Summary
The basic helix-loop-helix transcription factor Twist1 is essential for normal limb development. Twist1−/− embryos die at midgestation. However, studies on early limb buds found that Twist1−/− mutant limb mesenchyme has an impaired response to FGF signaling from the apical ectodermal ridge, which disrupts the feedback loop between the mesenchyme and AER, and reduces and shifts anteriorly Shh expression in the zone of polarizing activity. We have combined Twist1 null, hypomorph and conditional alleles to generate a Twist1 allelic series that survives to birth. As Twist1 activity is reduced, limb skeletal defects progress from preaxial polydactyly to girdle reduction combined with hypoplasia, aplasia or mirror symmetry of all limb segments. With reduced Twist1 activity there is striking and progressive upregulation of ectopic Shh expression in the anterior of the limb, combined with an anterior shift in the posterior Shh domain, which is expressed at normal intensity, and loss of the posterior AER. Consequently limb outgrowth is initially impaired, before an ectopic anterior Shh domain expands the AER, promoting additional growth and repatterning. Reducing the dosage of FGF targets of the Etv gene family, which are known repressors of Shh expression in the anterior limb mesenchyme, strongly enhances the anterior skeletal phenotype. Conversely this and other phenotypes are suppressed by reducing the dosage of the Twist1 antagonist Hand2. Our data support a model whereby multiple Twist1 activity thresholds contribute to early limb bud patterning, and suggest how particular combinations of skeletal defects result from differing amounts of Twist1 activity.
Keywords: limb, Twist1, pattern, signaling center, ZPA, AER
Introduction
Limb patterning is coordinated by discrete signaling centers, located within the limb mesenchyme or overlying ectoderm (reviewed in (Zeller et al., 2009)). Anteroposterior patterning is primarily regulated by Sonic hedgehog (Shh) produced within the posterior mesenchymal zone of polarizing activity (ZPA) (Riddle et al., 1993), while proximodistal outgrowth is regulated by fibroblast growth factor (FGF) signals produced by the distal ectodermal apical ectodermal ridge (AER) (Fallon et al., 1994; Niswander et al., 1993). Patterning requires both appropriate responses to these signals, and precise temporally and spatially regulated signal expression. Thus ectopic Shh expression in anterior limb mesenchyme causes preaxial polydactyly (Riddle et al., 1993), while AER disruptions lead to skeletal element loss (Boulet et al., 2004; Saunders, 1948; Sun et al., 2002).
Shh expression is regulated by combinatorial interactions between positive and negative transcription factors and growth factor signaling. Posterior mesenchyme is made competent to express Shh by multiple transcription factors (Davenport et al., 2003; te Welscher et al., 2002; Zakany et al., 2004). These work in concert with AER-derived FGF signals to induce Shh expression, although whether secondary signals downstream of FGF are required for Shh expression has not been determined. Shh expression is maintained in competent tissue by a positive feedback loop between Shh and Fgf4/Fgf8 in the posterior AER (Laufer et al., 1994; Niswander et al., 1994). Competence factors for Shh expression include Hand2 and Tbx3, which are expressed in posterior limb mesenchyme, initially in response to primary axial patterning cues, and later in response to Shh signaling (Charite et al., 2000; Davenport et al., 2003; Fernandez-Teran et al., 2000; te Welscher et al., 2002). Shh expression is prevented in anterior and distal limb mesenchyme by other transcription factors, including Alx4 (Qu et al., 1997), and the Ets family proteins Etv4 (Pea3) and Etv5 (Erm) that act downstream of AER-derived FGF signals (Mao et al., 2009; Zhang et al., 2009). Thus Shh expression reflects a balance between positively and negatively acting factors that position the Shh expression domain along the anteroposterior and proximodistal axes.
Twist1 is a bHLH transcription factor implicated as a regulator of limb development (Cai and Jabs, 2005; Chen and Behringer, 1995; O'Rourke et al., 2002; Zuniga et al., 2002). Twist1 is expressed in lateral plate mesenchyme prior to limb outgrowth and becomes progressively restricted to the peripheral mesenchyme within the limb. In Twist1−/− mice forelimb growth is stunted (Chen and Behringer, 1995), which correlates with failure of AER maintenance, and induction of ectopic posterior mesenchymal cell death (O'Rourke et al., 2002; Zuniga et al., 2002). Molecular marker analysis suggests a positive feedback loop between Fgf10 in the mesenchyme and Fgf8 in the AER breaks down, as Fgfr1 expression is lost from Twist1−/− limb mesenchyme. Shh expression is severely reduced and shifted to more distal mesenchyme in Twist1−/− forelimbs. Expression of several Shh- and FGF-dependent patterning genes is also abnormal (O'Rourke et al., 2002; Zuniga et al., 2002). A number of genes apparently downstream of Twist1, including Alx family genes, have been identified (Loebel et al., 2002). However the functional significance and specificity of these gene expression changes has not been assessed because Twist1−/− embryos die by embryonic day (E) 10.5, prior to significant limb outgrowth.
Genetic evidence suggests that Twist1 also negatively regulates Shh signaling and/or Shh expression in anterior limb mesenchyme. Twist1+/− mice or human Saethre-Chotzen syndrome (SCS) patients who display a TWIST1 haploinsufficiency have mild, and variable, distal limb abnormalities (Bourgeois et al., 1998; Cai and Jabs, 2005; Firulli et al., 2005). These include hindlimb preaxial polydactyly, which is bilateral in 25% of Twist1+/− mice (Bourgeois et al., 1998). The murine polydactyly is completely suppressed by reducing the gene dosage of Hand2, a member of the Twist bHLH family that is a positive regulator of Shh expression (Firulli et al., 2005). In chick limbs Twist1 overexpression can reduce the severity of Hand2-induced preaxial polydactyly (Firulli et al., 2005). While an antagonistic balance between Twist1 and Hand2 activities is thus required for normal anteroposterior patterning, the molecular changes are not well described because of the low expressivity of the Twist1+/− phenotype.
Comparing the Twist1+/− morphological phenotype with the Twist1−/− molecular phenotype raises inconsistencies that suggest relatively complex functions for Twist1 in early limb development. Notably, more severe phenotypes than mild hindlimb polydactyly would be predicted from the changes in both forelimb and hindlimb gene expression patterns observed in null mutant embryos. Recently a novel murine Twist1 allele was identified in an ENU mutagenesis screen. Charlie Chaplin (Twist1CC) encodes an S192P amino acid substitution that disrupts the function of a C-terminal protein interaction domain (Bialek et al., 2004). Fifty-two percent of Twist1CC/+ mice were reported to have bilateral hindlimb polydactyly, while Twist1CC/CC mice have short limbs and hindlimb polydactyly, but die perinatally. Thus Twist1CC mutants are affected relatively severely and survive long enough to allow the integrated analysis of Twist1 morphological and molecular phenotypes.
We generated a Twist1 allelic series using Twist1CC and Twist1− mutant alleles to ask how progressively reducing Twist1 activity affects limb development and patterning. Twist1 mutant phenotypes include discrete temporal and spatial molecular defects, with concordant and dramatic changes in limb and girdle cartilage pattern. Furthermore we find that Twist1 positions the posterior Shh domain along the anteroposterior axis in a dosage sensitive manner and is a major negative regulator of Shh expression in the anterior limb. Twist1 apparently exerts its effects through a network of transcription factors that include Etv and Alx family genes, each of which contributes to aspects of the Twist1 phenotype. We provide a model that shows how posterior defects in early signaling center regulation can cause loss of anterior skeletal elements, and that is consistent with different thresholds of Twist1 activity regulating different aspects of limb patterning.
Materials and Methods
Mouse strains and genotyping
Animal experiments were performed according to Columbia University Institutional Animal Care guidelines. Noon on the day of the mating plug was considered E0.5. Hand2− (Srivastava et al., 1997), Twist1tm1Bhr (Twist1−; (Chen and Behringer, 1995), ShhLacZ (Shh−; (Jeong et al., 2004), Twist1Skam10Jus (Twist1CC; (Bialek et al., 2004), Twist1flox (Chen et al., 2007), Prx1-creTg (Logan et al., 2002), Etv4tm1Arbr (Etv4−; (Livet et al., 2002) and Etv5LacZ (Etv5−; (Lu et al., 2009) alleles were maintained on B6, 129Sv or 129Sv.B6 backgrounds. Mice were genotyped by PCR, and Twist1CC T707C substitutions were confirmed by sequencing.
Skeletal stains, digit scoring and in situ hybridization
Bone/cartilage stains were performed using Alcian blue/alizarin red S staining as described (Webb and Byrd, 1994). Digits were scored on the basis of phalange number and morphology (Patton and Kaufman, 1995). Whole-mount and section in situ hybridization was performed as described (Laufer et al., 1997). All gene expression analyses were performed on 3 or more limbs. Riboprobes used: Fgf8 (Crossley and Martin, 1995), Shh (Roelink et al., 1994), Twist1 (Chen and Behringer, 1995), Hand2 (Srivastava et al., 1995), HoxA11 (Davis et al., 1995), HoxD11 (Burke et al., 1995), Ptch1 (Marigo et al., 1996), Tbx2 (Chapman et al., 1996), Tbx3 (Chapman et al., 1996), Spry1 (Zhang et al., 2001), Pax1 (Balling et al., 1988), Fgfr1 (Peters et al., 1992), Etv5 (Chotteau-Lelievre et al., 2001), Alx4 (te Welscher et al., 2002), Gli1 (Hui et al., 1994), cSpry1 (Minowada et al., 1999), cShh (Riddle et al., 1993).
Chick limb experiments
Fertile SPF White Leghorn chicken eggs (Charles River) were incubated at 37.5°C in a humidified incubator and staged according to Hamburger and Hamilton (Hamburger and Hamilton, 1951). Stage 20 forelimb buds were infected with replication-defective activated Fgfr1 virus (Fgfr1*, RIS-pm174HA; (Liu et al., 2001), and limbs harvested up to 42 h later. In some experiments the posterior AER was removed (Laufer et al., 1994) prior to retroviral infection. The AMV-3C2 viral gag antibody was obtained from the Developmental Studies Hybridoma Bank. Section immunohistochemistry was performed as described (Vargesson et al., 2001).
Results
Effects of altering Twist1 activity on limb cartilage patterns
We performed the appropriate genetic crosses to generate Twist1CC/+, Twist1+/−, Twist1CC/CC and Twist1CC/− mutant animals. We previously found that 25% of hindlimbs in our Twist1+/− animals were polydactylous with one ectopic preaxial digit (Firulli et al., 2005). By contrast 82% of Twist1CC/+ hindlimbs had one or two ectopic preaxial digits (Figs. 1M, Fig. 2Ac), whether on a 129, B6, or 129.B6 background. Thus while Twist1CC/+ and Twist1+/− heterozygote phenotypes are similar, the Twist1CC/+ phenotype is more penetrant.
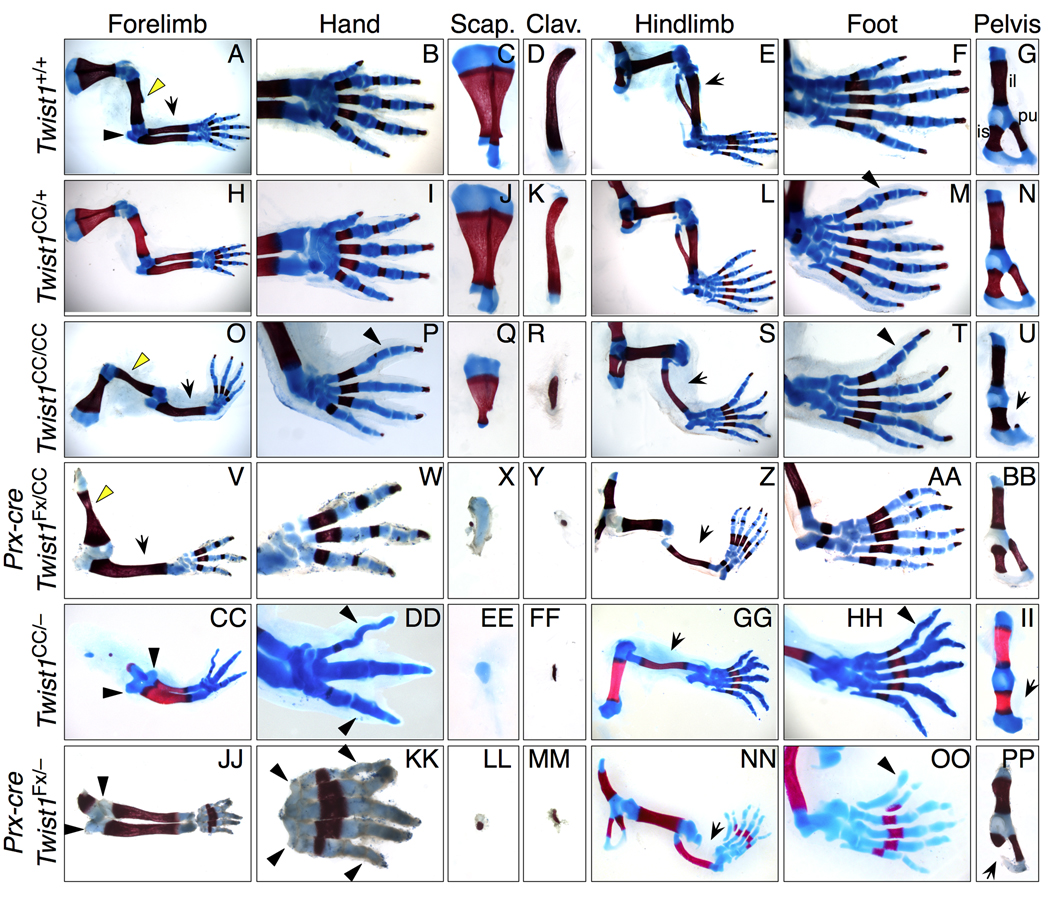
Figure 1. Limb and girdle skeletal phenotypes in Twist1CC compound mutant mice.
