Abstract
Vertebrate genomic assemblies were analyzed for endogenous sequences related to any known viruses with single-stranded DNA genomes. Numerous high-confidence examples related to the Circoviridae and two genera in the family Parvoviridae, the parvoviruses and dependoviruses, were found and were broadly distributed among 31 of the 49 vertebrate species tested. Our analyses indicate that the ages of both virus families may exceed 40 to 50 million years. Shared features of the replication strategies of these viruses may explain the high incidence of the integrations.
It has long been appreciated that retroviruses can contribute significantly to the genetic makeup of host organisms. Genes related to certain other viruses with single-stranded RNA genomes, formerly considered to be most unlikely candidates for such contribution, have recently been detected throughout the vertebrate phylogenetic tree (1, 6, 13). Here, we report that viruses with single-stranded DNA (ssDNA) genomes have also contributed to the genetic makeup of many organisms, stretching back as far as the Paleocene period and possibly the late Cretaceous period of evolution.
Determining the evolutionary ages of viruses can be problematic, as their mutation rates may be high and their replication may be rapid but also sporadic. To establish a lower age limit for currently circulating ssDNA viruses, we analyzed 49 published vertebrate genomic assemblies for the presence of sequences derived from the NCBI RefSeq database of 2,382 proteins from known viruses in this category, representing a total of 23 classified genera from 7 virus families. Our survey uncovered numerous high-confidence examples of endogenous sequences related to the Circoviridae and to two genera in the family Parvoviridae: the parvoviruses and dependoviruses (Fig. 1).
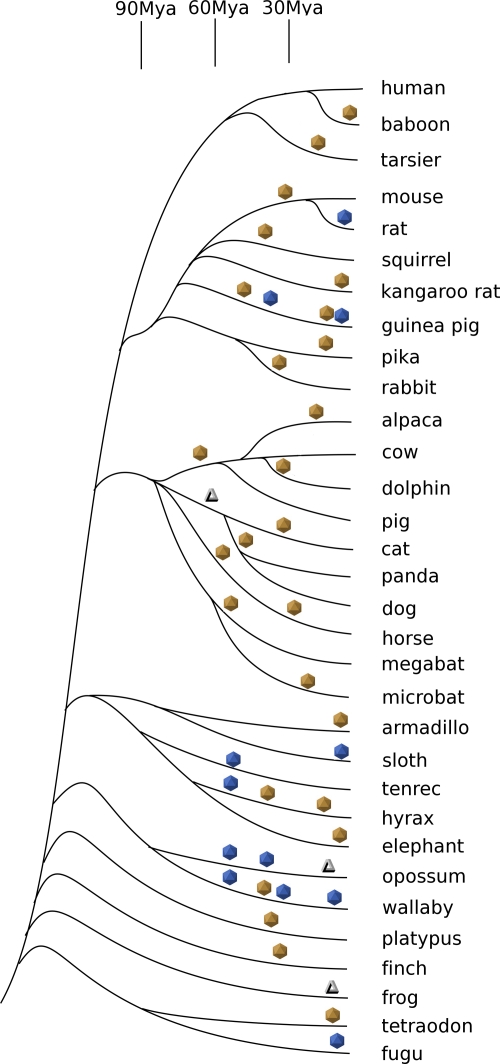
FIG. 1.
Phylogenetic tree of vertebrate organisms and history of ssDNA virus integrations. Times of integration of ancestral dependoviruses (yellow icosahedrons), parvoviruses (blue icosahedrons), and circoviruses (triangles) are approximate.
