Abstract
The N-terminal proline-rich domain of γ-zein (Zera) plays an important role in protein body (PB) formation not only in the original host (maize seeds) but in a broad spectrum of eukaryotic cells. However, the elements within the Zera sequence that are involved in the biogenesis of PBs have not been clearly identified. Here, we focused on amino acid sequence motifs that could be involved in Zera oligomerization, leading to PB-like structures in Nicotiana benthamiana leaves. By using fusions of Zera with fluorescent proteins, we found that the lack of the repeat region (PPPVHL)8 of Zera resulted in the secretion of the fusion protein but that this repeat by itself did not form PBs. Although the repeat region containing eight units was the most efficient for Zera self-assembly, shorter repeats of 4–6 units still formed small multimers. Based on site-directed mutagenesis of Zera cysteine residues and analysis of multimer formation, we conclude that the two N-terminal Cys residues of Zera (Cys7 and Cys9) are critical for oligomerization. Immunoelectron microscopy and confocal studies on PB development over time revealed that early, small, Zera-derived oligomers were sequestered in buds along the rough ER and that the mature size of the PBs could be attained by both cross-linking of preformed multimers and the incorporation of new chains of Zera fusions synthesized by active membrane-bound ribosomes. Based on these results and on the behavior of the Zera structure determined by molecular dynamics simulation studies, we propose a model of Zera-induced PB biogenesis.
Keywords: Molecular Dynamics, Plant, Protein Self-assembly, Protein Sorting, Protein-Protein Interactions, Endoplasmic Reticulum, Plant Transformation, Proline-rich Domain, Protein Aggregates, Protein Bodies
Introduction
The mechanisms by which prolamins, which lack the ER3 retention (H/K)DEL motif, are retained and assembled in the ER-derived PBs are only partially understood. Different factors act as determinants of PB biogenesis, some of them derived from cis-cargo properties and others with a cellular trans-origin (1). It has been proposed that the physico-chemical properties of prolamins, such as hydrophobicity and disulfide bond formation, promote specific interactions resulting in the formation of large self-assemblies that are responsible for their retention in the ER and PB formation (2–4). These polymers could be excluded because of their size from being carried by the COP II vesicles that transport cargo proteins to the Golgi complex (5). However, the generic COPII vesicles are flexible enough for large cargo loading, including that of procollagen (more than 300 nm in size) (6) and the collagen VII trimer (900 kDa) (7).
In maize, four distinct types of prolamins (α-, β-, γ-, and δ-zein) coexist in the PBs (8) and play distinct roles in PB formation. Recently, maize storage protein mutants created through RNAi showed that γ-RNAi maize mutant lines exhibited slightly altered protein body formation and that a more drastic effect was observed in the β-γ combined mutant, where protein bodies showed an irregular shape, particularly in their periphery (9).
Various studies on zein accumulation in heterologous expression systems suggest that γ-zein and β-zein mediate the retention of α- and δ-zeins in the ER lumen. Thus, although γ- and β-zeins accumulated in PBs in transgenic Arabidopsis and tobacco plants (10, 11), α- and δ-zeins only accumulated if they were coexpressed with γ- and β-zeins, respectively, in transgenic tobacco, indicating that γ-zein and β-zein have a stabilizing effect on other zeins (12, 13). Trans-acting interactions between prolamins and ER-residing chaperones, such as BiP, also play a role in prolamin retention by facilitating its folding and assembly (5, 14, 15). In rice, the localized targeting of prolamin mRNAs to distinct ER subdomains also facilitates the assembly and retention of these proteins (16, 17). An essential role of protein-disulfide isomerase in the segregation of rice storage proteins within the ER lumen has also been proposed as a trans-factor (18).
Previously, we have reported that γ-zein was sorted in the ER in leaf cells when expressed in Arabidopsis transgenic plants. Moreover, truncated γ-zeins containing the N-terminal proline-rich domain were sorted and retained in the ER inside the transformed leaf cells (10). These results contrast with what frequently occurs when other truncated proteins are overexpressed; they usually associate with BiP and are ultimately degraded in lytic vacuoles (19, 20) or enter the ER-associated protein degradation (ERAD) pathway (21, 22). The proline-rich N-terminal sequence of γ-zein comprising the first 112 amino acids (the Zera sequence) (23) fused to a variety of reporter and therapeutic proteins, was shown to induce the stable accumulation of the fusion proteins in ER-derived PB-like structures in plant vegetative tissues (24, 25). The ability of a similar sequence to redirect phaseolin, a vacuolar storage protein, to the ER compartment of transformed tobacco leaves in which the fusion protein, named zeolin, is retained and accumulated in PBs, has also been demonstrated (26). Interestingly, the PB-inducing capacity of Zera was also demonstrated in a variety of other (non-plant) eukaryotic cells, including mammalian, insect, and fungal cells (24). This ubiquitous behavior of Zera indicates that intrinsic molecular properties are responsible for the fusion protein assembly and that PB biogenesis is independent of specific seed or plant tissue mechanisms. In this context, a synthetic polypeptide composed of the (PPPVHL)8 repeat units present in the Zera sequence folds in vitro into an amphipathic poly-Pro II conformation (27) with the ability to self-assemble (28). In addition, cysteine residues have an important role in zeolin assembly (4). Recently, PB-like accretions have also been induced in plants overexpressing fusions of proteins with elastin-like polypeptides (29) and with a hydrophobin from Trichoderma reseei (HFBI) (30), both fusions bearing an ER retention motif. Proteins fused to elastin-like polypeptides can be recovered by using inverse transition cycling procedures (31), whereas hydrophobin fusions can be efficiently purified using a surfactant-based aqueous two-phase system (30). The encapsulation of these fusion proteins inside ER-derived PBs seems to be due to the unique intrinsic physico-chemical properties of the fusion partners. This phenomenon could be derived from general ER mechanisms that insulate the exogenous recombinant proteins and segregate them from both the secretory and the degradative vacuolar or ERAD pathways (32).
We report new data on PB biogenesis, obtained using Zera-induced PBs. Here we have attempted to define the minimum structural features of Zera necessary for PB induction. Various Zera-derived sequences were obtained by protein engineering, and their ability to induce protein oligomerization and PB-like organelles in agroinfiltrated tobacco (Nicotiana benthamiana) leaves was analyzed. Our results show the effect of factors derived from both the repeat (PPPVHL)8 region and the number and location of the Cys residues in Zera on PB biogenesis. Furthermore, we analyze the development of PBs over time as well as their ultrastructure. Finally, we propose a model of Zera sequence assembly interactions consistent with induced PB biogenesis.
EXPERIMENTAL PROCEDURES
Molecular Cloning
Plasmid pUC18Zera (24) was used as a template in PCRs designed to introduce a SalI site upstream of the Zera sequence, a SpeI restriction site at the end of the γ-zein signal peptide, and an AscI restriction site followed by a linker sequence encoding five Gly and a NcoI site downstream of the Zera sequence. The PCR fragment replaced the original Zera sequence in pUC18ZeraECFP (24). The cassette containing the new fusion sequence Zera-ECFP was transferred to the binary vector pC2300 (see the Cambia Web site) under the control of the enhanced 35S cauliflower mosaic virus, obtaining the plant transformation vector pCZera-ECFP (supplemental Fig. S1). Plasmids pCSPg-ECFP and pCSPg-ECFP-KDEL were obtained as described (24). Vector pCR8-ECFP containing the repeat region sequence fused to the ECFP sequence was obtained by replacing the Zera sequence in pCZera-ECFP by a synthetic DNA sequence coding for the R8 (PPPVHL)8 region flanked by SpeI and AscI restriction sites. Plasmid pCΔR8-ECFP containing the Zera-derived sequence complementary to pCR8-ECFP was obtained by deleting the repeat sequence from plasmid pKSG2 (33) by inverse PCR using primers pointing away from the repeat sequence. The blunt-ended amplified fragment was religated to obtain circular DNA from which the ΔR8 fragment, corresponding to Zera without the repeat sequence, was used to substitute Zera in pCZera-ECFP. pCR8CPPC-ECFP was obtained by replacing the Zera sequence in pCZera-ECFP with R8-CPPC synthetic DNA coding the N-terminal and repeat region of Zera elongated at the 3′-end by a CPPC coding sequence and flanked by 5′ SpeI and 3′ AscI restriction sites. Three internal AarI sites were also incorporated by degenerating the codon usage encoding VHL motifs of the repeat region to obtain R6-CPPC and R4-CPPC sequences in which the repeat sequence was reduced to encode seven, six, and four units respectively. These sequences were used to obtain pCR6CPPC-ECFP and pCR4CPPC-ECFP plasmids. Finally, Zera Cys-mutated constructs (pCZC7G-ECFP, pCZC9G-ECFP, pCCZ7,9G-ECFP, pCZC82,84,92G-ECFP, and pCZC64,82,84,92G-ECFP) were obtained by site-directed mutagenesis by PCR using PCZera-ECFP as the template and appropriate primers designed to mutate selected Cys residues to Gly. All constructs were verified by DNA sequencing.
