Abstract
Expression of dominant-negative (DN) versions of the Drosophila ortholog of the tumor suppressor p53 extends fly life span in a Calorie Restriction (CR) dependent manner. DN-Dmp53 expression furthermore leads to reduction of Drosophila insulin-like peptide (dILP) 2 mRNA levels and a decrease in insulin/insulin-like growth factor-signaling activity (IIS) in the fly fat body. It is unclear by which mechanisms DN-Dmp53 extends longevity, and whether modulation of insulin-signaling activity plays a pivotal role in life span regulation by Dmp53. Here we show that life span extension due to DN-Dmp53 expression is likely due to reduction of Dmp53 activity and that decreased Dmp53 activity does not extend life span when dILP2 is concomitantly over expressed. Furthermore, extended longevity due to DN-Dmp53 expression does not further extend the life span of flies over expressing the IIS associated transcription factor dFoxO, indicating that DN-Dmp53-dependent life span extension may be related to IIS. However, reduction of dFoxO levels does not decrease DN-Dmp53-dependent longevity extension. Interestingly, when DN-Dmp53 is expressed in flies lacking the translation initiation controlling factor Thor/4E-BP, the downstream target of dTOR signaling, no increase in life span is observed. Taken together, these data suggest that Dmp53 may affect life span by differentially engaging the IIS and dTor pathways.
Keywords: p53, Dmp53, calorie restriction, TOR, 4E-BP, Thor, Drosophila melanogaster
Introduction
Calorie restriction (CR), a technique that reduces calorie intake, has long been known to increase life span of mammals and of organisms as diverse as yeast, worms and flies (Masoro 2005). Mechanistically, CR may in fact be Dietary Restriction (DR), the specific reduction of micro-nutrients. In flies, the specific reduction of yeast extract in the food source (Mair et al. 2005), and especially the reduction of essential amino acids (Grandison et al. 2009), has been shown to underlie the effects of CR/DR.
CR in D. melanogaster has been associated with a decrease in the level of the histone deacetylase Rpd3 and an increase in the level of the sirtuin dSir2 (Rogina et al. 2002). Consequently, reducing Rpd3 function (Rogina et al. 2002) or increasing dSir2 activity (Rogina and Helfand 2004) increases life span in flies. The life span extending effects of CR in flies are dependent on dSir2, as dSir2 null flies do not show extended life span under CR conditions, and CR does not further extend the longer life span of flies over expressing dSir2 (Rogina and Helfand 2004).
Interestingly, mammalian Sir2 (SirT1) deacetylates diverse targets such as the DNA repair protein Ku70 (Cohen et al. 2004), the mitochondrial biogenesis controlling transcriptional co-factor PGC-1α (Nemoto et al. 2005), the forkhead transcription factor FoxO (Brunet et al. 2004; Daitoku et al. 2004; van der Horst et al. 2004) and the tumor suppressor p53 (Langley et al. 2002; Luo et al. 2001; Vaziri et al. 2001). Deacetylation of p53 by Sir2 inhibits p53 activity, suggesting that some of the Sir2 life span-extending effects might be mediated by inhibition of p53. We have recently shown that Drosophila dSir2 has similar functionality as mammalian SirT1. dSir2 binds to and deacetylates Drosophila melanogaster p53 (Dmp53), thereby inhibiting its activity (Bauer et al. 2009).
Altering Dmp53 function through the expression of different dominant-negative (DN) versions of D. melanogaster p53 (Dmp53) in the adult nervous system has been shown to extend the flies' life span by up to 26% (Bauer et al. 2005). These life span extension effects are not additive to the effects of CR, suggesting that DN-Dmp53 may be part of the CR pathway downstream of dSir2 (Bauer et al. 2009; Bauer et al. 2005). Interestingly, it is sufficient to express DN-Dmp53 in only the fourteen insulin-producing cells of the fly brain to achieve a similar life span increase as with pan-neuronal expression (Bauer et al. 2007). Importantly, DN-Dmp53-dependent life span extension is associated with a reduction in the levels of Drosophila insulin-like peptide 2 (dILP2) and a decrease in the activity of the insulin/insulin-like signaling pathway (IIS) in the fly fat body (Bauer et al. 2007). These data suggest that a reduction of IIS, or parts of IIS, may be related to DN-Dmp53-dependent longevity extension, and thus possibly play a role in CR-dependent life span regulation.
Here we further test this hypothesis by expressing DN-Dmp53 in mutants deficient in aspects of IIS. Surprisingly, we found that the IIS transcription factor dFoxO interacts with DN-Dmp53-dependent life span extension, but is not required for it. Instead, longevity control by Dmp53 requires downstream components of the related Target of Rapamycin (TOR) signaling pathway. Our data suggest that some elements of the TOR pathway, such as the translational inhibitor 4E-BP/Thor may be required for the life span extending effects of reduced Dmp53 activity and CR.
Materials and Methods
Fly culture and strains
All flies were kept in a humidified (50%), temperature-controlled incubator with 12 hour on/off light cycle at 25°C in vials containing standard cornmeal medium (Rogina and Helfand 2004). The ELAV-GeneSwitch line was from H. Keshishian (Yale University, New Haven, CT), S1-32, S1-106, dFoxO21 and dFoxO25 were from M. Tatar (Brown University, Providence, RI), the dILP2 driver line was from P. Shen (University of Georgia, Athens, GA) and the PH-domain-GFP fusion expressing line tGPH from B. Edgar (Fred Hutchinson Cancer Research Center, Seattle, WA). The 4E-BP lines Thor1, Thor1Δ and Thor1RV were from B. Zid (California Institute of Technology, Pasadena, CA). These fly strains were generated through remobilization of a P-element in the 4E-BP/Thor locus (Thor1) (Bernal et al. 2004). Remobilization of the P-element resulted in the isolation of a precise excision line, and thus a revertant line (Thor1RV), and a deletion of 4E-BP (Thor1Δ) (Bernal et al. 2004). These two stocks are in the same genetic background, and thus Thor1RV can serve as a genetically matched control for Thor1Δ. All other lines were from the Bloomington Drosophila Stockcenter at Indiana University (Bloomington, IN). All Dmp53 lines (UAS-Dmp53-R155H, UAS-Dmp53-H159N (Ollmann et al. 2000), UAS-Dmp53-259H (Brodsky et al. 2000) and Dmp53NS (Sogame et al. 2003)) were originally generated in a y1 w1118 or w1118 background.
For expression in a Dmp53 null background, ELAV-Switch and Dmp53-259H constructs were recombined onto Dmp53NS (Dmp53 null) chromosomes. The resulting fly lines (seven independent recombinant driver lines and five independent Dmp53-259H lines) were verified by PCR for the presence of the null allele and the UAS or GAL4 constructs, respectively. For expression in dFoxO null backgrounds, 2nd chromosome DN-Dmp53 (Dmp53-R155H and Dmp53-H159N) constructs were combined using classical Drosophila genetics with the dFoxO21 null alleles on the third chromosome, while the dILP2-Gal4 driver was combined with the dFoxO25 allele. For expression in 4E-BP null backgrounds, the 3rd chromosome Dmp53-259H was combined with the 2nd chromosome 4E-BP alleles.
