Abstract
The recently discovered heliobacteria are the only Gram-positive photosynthetic bacteria that have been cultured. One of the unique features of heliobacteria is that they have properties of both the photosynthetic green sulfur bacteria (containing the type I reaction center) and Clostridia (forming heat-resistant endospores). Most of the previous studies of heliobacteria, which are strict anaerobes and have the simplest known photosynthetic apparatus, have focused on energy and electron transfer processes. It has been assumed that like green sulfur bacteria, the major carbon flow in heliobacteria is through the (incomplete) reductive (reverse) tricarboxylic acid cycle, whereas the lack of CO2-enhanced growth has not been understood. Here, we report studies to fill the knowledge gap of heliobacterial carbon metabolism. We confirm that the CO2-anaplerotic pathway is active during phototrophic growth and that isoleucine is mainly synthesized from the citramalate pathway. Furthermore, to our surprise, our results suggest that the oxidative (forward) TCA cycle is operative and more active than the previously reported reductive (reverse) tricarboxylic acid cycle. Both isotopomer analysis and activity assays suggest that citrate is produced by a putative (Re)-citrate synthase and then enters the oxidative (forward) TCA cycle. Moreover, in contrast to (Si)-citrate synthase, (Re)-citrate synthase produces a different isomer of 2-fluorocitrate that is not expected to inhibit the activity of aconitase.
Keywords: Bacteria, Isotopic Tracers, Metabolism, Metabolomics, Photosynthesis, Tricarboxylic Acid (TCA) Cycle, Amino Acid Biosynthesis, Citrate Synthase, Fluoroacetate
Introduction
Heliobacteria are a relatively newly discovered group of anaerobic photosynthetic bacteria. All of the cultured heliobacteria require organic carbon for anoxygenic growth, and several of the species can fix nitrogen (1, 2). Heliobacteria are the only cultured Gram-positive photosynthetic bacteria and are phylogenetically related to the bacterial phylum Firmicutes, such as the aerobic Bacillus and anaerobic Clostridia (1). Among 10 cultured heliobacteria (2), Heliobacterium modesticaldum is the only one that can grow at temperatures of >50 °C. Madigan and co-workers (1, 3) reported that pyruvate, lactate, acetate (+HCO3−), or yeast extract can support the phototrophic growth of H. modesticaldum, and our recent studies demonstrated that d-sugars can also support the growth of H. modesticaldum (4).
An unusual feature of heliobacteria is that they have properties of both green sulfur bacteria (containing the type I reaction center) and Clostridia (forming heat-resistant endospores) (1). Heliobacteria have the simplest photosynthetic apparatus among the photosynthetic organisms (5), and most of the research on them has been focused on understanding the photosynthetic machinery and energy transfer processes (1, 6–8). In contrast, our knowledge of phototrophic carbon metabolism by heliobacteria is still poorly developed.
Only two reported studies have experimentally probed the carbon metabolism of heliobacteria; one is our recent study with H. modesticaldum (4), and the other is the 1994 report by Kelly and co-workers (9) on Heliobacterium strain HY-3. Our studies highlighted the critical roles that pyruvate plays during phototrophic and chemotrophic growth of H. modesticaldum, reported on three new carbon sources for H. modesticaldum, and verified the genomic information. Kelly and co-workers (9) assayed activity for the enzymes in the TCA cycle and employed 13C NMR to analyze 13C-labeled protein-based amino acids using [2-13C]pyruvate or [2-13C]acetate. Furthermore, the genes aclAB, encoding ATP citrate lyase (ACL),4 and gltA, encoding citrate synthase (CS), have not been annotated in the H. modesticaldum genome (5), and neither we (4) nor Kelly and co-workers (9) detected the enzymatic activities for ACL or CS. Together, previous studies suggested that the RTCA cycle is not complete in H. modesticaldum (Fig. 1A) (4, 5) and Heliobacterium strain HY-3 (9).
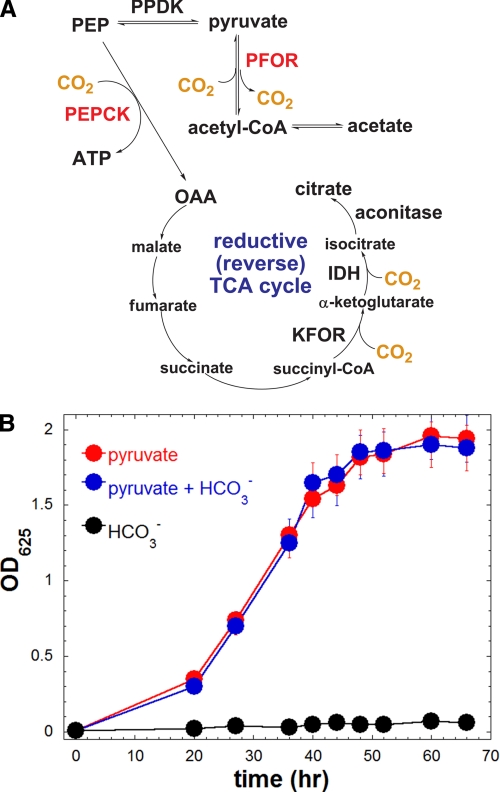
FIGURE 1.
A, previously proposed carbon flow in H. modesticaldum from genomic information and enzymatic activity assays. B, phototrophic growth curve of H. modesticaldum grown on pyruvate, HCO3−, or pyruvate and HCO3− as the defined carbon source. Cell density was estimated by measuring the optical density (OD) at 625 nm (see the “Experimental Procedures”). PFOR, pyruvate:ferredoxin oxidoreductase; PEPCK, phosphoenolpyruvate carboxykinase; KFOR, α-KG:ferredoxin oxidoreductase; IDH, isocitrate dehydrogenase.
Note that 13C NMR may be insufficient for generating complete labeling information of key metabolites, as 13C NMR cannot directly detect labeled carbon on the carboxyl group of an amino acid if its α-carbon is not labeled (10). More importantly, essential questions remain unresolved for carbon metabolism of heliobacteria. (a) If the (incomplete) RTCA cycle is employed for biomass production in heliobacteria (Fig. 1A), then CO2 is essential for phototrophic growth (a metabolic type similar to the green sulfur bacteria that employ an RTCA cycle) (11). However, no apparent CO2-enhanced growth has been observed for pyruvate-grown heliobacterial cultures. (b) Some 13C-labeled patterns in the study by Kelly and co-workers (9) suggested that the oxidative (forward) TCA (OTCA) cycle is operative. If this is the case, it is difficult to understand how the OTCA cycle could possibly be initiated without the encoding genes or enzymatic activities of ACL or CS.