Plant Material and Transient Expression System
N. benthamiana plants were agroinfiltrated using the syringe method. The binary plant vectors containing the construct of interest and the HC-Pro silencing suppressor construct (34) were introduced into Agrobacterium tumefaciens strain EHA 105 and used to agroinfiltrate N. benthamiana leaves of 4–6-week-old plants as described previously (25, 35).
Protein Extraction and Immunoblot Analysis
Total soluble proteins from transformed leaves were extracted in 100 mm Tris-HCl buffer, pH 7.5, containing 100 mm NaCl, 0.5% SDS, and 200 mm DTT for 1 h at room temperature. The resulting extracts were centrifuged at 10,000 × g for 30 min at 4 °C, and total soluble proteins were separated on 12.5% SDS-polyacrylamide gels. The proteins were detected by staining or by immunoblot using the anti-R8 antiserum (24) and anti-GFP raised in rabbits injected with a recombinant GFP protein expressed and purified from Escherichia coli cells.
Subcellular Fractionation
Agroinfiltrated tobacco leaf tissues were ground in a mortar at 0 °C in homogenization buffer (10 mm Tris, pH 8.0, 0.25 m sucrose, and protease inhibitors). The homogenate was filtered through two layers of Miracloth (22–24 μm; Calbiochem) to remove tissue debris before centrifuging at 50 × g for 5 min at 4 °C. The resulting clarified homogenates were loaded onto multistep iodixanol (Optiprep, Sigma) density-based gradients (steps: 1.11, 1.17, 1.19, 1.21, 1.23, and 1.25 g/cm3) buffered with homogenization buffer. The gradients were centrifuged at 4 °C for 2 h at 80,000 × g in a Beckman SW40 Ti rotor. Equivalent aliquots of supernatant, interphase fractions, and pellet were analyzed by SDS-PAGE and immunoblot using specific antibodies.
Immunocytochemistry and Imaging
Confocal Microscopy
Sections of leaf tobacco tissues transformed with sequences coding fluorescent proteins were mounted in water for direct confocal observation. Micrographs were obtained using a Leica TCS SP confocal laser-scanning microscope (Heidelberg, Germany). Yellow fluorescent images were collected after 515 nm excitation using a 530–630 nm emission window. Cyan fluorescent images were collected after 458 nm excitation with an emission window of 470–530 nm. Images presented in the microscopy figures are representative of at least five independent experiments.
Electron Microscopy
For immunocytochemistry studies, small sections of agroinfiltrated tobacco leaves were prepared as described previously (10, 25) using anti-R8 (1:1500) and protein A-colloidal gold (15 nm) antibodies. Sections were examined under an electron microscope (Phillips EM301, Eindhoven, The Netherlands). In all cases, non-immune serum was used as the control. For high pressure freezing and freeze substitution, small leaf pieces were frozen using high pressure freezing equipment (EMPact, Leica). The frozen samples were treated with acetone for 2 days at −80 °C and then warmed to −20 °C for 3 h, 4 °C for 2 h, and room temperature for 2 h prior to use. Fixed samples were dehydrated and embedded in Lowicryl K4M resin.
Quantitative RT-PCR
Total RNA was extracted from N. benthamiana leaves agroinfiltrated with pCZera-ECFP and collected at 0, 2, 4, 7 and 10 dpi using an RNeasy® plant minikit (Qiagen). cDNA was synthesized from 500 ng of RNA using the QuantiTect® reverse transcription kit (Qiagen) according to the manufacturer's instructions. RT-PCR assays were performed using LC480 SYBR Green I Master Mix (Roche Applied Science) according to the manufacturer's instructions, including appropriate primer pairs designed using the software Primer Express (Applied Biosystems). All samples were run in triplicate in a Roche Applied Science Light Cycler 480. Primer efficiency was calculated by the serial dilution method as described previously (36). The N. benthamiana EF1α mRNA was used as the internal control (37).
Sampling and Statistical Analysis
The distribution by size (in relative percentage) of PBs over time was determined by measuring the apparent diameters of around 500 PBs/time point (2, 4, 7, and 10 dpi). Three independent transformed plants were analyzed per time point, and the Olympus fluoview version 1.6a software was used to measure the fluorescent PBs observed under the FV1000 Olympus confocal microscope. To determine PB number, confocal projections corresponding to 105 μm3 (70 × 70 × 20) of transformed tissue were used. Forty confocal images from eight independently transformed plants were analyzed per time point. The results were analyzed statistically by one-way analysis of variance and the Bonferroni multiple comparisons test (p < 0.05 was considered significantly different).
Molecular Dynamics
A model of the Zera monomer was generated using the Biopolymer module of the InsightII molecular modeling software. The modeled sequence consisted of 93 residues (THTSGGCGCQP PPPVHL PPPVHL PPPVHL PPPVHL PPPVHL PPPVHL PPPVHV PPPVHL PPPPCHYPTQP PRPQPHPQPHP CPCQQPHPSPCQ), adopting a polyproline II (PPII) conformation with −75 and 145° for the φ and ψ dihedral angles, respectively. A model with two Zera molecules (Zera dimer) was also constructed by aligning two Zera molecules in parallel. The two Zera molecules were linked by a disulfide bond established between the two Cys9 residues. This cysteine link allows contact between the hydrophobic surfaces of both R8 repeats, whereas the hydrophilic surfaces of these repeats are surrounded by water molecules. The standard protonation state of ionizable residues at pH 7.0 was adopted. However, for histidine residues, both the positively charged and the neutral state were used in different MD calculations. MD simulations of a Zera monomer and Zera dimer in water, neutralized with chloride ions, were performed with NAMD 2.6 (38), using the CHARMM22 (39) force field and a 14 Å cut-off with a 2.0-A switch distance. The Zera monomer was placed in a box of 300 × 35 × 35 Å3, and the Zera dimer was placed in a box of 302 × 48 × 48 Å3. All bond distances were fixed by the SHAKE (40) algorithm. The simulations were performed using periodic boundary conditions, at constant temperature (25 °C) and pressure (1 atm) using the Langevin piston Nose-Hoover method (41, 42). A time step integration of 2 fs for MD equations was used. Equilibration was performed by restraining the coordinates of the backbone atoms by a harmonic potential with a force constant value of 1 kcal/mol for 250 ps at constant volume and for another 250 ps at constant pressure. Then three unrestrained MD simulations of 4 ns for the Zera monomer model and 10 ns for the Zera dimer model were performed using the charged state of all histidines in all cases. In addition, MD calculations of 3 ns for the neutral His form of both monomer and dimer were performed in order to analyze the effect of the histidine protonation state on the conformational properties. No significant differences were observed between the charged and neutral histidines. Here we only present the results of MD calculations using the charged state. Molecular graphics images were produced using the VMD 1.8.6 package (43).
RESULTS
Role of Zera Regions in PB Induction
Zera is the N-terminal proline-rich domain of γ-zein, a maize storage protein (23). The 112 amino acids of the Zera sequence include the signal peptide and the first 93 amino acids of mature γ-zein. Zera contains three proline-rich regions (supplemental Fig. S1): 10 non-proline amino acids containing a Cys-Gly-Cys motif preceded by the γ-zein signal peptide; a proline-rich repeat region containing eight units of the hexapeptide PPPVHL; and a proline-X sequence containing four cysteines. In the present work, Zera and Zera mutants were fused to the N terminus of a cyan fluorescent marker protein, ECFP. The fusions included a linker of five glycines inserted between Zera and ECFP (supplemental Fig. S1).
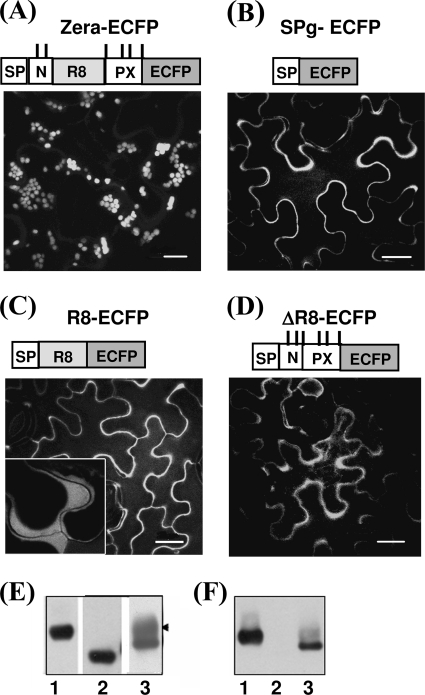
To explore the intrinsic features of Zera that are required for PB formation, we first attempted to establish whether PB could be induced in the absence of the Zera repeat region ((PPPVHL)8 (R8)) or in the absence of the N- and C-terminal sequences flanking it (Fig. 1). Two complementary deletions of the Zera sequence, termed R8 and ΔR8, were created and fused to a reporter fluorescent protein, ECFP. The R8-ECFP protein (Fig. 1C) contained only the eight units of (PPPVHL)8 directly fused to ECFP, whereas ΔR8-ECFP (Fig. 1D) lacked the entire R8 sequence but contained the flanking N-terminal (N) and C-terminal (PX) sequences of Zera. Two additional constructs were used: Zera-ECFP (Fig. 1A) as a positive control of PB induction and SP-ECFP lacking Zera as a control of ECFP secretion (25) (Fig. 1B). All constructs were agroinfiltrated into leaves of N. benthamiana plants. Fig. 1 (A–D) shows the fluorescence pattern displayed by proteins expressed in epidermal cells of tobacco leaves at 4 dpi. Cells expressing R8-ECFP and ΔR8-ECFP (Fig. 1, C and D) displayed a pattern of secretion, with fluorescence mainly located in the periphery of the cells, as occurred with the secreted SP-ECFP (Fig. 1B). Secretion of R8-ECFP and ΔR8-ECFP proteins was confirmed by plasmolysis of the epidermal cells. Plasmolysed cells showed fluorescence located between the membrane and the cell wall (Fig. 1C, inset), in both cases indicating that these fusions were competent for transport to the apoplast. In contrast, Zera-ECFP accumulated in PB-like organelles, visualized as spherical fluorescent spots of various sizes (Fig. 1A).
