Abstract
The asymmetric unit of the title compound, [Cd(C6H11Cl2NO6P2)(H2O)3]n, contains two octahedrally coordinated Cd atoms located in special positions, one on a twofold rotation axis and the other on a centre of symmetry. The metal atoms are connected by methyl morpholino dichloromethylenediphosphonate ligands into chains in the c-axis direction. These chains are further connected by O—H⋯O hydrogen bonds into a layer-like construction along (100).
Related literature
For applications of metal complexes of bisphosphonates, see: Clearfield (1998 ▶); Clearfield et al. (2001 ▶); Fu et al. (2007 ▶). For cadmium bisphosphonate complexes, see: Ying & Mao (2006 ▶); Man et al. (2006 ▶). For metal complexes of bisphosphonate ester derivatives, see: Jokiniemi et al. (2007 ▶, 2008 ▶). For Mg, Zn and Cd complexes of the symmetrical diethyl ester derivative of (dichloromethylene)bisphosphonate, see: Kontturi et al. (2002 ▶, 2005a
▶,b
▶).
Experimental
Crystal data
[Cd(C6H11Cl2NO6P2)(H2O)3]
M r = 492.45
Monoclinic,

a = 26.2488 (8) Å
b = 7.6578 (3) Å
c = 17.5445 (7) Å
β = 116.002 (3)°
V = 3169.6 (2) Å3
Z = 8
Mo Kα radiation
μ = 1.96 mm−1
T = 120 K
0.30 × 0.25 × 0.20 mm
Data collection
Nonius KappaCCD diffractometer
Absorption correction: multi-scan (XPREP in SHELXTL; Sheldrick, 2008 ▶) T min = 0.565, T max = 0.676
21828 measured reflections
4053 independent reflections
3370 reflections with I > 2σ(I)
R int = 0.038
Refinement
R[F 2 > 2σ(F 2)] = 0.027
wR(F 2) = 0.068
S = 1.06
4053 reflections
195 parameters
H-atom parameters constrained
Δρmax = 1.04 e Å−3
Δρmin = −1.12 e Å−3
Data collection: COLLECT (Nonius, 1997 ▶); cell refinement: DENZO/SCALEPACK (Otwinowski & Minor, 1997 ▶); data reduction: DENZO/SCALEPACK; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: DIAMOND (Brandenburg, 2005 ▶); software used to prepare material for publication: SHELXL97.
Supplementary Material
Crystal structure: contains datablocks I, global. DOI: 10.1107/S160053680901527X/er2063sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S160053680901527X/er2063Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Selected geometric parameters (Å, °).
| Cd1—O11 | 2.2256 (17) |
| Cd1—O21 | 2.3173 (16) |
| Cd1—O1 | 2.3409 (17) |
| Cd2—O12 | 2.1884 (17) |
| Cd2—O3 | 2.2795 (16) |
| Cd2—O2 | 2.3486 (16) |
Table 2. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| O1—H1A⋯O2i | 0.84 | 2.06 | 2.849 (2) | 156 |
| O1—H1B⋯O22ii | 0.88 | 2.12 | 2.990 (2) | 170 |
| O2—H2A⋯O21iii | 0.86 | 2.04 | 2.844 (2) | 155 |
| O2—H2B⋯O22 | 0.86 | 1.84 | 2.662 (2) | 159 |
| O3—H3A⋯O22 | 0.83 | 2.03 | 2.773 (2) | 149 |
| O3—H3B⋯O13iv | 0.90 | 1.87 | 2.745 (2) | 163 |
Symmetry codes: (i)  ; (ii)
; (ii)  ; (iii)
; (iii)  ; (iv)
; (iv)  .
.
supplementary crystallographic information
Comment
Metal complexes with bisphosphonic acids have interesting structures with various coordination architectures, and properties that offer practical applications in catalysis, ion-exchange and sorption (Clearfield et al., 2001, Clearfield, 1998, Fu et al., 2007). In our recent investigations, we studied the complexing properties of amide ester derivatives of (dichloromethylene)bisphosphonate, Cl2MBP (Jokiniemi et al., 2007, 2008). Introduction of these ester substituents to phosphorus groups can result in novel structures of metal bishosphonates and lead to interesting functionalities. We now present the crystal structure of the Cd(II) complex of the P-morpholinyl-P'-methyl ester derivative of Cl2MBP obtained by gel crystallization.