Skeletal preparations of E17.5-P2 limbs carrying combinations of Twist1CC, Twist1−, and Twist1Fx alleles show progressively more severe limb and girdle defects.
(A–G) Wild type, cartilage (blue) and bone (red).
(H–N) Twist1CC/+, hindlimb with preaxial polydactyly (M, arrowhead).
(O–U) Twist1CC/CC, forelimb with radial aplasia and absent deltoid tuberosity (O, dt, arrow and arrowhead), absent anterior digit and AP mirror symmetry (P, arrowhead), reduced scapula (Q) and clavicle (R); hindlimb with tibial aplasia (S, arrow), disorganized AP digit pattern (T, arrowhead) and hypoplastic pubis (U, arrow).
(V-BB) Prx1-cre; Twist1CC/Fx, forelimb with widened humerus and absent dt (V, arrowhead), radial aplasia (V, arrow), three digits (W) and severely hypoplastic scapula (X) and clavicle (Y); hindlimb with tibial aplasia (Z, arrow) and posteriorized AP digit pattern (AA).
(CC-II) Twist1CC/− forelimb with duplicated ulna and olecranon (CC, arrowheads), three digits in posteriorized mirror symmetric pattern (DD, arrowheads) and severely hypoplastic scapula (EE) and clavicle (FF); hindlimb with tibial aplasia (GG, arrow), disrupted AP digit pattern (HH, arrowhead) with bifurcation, and absent pubis (II, arrow).
(JJ-PP) Prx1-cre; Twist1Fx/− forelimb with duplicated ulna and olecranon (JJ, arrowheads), five digits with striking mirror symmetry (KK, arrowheads), severely hypoplastic scapula (LL) and clavicle (MM); hindlimb with tibial aplasia (NN, arrow), preaxial polydactyly with disorganized cartilage elements (OO, arrowhead), and hypoplastic ischium (PP, arrow).
Delays in ossification center formation due to reduced Twist1 activity were previously described (Bialek et al., 2004).
Scap: scapula, clav: clavicle, il: ilium, is: ischium, pu: pubis.
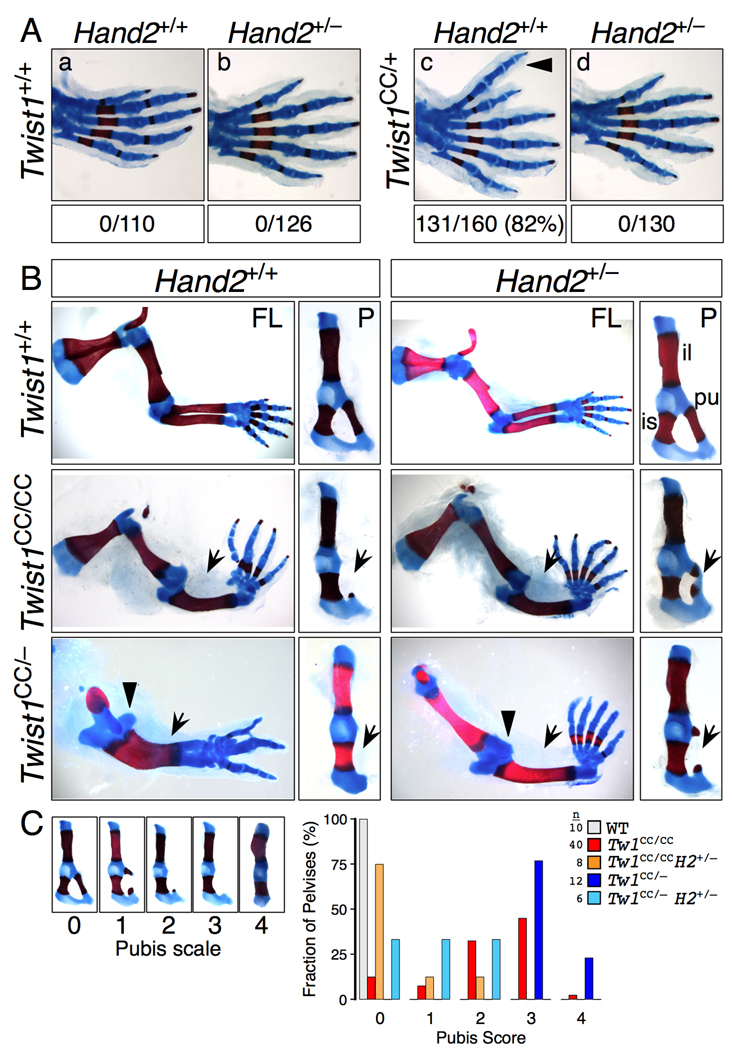
Figure 2. Genetic interactions between Twist1CC and Hand2.
(A) Twist1CC/+ hindlimb polydactyly is sensitive to Hand2 dosage. Hindlimbs of F1 progeny of Twist1CC/+; Hand2+/+ X Twist1+/+; Hand2+/− intercross scored for preaxial polydactyly (arrowhead) show complete suppression of polydactyly in double heterozygotes (p<0.001, chi squared).
(B) Twist1CC forelimb and pelvis phenotypes are sensitive to Hand2 dosage. Hand2 null allele was crossed onto compound Twist1 genotypes as indicated. Twist1CC/CC limbs with reduced Hand2 gene dosage (row 2) have less severe pubis hypoplasia (P columns, arrows) but still display radial aplasia (FL columns, arrows). Twist1CC/− limbs with reduced Hand2 gene dosage (row 3) resemble less severe Twist1CC/CC limbs, with duplicated ulnae replaced by radial aplasia (FL columns, arrows and arrowhead), increased digit number, and reduced severity of pubis hypoplasia (P columns, arrows). FL, forelimb; P, pelvis; il, ilium; pu, pubis; is, ischium.
(C) Twist1CC pubis defects were scored on a scale of increasing severity from 0 to 4 (left panels) and plotted as a percent fraction of total pelvises scored for each genotype. Reducing Hand2 gene dosage in Twist1CC/CC or Twist1CC/− pelvises significantly shifts pubis scores to lesser values (p<0.001 for each, Mann-Whitney).
Both Twist1CC/CC and Twist1CC/− mice died immediately after birth. They displayed gross limb, ventral body wall, skull and neural tube closure defects that were more severe in the Twist1CC/− animals (Figs. S1, Fig. S2). Skeletal preparations at E17.5 and E18.5 revealed multiple forelimb and hindlimb abnormalities, as well as defects in the scapula, clavicle and pelvic girdle (Fig. 1, Table 1). Twist1CC/CC forelimbs frequently had four digits with apparent posteriorized preaxial duplications (Fig. 1P). Their radii were hypoplastic or aplastic (Fig. 1O), and their humeruses lacked deltoid tuberosities (Fig. 1O). Twist1CC/− forelimbs had as few as 3 digits, with striking mirror symmetry along the anteroposterior axis (Fig. 1DD). They also had radial hypoplasia or aplasia, or an apparently duplicated ulna (Fig. 1CC). Twist1CC/− humeruses were severely hypoplastic (Fig. 1CC).
Table 1.
Twist1 and Twist1/Hand2 mutant limb and girdle phenotypes
| HUMERUS/FEMUR2 | RADIUS/TIBIA2 | SCAPULA/PUBIS2 | ||||||
|---|---|---|---|---|---|---|---|---|
| n1 | Hypoplastic | Absent | Hypoplastic | Absent | Dup. Ulna/Fibula | Hypoplastic | Absent | |
| Forelimb | ||||||||
| Twist1 +/+ | 10 | – | – | – | – | – | – | – |
| Twist1 CC/+ | 20 | – | – | – | – | – | – | – |
| Twist1 CC/CC | 42 | – | – | 19% | 81% | – | 100% | – |
| Twist1 CC/− | 12 | 100% | – | 25% | 8% | 67% | 100% | – |
| Twist1 CC/Fx, Prx-cre | 6 | 100% | – | – | 100% | – | 100% | – |
| Twist1 Fx/Fx, Prx-cre | 6 | 67% | 33% | – | 100% | 100% | 83% | 17% |
| Twist1 CC/CC, Hand2 +/− | 10 | – | – | 10% | 90% | – | 100% | – |
| Twist1 CC/−, Hand2 +/− | 7 | 71% | – | 14% | 72% | 14% | 100% | – |
| Hindlimb | ||||||||
| Twist1 +/+ | 10 | – | – | – | – | – | – | – |
| Twist1 CC/+ | 20 | – | – | – | – | – | – | – |
| Twist1 CC/CC | 37 | – | – | – | 100% | – | 87%4 | – |
| Twist1 CC/− | 11 | – | – | – | 100% | – | 100%5 | – |
| Twist1 CC/Fx, Prx-cre | 6 | – | – | – | 100% | – | 33% | – |
| Twist1 Fx/Fx, Prx-cre | 6 | – | – | 50% | 33% | – | 67% | – |
| Twist1 CC/CC, Hand2 +/− | 8 | – | – | – | 100% | – | 25% | – |
| Twist1 CC/−, Hand2 +/− | 5 | – | – | 40% | 60% | – | 67%6 | – |
| n1 | NUMBER OF DIGITS2 | Presence of digit 12 |
Normal A/P pattern2,3 |
||||
|---|---|---|---|---|---|---|---|
| 3 | 4 | 5 | 6 | ||||
| Forelimb | |||||||
| Twist1 +/+ | 10 | – | – | 100% | – | 100% | 100% |
| Twist1 CC/+ | 20 | – | – | 100% | – | 100% | 100% |
| Twist1 CC/CC | 42 | – | 64% | 36% | – | 10% | – |
| Twist1 CC/− | 12 | 50% | 25% | 25% | – | 9% | – |
| Twist1 CC/Fx, Prx-cre | 6 | 17% | 83% | – | – | – | – |
| Twist1 Fx/Fx, Prx-cre | 6 | – | 67% | 33% | – | – | – |
| Twist1 CC/CC, Hand2 +/− | 10 | – | 50% | 50% | – | – | – |
| Twist1 CC/−, Hand2 +/− | 7 | – | 20% | 70% | – | – | – |
| Hindlimb | |||||||
| Twist1 +/+ | 10 | – | – | 100% | – | 100% | 100% |
| Twist1 CC/+ | 20 | – | – | 25% | 75% | 100% | 25% |
| Twist1 CC/CC | 37 | – | – | 100% | – | – | – |
| Twist1 CC/− | 11 | – | 30% | 70% | – | – | – |
| Twist1 CC/Fx, Prx-cre | 6 | – | – | 100% | – | – | – |
| Twist1 Fx/Fx, Prx-cre | 6 | – | – | 50% | 50% | 33% | 17% |
| Twist1 CC/CC, Hand2 +/− | 8 | – | – | 100% | – | – | – |
| Twist1 CC/−, Hand2 +/− | 5 | – | 30% | 70% | – | – | – |
values represent individual limbs
– represents 0%
Abnormal patterns include ectopic preaxial digits (CC/+ hindlimbs), loss of asymmetry (CC/CC or CC/− hindlimbs), or digit reductions and/or fusions combined with mirror symmetry across the A/P axis (CC/CC, CC/Fx, Fx/Fx or CC/− forelimbs).
n=40 for pubis phenotypes
n=12 for pubis phenotypes
n=6 for pubis phenotypes
The difference between Twist1CC/CC and Twist1CC/− hindlimbs was less pronounced than for forelimbs. Hindlimbs had five or fewer digits, with digit I always replaced by a more posterior digit; other digit identities were hard to define (Fig. 1T,HH; Table 1). The tibia was always absent in Twist1CC/CC or Twist1CC/− animals, but the femurs appeared normal (Fig. 1S,GG).
