Abstract
Brassinosteroids (BRs) are important plant growth hormones that largely rely on transcription factors (TFs) to regulate a variety of plant physiological/developmental processes. Past genetic and biochemical studies have identified two key TFs and interacting partners that play major roles in regulating many BR-responsive genes, while genome-wide microarray experiments have discovered at least 50 BR-regulated TFs. However, little is known how these TFs function or whether additional TFs are involved in BR signaling. In the past few years, genetic studies and yeast one/two-hybrid screens coupled with microarray and chromatin immunoprecipitation experiments not only revealed new roles of the key regulatory TFs but also implicated additional TFs and other nuclear proteins in regulating the nuclear activities of BR signaling in Arabidopsis and rice.
Introduction
Brassinosteroids (BRs) are important plant growth hormones that regulate a variety of physiological and developmental processes [1]. In the last decade, genetic and biochemical studies in Arabidopsis have elucidated a phosphorylation-mediated signaling pathway that regulates the stability, subcellular localization, and/or DNA binding activity of two highly similar transcription factors (TFs), bri1-EMS-Suppressor 1 (BES1) and Brassinazol Resistant 1 (BZR1) [2,3]. By contrast, little is known how BRs regulate gene activities in rice despite a recent discovery demonstrating a role of a rice BZR1 homolog, OsBZR1, in feedback regulation of BR biosynthesis [4]. Genome-wide microarray analyses in Arabidopsis have shown that BRs affect the expression of hundreds, if not thousands, of genes involved in a wide range of cellular processes, and identified at least 50 BR-regulated TFs that include members of all major families of plant TFs, such as basic helix-loop-helix (bHLH) proteins, MYB, GRAS, AP2, NAC, WRKY, homeodomain-leucine-zipper (HD-ZIP), and basic leucine zipper (bZIP) TFs [5]. However, nothing is known whether these TFs function dependently or independently of BES1/BZR1 to regulate expression of certain BR-responsive genes or whether BR signaling involves additional nuclear proteins. This article reviews studies of the last several years that uncovered additional roles of BES1 and its interacting factor in specific developmental processes and implicated other TFs and nuclear proteins in regulating BR signaling in both Arabidopsis and rice.
New roles of BES1 and BIM1 in pollen/anther development and embryogenesis
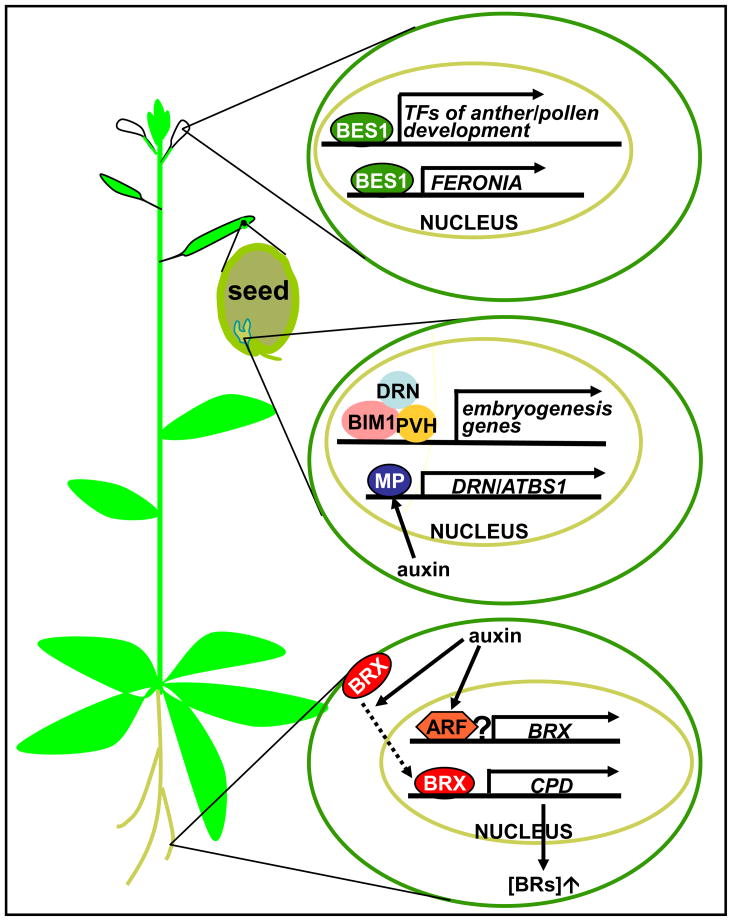
BR-deficient/signaling mutants are often male sterile, yet little is known about how BR regulates the pollen development/function, even though pollen is the tissue from which the first BR was discovered [6]. A systematic analysis of Arabidopsis BR-related mutants and transgenic lines revealed that BR signaling is important not only for tapetum and microspore development but also for pollen wall formation, which explains decreased number and reduced release efficiency of pollen in BR mutants [7]. Gene expression analyses and chromatin immunoprecipitation (ChIP) assays demonstrated that BES1 binds to the promoters of key regulatory TFs of anther/pollen development to control their expression (Figure 1A), thus revealing a molecular mechanism by which BR regulates anther/pollen development. BES1 is also involved in the BR-induction of several related receptor kinases, including FERONIA [8,9] that was previously known to mediate male-female interaction for releasing sperms from elongating pollen tubes [10], thus providing another possible explanation for reduced fertility of BR mutants (Figure 1A).
Figure 1.