This study is intended to apply both biochemical methods and 13C-based metabolite analysis via mass spectrometry to address those essential questions by investigating how the TCA cycle and amino acid biosynthesis are performed by H. modesticaldum. Our results identify the missing link for the carbon metabolism of H. modesticaldum (and perhaps Heliobacterium strain HY-3) and suggest that the carbon flow of H. modesticaldum is more akin to Clostridia than to the green sulfur bacteria.
EXPERIMENTAL PROCEDURES
Materials
Chemicals and enzymes were obtained from Sigma. The 13C-labeled sodium bicarbonate, [1-13C]- and [3-13C]pyruvate were obtained from Cambridge Isotope Laboratories, Inc. The DNA oligomers were obtained from Integrated DNA Technology without further purification.
Cell Cultures
The cell cultures employed in this study are listed below. Cell density was estimated by measuring the optical density (OD) of a suspension of cells at 625 nm (OD625) for the cell growth of both H. modesticaldum and Roseobacter denitrificans because absorbance of photosynthetic pigments is minimal around 625 nm (4), and OD600 was measured to estimate the growth of Desulfovibrio vulgaris Hildenborough (DvH), in which no photosynthetic pigment was present. Autoclaved subcultures were used as negative controls, and all of the experiments were performed in triplicate.
H. modesticaldum
The H. modesticaldum strain Ice1T culture was provided by Dr. Michael T. Madigan (Southern Illinois University, Carbondale) and was grown phototrophically in a minimal medium (1, 4) with pyruvate (20 mm) and (±)HCO3− (20 mm) as the carbon sources. The fresh medium was inoculated with 1–2% cell culture in the late exponential growth phase. Physiological studies with fluoroacetate (FAc) for the pyruvate-grown cultures were performed with 20 mm pyruvate and 20 mm FAc included in the growth medium. The cultures were grown inside the anaerobic chamber (Coy) in low intensity light (100 ± 10 μmol/m2/s) at 46–48 °C.
R. denitrificans
The R. denitrificans OCh114 culture, obtained from Dr. J. T. Beatty (University of British Columbia, Vancouver, Canada), was grown aerobically in a defined medium with pyruvate (20 mm), FAc (20 mm), or pyruvate (20 mm) + FAc (20 mm) as the sole carbon sources. The defined growth medium was prepared as reported (12). The cultures were grown at 28 °C (20–30%-filled Erlenmeyer flask) with shaking at 150 rpm in the dark.
D. vulgaris Hildenborough (DvH)
DvH was kindly provided by Dr. Terry C. Hazen (Lawrence Berkeley National Laboratory, Berkeley, CA). DvH was grown in a defined growth medium containing (per liter) NaCl (1.0 g), MgCl2·6H2O (0.5 g), KH2PO4 (0.2 g), NH4Cl (0.3 g), KCl (0.3 g), CaCl2·2H2O (0.015 g), MgSO4·7H2O (0.2 g), a trace element solution (1 ml) (13), a Na2SeO3/Na2WO4 solution (1 ml) (14), resazurin (10 mg), lactate (5 mm), Na2SO4 (5 mm), and with a N2/CO2 (80:20, v/v) headspace. After autoclaving and cooling to room temperature, the medium was supplied with a vitamin mixture solution (0.5 ml) (15). DvH was subcultured into fresh medium in triplicate with 1% (v/v) inoculate, and cell cultures were grown at 30 °C anaerobically. FAc amended cultures were prepared in the same content except FAc (1.5 mm) was added before inoculation.
RNA Extraction and Quantitative Real Time PCR
The transcriptomic profiling data were generated using the approaches described elsewhere (16). The primer sequences for QRT-PCR are listed in supplemental Table S1.
Isotopomer Analysis via GC-MS
The isotopomer analysis was performed using the methods reported previously (17) and is described in the supplemental material.
Activity Assays for Citrate Synthase
Enzymatic activities were performed in triplicate with cell-free extracts that were prepared using the procedure as described earlier (4). Protein concentration in cell extracts was determined by the Bradford assay (18) using bovine serum albumin as the standard. The reaction turnover for the reaction catalyzed by CS was followed by the formation of 5,5′-dithiobis-2-nitrobenzoic acid or Ellman's reagent-modified CoA using the method reported previously (9, 19) with minor modification. To minimize oxygen in the assay mixture for measuring CS activity, all of the reagents (5,5′-dithiobis-2-nitrobenzoic acid, metal ions), substrates (acetyl-CoA, oxaloacetate), and buffer were prepared with oxygen-free water in an anaerobic chamber (Coy), and the assay components were mixed in a tightly sealed cuvette inside the anaerobic chamber prior to removal for the spectral measurements. The reaction was initiated by the addition of oxaloacetate (OAA) (0.5 mm), and the increase at A412 was followed to monitor the activity. The amount of CoA production was estimated using a standard calibration curve with β-mercaptoethanol. The control experiment was performed with the same procedure with water rather than OAA added to initiate the reaction.
RESULTS
Carbon Metabolism in H. modesticaldum
In this study, we probed the central carbon metabolic pathway with the following approaches: (i) physiological studies with fluoroacetate; (ii) isotopomer data; (iii) mass spectrum of photosynthetic pigments; and (iv) transcriptomic profiles. Pyruvate is the best known organic carbon source for supporting the phototrophic growth of H. modesticaldum (3, 4) and several other heliobacteria (1), and it was used for probing the carbon metabolism of H. modesticaldum.