Twist1CC/CC and Twist1CC/− shoulder and pelvic girdles were also abnormal, with Twist1CC/− skeletons again more severely affected. Scapular phenotypes ranged from moderate hypoplasia with reduced acromion processes (Fig. 1Q), to almost complete aplasia (Fig. 1EE). Clavicles were also reduced, primarily within the medial endochondral segment (Fig. 1R,FF). Within the pelvic girdle the phenotypes ranged from hypoplasia to loss of the pubic bone exclusively (Fig. 1U,II). Taken together these data reveal a complex set of limb and girdle phenotypes that affect elements at all proximodistal levels and that also impact anteroposterior identities. The progressive severity of the phenotypes across this Twist1 allelic series is also consistent with a correlated progressive reduction in Twist1 function.
Despite the severity of the Twist1CC/− phenotype, these results do not reflect complete loss of Twist1 activity in the limb bud. We therefore used a floxed conditional null allele of Twist1 (Twist1Fx) in combination with the Prx1-cre transgene driver (Logan et al., 2002), to attempt complete removal of Twist1 activity from the limb mesenchyme. We generated both Prx1-cre;Twist1CC/Fx and Prx1-cre;Twist1Fx/− embryos. Prx1-cre;Twist1CC/Fx limbs resemble Twist1CC/CC limbs (Fig. 1V–BB, Table 1), whereas Prx1-cre;Twist1Fx/− limbs resemble Twist1CC/− limbs (Fig. 1JJ-PP, S3; Table 1). Notably Prx1-cre;Twist1Fx/− limbs can have a duplicated ulna and handplate, and severely reduced scapula and clavicle (Fig. 1JJ,LL,MM). Interestingly, pelvic defects are restricted to the ischium, rather than the pubis as in Twist1CC/CC and Twist1CC/− mutants (Fig. 1PP). If deletion of the conditional Twist1 allele is complete, then Prx1-cre;Twist1CC/Fx should phenocopy Twist1CC/− in the limb. That it more closely resembles Twist1CC/CC, suggests that some residual Twist1 activity is present. Nonetheless these data provide evidence that the Twist1CC allele is not significantly neomorphic, as the Prx1-cre;Twist1Fx/− phenotype recapitulates the range of limb and girdle defects associated with the Twist1CC allele.
Twist1CC phenotypes are sensitive to Hand2 gene dosage
If Twist1CC is functioning in normal Twist1 pathways, it should display genetic interactions similar to Twist1. We therefore asked if reducing Hand2 dosage in the context of Twist1CC allelic combinations reduced their severity, as Hand2 null heterozygosity completely suppresses Twist1+/− hindlimb polydactyly (Firulli et al., 2005).
We crossed a Hand2 null allele onto Twist1CC mutant backgrounds and scored the resultant skeletal phenotypes (Fig. 2, Table 1). Several aspects of the Twist1CC phenotype were sensitive to Hand2 dosage. Twist1CC/+ hindlimb polydactyly (n=131/160; 82% of hindlimbs) is completely suppressed by lowering Hand2 dosage (Twist1CC/+;Hand2+/−: 0/130; 0%; Fig. 2A). The Twist1CC/−;Hand2+/− forelimbs resembled those of the Twist1CC/CC genotype, as often digit number increased, the radius was absent, the ulna was not duplicated, and the humerus was more complete (Fig. 2B, Table 1). The Twist1CC/−;Hand2+/− pubis was more complete than in Twist1CC/−; Hand2+/+ animals (Fig. 2B,C; Table 1), and clavicle length was increased (data not shown). However Hand2 null heterozygosity never completely rescued a Twist1CC/CC or Twist1CC/− phenotype. Taken together these data provide additional evidence that the Twist1CC allele acts in similar pathways to wild type Twist1. They furthermore indicate that balanced antagonism between Twist1 and Hand2 is critical for regulating development along the entire limb proximodistal axis.
Twist1 and Hand2 activities converge to regulate Shh signaling
Preaxial polydactyly is a hallmark of ectopic Shh signaling. Thus reducing Twist1 activity might lead to either ectopic Shh expression or activation of Shh signaling. We therefore examined the expression of Shh and the Shh targets in Twist1CC/+ and Twist1CC/+;Hand2+/− hindlimbs at E11-E12, when Twist1CC/+ ectopic anterior outgrowths are first observed (Fig. 3). In Twist1CC/+ hindlimbs we detected ectopic anterior Shh expression at the base of the autopod at low frequency (n=2/23 limbs, Fig. 3A,B), similar to a previous report that a small subset of Twist1+/− hindlimbs had ectopic Shh expression (O'Rourke et al., 2002). By contrast the Shh targets Gli1 and Hand2 in limb mesenchyme and the indirect target Fgf4 in the overlying AER were each induced in 50%–75% of limbs (n=74/106 limbs in total, Fig. 3A,B). This suggests that ectopic Shh expression is induced in most Twist1CC/+ hindlimbs, but at a level that we cannot detect. We never observed ectopic gene expression in Twist1CC/+;Hand2+/− limbs (n=0/27 limbs; Fig. 3B), consistent with their normal skeletal morphology at later ages. The similar frequencies of ectopic Shh target expression and hindlimb polydactyly (82%, Fig. 2A) support the idea that Twist1 and Hand2 activities converge at the level of Shh signaling, and possibly upstream of ectopic Shh expression.
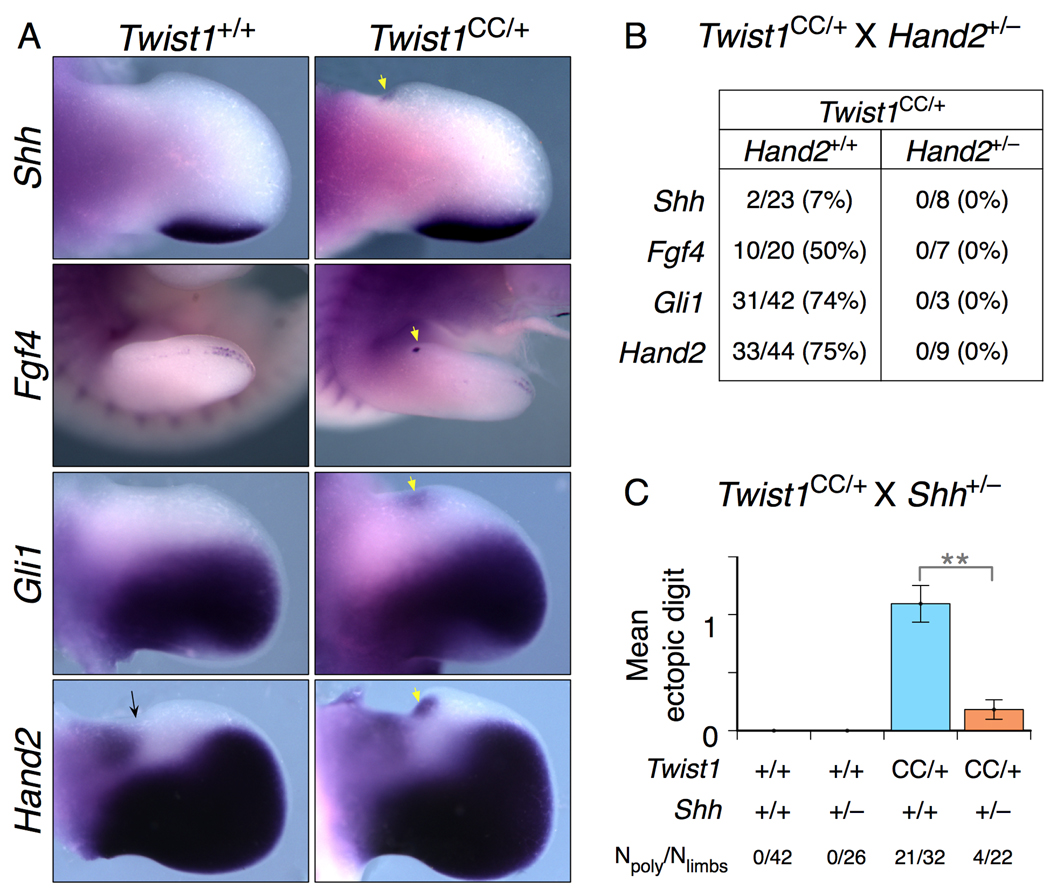
Figure 3. Twist1 and Hand2 activities in anterior limb bud converge at the regulation of Shh activity.
(A) E11.5 Twist1CC/+ hindlimb buds displaying ectopic anterior RNA expression of Shh and the Shh target genes Fgf4, Gli1 and Hand2 (white arrows). Note presence of proximal anterior Hand2 domain (black arrow).
(B) Reducing Hand2 gene dosage suppresses ectopic anterior gene expression in Twist1CC/+ embryos. Frequency of ectopic anterior gene expression detected in E11.0 – E13.0 Twist1CC/+; Hand2+/+ and Twist1CC/+; Hand2+/− hindlimbs (N=limbs with ectopic expression/limbs examined).
(C) Twist1CC/+; Shh+/− progeny of Twist1CC/+ X Shh+/− intercross show significantly reduced frequency and severity of hindlimb polydactyly, indicating the Twist1CC/+hindlimb polydactyly is Shh-dependent. **: p<0.001. (N=limbs with polydactyly/limbs examined).
To test this directly, we intercrossed Twist1CC/+ and Shh+/− animals, and scored the frequency of hindlimb polydactyly in their offspring (Fig. 3C). 21 of 32 Twist1CC/+ hindlimbs had preaxial polydactyly, while a significantly reduced 4 of 22 Twist1CC/+;Shh+/− hindlimbs (p<0.0001) were polydactylous. Reducing Shh dosage also reduced the severity of the polydactyly: none of the double mutant limbs had an ectopic digit 2, compared to two-thirds of the Twist1 single mutants. These results provide evidence that high levels of Twist1 activity are required primarily to repress Shh expression in anterior limb mesenchyme.
Signaling center defects in Twist1CC compound mutant limb buds
To better understand the changes underlying the more complex skeletal phenotypes of the compound Twist1 mutants, we analyzed limb buds during the time when pattern is established. From E9.5 Twist1CC/CC and Twist1CC/− forelimb buds were smaller along the anteroposterior axis, and tapered distally compared to controls. Subsequently at E11.5 they were narrowed and curved anteriorly (Fig. S3A). By E12.5, the anterior mesenchyme had expanded significantly, although more in Twist1CC/CC than Twist1CC/− limbs, and curved dramatically anteriorly while the zeugopod remained narrow (Fig. S3A,C). Hindlimb buds resembled forelimb buds, displaying an almost 90 degree turn towards the anterior by E12.5 (Fig. S3A,C).
Molecular analyses of signaling center marker expression revealed that prior to substantial limb outgrowth, from 20 to 25 somite stages, the anteroposterior extent of Fgf8 expression in the AER was reduced (Figs. 4Af,k). As forelimb outgrowth proceeded, this progressed into an absence of posterior Fgf8 expression (Figs. 4Ag,l). As the autopod curved anteriorly, Fgf8 expression expanded anteriorly (Figs. 4Ah,m). In the hindlimb the early absence of posterior expression was less pronounced (Figs. 4Ai,n), but the anterior Fgf8 boundary expanded and persisted longer than normal (Figs. 4Aj,o).
