Abstract
Background
A positive family history for myocardial infarction (MI) is known to be a major cardiovascular risk factor. The current European guidelines therefore recommend intensified primary prevention for the siblings and children of persons who have had an MI. Although the genes underlying the heritable component of MI were largely unknown previously, the development of new molecular genetic methods, and particularly the advent of genome-wide association (GWA) studies, has led to the discovery of numerous genetic variants that are associated with an elevated risk of MI.
Methods
In this article, we review GWA studies on MI and coronary heart disease (CHD) that were retrieved by a selective literature search from 2007 onward. We comment on their implications for clinical practice.
Results
In the last three years, GWA studies have enabled the identification of many alleles that confer a higher risk of MI. A total of eleven chromosomal regions have been replicated and associated with the disease, and their functional significance has been studied. Furthermore, it has been shown that some of the manifestations of CHD, e.g., calcification, ectasia and main-stem stenosis, are more strongly inherited than others.
Conclusion
The results of recent GWA studies for MI and CHD will aid in individual risk prediction for MI by molecular biological means. They will also permit the development of new approaches to research on the pathophysiology of myocardial infarction.
So-called candidate genes, i.e., molecular changes that lead to premature atherosclerosis, have been sought for the last two decades. All studies of this nature are based on the assumption that proteins involved in the pathophysiology of atherosclerosis carry mutations that make them a potential contributory cause of myocardial infarction. Although over 5000 publications have appeared in this field to date, only a few of them report a reproducible association of genes involved in lipid metabolism (table 1) with an elevated risk of myocardial infarction (1).
Table 1. Common genetic variations in genes involved in LDL-cholesterol metabolism that have an effect on the risk of myocardial infarction.
| Gene | Frequency of allele conferring elevated risk | Rise in LDL-cholesterol per copy of allele conferring elevated risk | Effect of allele on risk of myocardial infarction (odds ratio, OR) | p-value for association with myocardial infarction | Reference |
| PCSK9 | 80% | + 15% | OR 1.13 | 0.004 | (21) |
| Apo E | 18% | + 14% | OR 1.17 | 0.0001 | (21) |
| LDLR | 11% | + 6% | OR 1.18 | 0.0001 | (19) |
| Apo B | 33% | + 5% | OR 1.1 | 0.004 | (20) |
OR, odds ratio for one allele; PCSK9, proprotein convertase subtilisin/kexin type 9; Apo E, apolipoprotein E; LDLR, low-density lipoprotein receptor; Apo B, apolipoprotein B The differing values for statistical significance are due to large differences in sample size
From the genetic point of view, it should be pointed out that most candidate gene analyses have involved only one or a few of the many molecular variants of an individual gene. Moreover, a statistical problem besetting many such studies is the small size and heterogeneity of the sample. Thus, in retrospect, it comes as no surprise that only a small amount of valid data has been derived from studies of this type.
The breakthrough came in 2007 with the first use of genome-wide association (GWA) studies (figure 1). This type of analysis enables the characterization of approximately 2.4 million genetic markers in each subject. A systematic comparison of these markers in thousands of patients with myocardial infarction and in healthy control subjects has enabled the identification of gene segments whose influence on the risk of atherosclerosis is considered to be established beyond any doubt (table 2). Moreover, the main findings were replicated in many thousands of patients and control subjects. To date, the contribution of eleven chromosome segments (eleven genes) to the risk of myocardial infarction has been shown reproducibly and beyond doubt (2– 8). This review article summarizes all of the GWA studies on myocardial infarction that have been published up to the present; these studies were found by a selective literature search (www.ncbi.nih.gov/pubmed).
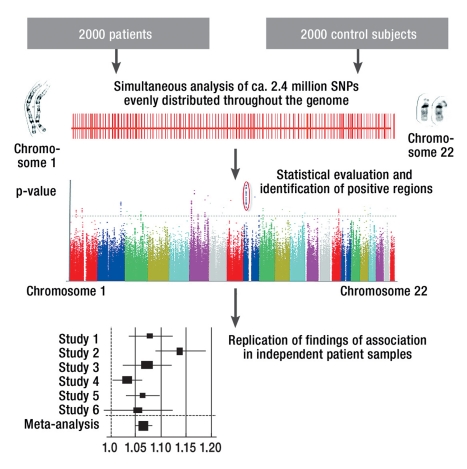
Figure 1.
The principle of genome-wide association studies
With the aid of DNA chips, the allele frequencies of up to 2.4 million polymorphisms are statistically compared in samples of patients and normal control subjects.
