Abstract
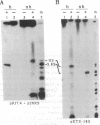
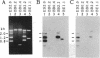
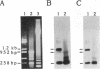
U3 small nuclear RNA is hydrogen-bonded to high molecular weight nucleolar RNA and can be isolated from greater than 60S pre-ribosomal ribonucleoprotein particles, suggesting that it is involved in processing of ribosomal RNA precursors (pre-rRNA) or in ribosome biogenesis. Here we have used in vivo psoralen cross-linking to identify the region of pre-rRNA interacting with U3 RNA. Quantitative hybridization selection/depletion experiments with clones of rRNA-encoding DNA (rDNA) and cross-linked nuclear RNA showed that all of the cross-linked U3 RNA was associated with a region that includes the external transcribed spacer (ETS) at the 5' end of the human rRNA precursor. To further identify the site of interaction within the approximately 3.7-kilobase ETS, Southern blots of rDNA clones were sandwich-hybridized with cross-linked RNA and then probed for cross-linked U3 RNA. These experiments showed that U3 RNA was cross-linked to a 258-base sequence between nucleotides +438 and +695, just downstream of the ETS early cleavage site (+414). The localization of U3 to this region of the rRNA precursor was not expected from previous models for a base-paired U3-rRNA interaction and suggests that U3 plays a role in the initial pre-rRNA processing event.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alwine J. C., Kemp D. J., Parker B. A., Reiser J., Renart J., Stark G. R., Wahl G. M. Detection of specific RNAs or specific fragments of DNA by fractionation in gels and transfer to diazobenzyloxymethyl paper. Methods Enzymol. 1979;68:220–242. doi: 10.1016/0076-6879(79)68017-5. [DOI] [PubMed] [Google Scholar]
- Bachellerie J. P., Michot B., Raynal F. Recognition signals for mouse pre-rRNA processing. A potential role for U3 nucleolar RNA. Mol Biol Rep. 1983 May;9(1-2):79–86. doi: 10.1007/BF00777477. [DOI] [PubMed] [Google Scholar]
- Bourbon H., Michot B., Hassouna N., Feliu J., Bachellerie J. P. Sequence and secondary structure of the 5' external transcribed spacer of mouse pre-rRNA. DNA. 1988 Apr;7(3):181–191. doi: 10.1089/dna.1988.7.181. [DOI] [PubMed] [Google Scholar]
- Bowman L. H., Rabin B., Schlessinger D. Multiple ribosomal RNA cleavage pathways in mammalian cells. Nucleic Acids Res. 1981 Oct 10;9(19):4951–4966. doi: 10.1093/nar/9.19.4951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calvet J. P., Meyer L. M., Pederson T. Small nuclear RNA U2 is base-paired to heterogeneous nuclear RNA. Science. 1982 Jul 30;217(4558):456–458. doi: 10.1126/science.6178162. [DOI] [PubMed] [Google Scholar]
- Calvet J. P., Myers J. A. In-vivo secondary structure analysis of the small nuclear RNA U1 using psoralen cross-linking. J Mol Biol. 1987 Oct 5;197(3):543–553. doi: 10.1016/0022-2836(87)90563-8. [DOI] [PubMed] [Google Scholar]
- Calvet J. P., Pederson T. Base-pairing interactions between small nuclear RNAs and nuclear RNA precursors as revealed by psoralen cross-linking in vivo. Cell. 1981 Nov;26(3 Pt 1):363–370. doi: 10.1016/0092-8674(81)90205-1. [DOI] [PubMed] [Google Scholar]
- Chabot B., Black D. L., LeMaster D. M., Steitz J. A. The 3' splice site of pre-messenger RNA is recognized by a small nuclear ribonucleoprotein. Science. 1985 Dec 20;230(4732):1344–1349. doi: 10.1126/science.2933810. [DOI] [PubMed] [Google Scholar]
- Choi Y. C. Structural organization of ribosomal RNAs from Novikoff hepatoma. II. Characterization of possible binding sites of 5 S rRNA and 5.8 S rRNA to 28 S rRNA. J Biol Chem. 1985 Oct 15;260(23):12773–12779. [PubMed] [Google Scholar]
- Cowley B. D., Jr, Smardo F. L., Jr, Grantham J. J., Calvet J. P. Elevated c-myc protooncogene expression in autosomal recessive polycystic kidney disease. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8394–8398. doi: 10.1073/pnas.84.23.8394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craig N., Kass S., Sollner-Webb B. Nucleotide sequence determining the first cleavage site in the processing of mouse precursor rRNA. Proc Natl Acad Sci U S A. 1987 Feb;84(3):629–633. doi: 10.1073/pnas.84.3.629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crouch R. J., Kanaya S., Earl P. L. A model for the involvement of the small nucleolar RNA (U3) in processing eukaryotic ribosomal RNA. Mol Biol Rep. 1983 May;9(1-2):75–78. doi: 10.1007/BF00777476. [DOI] [PubMed] [Google Scholar]
- England T. E., Uhlenbeck O. C. 3'-terminal labelling of RNA with T4 RNA ligase. Nature. 1978 Oct 12;275(5680):560–561. doi: 10.1038/275560a0. [DOI] [PubMed] [Google Scholar]
- Epstein P., Reddy R., Busch H. Multiple states of U3 RNA in Novikoff hepatoma nucleoli. Biochemistry. 1984 Nov 6;23(23):5421–5425. doi: 10.1021/bi00318a007. [DOI] [PubMed] [Google Scholar]
