. (2002) Acta Crystallogr. D Biol. Crystallogr. 58, 195–208 [DOI] [PubMed] [Google Scholar]