Abstract
In the crystal of the title compound, C8H10NO2 +·Cl−, alternating layers of hydrophobic and hydrophilic zones stack along the c axis. The chloride anions are sandwiched between the 4-(carboxymethyl)anilinium layers, forming intermolecular O—H⋯Cl and N—H⋯Cl hydrogen bonds with the ammonium and carboxyl groups of the cations. In addition, intermolecular N—H⋯O and weak C—H⋯O and C—H⋯Cl hydrogen bonds help stabilize the crystal structure.
Related literature
For our ongoing studies of hydrogen-bonding interactions in the crystal structures of protonated amines, see: Benslimane et al. (2007 ▶); Bouacida et al. (2005a
▶,b
▶,c
▶, 2006 ▶, 2007 ▶, 2008 ▶, 2009 ▶). For amino acids in which the amino N atom is protonated, see: Bouacida et al. (2006); Rademeyer (2004a
▶,b
▶). For a related structure, see: Benslimane et al. (2007 ▶). For bond-length data, see: Allen et al. (1987 ▶).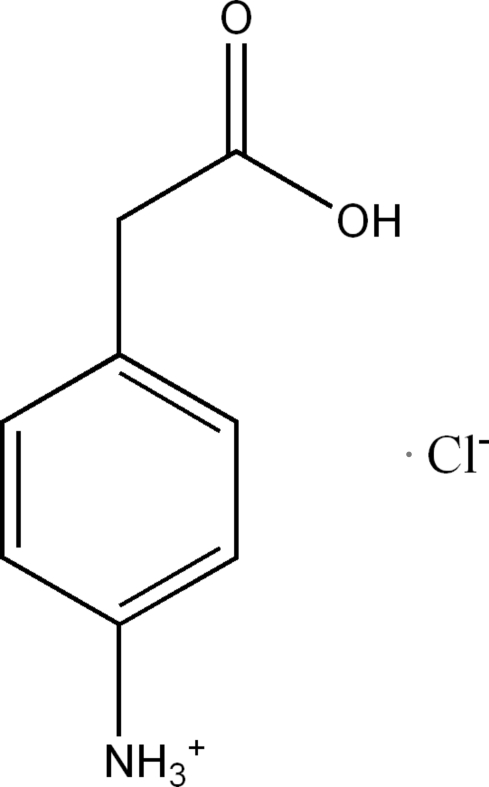
Experimental
Crystal data
C8H10NO2 +·Cl−
M r = 187.62
Monoclinic,

a = 4.4982 (4) Å
b = 11.0790 (11) Å
c = 17.7120 (17) Å
β = 95.429 (3)°
V = 878.73 (14) Å3
Z = 4
Mo Kα radiation
μ = 0.39 mm−1
T = 100 K
0.44 × 0.12 × 0.1 mm
Data collection
Bruker APEXII diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 1998 ▶) T min = 0.809, T max = 0.962
7536 measured reflections
2006 independent reflections
1785 reflections with I > 2σ(I)
R int = 0.040
Refinement
R[F 2 > 2σ(F 2)] = 0.033
wR(F 2) = 0.08
S = 1.03
2006 reflections
113 parameters
H-atom parameters constrained
Δρmax = 0.30 e Å−3
Δρmin = −0.22 e Å−3
Data collection: APEX2 (Bruker, 2001 ▶); cell refinement: SAINT (Bruker, 2001 ▶); data reduction: SAINT; program(s) used to solve structure: SIR2002 (Burla et al., 2005 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: ORTEP-3 for Windows (Farrugia, 1997 ▶) and DIAMOND (Brandenburg & Berndt, 2001 ▶); software used to prepare material for publication: WinGX (Farrugia, 1999 ▶).
Supplementary Material
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536809021849/lh2838sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536809021849/lh2838Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| O1—H1⋯Cl1i | 0.82 | 2.20 | 3.0087 (13) | 171 |
| N1—H1A⋯O2ii | 0.89 | 1.98 | 2.8517 (17) | 167 |
| N1—H1B⋯Cl1iii | 0.89 | 2.41 | 3.2285 (13) | 152 |
| N1—H1C⋯Cl1iv | 0.89 | 2.26 | 3.1516 (14) | 174 |
| C2—H2⋯O2ii | 0.93 | 2.49 | 3.2338 (18) | 137 |
| C3—H3⋯Cl1 | 0.93 | 2.82 | 3.7481 (15) | 175 |
Symmetry codes: (i)  ; (ii)
; (ii)  ; (iii)
; (iii)  ; (iv)
; (iv)  .
.
Acknowledgments
The authors are grateful to Dr Thierry Roisnel, Centre de Diffractométrie X (CDIFX) de Rennes, Université de Rennes 1, France, for data-collection facilities. SB thanks Université A. Mira de Béjaia, Algeria, for financial support.
supplementary crystallographic information
Comment
The title compound, was prepared as part of our ongoing studies of hydrogen-bonding interactions in the crystal structure of protonated amines (Bouacida et al., 2005a,b,c, 2006, 2007, 2008, 2009).