Life span analysis
Flies were collected under light anesthesia, randomly divided into treatment groups and housed at a density of 25 males and 25 females each per vial. At least ten such vials were used per treatment as per (Bauer et al. 2007). Flies were passed every other day and the number of dead flies recorded.
All life span experiments were performed on regular cornmeal food (2% yeast, 10% sucrose, 5% cornmeal (all w/v)) without added live yeast, except where indicated. For CR experiments, flies were raised on food containing 2%, 5%, 10%, 15%, 20%, respectively, yeast extract and sucrose (w/v), but no cornmeal. For sucrose DR experiments, yeast extract was kept constant at 15%, while varying the sucrose content (Chapman and Partridge 1996), while for yeast DR sucrose was kept constant at 15% and yeast extract was varied. For induction with the GeneSwitch system, RU486 (Sigma) was added directly to the food to a final concentration of 200μM. The same concentration of diluent was added to control food. RU486 was administered from the day of eclosion. For DN-Dmp53 expression in a Dmp53 null background, two of the recombinant driver lines were crossed to the recombinant UAS lines for a total of five independent driver/UAS combinations (out of a total possible 35). For expression with constitutive drivers, all lines were backcrossed to w1118 for 10 generations and isogenic controls were generated from the last backcross.
Fluorescent microscopy
For tGPH sub-cellular localization, adult fat body tissue was dissected into PBS from the abdomen of 10 day old female flies, as described (Bauer et al. 2007) and visualized using a Zeiss Axiovision Z1 fluorescent microscope with ApoTome™ optics (Zeiss). For each food condition at least five individual flies were used in at least two independent experiments.
Quantitative PCR
Total mRNA was isolated from at least 75 heads of 10-day old females using Trizol (Invitrogen) and further purified using the RNeasy kit (Qiagen). cDNA was generated with 0.5μg total mRNA in a 10μl reaction using the iScript cDNA synthesis kit (Bio-Rad). 0.8μl of the iScript reaction was used as QPCR template. QPCR was performed as described (Bauer et al. 2009) on an ABI 7500 Real-Time PCR machine using the ABI SYBR-Green PCR master mix following the manufacturers instructions. Each QPCR reaction was performed using four biological replicates in triplicate each.
Statistics
Statistical analyses, including log rank tests, were performed using the Prism suit of biostatistical software (GraphPad, San Diego). Maximum life span was calculated as the median age of the last surviving 10% of the population.
Results
Longevity extension through DN-Dmp53 is dependent on endogenous Dmp53
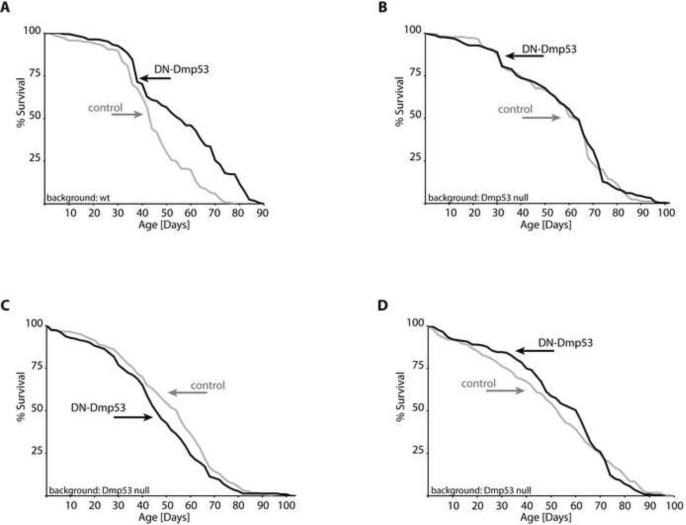
We have previously demonstrated that expression of DN-Dmp53 in the adult fly nervous system extends male and female life spans by up to 27% ((Bauer et al. 2005), Figure 1A). However, the use of the DN approach carries the risk of non-specific side effects, unrelated to a reduction in Dmp53 activity itself, which might account for the observed life span extension phenotype. To rule out these effects as the cause for life span extension, we expressed the DN-Dmp53 construct in the nervous system of adult flies in which the Dmp53 gene had been knocked out (Sogame et al. 2003). Expression of the DN-Dmp53 construct in the adult nervous system in the Dmp53 null background does not result in life span extension (Figure 1B, C and D). Averaged over five independent trials (Table 1), a mean life span extension of only ~4% is observed, compared to ~19% averaged over the four independent experiments with the same DN-Dmp53 construct shown in Table 1 in Dmp53 wt backgrounds (p=0.0232). When all experiments with the various DN-Dmp53 constructs in wt backgrounds are considered (shown in this report and previously published), mean life span extension averages to ~18% over thirteen trials (p=0.0018 for comparison with DN-Dmp53 expression in a Dmp53 null background). This indicates that endogenous Dmp53 is required for the life span extension phenotype of DN-Dmp53 expressing flies. As Dmp53 is normally expressed in the adult fly brain (data not shown), this suggests that it is indeed the down regulation of Dmp53 activity in the adult brain that is responsible for the life span extension, rather than non-specific effects of the DN construct.
Figure 1. Life span extension by DN-Dmp53 is dependent on endogenous Dmp53.
Survivorship curves of female ELAV-Switch-DN-Dmp53 flies are shown. (A) In a wt-Dmp53 background, expression of DN-Dmp53 (black) leads to extension of mean, median and maximum life span of 24%, 27% and 16%, respectively, over uninduced control flies (grey; for complete statistics, please see Table 1, line 2–7). (B) In a Dmp53 null background, induction of the DN-Dmp53 construct (black) does not lead to life span extension over uninduced controls (grey). (C and D) Two different independent experiments expressing DN-Dmp53 (black) in a Dmp53 null background also do not show life span extension over uninduced control (grey).
Table 1.
The effect of various proteins expressed in different expression patterns on female life span.