The Dependovirus and Parvovirus genomes are typically 4 to 6 kb in length, include 2 major open reading frames (encoding replicase proteins [Rep and NS1, respectively] and capsid proteins [Cap and VP1, respectively]), and have characteristic hairpin structures at both ends (Fig. 2). For replication, these viruses depend on host enzymes that are recruited by the viral replicase proteins to the hairpin regions, where self-primed viral DNA synthesis is initiated (2). Circovirus genomes are typically ∼2-kb circles. DNA of the type species, porcine circovirus 1 (PCV-1), contains a stem-loop structure within the origin of replication (Fig. 2), and the largest open reading frame includes sequences that are homologous to the Parvovirus replicase open reading frame (9, 11). The circoviruses also depend on host enzymes for replication, and DNA synthesis is self-primed from a 3′-OH end formed by endonucleolytic cleavage of the stem-loop structure (4). The frequency of Dependovirus infection is estimated to be as high as 90% within an individual's lifetime. None of the dependoviruses have been associated with human disease, but related viruses in the family Parvoviridae (e.g., erythrovirus B19 and possibly human bocavirus) are pathogenic for humans, and members of both the Parvoviridae and the Circoviridae can cause a variety of animal diseases (2, 4).
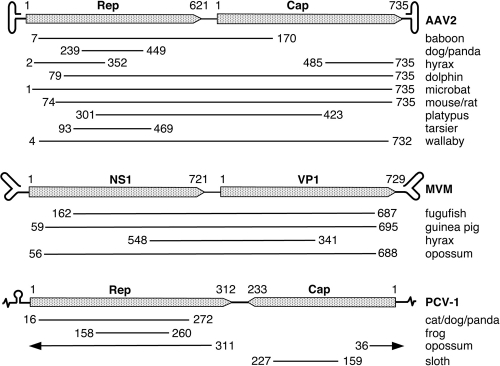
FIG. 2.
Schematics illustrating the structure and organization of Parvoviridae and Circoviridae genomes and origins of several of the longest-integrated ancestral viral sequences found in vertebrates. Integrations were aligned to the Dependovirus adeno-associated virus 2 (AAV2), the Parvovirus minute virus of mice (MVM), and the Circovirus porcine circovirus 1 (PCV-1). The inverted terminal repeat (ITR) sequences in the Dependovirus and Parvovirus genomes are depicted on an expanded scale. A linear representation of the circular genome of PCV-1 is shown with the 10-bp stem-loop structure on an expanded scale. Horizontal lines beneath the maps indicate the lengths of similar sequences that could be identified by BLAST. The numbers indicate the locations of amino acids in the viral proteins where the sequence similarities in the endogenous insertions start and end. The actual ancestral virus-derived integrated sequences may extend beyond the indicated regions.
With some ancestral endogenous sequences that we identified, phylogenetic comparisons can be used to estimate age. For example, as a Dependovirus-like sequence is present at the same location in the genomes of mice and rats, the ancestral virus must have existed before their divergence, more than 20 million years ago. Some Circovirus- and Dependovirus-related integrations also predate the split between dog and panda, about 42 million years ago. However, in most other cases, we rely on an indirect method for estimating age (1). As genomic sequences evolve, they accumulate new stop codons and insertion/deletion-induced frameshifts. The rates of these events can be tied directly to the rates of neutral sequence drift and, therefore, the time of evolution. To apply this method, we first performed a BLAST search of vertebrate genomes for all known ssDNA virus proteins (BLAST options, -p tblastn -M BLOSUM62 -e 1e−4). Candidate sequences were then recorded, along with 5 kb of flanking regions, and then again aligned against the database of ssDNA viruses to find the most complete alignment (BLAST options, -t blastx -F F -w 15 -t 1500 -Z 150 -G 13 -E 1 -e 1e−2). Detected alignments were then compared with a neutral model of genome evolution, as described in the supplemental material, and the numbers of stop codons and frameshifts were converted into the expected genomic drift undergone by the sequences. The age of integration was then estimated from the known phylogeny of vertebrates (7, 10). Using these methods, we discovered that as many as 110 ssDNA virus-related sequences have been integrated into the 49 vertebrate genomes considered, during a time period ranging from the present to over 40 to 60 million years ago (Table 1; see also Tables S1 to S3 in the supplemental material).
TABLE 1.