The molecular structure of (I), and the atomic numbering used, is illustrated in Fig. 1. All bond distances (Allen et al., 1987) and angles are within the ranges of accepted values. The amino N atom is protonated as in previously reported amino acids (Bouacida & al., 2006; Rademeyer, 2004a,b). The layered crystal packing of (I) is shown in Fig. 2, in which cations form alterning layers of 4-(carboxymethyl)anilinium of hydrophobic and hydrophilic zones along the c axis, and the chloride ions are located between these layers. In the structure, two types of classical hydrogen bonds are observed, viz. cation–anion and cation–cation (Fig. 3). The 4-(carboxymethyl)anilinium cations and the chloride anions form hydrogen-bonded double layers at z = 0 and z = 1/2, linked by N—H···Cl, C—H···Cl and O—H···Cl hydrogen bonds. Additional hydrogen-bonding parameters are listed in Table 1. These interaction bonds link the cations and the anions together, forming a three-dimensional network and reinforcing the cohesion of the ionic structure.
Experimental
The title compound was crystallized by slow evaporation of an aqueous solution of 4-aminophenyl acetic acid, tin(II) chloride dihydrate and hydrochloric acid in a molar ratio of 5:5:1. White stick-like crystals were obtained after two weeks.
Refinement
All H atoms were located in Fourier maps but introduced in calculated positions and treated as riding on their parent C, O and N atoms with C—H = 0.93–0.97 Å, O—H = 0.82Å and N—H = 0.89Å and Uiso(H) =1.5–1.2(carrier atom).
Figures
Fig. 1.
The asymmetric unit of the title compound with the atomic labelling scheme. Displacement are drawn at the 50% probability level.
Fig. 2.
Part of the crystal structure illustrating the molecular layers, viewed along the b axis.
Fig. 3.
Part of the crystal structure with hydrogen bonds shown as dashed lines, viewed along the a axis.
Crystal data
| C8H10NO2+·Cl− | F(000) = 392 |
| Mr = 187.62 | Dx = 1.418 Mg m−3 |
| Monoclinic, P21/n | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2yn | Cell parameters from 3208 reflections |
| a = 4.4982 (4) Å | θ = 2.3–27.4° |
| b = 11.0790 (11) Å | µ = 0.39 mm−1 |
| c = 17.7120 (17) Å | T = 100 K |
| β = 95.429 (3)° | Stick, white |
| V = 878.73 (14) Å3 | 0.44 × 0.12 × 0.1 mm |
| Z = 4 |
Data collection
| Bruker APEXII diffractometer | 1785 reflections with I > 2σ(I) |
| graphite | Rint = 0.040 |
| CCD rotation images, thin slices scans | θmax = 27.5°, θmin = 3.7° |
| Absorption correction: multi-scan (SADABS; Bruker, 1998) | h = −5→5 |
| Tmin = 0.809, Tmax = 0.962 | k = −14→14 |
| 7536 measured reflections | l = −22→22 |
| 2006 independent reflections |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.033 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.08 | H-atom parameters constrained |
| S = 1.03 | w = 1/[σ2(Fo2) + (0.0283P)2 + 0.5301P] where P = (Fo2 + 2Fc2)/3 |
| 2006 reflections | (Δ/σ)max = 0.001 |
| 113 parameters | Δρmax = 0.30 e Å−3 |
| 0 restraints | Δρmin = −0.22 e Å−3 |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| Cl1 | 0.33999 (7) | 0.14718 (3) | 0.035821 (18) | 0.01567 (11) | |