Physiological Studies with Fluoroacetate
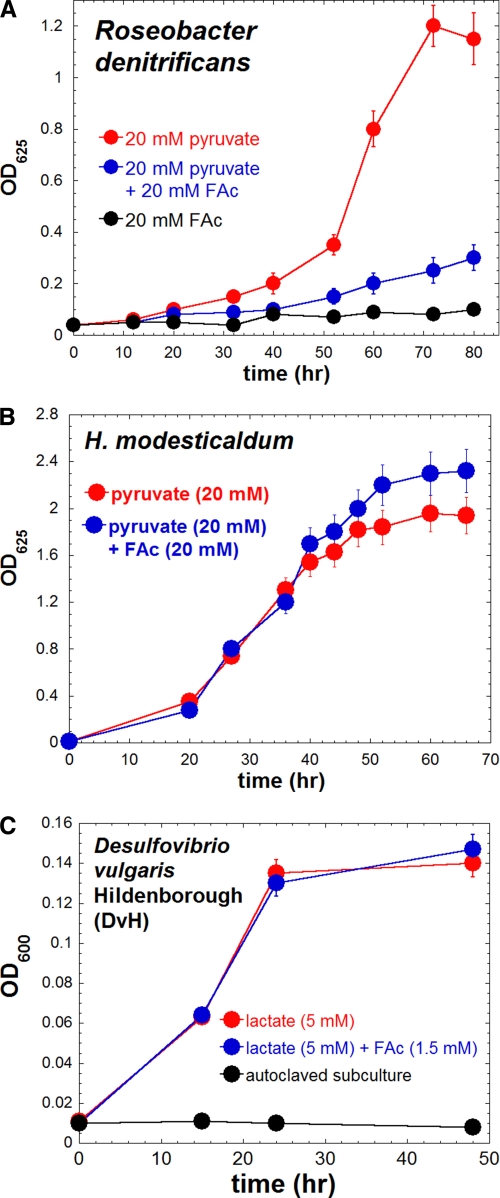
FAc has been reported as a metabolic toxin. The toxicity of FAc is generally recognized to arise from the fact that the carbon flow in the OTCA cycle is blocked through the inhibition of aconitase by (−)-erythro-(2R,3R)-2-F-citrate (2-FC), which is synthesized from F-acetyl-CoA and OAA by CS (20). Consistent with this hypothesis, an aerobic anoxygenic photoheterotrophic bacterium R. denitrificans, which has been known to have an active OTCA cycle (12), was notably inhibited by FAc (Fig. 2A). In contrast, the growth of H. modesticaldum on pyruvate was similar with FAc versus without FAc (Fig. 2B). It is known that FAc, an acetate analog, can be taken up by H. modesticaldum and converted into F-acetyl-CoA, because the acsA gene, encoding acetyl-CoA synthetase (ACS), has been annotated and the enzymatic activity of ACS has been reported (4). Furthermore, acetate (+HCO3−) is also known to support the phototrophic growth of H. modesticaldum (1, 3, 4). Consistent with the physiological studies, the transcript levels of genes for carbon metabolism are similar (within ∼2 ΔΔCT) for cultures grown on pyruvate (20 mm) and with or without the addition of FAc (20 mm) (Table 1). Together, our studies indicate little effect of FAc on the growth of H. modesticaldum, implying that (−)-erythro-2-FC cannot be synthesized in H. modesticaldum, in agreement with the findings that no genes encoding CS and ACL have been annotated in the genome (5), and no enzymatic activities of CS or ACL have been detected for H. modesticaldum (4).
FIGURE 2.
Effect of FAc for the growth of R. denitrificans, H. modesticaldum, and D. vulgaris Hildenborough (DvH). A, growth curve of R. denitrificans grown on pyruvate, FAc, or pyruvate and FAc as the carbon source; B, growth curve of H. modesticaldum during photoheterotrophic growth on pyruvate with or without FAc; C, growth curve of DvH grown on lactate with or without FAc.
TABLE 1.
The transcript level of the genes for carbon metabolism of H. modesticaldum during photoheterotrophic growth on pyruvate with or without FAc
The genes encoding the enzymes in the (R)TCA cycle are highlighted in boldface italic.
| Gene | ΔCTa (pyruvate) | ΔCTa (pyruvate + FAc) | ΔΔCT (ΔCT in pyruvate − ΔCT in pyruvate + FAc) | relative expression level (pyruvate vs. pyruvate + FAc) = 2ΔΔCT |
|---|---|---|---|---|
| pykA (HM1_0076, pyruvate kinase) | 11.0 ± 0.2 | 12.7 ± 0.2 | 1.7 ± 0.4 | 3.2 ± 1.3 |
| acn (HM1_0105, aconitase) | 10.7 ± 0.1 | 12.6 ± 0.1 | 1.8 ± 0.2 | 3.5 ± 1.1 |
| fdxR (HM1_0289, Fd-NADP+ reductase, FNR) | 12.2 ± 0.0 | 14.2 ± 0.1 | 2.0 ± 0.1 | 4.0 ± 1.0 |
| porA (HM1_0807, pyruvate, Fd oxidoreductase, PFOR) | 9.8 ± 0.2 | 11.7 ± 0.1 | 1.8 ± 0.3 | 3.5 ± 1.2 |
| acsA (HM1_0951, acetyl-CoA synthetase) | 12.1 ± 0.0 | 14.3 ± 0.2 | 2.1 ± 0.2 | 4.3 ± 1.1 |
| icd (HM1_1471, isocitrate dehydrogenase) | 10.7 ± 0.2 | 12.4 ± 0.0 | 1.6 ± 0.2 | 3.0 ± 1.1 |
| mdh (HM1_1472, malate dehydrogenase | 10.9 ± 0.0 | 12.4 ± 0.0 | 1.5 ± 0.0 | 2.8 ± 0.0 |
| ackA (HM1_2157, acetate kinase) | 11.6 ± 0.1 | 13.6 ± 0.0 | 2.0 ± 0.1 | 4.0 ± 1.0 |
| ppdK (HM1_2461, pyruvate phosphate dikinase) | 11.3 ± 0.1 | 13.2 ± 0.1 | 1.9 ± 0.2 | 3.7 ± 1.1 |
| korC (HM1_2762, KFOR, γ subunit) | 9.9 ± 0.0 | 10.8 ± 0.2 | 0.9 ± 0.2 | 1.9 ± 1.1 |
| oorB (HM1_2763, KFOR, β subunit) | 9.2 ± 0.0 | 10.9 ± 0.1 | 1.7 ± 0.1 | 3.2 ± 1.0 |
| korA (HM1_2766, KFOR, α subunit) | 8.9 ± 0.1 | 10.7 ± 0.0 | 1.8 ± 0.1 | 3.5 ± 1.0 |
| korD (HM1_2767, KFOR, δ subunit) | 8.9 ± 0.0 | 11.0 ± 0.1 | 2.1 ± 0.1 | 4.3 ± 1.0 |
| nifV (HM1_0858, homocitrate synthase) | 15.0 ± 0.1 | 15.4 ± 0.2 | 0.4 ± 0.3 | 1.3 ± 1.2 |
| aksA (HM1_2993, homocitrate synthase) | 10.5 ± 0.2 | 12.0 ± 0.2 | 1.5 ± 0.4 | 2.8 ± 1.3 |
| pckA (HM1_2773, PEP carboxykinase) | 11.9 ± 0.0 | 13.5 ± 0.1 | 1.6 ± 0.1 | 3.0 ± 1.0 |
a ΔCT = CT (the threshold cycle) of the target gene −CT of the 16 S rRNA gene.
