Abstract
Results of systematic base-substitution experiments suggest that the lambda repressor dimer, made of identical subunits, recognizes the "pseudo(2-fold)symmetric" operator sequence asymmetrically. Base substitutions within the consensus half of the operator affect binding more than base substitutions within the nonconsensus half of the operator. Furthermore, changing the nonconsensus base pairs to the consensus base pairs does not increase, but decreases, binding. Evidently, the two subunits of the lambda repressor dimer bind to the two halves of the operator differently. This is consistent with the recently determined crystal structure of the complex, which shows that the relative positioning of the amino acids to the DNA bases are slightly different in the two halves of the operator. The sequence-specific interactions indicated by the systematic base-substitution experiments correlate well with the locations of the specific contacts found in the complex. Thus, the amino acids of lambda repressor, mainly of alpha 3-helix and the N-terminus arm, seem to directly read-out the DNA sequence by forming specific hydrogen bonds and hydrophobic contacts to the DNA bases. The observed asymmetric recognition suggests that no recognition code governs amino acids and DNA bases in protein-DNA interactions.
Full text
PDF
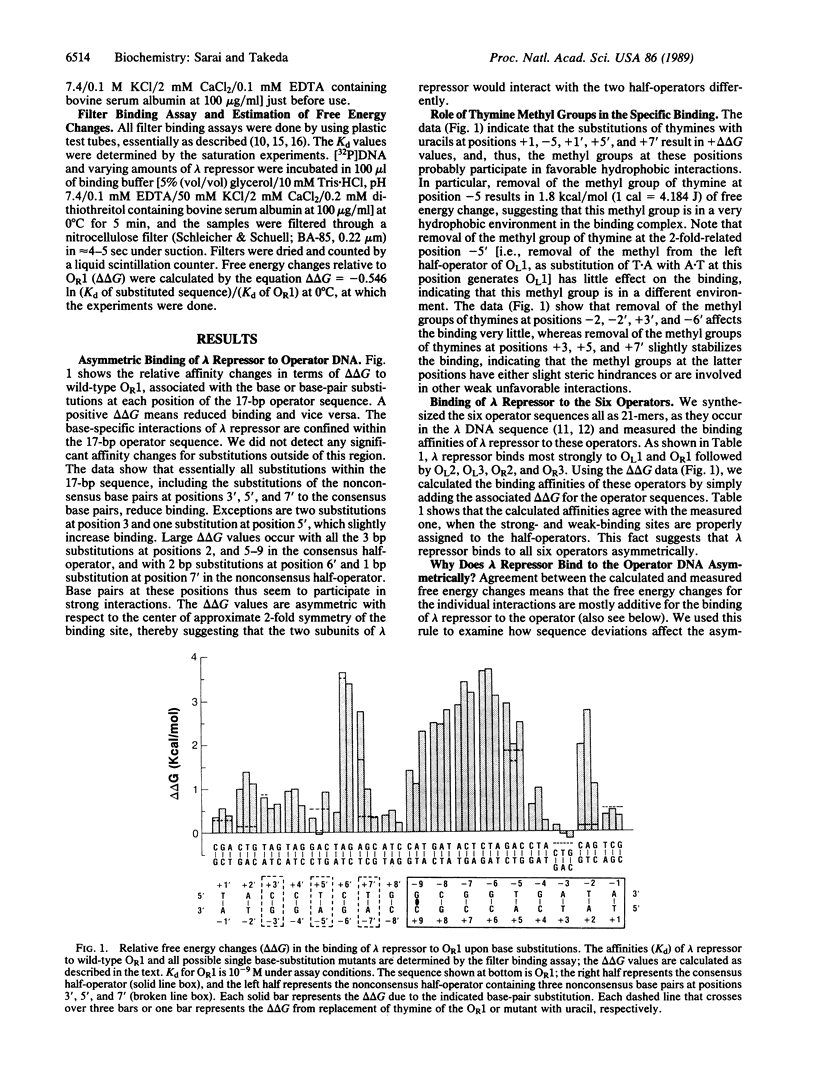

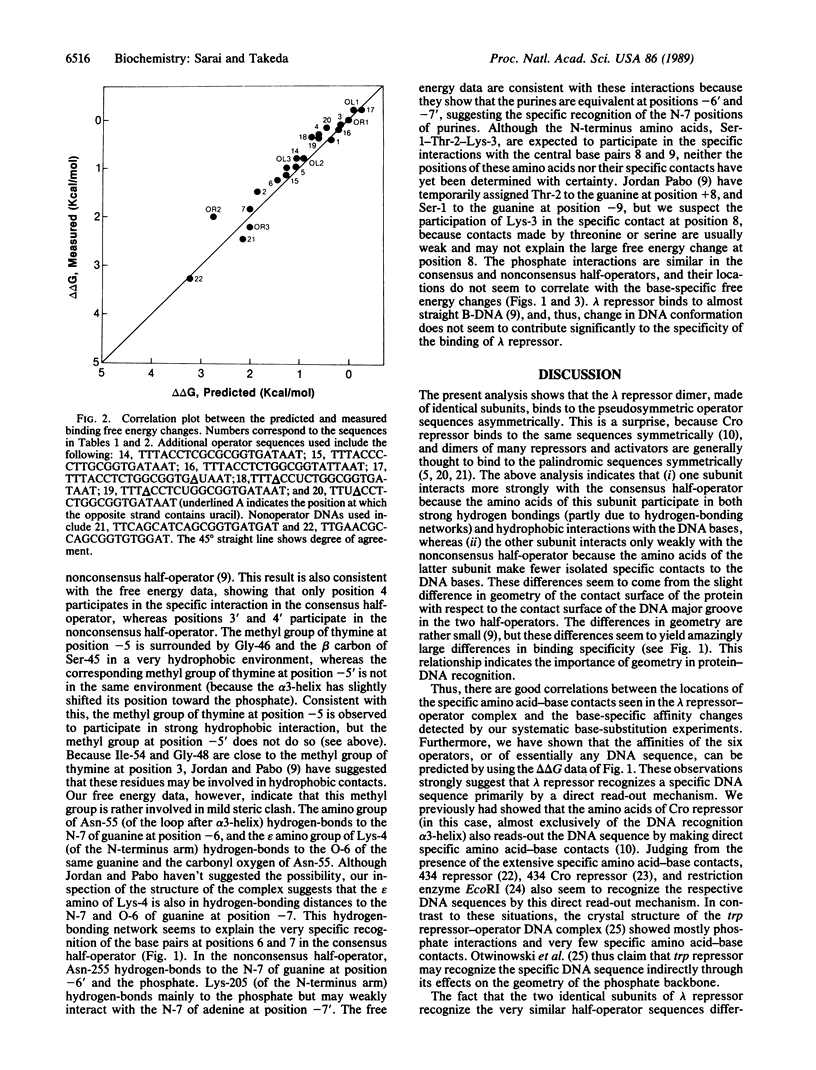

Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aggarwal A. K., Rodgers D. W., Drottar M., Ptashne M., Harrison S. C. Recognition of a DNA operator by the repressor of phage 434: a view at high resolution. Science. 1988 Nov 11;242(4880):899–907. doi: 10.1126/science.3187531. [DOI] [PubMed] [Google Scholar]
- Eliason J. L., Weiss M. A., Ptashne M. NH2-terminal arm of phage lambda repressor contributes energy and specificity to repressor binding and determines the effects of operator mutations. Proc Natl Acad Sci U S A. 1985 Apr;82(8):2339–2343. doi: 10.1073/pnas.82.8.2339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hecht M. H., Nelson H. C., Sauer R. T. Mutations in lambda repressor's amino-terminal domain: implications for protein stability and DNA binding. Proc Natl Acad Sci U S