The potential roles of BES1 and BR-related TFs in plant development. (A). In Arabidopsis flowers, BES1 likely binds to the promoters of known anther/pollen regulatory TFs and a female gametophytic receptor-like kinase FERONIA to affect anther/pollen development and pollen discharge, respectively. (B). In Arabidopsis embryos, BIM1 physically and genetically interact with DRN and PVH to regulate cotyledon development while an auxin-dependent MP directly activates the expression of DRN and ATBS1 (encoding an atypical bHLH protein recently implicated in BR signaling [33]) to control embryonic cotyledon and root development, respectively. (C). In Arabidopsis root, auxin induces the BRX expression likely though an ARF and promotes the translocation of BRX from the cell surface to the nucleus where it activates the CPD expression crucial for BR biosynthesis.
Despite its synergistic interaction with auxin, BR has so far not been implicated in embryogenesis that requires auxin signaling for pattern formation [11]. A recent study, however, uncovered a potential link between BR and embryogenesis (Figure 1B). BES1-interacting Myc-like 1 (BIM1) [12] physically binds and genetically interacts with two AP2-domain proteins, DORNRÖSCHEN (DRN) and its paralog DORNRÖSCHEN-LIKE (DRNL) [13] that are crucial for embryonic patterning by affecting cotyledon development in the apical embryo domain and cell division in the basal embryo domain [14].
Potential functions of ARF2, BRX, and DRN in the auxin-BR crosstalk
The auxin-BR interaction was thought to be mediated by auxin response factors (ARFs) that bind to the promoters of common target genes carrying the ARF core-binding element (TGTCT) [15]. Interestingly, BRASSINOSTEROID INSENSITIVE 2 (BIN2), an Arabidopsis GSK3-like kinase that inhibits BR signaling [16], was recently shown to interact with ARF2 [17], a known repressor of auxin signaling [18]. Genetic and microarray analyses suggested a role of ARF2 in BR signaling while biochemical studies demonstrated that BIN2 phosphorylates ARF2 in vitro to reduce its binding affinity to the SAUR-AC1 promoter, a shared target of auxin and BR signaling [15], thus inhibiting its repressor activity. However, It remains unclear how the BIN2-catalyzed ARF2 phosphorylation contributes to the auxin-BR synergistic interaction. BR is known to negatively regulate the kinase activity and protein abundance of BIN2 [19,20], implying ARF2 would be an active suppressor when the BR signaling is active. Perhaps the BIN2-catalyzed ARF2 phosphorylation inhibits ARF2 interaction with other ARFs that bind to the common targets of the two signaling pathways.
In the Arabidopsis roots, the auxin-BR interaction could be mediated by BREVIS RADIX (BRX) (Figure 1C), which was originally identified as a regulator of root cell division/elongation [21] and is predicted to regulate transcription [22]. Auxin induces the BRX expression and promotes BRX movement from the plasma membrane into the nucleus [23] where it activates the root-specific expression of the Constitutive Photomorphogenesis and Dwarf (CPD) gene [24] encoding a rate-limiting BR biosynthetic C23-hydroxylase [25]. Another potential mechanism of the auxin-BR crosstalk might be mediated by the BIM1-interacting DRN [13] that acts upstream of auxin signaling/transport [14] but is also a direct target of MONOPTEROS (MP) [26], a well-studied ARF (ARF5) regulating embryonic patterning [27] (Figure 1B).
Regulatory roles of atypical and non-canonical bHLH proteins
Consistent with enrichment of E-box element on many BR-responsive promoters [15], several bHLH proteins were implicated in BR signaling, including BR Enhanced Expression 1 (BEE1) to BEE3 [28] and 3 BIMs [12]. It is known that transcriptional activities of bHLH proteins are often negatively regulated through heterodimerization with a non-DNA-binding HLH protein (often called atypical bHLH proteins in the plant literature) to inhibit DNA binding [29]. The Arabidopsis and rice genomes encode at least 27 and 26 atypical bHLH proteins, respectively [30]. Three recent studies discovered regulatory functions of such atypical bHLH proteins in BR signaling [31–33].
Activation-Tagged bri1 Suppressor 1 (ATBS1), discovered as a suppressor of a weak BR receptor mutant bri1-301 [33,34], is a 93-amino-acid protein homologous to 5 other Arabidopsis proteins, including KIDARI and Paclobutrazol Resistance 1 (PRE1) previously implicated in light and gibberellin signaling, respectively [35,36]. Overexpression of ATBS1 or any of its 5 homologs suppressed bri1-301. Despite having a N-terminal basic region, all 6 proteins lack amino acids critical for DNA-binding [29] and are thus atypical bHLH proteins. Consistently, a truncated ATBS1 lacking the basic region still possess the bri1-301-rescuing activity, most likely via heterodimerization with one or more closely-related ATBS1-Interacting Factors (AIFs) (Figure 2A), which were discovered by a yeast two-hybrid screen and are also atypical bHLH proteins. Remarkably, overexpression of an AIF gene resulted in a severe dwarf phenotype indistinguishable from that of known BR-deficient mutants and suppressed the activation-tagged atbs1 mutant, whereas RNA interference (RNAi)-mediated silencing of AIF1 rescued bri1-301, suggesting their function redundancy in blocking BR signaling. Interestingly, both ATBS1 and AIF1 were previously discovered as BR-regulated genes with ATBS1 being down-regulated and AIF1 up-regulated by BR [15]. It remains to be determined how AIF1 and ATBS1 are regulated by BR and how AIF1 inhibits BR signaling. It is interesting to note that ATBS1 was recently discovered as a direct target of MP/ARF5 [26] (Figure 1B), revealing another potential mechanism mediating the auxin-BR crosstalk important for root initiation during embryogenesis.
Figure 2.