Moreover, when the anaerobic sulfate-reducing bacterium D. vulgaris Hildenborough (DvH) was grown with a 1:1 molar ratio of [2-13C]acetate and nonlabeled lactate, significant amounts of labeled carbon were detected in the biomass (supplemental Table S2), indicating that DvH can utilize acetate for producing biomass. Furthermore, isotopomer analysis using [2-13C]acetate (supplemental Table S2) and a previous DvH flux analysis (21) indicate that acetate ↔ acetyl-CoA is reversible in DvH. No FAc inhibition was detected during the growth of DvH (Fig. 2C).
Isotopomer Analysis by GC-MS
Previous studies have established that the cell growth of H. modesticaldum is best supported by pyruvate (1, 3, 4). We used 13C-labeled pyruvate and characterized the protein-based amino acids for probing the central carbon metabolic pathways. H. modesticaldum is recognized as a photoheterotrophic bacterium (3, 4), and CO2 neither supports nor enhances its phototrophic growth (Fig. 1B). When H. modesticaldum was grown on [1-13C]pyruvate, aspartate was mainly double-labeled, and glutamate was primarily single-labeled (Table 2). In the (incomplete) RTCA cycle (Fig. 1), α-KG, the precursor of glutamate, is synthesized from OAA, the precursor of aspartate, which results in the incorporation of all the carbons from aspartate into the carbon backbones of glutamate. Also, assuming that some of the CO2 molecules assimilated through the RTCA cycle are 13CO2 generated from the reaction [1-13C]pyruvate + CoA → acetyl-CoA + 13CO2 (Fig. 1), 13C-labeled content is expected to be higher for glutamate than for aspartate. Consequently, the multiply labeled instead of the singly labeled glutamate would eventually become dominant through the (incomplete) RTCA cycle. However, in this study, the observed lower 13C-labeled content of glutamate compared with aspartate with [1-13C]pyruvate-grown cells cannot have been generated by cells utilizing solely the (incomplete) RTCA cycle. Furthermore, the carbon flux of H. modesticaldum cannot primarily go through the (incomplete) RTCA cycle but must instead mainly transform via the OTCA cycle. Moreover, aspartate was dominantly single-labeled (M1 >0.8), and glutamate was mostly double-labeled (M2 >0.75) for cells using [3-13C]pyruvate (supplemental Table S3), in agreement with significant carbon flow through the OTCA cycle.
TABLE 2.
Isotopomer labeling patterns of protein-based amino acids in the cultures grown on [1-13C]pyruvate
| Amino acid | Precursor | Iona | M-57b | M-159c | f302d | Proposed 13C-enriched positions (shown as asterisk) |
|---|---|---|---|---|---|---|
| Ala | Pyruvate | M0 | 0.20 | 0.85 | 0.20 | C-C-*COOH |
| M1 | 0.77 | 0.09 | 0.79 | |||
| M2 | 0.03 | 0.06 | 0.01 | |||
| Gly | Serine | M0 | 0.24 | 0.98 | 0.12 | C-*COOH |
| M1 | 0.75 | 0.02 | 0.48 | |||
| M2 | 0.01 | 0.40 | ||||
| Val | Pyruvate | M0 | 0.20 | 0.89 | 0.28 | C-C-C-C-*COOH |
| M1 | 0.77 | 0.09 | 0.72 | |||
| M2 | 0.03 | 0.01 | 0.00 | |||
| Leu | Pyruvate | M0 | 0.60 | 0.83 | 0.63 | Nonlabeled |
| Acetyl-CoA | M1 | 0.36 | 0.14 | 0.36 | ||
| M2 | 0.03 | 0.02 | 0.01 | |||
| M3 | 0.00 | 0.00 | ||||
| Ile | Pyruvate | M0 | 0.66 | 0.83 | 0.69 | Nonlabeled |
| Threonine | M1 | 0.25 | 0.16 | 0.25 | ||
| M2 | 0.08 | 0.01 | 0.06 | |||
| M3 | 0.01 | 0.00 | ||||
| Met | Aspartate | M0 | 0.09 | 0.22 | 0.17 | C-S-*C-C-C-*COOH |
| Methyl-THFe | M1 | 0.24 | 0.58 | 0.51 | ||
| M2 | 0.51 | 0.20 | 0.32 | |||
| M3 | 0.15 | 0.01 | ||||
| Ser | 3-Phosphoglycerate | M0 | 0.19 | 0.94 | 0.21 | C-C-*COOH |
| M1 | 0.77 | 0.06 | 0.78 | |||
| M2 | 0.04 | 0.00 | 0.01 | |||
| Thr | Aspartate | M0 | 0.11 | 0.26 | 0.11 | *C-C-C-*COOH |
| M1 | 0.29 | 0.72 | 0.46 | |||
| M2 | 0.59 | 0.02 | 0.43 | |||
| Phe | PEP | M0 | 0.08 | 0.13 | 0.22 | C-C-*C-*C-C-C-C-C-*COOH |
| Erythrose 4-phosphate | M1 | 0.11 | 0.29 | 0.77 | ||
| M2 | 0.31 | 0.53 | 0.01 | |||
| M3 | 0.46 | 0.04 | ||||
| M4 | 0.04 | 0.01 | ||||
| Asp | OAA | M0 | 0.11 | 0.27 | 0.27 | *COOH-C-C-*COOH |
| M1 | 0.31 | 0.71 | 0.73 | |||
| M2 | 0.56 | 0.01 | 0.01 | |||
| Glu | α-Ketoglutarate | M0 | 0.26 | 0.28 | 0.82 | C-C-C-C-*COOH |
| M1 | 0.64 | 0.68 | 0.18 | |||
| M2 | 0.09 | 0.04 | 0.00 | |||
| M3 | 0.01 | 0.00 | ||||
| His | Ribose 5-phosphate | M0 | 0.13 | 0.13 | 0.92 | N-C-N-C-*C-*C-C-COOH |
| M1 | 0.41 | 0.41 | 0.06 | |||
| M2 | 0.32 | 0.34 | 0.02 | |||
| M3 | 0.10 | 0.07 | ||||
| Lys | Aspartate | M0 | 0.14 | 0.28 | 0.92 | C-C-*C-C-C-*COOH |
| Pyruvate | M1 | 0.26 | 0.68 | 0.08 | ||
| M2 | 0.58 | 0.03 | 0.00 | |||
| M3 | 0.02 | 0.00 | ||||
| M4 | 0.00 | 0.00 |
a M0, M1, M2, M3, and M4 indicate mass fraction of the unlabeled, single-labeled, double-labeled, triple-labeled, and quadruple-labeled amino acid.