If markers in a particular chromosome segment are present in significantly different frequency, this finding is replicated in further samples. Then, if a highly significant difference between cases and controls is found (p < 5 × 10–8), the finding is considered relevant, even though multiple statistical tests have been applied. At present, only the SNPs (single nucleotide polymorphisms) on the autosomal chromosomes, i.e., chromosomes 1 through 22, are analyzed in genome-wide association studies. The sex chromosomes X and Y are not included in the analyses, as there is no standardized analytic technique for these chromosomes to date.
Table 2. Overview of the gene regions associated with an elevated risk of myocardial infarction that have been identified to date by genome-wide association studies.
| Chromosome segment | SNP | Frequency of allele conferring increased risk in Europeans | OR (95% CI) | p-value | Genes | Association with traditional risk factor? | Function of genes | References |
| 1p13.3 | rs599839 | 77% | 1.13 (1.08–1.19) | 1.1 x 10–14 | PSCR1, CELSR2, SORT1, MYBPHL | yes/LDL | LDL increase | (21, 22) |
| 1q41 | rs3008621 | 72% | 1.10 (1.04–1.17) | 1.4 x 10–9 | MIA3 | no | Collagen processing | (3, 21, 22) |
| 2q33 | rs6725887 | 14% | 1.17 (1.11–1.23) | 1 x 10–8 | WDR12 | no | Apoptosis | (3) |
| 3q22.3 | rs9818870 | 15% | 1.15 (1.11–1.19) | 7.4 x 10–13 | MRAS | no | Adhesion signaling | (2) |
| 6p24 | rs12526453 | 65% | 1.13 (1.08–1.17) | 1 x 10–9 | PHACTR1 | no | Coronary calcification | (3) |
| 6q26–27 | rs2048327 | 18% | 1.20 (1.13–1.28) | 1.2 x 10–9 | SLC22A3, LPAL2, LPA | yes/Lp(a) | Lp(a) | (4) |
| rs3127599 | ||||||||
| rs7767084 | ||||||||
| rs10755578 | ||||||||
| 9p21.3 | rs1333049 | 52% | 1.36 (1.27–1.46) | 2.91 x 10–19 | MTAP, CDKN2A, CDKN2B, ANRIL | no | Unknown | (5, 6, 21, 23) |
| 10q11 | rs501120 | 84% | 1.11 (1.05–1.18) | 9.5 x 10–8 | SDF1 | no | EPC recruiting and inflammation | (21, 22) |
| 12q24 | rs11065987 | 34% | 1.14 (1.10–1.19) | 5.2 x 10–11 | SH2B3 | no | Unknown | (7, 8) |
| 12q24.3 | rs2259816 | 36% | 1.08 (1.05–1.11) | 4.8 x 10–7 | HNF1A, C12orf43 | no | Unknown | (2) |
| 21q22 | rs9982601 | 13% | 1.19 (1.14–1.27) | 6 x 10–11 | SLC5A3, MRPS6, KCNE2 | no | Unknown | (3) |
OR, odds ratio; CI, confidence interval; SNP, single nucleotide polymorphism
The significance of the new myocardial infarction genes
An overview of the findings enables us to make the following basic statements:
The strongest genetic effect on the risk of myocardial infarction is that of a segment on chromosome 9p21.3 (9). This risk locus for myocardial infarction has now been replicated in more than 45 000 cases and 85 000 controls. Interestingly, this chromosome region does not contain any gene coding for a protein. The underlying pathogenetic mechanism remains unclear.
Only two of the eleven chromosome segments associated with myocardial infarction are related to traditional risk factors (LDL-cholesterol and LP[a]). Thus, most of the genetic variants that have been described, including the one located on chromosome 9p21.3, elevate the risk of myocardial infarction through pathogenetic mechanisms that are not yet understood.
All of the risk alleles identified to date are relatively common. The probability of a person of European descent having one, or even two, risk alleles on chromosome 9p21.3 is about 50% and 25%, respectively. Only a minority of the European population does not possess a copy of this allele.
At the individual level, each affected chromosome segment has only a small positive effect on the risk of myocardial infarction, on the order of 10% to 30% for each risk allele. This means, for example, that the 25% of persons in the European population who are homozygous for the risk allele on chromosome 9p21.3 have a risk of myocardial infarction that is roughly 60% higher than that of the 25% of persons who do not bear the risk allele at all. Among men aged 40 to 45 who do not have any of the traditional risk factors (elevated LDL-cholesterol values, arterial hypertension, overweight, or smoking), the average 10-year risk of myocardial infarction is approximately 1% (PROCAM score), yet an average person in this group also possesses one risk allele of chromosome 9p21.3.