FIGURE 1.
Both the repeat region and its flanking sequences are necessary for PB biogenesis. A–D, confocal images of tobacco leaf epidermal cells at 4 dpi using constructs Zera-ECFP (A), SP-ECFP (B), R8-ECFP (C), and ΔR8-ECFP (D). Only the wild type Zera (A) sequence induced PBs in transformed cells. SP-ECFP (B) was used as a control of protein secretion. The truncated Zera protein fusions (C and D) were secreted. The inset in C indicates fluorescence in the apoplast after cell plasmolysis. A schematic diagram of each of the expressed constructs is shown above the corresponding confocal image. SP, γ-zein signal peptide; N-, first 10 amino acids of Zera sequence; PX, Pro-Xaa sequence. ECFP was used as reporter. The upper vertical lines in the diagrams represent Cys residues. Bar, 10 μm (A) or 20 μm (B–D). E and F, immunoblots of total protein extracts from tobacco leaves transformed with Zera-ECFP (lanes 1), ΔR8-ECFP (lanes 2), and R8-ECFP (lanes 3), using anti-GFP (E) and anti-R8 (F) antibodies.
To determine whether the absence of polymerization of the R8-ECFP and ΔR8-ECFP proteins was the consequence of low levels of protein expression, equal amounts of total protein extracted from agroinfiltrated tobacco leaves were analyzed by Western blot using both anti-GFP (Fig. 1E) and anti-R8 (Fig. 1F) antibodies. As shown in Fig. 1E, both the R8-ECFP (lane 3) and ΔR8-ECFP (lane 2) proteins accumulated to levels comparable with those of Zera-ECFP (lane 1), indicating that the expression levels did not justify the secretion of R8-ECFP and ΔR8-ECFP. An additional immunolabeled band was detected in R8-ECFP leaf extracts when using anti-GFP (Fig. 1E, arrowhead) but not with anti-R8, suggesting that the repeat region of R8-ECFP could be modified post-translationally during transit to the apoplast, masking the epitopes recognized by the anti-R8 antibody. Taken together, these results indicate that important roles are played by two Zera regions: the R8 repeat and the flanking amino acid residues.
Zera Cysteine Residues Play Different Roles in PB Biogenesis
The above data prompted us to study whether the importance of sequences flanking the R8 repeat is attributable to the presence of Cys residues in the N-terminal and Pro-X region. Two cysteine residues, Cys7 and Cys9, are present in the N-terminal region preceding the repeat sequence, whereas Cys64, Cys82, Cys84, and Cys92 are present in the C-terminal PX sequence (see scheme in Fig. 2A). Simultaneous mutation of all six cysteines to glycines in the Zera-ECFP sequence resulted in the secretion of the recombinant protein when expressed in N. benthamiana epidermal cells (not shown). Similar results were previously described for zeolin, a fusion of the bean phaseoline with the N-terminal γ-zein sequence (4).
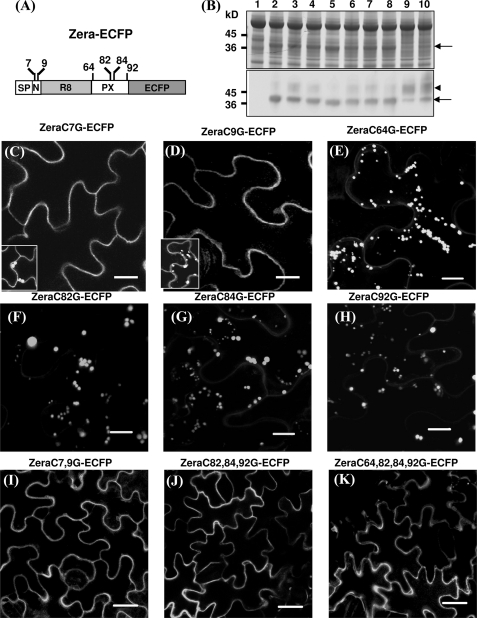
FIGURE 2.
Role of Cys residues in Zera-PB formation. A, diagram showing the positions of the cysteine residues in Zera fused to ECFP. B, total protein analysis of tobacco leaves transformed with Zera-Cys mutants fused to ECFP by SDS-PAGE/Coomassie Blue staining (top) and immunoblot using an anti-GFP antibody (bottom). Shown are untransformed tobacco protein extracts (lane 1), tobacco extracts of non-mutated Zera-ECFP (lane 2), and protein extracts of transformed tobacco with Zera Cys7 mutant (lane 3), Zera Cys9 (lane 4), Zera Cys64 (lane 5), Zera Cys82 (lane 6), Zera Cys84 (lane 7), Zera Cys92 (lane 8), Zera Cys7-Cys9 (lane 9), and Zera Cys82-Cys84-Cys92 (lane 10). The arrows indicate the electrophoretic bands of Zera-ECFP and Zera-ECFP Cys mutants, and the arrowhead shows an additional immunoreactive band. C–K, confocal images showing the fluorescence pattern of epidermal cells transformed with Zera-Cys mutants. Single Cys7 and Cys9 mutants were mainly secreted (C and D), but some cells were able to induce PBs (insets in C and D). E–H, single mutation of Cys64, Cys82, Cys84, or Cys92 had no effect on PB formation. I–K, Zera-ECFP containing multiple Cys mutations was not able to oligomerize and was secreted. Scale bars, 10 μm (C–H) or 20 μm (I–K).
The next question was whether each Cys residue contributes equally to Zera-ECFP polymerization. We analyzed the effect of perturbing the formation of disulfide bridges by generating point mutations of cysteine in the Zera sequence (Fig. 2) that were expressed in tobacco leaves and subsequently analyzed by SDS-PAGE and immunoblot (Fig. 2B) and confocal microscopy (Fig. 2, C–K). Fusion proteins carrying a single cysteine mutation in Zera (Fig. 2B, lanes 3–8) showed accumulation levels of protein similar to native Zera fusion proteins (lane 2). Similar fluorescent patterns were observed by confocal microscopy in cells expressing ZeraC7G-ECFP and ZeraC9G-ECFP. A consistent observation was that both proteins were mainly secreted, indicating that these mutations strongly impaired the capacity for multimerization (Fig. 2, C and D). However, as shown in Fig. 2, C and D (insets), we also observed that highly fluorescent cells (around 30% of the transformed cells) induced PBs that were similar in shape and size to those observed in Zera-ECFP cells (Fig. 1A). It appears that individual cells with very high rates of recombinant protein biosynthesis are able to proceed with protein oligomerization. We also engineered point mutations in the last four cysteines of the Zera sequence (ZeraC64G-ECFP, ZeraC82G-ECFP, ZeraC84G-ECFP, and ZeraC92G-ECFP). Interestingly, in contrast to Cys7 and Cys9, single substitutions of the C-terminal cysteine residues did not appear to interfere with multimerization significantly because most epidermal cells displayed a regular PB pattern (Fig. 2, E–H). We then examined the effect when both N-terminal Cys7 and Cys9 were mutated (ZeraC7,9G-ECFP). As shown in Fig. 2I, the double mutation dramatically interfered with the ability to oligomerize, and fusion protein was always secreted. This result indicates that the combination of Cys7 and Cys9 is essential for Zera-ECFP oligomerization and suggests that both cysteines could act in a cooperative way. We subsequently investigated the effect of multiple mutations of Cys residues located downstream of the repeat region. Two more constructs were generated, ZeraC82,84,92G-ECFP and ZeraC64,82,84,92G-ECFP (Fig. 2, J and K). As shown, PB formation was strongly impaired in the triple and quadruple mutants, and the fusion proteins were secreted. However, a few sporadic cells of the Zera triple Cys mutant displayed numerous spherical fluorescent foci (data not shown). In those Zera Cys mutants that were mainly secreted, the immunoblot showed a clear high molecular weight diffuse band recognized by the anti-GFP antibody (Fig. 2B, arrowhead), as previously observed in Fig. 1E (lane 3) for secreted truncated Zera fusions. These findings indicate that the relevance of single cysteine mutations in terms of protein oligomerization depends on their location in the Zera sequence, with Cys7 and Cys9 being necessary but not sufficient for determining the fate of polymerization-prone Zera-ECFP chains.