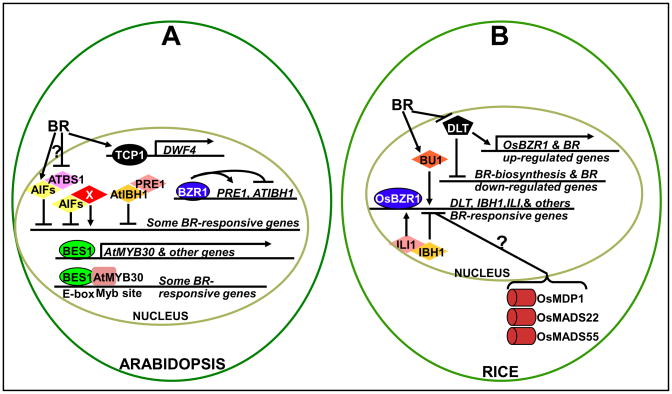
The involvement of bHLH proteins and TFs of other families in BR signaling. (A). In Arabidopsis, BR induces AIF1 and PRE1 but inhibits ATBS1 and AtIBH1 with BZR1 binding to both PRE1 and AtIBH1 promoters. ATBS1 binds AIF1, thus inhibiting AIF1 dimerization with yet to be discovered bHLH proteins (X) that activate a subset of BR-responsive genes, while PRE1 heterodimerizes with AtIBH1 that functions as a negative regulator of BR-mediated cell elongation. BR also induces the expression of TCP1, which encodes a non-canonical bHLH protein that binds and activates the DWF4 promoter to stimulate BR biosynthesis, and stimulate the AtMYB30 gene, a directly target of BES1. BES1 was shown to interact with AtMYB30 on the promoters of some BR-responses genes to amplify BR signaling. (B). In rice, BR induces BU1 and ILI1 but suppresses IBH1 and DLT with OsBZR1 binding to the promoters of ILI1, IBH1, and DLT. ILI1 heterodimerizes with and thus inhibits IBH1 that blocks BR signaling; while DLT promotes BR signaling and plays a role in feedback regulation, likely by activating BR-upregulated genes and OsBZR1. The BR signaling in rice might also be negatively regulated by three SVP-type MADS proteins that likely localize in the cytosol [60].
Support for a role of ATBS1 in BR signaling came from independent studies on two rice genes encoding ATBS1/PRE1-like proteins [31,32] (Figure 2B). BRASSINOSTEROID UPREGULATED 1 (BU1) was discovered as a BR-induced bHLH gene, while INCREASED LAMINA INCLINATION (ILI1) was identified by its activation-tagged mutant phenotype. Overexpression of BU1 or ILI1 resulted in enhanced lamina inclination [31], a characteristic phenotype of enhanced BR signaling in rice, whereas RNAi-triggered silencing of BU1 and its homologs or antisense-mediated inhibition of ILI1 led to reduced lamina inclination and decreased BR sensitivity. How BU1 stimulates BR signaling remains unknown but genetic and biochemical studies of ILI1 revealed a potential mechanism of ILI1 function that seems different from that of ATBS1 [32]. Despite the presence of several AIF homologs in rice, a yeast two-hybrid screen identified a typical bHLH protein as an ILI1-binding bHLH protein (hence the name IBH1) that has a single homolog in Arabidopsis, AtIBH1. Biochemical assays showed that ILI1 and its Arabidopsis homolog PRE1 interact with IBH1 and AtIBH1, while transgenic studies supported a role of the ILI1/PRE1-IBH1/AtIBH1 interaction in modulating BR responses in both rice and Arabidopsis. ChIP and gene expression analyses indicated that BZR1/OsBZR1 binds to the promoters of PRE1/ILI1 and AtIBH1/IBH1, stimulating expression of PRE1/ILI1 but inhibiting transcription of AtIBH1/IBH1.
Given the interactions of ATBS1-AIFs, PRE1-AtIBH1, and a previously reported interaction of KIDARI with HFR1, a well-studied atypical bHLH protein involved in light signaling [37], it will be interesting to investigate if each member of the ATBS1/PRE1 family has specific interaction partners or whether heterodimerization between an ATBS1/PRE1 family member with bHLH proteins is mainly dictated by their tissue/organ-specific expression patterns. Further studies are also needed to determine the relative importance of each member of the ATBS1/PRE1 or BU1/ILI1 family in BR signaling or in crosstalk with other signaling pathways.
Besides these canonical bHLH proteins, a recent study implicated a regulatory role for a non-canonical bHLH protein TCP1 in BR biosynthesis [38], which was previously known to regulate floral symmetry [39]. TCP1 is a member of the plant-specific TFs sharing a highly conserved ~ 60-amino-acid TCP domain with a bHLH-like structure involved in DNA-binding and protein dimerization [39,40]. Activation-tagging of TCP1 suppresses the weak bri1-5 mutation whereas overexpression of a dominant-negative form of TCP1 resulted in a bri1-5-like dwarf phenotype that could be rescued by BR application. Metabolic profiling and gene expression analysis suggested that TCP1 activates the DWARF4 (DWF4) gene encoding a rate-limiting BR biosynthetic C22-hydroxylase [41], while a ChIP assay demonstrated a direct TCP1 binding to the DWF4 promoter (Figure 2A). Interestingly, TCP1 was induced by BR but inhibited by brassinazol (Brz), a specific inhibitor of BR biosynthesis [42], suggesting that TCP1 is not involved in feedback regulation of BR biosynthesis but might play a role to regulate BR biosynthesis and plant growth in response to certain environmental and developmental signals.