| Driver/Protein | Mean LS (vs. ctrl) | Mean LS extension | Median LS (vs. ctrl) | Median LS extension | Max LS (vs. ctrl) | Max LS extension | Number of flies (control;experimental) | χ 2 | p-value | |
|---|---|---|---|---|---|---|---|---|---|---|
| ELAV-Switch | ||||||||||
| 1 | Dmp53-259H | 44/40 | 10% | 42/38 | 11% | 68/66 | 3% | 242 247 |
4.197 | 0.0405 |
| 2 | Dmp53-259H | 56/45 | 24% | 56/44 | 27% | 84/72 | 16% | 136 161 |
37.41 | <0.0001 |
| 3 | Dmp53-259H (Dmp53 null background) | 57/57 | 0% | 64/63 | 2% | 88/84 | 5% | 176 250 |
0.267 7 |
0.6049 |
| 4 | Dmp53-259H (Dmp53 null background) | 54/51 | 6% | 62/54 | 15% | 82/86 | −5% | 211 228 |
0.025 3 |
0.8736 |
| 5 | Dmp53-259H (Dmp53 null background) | 49/44 | 11% | 48/46 | 4% | 84/80 | 5% | 190 185 |
6.405 | 0.0114 |
| 6 | Dmp53-259H (Dmp53 null background) | 54/49 | 10% | 60/52 | 15% | 82/80 | 3% | 200 208 |
11.41 | 0.0007 |
| 7 | Dmp53-259H (Dmp53 null background) | 47/51 | −8% | 46/56 | −18% | 76/82 | −7% | 222 220 |
4.55 | 0.0329 |
| 8 | Dmp53-H159N | 34/29 | 17% | 32/30 | 7% | 54/51 | 6% | 295 281 |
14.65 | 0.0001 |
| 9 | Dmp53-H159N | 41/36 | 14% | 42/34 | 24% | 68/56 | 21 | 245 247 |
18.7 | <0.0001 |
| 10 | Dmp53-R155H | 40/36 | 11% | 40/34 | 18% | 58/60 | −3% | 256 232 |
1.537 | 0.215 |
| 11 | Dmp53-R155H | 51/42 | 21% | 52/42 | 24% | 72/68 | 6% | 245 248 |
30.15 | <0.0001 |
| 12 | Dmp53-H159N (dFoxo21/+ background) | 35/31 | 13% | 34/32 | 6% | 52/50 | 4% | 244 249 |
12.76 | 0.0004 |
| 13 | Dmp53-R155H (dFoxo21/+ background) | 36/25 | 44% | 34/22 | 55% | 58/50 | 16% | 255 255 |
58.09 | <0.0001 |
| 14 | Dmp53-259H (Thor1RV background) | 63/50 | 26% | 68/52 | 31% | 80/70 | 14% | 247 239 |
114.6 | <0.0001 |
| 15 | Dmp53-259H (Thor1RV background) | 61/53 | 15% | 62/55 | 13% | 82/78 | 5% | 248 246 |
19.9 | <0.0001 |
| 16 | Dmp53-259H (Thor1Δbackground) | 46/47 | −2% | 48/48 | 0% | 66/68 | −3% | 247 232 |
0.353 9 |
0.5519 |
| 17 | Dmp53-259H (Thor1Δbackground) | 51/54 | −5% | 53/56 | −5% | 75/76 | −1% | 243 238 |
2.968 | 0.085 |
| 18 | Dmp53-259H (Thor1 background) | 57/45 | 27% | 62/44 | 41% | 78/74 | 5% | 251 243 |
47.52 | <0.0001 |
| 19 | Dmp53-259H/dFoxo | 40/33 | 21% | 39/30 | 30% | 60/56 | 7% | 241 251 |
31.07 | <0.0001 |
| S1-32 | ||||||||||
| 20 | dFoxo | 48/38 | 26% | 50/36 | 39% | 74/64 | 16% | 246 240 |
38.35 | <0.0001 |
| 21 | Dmp53-259H/dFoxo | 51/44 | 16% | 52/42 | 24% | 83/68 | 22% | 232 237 |
22.69 | <0.0001 |
| 22 | 4E-BP | 49/45 | 9% | 50/44 | 14% | 78/76 | 3% | 245 248 |
4.003 | 0.0454 |
| S1-106 | ||||||||||
| 23 | 4E-BP | 46/46 | 0% | 50/46 | 9% | 70/70 | 0% | 252 246 |
0.686 9 |
0.4063 |
| S1-32-ELAV-Switch | ||||||||||
| 24 | Dmp53-259H | 47/41 | 15% | 47/42 | 12% | 73/60 | 22% | 249 248 |
40.37 | <0.0001 |
| 25 | dFoxo | 52/38 | 37% | 54/36 | 50% | 84/68 | 24% | 216 244 |
71.69 | <0.0001 |
| 26 | Dmp53-259H/ dFoxo | 48/43 | 12% | 47/42 | 12% | 75/72 | 4% | 224 245 |
9.212 | 0.0024 |
| 27 | Dmp53-259H/ dFoxo | 43/49 | −12% | 42/48 | −12% | 72/81 | −11% | 237 230 |
18.86 | <0.0001 |
| 28 | Dmp53-259H/ dFoxo | 49/42 | 17% | 51/42 | 21% | 76/80 | −5% | 219 184 |
5.302 | 0.0213 |
| dILP2 | ||||||||||
| 29 | Dmp53-259H | 56/54 | 4% | 58/52 | 12% | 86/82 | 5% | 165 199 |
4.691 | 0.0303 |
| 30 | dILP2/ Dmp53-259H | 41/44 | −7% | 40/42 | −5% | 72/74 | −3% | 226 249 |
2.989 | 0.0838 |
| 31 | dILP2/ Dmp53-259H | 61/70 | −13% | 64/74 | −14% | 88/96 | −8% | 242 253 |
27.41 | <0.0001 |
| 32 | Dmp53-R155H (dFoxo21/dFoxo25 background) | 45/33 | 36% | 50/32 | 56% | 58/46 | 26% | 24 19 |
15.46 | <0.0001 |
The indicated drivers were crossed to the listed UAS flies. Backgrounds other than wt are shown in brackets. Offspring were collected and lifespan was determined on standard cornmeal food (except where indicated) in the presence of the inducing agent RU486 or diluent in the food. Only females are shown. Mean, median and maximum lifespan, log rank analysis, p-value, percent change in mean, median and maximum lifespan as compared to controls (w1118 for constitutive expression, without RU486 for GeneSwitch experiments), Chi-square and p-values derived from survivorship curves of each protein and driver are shown. Maximum life span was calculated as the median life span of the longest surviving 10%
DN-Dmp53 does not extend life span when dILP2 is over expressed in insulin-producing cells
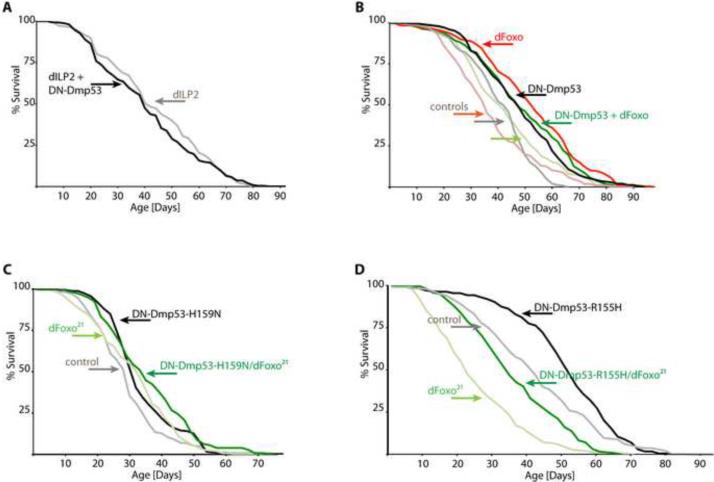
DN-Dmp53 activity reduction in the fourteen insulin-producing cells of the fly brain (IPC) is sufficient to extend life span and is associated with reduction of IIS activity in the fat body (Bauer et al. 2007). Furthermore, DN-Dmp53 long-lived flies have a 60% decrease in dILP2 mRNA levels. These data suggest that DN-Dmp53 may increase longevity through a decrease of dILP2 levels and down regulation of IIS in dILP2 responsive tissues. In order to test this hypothesis, we over expressed dILP2 and simultaneously expressed DN-Dmp53 in IPC. When DN-Dmp53 is thus expressed in dILP2 over expressing flies, flies no longer have extended longevity and may even have reduced life span (Figure 2A and Table 1), indicating that a reduction of dILP2 may be a key signal for longevity regulation by Dmp53.