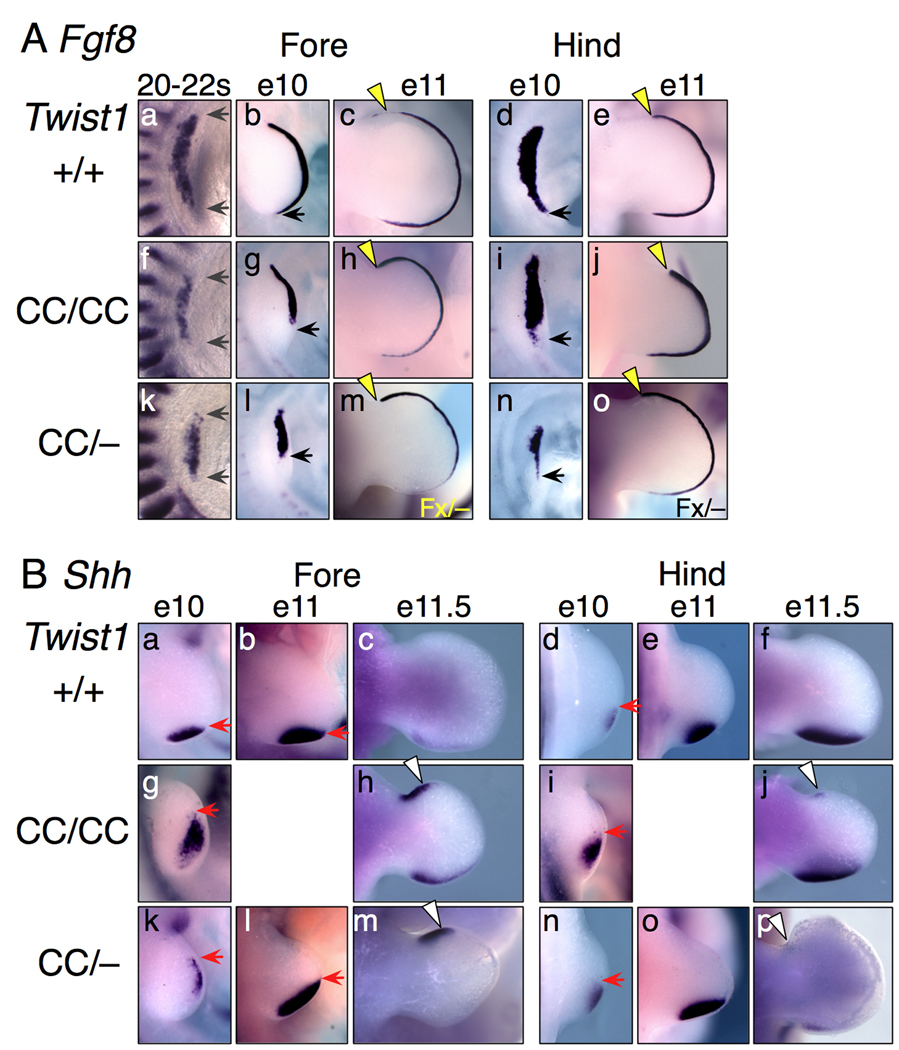
Figure 4. Fgf8 and Shh expression defects in Twist1 mutant limbs.
(A) Fgf8 RNA expression in whole-mount limb buds. At 20–22 somites, the AP length of Fgf8 expression is significantly narrowed (gray arrows) in Twist1CC mutant forelimb buds. At E10, posterior Fgf8 expression is absent (forelimb) or reduced (hindlimb) in the small Twist1CC mutant buds (black arrows). At E11 Twist1CC/CC and Prx1-cre; Twist1Fx/− (Fx/−; panels m,o) limb buds have increased anterior Fgf8 expression extending to the anterior limb boundary (yellow arrowheads).
(B) Shh RNA expression in whole-mount limb buds. At E10, Shh expression is shifted anteriorly and distally (red arrows) and is less organized in Twist1 mutant limb buds. By E11, Shh expression encompasses the entire posterior and distal regions of the small Twist1CC/− forelimb bud (red arrows), and encompasses much of the posterior region in hindlimb buds. Subsequently, at E11.5, a strong ectopic anterior Shh domain is detected in the mutant forelimbs, with a lesser ectopic domain in hindlimbs (white arrowheads).
Shh expression in the mutant embryos was dynamic (Fig. 4B). Early forelimb bud expression shifted distally, compared to normal posterior mesenchyme expression, and Shh+ cells were slightly dispersed (Figs. 4Ba,g). In early hindlimbs, expression was slightly elevated, and possibly anteriorly shifted, but not as severely as in mutant forelimbs (Figs. 4Bi,n). Shh expression then shifted to the posterior margin of the narrow, tapered limb buds (Figs. 4Bl,o). As the autopod expanded, an additional anterior Shh domain appeared. This domain was very robust in mutant forelimbs, but weak in mutant hindlimbs (Figs. 4Bh,f,m,p). Thus Twist1 compound mutants display complex multiphasic changes in gene expression, with posterior effects followed by anterior ones.
FGF signaling is reduced in Twist1 mutant limbs
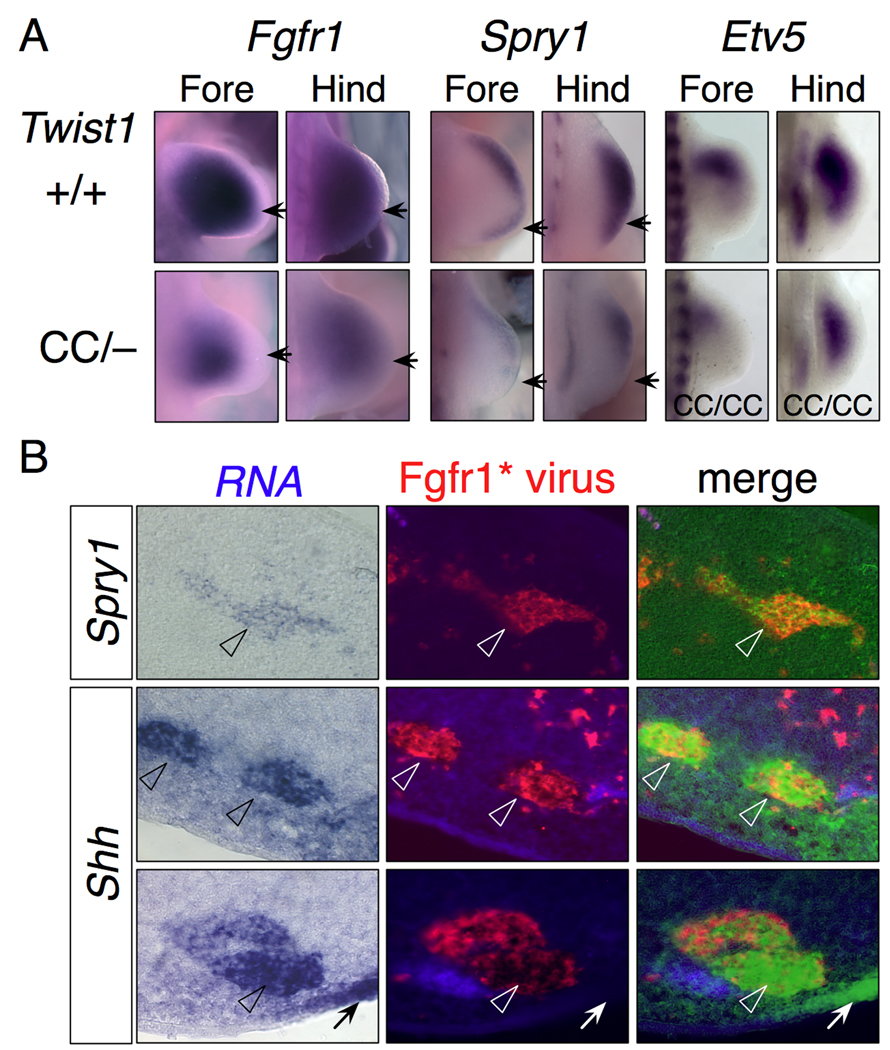
The early reduction in the size of the Fgf8 expression domain in the AER suggested that overall FGF signaling might be reduced in the Twist1CC compound mutants. This would be consistent with previous reports that Fgfr1 expression was severely reduced in Twist1−/− mesenchyme (O'Rourke et al., 2002; Zuniga et al., 2002). We therefore examined expression of Fgfr1, and the FGF targets Spry1 (Mason et al., 2006) and Etv5 (Mao et al., 2009; Zhang et al., 2009). Fgfr1 expression was reduced, but present, in both forelimb and hindlimb mutant mesenchyme (Fig. 5A). Spry1 was expressed in the subridge mesenchyme in wild type limbs, but was absent from posterior mesenchyme and reduced anteriorly and distally in mutant limbs (Fig. 5A). Similarly, Etv5 expression was reduced, most strongly in forelimb posterior mesenchyme. Thus there is reduced but active FGF signaling in much of the Twist1 compound mutant mesenchyme.
Figure 5. FGF signaling is compromised in Twist1 mutants.
(A) Fgfr1 and Spry1 expression in wild type and Twist1CC/− limb buds. Fgfr1 expression is reduced but not absent in mutant limb buds, particularly in the distal and posterior margins (arrows). Spry1 expression is substantially reduced in Twist1CC/− limb buds with less expression in the posterior (arrow), and overall expression in hindlimb higher than forelimb (arrow). Etv5 expression is also significantly reduced in Twist1CC/CC limb buds.
(B) Shh is induced cell-autonomously by FGF signaling. Sections of limb buds infected at stage 20 with a replication-defective retrovirus expressing constitutively active Fgfr1, harvested 36 h post-infection, and sequentially processed by in situ hybridization for target gene expression and immunostained for retroviral infection.
(Row 1) Induction of Spry1 expression in anterior limb mesenchyme. Overlay of bright field image (false colored green; original, left panel) and Fgfr1* virus show a large clone of Spry1+ cells coincident with the infected cell clone (arrowheads).
(Rows 2,3) Induction of Shh expression in posterior limb mesenchyme. Overlay of bright field (false-colored green; original, left) and Fgfr1* virus show Shh+ cells coincident with the infected cell clones (arrowheads). Endogenous Shh domain is marked in Row 3 (arrow).
It is striking that there is no Shh expression in the posterior of mutant forelimb buds, even though there is posterior Fgfr1 expression. While it is known that FGF signals are required for Shh expression, it is unclear whether Shh is induced autonomously in cells that transduce the FGF signal, or if FGF-dependent secondary signals are required. If Shh is induced autonomously, then the lack of Fgf8 expression in the overlying posterior ectoderm might explain the lack of posterior Shh expression. To test how directly Shh is induced, we infected small groups of cells in chick limb buds with a replication defective virus expressing constitutively active Fgfr1 (Fgfr1*(Liu et al., 2001)) and assayed for Shh expression in infected cells. Sections were processed sequentially by in situ hybridization for Spry1 or Shh expression, then immunostained for retroviral gag expression to identify infected cells. The Fgfr1* virus induced Spry1 expression in anterior, central and posterior limb mesenchyme, whereas it induced Shh expression only near the posterior margin (Fig. 5B). This restriction is consistent with other observations that the competence for Shh expression is limited to posterior mesenchyme by additional inputs that include both cell-autonomous transcription factors and non-autonomous signals from the overlying ectoderm (Zeller et al., 2009). In image overlays comparing the distribution of infected cells with the induced mRNA, there was excellent concordance between retroviral infection and induced patches of either Spry1 or Shh mRNA (Fig. 5B, Fig. S4A,B). In some experiments the posterior AER was removed prior to infection to eliminate the major endogenous source of posterior FGF activity (Laufer et al., 1994), and Shh expression was still detected in some infected cells (Fig. S4C). These data provide evidence that Shh, like Spry1, is expressed by cells that receive FGF signals. While these data imply that no intermediary signal downstream of FGF is involved, they do not obviate the need for additional factors to impart competence for Shh expression to limb mesenchymal cells.
Twist1 regulates transcription factor expression
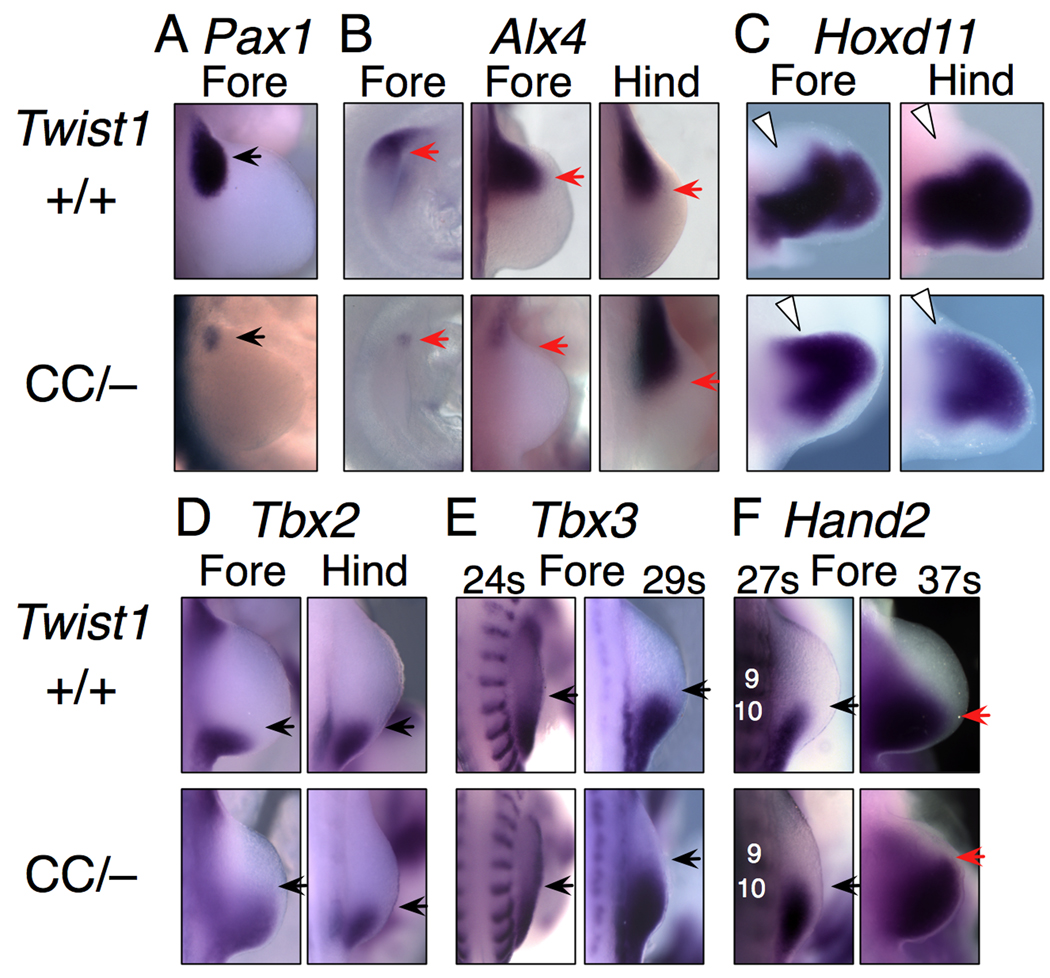
Null mutant mice for other transcription factors, notably within the Pax and Alx gene families, have limb and girdle cartilage phenotypes similar to the Twist1CC mutants (Balling et al., 1988; Kuijper et al., 2005). While none is as severe as the Twist1CC mutant phenotype, together they share many aspects, including preaxial polydactyly, anterior element aplasia and girdle defects. Previous reports showed that several of the genes are downregulated in Twist1−/− mutants. Pax1, which is required for scapula development, was also significantly reduced in Twist1CC mutants (Fig. 6A,Fig. S3). We also found that Alx4, which is normally expressed in anterior limb mesenchyme, is markedly reduced in forelimb buds of Twist1CC mutants, but is affected only mildly if at all in hindlimb buds (Fig. 6B,Fig. S3). These results are consistent with the idea that at least part of the Twist1CC phenotype is due to misregulation of a network of downstream transcription factors.