Involvement of members of the GRAS, MADS, and MYB families in BR signaling
In addition to bHLH proteins, TFs of other families were recently implicated in BR signaling, which include a GRAS and three MADS proteins in rice and a MYB protein in Arabidopsis [43-45] (Figure 2B). The rice DWARF and LOW TILLERING (DLT) protein and its Arabidopsis homolog AtGRAS8 constitute a unique subgroup of the GRAS family of the plant-specific TFs [46]. Loss-of-function dlt mutation not only results in a phenotype resembling weak BR-deficient rice mutants, but also reduces BR sensitivity and impairs the BR feedback regulation likely via inhibiting OsBZR1 expression. Intriguingly, the DLT itself is negatively regulated by BRs and its promoter binds OsBRZ1 in vitro, thus revealing a potential regulatory loop likely for maintaining appropriate levels of OsBZR1 and DLT. However, it remains unknown how DLT affects OsBZR1 expression and whether the Arabidopsis homolog of DLT, AtGRAS8, plays a similar role in BR signaling.
Besides OsBZR1 and DLT, the BR signaling in rice might also involve three related MADS proteins [44,45] (Figure 2B) that are members of the SHORT VEGETATIVE PHASE (SVP)-subfamily of the MADS TFs, named after the Arabidopsis SVP known to regulate flowering time and floral meristem identity [47]. Oryza sativa MADS-domain-containing protein 1 (OsMDP1) was identified as a BR-repressed gene and its silencing resulted in enhanced BR signaling [45]. Mutations in one of its homolog, OsMADS22, had no detectable growth defect, but silencing of the other homolog, OsMADS55, led to an increased lamina inclination with normal sensitivity to exogenously BR [44]. Interestingly, simultaneously silencing of all three MADS genes gave rise to a larger angle of the lamina inclination with enhanced BR sensitivity, arguing for their redundant roles in regulating BR responses. Further studies are needed to investigate how they regulate BR signaling and to determine whether the involvement of the SVP-type MADS proteins in BR signaling is evolutionarily conserved in Arabidopsis.
As mentioned above, at least 50 TFs are regulated by BR in Arabidopsis [5]. Using a ChIP assay, Li et al showed that one of these TFs, AtMYB30, is a direct BES1 target [48]. Genetic, physiology, and microarray analyses suggested that AtMYB30 plays a role in regulating a subset of BR-responsive genes. Remarkably, AtMYB30 also interacts with BES1 and both proteins can bind to promoters of two well-studied BR-responsive genes, suggesting an interesting model by which BES1 acts cooperatively with its target to amplify BR signaling (Figure 2B). Because AtMYB30 is involved in the hypersensitive response to pathogen attack [49], the BES1-mediated up-regulation of AtMYB30 provides a plausible mechanism by which BR enhances pathogen resistance. In addition to AtMYB30, BR also induces the expression of another well-studied MYB TF, WEREWOLF (WER) [50], to influence root hair formation in Arabidopsis [51], but it remains to be determined whether WER is another target of BES1.
BES1-mediated recruitment of other nuclear proteins
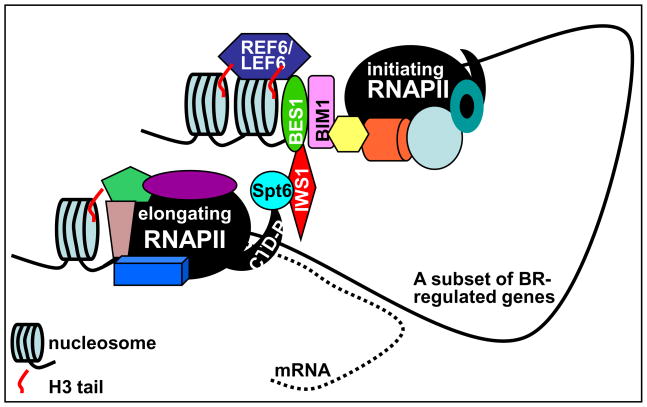
It is well known that eukaryotic gene regulation involves proteins other than TFs, such as transcription elongation factors and chromatin-modifying proteins. The nuclear regulation of BR signaling is no exception (Figure 3). It was shown recently that the full transcriptional activity of BES1 requires AtIWS1 [52], an Arabidopsis homolog of the yeast/human Interacting-With-Spt6 1 (IWS1) known to regulate mRNA export and histone modification [53]. A loss-of-AtIWS1 mutation alters expression of many BR-responsive genes, giving rise to a semi-dwarf phenotype with reduced BR sensitivity. AtIWS1 interacts not only with BES1 but also with the Arabidopsis homolog of the yeast Spt6, a well-studied transcription elongation factor [53], suggesting that AtIWS1 is likely recruited by BES1 to promote transcription elongation of BR-responsive genes. Indeed, AtIWS1 binds preferentially to the transcribed region of a BES1 target gene and that a loss-of-AtIWS1 mutant is hypersensitive to a transcription elongation inhibitor. Interestingly, the human IWS1 recruits a histone methyltransferase to the elongating RNA polymerase II complex, thus stimulating trimethylation of lysine 36 on histone 3 (H3K36me3) crucial for nucleosome reassembly during transcription elongation [54]. It will be interestingly to know if recruitment of a H3K36 methyltransferase by IWS1 is evolutionarily conserved in plants.
Figure 3.
BES1-mediated recruitment of elongation factors and histone-modifying proteins. BES1 not only binds histone-modifying proteins ELF6 and REF6 in the promoters of BR-responsive genes to initiate their transcription, most likely through demethylating histone H3 and altering chromatin structure, but also recruits elongation factors to the phosphorylated C-terminal domain (CTD-P) of the RNA polymerase II (RNAPII) within the transcribed regions of BR-regulated genes to promote the production and export of their mRNAs.