Selected endogenous sequences in vertebrate genomes related to single-stranded DNA viruses
| Virus group and vertebrate species | Initial genomic search using TBLASTN |
Best sequence homology identified using BLASTX |
Predicted nucleotide drift (%) | Integration label | Age (million yr) or timing of integration based on sequence aging | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Chromosomal or scaffold location | Protein | BLAST E value/% sequence identity | Most similar virusa | Protein | Coordinates | No. of stop codons/frameshifts | ||||
| Circoviruses | ||||||||||
| Cat | Scaffold_62068 | Rep | 6E−05/37 | Canary circovirus | Rep | 4-283 | 3/7 in 268 aab | 14.2 | fcECLG-1 | 82 |
| Scaffold_24038 | Rep | 6E−06/51 | Columbid circovirus | Rep | 44-317 | 4/5 in 231 aac | 15.2 | fcECLG-2 | 87 | |
| Dog | Chr5d | Rep | 7E−16/46 | Raven circovirus | Rep | 16-263 | 6/5 in 250 aa | 17.6 | cfECLG-1 | 98 |
| Chr22 | Rep | 1E−14/43 | Beak and feather disease virus | Rep | 7-264 | 2/1 in 261 aac | 4.5 | cfECLG-2 | 54 | |
| Opossum | Chr3 | Rep | 4E−46/44 | Finch circovirus | Rep | 2-291 | 0/2 in 282 aa | 2.3 | mdECLG | 12 |
| Cap | 6-36 | 0/0 in 30 aa | ||||||||
| Dependoviruses | ||||||||||
| Dog | ChrX | Rep | 6E−05/55 | AAV5 | Rep | 239-445 | 3/4 in 200 aa | 14.0 | cfEDLG-1 | 78 |
| Dolphin | GeneScaffold1475 | Rep | 8E−39/39 | Avian AAV DA1 | Rep | 79-486 | 3/4 in 379 aac | 6.6 | ttEDLG-2 | 55 |
| Cap | 4E−61/47 | Cap | 1-738 | 4/7 in 678 aac | ||||||
| Elephant | Scaffold_4 | Rep | 0/55 | AAV5 | Rep | 3-589 | 0/0 in 579 aa | 0.0 | laEDLG | Recent |
| Hyrax | GeneScaffold5020 | Cap | 3E−34/53 | AAV3 | Cap | 485-735 | 0/5 in 256 aa | 7.0 | pcEDLG-1 | 29 |
| Scaffold_19252 | Rep | 9E−72/47 | Bovine AAV | Rep | 2-348 | 8/4 in 348 aa | 14.3 | pcEDLG-2 | 60 | |
| Megabat | Scaffold_5601 | Rep | 2E−13/31 | AAV2 | Rep | 315-479 | 1/5 in 175 aa | 13.1 | pvEDLG-3 | 76 |
| Microbat | GeneScaffold2026 | Rep | 1E−117/50 | AAV2 | Rep | 1-617 | 2/5 in 612 aa | 5.8 | mlEDLG-1 | 27 |
| Cap | 9E−33/51 | Cap | 1-731 | 2/9 in 509 aac | ||||||
| Scaffold_146492 | Cap | 6E−32/42 | AAV2 | Cap | 479-732 | 0/3 in 252 aa | 4.2 | mlEDLG-2 | 19 | |
| Mouse | Chr1 | Rep | 2E−06/34 | AAV2 | Rep | 4-206 | 3/5 in 191 aa | 17.1 | mmEDLG-1 | 39 |
| Chr3 | Rep | 2E−24/31 | AAV5 | Rep | 71-478 | 12/7 in 389 aa | 16.5 | mmEDLG-2 | 37 | |
| Cap | 2E−22/45 | Cap | 22-724 | 12/10 in 649aac | ||||||
| Chr8 | Rep | 1E−08/46 | AAV2 | Rep | 314-473 | 3/3 in 147 aa | 13.8 | mmEDLG-3 | 31 | |
| Cap | 1-137 | 1/2 in 114 aa | ||||||||
| Panda | Scaffold2359 | Rep | 2E−06/37 | Bovine AAV | Rep | 238-426 | 2/3 in 186 aa | 10.4 | amEDLG-1 | 59 |
| Pika | Scaffold_9941 | Rep | 4E−14/28 | AAV5 | Rep | 126-415 | 2/2 in 282 aa | 5.4 | opEDLG | 14 |