Minimal Sequence Regions Required for Zera Multimerization
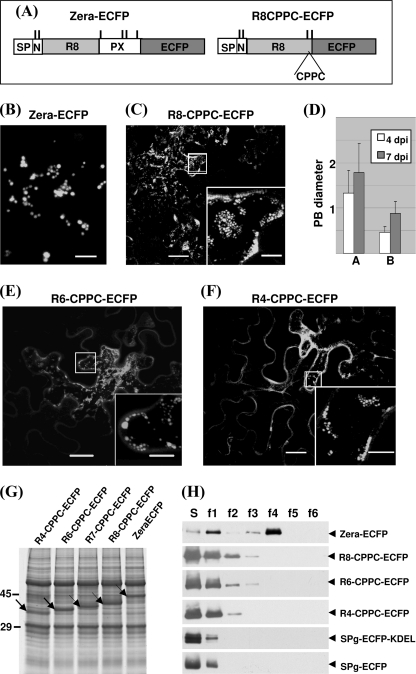
From the above results, we aimed to determine the minimal requirements for the Zera sequence to induce PBs. Thus, we generated a minimal Zera-ECFP-derived protein, R8-CPPC-ECFP, by deleting the PX sequence, which thus lacked Cys82, Cys84, and Cys92 but contained an additional new cysteine residue linked to Cys64 by two prolines to reinforce the interchain disulfide bridges (Fig. 3A). The distribution of the R8-CPPC-ECFP protein when expressed in N. benthamiana leaves, instead of being secreted as R8-ECFP (Fig. 1C) or accumulated in spherical fluorescent spots (Fig. 3B), displayed a punctuate fluorescent pattern (Fig. 3C), which at high magnification appeared to consist of small protein polymers (Fig. 3C, inset). Although PBs induced by Zera-ECFP reached an average diameter of 1.4 μm at 4 dpi and increased progressively to up to 2 μm at 7 dpi, the fluorescent spots induced by R8-CPPC-ECFP rarely reached diameters of 1 μm at 7 dpi (Fig. 3D). Therefore, the cysteines on each side of the repeat region in R8-CPPC-ECFP and the repeat sequence itself are probably sufficient to nucleate protein oligomerization, but they remain smaller than Zera-ECFP-induced PBs. These results suggest that the strong tendency of Zera-ECFP to form large polymers (PBs) compared with that of R8-CPPC-ECFP is related to the fact that Zera-ECFP oligomers have more opportunity to grow by increasing the interchain cross-linking by virtue of their six cysteine residues.
FIGURE 3.
Minimal Zera sequence required for PB induction. A, diagram of Zera-ECFP and R8-CPPC-ECFP constructs. B and C, confocal images of Zera-ECFP and R8-CPPC-ECFP accumulation patterns in 4 dpi transformed leaves. Small fluorescent spots were visualized in R8-CPPC-ECFP (C, inset). D, diameters (average ± S.D.) of Zera-ECFP PBs (histograms A) and R8-CPPC-ECFP PBs (histograms B) at 4 and 7 dpi. E and F, representative confocal images obtained when the eight units of the repeat sequence were shortened to six (E) and four (F) repeat units. In the insets, high magnifications of the indicated areas show small aggregates. G, SDS-PAGE/Coomassie Blue staining of total proteins extracted from leaf tissues transformed with the indicated Zera-truncated constructs fused to ECFP. The accumulated recombinant proteins are indicated by arrows. H, subcellular fractionation of homogenates from tobacco leaves transformed with the constructs indicated at the right. Supernatants (S) and density-increasing interface fractions (f1–f6) from gradients were analyzed by immunoblot using anti-GFP antibodies. SP-ECFP and SP-ECFP-KDEL were used, respectively, as control of secreted and ER-retained proteins. Scale bars, 10 μm (B), 20 μm (C–F), or 5 μm (insets).
The (PPPVHL)8 sequence has previously been shown to self-interact in vitro (28), and it seems to be a key piece in the Zera oligomerization process because, when deleted, the resulting fusion protein fails to form PBs (Fig. 1D). Thus, we investigated whether polymerization was related to the length of the repeat region. From R8-CPPC-ECFP, we generated two more constructs in which the repeat sequence was shortened to seven, six, and four PPPVHL units. As shown in Fig. 3 (E and F), overexpressed fusion proteins containing six or four repeat units (R6-CPPC-ECFP and R4-CPPC-ECFP, respectively) were still able to form small oligomers, but secretion increased as the repeat sequence was shortened (Fig. 3, E and F, inset). SDS-PAGE and Coomassie Blue stain analysis of total proteins extracted from leaves overexpressing truncated Zera-ECFP (Fig. 3G) showed that the concentration of recombinant proteins was similar in all cases, indicating that they were stable and that the lower polymerization efficiency in mutants was not related to their protein expression levels. These results suggest that there is a critical number of repeats that favors optimal Zera self-interactions, determining the efficiency of PB formation.
In a previous study, we showed by subcellular fractionation in density gradients that PBs containing Zera fusions sediment at high densities (24, 25). To determine how dense the protein polymers or the PB-like structures are when induced by truncated Zera-ECFP proteins, we analyzed homogenates of Zera-ECFP-, R8-CPPC-ECFP-, R6-CPPC-ECFP-, and R4-CPPC-ECFP-expressing leaves using iodixanol density gradients (Fig. 3H). Soluble secreted (SP-ECFP) and ER-resident (SP-ECFP-KDEL) proteins were used as controls. As shown in Fig. 3H and consistent with the confocal images, truncated Zera-ECFP proteins were mostly recovered in the soluble fractions (S), like the control soluble proteins. However, some fusions with truncated Zera were able to oligomerize because traces of the proteins were recovered in the density ranges of 1.17–1.19 g/ml (f2 interface) and 1.19–1.21 g/ml (f3 interface).
From Zera Oligomers to PB Formation
Next we investigated the kinetics of PB biogenesis from the early stages of protein oligomerization to mature Zera-ECFP PBs. First we analyzed Zera-ECFP RNA stability and protein accumulation at 2, 4, 7, and 10 dpi by quantitative RT-PCR. As shown in Fig. 4, both the RNA and protein were stably accumulated until 7 dpi. Due to the transient expression system, the RNA had already been degraded at 10 dpi (Fig. 4A), but the protein was still stably accumulated (Fig. 4B). Therefore, we assume that during the period of overexpression, the recombinant protein did not enter the ERAD pathway or the secretory pathway but was stored as large polymers in the ER (24).
FIGURE 4.
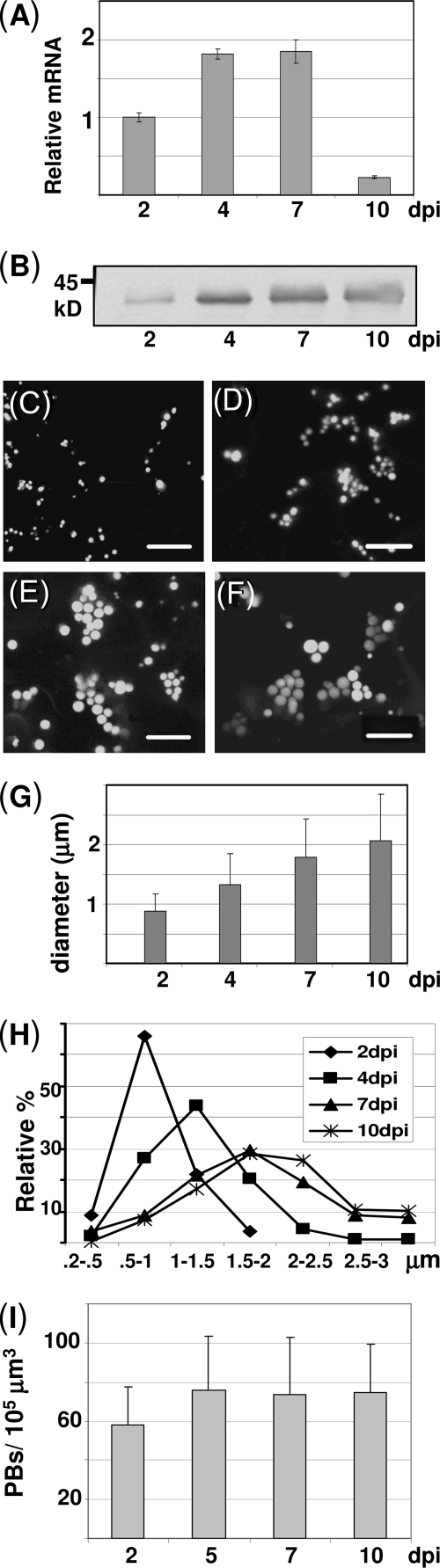
Zera-ECFP PBs growth. A, Zera-ECFP RNA analysis at 2, 4, 7, and 10 dpi. The relative Zera-ECFP mRNA content was determined by quantitative RT-PCR using EF1α as a housekeeping reference. B, Zera-ECFP protein accumulation over time after tobacco transformation. Total protein extracts of transformed leaves were analyzed by immunoblot using anti-R8 antibody. C–F, confocal images showing Zera-ECFP-induced PBs at 2 (C), 4 (D), 7 (E), and 10 (F) dpi. Scale bars, 10 μm. G, PB apparent diameter size measurements (average ± S.D.) at various dpi showing the PB growth. H, PB size distribution profiles from 2 to 10 dpi. I, quantification of PB number per leaf volume unit as a function of dpi. Error bars, S.D.