A. 1983 May;80(9):2676–2680. doi: 10.1073/pnas.80.9.2676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Humayun Z., Jeffrey A., Ptashne M. Completed DNA sequences and organization of repressor-binding sites in the operators of phage lambda. J Mol Biol. 1977 May 15;112(2):265–277. doi: 10.1016/s0022-2836(77)80143-5. [DOI] [PubMed] [Google Scholar]
- Johnson A. D., Pabo C. O., Sauer R. T. Bacteriophage lambda repressor and cro protein: interactions with operator DNA. Methods Enzymol. 1980;65(1):839–856. doi: 10.1016/s0076-6879(80)65078-2. [DOI] [PubMed] [Google Scholar]
- Johnson A. D., Poteete A. R., Lauer G., Sauer R. T., Ackers G. K., Ptashne M. lambda Repressor and cro--components of an efficient molecular switch. Nature. 1981 Nov 19;294(5838):217–223. doi: 10.1038/294217a0. [DOI] [PubMed] [Google Scholar]
- Johnson A., Meyer B. J., Ptashne M. Mechanism of action of the cro protein of bacteriophage lambda. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1783–1787. doi: 10.1073/pnas.75.4.1783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan S. R., Pabo C. O. Structure of the lambda complex at 2.5 A resolution: details of the repressor-operator interactions. Science. 1988 Nov 11;242(4880):893–899. doi: 10.1126/science.3187530. [DOI] [PubMed] [Google Scholar]
- Kim J. G., Takeda Y., Matthews B. W., Anderson W. F. Kinetic studies on Cro repressor-operator DNA interaction. J Mol Biol. 1987 Jul 5;196(1):149–158. doi: 10.1016/0022-2836(87)90517-1. [DOI] [PubMed] [Google Scholar]
- Lewis M., Jeffrey A., Wang J., Ladner R., Ptashne M., Pabo C. O. Structure of the operator-binding domain of bacteriophage lambda repressor: implications for DNA recognition and gene regulation. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 1):435–440. doi: 10.1101/sqb.1983.047.01.051. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Ptashne M., Backman K., Kield D., Flashman S., Jeffrey A., Maurer R. Recognition sequences of repressor and polymerase in the operators of bacteriophage lambda. Cell. 1975 Jun;5(2):109–113. doi: 10.1016/0092-8674(75)90018-5. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClarin J. A., Frederick C. A., Wang B. C., Greene P., Boyer H. W., Grable J., Rosenberg J. M. Structure of the DNA-Eco RI endonuclease recognition complex at 3 A resolution. Science. 1986 Dec 19;234(4783):1526–1541. doi: 10.1126/science.3024321. [DOI] [PubMed] [Google Scholar]