b [M-57]+ indicates un-fragmented amino acid detected by GC-MS.
c [M-159]+ indicates an amino acid minus the α−carboxyl group detected by GC-MS.
d f302 is the fragment of the first two carbons in a derivatized amino acid (17).
e THF is tetrahydrofolate.
Mass Spectrum of Photosynthetic Pigments
We further analyzed the mass spectrum of BChls we reported earlier using MALDI-TOF (matrix-assisted laser desorption ionization time-of-flight) mass spectrometry (4), in which 13C-labeled photosynthetic pigments (bacteriochlorophyll g (BChl g) and 81-hydroxychlorophyll a with a farnesol tail (81-OH-Chl aF)) were detected using [3-13C]pyruvate. Both BChl g and 81-OH-Chl aF were synthesized from eight molecules of glutamate and eight molecules of acetyl-CoA (for generating the farnesyl tail with a 15-carbon unit at C173 position), and thus 13C-labeled BChl g and 81-OH-Chl aF are expected using [3-13C]pyruvate. The m/e for BChl g and 81-OH-Chl aF is 796.7 and 812.7, respectively, with unlabeled pyruvate, and for BChl g and 81-OH-Chl aF is 816.8 and 832.8, respectively, with [3-13C]pyruvate. The number of labeled carbons for the labeled BChl g or 81-OH-Chl aF was estimated to be 21–22 using the program IsoPro 3.1. Using [3-13C]pyruvate as the carbon source, glutamate is expected to be double-labeled through the OTCA cycle because citrate is double-labeled via condensation of [3-13C]OAA and [2-13C]acetyl-CoA, where glutamate is single-labeled through the RTCA cycle (from [3-13C]OAA). Thus, 16 carbons are labeled through the RTCA cycle (eight single-labeled glutamates and eight single-labeled acetyl-CoAs) and 24-labeled carbons through the OTCA cycle (eight double-labeled glutamates and eight single-labeled acetyl-CoAs) using [3-13C]pyruvate. Thus, observations of higher 13C contents on BChl g and 81-OH-Chl aF in the mass spectrum than expected from the RTCA cycle would suggest that the OTCA cycle contributes to the formation of glutamate and α-KG. Consistent with this hypothesis, glutamate was found to be mostly double-labeled (M2 >75%) using [3-13C]pyruvate (supplemental Table S3).
Transcriptomic Profiles
All of the genes encoding the enzymes in the RTCA cycle, except for ACL, have been annotated in the H. modesticaldum genome (5). Table 1 shows that genes in the RTCA cycle (highlighted in boldface italic) are expressed and that the transcript level for the genes responsible for carbon metabolism is similar (at most a 4-fold difference) in the pyruvate-grown cultures with and without FAc.
Taking all of the experimental evidence together, our studies indicate that in addition to the incomplete RTCA cycle, the OTCA cycle is also employed by H. modesticaldum. Like the RTCA cycle, the OTCA cycle is not complete but mainly contributes to the formation of α-KG, as indicated by isotopomer analysis and mass spectrometry of photosynthetic pigments. The active OTCA cycle is consistent with the observed lack of CO2-enhanced pyruvate growth of H. modesticaldum (Fig. 1B), because CO2 is produced through the OTCA cycle.
Citrate Synthase Activity Detected Anaerobically
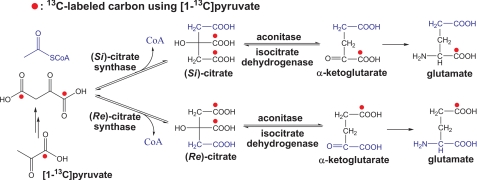
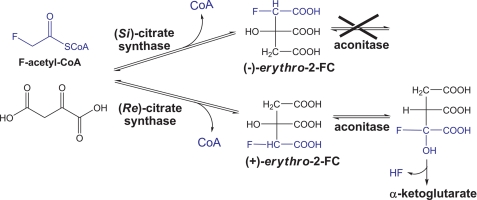
Table 2 shows that over 50% aspartate and 60% glutamate were labeled in the β-carboxyl group with [1-13C]pyruvate as the carbon source, implying that citrate formation is possibly catalyzed by (Re)-citrate synthase ((Re)-CS) (Fig. 3). Both (Re)-CS and normal CS (i.e. (Si)-CS) catalyze the formation of citrate through aldol condensation of OAA and acetyl-CoA, whereas the acetyl-CoA moiety is added to the “pro-R” and “pro-S” arm of citrate through catalysis of (Re)-CS and (Si)-CS, respectively (Fig. 3).
FIGURE 3.
Reactions catalyzed by the (Re)- versus (Si)-citrate synthase, aconitase, and isocitrate dehydrogenase. 13C labeling distributions in citrate, α-ketoglutarate, and glutamate using [1-13C]pyruvate are shown.
We have previously showed that no CoA production was detected with acetyl-CoA and OAA added in the cell extracts of H. modesticaldum under aerobic conditions and without divalent metal ion supplied, confirming that (Si)-CS is not produced by H. modesticaldum (4). Anaerobic conditions have not been reported to be required for the activity of (Si)-CS. To test if citrate can be produced by H. modesticaldum via the catalysis of (Re)-CS, rigorous efforts were made to minimize the oxygen content in the assay mixtures (described under “Experimental Procedures”), because oxygen-sensitive and divalent metal ion (Mn2+, Mg2+, or Co2+) dependences were reported for the activity measurements of (Re)-CS (19). The increase of A412 for 5,5′-dithiobis-2-nitrobenzoic acid-modified CoA can be detected with Mn2+, OAA, and acetyl-CoA under anaerobic conditions, suggesting the presence of the CS activity in H. modesticaldum. The catalytic activity of the novel CS, likely (Re)-CS, was estimated to be 50 ± 20 nmol/min/mg protein.