Although the risk increase for an individual person is relatively small, the high frequency of the risk alleles leads to an enormous increase in the incidence of myocardial infarction at the level of the entire population. For example, among one million homozygous carriers of the risk allele on chromosome 9p21.3 aged 40 to 45, the absolute risk increase of 0.6% leads to 6000 additional heart attacks over a ten-year period. Practically all persons of European descent have one or more of the risk alleles listed in table 2 in varying configurations, and thus a higher or lower inherited risk of myocardial infarction.
The genetic predisposition conferred by the risk alleles is seen both in persons with a positive family history and in persons with a negative family history. Thus, the genetic effect of the newly identified variants adds to that of a positive family history. Stated in another way, a positive family history reflects a genetic effect of a type that has not yet been explained (figure 2).
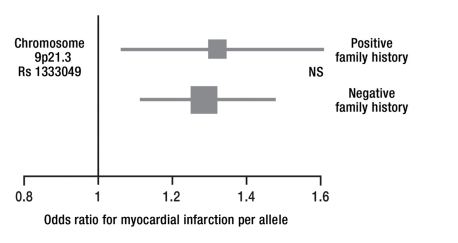
Figure 2.
Chromosome 9p21.3 and family history
One risk allele raises the risk of myocardial infarction by a factor of 1.3, i.e., by approximately 30%. The strength of this effect is comparable in persons with a positive or negative family history; in other words, the effect of the risk allele is additive to that of a positive family history. The size of the squares in the diagram reflects the sample size. A total of 4500 persons were studied. NS, not significant.
Genetic prediction
Despite the enormous progress made in the last two years, only 10% of the genetic influence on myocardial infarction has been accounted for to date (3). Under the leadership of the Lübeck team, a globally operating consortium called CARDIoGRAM has been established, which collates the genome-wide association studies performed to date on a total of some 100 000 subjects, thus enabling a new quantitative dimension of analysis. The list of known and definitively evaluated genetic variants that are associated with myocardial infarction can be expected to become much longer in the near future.
The systematic, genome-wide analysis of cardiovascular risk factors has also led to the identification of many genes that influence lipid metabolism, arterial blood pressure, the risk of diabetes mellitus, and the tendency toward obesity. Thus, light has not just been shed on the genetics of myocardial infarction; it is now possible to predict more accurately, on a genetic basis, whether an individual will develop one of the classic cardiovascular risk factors over the course of a lifetime. Taking all of the data into consideration, one can conclude that it should now be possible to predict cardiovascular risk more accurately than is currently done on the basis of the family history alone (10). At present, large-scale epidemiological studies are being performed to determine whether the knowledge gained from GWA studies of myocardial infarction can improve the prognostication of future events beyond what is already possible from consideration of the traditional risk factors. If genetic information does turn out to have additional predictive value, then, in the future, it should be possible to develop risk scores that discriminate low-, middle-, and high-risk patients more accurately than currently possible, with the result that primary preventive measures can be applied in a more targeted fashion. We must await the results of these prospective studies before we can say whether the knowledge of the risk alleles that a person carries will lead to a clinically relevant improvement in our ability to predict myocardial infarction.
Myocardial infarction families
The genome-wide association studies have thus succeeded in identifying a large number of factors that make a major contribution to the risk of myocardial infarction at the population level. These findings do not yet account for the small number of families in which an autosomal dominant inheritance pattern for myocardial infarction has been found (11– 13). In these families, a genetic factor must be at work that elevates the risk of myocardial infarction so massively that nearly every individual possessing the risk-conferring gene, i.e., every other child of an affected person, actually goes on to have a myocardial infarction (figure 3).
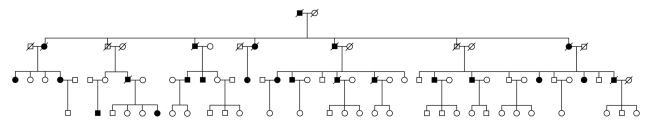
Figure 3.
A family tree from the German Myocardial Infarction Families Study (Deutsche Herzinfarktfamilienstudie)
Roughly every other child of an affected person of either sex develops coronary heart disease. Thus, autosomal dominant inheritance is suspected.
Symbols: squares/circles = men/women; filled/open symbols = affected/unaffected individuals; line drawn through symbol = deceased
Although it has been possible in these families, as in others, to identify chromosome segments associated with a heritable risk of myocardial infarction, the genes and genetic variants responsible for the autosomal dominant inheritance of myocardial infarction in these families have not yet been found. This will be a major challenge for the future, as a better understanding of the underlying mechanisms in these families might turn out to be of profound importance for the prevention of myocardial infarction in general.