The appearance and development of induced PBs were examined over time by confocal microscopy in cells expressing Zera-ECFP at 2, 4, 7, and 10 dpi (Fig. 4, C–F). As shown in Fig. 4C, numerous small punctuate structures were detected as early as 2 dpi. From 4 to 10 dpi (Fig. 4, D–F), abundant round fluorescent spots were observed adjacent to the plasma membrane and surrounding the large central vacuole of epidermal cells. PBs increased progressively in size and were frequently seen forming clusters. Zera-ECFP PB growth was quantified by measuring the apparent diameter of about 500 PBs/time point in three independently transformed leaves. Despite the variability in PB size, there was a progressive increase over time (Fig. 4G) that was reflected by a clear shift in the size distribution profiles of PB populations (Fig. 4H). At 2 dpi, the most abundant PB diameters ranged from 0.5 to 1 μm, whereas at 7 dpi, around 70% of the PBs were larger than 1.5 μm.
We also quantified the number of these structures (Fig. 4I) per leaf arbitrary volume unit over time. The quantitative data were statistically analyzed by analysis of variance and Bonferroni test (p < 0.05). Accordingly, the number of PBs at 2 dpi was significantly lower compared with that at 5, 7, and 10 dpi, which apparently remained stable over time. Whereas at early stages of Zera-ECFP expression (2–5 dpi), recombinant protein polymers increased in number and size, at later stages (5–10 dpi), they increased in size but not in number. Apparently, Zera-ECFP synthesized after 5 dpi mainly contribute to PB growth rather than being involved in forming new PBs. The increase in size is the result of feeding of preformed PBs by newly biosynthesized Zera-ECFP polypeptide chains from active ER ribosomes.
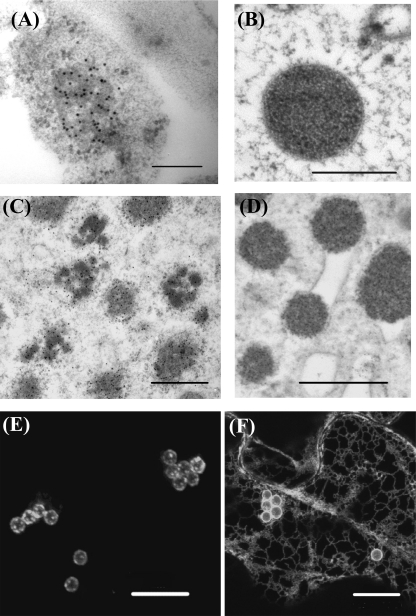
We then used immunoelectron and transmission electron microscopy to assess the presence of small Zera-ECFP oligomers within the ER at the early stages of PB biogenesis. Fig. 5 shows thin sections of epidermal cells expressing either Zera-ECFP (A and B) or R8-CPPC-ECFP (C and D) at 2 dpi (A and C) and 7 dpi (B and D). At 2 dpi, epidermal cells expressing Zera-ECFP contained small non-homogeneous electron-dense accretions labeled with anti-R8 antibody and surrounded by membrane-bound ribosomes. The immunolabel in these structures suggests that small neighboring Zera-ECFP oligomers inside the ER could cross-link to form larger denser structures. We took advantage of the weak capacity of R8-CPPC-ECFP to form large polymers (see Fig. 3C) to better visualize small protein polymers created at early stages of R8-CPPC-ECFP expression. Immunolabeling of R8-CPPC-ECFP cells revealed numerous clusters of small protein assemblies (Fig. 5C), not completely structured as dense organelles but closely connected inside the ER. We also performed morphological studies by EM on Zera-ECFP (Fig. 5B) and R8-CPPC-ECFP PBs (Fig. 5D) using sections of transformed leaves subjected to high pressure freezing/freeze substitution at 7 dpi. We found round aggregates ranging in size from 1.2 to 2.4 μm for Zera-ECFP and from 0.6 to 1.1 μm for R8-CPPC-ECFP, which were in both cases localized in a dilated rough ER and frequently surrounded by the ER membrane (see Fig. 5D). Apparently, the clusters of small aggregates observed at 2 dpi could promote swelling in regions of the rough ER, which, after growing, remained connected in the lumen of the ER instead of forming independent organelles. This connection was verified by the expression of the ER-soluble protein YFP-KDEL in epidermal cells transformed with only the Zera sequence (Zera-stop), in which YFP-KDEL fluorescence was observed surrounding Zera-induced PBs. Fluorescence recovery after photobleaching experiments showed that YFP-KDEL diffused through the ER, rapidly reaching the periphery of bleached PBs, indicating the connection of mature PBs with the ER (supplemental Fig. S2).
FIGURE 5.
Zera-ECFP PB formation and growing takes place within the ER. A–D, electron microscopy images showing oligomers and PBs of Zera-ECFP (A and B) and of R8-CPPC-ECFP (C and D) in transformed tobacco leaf cells at 2 dpi (A and C) and at 7 dpi (B and D). Samples from 2 dpi were immunolabeled using anti-GFP antibody and protein A coupled to gold particles (10 nm). 7 dpi samples were subjected to high pressure freezing techniques to preserve morphology. E and F, confocal images at 7 dpi after sequential transformation of N. benthamiana leaves with, first, Zera-stop and, at 4 dpi of Zera-stop expression, with Zera-GFP (E) or SPg-YFP-KDEL (F).
We next examined how preformed oligomers of Zera-ECFP grow to yield large polymers like PBs. To this end, we sequentially expressed a construct containing Zera-stop, which forms unlabeled PBs, followed by a construct corresponding to Zera-GFP, which was agroinfiltrated 4 days later. As a control, the same tissue was agroinfiltrated with YFP-KDEL construct after 4 days of Zera-stop expression. Fluorescent cells were observed by confocal microscopy at 7 days after the first transformation event (Fig. 5, E and F). When Zera-GFP was infiltrated after Zera-stop, we detected a green label specifically incorporated within the unlabeled Zera-stop PBs. Interestingly, the majority of green label was condensed in punctuate spots surrounding the Zera-stop PBs (Fig. 5E). Oligomers of Zera-GFP and Zera-stop polymerized in the preformed Zera-stop PBs. In contrast, when YFP-KDEL was infiltrated after 4 days of Zera-stop expression, cells displayed a clear ER pattern, and unlabeled PBs were surrounded by homogeneous fluorescent label (Fig. 5F). Thus, whereas new Zera-GFP and Zera-stop polypeptide chains were able to efficiently self-assemble and polymerize to give rise to fluorescent foci, YFP-KDEL was incorporated as a soluble protein and diffused through the ER. These data suggest that PB growth either occurs by diffusion of undetectable oligomers into preformed PBs or by the activity of ribosomes present in the ER membranes surrounding Zera-stop PBs that would feed the pre-formed organelles with newly synthesized Zera-GFP and Zera-stop polypeptide chains.