| Platypus | Chr2 | Rep | 9E−10/35 | Bovine AAV | Rep | 297-437 | 4/3 in 138 aa | 17.1 | oaEDLG-1 | 79 |
| Cap | 272-419 | 1/2 in 150 aac | ||||||||
| Contig12430 | Rep | 2E−09/47 | Bovine AAV | Rep | 353-450 | 3/1 in 123 aa | 12.0 | oaEDLG-2 | 55 | |
| Cap | 2E−05/32 | Cap | 253-367 | 2/1 in 116 aa | ||||||
| Rabbit | Chr10 | Rep | 3E−97/39 | AAV2 | Rep | 1-619 | 3/9 in 613 aa | 9.3 | ocEDLG | 43 |
| Cap | 5E−50/45 | Cap | 1-723 | 10/9 in 675 aa | ||||||
| Rat | Chr13 | Rep | 2E−09/33 | AAV2 | Rep | 4-175 | 2/4 in 177 aa | 13.3 | rnEDLG-1 | 28 |
| Chr2 | Rep | 4E−18/40 | AAV5 | Rep | 1-461 | 12/12 in 454 aa | 22.7 | rnEDLG-2 | 51 | |
| Chr19 | Rep | 2E−07/33 | AAV5 | Rep | 329-464 | 2/4 in 136 aa | 16.1 | rnEDLG-3 | 35 | |
| Cap | 31-133 | 2/1 in 93 aa | ||||||||
| Tarsier | Scaffold_178326 | Rep | 4E−14/23 | AAV5 | Rep | 96-465 | 2/3 in 356 aa | 5.3 | tsEDLG | 23 |
| Parvoviruses | ||||||||||
| Guinea pig | Scaffold_188 | Rep | 3E−24/46 | Porcine parvovirus | Rep | 313-567 | 5/3 in 250 aa | 12.3 | cpEPLG-1 | 40 |
| Cap | 1E−16/36 | Cap | 10-689 | 11/12 in 672 aa | ||||||
| Scaffold_27 | Rep | 1E−50/39 | Canine parvovirus | Rep | 11-640 | 1/4 in 616 aa | 5.3 | cpEPLG-2 | 17 | |
| Cap | 1E−38/39 | Porcine parvovirus | Cap | 3-719 | 2/14 in 700 aa | |||||
| Tenrec | Scaffold_260946 | Rep | 2E−20/38 | LuIII virus | Rep | 406-598 | 4/4 in 190 aa | 19.0 | etEPLG-2 | 60 |
| Cap | 11-639 | 16/15 in 595 aa | ||||||||
| Rat | Chr5 | Rep | 6E−10/56 | Canine parvovirus | Rep | 1-282 | 0/0 in 312 aa | 0.6 | rnEPLG | Recent |
| Cap | 0/62 | Cap | 637-667 | 0/2 in 760 aa | ||||||
| Rep | 0/63 | 1-751 | ||||||||
| Opossum | Chr3 | Rep | 2E−39/33 | LuIII virus | Rep | 7-570 | 11/3 in 502 aa | 10.9 | mdEPLG-2 | 56 |
| Cap | 7E−8/33 | Cap | 11-729 | 14/7 in 704 aa | ||||||
| Chr6 | Rep | 6E−58/44 | Porcine parvovirus | Rep | 16-563 | 3/7 in 534 aac | 4.6 | mdEPLG-3 | 24 | |
| Cap | 6E−60/38 | Cap | 10-715 | 2/5 in 707 aac | ||||||
| Wallaby | Scaffold_108040 | Rep | 4E−74/62 | Canine parvovirus | Rep | 341-645 | 0/0 in 287 aa | 1.3 | meEPLG-3 | 7 |
| Cap | 8E−37/32 | Cap | 35-738 | 0/4 in 687 aa | ||||||
| Scaffold_72496 | Rep | 2E−61/42 | Porcine parvovirus | Rep | 23-567 | 4/3 in 531 aa | 5.7 | meEPLG-6 | 30 | |
| Cap | 2E−31/38 | Cap | 10-532 | 6/4 in 514 aa | ||||||
| Scaffold_88340 | Rep | 7E−37/55 | Mouse parvovirus 1 | Rep | 344-566 | 0/3 in 223 aa | 6.7 | meEPLG-16 | 36 | |
| Cap | 7E−22/33 | Cap | 11-713 | 6/9 in 700 aa | ||||||
Some ambiguity in choosing the most similar virus is possible. We generally used the alignment with the lowest E value in the BLAST results. However, one or two points in the exponent of an E value were sometimes sacrificed to achieve a longer sequence alignment.