The title compound is isomorphous with the earlier reported Mg complexes of (dichloromethylene)bisphosphonic acid methyl esters of piperidinyl and morpholinyl derivatives (Jokiniemi et al., 2007, 2008). The title compound is polymeric, consisting of chains in the direction of the c-axis. There are two crystallographically independent six-coordinated Cd2+ cations in the asymmetric unit, located in special positions: Cd1 on the twofold rotation axis and Cd2 on the centre of symmetry (Fig. 1). Two symmetrically related L1 ligands, L1 = (Cl2CP2O5MeNC4H8O), around the Cd1 atom form six-membered chelate rings. The L1 ligand is further connected to Cd2 through one O atom, and thus acts as a triatomic bridge between the adjacent Cd atoms. The fourth phosphonate O atom remains non-coordinated but is involved in hydrogen bonding. The remaining coordination sites around the Cd1 atoms are occupied by aqua ligands in cis position; the geometry is a significantly distorted octahedron having Cd1–O bond distances 2.226 (2)–2.341 (2) Å (Table 1). The three trans angles are O11–Cd1–O11A 166.14 (9), O21–Cd1–O1A 174.23 (5) and O21A–Cd1–O1 174.23 (5)°. The Cd2 atom has a distorted octahedral geometry, and the binding sites around the metal atom are occupied by two phosphonate O atoms in axial positions and four aqua ligands having Cd2–O bond lengths 2.188 (2)–2.349 (2) Å. The three trans bond angles are 180°, while the cis bond angles around the Cd2 atom range from 82.50 (6) to 97.50 (6)°. In addition to isomorphous Mg complexes of monomethyl ester of morpholinyl and piperidinyl derivatives of Cl2MBP (Jokiniemi et al., 2007 and 2008), the same kind of chain construction is found in Mg, Zn and Cd complexes of the symmetrical diethyl ester derivative of Cl2MBP (Kontturi et al., 2002, 2005a and 2005b).
The polymeric chains are connected, in a layer-like structure parallel to the (100) plane, by hydrogen bonds [O···O 2.745 (2)–2.990 (2) Å, 149–170°, Table 2]. The morpholinyl rings and chlorine atoms of the L1 ligands point out from the layers (Fig. 2), which are held together solely by weak Van der Waals interactions, with an interlayer distance of 11.7959 Å.
Experimental
(H2N[(CH2)2]2O)2CH3PO3CCl2PO2NC4H8O (19.8 mg, 0.039 mmol) and Cd(NO3)2×4H2O (12.1 mg, 0.039 mmol) were dissolved separately in water (0.45 ml), the solutions were mixed, and tetramethoxysilane (TMOS 0.1 ml) was added. The two-phase system was shaken until homogeneous. After gel formation, a precipitant, acetone (1.0 ml), was added above the gel to induce crystallization. After about three weeks, large, colourless plank-shaped crystals suitable for X-ray analysis formed at the gel-liquid boundary. Anal. Found: C, 14.63; H, 3.48; N, 2.84; Cd, 22.83%. Calc. for C6H17Cl2CdNO9P2: C, 14.74; H, 3.48; N, 2.86; Cd, 22.45%. Main IR absorptions (KBr pellet, cm-1): 3432 (s), 2961 (m), 2926 (m), 2854 (m), 1627 (m), 1204 (versus), 1145 (m), 1101 (versus), 1072 (s), 1056 (s), 981 (s), 869 (m), 843 (m). 31P CP/MAS NMR: δP 8.4 and 4.3 p.p.m.. TGA (25–900 °C under a synthetic air): 30–110 °C 12.6% (calculated 11.0% for the loss of three aqua ligands). The second step (190–900 °C) is attributed to the release of organic groups, chlorine atoms and a methylene carbon atom. The observed total weight loss is 47.0% (calculated 45.1% if the final product is assumed to be Cd(PO3)2).