Figure 6. Regulation of transcription factor mRNA expression by Twist1.
(A) Anterior-proximal Pax1 is strongly downregulated (arrow) in mutant forelimb buds.
(B) Alx4 is substantially downregulated at E9.25 (left column) and excluded from the forelimb mesenchyme by E10 (center), but is grossly unaffected in hindlimbs.
(C) Hoxd11 is induced in anterior-proximal mutant limb buds at E11 (white arrowheads).
(D) Anterior boundary of Tbx2 expression at e10 (black arrow) is shifted anteriorly in mutant forelimb, but not hindlimb.
(E) The anterior boundary of Tbx3 is unchanged in Twist1CC/− forelimb at 24 somites, but is shifted anteriorly at 29 somites (E9.5–10).
(F) The anterior boundary of Hand2 is unchanged in Twist1CC/− forelimb at 27 somites (black arrows), but is shifted anteriorly (red arrows) at 37 somites (E10.5). Somites 9 and 10 are indicated.
We next examined whether the expression boundaries of genes implicated in positioning Shh expression in the early limb bud, Tbx2, Tbx3 and Hand2 (Davenport et al., 2003; te Welscher et al., 2002), might be altered in Twist1CC mutants. As Shh expression is initiated, Tbx3 and Hand2 are expressed in approximately the posterior one-third of normal forelimb buds. In Twist1CC/− forelimbs both the Tbx3 and Hand2 expression boundaries are apparently unaffected through the 27 somite stage, prior to the onset of Shh expression (Fig. 6E,F). These borders do, however, shift anteriorly after Shh expression is initiated (Fig. 6E,F). The anterior expression border of Tbx2 is also extended anteriorly in the Twist1CC mutants after Shh expression begins (Fig. 6D). As each of these genes is responsive to Shh signaling (Lu et al., 2009), these changes are likely secondary to the shifting of the Shh expression domain.
We also asked whether expression of more downstream effectors of limb patterning, such as HoxA or HoxD cluster genes was normal in Twist1CC mutants. HoxD11, which is normally expressed in posterior and distal limb mesenchyme, is expressed ectopically in anterior proximal forelimb and hindlimb buds of E10.5 Twist1CC compound mutant embryos (Fig. 6C, Fig. S3). By contrast HoxA11 is expressed normally in a zeugopodal domain in the mutants (Fig. S3). These results are consistent with posteriorization of the anterior mesenchyme, but normal proximodostal specification of the zeugopod.
Twist1 interacts genetically with Etv genes
Our molecular analyses and previous studies highlight the interaction of Twist1 function with FGF signaling. Interestingly, we detect ectopic anterior Shh expression concomitant with reduced Etv5 expression, which was recently identified as a negative regulator of anterior and distal limb Shh expression downstream of FGF signaling (Mao et al., 2009; Zhang et al., 2009). Thus Twist1 and Etv genes might function in the same genetic pathway. To test this, we crossed Twist1CC/+ mice with Etv4 and Etv5 null mutant mice (Livet et al., 2002; Lu et al., 2009) to progressively reduce Etv gene dosage, as Etv4 and Etv5 have overlapping functions (Lu et al., 2009; Mao et al., 2009; Zhang et al., 2009). Mice carrying up to two Etv4 null alleles, one Etv5 null allele and one Twist1CC allele were viable, and their limb and girdle phenotypes were scored between E17.5 and P14.
Reducing Etv gene dosage had a dramatic effect on the distal forelimb skeleton (Fig. 7, Fig. S5; Table 2). Phenotypes ranged from simple preaxial polydactyly in 17% of Etv4+/−;Twist1CC/+ or Etv5+/−;Twist1CC/+ limbs, to radial aplasia and polydactyly with up to 7 forelimb digits in Etv4−/−;Etv5+/−;Twist1CC/+ limbs (Fig. 7U,V; Table 2). Hindlimb phenotypes were also enhanced, with the strongest phenotypes including tibial aplasia and polydactyly (Fig. 7W,X). Strikingly we did not observe any defects in more proximal limb elements or the girdles. Removing up to 3 copies of the Etv genes in the context of two wild type Twist1 alleles had almost no effect on skeletal pattern. The only limb defect we observed was a small preaxial cartilage digit that formed in one forelimb of a Etv4−/−;Etv5+/− pup (Fig. 7F). These data provide evidence of strong dosage-sensitive interactions between Twist1 and Etv genes to pattern the more distal limb skeleton.
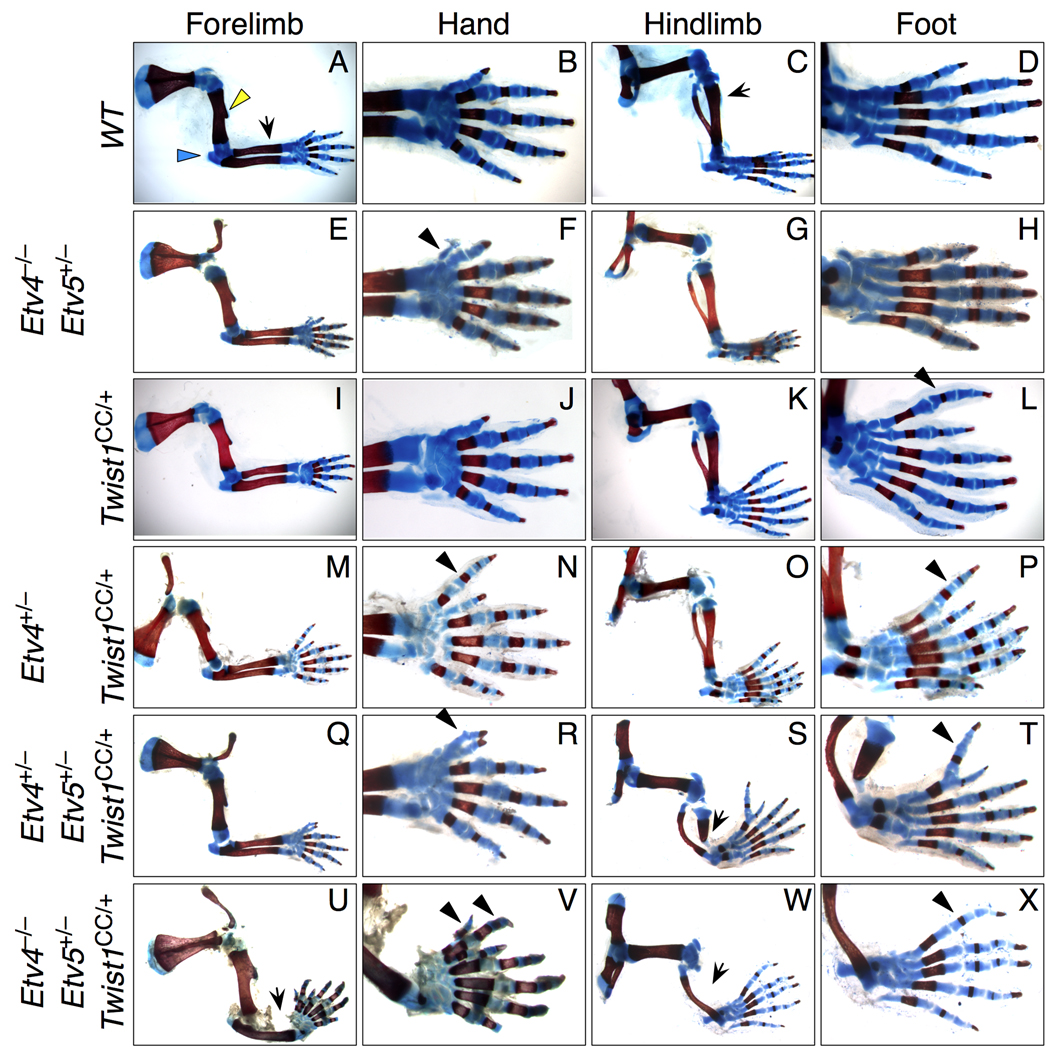
Figure 7. Twist1 interacts genetically with Etv family genes.
Limb skeletal patterns in progeny of Twist1CC/+ intercrosses with Etv4+/− and Etv5+/− mice.
(A–D) Wild-type morphology, duplicated from Fig. 1.
(E–H) A small preaxial digit is occasionally present in Etv4−/−Etv5+/− forelimbs (arrowhead).
(I–L) Twist1CC/+ hindlimb preaxial polydactyly, duplicated from Fig. 1.
(M–P) Reducing Etv4 or Etv5 (see also Table 2) dosage in Twist1CC/+ mutants causes posteriorized preaxial polydactyly of both forelimb and hindlimb (arrowheads).
(Q–R) Twist1CC/+ mice lacking two Etv4 or Etv5 alleles have enhanced limb phenotypes, with preaxial polydactyly (arrowheads) and tibial hypoplasia (arrow).
(U–X) Removing three Etv gene copies phenocopies Twist1CC/CC limbs, with radial and tibial aplasia (arrows) and preaxial polydactyly (arrowheads). Note the humerus, including the deltoid tuberosity (yellow arrow, A), and proximal elements appear normal.
Table 2.