Toward a Model of Zera-PB Biogenesis by Molecular Dynamics Simulation
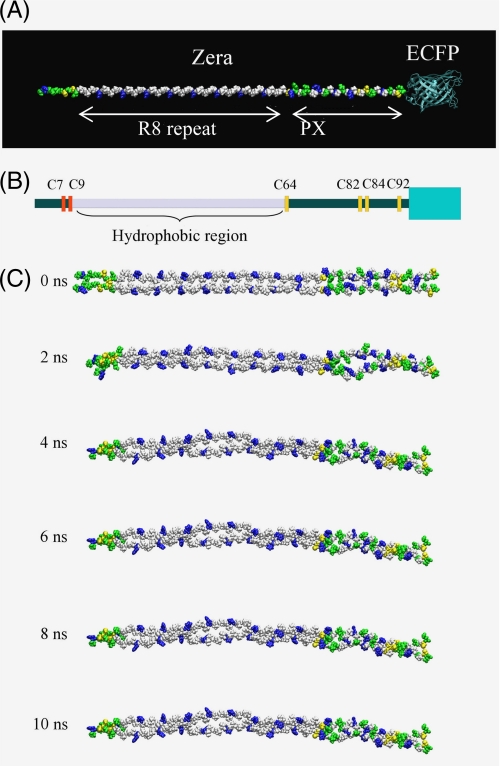
Our goal in this study was to investigate the driving forces involved in the organization of Zera into oligomers and to provide a rational understanding of the chemical details of Zera self-assembly. Molecular dynamics (MD) simulations offer the opportunity to overcome the problems derived from working with insoluble polymers using conventional biochemical techniques. Thus, we performed an MD study of Zera molecule lacking the signal peptide to analyze the stability of putative Zera monomer and dimer structures. Intermolecular Zera interactions were studied using a model of a Zera dimer constructed using an interchain disulfide bond between the two Cys9 residues of adjacent Zera molecules (Fig. 6C). Based on previous work describing the presence of a PPII helix in the repeat (PPPVHL)8 region and the tendency of the PX sequence to adopt the PPII conformation (44–46), we used the PPII helix as the elementary conformation of Zera (Fig. 6A). Because the oligomerization of Zera occurs inside the ER lumen, water was used as the solvent in all simulations. Analysis of the MD calculations of the monomer indicated that Zera containing both the repeat and the PX regions conserved the PPII conformation. Moreover, no bending sites were observed during the whole trajectory of MD simulations. Only the first residues of the N terminus of Zera (from Thr1 to Gly8) and the last two residues of the C terminus appeared to be flexible and did not preserve the PPII helix conformation (supplemental Fig. S3). This conformational rigidity of the Zera backbone could facilitate the formation of oligomers. In addition, we have shown that side chains, such as those of cysteines, enormously influence PB formation (Fig. 2). It has previously been postulated (28) that histidines in the repeat sequence are all oriented on one side of the helix and that their protonation at neutral pH creates an amphipathic PPII helix that could contribute to the orientation of the oligomer. As shown in the Zera dimer simulation (Fig. 6C and supplemental Fig. S4), no contact between the polar side chains of His (in blue) in (PPPVHL)8 was observed. In contrast, the number of contacts among hydrophobic leucine and valine residues was conserved during the simulation. In the PX region, there were no fully hydrophobic or polar contacts or solvent-accessible surfaces. Polar amino acids in the PX region were oriented in the three possible directions of the helix, making this proline-rich region of Zera non-amphipathic. The hydrophobic profile analysis of Zera (supplemental Fig. S5) demonstrated that the repeat region (PPPVHL)8 is more hydrophobic than the PX region, indicating that both hydrophobic interactions of the amphipathic repeat and side chain interactions of the N and PX regions facilitate the lateral interchain assembly of Zera molecules. In addition, analysis of the evolution of distances between cysteine pairs along the Zera dimer MD (data not shown) allowed us to estimate the tendency of each Zera cysteine residue to form interchain disulfide bonds. Internal cysteine residues (see Fig. 6B, Cys64, Cys82, and Cys84) have a high probability of linking the two parallel aligned Zera molecules. In contrast, terminal Cys7 and Cys92 do not participate in the covalent stabilization of the dimer but are free to participate in linking other Zera molecules.
FIGURE 6.
Molecular dynamics calculations. A, initial disposition of Zera molecule in PPII conformation, indicating the R8 and PX regions. ECFP was also shown. B, schematic representation of a Zera monomer, taking into account the disposition of cysteine residues and the relative size of Zera and ECFP (green box). C, snapshots taken from the 10-ns MD simulation of the Zera dimer model. Hydrophobic residues are shown in white, polar residues in green, positively charged residues in blue, and cysteine residues in yellow.
DISCUSSION
Previous work demonstrated that the proline-rich N-terminal domain of γ-zein (Zera) fused to a protein induced PB formation in plant vegetative tissues (24, 25, 47).
The results presented here provide key insights into specific traits of Zera that when fused to a marker protein lead to multimerization of fusion protein and PB formation. The most important trait is the presence of cysteine residues in Zera, which participate in the interdisulfide bonds between Zera sequences. Previous evidence of a role for disulfide bonds in Zera polymerization came from zeolin expression studies in tobacco (4). Zeolin, a fusion of phaseolin and a γ-zein sequence equivalent to Zera, was insoluble and formed PBs in the ER when expressed in tobacco (26). When protoplasts expressing zeolin were treated with 2-ME or when a zeolin devoid of the six Cys residues was expressed in the protoplasts, the solubility of the protein was enhanced, allowing its trafficking along the secretory pathway (4). Similarly, a fluorescent protein fused to rice prolamine accumulated in the ER of leaf cells by forming polymers that required reducing agents for complete solubilization (48). The formation of disulfide-linked oligomers or large aggregates does not exclusively occur in plants. Diabetes insipidus disease is caused by dominant mutations in provasopressin, which, instead of being secreted, is retained in the ER, forming disulfide-linked homo-oligomers and large polymers (49). However, it should be noted that in this case, assembly results from a pathological situation caused by protein misfolding.
Here we show the effect of point mutations on each of the six cysteines of Zera as a way of elucidating Zera self-polymerization by disulfide bonds. The overexpression of Zera-ECFP devoid of Cys7 or Cys9 clearly disturbed the multimerization of the fusion protein because both mutants were secreted at high levels. In contrast, individual mutations of the other four cysteine residues did not have any significant effect on the oligomerization of fusion proteins, and the proteins were stored in PBs. It is worth noting that a huge amount of Zera-ECFP enters the ER in the transient expression system used and that the ER needs to accommodate the redox homeostasis by maintaining sufficient oxidizing conditions within the lumen. This is achieved by interplay between the enzyme protein-disulfide isomerase (50, 51), oxidoreductases, such as the flavoprotein Ero1 recently described in rice (52), and glutathione as the redox buffer.
γ-Zein, the origin of the Zera sequence, self-assembles and aggregates with α-zeins in its natural maize seed expression system to form PBs, a process that is probably assisted by BiP and protein-disulfide isomerase (53). One explanation for the biased effect of individual Cys7 and Cys9 mutations compared with the other Zera Cys mutants could be that these Cys7 and Cys9 mutations disrupt the first oxidation steps of the newly synthesized fusion proteins inside the lumen of the ER. Therefore, individual mutations of N-terminal cysteine residues result in deficient cross-linking between Zera chains, which, in turn, would impair their condensation, thus favoring secretion. The normal oligomerization capacities of Zera Cys64 to Zera Cys92 single mutants support the hypothesis that the two N-terminal cysteine residues are crucial for nucleating the oligomerization process of fusion proteins. However, since multiple C-terminal Cys mutants did not proceed with PB biogenesis, it is clear that Zera requires more than the early formed Cys7- and Cys9-linked homo-oligomers. The former oligomers of Zera might grow further to form polymers by continuous interdisulfide bond formation involving free C-terminal SH groups of new Zera chains.
Protein concentration is a key parameter controlling the oligomerization of self-assembling peptides (54). The expression levels of Zera-ECFP and Zera Cys mutants-ECFP were clearly visualized in SDS-PAGE Coomassie Blue-stained gels, and no significant differences in protein levels were observed between those proteins that formed PBs and those that were not able to polymerize and were secreted. This suggests that differences in protein multimerization of fusions containing different Zera mutants are related to differences in their intrinsic properties that affect their assembly capacity. Modification of the Zera sequence would either impair this polymerization capacity or raise the protein concentration threshold required for oligomerization. In fact, in cell-by-cell image analysis of epidermal cells transformed with individual Cys7 or Cys9 mutants, fusion proteins were generally secreted, but a few single, highly fluorescent cells displayed fluorescent PB-like spots. This observation suggests that high rates of expression of the recombinant protein in the ER result in an increase in the polymerization efficiency, indicating that a higher critical protein concentration is required for protein assembly compared with protein fusions containing wild type Zera. We cannot rule out the possibility, however, that in addition to protein concentration and intrinsic Zera properties, extrinsic factors, such as chaperone concentration, might also play a role in PB formation. It is likely that PB formation is the result of a three-sided balance between (i) the aggregation capacity of the Zera sequence, as an intrinsic property of the system; (ii) the recombinant fusion protein concentration; and (iii) the activity of the chaperones, as an extrinsic property.
The development of Zera-ECFP-induced PBs over time indicated that after an initial increase in PB number, the population of PBs remained constant in number but increased in size. This suggests two steps in PB formation: (i) early synthesized Zera-ECFP polypeptide chains promote nucleation of Zera self-assembly and constitute the origin of PBs, and (ii) continued synthesis of Zera-ECFP at later stages contributes to PB growth by incorporation of Zera fusions in the above preformed PBs. Therefore, as mentioned earlier, PB formation could be driven by a nucleation-dependent Zera self-assembling mechanism that is expected to depend on protein concentration. Recently, Foresti et al. (55) reported that a sequence similar to Zera, named “zein” by the authors, when fused to GFP (zein-GFP) and expressed in tobacco transgenic lines, was not assembled into stable PBs. The fusion underwent post-translational fragmentation of the GFP “core,” probably through its delivery to the vacuole. Although we cannot rule out the possibility that a small proportion of the fusions containing Zera or Zera Cys mutants were degraded by the ERAD pathway (22), by delivery to vacuoles in a Golgi-mediated pathway, or by autophagy (12, 56), the bulk of the recombinant Zera proteins was stably accumulated over 10 days, and protein levels remained high even when RNA levels decreased. The results of Foresti et al. (55) might be explained by low levels of expression of zein-GFP in their transgenic lines, thus not reaching the threshold required for protein oligomerization.