Anaplerotic-CO2 Fixation Pathways
The isotopomer labeling experiment with 13C-labeled HCO3− and unlabeled pyruvate as the carbon source showed that over 50% alanine was labeled (supplemental Table S4) and that all the labeled alanine was labeled at the carboxyl group. The labeling pattern suggests that the reaction catalyzed by pyruvate:ferredoxin oxidoreductase (acetyl-CoA + CO2 + 2Fdred + 2H+ ↔ pyruvate + CoA + 2Fdox) is freely reversible and very active so that the labeled bicarbonate is incorporated into pyruvate. The isotopomer analysis is in agreement with our recent physiological studies that pyruvate:ferredoxin oxidoreductase plays a central role in carbon metabolism of H. modesticaldum (4). Moreover, when using [1-13C]pyruvate as the carbon source, aspartate can be synthesized either from the CO2-anaplerotic pathways with pyruvate and/or phosphoenolpyruvate (PEP) and 13CO2 (from decarboxylation of [1-13C] pyruvate), leading to double-labeled aspartate, or through the OTCA cycle, in which aspartate is not expected to be double labeled. The predominance of double-labeled aspartate indicates the high carbon flow via the CO2-anaplerotic pathway and low flux from the OTCA cycle, consistent with observed activity of PEP carboxykinase illustrated in our recent studies (4), and confirming that the OTCA cycle is not complete as genes encoding the enzymes specific for the OTCA cycle have not been annotated in the genome (5).
Amino Acid Biosynthesis
The isotopomer analysis for biosynthesis of several amino acids is illustrated as follows.
Alternative Isoleucine Biosynthesis Pathway
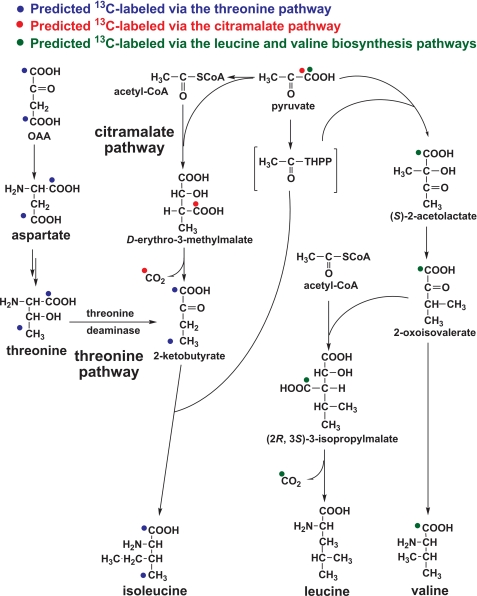
Isoleucine is typically synthesized through the threonine pathway (Fig. 4). Using [1-13C]pyruvate as the carbon source, threonine was mainly double-labeled (M2 >58%), so isoleucine would be expected to be double-labeled via the threonine pathway. Table 2 shows that isoleucine was largely nonlabeled (M0 >60%) with [1-13C]pyruvate, suggesting that isoleucine was mostly synthesized through the citramalate pathway, although all of the genes in the threonine pathway have been annotated. In the citramalate pathway, citramalate synthase (CimA) catalyzes the formation of d-erythro-3-methylmalate (i.e. citramalate) through condensation of pyruvate and acetyl-CoA (Fig. 4). Two gene loci, HM1_1519 and HM1_1515, encoding putative CimA in the citramalate pathway have been annotated in the genome. The amino acid sequence encoding by gene locus HM1_1519 shows >50% identity to the recently reported CimA (Teth514_1204) in Thermoanaerobacter sp. X514 (22), which is a close relative to the genus Clostridium. The supplemental Table S5 lists the bacteria with the citramalate pathway identified.
FIGURE 4.
Proposed biosynthesis pathways for isoleucine, including the citramalate and threonine pathways, leucine, and valine. Predicted 13C labeling distributions using [1-13C]pyruvate are shown.
Normal Pathways for Alanine, Serine, Phenylalanine, and Lysine Biosynthesis
The isotopomer pattern suggested alanine and serine are synthesized from pyruvate and PEP, respectively. Also, using [3-13C]pyruvate as the carbon source, the labeling pattern in supplemental Table S3 indicates that phenylalanine is synthesized from the common biosynthetic pathway with erythrose 4-phosphate and PEP as precursors in agreement with the genomic information (5). Furthermore, Pickett et al. (9) also reported the same results regarding pathways of alanine, serine, and phenylalanine biosynthesis for Heliobacterium strain HY-3.
The labeling patterns of lysine in the isotopomer analysis suggest that lysine is synthesized through the common diaminopimelate pathway with pyruvate and aspartate as the precursors, rather than through the α-amino adipate pathway (supplemental Fig. S1), in which 2-ketoglutarate and acetyl-CoA are condensed and converted to lysine. This conclusion is consistent with the fact that all genes in the α-aminoadipate pathway are missing, except for the genes encoding putative homocitrate synthase (nifV (HM1_0858) and aksA (HM1_2993)). This enzyme catalyzes the formation of homocitrate ((R)-2-hydroxybutane-1,2,4-tricarboxylate) by condensing α-KG and acetyl-CoA. Note that the askA gene is expressed with higher transcript level (Table 1). The possible function of the homocitrate synthase for carbon metabolism will be discussed later.
DISCUSSION
Growth of H. modesticaldum with FAc
Fig. 2 shows that there is no detectable difference in the growth of H. modesticaldum with or without FAc, suggesting that (−)-erythro-2-FC cannot be synthesized by CS or/and ACL in H. modesticaldum. However, the lack of genes encoding ACL and CS along with the lack of the enzymatic activities is inconsistent with our presented isotopomer data, which suggest that at least a partial OTCA cycle is active and an oxygen-sensitive novel CS is produced by H. modesticaldum to initiate the OTCA cycle. If a novel CS is produced by H. modesticaldum, then the lack of FAc inhibition during the growth of H. modesticaldum needs to be explained. Two working hypotheses can be considered.