Myocardial infarction
Coronary heart disease as a specific illness, and atherosclerosis in general, can affect patients in a variety of characteristic patterns. It has been demonstrated that the risk allele on chromosome 9p21.3 raises the risk not only of coronary heart disease, but also of stroke, aneurysms, and peripheral vascular occlusive disease (14, 15). These diseases probably all have their own, individual genetic risk factors, in addition to the common genetic risk factor that has been identified. Thus, a number of studies have revealed that calcification of the coronary arteries or of the aorta is subject to a major influence from genetic factors (16). Although no particular genetic factor of this type has yet been identified in man, the murine ABCC6 gene has been shown to cause vascular calcification by way of an abnormality of cellular transport (17, 18). It is hoped that a better understanding of the underlying mechanisms will help explain why some patients have severe vascular calcification and will lead the way toward effective prevention of this problem.
Coronary main-stem stenosis has been found to be a particularly heritable condition (figure 4). That is, narrowing at the ostia or in the first centimetres of the right or left coronary artery is subject to a stronger genetic influence than more distally located stenosis. As a result, this highly dangerous manifestation of coronary heart disease also has a high rate of occurrence in siblings and other first-degree relatives of affected persons. Unfortunately, the mechanisms of inheritance of main-stem stenosis remains unknown. Thus, the relatives of affected persons can be advised at present only to have themselves thoroughly evaluated and to undertake a strict regimen to modify any risk factors that may be present.
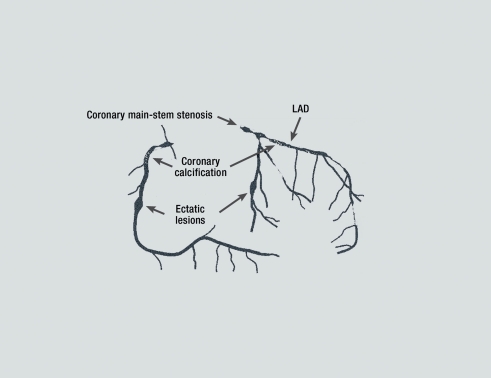
Figure 4.
Heritability of main-stem stenosis. Main-stem stenosis, coronary calcification, and ectatic coronary lesions are highly heritable (16); LAD, left anterior descending coronary artery
Future prospects
These rapid developments in the discovery of molecular causes of myocardial infarction lead us to expect new means of risk prediction to become available in the near future. The current data show that most of these genetic effects operate independently of the traditional risk factors (lipid metabolism, arterial hypertension, diabetes, body-mass index, and smoking). The scientific data have already improved our ability to prognosticate the risk of myocardial infarction in a population sample, but it remains unclear at present whether knowledge of the associated genetic variants will be useful for risk prediction, and thus for prevention, for individual patients.
The findings described here have shed new light on the pathophysiology of myocardial infarction. In principle, genetic variants that confer risk are always to be considered as causal cofactors (like the risk alleles listed in table 2), or even as precipitating factors (in the case of the autosomal dominant forms), of atherosclerosis and myocardial infarction. But how do these risk alleles work, if not by way of the traditional risk factors? Once the underlying mechanisms are deciphered, it will probably be possible to develop new methods of treating and preventing coronary heart disease. The current spirit of optimism among researchers in the field is reminiscent of the time when LDL-cholesterol had been identified as a risk factor for myocardial infarction, but no treatment was yet available to lower the associated risk. The total, summated effect of the genetic risk variants that have now been discovered seems to be far greater than that of an elevated LDL-cholesterol level. It remains to be seen, however, whether a better understanding of the genetics of myocardial infarction will lead to the development of new preventive measures whose effect will be comparable to that of the statins in lowering the risk due to high LDL-cholesterol.
Key Messages.
Genome-wide association studies have led to the discovery of a large number of new myocardial infarction genes.
The strongest genetic effect on the risk of myocardial infarction is that of a variant found on chromosome 9p21.
Only a few of the myocardial infarction genes are associated with classic risk factors.
Most of the risk-conferring variants are common (frequency up to 84% for the risk allele, compared with 16% for the “protective” allele) and lead to a low individual elevation of risk (relative risk increase, 10% to 30%).
Prospective studies on risk prediction with genetic scores have yet to be published.
Acknowledgments
Translated from the original German by Ethan Taub, M.D.
Footnotes
Conflict of interest statement
The authors state that they have no conflict of interest as defined by the guidelines of the International Committee of Medical Journal Editors.