Refinement
H atoms of the methyl and morpholinyl groups were placed at calculated positions in the riding-model approximation with C–H = 0.99 Å (morpholinyl) [UISO(H) = 1.2Ueq(C)] and C–H = 0.98 Å (methyl) [UISO(H) = 1.5Ueq(C)]. H atoms of the aqua ligands were located in a difference map and treated as riding, with O–H bond lengths constrained to 0.83–0.90 Å and with UISO(H) = 1.5Ueq(O).
Figures
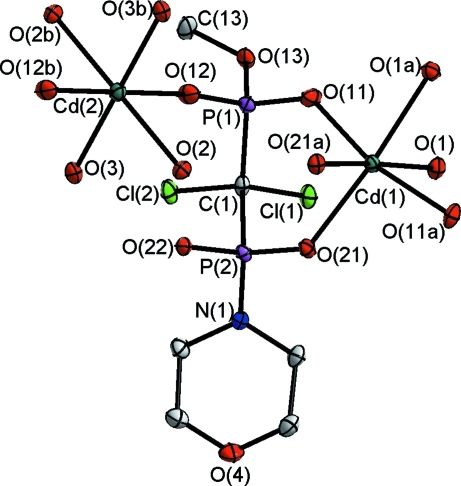
Fig. 1.
Structure of the title compound showing the atomic numbering scheme and 50% probability displacement ellipsoids. Hydrogen atoms are omitted for clarity. Atoms labelled with suffixes A and B are at the symmetry postitions (1 - x, y, 1/2 - z) and (1 - x, 1 - y, - z) respectively.
Fig. 2.
Packing of the title compound viewed along the b-axis. CdO6 octahedra are presented in medium grey and PO3C and NPO2C tetrahedra in dark grey. Hydrogen atoms are omitted for clarity.
Crystal data
| [Cd(C6H11Cl2NO6P2)(H2O)3] | F(000) = 1952 |
| Mr = 492.45 | Dx = 2.064 Mg m−3 |
| Monoclinic, C2/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -C 2yc | Cell parameters from 21828 reflections |
| a = 26.2488 (8) Å | θ = 2.8–28.7° |
| b = 7.6578 (3) Å | µ = 1.96 mm−1 |
| c = 17.5445 (7) Å | T = 120 K |
| β = 116.002 (3)° | Plank, colourles |
| V = 3169.6 (2) Å3 | 0.30 × 0.25 × 0.20 mm |
| Z = 8 |
Data collection
| Nonius KappaCCD diffractometer | 4053 independent reflections |
| Radiation source: fine-focus sealed tube | 3370 reflections with I > 2σ(I) |
| graphite | Rint = 0.038 |
| multi–scan | θmax = 28.7°, θmin = 2.8° |
| Absorption correction: multi-scan (XPREP in SHELXTL; Sheldrick, 2008) | h = −35→35 |
| Tmin = 0.565, Tmax = 0.676 | k = −10→10 |
| 21828 measured reflections | l = −23→21 |
Refinement
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.027 | H-atom parameters constrained |
| wR(F2) = 0.068 | w = 1/[σ2(Fo2) + (0.04P)2] where P = (Fo2 + 2Fc2)/3 |
| S = 1.06 | (Δ/σ)max < 0.001 |
| 4053 reflections | Δρmax = 1.03 e Å−3 |
| 195 parameters | Δρmin = −1.12 e Å−3 |
| 0 restraints | Extinction correction: SHELXL97 (Sheldrick, 2008), Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| Primary atom site location: structure-invariant direct methods | Extinction coefficient: 0.00053 (7) |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| Cl1 | 0.33755 (3) | 0.03444 (8) | 0.08713 (4) | 0.01735 (14) | |