Compound Twist1CC and Etv4/Etv5 mutant limb and girdle phenotypes
| Genotype | HUMERUS/FEMUR1 | RADIUS/TIBIA1 | SCAPULA/PUBIS1 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Twist1 (C) | Etv4 (4) | Etv5 (5) | Abbreviation | n limbs | Hypoplastic | Hypoplastic | Absent | Dup. Ulna/Fibula | Hypoplastic | Absent |
| Forelimb | ||||||||||
| +/+ | +/+ | +/+ | Wild-type | 8 | – | –– | – | – | – | – |
| +/+ | +/− | +/+ | 4 | 8 | – | –– | – | – | – | – |
| +/+ | +/+ | +/− | 5 | 8 | – | –– | – | – | – | – |
| +/+ | −/− | +/+ | 44 | 8 | – | –– | – | – | – | – |
| +/+ | −/− | +/− | 445 | 4 | – | –– | – | – | – | – |
| CC/+ | +/+ | +/+ | C | 22 | – | –– | – | – | – | – |
| CC/+ | +/− | +/+ | C4 | 22 | – | –– | – | – | – | – |
| CC/+ | +/+ | +/− | C5 | 8 | – | –– | – | – | – | – |
| CC/+ | −/− | +/+ | C44 | 14 | – | –– | – | – | – | – |
| CC/+ | +/− | +/− | C45 | 10 | – | 13% | – | – | – | – |
| CC/+ | −/− | +/− | C445 | 4 | – | 25% | 25% | – | – | – |
| Hindlimb | ||||||||||
| +/+ | +/+ | +/+ | Wild-type | 8 | – | –– | – | – | – | – |
| +/+ | +/− | +/+ | 4 | 8 | – | –– | – | – | – | – |
| +/+ | +/+ | +/− | 5 | 8 | – | –– | – | – | – | – |
| +/+ | −/− | +/+ | 44 | 8 | – | –– | – | – | – | – |
| +/+ | −/− | +/− | 445 | 4 | – | –– | – | – | – | – |
| CC/+ | +/+ | +/+ | C | 22 | – | –– | – | – | – | – |
| CC/+ | +/− | +/+ | C4 | 22 | – | –– | – | – | – | – |
| CC/+ | +/+ | +/− | C5 | 8 | – | 25% | – | – | – | – |
| CC/+ | −/− | +/+ | C44 | 14 | – | 29% | 7% | – | – | – |
| CC/+ | +/− | +/− | C45 | 10 | – | 50% | 50% | – | – | – |
| CC/+ | −/− | +/− | C445 | 4 | – | – | 100% | – | – | – |
| NUMBER OF DIGITS1 | Presence of digit 1 |
Normal A/P pattern1 |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 4 | 5 | 6 | 7 | |||||||
| Forelimb | ||||||||||
| +/+ | +/+ | +/+ | Wild-type | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | +/− | +/+ | 4 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | +/+ | +/− | 5 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | −/− | +/+ | 44 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | −/− | +/− | 445 | 4 | – | 75% | 25% | – | 100% | 75% |
| CC/+ | +/+ | +/+ | C | 22 | – | 100% | – | – | 100% | 100% |
| CC/+ | +/− | +/+ | C4 | 22 | – | 82% | 18% | – | 100% | 82% |
| CC/+ | +/+ | +/− | C5 | 8 | – | 87% | 13% | – | 100% | 87% |
| CC/+ | −/− | +/+ | C44 | 14 | – | 93% | 7% | – | 100% | 93% |
| CC/+ | +/− | +/− | C45 | 10 | – | 30% | 70% | 30% | 100% | 30% |
| CC/+ | −/− | +/− | C445 | 4 | – | – | 50% | 50% | 100% | – |
| Hindlimb | ||||||||||
| +/+ | +/+ | +/+ | Wild-type | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | +/− | +/+ | 4 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | +/+ | +/− | 5 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | −/− | +/+ | 44 | 8 | – | 100% | – | – | 100% | 100% |
| +/+ | −/− | +/− | 445 | 4 | – | 100% | – | – | 100% | 100% |
| CC/+ | +/+ | +/+ | C | 22 | – | 14% | 86% | – | 100% | 14% |
| CC/+ | +/− | +/+ | C4 | 22 | – | 9% | 86% | 5% | 100% | 9% |
| CC/+ | +/+ | +/− | C5 | 8 | – | – | 100% | – | 100% | – |
| CC/+ | −/− | +/+ | C44 | 14 | – | 7% | 86% | 7% | 100% | 7% |
| CC/+ | +/− | +/− | C45 | 10 | – | – | 70% | 30% | 90% | – |
| CC/+ | −/− | +/− | C445 | 4 | – | – | 100% | – | 50% | – |
– represents 0%
Discussion
Previous studies aimed at understanding the role of Twist1 during limb development have been hampered by impaired limb bud outgrowth and early embryonic lethality of Twist1−/− embryos (Chen and Behringer, 1995; O'Rourke et al., 2002; Zuniga et al., 2002). Using an array of Twist1 mutant alleles we generated an allelic series, which reveals that Twist1 has a range of previously unidentified roles in limb and limb girdle development. We demonstrate that Twist1 regulates limb signaling center function through precise positioning of the ZPA and both inhibition and maintenance of Shh expression, as well as influencing the anterior-posterior extent of the AER. Furthermore, we provide evidence that Twist1 acts through a gene network that includes Hand2 and members of the Etv and aristaless families. Unexpectedly, we find that anterior limb skeletal morphology is most strongly affected despite severe defects in the posterior of Twist1 mutant limb buds. We present a model whereby discrete thresholds of Twist1 activity contribute to early limb bud patterning, and suggest how particular combinations of skeletal defects result from differing Twist1 levels.
Our results implicate Twist1 activity as critical for repressing, positioning and maintaining Shh expression in the developing limb bud. Progressively reducing Twist1 activity, as in the transition from Twist1CC/+ to Twist1CC/− embryos, leads to progressively increasing amounts of ectopic Shh expression in the anterior of the bud. An anterior Shh domain was previously observed at low frequency in Twist1+/− hindlimbs (Bourgeois et al., 1998). Twist1CC/+ hindlimbs display ectopic Shh expression and preaxial polydactyly at higher frequency, which is suppressed by reducing either Shh or Hand2 dosage. Hand2 is expressed strongly in posterior limb mesenchyme, and is required for Shh expression (Charite et al., 2000). The ectopic Shh expression we observe in Twist1 mutants initiates at the far anterior margin of the limb bud, yet it is extremely sensitive to Hand2 activity. This raises the question of how Hand2 might act at such a long range. One possibility is that the small, weak domain of Hand2 expressed at the base of the anterior autopod mediates this activity; perhaps this level of Hand2 expression is normally insufficient to promote Shh expression, but can do so if Twist1 activity is modestly reduced.
There are interesting differences between the consequences of substantially reducing and completely removing Twist1 activity. In Twist1CC/CC or Twist1CC/− forelimb and hindlimb buds, there is a large ectopic Shh domain. This was unexpected, as there is no ectopic domain in Twist1−/− limb buds. One possible explanation for this is that null mutants die before ectopic Shh expression is initiated. However we have observed ectopic Shh as early as e10.0 in Twist1CC/− forelimbs, when null embryos are viable (Chen and Behringer, 1995). Thus there is a sharp threshold of Twist1 activity that can support anterior Shh expression. Another activity threshold is apparent in the posterior of Twist1CC/CC or Twist1CC/− limbs, where reducing Twist1 activity shifts the normal Shh domain anteriorly and distally and expression is robust. A similar shift occurs in Twist1−/− embryos, however there Shh expression is weak (O'Rourke et al., 2002; Zuniga et al., 2002). This shift might be caused by anteriorly repositioning the field of cells competent to express Shh. Interestingly we found that the expression of positive factors such as Hand2 did not shift prior to the onset of Shh expression, which suggests that Twist1 is acting more directly as an inhibitor of Shh expression.
These changes in Shh expression can be attributed at least in part to alterations in FGF signaling from the AER to the underlying mesenchyme. Previous studies established that Twist1 expression is downstream of FGF signaling from the AER, and is also required to maintain FGF receptor expression in the limb mesenchyme (Isaac et al., 2000; O'Rourke et al., 2002; Zuniga et al., 2002). Consistent with these results, we find that partial reductions in Twist1 activity lead to reduced FGF receptor expression and FGF pathway activity. There is also a secondary breakdown in signaling from the limb mesenchyme to the ectoderm that leads to a reduced AER, even before Shh expression is initiated. We also found that Shh expression is apparently induced autonomously in cells that receive FGF signals, without requiring a secondary signal downstream of FGF. This contrasts with the reciprocal signaling pathway from Shh to the overlying AER, which involves intermediary Gremlin antagonism of BMP activity (Zeller et al., 2009). Thus in Twist1 mutants the anterior shift of the posterior AER border and reduced levels of FGF signaling activity together likely contribute to the anterior shift in the posterior Shh domain.
Recent studies also implicate FGF signaling as a major antagonist of Shh expression in anterior limb mesenchyme, with a critical role played by FGF targets of the Etv gene family (Mao et al., 2009; Zhang et al., 2009). Consistent with these observations Etv gene expression is lower when Twist1 activity is strongly reduced. By contrast in Etv conditional mutant limbs, Twist1 expression and FGF signaling activity are unaffected, suggesting that Etv function lies downstream and possibly parallel to that of Twist1 (Zhang et al., 2009). We find that Twist1CC/+;Etv mutant combinations cause preaxial polydactyly plus anterior zeugopod defects, each of which we detect in Twist1CC/CC or Twist1CC/− mutants. This synergistic interaction provides strong evidence in support of Etv genes mediating Twist1 activity. However in Etv mutants, the zeugopod is either unaffected, or both elements are modestly reduced (Mao et al., 2009; Zhang et al., 2009). Furthermore in Twist1 mutants we observe as few as three digits in the forelimb autopod, consistent with a shortened AER, while Etv mutants have a normal or extended AER and do not have reduced digit numbers (Mao et al., 2009; Zhang et al., 2009). Thus changes in Etv activity account for only part of the Twist1 mutant phenotype.
Our results and previous observations show that Twist1 is required for the expression of Alx3 and Alx4, members of the aristaless gene family (O'Rourke et al., 2002). Twist1 might be required for expression of another family member, Cart1, but we were unable to detect any obvious differences in Cart1 expression in Twist1 mutants (not shown), because Cart1 is at low levels in the limb mesenchyme (Beverdam and Meijlink, 2001). Single and compound loss-of-function mutants for Alx4, Alx3 and Cart1 cause progressive limb phenotypes that overlap with the progression of Twist1 mutant phenotypes. These range from weak preaxial polydactyly and missing deltoid tuberosities in single mutants to extreme polydactyly with tibial hypoplasia or aplasia in compound mutants (Beverdam et al., 2001; Qu et al., 1999). They can also exhibit shortened clavicles, abnormal scapulas, and missing or hypoplastic pubic bones (Kuijper et al., 2005). Thus changes in aristaless-family gene activity might contribute to both limb and girdle aspects of the Twist1 mutant phenotype.
Interestingly aristaless gene expression is dependent on Twist1 primarily in the forelimb. While expansion of Hand2 into the anterior limb can repress Alx4 expression, we detect loss of Alx4 expression prior to any shift in Hand2 expression. This suggests that Shh derepression in Twist1 mutant forelimbs is affected by both aristaless and Etv activity, while in hindlimbs it is due mostly to reduced Etv function. This would be consistent with the relatively stronger skeletal phenotypes such as the ulnar duplications in Twist1 mutant forelimbs.
While Twist1 clearly modulates Shh expression and FGF signaling from the AER, its activity is not restricted to this distal mesenchyme patterning system. The limb girdles develop independent of the ZPA or AER, as neither pelvis nor scapula is affected in Shh−/− or Fgf4−/−;Fgf8−/− mutant animals (Boulet et al., 2004; Chiang et al., 2001; Sun et al., 2002). Furthermore the antagonism between Hand2 and Twist1 is not unique to distal limb mesenchyme, as the Twist1 pubic bone and clavicle defects are sensitive to Hand2 dosage.
Our studies show that Twist1 levels are critically important for exerting different aspects of its function (Fig. 8). High Twist1 activity is required to repress Shh expression in the anterior limb bud, primarily through FGF signaling and Etv function and secondarily via aristaless gene activity. It also involves competitive antagonism with Hand2. Thus in limbs with only modestly reduced Twist1 activity, such as in Twist1CC/+ embryos, a late domain of ectopic Shh expression induces weak preaxial polydactyly.
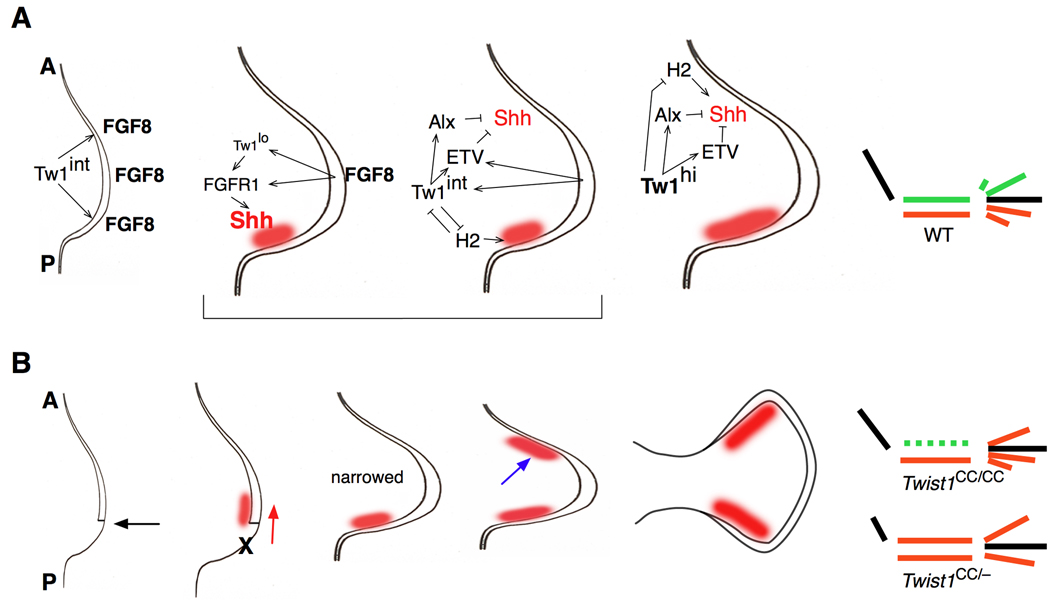
Figure 8. Model of Twist1 function in early limb development.