High Flux from Pyruvate to Acetyl-CoA to Compete with F-acetyl-CoA
Similar activity of F-acetyl-CoA versus acetyl-CoA for normal CS has been reported previously (23). Assuming that (−)-erythro-2-FC is synthesized by a putative novel CS, acetyl-CoA from pyruvate could compete with F-acetyl-CoA from FAc for the interactions of the putative CS. A high flux from pyruvate to acetyl-CoA is suggested from our recent studies showing that conversion of pyruvate to acetyl-CoA, catalyzed by pyruvate:ferredoxin oxidoreductase, is very active in pyruvate-grown H. modesticaldum (4) and the isotopomer data presented in this report. However, Fig. 2A shows that the pyruvate-grown R. denitrificans, which is also known to exhibit high flux from pyruvate to acetyl-CoA (12), was notably reduced by the presence of FAc, suggesting that other factors must contribute to the lack of FAc-inhibition on the phototrophic growth of H. modesticaldum.
Different Stereoisomer of 2-FC Is Synthesized
Alternatively, it is possible that the putative CS catalyzes the formation of a 2-FC isomer other than (−)-erythro-2-FC. Note that only one of four possible 2-FC isomers, (−)-erythro-2-FC, catalyzed by (Si)-CS, is a potent inhibitor of aconitase (20). Thus, if the putative CS does not catalyze the formation of (−)-erythro-2-FC but a different 2-FC isomer, then the toxicity of FAc is not expected to be observed for H. modesticaldum. This hypothesis is further elaborated below.
Different Isomers of 2-FC Produced by (Re)- Versus (Si)-CS
Although there is no difference in the stereochemistry of citrate from condensation of acetyl-CoA and OAA catalyzed by (Si)-CS versus (Re)-CS (Fig. 3), different isomers of 2-F-citrate (2-FC) are produced from condensation of F-acetyl-CoA and OAA catalyzed by (Si)-CS versus (Re)-CS (Fig. 5). Lauble et al. (20) proposed that interactions of (−)-erythro-2-FC with aconitase produce an intermediate that inhibits aconitase, whereas interactions of (+)-erythro-2-FC with aconitase leads to formation of α-KG, instead of inhibiting aconitase.
FIGURE 5.
Reaction of F-acetyl-CoA and OAA catalyzed by (Re)- versus (Si)-citrate synthase, and interactions of (−)-erythro-2-FC versus (+)-erythro-2-FC with aconitase.
To test this hypothesis, we performed physiological studies with the sulfate-reducing bacterium D. vulgaris Hildenborough (DvH) using FAc. Acetate can serve as a carbon source for DvH in the presence of reducing power, which can be generated during lactate-supported growth (supplemental Table S2). Thus, FAc, like acetate, can be assimilated by DvH using the reducing power generated by lactate oxidation. Like H. modesticaldum, no genes encoding ACL and (Si)-CS have been annotated in the DvH genome, whereas the activity of (Re)-CS and contribution of CS to the carbon flow of the OTCA cycle have been identified for DvH (24). Fig. 2C shows no inhibition for the growth of DvH with FAc, consistent with the hypothesis by Lauble et al. (20). Consequently, we believe that the lack of FAc inhibition on the growth of H. modesticaldum is in agreement with generation of (+)-erythro-2-FC, instead of (−)-erythro-2-FC, by (Re)-CS.
Homocitrate Synthase May Function as (Re)-CS in H. modesticaldum
(Re)-CS has been identified in Clostridium kluyveri (19), and the activity of (Re)-CS has been reported in Thermoanaerobacter sp. X514 (22), DvH (24), Dehalococcoides ethenogenes 195 (26), Ignicoccus hospitalis (27), and several anaerobic bacteria. Because different stereoisomers of citrates are generated through the catalysis of (Re)- versus (Si)-CS, it is not surprising that these two types of CS are rather phylogenetically distinct (19, 28).
As mentioned under “Results,” the product turnover catalyzed by an oxygen-sensitive CS, possibly a putative (Re)-CS, has been detected with Mn2+ ions supplied while minimizing oxygen content in the assay solution. Although the gene encoding (Re)-CS has not been annotated in the H. modesticaldum genome, homocitrate synthase has been suggested to be phylogenetically related to (Re)-CS (19). It is possible that homocitrate synthase catalyzes not only the formation of homocitrate from acetyl-CoA and α-KG but also the formation of citrate from acetyl-CoA and OAA (i.e. (Re)-CS). Two genes in the H. modesticaldum genome possibly encode homocitrate synthase, nifV (HM1_0858) and aksA (HM1_2993). A BLAST search shows that proteins encoded by both genes, particularly by askA, share high identity with the reported putative homocitrate synthase/(Re)-CS, in agreement with the observed higher transcript level (i.e. lower ΔCT) of askA compared with that of nifV (Table 1). Given the data presented in this study, we propose that the putative homocitrate synthase, an enzyme that has been reported to function as (Re)-CS in several bacteria, is likely responsible for synthesizing citrate in H. modesticaldum.
New View for Carbon Flow of H. modesticaldum
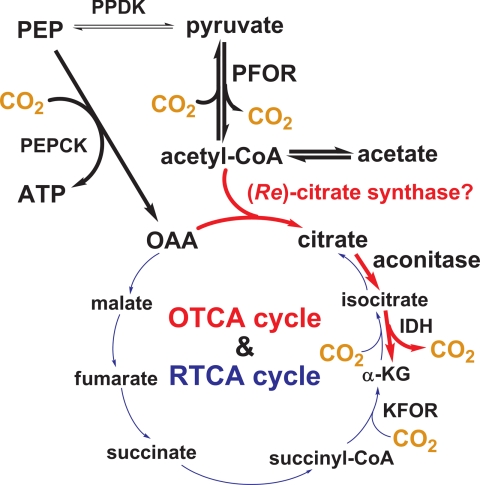
Fig. 6 represents a new model for the carbon flow of H. modesticaldum that reflects improved understanding resulting from isotopomer and activity assays; the major carbon flux of H. modesticaldum is through the OTCA cycle. (Re)-citrate synthase, identified in several Clostridia and other anaerobic bacteria, is likely employed by H. modesticaldum to produce citrate for entering the OTCA cycle. The finding of the OTCA cycle herein fills the knowledge gap for the carbon flow of H. modesticaldum. It is intriguing to learn that the major carbon flux of H. modesticaldum is switched from the RTCA cycle to the OTCA cycle when the only gene (ATP citrate lyase) missing in the RTCA cycle is replaced by a gene with novel function ((Re)-citrate synthase) in the OTCA cycle. Together, our studies suggest that the carbon flow of H. modesticaldum (and perhaps Heliobacterium strain HY-3) is more akin to Clostridia than to the green sulfur bacteria, which employ the RTCA cycle for CO2 assimilation and biomass production (25).