| Cl2 | 0.31951 (3) | 0.31865 (8) | −0.03219 (3) | 0.01910 (14) | |
| P1 | 0.42845 (3) | 0.12553 (8) | 0.03696 (4) | 0.01308 (14) | |
| P2 | 0.39459 (2) | 0.37248 (8) | 0.15068 (4) | 0.01129 (13) | |
| Cd1 | 0.5000 | 0.05298 (3) | 0.2500 | 0.01210 (8) | |
| Cd2 | 0.5000 | 0.5000 | 0.0000 | 0.01366 (8) | |
| O1 | 0.43819 (7) | −0.1737 (2) | 0.24266 (10) | 0.0170 (4) | |
| H1A | 0.4474 | −0.2385 | 0.2856 | 0.025* | |
| H1B | 0.4298 | −0.2517 | 0.2023 | 0.025* | |
| O2 | 0.52362 (7) | 0.5362 (2) | 0.14457 (10) | 0.0149 (4) | |
| H2A | 0.5420 | 0.4483 | 0.1748 | 0.022* | |
| H2B | 0.4929 | 0.5313 | 0.1507 | 0.022* | |
| O3 | 0.41722 (7) | 0.6409 (2) | −0.02767 (10) | 0.0182 (4) | |
| H3A | 0.4041 | 0.6034 | 0.0045 | 0.027* | |
| H3B | 0.4177 | 0.7579 | −0.0243 | 0.042 (9)* | |
| O4 | 0.26897 (8) | 0.6093 (3) | 0.22336 (12) | 0.0281 (4) | |
| O11 | 0.46611 (8) | 0.0179 (2) | 0.11038 (11) | 0.0182 (4) | |
| O12 | 0.45203 (8) | 0.2725 (2) | 0.00769 (11) | 0.0220 (4) | |
| O13 | 0.39645 (7) | −0.0065 (2) | −0.03937 (10) | 0.0172 (4) | |
| O21 | 0.43465 (7) | 0.2737 (2) | 0.22805 (9) | 0.0138 (3) | |
| O22 | 0.41655 (7) | 0.5285 (2) | 0.12215 (10) | 0.0142 (4) | |
| N1 | 0.33955 (8) | 0.4329 (3) | 0.16396 (12) | 0.0147 (4) | |
| C1 | 0.37131 (10) | 0.2129 (3) | 0.06118 (14) | 0.0130 (5) | |
| C2 | 0.29966 (10) | 0.5686 (3) | 0.11230 (15) | 0.0187 (5) | |
| H2E | 0.2634 | 0.5141 | 0.0731 | 0.022* | |
| H2F | 0.3155 | 0.6307 | 0.0781 | 0.022* | |
| C3 | 0.28936 (12) | 0.6969 (4) | 0.16995 (17) | 0.0246 (6) | |
| H3E | 0.3251 | 0.7585 | 0.2056 | 0.029* | |
| H3F | 0.2612 | 0.7851 | 0.1351 | 0.029* | |
| C5 | 0.30950 (12) | 0.4819 (4) | 0.27479 (17) | 0.0240 (6) | |
| H5E | 0.2955 | 0.4245 | 0.3125 | 0.029* | |
| H5F | 0.3457 | 0.5408 | 0.3109 | 0.029* | |
| C6 | 0.31968 (11) | 0.3450 (3) | 0.22048 (15) | 0.0188 (5) | |
| H6E | 0.3485 | 0.2601 | 0.2570 | 0.023* | |
| H6F | 0.2841 | 0.2808 | 0.1865 | 0.023* | |
| C13 | 0.37843 (13) | 0.0444 (4) | −0.12767 (15) | 0.0267 (6) | |
| H13A | 0.3832 | 0.1707 | −0.1308 | 0.040* | |
| H13B | 0.3384 | 0.0137 | −0.1610 | 0.040* | |
| H13C | 0.4015 | −0.0171 | −0.1504 | 0.040* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Cl1 | 0.0178 (3) | 0.0143 (3) | 0.0189 (3) | −0.0048 (2) | 0.0070 (2) | 0.0009 (2) |