- Gerbi S. A., Jeppesen C., Stebbins-Boaz B., Ares M., Jr Evolution of eukaryotic rRNA: constraints imposed by RNA interactions. Cold Spring Harb Symp Quant Biol. 1987;52:709–719. doi: 10.1101/sqb.1987.052.01.080. [DOI] [PubMed] [Google Scholar]
- Gurney T., Jr Characterization of mouse 45S ribosomal RNA subspecies suggests that the first processing cleavage occurs 600 +/- 100 nucleotides from the 5' end and the second 500 +/- 100 nucleotides from the 3' end of a 13.9 kb precursor. Nucleic Acids Res. 1985 Jul 11;13(13):4905–4919. doi: 10.1093/nar/13.13.4905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeppesen C., Stebbins-Boaz B., Gerbi S. A. Nucleotide sequence determination and secondary structure of Xenopus U3 snRNA. Nucleic Acids Res. 1988 Mar 25;16(5):2127–2148. doi: 10.1093/nar/16.5.2127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass S., Craig N., Sollner-Webb B. Primary processing of mammalian rRNA involves two adjacent cleavages and is not species specific. Mol Cell Biol. 1987 Aug;7(8):2891–2898. doi: 10.1128/mcb.7.8.2891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kupriyanova N. S., Timofeeva MYa 32S pre-rRNA processing: a dynamic model for interaction with U3RNA and structural rearrangements of spacer regions. Mol Biol Rep. 1988;13(2):91–96. doi: 10.1007/BF00539056. [DOI] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ochs R. L., Lischwe M. A., Spohn W. H., Busch H. Fibrillarin: a new protein of the nucleolus identified by autoimmune sera. Biol Cell. 1985;54(2):123–133. doi: 10.1111/j.1768-322x.1985.tb00387.x. [DOI] [PubMed] [Google Scholar]
- Parker K. A., Bruzik J. P., Steitz J. A. An in vitro interaction between the human U3 snRNP and 28S rRNA sequences near the alpha-sarcin site. Nucleic Acids Res. 1988 Nov 25;16(22):10493–10509. doi: 10.1093/nar/16.22.10493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker K. A., Steitz J. A. Structural analysis of the human U3 ribonucleoprotein particle reveal a conserved sequence available for base pairing with pre-rRNA. Mol Cell Biol. 1987 Aug;7(8):2899–2913. doi: 10.1128/mcb.7.8.2899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porter G. L., Brennwald P. J., Holm K. A., Wise J. A. The sequence of U3 from Schizosaccharomyces pombe suggests structural divergence of this snRNA between metazoans and unicellular eukaryotes. Nucleic Acids Res. 1988 Nov 11;16(21):10131–10152. doi: 10.1093/nar/16.21.10131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prestayko A. W., Tonato M., Busch H. Low molecular weight RNA associated with 28 s nucleolar RNA. J Mol Biol. 1970 Feb 14;47(3):505–515. doi: 10.1016/0022-2836(70)90318-9. [DOI] [PubMed] [Google Scholar]
- Reimer G., Pollard K. M., Penning C. A., Ochs R. L., Lischwe M. A., Busch H., Tan E. M. Monoclonal autoantibody from a (New Zealand black x New Zealand white)F1 mouse and some human scleroderma sera target an Mr 34,000 nucleolar protein of the U3 RNP particle. Arthritis Rheum. 1987 Jul;30(7):793–800. doi: 10.1002/art.1780300709. [DOI] [PubMed] [Google Scholar]
- Shumard C. M., Eichler D. C. Ribosomal RNA processing. Limited cleavages of mouse preribosomal RNA by a nucleolar endoribonuclease include the early +650 processing site. J Biol Chem. 1988 Dec 25;263(36):19346–19352. [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Tague B. W., Gerbi S. A. Processing of the large rRNA precursor: two proposed categories of RNA-RNA interactions in eukaryotes. J Mol Evol. 1984;20(3-4):362–367. doi: 10.1007/BF02104742. [DOI] [PubMed] [Google Scholar]
- Thompson J. F., Hearst J. E. Structure of E. coli 16S RNA elucidated by psoralen crosslinking. Cell. 1983 Apr;32(4):1355–1365. doi: 10.1016/0092-8674(83)90316-1. [DOI] [PubMed] [Google Scholar]
- Tollervey D. A yeast small nuclear RNA is required for normal processing of pre-ribosomal RNA. EMBO J. 1987 Dec 20;6(13):4169–4175. doi: 10.1002/j.1460-2075.1987.tb02763.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wellauer P. K., Dawid I. B. Isolation and sequence organization of human ribosomal DNA. J Mol Biol. 1979 Mar 5;128(3):289–303. doi: 10.1016/0022-2836(79)90089-5. [DOI] [PubMed] [Google Scholar]
- Wellauer P. K., Dawid I. B., Kelley D. E., Perry R. P. Secondary structure maps of ribosomal RNA. II. Processing of mouse L-cell ribosomal RNA and variations in the processing pathway. J Mol Biol. 1974 Oct 25;89(2):397–407. doi: 10.1016/0022-2836(74)90527-0. [DOI] [PubMed] [Google Scholar]
- Zieve G., Penman S. Small RNA species of the HeLa cell: metabolism and subcellular localization. Cell. 1976 May;8(1):19–31. doi: 10.1016/0092-8674(76)90181-1. [DOI] [PubMed] [Google Scholar]