It has been suggested that the repetitive proline-rich region of Zera is responsible for the “in vivo” oligomerization of γ-zein at the surface of the PB (28). Analysis of the secondary structure of the entire γ-zein protein by CD spectra in aqueous media showed that the whole protein has a high helical content but a low PPII conformation content, at 55 and 7%, respectively (57). In contrast, the synthetic peptide (PPPVHL)8, corresponding to the repeat region, has been shown to adopt an amphipathic polyproline II conformation (27, 44) and has been linked to a new family of cell-penetrating peptides (58). We demonstrate here that the repeat region of Zera is required for Zera-ECFP assembly, suggesting that the amphipathic character of Zera facilitates proper molecular orientation in the aqueous environment of the ER to optimize interchain disulfide bond cross-linking. The eight units of PPPVHL in Zera seem to provide the optimum length for self-assembly because six and four units result in a significant decrease in polymerization capacity. Shorter repeat sequences lead to smaller and less dense polymers. Although it is known that the synthetic peptide (PPPVHL)8 has the capacity to interact with liposomes (59), we did not detect Zera-ECFP interactions with the ER membranes using high pressure EM observations. Thus, if Zera sequence interacts with the membrane, it must be a transient phenomenon that could occur at early stages of the multimerization process.
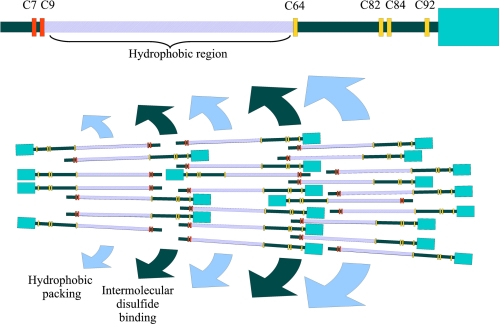
Based on our results, we propose a molecular model to explain the initial process of Zera oligomer stabilization and subsequent growth of these oligomers into large polymers, namely PB formation (Fig. 7). The model was constructed by considering different factors: (i) the fact that the Zera molecule appears to have a sticklike conformation (Zera MD simulations rule out bending that would distort the PPII helix), (ii) the essential role of the N-terminal and C-terminal cysteine residues for interchain cross-linking by disulfide bond formation, and (iii) the amphipathicity of the (PPPVHL)8 region of Zera, which facilitates the lateral protein-protein interaction and consequently the alignment of Zera molecules. Given that a synthetic Cys-free (PPPVHL)8 peptide has been shown to self-assemble in a water solution (45), in the absence of Cys residues, it is likely that, in the initial steps, dimers and higher order oligomers are produced through interchain hydrophobic interactions between the (PPPVHL)8 region of two or more Zera chains. These oligomers could be stabilized and strengthened by interchain disulfide bonds involving Cys7 or Cys9, resulting in polymer nucleation. Hydrophobic interactions and disulfide bond formation are the two main driving forces for self-assembly in this model. Hydrophobic interactions are facilitated by high ionic strength (60), and disulfide bond formation is facilitated by an oxidizing environment. Thus, the high ionic strength and oxidizing environment in the ER lumen (11, 61) could also be a favorable factor for PB self-assembly.
FIGURE 7.
Schematic representation of the proposed molecular model of PB biogenesis and development. The hydrophobic region (violet), the N-terminal Cys (red), and the C-terminal Cys (orange) of one Zera molecule are indicated at the top. The hydrophobic region of Zera molecules facilitates the lateral protein-protein interaction and consequently the alignment of Zera molecules (hydrophobic packing). Parallel arrangement between Zera molecules is also stabilized with intermolecular disulfide binding between red Cys and red Cys and between orange Cys and orange Cys. To attain PB growing, crossed intermolecular disulfide binding (red with orange disulfide bridges) is also required. Thus, PB development takes place by the formation of alternate layers stabilized, respectively, by hydrophobic packing and intermolecular disulfide binding.
The results obtained with Zera cysteine mutants fused to ECFP support this model. Fusion proteins containing the double Cys7-Cys9 mutant of Zera (Fig. 2I) were always secreted, but when only one of these Cys residues was mutated (Fig. 2, C and D), the oligomerization process of fusion protein also took place because the other Cys residue was still able to form a disulfide bond with another Zera partner chain. Efficient growth of the polymer would be achieved by the sequential addition of monomers or oligomers that benefit from the remaining unoxidized sulfhydryl groups to establish more disulfide bonds. A deterministic model of Cys-Cys linkages in a parallel orientation would not produce spherical shaped polymers. We suggest that in addition to the parallel orientation, head-to-tail interactions also take place (e.g. between one C-terminal cysteine of an incoming Zera chain and an N-terminal cysteine of a contiguous Zera molecule already present in the oligomer). For this reason, when all N-terminal Cys residues or all C-terminal Cys residues were simultaneously mutated, no PB formation was observed (Fig. 2, I and K). Thus, the increasing number of hydrophobic interactions and disulfide bonds results in an unordered net of disulfide bonds in the Zera-ECFP self-assembly that leads to PB formation. A schematic representation of this molecular self-assembly is sketched in Fig. 7, which shows a small wedge-shaped portion of the entire spherical PB.
On the basis of the identification of the relevant traits involved in PB formation induced by the Zera sequence, we show that, whereas the presence of the six cysteines optimizes the formation of disulfide-linked Zera polymer, Cys7 and Cys9 are critical for homo-oligomer nucleation. The repeat region length and its amphipathic feature determine the efficiency of Zera-Zera self-assembly by hydrophobic interactions. Molecular dynamics studies are consistent with a model of oligomer growth in which disulfide cross-linking in an oxidizing redox context and hydrophobic interactions between parallel and anti-parallel Zera-ECFP chains are the crucial factors for PB formation. In steady-state confocal analysis of epidermal cells during Zera-ECFP expression over time, fluorescence was always visualized as concrete dot spots but was barely visible through the ER network, with Zera-ECFP accretions increasing in size but not in number. One explanation could be that Zera-ECFP mRNA localizes in specific sites, driving recombinant protein synthesis to specific subdomains of the ER. It is well established that the mRNA may contain signals that direct it to discrete locations within the cell (62). Recently, cis-localization elements have been identified in maize 10-kDa δ-zein RNA that are responsible for its targeting to cortical ER (PB-ER) subdomains (63) in transgenic rice plants. If Zera constructs contain cis-elements that drive RNA localization, these might reside in the coding region because the gene fusions do not contain the original 5′- and 3′-untranslated sequences of γ-zein. New experimental approaches are currently being used to explore this latter hypothesis.
Supplementary Material
Acknowledgments
We thank Miriam Ortiz and Esther Piñeiro for excellent technical support; J. J. Lopez-Moya for providing HcPro vector; and J. Casacuberta, Minu Joseph, and P. Mas for critical reading of the manuscript. The simulations were done on the supercomputer MareNostrum at the Barcelona Supercomputing Center, Centro Nacional de Supercomputación (Spanish National Supercomputing Center).
This work was supported by grants from the Generalitat de Catalunya XRB and SGR00182 (to M. D. L) and XRB and Grup Consolidat (to E. G.). This work was also supported by Ministerio de Ciencia y Tecnología-Fondo Europeo de Desarrollo Regional (MCYT-FEDER) Grant Bio2008-00799 and ERA Biotech S.A. (to M. D. L).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S5.
- ER
- endoplasmic reticulum
- ERAD
- ER-associated protein degradation
- MD
- molecular dynamics
- ECFP
- enhanced cyan fluorescent protein
- YFP
- yellow fluorescent protein
- PB
- protein body
- dpi
- days postinfiltration
- R8
- (PPPVHL)8 region
- PPII
- polyproline II helix.