Figure 2. Longevity increase by DN-Dmp53 requires dILP2, but not dFoxO.
Survivorship curves of female flies are shown (for complete statistical analysis, including p-values, mean, median and maximum life span, as well as replicate experiments, please refer to Table 1, lines 8–13, 24–31). (A) IPC-directed expression of DN-Dmp53 and simultaneous over expression of dILP2 using a dILP2 driver (black) fails to increase life span over that of control flies over expressing dILP2 only (grey). (B) Expression of DN-Dmp53 (black) or of dFoxO (red) using the combined ELAV-Switch/ S1-32 driver leads to significant life span extension (statistical analysis can be found in Table 1). Expression of both dFoxO and DN-Dmp53 together using the combined drivers (green) does not lead to additive effects on longevity extension (all respective controls are in lighter colors). (C–D) Expression of two different DN-Dmp53 constructs using the ELAV-Switch driver leads to extended life spans (grey: controls; black: DN-Dmp53; panel C: Dmp53-H159N; panel D: Dmp53-R155H). This life span extension is still observed when the DN-Dmp53 constructs are expressed in the dFoxO deficient background dFoxO21 (light green: controls; green: DN-Dmp53).
Life span extension by dFoxO and DN-Dmp53 expression are not additive
Since the DN-Dmp53 longevity phenotype can be prevented through over expression of dILP2, we hypothesized that down regulation of IIS in fat body cells may play an important role in DN-Dmp53-dependent life span extension. IIS down regulation leads to activation and nuclear accumulation of the dFoxO transcription factor. Over expression of dFoxO in the fly head fat body, but not the nervous system, leads to extended longevity (Hwangbo et al. 2004), while DN-Dmp53 only extends life span when expressed in neurons, but not in the fat body (Bauer et al. 2005). If DN-Dmp53 expression increases longevity through activation of dFoxO, combining these two interventions is not expected to have additive effects on life span extension. We therefore constructed flies that expressed dFoxO and DN-Dmp53 in both the adult nervous system and the fly head fat body. Expressing dFoxO alone (ELAV-Switch/S1-32 driver) leads to a ~37% increase in mean life span, while only expressing DN-Dmp53 leads to a ~15% increase in life span (Figure 2B, Table 1). When both DN-Dmp53 and dFoxO are expressed, the resulting longevity extension is not additive, as life span is only increased by ~12%, suggesting both interventions may share similar pathways (Figure 2B, Table 1). Similarly, when both constructs are expressed together with either driver (S1-32 or ELAV-Switch) no additive effects on life span are observed (Supplemental Figure S1 and Table 1).
Life span extension by DN-Dmp53 is not affected by reduction of dFoxO activity
The previous data suggest that DN-Dmp53 and dFoxO may engage similar longevity regulating pathways. To test whether dFoxO mediated the life span increasing effects of DN-Dmp53 expression, we over expressed DN-Dmp53 in flies in which dFoxO had been functionally disabled. Surprisingly, when DN-Dmp53 is expressed in the nervous system of flies with only one functional copy of dFoxO, the same life span extension as in wt flies is observed (Figure 2C and D, Table 1). Similar results are obtained when the dILP2 driver is used (data not shown). Even when the function of both copies of dFoxO is compromised, DN-Dmp53 is still capable of extending adult fly life span (Supplemental Figure S2). These data suggest that dFoxO may not be required for longevity regulation by Dmp53.
DN-Dmp53 dependent life span extension is abolished in 4E-BP deficient flies
The results presented above indicate that, while reduction in IIS in general may be an important downstream event, activation of dFoxO is dispensable for the longevity effects of Dmp53 activity reduction. Therefore, the signaling pathway leading to extended longevity may branch off the canonical IIS at a point before dFoxO activation. The TOR signaling pathway is a good candidate for such divergence of pathways. Reducing the activity of this amino acid responsive pathway in the fat body has been demonstrated to extend Drosophila life span in a CR-dependent manner (Kapahi et al. 2004). Importantly, the activity of the TOR pathway is regulated by the Tsc kinase complex, which in turn is controlled by the Akt protein kinase (Huang and Manning 2009), which also directly controls dFoxo activity (Gingras et al. 1998; Tatar et al. 2003).
We therefore investigated whether activity of the TOR pathway is required for life span extension by DN-Dmp53. Activation of TOR kinase leads to phosphorylation and inhibition of 4E-BP/Thor, an inhibitor of the translation initiation factor elF-4E (Thomas and Hall 1997). Interestingly, 4E-BP is furthermore a target of dFoxO transcriptional activity (Junger et al. 2003; Wang et al. 2005) and is thus an ideal target to investigate the interplay between canonical, dFoxO-dependent IIS and TOR signaling in Dmp53-dependent life span regulation.
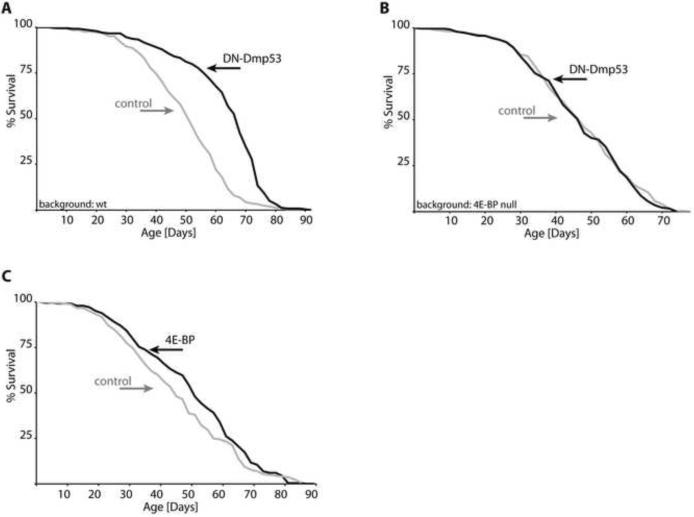
First, we investigated whether 4E-BP mRNA levels are changed in DN-Dmp53 expressing flies, but did not observe any significant changes (Supplemental Figure S3). Next, we investigated if the presence of 4E-BP was required for DN-Dmp53-dependent longevity extension. When DN-Dmp53 is expressed in the control (revertant) Thor1RV background, flies have mean extended life span of up to 26% over control flies. However, no extension of life span is observed when DN-Dmp53 is expressed in the deletion Thor1Δ background (Figure 3A and B, Table 1). Interestingly, flies carrying the original Thor1 allele of 4E-BP show life span extension upon DN-Dmp53 expression that is similar to flies carrying the Thor1RV allele, suggesting that even a small amount of 4E-BP activity is sufficient to mediate the longevity effects of DN-Dmp53 (Supplemental Figure S4). These data indicate that 4E-BP is required for DN-Dmp53-dependent life span extension.