| O1 | 0.7756 (3) | 0.59938 (11) | 0.02054 (6) | 0.0270 (3) | |
| H1 | 0.7238 | 0.6660 | 0.0040 | 0.040* | |
| O2 | 0.5016 (3) | 0.65210 (10) | 0.11399 (6) | 0.0257 (3) | |
| N1 | 0.3016 (3) | 0.37706 (11) | 0.40929 (7) | 0.0145 (3) | |
| H1A | 0.1824 | 0.3128 | 0.4051 | 0.022* | |
| H1B | 0.1969 | 0.4414 | 0.4212 | 0.022* | |
| H1C | 0.4494 | 0.3642 | 0.4455 | 0.022* | |
| C1 | 0.4271 (3) | 0.39836 (13) | 0.33671 (8) | 0.0126 (3) | |
| C2 | 0.3601 (3) | 0.32006 (13) | 0.27659 (8) | 0.0148 (3) | |
| H2 | 0.2377 | 0.2535 | 0.2817 | 0.018* | |
| C3 | 0.4804 (3) | 0.34312 (13) | 0.20801 (8) | 0.0155 (3) | |
| H3 | 0.4373 | 0.2911 | 0.1672 | 0.019* | |
| C4 | 0.6631 (3) | 0.44257 (13) | 0.19984 (8) | 0.0145 (3) | |
| C5 | 0.7279 (3) | 0.51913 (14) | 0.26215 (8) | 0.0176 (3) | |
| H5 | 0.8524 | 0.5853 | 0.2576 | 0.021* | |
| C6 | 0.6098 (3) | 0.49813 (14) | 0.33049 (8) | 0.0166 (3) | |
| H6 | 0.6522 | 0.5499 | 0.3714 | 0.020* | |
| C7 | 0.7903 (3) | 0.46817 (14) | 0.12557 (8) | 0.0180 (3) | |
| H7A | 0.7456 | 0.4006 | 0.0916 | 0.022* | |
| H7B | 1.0060 | 0.4744 | 0.1347 | 0.022* | |
| C8 | 0.6709 (3) | 0.58249 (14) | 0.08711 (8) | 0.0157 (3) |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Cl1 | 0.01972 (19) | 0.01379 (19) | 0.01353 (18) | 0.00041 (13) | 0.00178 (13) | 0.00186 (12) |
| O1 | 0.0392 (7) | 0.0206 (6) | 0.0239 (6) | 0.0128 (5) | 0.0181 (5) | 0.0103 (5) |
| O2 | 0.0361 (7) | 0.0226 (6) | 0.0203 (6) | 0.0151 (5) | 0.0123 (5) | 0.0061 (5) |
| N1 | 0.0169 (6) | 0.0146 (6) | 0.0124 (6) | −0.0011 (5) | 0.0034 (5) | −0.0013 (5) |
| C4 | 0.0141 (6) | 0.0146 (7) | 0.0150 (7) | 0.0054 (5) | 0.0026 (5) | 0.0034 (5) |
| C1 | 0.0131 (6) | 0.0142 (7) | 0.0108 (6) | 0.0020 (5) | 0.0025 (5) | 0.0023 (5) |
| C6 | 0.0173 (7) | 0.0163 (7) | 0.0160 (7) | −0.0025 (6) | 0.0003 (5) | −0.0027 (6) |
| C3 | 0.0190 (7) | 0.0138 (7) | 0.0137 (7) | 0.0024 (6) | 0.0011 (5) | −0.0020 (5) |
| C7 | 0.0199 (7) | 0.0173 (7) | 0.0178 (7) | 0.0043 (6) | 0.0071 (6) | 0.0034 (6) |
| C8 | 0.0156 (7) | 0.0161 (7) | 0.0157 (7) | 0.0003 (6) | 0.0034 (5) | 0.0007 (6) |
| C5 | 0.0152 (7) | 0.0173 (7) | 0.0206 (7) | −0.0039 (6) | 0.0032 (5) | 0.0013 (6) |
| C2 | 0.0161 (6) | 0.0123 (7) | 0.0159 (7) | −0.0021 (6) | 0.0014 (5) | 0.0000 (6) |
Geometric parameters (Å, °)
| O1—C8 | 1.3234 (17) | C1—C6 | 1.388 (2) |
| O1—H1 | 0.8200 | C6—C5 | 1.387 (2) |
| O2—C8 | 1.2121 (18) | C6—H6 | 0.9300 |
| N1—C1 | 1.4709 (17) | C3—C2 | 1.399 (2) |
| N1—H1A | 0.8900 | C3—H3 | 0.9300 |
| N1—H1B | 0.8900 | C7—C8 | 1.512 (2) |
| N1—H1C | 0.8900 | C7—H7A | 0.9700 |
| C4—C3 | 1.390 (2) | C7—H7B | 0.9700 |
| C4—C5 | 1.401 (2) | C5—H5 | 0.9300 |
| C4—C7 | 1.5100 (19) | C2—H2 | 0.9300 |
| C1—C2 | 1.384 (2) | ||
| C8—O1—H1 | 109.5 | C4—C3—H3 | 119.5 |
| C1—N1—H1A | 109.5 | C2—C3—H3 | 119.5 |
| C1—N1—H1B | 109.5 | C4—C7—C8 | 113.72 (12) |
| H1A—N1—H1B | 109.5 | C4—C7—H7A | 108.8 |
| C1—N1—H1C | 109.5 | C8—C7—H7A | 108.8 |
| H1A—N1—H1C | 109.5 | C4—C7—H7B | 108.8 |
| H1B—N1—H1C | 109.5 | C8—C7—H7B | 108.8 |
| C3—C4—C5 | 118.62 (13) | H7A—C7—H7B | 107.7 |
| C3—C4—C7 | 121.06 (13) | O2—C8—O1 | 123.32 (14) |