A. Summary of regulatory relationships between Twist1 and downstream genes in the limb bud.
1st panel: Intermediate levels of Twist1 (Tw1int) are required for Fgf8 expression in the AER before Shh expression begins. 2nd and 3rd panels: FGF signals from the AER induce Twist1 expression which itself maintains FGF receptor (FGFR1) expression. A minimal level of Twist1 activity (Tw1lo) is required for FGF to induce robust Shh expression (red area). At the same time, intermediate Twist1 levels act through aristaless (Alx) and Etv family genes to suppress Shh expression in the anterior limb bud, while also maintaining Shh expression at the posterior of the bud by antagonizing Hand2 activity. 4th panel: High levels of Twist1 activity (Tw1hi) suppress Shh expression in the anterior limb bud through the same gene network. Diagram: The result is a normally patterned limb, with an intact stylopod (black), two zeugopod bones, and five digits in the autopod. Anterior elements in green, posterior ones in red. A: anterior, P: posterior.
B. Developmental progression of Twist1CC/CC and Twist1CC/− limb phenotypes.
1st panel: When Twist1 activity is substantially reduced the AER is shortened, most importantly at the posterior (black arrow). 2nd panel: the reduced range and intensity of FGF signaling result in an anterior/distal shift in Shh expression (red arrow), while the posterior mesenchyme does not grow out (X). 3rd panel: Failed outgrowth of the limb margins leads to a narrower bud, with Shh at the new posterior margin. 4th panel: Derepressing Shh expression leads to an ectopic anterior Shh domain (blue arrow), with timing and magnitude proportional to reduction in Twist1 activity. 5th: This ectopic Shh extends the anterior AER, thus expanding the distal bud.
Diagram: In Twist1CC/CC mutants, the smaller bud supports only one zeugopod condensation, which is specified as posterior by Shh. In the forelimb autopod fewer elements condense, although this is partially rescued by anterior Shh, which posteriorizes these digits. In Twist1CC/− mutants, the initial reduction in the bud is more severe, but ectopic Shh initiates earlier. This expands the presumptive zeugopod region allowing two condensations, along with posteriorization of the anterior element, resulting in a duplicated ulna. Anterior elements in the autopod are also posteriorized.
In Twist1CC/CC or Twist1CC/− embryos that have less Twist1 activity, the molecular defects are more severe. Prior to the onset of Shh expression the anterior-posterior extent of the AER is reduced. This leads to an anterior shift in the strongest region of FGF signaling, and reduction in the antagonistic activity between Twist1/Etv and Hand2, thereby resulting in an anterior shift in initial Shh expression. Furthermore because the AER is smaller, the most posterior (and probably anterior) limb mesenchyme regions have reduced outgrowth. There might be increased apoptosis in posterior mesenchyme, as occurs in Twist1 null limb buds (O'Rourke et al., 2002; Zuniga et al., 2002), although we have not consistently detected this in Twist1CC/CC or Twist1CC/− limb buds (not shown). As the limb grows out, the Shh domain becomes restricted to the posterior margin of the now smaller bud. Subsequently on the anterior, there is earlier Shh derepression. This anterior Shh domain then rescues and extends the anterior AER, expanding the distal limb bud.
The morphological consequences of these changes reflect both the degree of AER contraction and the timing of anterior Shh expression. In Twist1CC/CC limbs the most pronounced early signaling defects are on the posterior, yet only the anterior zeugopodal elements are affected. This is probably because when the limb bud narrows, the zeugopodal primordium supports formation of only one cartilage condensation, and as the Shh domain is strong and nearby, this tissue is specified as posterior. Consistent with this idea, fate-mapping experiments reveal a Shh-response only in posterior zeugopod (Harfe et al., 2004). If Twist1 activity is further reduced, as in Twist1CC/− limbs, then initial outgrowth is more severely compromised but is rescued by more robust and earlier ectopic Shh as well as rescue of the overlying AER. This expands the zeugopodal precursor region sufficiently to support two condensations, both of which will be specified as posterior. However this rescue occurs too late to promote normal stylopod development. This mechanism would account for both the hypoplastic humerus and duplicated ulna found in Twist1CC/− forelimbs. Interestingly in these animals the hindlimbs have a normal femur, and no duplication of the fibula, although the tibia and autopod are severely affected. This might reflect the minimal change in Alx4 expression in the anterior hindlimb bud.
Shh expression in the limb is controlled by a conserved enhancer element that is required for expression in the posterior limb bud (Ros et al., 2003; Sagai et al., 2005), but that when mutated can also direct expression to the limb anterior (Sagai et al., 2004; Sharpe et al., 1999). The protein complexes that interact with this element are not yet well defined (Amano et al., 2009). But as Twist1, Hand2, Etv and aristaless family genes all encode transcription factors, they might directly regulate this enhancer element. Furthermore, these proteins have the potential to interact physically as well as genetically. Physical interaction between Twist1 and Hand2 is already well documented, and their choice of binding partners is critical for in vivo function (Barnes and Firulli, 2009; Firulli et al., 2005; Firulli et al., 2007). It will be interesting to learn whether proteins of these other families bind to Twist1 or Hand2, or if they act less directly to mediate Twist1 regulation of Shh expression, and ultimately limb development.
Supplementary Material
Acknowledgments
We thank Alex Paul, Amy Daniel, Shannon Giorgianni for experimental support, and Andreas Kottmann, Cliff Tabin, Peter Cserjesi and Monica Justice for mice. Funding was provided by the NIH (R01-HD30284 to RRB, R01-DK075578 to FC, F31-DE1453 to DK, P30-AR044535 and R01-DK081515 to EL) and March of Dimes (5-FY99-855 to EL).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Amano T, Sagai T, Tanabe H, Mizushina Y, Nakazawa H, Shiroishi T. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev Cell. 2009;16:47–57. doi: 10.1016/j.devcel.2008.11.011. [DOI] [PubMed] [Google Scholar]
- Balling R, Deutsch U, Gruss P. undulated, a mutation affecting the development of the mouse skeleton, has a point mutation in the paired box of Pax 1. Cell. 1988;55:531–535. doi: 10.1016/0092-8674(88)90039-6. [DOI] [PubMed] [Google Scholar]
- Barnes RM, Firulli AB. A twist of insight - the role of Twist-family bHLH factors in development. Int J Dev Biol. 2009;53:909–924. doi: 10.1387/ijdb.082747rb. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beverdam A, Brouwer A, Reijnen M, Korving J, Meijlink F. Severe nasal clefting and abnormal embryonic apoptosis in Alx3/Alx4 double mutant mice. Development. 2001;128:3975–3986. doi: 10.1242/dev.128.20.3975. [DOI] [PubMed] [Google Scholar]
- Beverdam A, Meijlink F. Expression patterns of group-I aristaless-related genes during craniofacial and limb development. Mech Dev. 2001;107:163–167. doi: 10.1016/s0925-4773(01)00450-6. [DOI] [PubMed] [Google Scholar]
- Bialek P, Kern B, Yang X, Schrock M, Sosic D, Hong N, Wu H, Yu K, Ornitz DM, Olson EN, Justice MJ, Karsenty G. A twist code determines the onset of osteoblast differentiation. Dev Cell. 2004;6:423–435. doi: 10.1016/s1534-5807(04)00058-9. [DOI] [PubMed] [Google Scholar]
- Boulet AM, Moon AM, Arenkiel BR, Capecchi MR. The roles of Fgf4 and Fgf8 in limb bud initiation and outgrowth. Dev Biol. 2004;273:361–372. doi: 10.1016/j.ydbio.2004.06.012. [DOI] [PubMed] [Google Scholar]
- Bourgeois P, Bolcato-Bellemin AL, Danse JM, Bloch-Zupan A, Yoshiba K, Stoetzel C, Perrin-Schmitt F. The variable expressivity and incomplete penetrance of the twist-null heterozygous mouse phenotype resemble those of human Saethre-Chotzen syndrome. Hum Mol Genet. 1998;7:945–957. doi: 10.1093/hmg/7.6.945. [DOI] [PubMed] [Google Scholar]
- Burke AC, Nelson CE, Morgan BA, Tabin C. Hox genes and the evolution of vertebrate axial morphology. Development. 1995;121:333–346. doi: 10.1242/dev.121.2.333. [DOI] [PubMed] [Google Scholar]
- Cai J, Jabs EW. A twisted hand: bHLH protein phosphorylation and dimerization regulate limb development. Bioessays. 2005;27:1102–1106. doi: 10.1002/bies.20313. [DOI] [PubMed] [Google Scholar]
- Chapman DL, Garvey N, Hancock S, Alexiou M, Agulnik SI, Gibson-Brown JJ, Cebra-Thomas J, Bollag RJ, Silver LM, Papaioannou VE. Expression of the T-box family genes, Tbx1-Tbx5, during early mouse development. Dev Dyn. 1996;206:379–390. doi: 10.1002/(SICI)1097-0177(199608)206:4<379::AID-AJA4>3.0.CO;2-F. [DOI] [PubMed] [Google Scholar]
- Charite J, McFadden DG, Olson EN. The bHLH transcription factor dHAND controls Sonic hedgehog expression and establishment of the zone of polarizing activity during limb development. Development. 2000;127:2461–2470. doi: 10.1242/dev.127.11.2461. [DOI] [PubMed] [Google Scholar]
- Chen YT, Akinwunmi PO, Deng JM, Tam OH, Behringer RR. Generation of a Twist1 conditional null allele in the mouse. Genesis. 2007;45:588–592. doi: 10.1002/dvg.20332. [DOI] [PubMed] [Google Scholar]
- Chen ZF, Behringer RR. Twist is required in head mesenchyme for cranial neural tube morphogenesis. Genes Dev. 1995;9:686–699. doi: 10.1101/gad.9.6.686. [DOI] [PubMed] [Google Scholar]
- Chiang C, Litingtung Y, Harris MP, Simandl BK, Li Y, Beachy PA, Fallon JF. Manifestation of the limb prepattern: limb development in the absence of sonic hedgehog function. Dev Biol. 2001;236:421–435. doi: 10.1006/dbio.2001.0346. [DOI] [PubMed] [Google Scholar]
- Chotteau-Lelievre A, Dolle P, Peronne V, Coutte L, de Launoit Y, Desbiens X. Expression patterns of the Ets transcription factors from the PEA3 group during early stages of mouse development. Mech Dev. 2001;108:191–195. doi: 10.1016/s0925-4773(01)00480-4. [DOI] [PubMed] [Google Scholar]
- Crossley PH, Martin GR. The mouse Fgf8 gene encodes a family of polypeptides and is expressed in regions that direct outgrowth and patterning in the developing embryo. Development. 1995;121:439–451. doi: 10.1242/dev.121.2.439. [DOI] [PubMed] [Google Scholar]
- Davenport TG, Jerome-Majewska LA, Papaioannou VE. Mammary gland, limb and yolk sac defects in mice lacking Tbx3, the gene mutated in human ulnar mammary syndrome. Development. 2003;130:2263–2273. doi: 10.1242/dev.00431. [DOI] [PubMed] [Google Scholar]
- Davis AP, Witte DP, Hsieh-Li HM, Potter SS, Capecchi MR. Absence of radius and ulna in mice lacking hoxa-11 and hoxd-11. Nature. 1995;375:791–795. doi: 10.1038/375791a0. [DOI] [PubMed] [Google Scholar]
- Fallon JF, Lopez A, Ros MA, Savage MP, Olwin BB, Simandl BK. FGF-2: apical ectodermal ridge growth signal for chick limb development. Science. 1994;264:104–107. doi: 10.1126/science.7908145. [DOI] [PubMed] [Google Scholar]
- Fernandez-Teran M, Piedra ME, Kathiriya IS, Srivastava D, Rodriguez-Rey JC, Ros MA. Role of dHAND in the anterior-posterior polarization of the limb bud: implications for the Sonic hedgehog pathway. Development. 2000;127:2133–2142. doi: 10.1242/dev.127.10.2133. [DOI] [PubMed] [Google Scholar]