Figure 3. 4E-BP is required, but not sufficient for DN-Dmp53-dependent longevity extension.
Survivorship curves of female flies are shown (for complete statistics, please see Table 1, lines 14–17, 22). (A) Pan-neuronal adult expression of DN-Dmp53 using the ELAV-Switch driver extends life span in a 4E-BP wild-type background (grey: controls; black: DN-Dmp53; background: Thor1RV, a precise P-element excision line of the P-element insertion Thor1 line). (B) When 4E-BP is deleted, DN-Dmp53 fails to increase longevity (grey: controls; black: DN-Dmp53; background: Thor1Δ, an imprecise P-element excision line of the P-element insertion Thor1 line). (C) Over expression of 4E-BP in the adult peri-cerebral fat body using the S1-32 driver marginally extends fly life span (grey: controls; black: 4E-BP).
Next, we investigated whether over expression of 4E-BP alone was sufficient to extend life span. When over expressed in the head fat body, 4E-BP marginally extended longevity (Figure 3C), but not when over expressed in the abdominal fat body (Supplemental Figure S5). This suggests that 4E-BP is required, but may not be sufficient for life span extension by DN-Dmp53.
Calorie restriction changes the activity of IIS in fat body cells
We have recently shown that Dmp53 regulates Drosophila life span in a pathway that is related to CR and downstream of dSir2 (Bauer et al. 2009). Furthermore, DN-Dmp53 expression is associated with a decrease in dILP2 levels and a reduction of IIS activity (Bauer et al. 2007). Together with the data presented here, this suggests that the events downstream of Dmp53 activity reduction include a decrease of IIS activity. Down regulation of IIS activity in the fat body upon DN-Dmp53 expression in the nervous system or IPC only is evidenced by increased nuclear localization of the downstream transcription factor dFoxO (Bauer et al. 2007). In addition, activity of phosphoinositide-3-kinase (PI3K) is reduced and accumulation of phosphatidylinositol-3, 4, 5-trisphosphate (PIP3) is decreased (Bauer et al. 2007). Since dFoxO is not required for the life span extending benefits of CR (Giannakou et al. 2008; Min et al. 2008), we examined whether more upstream components of IIS take part in the CR response. We measured PI3K activity in response to differing caloric food content by using GFP fused to a pleckstrin homology domain (PH) (Britton et al. 2002). When IIS is active, this GPH reporter protein binds to newly generated PIP3 and thus localizes to the plasma membrane (Bauer et al. 2007). Under very low and very high calorie conditions (2% and 20%, respectively), strong cytosolic staining is observed indicating low IIS activity (Supplemental Figure S6). This may represent a pathologic state in response to under feeding (2% food), or over feeding (20%), respectively. Strongest plasma membrane staining, and thus highest IIS activity, is observed at food conditions between 10% and 15%, while 5% (near the life span maximum) has slightly less plasma membrane staining. These data indicate that PI3K activity is affected by nutritional input.
Life span extension by dietary restriction is partially dependent on 4E-BP
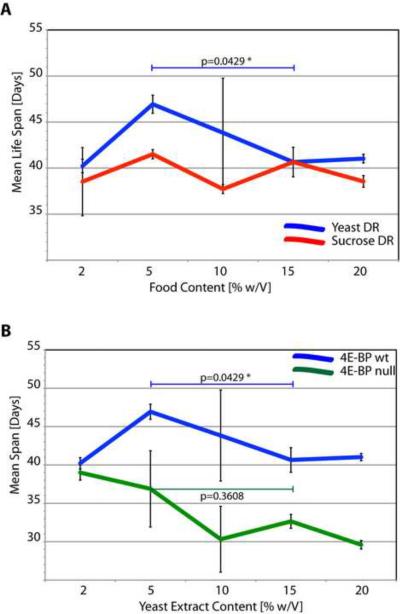
Our data show that life span increase by DN-Dmp53 is dependent on 4E-BP and the TOR pathway. We therefore examined whether the TOR pathway may also function as a component of the CR life span extending response. Since the TOR pathway is an amino acid responsive pathway, while IIS is a glucose responsive pathway, we thus investigated whether flies lacking functional 4E-BP responded to Dietary Restriction (DR), the reduction of specific food components, with extension of life span. First, we investigated the effect of either yeast extract content reduction (yeast DR) or sucrose content reduction (sucrose DR) on life span of wild type flies. As shown in Figure 5A, the control Thor1RV fly strain does not respond to a reduction in sucrose content with an extension of life span. However, when yeast extract content is reduced from 15% w/v to 5% w/v, control Thor1RV flies demonstrate a ~15% increase of longevity. At higher yeast content life span is unchanged, while at even lower yeast conditions life span is shortened. Thus, 5% yeast extract defines the point of optimal life span for this particular fly strain.
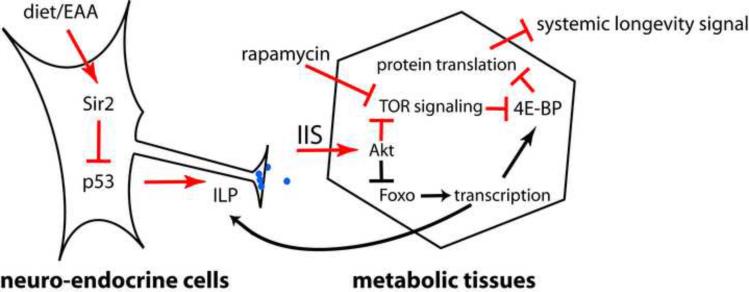
Figure 5. A model for the longevity regulating pathways connected to Dmp53.
Activity reduction of Dmp53 in the Drosophila insulin-producing cells leads to down regulation of dILP2 abundance by unknown mechanisms. Dmp53 activity can be reduced through deacetylation by dSir2, the activity of which is increased under CR/DR conditions. How dSir2 activity is increased by CR/DR is unclear, but it may be that the reduction of specific nutrient components, particularly essential amino acids (EAA), underlies the CR/DR effects.
Reduced Dmp53 activity inhibits PI3K and Akt activity in the fly fat body. As a consequence, dFoxo is predominantly nuclear to control transcription, but this event does not control life span. The longevity response is transmitted through inhibition of the TOR pathway and the translational inhibitor 4E-BP. dFoxo may modify the overall magnitude of the 4E-BP response and provide feedback regulation of dILPs (the signaling events underlying CR/DR are in red).
This model may also apply to mammalian systems, in which the relevant tissues and signaling pathways are conserved.
Next, we exposed flies lacking 4E-BP to yeast DR conditions. As shown in Figure 5B, Thor1Δ 4E-BP null flies generally have shortened life spans compared to wt flies, and do not respond to optimal yeast DR from 15% to 5% (as determined with the control Thor1RV flies) with increased longevity. However, when lower and higher content food conditions are included, an increase in longevity is observed, but it is unclear whether this merely represents a reversion to normal, healthy life span at 2% yeast conditions, rather than a genuine DR response.