A role of histone modification in BR-regulated gene expression was revealed by a recent exciting discovery of enhancement of the BES1 transcriptional activity by ELF6 and REF6 [55], two similar proteins known to antagonistically regulate Arabidopsis flowering [56]. Both proteins contain the highly conserved Jumonji N/C domain present in a group of histone H3 demethylases that demethylate di or trimethylated lysine 9 or lysine 36 on histone H3 (H3K9me2/3 preventing transcription or H3K36me2/3 activating transcription) [57]. BES1 interacts with ELF6 and REF6 to regulate the expression of ~13% of the previously identified BR-induced genes, while a ChIP experiment detected elevated H3K9m3 levels in the promoter region of a BES1 target gene in both elf6 and ref6 mutants, supporting a role of ELF6 and REF6 in posttranslational H3 modification. The discovery of ELF6 and REF6 as BES1-intercating factors also suggested a potential molecular link between BR signaling and plant flowering [58].
Conclusions and perspectives
These studies revealed that BR-regulated gene expression is much more complicated than what is expected from a simple linear signaling pathway involving only two highly similar TFs. Some of the discussed TFs were potential targets of BES1 or BZR1/OsBZR1 based on ChIP experiments, but it is important to experimentally verify that the detected TF-promoter binding is necessary for the BR-regulated expression of a target gene of interest. It is expected that other TFs are likely regulated independently of BES1/BZR1 and further studies are needed to elucidate their regulatory mechanisms. For example, a preliminary assay showed that AIF1 was phosphorylated by BIN2 [33], but it remains unknown if AIF1 is phosphorylated by BIN2 in vivo and if so, how the BIN2-catalyzed AIF1 phosphorylation affects its inhibitory activity. Regardless of their regulatory mechanisms, a full understanding of BR signaling demands identification of the target genes of not only BES1 and BZR1 but also the BR-regulated TFs. The ChIP method coupled with the whole genome-tiling arrays (ChIP-CHIP) or next-generation sequencing technologies (ChIP-SEQ) [59] will generate genome-wide binding profiles for each of these BR-regulated TFs and reveal interesting transcriptional networks of BR-regulated processes. It is anticipated that classical and novel genetic approaches, yeast one/two-hybrid screens, and proteomic experiments will uncover additional proteins that regulate the nuclear activities of the BR signaling.
Acknowledgments
I apologize to those colleagues whose works I was unable to cite due to the space limitation of this article. Research on BR signaling in my lab is supported by grants from National Institutes of Health (GM060519) and Department of Energy (ER15672).
Footnotes
Competing interest
I declare that I have no competing financial interest
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and recommended reading
Papers of particular interest, published within the annual period of review, have been highlighted as:
• of special interest
•• of outstanding interest
- 1.Clouse SD, Sasse JM. BRASSINOSTEROIDS: Essential Regulators of Plant Growth and Development. Annu Rev Plant Physiol Plant Mol Biol. 1998;49:427–451. doi: 10.1146/annurev.arplant.49.1.427. [DOI] [PubMed] [Google Scholar]
- 2.Kim TW, Wang ZY. Brassinosteroid Signal Transduction from Receptor Kinases to Transcription Factors. Annu Rev Plant Biol. 2010;61:681–704. doi: 10.1146/annurev.arplant.043008.092057. [DOI] [PubMed] [Google Scholar]
- 3.Li J, Jin H. Regulation of brassinosteroid signaling. Trends Plant Sci. 2007;12:37–41. doi: 10.1016/j.tplants.2006.11.002. [DOI] [PubMed] [Google Scholar]
- 4.Bai MY, Zhang LY, Gampala SS, Zhu SW, Song WY, Chong K, Wang ZY. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc Natl Acad Sci U S A. 2007;104:13839–13844. doi: 10.1073/pnas.0706386104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vert G, Nemhauser JL, Geldner N, Hong F, Chory J. Molecular mechanisms of steroid hormone signaling in plants. Annu Rev Cell Dev Biol. 2005;21:177–201. doi: 10.1146/annurev.cellbio.21.090704.151241. [DOI] [PubMed] [Google Scholar]
- 6.Grove MD, Spencer GF, Rohwedder WK, Mandava NB, Worley JF, Warthen JDJ, Steffens GL, Flippen-Anderson JL, Cook JCJ. Brassinolide, a plant growth-promoting steroid isolated from Brassica napus pollen. Nature. 1979;281:216–217. [Google Scholar]
- 7••.Ye Q, Zhu W, Li L, Zhang S, Yin Y, Ma H, Wang X. Brassinosteroids control male fertility by regulating the expression of key genes involved in Arabidopsis anther and pollen development. Proc Natl Acad Sci U S A. 2010;107:6100–6105. doi: 10.1073/pnas.0912333107. This study shows that mutations blocking BR biosynthesis/signaling accumulated lower transcript levels of key transcription factors critical for anther/pollen development and uses ChIP assays to demonstrate BES1 binding of their promoter, thus revealing a potential molecular mechanism by which BR affects the male fertility. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Guo H, Li L, Ye H, Yu X, Algreen A, Yin Y. Three related receptor-like kinases are required for optimal cell elongation in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2009;106:7648–7653. doi: 10.1073/pnas.0812346106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Guo H, Ye H, Li L, Yin Y. A family of receptor-like kinases are regulated by BES1 and involved in plant growth in Arabidopsis thaliana. Plant Signal Behav. 2009;4:784–786. doi: 10.4161/psb.4.8.9231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Escobar-Restrepo JM, Huck N, Kessler S, Gagliardini V, Gheyselinck J, Yang WC, Grossniklaus U. The FERONIA receptor-like kinase mediates male-female interactions during pollen tube reception. Science. 2007;317:656–660. doi: 10.1126/science.1143562. [DOI] [PubMed] [Google Scholar]