aa, amino acids.
These sequences have long insertions compared to the present-day viruses. In all cases tested, these insertions originated from short interspersed elements (SINEs). These insertions were excluded from the counts of stop codons and frameshifts and the estimation of integration age.
Chr, chromosome.
It is important to recognize that there is an intrinsic limit on how far back in time we can reach to identify ancient endogenous viral sequences. First, the sequences must be identified with confidence by BLAST or similar programs. This requirement places a lower limit on sequence identity at about 20 to 30% of amino acids, or about 75% of nucleotides (nucleotides evolve nearly 2.5 times slower than the amino acid sequence they encode). Second, the related, present-day virus must have evolved at a rate that is not much higher than that of the endogenous sequences. The viruses for which ancestral endogenous sequences were identified in this study exhibit sequence drift similar to that associated with mammalian genomes. Setting this rate at 0.14% per million years of evolution (8), we arrive at 90 million years as the theoretical limit for the oldest sequences that can be identified using our methods. This limit drops to less than 35 million years for endogenous viral sequences in rodents and even lower for sequences related to viruses that evolve faster than mammalian genomes.
The most widespread integrations found in our survey are derived from the dependoviruses. These include nearly complete genomes related to adeno-associated virus (AAV) in microbat, wallaby, dolphin, rabbit, mouse, and baboon (Fig. 2). We did not detect inverted terminal repeats in several integrations tested, even though repeats are common in the present-day dependoviruses. This result could be explained by sequence decay or the absence of such structures in the ancestral viruses. However, we do see sequences that resemble degraded hairpin structures to which Dependovirus Rep proteins bind, with an example from microbat integration mlEDLG-1 shown in Fig. 3. The second most widespread endogenous sequences are related to the parvoviruses. They are found in 6 of 49 vertebrate species considered, with nearly complete genomes in rat, opossum, wallaby, and guinea pig (Fig. 2).
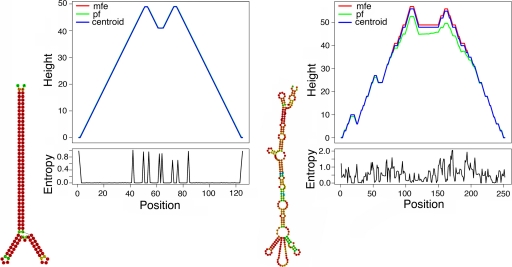
FIG. 3.
Hairpin structure of the inverted terminal repeat of adeno-associated virus 2 (left) and a candidate degraded hairpin structure located close to the 5′ end of the mlEDLG-1 integration in microbats (right). Structures and mountain plots were generated using default parameters of the RNAfold program (5), with nucleotide coloring representing base-pairing probabilities: blue is below average, green is average, and red is above average. Mountain plots represent hairpin structures based on minimum free energy (mfe) calculations and partition function (pf) calculations, as well as the centroid structure (5). Height is expressed in numbers of nucleotides; position represents nucleotide.