- Firulli BA, Krawchuk D, Centonze VE, Vargesson N, Virshup DM, Conway SJ, Cserjesi P, Laufer E, Firulli AB. Altered Twist1 and Hand2 dimerization is associated with Saethre-Chotzen syndrome and limb abnormalities. Nat Genet. 2005;37:373–381. doi: 10.1038/ng1525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Firulli BA, Redick BA, Conway SJ, Firulli AB. Mutations within helix I of Twist1 result in distinct limb defects and variation of DNA binding affinities. J Biol Chem. 2007;282:27536–27546. doi: 10.1074/jbc.M702613200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamburger V, Hamilton HL. A series of normal stages in the development of the chick embryo. J Exp Morph. 1951;88:49–92. [PubMed] [Google Scholar]
- Harfe BD, Scherz PJ, Nissim S, Tian H, McMahon AP, Tabin CJ. Evidence for an expansion-based temporal Shh gradient in specifying vertebrate digit identities. Cell. 2004;118:517–528. doi: 10.1016/j.cell.2004.07.024. [DOI] [PubMed] [Google Scholar]
- Hui CC, Slusarski D, Platt KA, Holmgren R, Joyner AL. Expression of three mouse homologs of the Drosophila segment polarity gene cubitus interruptus, Gli, Gli-2, and Gli-3, in ectoderm- and mesoderm-derived tissues suggests multiple roles during postimplantation development. Dev Biol. 1994;162:402–413. doi: 10.1006/dbio.1994.1097. [DOI] [PubMed] [Google Scholar]
- Isaac A, Cohn MJ, Ashby P, Ataliotis P, Spicer DB, Cooke J, Tickle C. FGF and genes encoding transcription factors in early limb specification. Mech Dev. 2000;93:41–48. doi: 10.1016/s0925-4773(00)00261-6. [DOI] [PubMed] [Google Scholar]
- Jeong J, Mao J, Tenzen T, Kottmann AH, McMahon AP. Hedgehog signaling in the neural crest cells regulates the patterning and growth of facial primordia. Genes Dev. 2004;18:937–951. doi: 10.1101/gad.1190304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuijper S, Beverdam A, Kroon C, Brouwer A, Candille S, Barsh G, Meijlink F. Genetics of shoulder girdle formation: roles of Tbx15 and aristaless-like genes. Development. 2005;132:1601–1610. doi: 10.1242/dev.01735. [DOI] [PubMed] [Google Scholar]
- Laufer E, Dahn R, Orozco OE, Yeo CY, Pisenti J, Henrique D, Abbott UK, Fallon JF, Tabin C. Expression of Radical fringe in limb-bud ectoderm regulates apical ectodermal ridge formation. Nature. 1997;386:366–373. doi: 10.1038/386366a0. [DOI] [PubMed] [Google Scholar]
- Laufer E, Nelson CE, Johnson RL, Morgan BA, Tabin C. Sonic hedgehog and Fgf-4 act through a signaling cascade and feedback loop to integrate growth and patterning of the developing limb bud. Cell. 1994;79:993–1003. doi: 10.1016/0092-8674(94)90030-2. [DOI] [PubMed] [Google Scholar]
- Liu JP, Laufer E, Jessell TM. Assigning the positional identity of spinal motor neurons: rostrocaudal patterning of Hox-c expression by FGFs, Gdf11, and retinoids. Neuron. 2001;32:997–1012. doi: 10.1016/s0896-6273(01)00544-x. [DOI] [PubMed] [Google Scholar]
- Livet J, Sigrist M, Stroebel S, De Paola V, Price SR, Henderson CE, Jessell TM, Arber S. ETS gene Pea3 controls the central position and terminal arborization of specific motor neuron pools. Neuron. 2002;35:877–892. doi: 10.1016/s0896-6273(02)00863-2. [DOI] [PubMed] [Google Scholar]
- Loebel DA, O'Rourke MP, Steiner KA, Banyer J, Tam PP. Isolation of differentially expressed genes from wild-type and Twist mutant mouse limb buds. Genesis. 2002;33:103–113. doi: 10.1002/gene.10091. [DOI] [PubMed] [Google Scholar]
- Logan M, Martin JF, Nagy A, Lobe C, Olson EN, Tabin CJ. Expression of Cre Recombinase in the developing mouse limb bud driven by a Prxl enhancer. Genesis. 2002;33:77–80. doi: 10.1002/gene.10092. [DOI] [PubMed] [Google Scholar]
- Lu BC, Cebrian C, Chi X, Kuure S, Kuo R, Bates CM, Arber S, Hassell J, MacNeil L, Hoshi M, Jain S, Asai N, Takahashi M, Schmidt-Ott KM, Barasch J, D’Agati V, Costantini F. Etv4 and Etv5 are required downstream of GDNF and Ret for kidney branching morphogenesis. Nat Genet. 2009;41:1295–1302. doi: 10.1038/ng.476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao J, McGlinn E, Huang P, Tabin CJ, McMahon AP. Fgf-dependent Etv4/5 activity is required for posterior restriction of Sonic Hedgehog and promoting outgrowth of the vertebrate limb. Dev Cell. 2009;16:600–606. doi: 10.1016/j.devcel.2009.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marigo V, Davey RA, Zuo Y, Cunningham JM, Tabin CJ. Biochemical evidence that patched is the Hedgehog receptor. Nature. 1996;384:176–179. doi: 10.1038/384176a0. [DOI] [PubMed] [Google Scholar]
- Mason JM, Morrison DJ, Basson MA, Licht JD. Sprouty proteins: multifaceted negative-feedback regulators of receptor tyrosine kinase signaling. Trends Cell Biol. 2006;16:45–54. doi: 10.1016/j.tcb.2005.11.004. [DOI] [PubMed] [Google Scholar]
- Minowada G, Jarvis LA, Chi CL, Neubuser A, Sun X, Hacohen N, Krasnow MA, Martin GR. Vertebrate Sprouty genes are induced by FGF signaling and can cause chondrodysplasia when overexpressed. Development. 1999;126:4465–4475. doi: 10.1242/dev.126.20.4465. [DOI] [PubMed] [Google Scholar]
- Niswander L, Jeffrey S, Martin GR, Tickle C. A positive feedback loop coordinates growth and patterning in the vertebrate limb. Nature. 1994;371:609–612. doi: 10.1038/371609a0. [DOI] [PubMed] [Google Scholar]
- Niswander L, Tickle C, Vogel A, Booth I, Martin GR. FGF-4 replaces the apical ectodermal ridge and directs outgrowth and patterning of the limb. Cell. 1993;75:579–587. doi: 10.1016/0092-8674(93)90391-3. [DOI] [PubMed] [Google Scholar]
- O'Rourke MP, Soo K, Behringer RR, Hui CC, Tam PP. Twist plays an essential role in FGF and SHH signal transduction during mouse limb development. Dev Biol. 2002;248:143–156. doi: 10.1006/dbio.2002.0730. [DOI] [PubMed] [Google Scholar]
- Patton JT, Kaufman MH. The timing of ossification of the limb bones, and growth rates of various long bones of the fore and hind limbs of the prenatal and early postnatal laboratory mouse. J Anat. 1995;186(Pt 1):175–185. [PMC free article] [PubMed] [Google Scholar]
- Peters KG, Werner S, Chen G, Williams LT. Two FGF receptor genes are differentially expressed in epithelial and mesenchymal tissues during limb formation and organogenesis in the mouse. Development. 1992;114:233–243. doi: 10.1242/dev.114.1.233. [DOI] [PubMed] [Google Scholar]
- Qu S, Niswender KD, Ji Q, van der Meer R, Keeney D, Magnuson MA, Wisdom R. Polydactyly and ectopic ZPA formation in Alx-4 mutant mice. Development. 1997;124:3999–4008. doi: 10.1242/dev.124.20.3999. [DOI] [PubMed] [Google Scholar]
- Qu S, Tucker SC, Zhao Q, deCrombrugghe B, Wisdom R. Physical and genetic interactions between Alx4 and Cart1. Development. 1999;126:359–369. doi: 10.1242/dev.126.2.359. [DOI] [PubMed] [Google Scholar]
- Riddle RD, Johnson RL, Laufer E, Tabin C. Sonic hedgehog mediates the polarizing activity of the ZPA. Cell. 1993;75:1401–1416. doi: 10.1016/0092-8674(93)90626-2. [DOI] [PubMed] [Google Scholar]
- Roelink H, Augsburger A, Heemskerk J, Korzh V, Norlin S, Ruiz i Altaba A, Tanabe Y, Placzek M, Edlund T, Jessell TM, et a. Floor plate and motor neuron induction by vhh-1, a vertebrate homolog of hedgehog expressed by the notochord. Cell. 1994;76:761–775. doi: 10.1016/0092-8674(94)90514-2. [DOI] [PubMed] [Google Scholar]
- Ros MA, Dahn RD, Fernandez-Teran M, Rashka K, Caruccio NC, Hasso SM, Bitgood JJ, Lancman JJ, Fallon JF. The chick oligozeugodactyly (ozd) mutant lacks sonic hedgehog function in the limb. Development. 2003;130:527–537. doi: 10.1242/dev.00245. [DOI] [PubMed] [Google Scholar]
- Sagai T, Hosoya M, Mizushina Y, Tamura M, Shiroishi T. Elimination of a long-range cis-regulatory module causes complete loss of limb-specific Shh expression and truncation of the mouse limb. Development. 2005;132:797–803. doi: 10.1242/dev.01613. [DOI] [PubMed] [Google Scholar]
- Sagai T, Masuya H, Tamura M, Shimizu K, Yada Y, Wakana S, Gondo Y, Noda T, Shiroishi T. Phylogenetic conservation of a limb-specific, cis-acting regulator of Sonic hedgehog ( Shh) Mamm Genome. 2004;15:23–34. doi: 10.1007/s00335-033-2317-5. [DOI] [PubMed] [Google Scholar]
- Saunders J., Jr The proximo-distal sequence of origin of the parts of the chick wing and the role of the ectoderm. J Exp Zool. 1948;108:363–403. doi: 10.1002/jez.1401080304. [DOI] [PubMed] [Google Scholar]
- Sharpe J, Lettice L, Hecksher-Sorensen J, Fox M, Hill R, Krumlauf R. Identification of sonic hedgehog as a candidate gene responsible for the polydactylous mouse mutant Sasquatch. Curr Biol. 1999;9:97–100. doi: 10.1016/s0960-9822(99)80022-0. [DOI] [PubMed] [Google Scholar]
- Srivastava D, Cserjesi P, Olson EN. A subclass of bHLH proteins required for cardiac morphogenesis. Science. 1995;270:1995–1999. doi: 10.1126/science.270.5244.1995. [DOI] [PubMed] [Google Scholar]
- Srivastava D, Thomas T, Lin Q, Kirby ML, Brown D, Olson EN. Regulation of cardiac mesodermal and neural crest development by the bHLH transcription factor, dHAND. Nat Genet. 1997;16:154–160. doi: 10.1038/ng0697-154. [DOI] [PubMed] [Google Scholar]
- Sun X, Mariani FV, Martin GR. Functions of FGF signalling from the apical ectodermal ridge in limb development. Nature. 2002;418:501–508. doi: 10.1038/nature00902. [DOI] [PubMed] [Google Scholar]
- te Welscher P, Fernandez-Teran M, Ros MA, Zeller R. Mutual genetic antagonism involving GLI3 and dHAND prepatterns the vertebrate limb bud mesenchyme prior to SHH signaling. Genes Dev. 2002;16:421–426. doi: 10.1101/gad.219202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vargesson N, Luria V, Messina I, Erskine L, Laufer E. Expression patterns of Slit and Robo family members during vertebrate limb development. Mech Dev. 2001;106:175–180. doi: 10.1016/s0925-4773(01)00430-0. [DOI] [PubMed] [Google Scholar]
- Webb GN, Byrd RA. Simultaneous differential staining of cartilage and bone in rodent fetuses: an alcian blue and alizarin red S procedure without glacial acetic acid. Biotech Histochem. 1994;69:181–185. doi: 10.3109/10520299409106284. [DOI] [PubMed] [Google Scholar]
- Zakany J, Kmita M, Duboule D. A dual role for Hox genes in limb anterior-posterior asymmetry. Science. 2004;304:1669–1672. doi: 10.1126/science.1096049. [DOI] [PubMed] [Google Scholar]
- Zeller R, Lopez-Rios J, Zuniga A. Vertebrate limb bud development: moving towards integrative analysis of organogenesis. Nat Rev Genet. 2009;10:845–858. doi: 10.1038/nrg2681. [DOI] [PubMed] [Google Scholar]
- Zhang S, Lin Y, Itaranta P, Yagi A, Vainio S. Expression of Sprouty genes 1, 2 and 4 during mouse organogenesis. Mech Dev. 2001;109:367–370. doi: 10.1016/s0925-4773(01)00526-3. [DOI] [PubMed] [Google Scholar]
- Zhang Z, Verheyden JM, Hassell JA, Sun X. FGF-regulated Etv genes are essential for repressing Shh expression in mouse limb buds. Dev Cell. 2009;16:607–613. doi: 10.1016/j.devcel.2009.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuniga A, Quillet R, Perrin-Schmitt F, Zeller R. Mouse Twist is required for fibroblast growth factor-mediated epithelial-mesenchymal signalling and cell survival during limb morphogenesis. Mech Dev. 2002;114:51–59. doi: 10.1016/s0925-4773(02)00048-5. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.