These data nonetheless suggest that 4E-BP may be important for aspects of the CR/DR response downstream of Dmp53.
Discussion
We have recently shown that expression of DN-Dmp53 in the adult nervous system extends Drosophila life span in a CR-dependent manner (Bauer et al. 2005). The question of non-specific effects, often encountered when using DN constructs, prompted us to investigate the longevity effects of DN-Dmp53 expression in a Dmp53 null background. As shown in Figure 1, under these conditions DN-Dmp53 expression does not extend fly life span, suggesting that inhibition of Dmp53 function is responsible for the observed positive effects on life span by DN-Dmp53. Over expression of dSir2 in the fly nervous system also leads to extended longevity (Rogina and Helfand 2004). This function of dSir2 is related to CR, as life span extension by dSir2 is not additive to the extended life span associated with CR treatment and loss of dSir2 abrogates the life span extending effects of CR. dSir2 can deacetylate Dmp53 and inhibits its activity (as can their mammalian counterparts) (Bauer et al. 2009). Furthermore, simultaneous expression of dSir2 and DN-Dmp53 in the adult nervous system does not result in additive effects on longevity extension (Bauer et al. 2009). It is unclear, how reduced Dmp53 activity, either through expression of DN-Dmp53 or deacetylation by dSir2, initiates cellular events leading to extended longevity. One possibility may be that the Dmp53 transcriptional profile changes, specifically in IPC. Interesting candidate genes may be dILP2, or an as yet unidentified fly ortholog of mammalian TIGAR, a protein that can influence cellular energy balance through its regulation of glycolysis (Bensaad et al. 2006). This altered energy balance may thus in turn be responsible for changed secretion dynamics of IPC.
The placement of Dmp53 distal to dSir2 in the CR/DR life span extending pathway suggests that knowledge of the elements downstream of Dmp53 life span extension may be of great interest in understanding the molecular elements of CR/DR life span extension. We have shown that flies expressing DN-Dmp53 have lowered mRNA levels of dILP2 and decreased IIS activity in the fat body (Bauer et al. 2007), suggesting that inhibition of IIS may contribute to DN-Dmp53-dependent extended longevity. However, the relationship between CR/DR and IIS in flies is unclear. Mutations in the Drosophila insulin-receptor substrate chico have also been linked to CR/DR (Clancy et al. 2002). The life span extension associated with over expression of dFoxO is not additive with DR (Giannakou et al. 2008), suggesting IIS and CR/DR life span extension may be associated. Nonetheless, absence of dFoxO does not prevent DR life span extension (Giannakou et al. 2008; Min et al. 2008). The finding that dFoxO null flies respond to yeast DR with an increase in life span (Giannakou et al. 2008; Min et al. 2008) indicates that IIS, or at the very least its most downstream component dFoxO, is not absolutely required for the CR/DR response.
Here, we further investigated the molecular signaling pathways leading to extended longevity following Dmp53 inhibition. Flies expressing DN-Dmp53 have reduced dILP2 mRNA levels (Bauer et al. 2007), and adding back dILP2 into DN-Dmp53 long-lived flies did prevent the longevity phenotype of DN-Dmp53 expression. These observations suggest that a decrease in IIS, possibly through reduction of dILP2, may play an important role in mediating the longevity effects of decreased Dmp53. Notwithstanding, the role of dILPs in CR/DR and life span determination is unclear. Partial IPC ablation or expression of JNK or DN-Dmp53 in IPC reduces dILP2 mRNA levels between 50–70% and leads to increased longevity (Bauer et al. 2007; Broughton et al. 2005; Wang et al. 2005), whereas complete elimination of dILP1-5 yields small flies with decreased viability (Zhang et al. 2009). In contrast, specific reduction of dILP2 levels by RNAi does not influence life span (Broughton et al. 2008). However, this RNAi-mediated dILP2 down regulation is accompanied by a dFoxo-dependent up regulation of dILP3 that might be compensatory. This effect is not observed in flies expressing DN-Dmp53 (Bauer et al. 2007). These data reveal a complicated interplay between the various dILPs and suggest that ratios of dILPs relative to each other may be more important for life span determination than absolute levels of specific dILPs. Different upstream signaling mechanisms may therefore differentially change dILP abundance to control which downstream signaling mechanisms are engaged.
Nonetheless, when DN-Dmp53 is expressed in flies with reduced dFoxO levels life span is increased to the same relative extent as when expressed in a wt background. In contrast, JNK over expression-dependent longevity is abolished when one copy of dFoxO is disabled (Wang et al. 2005). Taken together, these data suggest that Dmp53-dependent regulation of longevity may employ upstream components of IIS, but branches off along the pathway and does not appear to utilize the downstream target dFoxO. A possible branch may be at the level of the protein kinase Akt. Among other targets, Akt phosphorylates and inactivates the Tsc kinase complex (Huang and Manning 2009), which in turn controls the activity of the TOR kinase. TOR phosphorylates and inhibits 4E-BP, a protein binding to and inhibiting the translation initiation factor elF4E. Furthermore, 4E-BP is controlled at the transcriptional level by Foxo (Junger et al. 2003; Wang et al. 2005). In fact, when DN-Dmp53 is expressed in flies lacking 4E-BP, no life span increase is observed, suggesting that 4E-BP and the dTOR pathway play an important role in life span regulation by Dmp53.
Evidence obtained in the yeast aging model suggests that CR/DR, Sir2 and the TOR pathway may be linked (Medvedik et al. 2007). Since the mechanism of life span extension in flies by dSir2 over expression or DN-Dmp53 expression share similar mechanism to CR (Bauer et al. 2009; Bauer et al. 2005), we hypothesized that 4E-BP may therefore also be required to mediate the longevity effects of CR/DR. When the functional 4E-BP gene is removed (Thor1Δ), yeast DR (between 15% and 5%) no longer leads to extended longevity. When flies are kept on very high caloric food (20%), life span of 4E-BP null flies (Thor1Δ) decreases dramatically, while under very low calorie conditions their life span approaches that of normally fed control flies (Thor1RV). Therefore, this life span extending effect observed between 20% and 2% may reflect toxicity issues of very high calorie food, and not a CR response per se. Sucrose DR does not increase life span, even of control flies, which is in agreement with what has been reported previously (Mair et al. 2005; Skorupa et al. 2008).