| Cl2 | 0.0207 (3) | 0.0164 (3) | 0.0133 (3) | 0.0014 (2) | 0.0010 (2) | 0.0018 (2) |
| P1 | 0.0172 (3) | 0.0098 (3) | 0.0123 (3) | −0.0019 (2) | 0.0065 (2) | −0.0020 (2) |
| P2 | 0.0116 (3) | 0.0097 (3) | 0.0117 (3) | −0.0005 (2) | 0.0042 (2) | −0.0006 (2) |
| Cd1 | 0.01370 (13) | 0.00967 (13) | 0.01166 (13) | 0.000 | 0.00440 (9) | 0.000 |
| Cd2 | 0.01748 (14) | 0.01108 (14) | 0.01369 (14) | −0.00167 (9) | 0.00800 (10) | −0.00030 (9) |
| O1 | 0.0202 (9) | 0.0148 (9) | 0.0144 (8) | −0.0028 (7) | 0.0062 (7) | 0.0010 (7) |
| O2 | 0.0155 (9) | 0.0161 (9) | 0.0129 (9) | 0.0000 (7) | 0.0060 (7) | 0.0004 (7) |
| O3 | 0.0229 (10) | 0.0124 (9) | 0.0210 (9) | 0.0016 (7) | 0.0112 (7) | 0.0029 (7) |
| O4 | 0.0294 (11) | 0.0286 (11) | 0.0380 (11) | 0.0081 (9) | 0.0254 (9) | 0.0052 (9) |
| O11 | 0.0201 (9) | 0.0220 (10) | 0.0104 (9) | 0.0041 (7) | 0.0047 (7) | −0.0021 (7) |
| O12 | 0.0316 (11) | 0.0129 (9) | 0.0309 (10) | −0.0063 (8) | 0.0225 (9) | −0.0042 (7) |
| O13 | 0.0257 (10) | 0.0107 (9) | 0.0117 (9) | −0.0015 (7) | 0.0050 (7) | −0.0020 (6) |
| O21 | 0.0137 (8) | 0.0132 (9) | 0.0123 (8) | 0.0013 (6) | 0.0036 (6) | −0.0002 (6) |
| O22 | 0.0165 (9) | 0.0120 (9) | 0.0165 (9) | −0.0007 (7) | 0.0094 (7) | 0.0003 (6) |
| N1 | 0.0172 (11) | 0.0143 (11) | 0.0150 (10) | 0.0018 (8) | 0.0092 (8) | 0.0031 (8) |
| C1 | 0.0142 (11) | 0.0108 (12) | 0.0111 (11) | −0.0021 (9) | 0.0030 (9) | 0.0015 (9) |
| C2 | 0.0159 (12) | 0.0208 (14) | 0.0187 (13) | 0.0049 (10) | 0.0071 (10) | 0.0043 (10) |
| C3 | 0.0276 (15) | 0.0210 (14) | 0.0301 (15) | 0.0069 (11) | 0.0173 (12) | 0.0027 (11) |
| C5 | 0.0303 (16) | 0.0247 (15) | 0.0247 (15) | −0.0005 (11) | 0.0192 (12) | 0.0025 (11) |
| C6 | 0.0202 (13) | 0.0185 (13) | 0.0212 (13) | −0.0030 (10) | 0.0124 (10) | 0.0025 (10) |
| C13 | 0.0457 (18) | 0.0198 (14) | 0.0107 (13) | 0.0026 (12) | 0.0088 (12) | −0.0007 (10) |
Geometric parameters (Å, °)
| Cl1—C1 | 1.792 (2) | O1—H1B | 0.8775 |
| Cl2—C1 | 1.799 (2) | O2—H2A | 0.8617 |
| P1—O12 | 1.4803 (18) | O2—H2B | 0.8598 |
| P1—O11 | 1.4829 (18) | O3—H3A | 0.8298 |
| P1—O13 | 1.5922 (17) | O3—H3B | 0.8982 |
| P1—C1 | 1.854 (2) | O4—C3 | 1.433 (3) |
| P2—O22 | 1.5044 (17) | O4—C5 | 1.434 (3) |
| P2—O21 | 1.5062 (16) | O13—C13 | 1.460 (3) |
| P2—N1 | 1.628 (2) | N1—C6 | 1.470 (3) |
| P2—C1 | 1.869 (2) | N1—C2 | 1.472 (3) |
| Cd1—O11 | 2.2256 (17) | C2—C3 | 1.517 (3) |