- Otwinowski Z., Schevitz R. W., Zhang R. G., Lawson C. L., Joachimiak A., Marmorstein R. Q., Luisi B. F., Sigler P. B. Crystal structure of trp repressor/operator complex at atomic resolution. Nature. 1988 Sep 22;335(6188):321–329. doi: 10.1038/335321a0. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Krovatin W., Jeffrey A., Sauer R. T. The N-terminal arms of lambda repressor wrap around the operator DNA. Nature. 1982 Jul 29;298(5873):441–443. doi: 10.1038/298441a0. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Lewis M. The operator-binding domain of lambda repressor: structure and DNA recognition. Nature. 1982 Jul 29;298(5873):443–447. doi: 10.1038/298443a0. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Ptashne M., Jeffrey A., Johnson A. D., Maurer R., Meyer B. J., Pabo C. O., Roberts T. M., Sauer R. T. How the lambda repressor and cro work. Cell. 1980 Jan;19(1):1–11. doi: 10.1016/0092-8674(80)90383-9. [DOI] [PubMed] [Google Scholar]
- Riggs A. D., Suzuki H., Bourgeois S. Lac repressor-operator interaction. I. Equilibrium studies. J Mol Biol. 1970 Feb 28;48(1):67–83. doi: 10.1016/0022-2836(70)90219-6. [DOI] [PubMed] [Google Scholar]
- Takeda Y., Ohlendorf D. H., Anderson W. F., Matthews B. W. DNA-binding proteins. Science. 1983 Sep 9;221(4615):1020–1026. doi: 10.1126/science.6308768. [DOI] [PubMed] [Google Scholar]
- Takeda Y., Sarai A., Rivera V. M. Analysis of the sequence-specific interactions between Cro repressor and operator DNA by systematic base substitution experiments. Proc Natl Acad Sci U S A. 1989 Jan;86(2):439–443. doi: 10.1073/pnas.86.2.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolberger C., Dong Y. C., Ptashne M., Harrison S. C. Structure of a phage 434 Cro/DNA complex. Nature. 1988 Oct 27;335(6193):789–795. doi: 10.1038/335789a0. [DOI] [PubMed] [Google Scholar]
- Woodbury C. P., Jr, Hagenbüchle O., von Hippel P. H. DNA site recognition and reduced specificity of the Eco RI endonuclease. J Biol Chem. 1980 Dec 10;255(23):11534–11548. [PubMed] [Google Scholar]