- 11.Moller B, Weijers D. Auxin control of embryo patterning. Cold Spring Harb Perspect Biol. 2009;1:a001545. doi: 10.1101/cshperspect.a001545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yin Y, Vafeados D, Tao Y, Yoshida S, Asami T, Chory J. A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell. 2005;120:249–259. doi: 10.1016/j.cell.2004.11.044. [DOI] [PubMed] [Google Scholar]
- 13•.Chandler JW, Cole M, Flier A, Werr W. BIM1, a bHLH protein involved in brassinosteroid signaling, controls Arabidopsis embryonic patterning via interaction with DORNROSCHEN and DORNROSCHEN-LIKE. Plant Mol Biol. 2009;69:57–68. doi: 10.1007/s11103-008-9405-6. This study reports biochemical and genetic interaction between BIM1 and three other transcription factors that were previously implicated in embryogenesis, thus revealing a potential functional link between the plant steroid hormone and embryogenesis. [DOI] [PubMed] [Google Scholar]
- 14.Chandler JW, Cole M, Flier A, Grewe B, Werr W. The AP2 transcription factors DORNROSCHEN and DORNROSCHEN-LIKE redundantly control Arabidopsis embryo patterning via interaction with PHAVOLUTA. Development. 2007;134:1653–1662. doi: 10.1242/dev.001016. [DOI] [PubMed] [Google Scholar]
- 15.Nemhauser JL, Mockler TC, Chory J. Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol. 2004;2:E258. doi: 10.1371/journal.pbio.0020258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li J, Nam KH. Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science. 2002;295:1299–1301. doi: 10.1126/science.1065769. [DOI] [PubMed] [Google Scholar]
- 17•.Vert G, Walcher CL, Chory J, Nemhauser JL. Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc Natl Acad Sci U S A. 2008;105:9829–9834. doi: 10.1073/pnas.0803996105. The authors took a combinatory approach of genetics, genomics, molecular biology, and biochemistry to suggest a potential role of the BIN2-catalyzed phosphorylation of ARF2 in the well-established auxin-BR interaction. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Okushima Y, Mitina I, Quach HL, Theologis A. AUXIN RESPONSE FACTOR 2 (ARF2): a pleiotropic developmental regulator. Plant J. 2005;43:29–46. doi: 10.1111/j.1365-313X.2005.02426.x. [DOI] [PubMed] [Google Scholar]
- 19.Peng P, Yan Z, Zhu Y, Li J. Regulation of the Arabidopsis GSK3-like kinase BRASSINOSTEROID-INSENSITIVE 2 through proteasome-mediated protein degradation. Mol Plant. 2008;1:338–346. doi: 10.1093/mp/ssn001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kim TW, Guan S, Sun Y, Deng Z, Tang W, Shang JX, Burlingame AL, Wang ZY. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat Cell Biol. 2009;11:1254–1260. doi: 10.1038/ncb1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mouchel CF, Briggs GC, Hardtke CS. Natural genetic variation in Arabidopsis identifies BREVIS RADIX, a novel regulator of cell proliferation and elongation in the root. Genes Dev. 2004;18:700–714. doi: 10.1101/gad.1187704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Briggs GC, Mouchel CF, Hardtke CS. Characterization of the plant-specific BREVIS RADIX gene family reveals limited genetic redundancy despite high sequence conservation. Plant Physiol. 2006;140:1306–1316. doi: 10.1104/pp.105.075382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Scacchi E, Osmont KS, Beuchat J, Salinas P, Navarrete-Gomez M, Trigueros M, Ferrandiz C, Hardtke CS. Dynamic, auxin-responsive plasma membrane-to-nucleus movement of Arabidopsis BRX. Development. 2009;136:2059–2067. doi: 10.1242/dev.035444. [DOI] [PubMed] [Google Scholar]
- 24.Mouchel CF, Osmont KS, Hardtke CS. BRX mediates feedback between brassinosteroid levels and auxin signalling in root growth. Nature. 2006;443:458–461. doi: 10.1038/nature05130. [DOI] [PubMed] [Google Scholar]
- 25.Szekeres M, Nemeth K, Koncz-Kalman Z, Mathur J, Kauschmann A, Altmann T, Redei GP, Nagy F, Schell J, Koncz C. Brassinosteroids rescue the deficiency of CYP90, a cytochrome P450, controlling cell elongation and de-etiolation in Arabidopsis. Cell. 1996;85:171–182. doi: 10.1016/s0092-8674(00)81094-6. [DOI] [PubMed] [Google Scholar]
- 26.Schlereth A, Moller B, Liu W, Kientz M, Flipse J, Rademacher EH, Schmid M, Jurgens G, Weijers D. MONOPTEROS controls embryonic root initiation by regulating a mobile transcription factor. Nature. 2010;464:913–916. doi: 10.1038/nature08836. [DOI] [PubMed] [Google Scholar]