| Cd1—O11i | 2.2256 (17) | C2—H2E | 0.9900 |
| Cd1—O21i | 2.3173 (16) | C2—H2F | 0.9900 |
| Cd1—O21 | 2.3173 (16) | C3—H3E | 0.9900 |
| Cd1—O1 | 2.3409 (17) | C3—H3F | 0.9900 |
| Cd1—O1i | 2.3409 (17) | C5—C6 | 1.517 (4) |
| Cd2—O12ii | 2.1884 (17) | C5—H5E | 0.9900 |
| Cd2—O12 | 2.1884 (17) | C5—H5F | 0.9900 |
| Cd2—O3ii | 2.2795 (16) | C6—H6E | 0.9900 |
| Cd2—O3 | 2.2795 (16) | C6—H6F | 0.9900 |
| Cd2—O2 | 2.3486 (16) | C13—H13A | 0.9800 |
| Cd2—O2ii | 2.3486 (16) | C13—H13B | 0.9800 |
| O1—H1A | 0.8439 | C13—H13C | 0.9800 |
| O12—P1—O11 | 120.19 (11) | Cd2—O3—H3A | 109.4 |
| O12—P1—O13 | 109.73 (10) | Cd2—O3—H3B | 118.2 |
| O11—P1—O13 | 106.42 (10) | H3A—O3—H3B | 107.4 |
| O12—P1—C1 | 107.98 (10) | C3—O4—C5 | 110.07 (19) |
| O11—P1—C1 | 107.43 (10) | P1—O11—Cd1 | 132.97 (10) |
| O13—P1—C1 | 103.90 (10) | P1—O12—Cd2 | 165.00 (12) |
| O22—P2—O21 | 118.72 (10) | C13—O13—P1 | 121.93 (16) |
| O22—P2—N1 | 108.40 (10) | P2—O21—Cd1 | 133.58 (9) |
| O21—P2—N1 | 109.04 (10) | C6—N1—C2 | 111.82 (19) |
| O22—P2—C1 | 105.77 (10) | C6—N1—P2 | 124.48 (17) |
| O21—P2—C1 | 105.72 (10) | C2—N1—P2 | 123.28 (16) |
| N1—P2—C1 | 108.80 (11) | Cl1—C1—Cl2 | 108.31 (12) |
| O11—Cd1—O11i | 166.14 (9) | Cl1—C1—P1 | 108.85 (12) |
| O11—Cd1—O21i | 100.40 (6) | Cl2—C1—P1 | 108.55 (12) |
| O11i—Cd1—O21i | 89.75 (6) | Cl1—C1—P2 | 107.53 (11) |
| O11—Cd1—O21 | 89.75 (6) | Cl2—C1—P2 | 107.98 (12) |
| O11i—Cd1—O21 | 100.40 (6) | P1—C1—P2 | 115.42 (12) |
| O21i—Cd1—O21 | 86.31 (8) | N1—C2—C3 | 109.5 (2) |
| O11—Cd1—O1 | 85.24 (6) | N1—C2—H2E | 109.8 |
| O11i—Cd1—O1 | 84.49 (6) | C3—C2—H2E | 109.8 |
| O21i—Cd1—O1 | 174.23 (5) | N1—C2—H2F | 109.8 |
| O21—Cd1—O1 | 95.00 (6) | C3—C2—H2F | 109.8 |
| O11—Cd1—O1i | 84.49 (6) | H2E—C2—H2F | 108.2 |
| O11i—Cd1—O1i | 85.24 (6) | O4—C3—C2 | 111.1 (2) |
| O21i—Cd1—O1i | 95.00 (6) | O4—C3—H3E | 109.4 |
| O21—Cd1—O1i | 174.23 (5) | C2—C3—H3E | 109.4 |
| O1—Cd1—O1i | 84.25 (8) | O4—C3—H3F | 109.4 |
| O12ii—Cd2—O12 | 180.00 (9) | C2—C3—H3F | 109.4 |
| O12ii—Cd2—O3ii | 82.50 (6) | H3E—C3—H3F | 108.0 |
| O12—Cd2—O3ii | 97.50 (6) | O4—C5—C6 | 111.2 (2) |
| O12ii—Cd2—O3 | 97.50 (6) | O4—C5—H5E | 109.4 |
| O12—Cd2—O3 | 82.50 (6) | C6—C5—H5E | 109.4 |
| O3ii—Cd2—O3 | 180.00 (8) | O4—C5—H5F | 109.4 |
| O12ii—Cd2—O2 | 94.94 (6) | C6—C5—H5F | 109.4 |
| O12—Cd2—O2 | 85.06 (6) | H5E—C5—H5F | 108.0 |