| C5—C4—C7 | 120.33 (13) | O2—C8—C7 | 124.41 (13) |
| C2—C1—C6 | 121.69 (13) | O1—C8—C7 | 112.27 (12) |
| C2—C1—N1 | 119.89 (12) | C6—C5—C4 | 121.20 (14) |
| C6—C1—N1 | 118.42 (12) | C6—C5—H5 | 119.4 |
| C5—C6—C1 | 118.77 (13) | C4—C5—H5 | 119.4 |
| C5—C6—H6 | 120.6 | C1—C2—C3 | 118.67 (13) |
| C1—C6—H6 | 120.6 | C1—C2—H2 | 120.7 |
| C4—C3—C2 | 121.05 (13) | C3—C2—H2 | 120.7 |
| C2—C1—C6—C5 | −0.1 (2) | C4—C7—C8—O1 | −176.97 (13) |
| N1—C1—C6—C5 | −179.72 (13) | C1—C6—C5—C4 | 0.7 (2) |
| C5—C4—C3—C2 | 0.7 (2) | C3—C4—C5—C6 | −1.0 (2) |
| C7—C4—C3—C2 | −179.38 (13) | C7—C4—C5—C6 | 179.07 (13) |
| C3—C4—C7—C8 | 113.71 (16) | C6—C1—C2—C3 | −0.2 (2) |
| C5—C4—C7—C8 | −66.37 (18) | N1—C1—C2—C3 | 179.42 (12) |
| C4—C7—C8—O2 | 3.9 (2) | C4—C3—C2—C1 | −0.1 (2) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···Cl1i | 0.82 | 2.20 | 3.0087 (13) | 171 |
| N1—H1A···O2ii | 0.89 | 1.98 | 2.8517 (17) | 167 |
| N1—H1B···Cl1iii | 0.89 | 2.41 | 3.2285 (13) | 152 |
| N1—H1C···Cl1iv | 0.89 | 2.26 | 3.1516 (14) | 174 |
| C2—H2···O2ii | 0.93 | 2.49 | 3.2338 (18) | 137 |
| C3—H3···Cl1 | 0.93 | 2.82 | 3.7481 (15) | 175 |
Symmetry codes: (i) −x+1, −y+1, −z; (ii) −x+1/2, y−1/2, −z+1/2; (iii) −x+1/2, y+1/2, −z+1/2; (iv) x+1/2, −y+1/2, z+1/2.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: LH2838).
References
- Allen, F. H., Kennard, O., Watson, D. G., Brammer, L., Orpen, A. G. & Taylor, R. (1987). J. Chem. Soc. Perkin Trans. 2, pp. S1–19.
- Benslimane, M., Merazig, H., Bouacida, S., Denbri, S., Beghidja, A. & Ouahab, L. (2007). Acta Cryst. E63, o3682–o3683.
- Bouacida, S. (2008). PhD thesis, Montouri–Constantine University, Algeria.
- Bouacida, S., Belhouas, R., Kechout, H., Merazig, H. & Bénard-Rocherullé, P. (2009). Acta Cryst. E65, o628–o629. [DOI] [PMC free article] [PubMed]
- Bouacida, S., Merazig, H., Beghidja, A. & Beghidja, C. (2005a). Acta Cryst. E61, m1153–m1155.
- Bouacida, S., Merazig, H., Beghidja, A. & Beghidja, C. (2005b). Acta Cryst. E61, m2072–m2074.
- Bouacida, S., Merazig, H., Beghidja, A. & Beghidja, C. (2005c). Acta Cryst. E61, m577–m579.
- Bouacida, S., Merazig, H. & Benard-Rocherulle, P. (2006). Acta Cryst. E62, o838–o840.
- Bouacida, S., Merazig, H., Benard-Rocherulle, P. & Rizzoli, C. (2007). Acta Cryst. E63, m379–m381.
- Brandenburg, K. & Berndt, M. (2001). DIAMOND Version 3.1e. Crystal Impact, Bonn, Germany.
- Bruker (1998). SADABS Bruker AXS Inc., Madison, Wisconsin, USA.
- Bruker (2001). APEX2 and SAINT Bruker AXS Inc., Madison, Wisconsin, USA.
- Burla, M. C., Caliandro, R., Camalli, M., Carrozzini, B., Cascarano, G. L., De Caro, L., Giacovazzo, C., Polidori, G. & Spagna, R. (2005). J. Appl. Cryst.38, 381–388.
- Farrugia, L. J. (1997). J. Appl. Cryst.30, 565.
- Farrugia, L. J. (1999). J. Appl. Cryst.32, 837–838.
- Rademeyer, M. (2004a). Acta Cryst. C60, m55–m56. [DOI] [PubMed]
- Rademeyer, M. (2004b). Acta Cryst. E60, m345–m347.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536809021849/lh2838sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536809021849/lh2838Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report