- 27.Hardtke CS, Berleth T. The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J. 1998;17:1405–1411. doi: 10.1093/emboj/17.5.1405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Friedrichsen DM, Nemhauser J, Muramitsu T, Maloof JN, Alonso J, Ecker JR, Furuya M, Chory J. Three redundant brassinosteroid early response genes encode putative bHLH transcription factors required for normal growth. Genetics. 2002;162:1445–1456. doi: 10.1093/genetics/162.3.1445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Toledo-Ortiz G, Huq E, Quail PH. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell. 2003;15:1749–1770. doi: 10.1105/tpc.013839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li X, Duan X, Jiang H, Sun Y, Tang Y, Yuan Z, Guo J, Liang W, Chen L, Yin J, et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol. 2006;141:1167–1184. doi: 10.1104/pp.106.080580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31•.Tanaka A, Nakagawa H, Tomita C, Shimatani Z, Ohtake M, Nomura T, Jiang CJ, Dubouzet JG, Kikuchi S, Sekimoto H, et al. BRASSINOSTEROID UPREGULATED 1, Encoding a Helix-Loop-Helix Protein, is a Novel Gene Involved in Brassinosteroid Signaling and Controls Bending of the Lamina Joint in Rice. Plant Physiol. 2009;151:669–680. doi: 10.1104/pp.109.140806. In this paper, the authors describe the discovery and functional analysis of a BR-induced atypical bHLH protein and suggest that the BR-induced BU1 expression is mediated by two BR signaling mechanism through a rice BRI1 homolog OsBRI1 and the rice heterotrimeric G protein alpha subunit RGA1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32••.Zhang LY, Bai MY, Wu J, Zhu JY, Wang H, Zhang Z, Wang W, Sun Y, Zhao J, Sun X, et al. Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis. Plant Cell. 2009;21:3767–3780. doi: 10.1105/tpc.109.070441. This study combines a rice activation tagging-based genetic screen with the yeast two-hybrid approach to discovery a rice BU1-like atypical bHLH protein Increased Leaf Inclination1 (ILI1) and its interacting partner ILI1-Binding bHLH Protein1 (IBH1) and demonstrate the evolutionarily conserved function and regulation of the antagonistic bHLH pair in rice and Arabidopsis. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33••.Wang H, Zhu Y, Fujioka S, Asami T, Li J. Regulation of Arabidopsis brassinosteroid signaling by atypical basic helix-loop-helix proteins. Plant Cell. 2009;21:3781–3791. doi: 10.1105/tpc.109.072504. This study not only revealed regulatory roles of several Arabidopsis BU1-like atypical bHLH proteins in BR signaling but also identified their interacting proteins and suggested potential biochemical mechanisms by which the atypical bHLH proteins affect BR-mediated plant growth. This study along with the studies of [31•] and [32••] also raised an interesting question on how the signaling specificity of each atypical bHLH protein is achieved. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kang B, Wang H, Nam KH, Li J. Activation-tagged suppressors of a weak brassinosteroid receptor mutant. Mol Plant. 2010;3:260–268. doi: 10.1093/mp/ssp099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hyun Y, Lee I. KIDARI, encoding a non-DNA Binding bHLH protein, represses light signal transduction in Arabidopsis thaliana. Plant Mol Biol. 2006;61:283–296. doi: 10.1007/s11103-006-0010-2. [DOI] [PubMed] [Google Scholar]
- 36.Lee S, Yang KY, Kim YM, Park SY, Kim SY, Soh MS. Overexpression of PRE1 and its homologous genes activates Gibberellin-dependent responses in Arabidopsis thaliana. Plant Cell Physiol. 2006;47:591–600. doi: 10.1093/pcp/pcj026. [DOI] [PubMed] [Google Scholar]
- 37.Fairchild CD, Schumaker MA, Quail PH. HFR1 encodes an atypical bHLH protein that acts in phytochrome A signal transduction. Genes Dev. 2000;14:2377–2391. [PMC free article] [PubMed] [Google Scholar]
- 38••.Guo Z, Fujioka S, Blancaflor EB, Miao S, Guo X, Li J. TCP1 controls brassinosteroid biosynthesis by regulating the expression of the key biosynthetic gene DWARF4 in Arabidopsis thaliana. Plant Cell. 2010;22:1161–1173. doi: 10.1105/tpc.109.069203. This study identified a potential transcriptional factor that is required for activating the transcription of the rate-liming BR biosynthetic DWF4 gene, thus increasing our understanding of BR homeostasis. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Martin-Trillo M, Cubas P. TCP genes: a family snapshot ten years later. Trends Plant Sci. 2010;15:31–39. doi: 10.1016/j.tplants.2009.11.003. [DOI] [PubMed] [Google Scholar]
- 40.Aggarwal P, Das Gupta M, Joseph AP, Chatterjee N, Srinivasan N, Nath U. Identification of Specific DNA Binding Residues in the TCP Family of Transcription Factors in Arabidopsis. Plant Cell. 2010 doi: 10.1105/tpc.1109.066647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Choe S, Dilkes BP, Fujioka S, Takatsuto S, Sakurai A, Feldmann KA. The DWF4 gene of Arabidopsis encodes a cytochrome P450 that mediates multiple 22alpha-hydroxylation steps in brassinosteroid biosynthesis. Plant Cell. 1998;10:231–243. doi: 10.1105/tpc.10.2.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Nagata N, Asami T, Yoshida S. Brassinazole, an inhibitor of brassinosteroid biosynthesis, inhibits development of secondary xylem in cress plants (Lepidium sativum) Plant Cell Physiol. 2001;42:1006–1011. doi: 10.1093/pcp/pce122. [DOI] [PubMed] [Google Scholar]
- 43••.Tong H, Jin Y, Liu W, Li F, Fang J, Yin Y, Qian Q, Zhu L, Chu C. DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J. 2009;58:803–816. doi: 10.1111/j.1365-313X.2009.03825.x. This study identified a unique GRAS protein as a potential target of the rice homolog of OsBZR1 and showed that this GRAS protein plays a key regulator role in BR signaling in rice. It remains unknown if its Arabidopsis homolog has a similar function. [DOI] [PubMed] [Google Scholar]
