Abstract
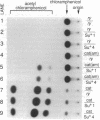
Drosophila melanogaster strains with a stably incorporated amber suppressor tRNA gene have been generated. A tRNATyr gene was site specifically mutated to produce an anticodon sequence that recognizes the amber codon and then introduced into Drosophila by using P-element-mediated transformation. Transformants from four integration events were recovered. Two integrations resulted in both male and female sterility, whereas the other two resulted in male sterility but female fertility. Strains derived from the two female-fertile integration events were shown to have a low level of amber-suppressing activity by their ability to suppress an amber mutation in a chloramphenicol acetyltransferase gene.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alton N. K., Vapnek D. Nucleotide sequence analysis of the chloramphenicol resistance transposon Tn9. Nature. 1979 Dec 20;282(5741):864–869. doi: 10.1038/282864a0. [DOI] [PubMed] [Google Scholar]
- Capone J. P., Sedivy J. M., Sharp P. A., RajBhandary U. L. Introduction of UAG, UAA, and UGA nonsense mutations at a specific site in the Escherichia coli chloramphenicol acetyltransferase gene: use in measurement of amber, ochre, and opal suppression in mammalian cells. Mol Cell Biol. 1986 Sep;6(9):3059–3067. doi: 10.1128/mcb.6.9.3059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capone J. P., Sharp P. A., RajBhandary U. L. Amber, ochre and opal suppressor tRNA genes derived from a human serine tRNA gene. EMBO J. 1985 Jan;4(1):213–221. doi: 10.1002/j.1460-2075.1985.tb02338.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chanet R., Magana-Schwencke N., Fabre F. Potential DNA-binding domains in the RAD18 gene product of Saccharomyces cerevisiae. Gene. 1988 Dec 30;74(2):543–547. doi: 10.1016/0378-1119(88)90187-4. [DOI] [PubMed] [Google Scholar]
- Doerig R. E., Suter B., Gray M., Kubli E. Identification of an amber nonsense mutation in the rosy516 gene by germline transformation of an amber suppressor tRNA gene. EMBO J. 1988 Aug;7(8):2579–2584. doi: 10.1002/j.1460-2075.1988.tb03107.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gelbart W. M., Irish V. F., St Johnston R. D., Hoffmann F. M., Blackman R. K., Segal D., Posakony L. M., Grimaila R. The decapentaplegic gene complex in Drosophila melanogaster. Cold Spring Harb Symp Quant Biol. 1985;50:119–125. [PubMed] [Google Scholar]
- Ho Y. S., Norton G. P., Palese P., Dozy A. M., Kan Y. W. Expression and function of suppressor tRNA genes in mammalian cells. Cold Spring Harb Symp Quant Biol. 1986;51(Pt 2):1033–1040. doi: 10.1101/sqb.1986.051.01.119. [DOI] [PubMed] [Google Scholar]
- Hudziak R. M., Laski F. A., RajBhandary U. L., Sharp P. A., Capecchi M. R. Establishment of mammalian cell lines containing multiple nonsense mutations and functional suppressor tRNA genes. Cell. 1982 Nov;31(1):137–146. doi: 10.1016/0092-8674(82)90413-5. [DOI] [PubMed] [Google Scholar]
- Hutchison C. A., 3rd, Phillips S., Edgell M. H., Gillam S., Jahnke P., Smith M. Mutagenesis at a specific position in a DNA sequence. J Biol Chem. 1978 Sep 25;253(18):6551–6560. [PubMed] [Google Scholar]
- Karess R. E., Rubin G. M. Analysis of P transposable element functions in Drosophila. Cell. 1984 Aug;38(1):135–146. doi: 10.1016/0092-8674(84)90534-8. [DOI] [PubMed] [Google Scholar]
- Kudo I., Leineweber M., RajBhandary U. L. Site-specific mutagenesis on cloned DNAs: generation of a mutant of Escherichia coli tyrosine suppressor tRNA in which the sequence G-T-T-C corresponding to the universal G-T-pseudouracil-C sequence of tRNAs is changed to G-A-T-C. Proc Natl Acad Sci U S A. 1981 Aug;78(8):4753–4757. doi: 10.1073/pnas.78.8.4753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laski F. A., Belagaje R., Hudziak R. M., Capecchi M. R., Norton G. P., Palese P., RajBhandary U. L., Sharp P. A. Synthesis of an ochre suppressor tRNA gene and expression in mammalian cells. EMBO J. 1984 Nov;3(11):2445–2452. doi: 10.1002/j.1460-2075.1984.tb02154.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laski F. A., Belagaje R., RajBhandary U. L., Sharp P. A. An amber suppressor tRNA gene derived by site-specific mutagenesis: cloning and function in mammalian cells. Proc Natl Acad Sci U S A. 1982 Oct;79(19):5813–5817. doi: 10.1073/pnas.79.19.5813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laski F. A., Rio D. C., Rubin G. M. Tissue specificity of Drosophila P element transposition is regulated at the level of mRNA splicing. Cell. 1986 Jan 17;44(1):7–19. doi: 10.1016/0092-8674(86)90480-0. [DOI] [PubMed] [Google Scholar]
- Mismer D., Rubin G. M. Analysis of the promoter of the ninaE opsin gene in Drosophila melanogaster. Genetics. 1987 Aug;116(4):565–578. doi: 10.1093/genetics/116.4.565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullins M. C., Rio D. C., Rubin G. M. cis-acting DNA sequence requirements for P-element transposition. Genes Dev. 1989 May;3(5):729–738. doi: 10.1101/gad.3.5.729. [DOI] [PubMed] [Google Scholar]
- Pelham H. R. A regulatory upstream promoter element in the Drosophila hsp 70 heat-shock gene. Cell. 1982 Sep;30(2):517–528. doi: 10.1016/0092-8674(82)90249-5. [DOI] [PubMed] [Google Scholar]
- Razin A., Hirose T., Itakura K., Riggs A. D. Efficient correction of a mutation by use of chemically synthesized DNA. Proc Natl Acad Sci U S A. 1978 Sep;75(9):4268–4270. doi: 10.1073/pnas.75.9.4268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sedivy J. M., Capone J. P., RajBhandary U. L., Sharp P. A. An inducible mammalian amber suppressor: propagation of a poliovirus mutant. Cell. 1987 Jul 31;50(3):379–389. doi: 10.1016/0092-8674(87)90492-2. [DOI] [PubMed] [Google Scholar]
- Temple G. F., Dozy A. M., Roy K. L., Kan Y. W. Construction of a functional human suppressor tRNA gene: an approach to gene therapy for beta-thalassaemia. Nature. 1982 Apr 8;296(5857):537–540. doi: 10.1038/296537a0. [DOI] [PubMed] [Google Scholar]
- Wills N., Gesteland R. F., Karn J., Barnett L., Bolten S., Waterston R. H. The genes sup-7 X and sup-5 III of C. elegans suppress amber nonsense mutations via altered transfer RNA. Cell. 1983 Jun;33(2):575–583. doi: 10.1016/0092-8674(83)90438-5. [DOI] [PubMed] [Google Scholar]
- Young J. F., Capecchi M., Laski F. A., RajBhandary U. L., Sharp P. A., Palese P. Measurement of suppressor transfer RNA activity. Science. 1983 Aug 26;221(4613):873–875. doi: 10.1126/science.6308765. [DOI] [PubMed] [Google Scholar]