REFERENCES
- 1.Herman E. M. (2008) Curr. Opin. Plant Biol. 11, 672–679 [DOI] [PubMed] [Google Scholar]
- 2.Müntz K. (1998) Plant Mol. Biol. 38, 77–99 [PubMed] [Google Scholar]
- 3.Kawagoe Y., Suzuki K., Tasaki M., Yasuda H., Akagi K., Katoh E., Nishizawa N. K., Ogawa M., Takaiwa F. (2005) Plant Cell 17, 1141–1153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pompa A., Vitale A. (2006) Plant Cell 18, 2608–2621 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vitale A., Ceriotti A. (2004) Plant Physiol. 136, 3420–3426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Starkuviene V., Pepperkok R. (2007) Traffic 8, 1035–1051 [DOI] [PubMed] [Google Scholar]
- 7.Saito K., Chen M., Bard F., Chen S., Zhou H., Woodley D., Polischuk R., Schekman R., Malhotra V. (2009) Cell 136, 891–902 [DOI] [PubMed] [Google Scholar]
- 8.Larkins B. A., Hurkman W. J. (1978) Plant Physiol. 62, 256–263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wu Y., Messing J. (2010) Plant Physiol. 153, 337–347 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Geli M. I., Torrent M., Ludevid D. (1994) Plant Cell 6, 1911–1922 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bagga S., Adams H., Kemp J. D., Sengupta-Gopalan C. (1995) Plant Physiol. 107, 13–23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Coleman C. E., Herman E. M., Takasaki K., Larkins B. A. (1996) Plant Cell 8, 2335–2345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bagga S., Adams H. P., Rodriguez F. D., Kemp J. D., Sengupta-Gopalan C. (1997) Plant Cell 9, 1683–1696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Boston R. S., Viitanen P. V., Vierling E. (1996) Plant. Mol. Biol. 32, 191–222 [DOI] [PubMed] [Google Scholar]
- 15.Muench D. G., Wu Y., Zhang Y., Li X., Boston R. S., Okita T. W. (1997) Plant Cell Physiol. 38, 404–412 [DOI] [PubMed] [Google Scholar]
- 16.Li X., Franceschi V. R., Okita T. W. (1993) Cell 72, 869–879 [DOI] [PubMed] [Google Scholar]
- 17.Crofts A. J., Washida H., Okita T. W., Ogawa M., Kumamaru T., Satoh H. (2004) Plant Physiol. 136, 3414–3419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Takemoto Y., Coughlan S. J., Okita T. W., Satoh H., Ogawa M., Kumamaru T. (2002) Plant Physiol. 128, 1212–1222 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pedrazzini E., Giovinazzo G., Bollini R., Ceriotti A., Vitale A. (1994) Plant J. 5, 103–110 [Google Scholar]
- 20.Pueyo J. J., Chrispeels M. J., Herman E. M. (1995) Planta 196, 586–596 [DOI] [PubMed] [Google Scholar]
- 21.Di Cola A., Frigerio L., Lord J. M., Ceriotti A., Roberts L. M. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 14726–14731 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Brandizzi F., Hanton S., DaSilva L. L., Boevink P., Evans D., Oparka K., Denecke J., Hawes C. (2003) Plant J. 34, 269–281 [DOI] [PubMed] [Google Scholar]
- 23.Ludevid M. D., Torrent M., Lasserre-Ramassamy S. (August1, 2004) International Patent WO2004003207
- 24.Torrent M., Llompart B., Lasserre-Ramassamy S., Llop-Tous I., Bastida M., Marzabal P., Westerholm-Parvinen A., Saloheimo M., Heifetz P. B., Ludevid M. D. (2009) BMC Biol. 7, 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Torrent M., Llop-Tous I., Ludevid M. D. (2009) Methods Mol. Biol. 483, 193–208 [DOI] [PubMed] [Google Scholar]
- 26.Mainieri D., Rossi M., Archinti M., Bellucci M., De Marchis F., Vavassori S., Pompa A., Arcioni S., Vitale A. (2004) Plant Physiol. 136, 3447–3456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Rabanal F., Ludevid M. D., Pons M., Giralt E. (1993) Biopolymers 33, 1019–1028 [DOI] [PubMed] [Google Scholar]
- 28.Kogan M. J., Dalcol I., Gorostiza P., López-Iglesias C., Pons M., Sanz F., Ludevid D., Giralt E. (2001) J. Mol. Biol. 312, 907–913 [DOI] [PubMed] [Google Scholar]
- 29.Conley A. J., Joensuu J. J., Menassa R., Brandle J. E. (2009) BMC Biol. 7, 48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Joensuu J. J., Conley A. J., Lienemann M., Brandle J. E., Linder M. B., Menassa R. (2010) Plant Physiol. 152, 622–633 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Conley A. J., Joensuu J. J., Jevnikar A. M., Menassa R., Brandle J. E. (2009) Biotechnol. Bioeng. 103, 562–573 [DOI] [PubMed] [Google Scholar]
- 32.Vitale A., Boston R. S. (2008) Traffic 9, 1581–1588 [DOI] [PubMed] [Google Scholar]
- 33.Torrent M., Geli M. I., Ruiz-Avila L., Canals J. M., Puigdomènech P., Ludevid D. (1994) Planta 192, 512–518 [DOI] [PubMed] [Google Scholar]
- 34.Goytia E., Fernández-Calvino L., Martínez-García B., López-Abella D., López-Moya J. J. (2006) J. Gen. Virol. 87, 3413–3423 [DOI] [PubMed] [Google Scholar]
- 35.Voinnet O., Rivas S., Mestre P., Baulcombe D. (2003) Plant J. 33, 949–956 [DOI] [PubMed] [Google Scholar]
- 36.Pfaffl M. W. (2001) Nucleic Acids Res. 29, e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Qu J., Ye J., Fang R. (2007) J. Virol. 81, 6690–6699 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Phillips J. C., Braun R., Wang W., Gumbart J., Tajkhorshid E., Villa E., Chipot C., Skeel R. D., Kalé L., Schulten K. (2005) J. Comput. Chem. 26, 1781–1802 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.MacKerell A. D., Bashford D., Bellott M., Dunbrack R. L., Evanseck J. D., Field M. J., Fischer S., Gao J., Guo H., Ha S., McCarthy D. J., Kuchnir L., Kuczera K., Lau F. T., Mattos C., Michnick S., Ngo T., Nguyen D. T., Prodhom B., Reiher W. E., Roux B., Schlenkrich M., Smith J. C., Stote R., Straub J., Watanabe M., Wiorkiewicz-Kuczera J., Yin D., Karplus M. (1998) J. Phys. Chem. B 102, 3586–3616 [DOI] [PubMed] [Google Scholar]
- 40.Ryckaert J. P., Ciccotti G., Berendsen H. J. (1977) J. Comp. Phys. 23, 327–341 [Google Scholar]
- 41.Martyna G. J., Tobias D. J., Klein M. L. (1994) J. Chem. Physiol. 101, 4177–4189 [Google Scholar]
- 42.Feller S. E., Zhang Y., Pastor R. W., Brooks B. R. (1995) J. Chem. Phys. 103, 4613–4621 [Google Scholar]
- 43.Humphrey W., Dalke A., Schulten K. (1996) J. Mol. Graph. 14, 33–38 [DOI] [PubMed] [Google Scholar]
- 44.Dalcol I., Pons M., Ludevid M. D., Giralt E. (1996) J. Org. Chem. 61, 6775–6782 [DOI] [PubMed] [Google Scholar]
- 45.Kogan M. J., Dalcol I., Gorostiza P., Lopez-Iglesias C., Pons R., Pons M., Sanz F., Giralt E. (2002) Biophys. J. 83, 1194–1204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Vila J. A., Baldoni H. A., Ripoll D. R., Ghosh A., Scheraga H. A. (2004) Biophys. J. 86, 731–742 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Alvarez M. L., Topal E., Martin F., Cardineau G. A. (2010) Plant Mol. Biol. 72, 75–89 [DOI] [PubMed] [Google Scholar]
- 48.Saito Y., Kishida K., Takata K., Takahashi H., Shimada T., Tanaka K., Morita S., Satoh S., Masumura T. (2009) J. Exp. Bot. 60, 615–627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Birk J., Friberg M. A., Prescianotto-Baschong C., Spiess M., Rutishauser J. (2009) J. Cell Sci. 122, 3994–4002 [DOI] [PubMed] [Google Scholar]
- 50.Shorrosh B. S., Subramaniam J., Schubert K. R., Dixon R. A. (1993) Plant Physiol. 103, 719–726 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wilkinson B., Gilbert H. F. (2004) Biochim. Biophys. Acta 1699, 35–44 [DOI] [PubMed] [Google Scholar]
- 52.Onda Y., Kumamaru T., Kawagoe Y. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 14156–14161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li C. P., Larkins B. A. (1996) Plant Mol. Biol. 30, 873–882 [DOI] [PubMed] [Google Scholar]
- 54.Wetzel R. (1999) Methods Enzymol. 309, 189–204 [DOI] [PubMed] [Google Scholar]
- 55.Foresti O., De Marchis F., de Virgilio M., Klein E. M., Arcioni S., Bellucci M., Vitale A. (2008) Mol. Plant 1, 1067–1076 [DOI] [PubMed] [Google Scholar]
- 56.Pimpl P., Taylor J. P., Snowden C., Hillmer S., Robinson D. G., Denecke J. (2006) Plant Cell 18, 198–211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Bicudo T. C., Bicudo R. C., Forato L. A., Beltramini L. M., Batista L. A., Filho R. B., Colnago L. A. (2008) Biopolymers 89, 175–178 [DOI] [PubMed] [Google Scholar]
- 58.Pujals S., Fernández-Carneado J., López-Iglesias C., Kogan M. J., Giralt E. (2006) Biochim. Biophys. Acta. 1758, 264–279 [DOI] [PubMed] [Google Scholar]
- 59.Kogan M. J., López O., Cocera M., López-Iglesias C., De La Maza A., Giralt E. (2004) Biopolymers 73, 258–268 [DOI] [PubMed] [Google Scholar]
- 60.Curtis R. A., Steinbrecher C., Heinemann M., Blanch H. W., Prausnitz J. M. (2002) Biophys. Chem. 98, 249–265 [DOI] [PubMed] [Google Scholar]
- 61.Hwang C., Sinskey A. J., Lodish H. F. (1992) Science 257, 1496–1502 [DOI] [PubMed] [Google Scholar]
- 62.St Johnston D. (2005) Nat. Rev. Mol. Cell. Biol. 6, 363–375 [DOI] [PubMed] [Google Scholar]
- 63.Washida H., Sugino A., Kaneko S., Crofts N., Sakulsingharoj C., Kim D., Choi S. B., Hamada S., Ogawa M., Wang C., Esen A., Higgins T. J., Okita T. W. (2009) Plant J. 60, 146–155 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.