- 44.Lee S, Choi SC, An G. Rice SVP-group MADS-box proteins, OsMADS22 and OsMADS55, are negative regulators of brassinosteroid responses. Plant J. 2008;54:93–105. doi: 10.1111/j.1365-313X.2008.03406.x. [DOI] [PubMed] [Google Scholar]
- 45.Duan K, Li L, Hu P, Xu SP, Xu ZH, Xue HW. A brassinolide-suppressed rice MADS-box transcription factor, OsMDP1, has a negative regulatory role in BR signaling. Plant J. 2006;47:519–531. doi: 10.1111/j.1365-313X.2006.02804.x. [DOI] [PubMed] [Google Scholar]
- 46.Tian C, Wan P, Sun S, Li J, Chen M. Genome-wide analysis of the GRAS gene family in rice and Arabidopsis. Plant Mol Biol. 2004;54:519–532. doi: 10.1023/B:PLAN.0000038256.89809.57. [DOI] [PubMed] [Google Scholar]
- 47.Liu C, Xi W, Shen L, Tan C, Yu H. Regulation of floral patterning by flowering time genes. Dev Cell. 2009;16:711–722. doi: 10.1016/j.devcel.2009.03.011. [DOI] [PubMed] [Google Scholar]
- 48•.Li L, Yu X, Thompson A, Guo M, Yoshida S, Asami T, Chory J, Yin Y. Arabidopsis MYB30 is a direct target of BES1 and cooperates with BES1 to regulate brassinosteroid-induced gene expression. Plant J. 2009;58:275–286. doi: 10.1111/j.1365-313X.2008.03778.x. Combining microarray and ChIP experiments, the authors showed that AtMYB30 is likely a direct target of BES1 and that it works together with BES1 to promote the expression of a subset of BR-responsive genes. Given the previous findings implicating AtMYB30 in plant pathogen resistance, this work also revealed a potential mechanism for BR enhanced pathogen resistance. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Vailleau F, Daniel X, Tronchet M, Montillet JL, Triantaphylides C, Roby D. A R2R3-MYB gene, AtMYB30, acts as a positive regulator of the hypersensitive cell death program in plants in response to pathogen attack. Proc Natl Acad Sci U S A. 2002;99:10179–10184. doi: 10.1073/pnas.152047199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lee MM, Schiefelbein J. WEREWOLF, a MYB-related protein in Arabidopsis, is a position-dependent regulator of epidermal cell patterning. Cell. 1999;99:473–483. doi: 10.1016/s0092-8674(00)81536-6. [DOI] [PubMed] [Google Scholar]
- 51.Kuppusamy KT, Chen AY, Nemhauser JL. Steroids are required for epidermal cell fate establishment in Arabidopsis roots. Proc Natl Acad Sci U S A. 2009;106:8073–8076. doi: 10.1073/pnas.0811633106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52••.Li L, Ye H, Guo H, Yin Y. Arabidopsis IWS1 interacts with transcription factor BES1 and is involved in plant steroid hormone brassinosteroid regulated gene expression. Proc Natl Acad Sci U S A. 2010;107:3918–3923. doi: 10.1073/pnas.0909198107. This study indicated that the BR-regulated gene expression not only requires common DNA-binding transcriptional factors to initiate transcription but also need transcriptional elongation factors to ensure successful biosynthesis and export of mRNAs. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Bres V, Yoh SM, Jones KA. The multi-tasking P-TEFb complex. Curr Opin Cell Biol. 2008;20:334–340. doi: 10.1016/j.ceb.2008.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yoh SM, Lucas JS, Jones KA. The Iws1:Spt6:CTD complex controls cotranscriptional mRNA biosynthesis and HYPB/Setd2-mediated histone H3K36 methylation. Genes Dev. 2008;22:3422–3434. doi: 10.1101/gad.1720008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55••.Yu X, Li L, Guo M, Chory J, Yin Y. Modulation of brassinosteroid-regulated gene expression by Jumonji domain-containing proteins ELF6 and REF6 in Arabidopsis. Proc Natl Acad Sci U S A. 2008;105:7618–7623. doi: 10.1073/pnas.0802254105. The study suggested that the full transcriptional activity of BES1 requires involvement of two Jumonji N/C-domain containing proteins that likely alter the methylation status of histone 3. This is the first report that implicated chromatin-modifying proteins in BR-regulated gene expression. Further studies will likely uncover additional nuclear proteins that are recruited by BR-specific transcription factor to regulate the accessibility of chromatin DNAs. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Noh B, Lee SH, Kim HJ, Yi G, Shin EA, Lee M, Jung KJ, Doyle MR, Amasino RM, Noh YS. Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell. 2004;16:2601–2613. doi: 10.1105/tpc.104.025353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Klose RJ, Kallin EM, Zhang Y. JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet. 2006;7:715–727. doi: 10.1038/nrg1945. [DOI] [PubMed] [Google Scholar]
- 58.Clouse SD. The molecular intersection of brassinosteroid-regulated growth and flowering in Arabidopsis. Proc Natl Acad Sci U S A. 2008;105:7345–7346. doi: 10.1073/pnas.0803552105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Aleksic J, Russell S. ChIPing away at the genome: the new frontier travel guide. Mol Biosyst. 2009;5:1421–1428. doi: 10.1039/B906179G. [DOI] [PubMed] [Google Scholar]
- 60.Lee S, Jeong DH, An G. A possible working mechanism for rice SVP-group MADS-box proteins as negative regulators of brassinosteroid responses. Plant Signal Behav. 2008;3:471–474. doi: 10.4161/psb.3.7.5677. [DOI] [PMC free article] [PubMed] [Google Scholar]