Inhibition of the dTOR pathway through expression of dTsc or DN-S6K(Kapahi et al. 2004), or by over expression of 4E-BP (Zid et al. 2009) has previously been shown to extend life span. These increases in longevity are not additive to the effects of DR. Importantly, flies in which 4E-BP has been disabled have independently been shown to be incapable of responding to DR with extended life spans (Zid et al. 2009). Interestingly, over expression of 4E-BP in the peri-cerebral, but not in the abdominal fat body was capable of extending life span. This may represent different functionalities of these two distinct types of fat body. On the other hand, it may indicate that 4E-BP is required, but not sufficient for longevity regulation by CR/DR. It may be that the dFoxO branch of IIS, although not required, plays a supporting role in control of CR/DR mediated life span extension. 4E-BP is a known target of dFoxO transcriptional activity (Junger et al. 2003), and expression of dFoxO in the fat body is sufficient to increase life span (Giannakou et al. 2004; Hwangbo et al. 2004). Alternatively, over expressing 4E-BP may not activate it sufficiently to suppress translation initiation, or 4E-BP may need to be transcribed at higher levels to inhibit elF4E sufficiently. When 4E-BP is over expressed using the armadillo-driver, life span extension of up to 22% is observed (Zid et al. 2009), suggesting that 4E-BP expression and activation levels may be crucial to its ability to suppress elF4E and increase longevity.
Taken together, these data suggest that CR/DR influences life span mainly through the TOR pathway, while the Foxo branch of IIS is utilized to mediate other aspects of CR/DR and/or to support TOR signaling. Foxo signaling may furthermore provide critical feedback regulation of dILP levels (Broughton et al. 2008).
The mechanisms of TOR-dependent life span regulation are the subject of intense research efforts. Zid et. al. have shown that DR in flies increases mitochondrial density and the activity of components of the mitochondrial electron transport chain. These changes are dependent on the presence of 4E-BP and presumably mediated through 4EBP-dependent alterations in protein translation (Zid et al. 2009). Flies treated with the TOR inhibitor rapamycin have extended longevity, which is dependent on functional 4EBP. Furthermore, these flies have lower levels of protein translation and an increase in the levels of autophagy (Bjedov et al.). Increased autophagy extends the longevity of C. elegans (Hansen et al. 2008), as does inhibition of protein synthesis (Pan et al. 2007). Likewise, reduction of methionine content in the food has been demonstrated to extend life span of mice (Miller et al. 2005), rats (Orentreich et al. 1993) and flies (Grandison et al. 2009; Troen et al. 2007), and may be the cause for the life span extending effects of DR (Grandison et al. 2009) by reducing protein synthesis (Miller et al. 2005).
Interestingly, mice fed late in life with rapamycin show remarkable life span extension that may be related to CR/DR (Harrison et al. 2009) which suggest that inhibition of the TOR pathway is a conserved CR/DR-related mechanism that increases longevity. Nonetheless, the use of blunt tools, such as CR/DR or drug treatment, ignores tissue-specific effects and carries the risk of inhibiting molecular pathways in tissues in which inhibition may be detrimental. For example, general inhibition of p53 leads to cancer, whereas neuronal (or neuroendocrine-specific) p53 inhibition contributes to longevity. On the other hand, mTOR stimulation protects cone cells from death in a mouse model of retinal degeneration (Punzo et al. 2009). Clearly, tissue-specific evaluation of signaling pathways is required to fully understand the CR/DR response.
Taken together, our data suggest a model for Dmp53-dependent life span regulation (Figure 5). CR/DR increases dSir2 activity, which in turn inhibits Dmp53. Decreased activity of Dmp53 in neuroendocrine cells leads to a decrease of dILP2 levels by an as yet unclear mechanism and down regulation of IIS in the fat body. While decreased IIS leads to accumulation of dFoxo in the nucleus, dFoxO itself does not mediate the life span extending effects of decreased Dmp53 activity, but may be responsible for other effects. The life span extending effects may be mediated through the TOR pathway, likely due to a branch in IIS at the level of Akt. Inhibition of the TOR pathway in turn decreases protein translation through increased 4E-BP activity. It will be interesting to untangle the molecular and cellular events leading to increased life span that are a consequence of decreased TOR pathway signaling.
Supplementary Material
Figure 4. 4E-BP is required for the longevity response under standard DR conditions.
Mean life span as a function of food content. (A) Wild type flies (Thor1RV) have shortened life span in the pathological zone (2% and 20%), but respond vigorously with a longevity increase when yeast extract content is reduced in the healthy zone from 15% to 5% (blue line). When sucrose content is varied (red line), mean life span varies around the 40 day mark with no significant increase when sucrose is reduced from 15% to 5%. (B) Flies missing 4E-BP (Thor1Δ) do not respond to a decrease of yeast extract content from 15% to 5% in the food in the healthy food conditions zone with a significant increase in mean life span (green). There is an increase in life span in the pathological zone when yeast extract is reduced from 20% to 2% back to the mean life span of wt flies (shown are the averages of two independent experiments; the asterisks denote statistically significant mean life span differences).
Acknowledgements
The authors would like to thank M. Tatar, P. Shen, B. Zid, B. Edgar and H. Keshishian for the kind gift of fly stocks. We also thank Will Lightfoot and Santharam Kolli for technical assistance and the anonymous reviewers for their helpful comments on the manuscript. This work was supported by NIA grants AG16667, AG24353 and AG25277 to SLH and NIA AG029723 to JHB. SLH is an Ellison Medical Research Foundation Senior Investigator and recipient of a Glenn Award for Research in Biological Mechanisms of Aging.