The Dependovirus AAV2 has strong bias for integration into human chromosome 19 during infection, driven by a host sequence that is recognized by the viral Rep protein(s). Rep mediates the formation of a synapse between viral and cellular sequences, and the cellular sequences are nicked to serve as an origin of viral replication (14). The related integrations in mice and rats, located in the same chromosomal locations, might be explained by such a mechanism. However, the extent of endogenous sequence decay and the frequency of stop codons indicate that these integrations occurred some 30 to 35 million years ago, implying that they are derived from a single event in a rodent ancestor rather than two independent integration events at the same location. Similarly, integrations EDLG-1 in dog and panda lie in chromosomal regions that can be readily aligned (based on University of California—Santa Cruz [UCSC] genome assemblies) and show sequence decay consistent with the age of the common ancestor, about 42 million years. Endogenous sequences related to the family Parvoviridae can thus be traced to over 40 million years back in time, and viral proteins related to this family have remained over 40% conserved.
Sequences related to circoviruses were detected in five vertebrate species (Table 1 and Table S1 in the supplemental material). At least one of these sequences, the endogenous sequence in opossum, likely represents a recent integration. Several integrations in dog, cat, and panda, on the other hand, appear to date from at least 42 million years ago, which is the last time when pandas and dogs shared a common ancestor. We see evidence for this age in data from sequence degradation (Table 1), phylogenetic analyses of endogenous Circovirus-like genomes (see Fig. S2 in the supplemental material), and genomic synteny where integration ECLG-3 is surrounded by genes MTA3 and ARID5A in both dog and panda and integration ECLG-2 lies 35 to 43 kb downstream of gene UPF3A. In fact, Circovirus integrations may even precede the split between dogs and cats, about 55 million years ago, although the preliminary assembly and short genomic contigs for cats make synteny analysis impossible.
The most common Circovirus-related sequences detected in vertebrate genomes are derived from the rep gene. We speculate that, like those of the Parvoviridae, the ancestral Circoviridae sequences might have been copied using a primer sequence in the host DNA that resembled the viral origin and was therefore recognized by the virus Rep protein. Higher incidence of rep gene identifications may represent higher conservation of this gene with time, or alternatively, possession of these sequences may impart some selective advantage to the host species. The largest Circovirus-related integration detected, in the opossum, comprises a short fragment of what may have been the cap gene immediately adjacent to and in the opposite orientation from the rep gene. This organization is similar to that of the present day Circovirus genome in which these genes share a promoter in the hairpin regions but are translated in opposite directions (Fig. 2).
In summary, our results indicate that sequences derived from ancestral members of the families Parvoviridae and Circoviridae were integrated into their host's genomes over the past 50 million years of evolution. Features of their replication strategies suggest mechanisms by which such integrations may have occurred. It is possible that some of the endogenous viral sequences could offer a selective advantage to the virus or the host. We note that rep open reading frame-derived proteins from some members of these families kill tumor cells selectively (3, 12). The genomic “fossils” we have discovered provide a unique glimpse into virus evolution but can give us only a lower estimate of the actual ages of these families. However, numerous recent integrations suggest that their germ line transfer has been continuing into present times.
Supplementary Material
Acknowledgments
A.M.S. was funded by National Institutes of Health grants CA71515 and CA06927 and also by an appropriation from the Commonwealth of Pennsylvania. V.A.B. is supported by membership from Martin A. and Helen Chooljian at the Institute for Advanced Study.
We thank Marie Estes for excellent secretarial assistance and Karen Trush for help with the figures.
The contents of this paper are the responsibility solely of the authors and do not necessarily represent the official views of the National Cancer Institute or any other sponsoring organization.
Footnotes
Published ahead of print on 22 September 2010.
Supplemental material for this article may be found at http://jvi.asm.org/.
REFERENCES
- 1.Belyi, V. A., A. J. Levine, and A. M. Skalka. 2010. Unexpected inheritance: multiple integrations of ancient Bornavirus and Ebolavirus/Marburgvirus sequences in vertebrate genomes. PLoS Pathog. 6:e1001030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Berns, K., and C. R. Parrish. 2007. Parvoviridae, p. 2437-2478. In D. M. Knipe and P. M. Howley (ed.), Fields virology. Lippincott Williams & Wilkins, Philadelphia, PA.