| O3ii—Cd2—O2 | 92.88 (6) | N1—C6—C5 | 108.6 (2) |
| O3—Cd2—O2 | 87.12 (6) | N1—C6—H6E | 110.0 |
| O12ii—Cd2—O2ii | 85.06 (6) | C5—C6—H6E | 110.0 |
| O12—Cd2—O2ii | 94.94 (6) | N1—C6—H6F | 110.0 |
| O3ii—Cd2—O2ii | 87.12 (6) | C5—C6—H6F | 110.0 |
| O3—Cd2—O2ii | 92.88 (6) | H6E—C6—H6F | 108.3 |
| O2—Cd2—O2ii | 180.0 | O13—C13—H13A | 109.5 |
| Cd1—O1—H1A | 117.7 | O13—C13—H13B | 109.5 |
| Cd1—O1—H1B | 118.1 | H13A—C13—H13B | 109.5 |
| H1A—O1—H1B | 101.1 | O13—C13—H13C | 109.5 |
| Cd2—O2—H2A | 112.6 | H13A—C13—H13C | 109.5 |
| Cd2—O2—H2B | 108.2 | H13B—C13—H13C | 109.5 |
| H2A—O2—H2B | 101.1 | ||
| O12—P1—O11—Cd1 | 82.84 (17) | O21—P2—N1—C2 | 163.49 (18) |
| O13—P1—O11—Cd1 | −151.80 (14) | C1—P2—N1—C2 | −81.7 (2) |
| C1—P1—O11—Cd1 | −41.01 (18) | O12—P1—C1—Cl1 | 174.59 (11) |
| O11i—Cd1—O11—P1 | 151.44 (15) | O11—P1—C1—Cl1 | −54.41 (14) |
| O21i—Cd1—O11—P1 | −72.10 (16) | O13—P1—C1—Cl1 | 58.11 (13) |
| O21—Cd1—O11—P1 | 14.09 (16) | O12—P1—C1—Cl2 | 56.89 (14) |
| O1—Cd1—O11—P1 | 109.13 (16) | O11—P1—C1—Cl2 | −172.11 (11) |
| O1i—Cd1—O11—P1 | −166.20 (16) | O13—P1—C1—Cl2 | −59.59 (13) |
| O11—P1—O12—Cd2 | −39.7 (5) | O12—P1—C1—P2 | −64.44 (15) |
| O13—P1—O12—Cd2 | −163.4 (4) | O11—P1—C1—P2 | 66.56 (15) |
| C1—P1—O12—Cd2 | 83.9 (5) | O13—P1—C1—P2 | 179.07 (11) |
| O3ii—Cd2—O12—P1 | 77.6 (4) | O22—P2—C1—Cl1 | −173.74 (11) |
| O3—Cd2—O12—P1 | −102.4 (4) | O21—P2—C1—Cl1 | 59.52 (13) |
| O2—Cd2—O12—P1 | −14.7 (4) | N1—P2—C1—Cl1 | −57.47 (14) |
| O2ii—Cd2—O12—P1 | 165.3 (4) | O22—P2—C1—Cl2 | −57.06 (14) |
| O12—P1—O13—C13 | −18.2 (2) | O21—P2—C1—Cl2 | 176.19 (10) |
| O11—P1—O13—C13 | −149.7 (2) | N1—P2—C1—Cl2 | 59.21 (14) |
| C1—P1—O13—C13 | 97.1 (2) | O22—P2—C1—P1 | 64.58 (14) |
| O22—P2—O21—Cd1 | −84.54 (15) | O21—P2—C1—P1 | −62.16 (14) |
| N1—P2—O21—Cd1 | 150.72 (12) | N1—P2—C1—P1 | −179.15 (11) |
| C1—P2—O21—Cd1 | 33.90 (15) | C6—N1—C2—C3 | 55.4 (3) |
| O11—Cd1—O21—P2 | −10.33 (14) | P2—N1—C2—C3 | −131.8 (2) |
| O11i—Cd1—O21—P2 | 179.17 (13) | C5—O4—C3—C2 | 59.5 (3) |
| O21i—Cd1—O21—P2 | 90.11 (13) | N1—C2—C3—O4 | −56.6 (3) |
| O1—Cd1—O21—P2 | −95.53 (13) | C3—O4—C5—C6 | −60.6 (3) |
| O22—P2—N1—C6 | −155.22 (19) | C2—N1—C6—C5 | −55.9 (3) |
| O21—P2—N1—C6 | −24.6 (2) | P2—N1—C6—C5 | 131.4 (2) |
| C1—P2—N1—C6 | 90.2 (2) | O4—C5—C6—N1 | 58.1 (3) |
| O22—P2—N1—C2 | 32.9 (2) |
Symmetry codes: (i) −x+1, y, −z+1/2; (ii) −x+1, −y+1, −z.