References
- 1.Mayer B, Erdmann J, Schunkert H. Genetics and heritability of coronary artery disease and myocardial infarction. Clin Res Cardiol. 2007;96:1–7. doi: 10.1007/s00392-006-0447-y. [DOI] [PubMed] [Google Scholar]
- 2.Erdmann J, et al. New susceptibility locus for coronary artery disease on chromosome 3q223. Nat Genet. 2009;41:280–282. doi: 10.1038/ng.307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kathiresan S, et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat Genet. 2009;41:334–341. doi: 10.1038/ng.327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tregouet DA, et al. Genome-wide haplotype association study identifies the SLC22A3-LPAL2-LPA gene cluster as a risk locus for coronary artery disease. Nat Genet. 2009;41:283–285. doi: 10.1038/ng.314. [DOI] [PubMed] [Google Scholar]
- 5.Helgadottir A, et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science. 2007;316:1491–1493. doi: 10.1126/science.1142842. [DOI] [PubMed] [Google Scholar]
- 6.McPherson R, et al. A common allele on chromosome 9 associated with coronary heart disease. Science. 2007;316:1488–1491. doi: 10.1126/science.1142447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gudbjartsson DF, et al. Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat Genet. 2009;41:342–347. doi: 10.1038/ng.323. [DOI] [PubMed] [Google Scholar]
- 8.Soranzo N, et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat Genet. 2009;41:1182–1190. doi: 10.1038/ng.467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Schunkert H, et al. Repeated replication and a prospective meta-analysis of the association between chromosome 9p213. and coronary artery disease. Circulation. 2008;117:1675–1684. doi: 10.1161/CIRCULATIONAHA.107.730614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kathiresan S, et al. Polymorphisms associated with cholesterol and risk of cardiovascular events. N Engl J Med. 2008;358:1240–1249. doi: 10.1056/NEJMoa0706728. [DOI] [PubMed] [Google Scholar]
- 11.Lieb W, et al. Lack of association between the MEF2A gene and myocardial infarction. Circulation. 2008;117:185–191. doi: 10.1161/CIRCULATIONAHA.107.728485. [DOI] [PubMed] [Google Scholar]
- 12.Wang L, Fan C, Topol SE, Topol EJ, Wang Q. Mutation of MEF2A in an inherited disorder with features of coronary artery disease. Science. 2003;302:1578–1581. doi: 10.1126/science.1088477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mani A, et al. LRP6 mutation in a family with early coronary disease and metabolic risk factors. Science. 2007;315:1278–1282. doi: 10.1126/science.1136370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gschwendtner A, et al. Sequence variants on chromosome 9p213. confer risk for atherosclerotic stroke. Ann Neurol. 2009;65:531–539. doi: 10.1002/ana.21590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Helgadottir A, et al. The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat Genet. 2008;40(2):217–224. doi: 10.1038/ng.72. [DOI] [PubMed] [Google Scholar]
- 16.Fischer M, et al. Distinct heritable patterns of angiographic coronary artery disease in families with myocardial infarction. Circulation. 2005;111:855–862. doi: 10.1161/01.CIR.0000155611.41961.BB. [DOI] [PubMed] [Google Scholar]
- 17.Aherrahrou Z, et al. An alternative splice variant in Abcc6, the gene causing dystrophic calcification, leads to protein deficiency in C3H/He mice. J Biol Chem. 2008;283:7608–7615. doi: 10.1074/jbc.M708290200. [DOI] [PubMed] [Google Scholar]
- 18.Aherrahrou Z, et al. Ultrafine mapping of Dyscalc1 to an 80-kb chromosomal segment on chromosome 7 in mice susceptible for dystrophic calcification. Physiol Genomics. 2007;28:203–212. doi: 10.1152/physiolgenomics.00133.2006. [DOI] [PubMed] [Google Scholar]
- 19.Linsel-Nitschke P, et al. Lifelong reduction of LDL-cholesterol related to a common variant in the LDL-receptor gene decreases the risk of coronary artery disease—a Mendelian Randomisation study. PLoS One. 2008;3 doi: 10.1371/journal.pone.0002986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Willer CJ, et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat Genet. 2008;40:161–169. doi: 10.1038/ng.76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Samani NJ, et al. Genomewide association analysis of coronary artery disease. N Engl J Med. 2007;357:443–453. doi: 10.1056/NEJMoa072366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Samani NJ, et al. Large scale association analysis of novel genetic loci for coronary artery disease. Arterioscler Thromb Vasc Biol. 2009;29:774–780. doi: 10.1161/ATVBAHA.108.181388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Anonymous. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661–678. doi: 10.1038/nature05911. [DOI] [PMC free article] [PubMed] [Google Scholar]