FIGURE 6.
New view for carbon flow in H. modesticaldum. The proposed carbon flow is through the OTCA cycle (red) and the RTCA cycle (blue) with a stronger flux through the OTCA cycle (acetyl-CoA → α-KG, shown in bold). The proposed role of the putative (Re)-citrate synthase is shown.
Supplementary Material
This work was supported by National Science Foundation Career Grant MCB0954016 (to Y. J. T.) and Exobiology Program of NASA Grant NNX08AP62G (to R. E. B.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Tables S1–S5, Fig. S1, and additional references.
- ACL
- ATP citrate lyase
- CS
- citrate synthase
- FAc
- fluoroacetate
- 2-FC
- 2-fluorocitrate
- α-KG
- α-ketoglutarate
- OAA
- oxaloacetate
- OTCA cycle
- oxidative (forward) TCA cycle
- PEP
- phosphoenolpyruvate
- RTCA cycle
- reductive (reverse) TCA cycle
- BChl g
- bacteriochlorophyll g
- 81-OH-Chl aF
- 81-hydroxychlorophyll a with a farnesol tail.
REFERENCES
- 1.Madigan M. T. (2006) Prokaryotes 4, 951–964 [Google Scholar]
- 2.Asao M., Madigan M. T. (2010) Photosynth. Res. 104, 103–111 [DOI] [PubMed] [Google Scholar]
- 3.Kimble L. K., Mandelco L., Woese C. R., Madigan M. T. (1995) Arch. Microbiol. 163, 259–267 [Google Scholar]
- 4.Tang K. H., Yue H., Blankenship R. E. (2010) BMC Microbiol. 10, 150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sattley W. M., Madigan M. T., Swingley W. D., Cheung P. C., Clocksin K. M., Conrad A. L., Dejesa L. C., Honchak B. M., Jung D. O., Karbach L. E., Kurdoglu A., Lahiri S., Mastrian S. D., Page L. E., Taylor H. L., Wang Z. T., Raymond J., Chen M., Blankenship R. E., Touchman J. W. (2008) J. Bacteriol. 190, 4687–4696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Heinnickel M., Golbeck J. H. (2007) Photosynth. Res. 92, 35–53 [DOI] [PubMed] [Google Scholar]
- 7.Baymann F., Nitschke W. (2010) Photosynth. Res. 104, 177–187 [DOI] [PubMed] [Google Scholar]
- 8.Golbeck J. H. (2010) Photosynth. Res. 104, 101–102 [DOI] [PubMed] [Google Scholar]
- 9.Pickett M. W., Williamson M. P., Kelly D. J. (1994) Photosynth. Res. 41, 75–88 [DOI] [PubMed] [Google Scholar]
- 10.Tang Y. J., Hwang J. S., Wemmer D. E., Keasling J. D. (2007) Appl. Environ. Microbiol. 73, 718–729 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Overmann J. (2006) Prokaryotes 7, 359–378 [Google Scholar]
- 12.Tang K. H., Feng X., Tang Y. J., Blankenship R. E. (2009) PLoS One. 4, e7233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Widdel F., Pfennig N. (1984) in Bergey's Manual of Systematic Bacteriology (Krieg N. R., Holt J. G. eds) Vol. 1, pp. 663–679, Williams & Wilkins, Baltimore [Google Scholar]
- 14.Brysch K., Schneider C., Fuchs G., Widdel F. (1987) Arch. Microbiol. 148, 264–274 [Google Scholar]
- 15.Wolin E. A., Wolin M. J., Wolfe R. S. (1963) J. Biol. Chem. 238, 2882–2886 [PubMed] [Google Scholar]
- 16.Tang K. H., Wen J., Li X., Blankenship R. E. (2009) J. Bacteriol. 191, 3580–3587 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wahl S. A., Dauner M., Wiechert W. (2004) Biotechnol. Bioeng. 85, 259–268 [DOI] [PubMed] [Google Scholar]
- 18.Bradford M. M. (1976) Anal. Biochem. 72, 248–254 [DOI] [PubMed] [Google Scholar]
- 19.Li F., Hagemeier C. H., Seedorf H., Gottschalk G., Thauer R. K. (2007) J. Bacteriol. 189, 4299–4304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lauble H., Kennedy M. C., Emptage M. H., Beinert H., Stout C. D. (1996) Proc. Natl. Acad. Sci. U.S.A. 93, 13699–13703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tang Y., Pingitore F., Mukhopadhyay A., Phan R., Hazen T. C., Keasling J. D. (2007) J. Bacteriol. 189, 940–949 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Feng X., Mouttaki H., Lin L., Huang R., Wu B., Hemme C. L., He Z., Zhang B., Hicks L. M., Xu J., Zhou J., Tang Y. J. (2009) Appl. Environ. Microbiol. 75, 5001–5008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Marcus A., Elliott W. B. (1956) J. Biol. Chem. 218, 823–830 [PubMed] [Google Scholar]
- 24.Pingitore F., Tang Y., Kruppa G. H., Keasling J. D. (2007) Anal. Chem. 79, 2483–2490 [DOI] [PubMed] [Google Scholar]
- 25.Tang K. H., Blankenship R. E. (July22, 2010) J. Biol. Chem. 10.1074/jbc.M110.157834 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tang Y. J., Yi S., Zhuang W. Q., Zinder S. H., Keasling J. D., Alvarez-Cohen L. (2009) J. Bacteriol. 191, 5224–5231 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jahn U., Huber H., Eisenreich W., Hügler M., Fuchs G. (2007) J. Bacteriol. 189, 4108–4119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lamzin V. S., Dauter Z., Wilson K. S. (1995) Curr. Opin. Struct. Biol. 5, 830–836 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.