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1A···O2iii | 0.84 | 2.06 | 2.849 (2) | 156 |
| O1—H1B···O22iv | 0.88 | 2.12 | 2.990 (2) | 170 |
| O2—H2A···O21i | 0.86 | 2.04 | 2.844 (2) | 155 |
| O2—H2B···O22 | 0.86 | 1.84 | 2.662 (2) | 159 |
| O3—H3A···O22 | 0.83 | 2.03 | 2.773 (2) | 149 |
| O3—H3B···O13v | 0.90 | 1.87 | 2.745 (2) | 163 |
Symmetry codes: (iii) −x+1, y−1, −z+1/2; (iv) x, y−1, z; (i) −x+1, y, −z+1/2; (v) x, y+1, z.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: ER2063).
References
- Brandenburg, K. (2005). DIAMOND Crystal Impact GbR, Bonn, Germany.
- Clearfield, A. (1998). Progress in Inorganic Chemistry: Metal Phosphonate Chemistry, edited by K. D. Karlin, Vol. 47, pp. 371–510, and references therein. New York: Wiley.
- Clearfield, A., Krishnamohan Sharma, C. V. & Zhang, B. (2001). Chem. Mater.13, 3099–3112.
- Fu, R., Hu, S. & Wu, X. (2007). Cryst. Crowth. Des.7, 1134–1144.
- Jokiniemi, J., Peräniemi, S., Vepsäläinen, J. J. & Ahlgrén, M. (2008). CrystEngComm, 10, 1011–1017.
- Jokiniemi, J., Vuokila-Laine, E., Peräniemi, S., Vepsäläinen, J. J. & Ahlgrén, M. (2007). CrystEngComm, 9, 158–164.
- Kontturi, M., Peräniemi, S., Vepsäläinen, J. J. & Ahlgrén, M. (2005a). Acta Cryst. E61, m635–m637. [DOI] [PubMed]
- Kontturi, M., Peräniemi, S., Vepsäläinen, J. J. & Ahlgrén, M. (2005b). Acta Cryst. E61, m638–m640. [DOI] [PubMed]
- Kontturi, M., Vuokila-Laine, E., Peräniemi, S., Pakkanen, T. T., Vepsäläinen, J. J. & Ahlgrén, M. (2002). J. Chem. Soc. Dalton Trans. pp. 1969–1973.
- Man, S. P., Motevalli, M., Gardiner, S., Sullivan, A. & Wilson, J. (2006). Polyhedron, 25, 1017–1032.
- Nonius (1997). COLLECT Nonius BV, Delft, The Netherlands.
- Otwinowski, Z. & Minor, W. (1997). Methods in Enzymology, Vol. 276, Macromolecular Crystallography, Part A, edited by C. W. Carter Jr & R. M. Sweet, pp. 307–326. New York: Academic Press.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Ying, S.-M. & Mao, J.-G. (2006). J. Mol. Struct.783, 13–20.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks I, global. DOI: 10.1107/S160053680901527X/er2063sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S160053680901527X/er2063Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report