- 3.de Smith, M. H., and M. H. M. Noteborn. 2009. Apoptosis-inducing proteins in chicken anemia virus and TT virus. In E.-M. de Villiers and H. zur Hausen (ed.), TT viruses: the still elusive human pathogens. Springer-Verlag, Berlin, Germany. [PubMed]
- 4.Faurez, F., D. Dory, B. Grasland, and A. Jestin. 2009. Replication of porcine circoviruses. Virol. J. 6:60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gruber, A. R., R. Lorenz, S. H. Bernhart, R. Neubock, and I. L. Hofacker. 2008. The Vienna RNA websuite. Nucleic Acids Res. 36:W70-W74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Horie, M., T. Honda, Y. Suzuki, Y. Kobayashi, T. Daito, T. Oshida, K. Ikuta, P. Jern, T. Gojobori, J. M. Coffin, and K. Tomonaga. 2010. Endogenous non-retroviral RNA virus elements in mammalian genomes. Nature 463:84-87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hubbard, T. J., B. L. Aken, S. Ayling, B. Ballester, K. Beal, E. Bragin, S. Brent, Y. Chen, P. Clapham, L. Clarke, G. Coates, S. Fairley, S. Fitzgerald, J. Fernandez-Banet, L. Gordon, S. Graf, S. Haider, M. Hammond, R. Holland, K. Howe, A. Jenkinson, N. Johnson, A. Kahari, D. Keefe, S. Keenan, R. Kinsella, F. Kokocinski, E. Kulesha, D. Lawson, I. Longden, K. Megy, P. Meidl, B. Overduin, A. Parker, B. Pritchard, D. Rios, M. Schuster, G. Slater, D. Smedley, W. Spooner, G. Spudich, S. Trevanion, A. Vilella, J. Vogel, S. White, S. Wilder, A. Zadissa, E. Birney, F. Cunningham, V. Curwen, R. Durbin, X. M. Fernandez-Suarez, J. Herrero, A. Kasprzyk, G. Proctor, J. Smith, S. Searle, and P. Flicek. 2009. Ensembl 2009. Nucleic Acids Res. 37:D690-D697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li, W. H., D. L. Ellsworth, J. Krushkal, B. H. Chang, and D. Hewett-Emmett. 1996. Rates of nucleotide substitution in primates and rodents and the generation-time effect hypothesis. Mol. Phylogenet. Evol. 5:182-187. [DOI] [PubMed] [Google Scholar]
- 9.Mankertz, A., J. Mankertz, K. Wolf, and H. J. Buhk. 1998. Identification of a protein essential for replication of porcine circovirus. J. Gen. Virol. 79:381-384. [DOI] [PubMed] [Google Scholar]
- 10.Murphy, W. J., T. H. Pringle, T. A. Crider, M. S. Springer, and W. Miller. 2007. Using genomic data to unravel the root of the placental mammal phylogeny. Genome Res. 17:413-421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Niagro, F. D., A. N. Forsthoefel, R. P. Lawther, L. Kamalanathan, B. W. Ritchie, K. S. Latimer, and P. D. Lukert. 1998. Beak and feather disease virus and porcine circovirus genomes: intermediates between the geminiviruses and plant circoviruses. Arch. Virol. 143:1723-1744. [DOI] [PubMed] [Google Scholar]
- 12.Nuesch, J. P., and J. Rommelaere. 2007. A viral adaptor protein modulating casein kinase II activity induces cytopathic effects in permissive cells. Proc. Natl. Acad. Sci. U. S. A. 104:12482-12487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Taylor, D. J., R. W. Leach, and J. Bruenn. 2010. Filoviruses are ancient and integrated into mammalian genomes. BMC Evol. Biol. 10:193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Urcelay, E., P. Ward, S. M. Wiener, B. Safer, and R. M. Kotin. 1995. Asymmetric replication in vitro from a human sequence element is dependent on adeno-associated virus Rep protein. J. Virol. 69:2038-2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.