Abbreviations
- CR
calorie restriction
- DR
dietary restriction
- DN
dominant-negative
- Dmp53
Drosophila p53
- wt
wild type
- IIS
insulin/insulin-like growth factor signaling
- dILP
Drosophila insulin-like peptide
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Bauer JH, Chang C, Morris SN, Hozier S, Andersen S, et al. Expression of dominant-negative Dmp53 in the adult fly brain inhibits insulin signaling. Proc Natl Acad Sci U S A. 2007;104:13355–13360. doi: 10.1073/pnas.0706121104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bauer JH, Morris SN, Chang C, Flatt T, Wood JG, et al. dSir2 and Dmp53 interact to mediate aspects of CR-dependent life span extension in D. melanogaster. Impact: Aging. 2009;1:38–48. doi: 10.18632/aging.100001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bauer JH, Poon PC, Glatt-Deeley H, Abrams JM, Helfand SL. Neuronal expression of p53 dominant-negative proteins in adult Drosophila melanogaster extends life span. Curr Biol. 2005;15:2063–2068. doi: 10.1016/j.cub.2005.10.051. [DOI] [PubMed] [Google Scholar]
- Bensaad K, Tsuruta A, Selak MA, Vidal MN, Nakano K, et al. TIGAR, a p53-inducible regulator of glycolysis and apoptosis. Cell. 2006;126:107–120. doi: 10.1016/j.cell.2006.05.036. [DOI] [PubMed] [Google Scholar]
- Bernal A, Schoenfeld R, Kleinhesselink K, Kimbrell DA. Loss of Thor, the single 4E-BP gene of Drosophila, does not result in lethality. Dros. Inf. Serv. 2004;87:81–84. [Google Scholar]
- Bjedov I, Toivonen JM, Kerr F, Slack C, Jacobson J, et al. Mechanisms of Life Span Extension by Rapamycin in the Fruit Fly Drosophila melanogaster. 2010;11:35–46. doi: 10.1016/j.cmet.2009.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Britton JS, Lockwood WK, Li L, Cohen SM, Edgar BA. Drosophila's insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev Cell. 2002;2:239–249. doi: 10.1016/s1534-5807(02)00117-x. [DOI] [PubMed] [Google Scholar]
- Brodsky MH, Nordstrom W, Tsang G, Kwan E, Rubin GM, et al. Drosophila p53 binds a damage response element at the reaper locus. Cell. 2000;101:103–113. doi: 10.1016/S0092-8674(00)80627-3. [DOI] [PubMed] [Google Scholar]
- Broughton S, Alic N, Slack C, Bass T, Ikeya T, et al. Reduction of DILP2 in Drosophila triages a metabolic phenotype from lifespan revealing redundancy and compensation among DILPs. PLoS ONE. 2008;3:e3721. doi: 10.1371/journal.pone.0003721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broughton SJ, Piper MD, Ikeya T, Bass TM, Jacobson J, et al. Longer lifespan, altered metabolism, and stress resistance in Drosophila from ablation of cells making insulin-like ligands. Proc Natl Acad Sci U S A. 2005;102:3105–3110. doi: 10.1073/pnas.0405775102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brunet A, Sweeney LB, Sturgill JF, Chua KF, Greer PL, et al. Stress-dependent regulation of FOXO transcription factors by the SIRT1 deacetylase. Science. 2004;303:2011–2015. doi: 10.1126/science.1094637. [DOI] [PubMed] [Google Scholar]
- Chapman T, Partridge L. Female fitness in Drosophila melanogaster: an interaction between the effect of nutrition and of encounter rate with males. Proc R Soc Lond B Biol Sci. 1996;263:755–759. doi: 10.1098/rspb.1996.0113. [DOI] [PubMed] [Google Scholar]
- Clancy DJ, Gems D, Hafen E, Leevers SJ, Partridge L. Dietary restriction in long-lived dwarf flies. Science. 2002;296:319. doi: 10.1126/science.1069366. [DOI] [PubMed] [Google Scholar]
- Cohen HY, Miller C, Bitterman KJ, Wall NR, Hekking B, et al. Calorie restriction promotes mammalian cell survival by inducing the SIRT1 deacetylase. Science. 2004;305:390–392. doi: 10.1126/science.1099196. [DOI] [PubMed] [Google Scholar]
- Daitoku H, Hatta M, Matsuzaki H, Aratani S, Ohshima T, et al. Silent information regulator 2 potentiates Foxo1-mediated transcription through its deacetylase activity. Proc Natl Acad Sci U S A. 2004;101:10042–10047. doi: 10.1073/pnas.0400593101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giannakou ME, Goss M, Junger MA, Hafen E, Leevers SJ, et al. Long-lived Drosophila with overexpressed dFOXO in adult fat body. Science. 2004;305:361. doi: 10.1126/science.1098219. [DOI] [PubMed] [Google Scholar]
- Giannakou ME, Goss M, Partridge L. Role of dFOXO in lifespan extension by dietary restriction in Drosophila melanogaster: not required, but its activity modulates the response. Aging Cell. 2008;7:187–198. doi: 10.1111/j.1474-9726.2007.00362.x. [DOI] [PubMed] [Google Scholar]
- Gingras AC, Kennedy SG, O'Leary MA, Sonenberg N, Hay N. 4E-BP1, a repressor of mRNA translation, is phosphorylated and inactivated by the Akt(PKB) signaling pathway. Genes Dev. 1998;12:502–513. doi: 10.1101/gad.12.4.502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandison RC, Piper MD, Partridge L. Amino-acid imbalance explains extension of lifespan by dietary restriction in Drosophila. Nature. 2009;462:1061–1064. doi: 10.1038/nature08619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen M, Chandra A, Mitic LL, Onken B, Driscoll M, et al. A role for autophagy in the extension of lifespan by dietary restriction in C. elegans. PLoS Genet. 2008;4:e24. doi: 10.1371/journal.pgen.0040024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrison DE, Strong R, Sharp ZD, Nelson JF, Astle CM, et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature. 2009 doi: 10.1038/nature08221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang J, Manning BD. A complex interplay between Akt, TSC2 and the two mTOR complexes. Biochem Soc Trans. 2009;37:217–222. doi: 10.1042/BST0370217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwangbo DS, Gersham B, Tu MP, Palmer M, Tatar M. Drosophila dFOXO controls lifespan and regulates insulin signalling in brain and fat body. Nature. 2004;429:562–566. doi: 10.1038/nature02549. [DOI] [PubMed] [Google Scholar]
- Junger MA, Rintelen F, Stocker H, Wasserman JD, Vegh M, et al. The Drosophila Forkhead transcription factor FOXO mediates the reduction in cell number associated with reduced insulin signaling. J Biol. 2003;2:20. doi: 10.1186/1475-4924-2-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kapahi P, Zid BM, Harper T, Koslover D, Sapin V, et al. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr Biol. 2004;14:885–890. doi: 10.1016/j.cub.2004.03.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langley E, Pearson M, Faretta M, Bauer UM, Frye RA, et al. Human SIR2 deacetylates p53 and antagonizes PML/p53-induced cellular senescence. Embo J. 2002;21:2383–2396. doi: 10.1093/emboj/21.10.2383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo J, Nikolaev AY, Imai S, Chen D, Su F, et al. Negative control of p53 by Sir2alpha promotes cell survival under stress. Cell. 2001;107:137–148. doi: 10.1016/s0092-8674(01)00524-4. [DOI] [PubMed] [Google Scholar]
- Mair W, Piper MD, Partridge L. Calories do not explain extension of life span by dietary restriction in Drosophila. PLoS Biol. 2005;3:e223. doi: 10.1371/journal.pbio.0030223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masoro EJ. Overview of caloric restriction and ageing. Mech Ageing Dev. 2005;126:913–922. doi: 10.1016/j.mad.2005.03.012. [DOI] [PubMed] [Google Scholar]
- Medvedik O, Lamming DW, Kim KD, Sinclair DA. MSN2 and MSN4 link calorie restriction and TOR to sirtuin-mediated lifespan extension in Saccharomyces cerevisiae. PLoS Biol. 2007;5:e261. doi: 10.1371/journal.pbio.0050261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller RA, Buehner G, Chang Y, Harper JM, Sigler R, et al. Methionine-deficient diet extends mouse lifespan, slows immune and lens aging, alters glucose, T4, IGF-I and insulin levels, and increases hepatocyte MIF levels and stress resistance. Aging Cell. 2005;4:119–125. doi: 10.1111/j.1474-9726.2005.00152.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Min KJ, Yamamoto R, Buch S, Pankratz M, Tatar M. Drosophila life span control by dietary restriction independent of insulin-like signaling. Aging Cell. 2008 doi: 10.1111/j.1474-9726.2008.00373.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nemoto S, Fergusson MM, Finkel T. SIRT1 functionally interacts with the metabolic regulator and transcriptional coactivator PGC-1α. J Biol Chem. 2005;280:16456–16460. doi: 10.1074/jbc.M501485200. [DOI] [PubMed] [Google Scholar]
- Ollmann M, Young LM, Di Como CJ, Karim F, Belvin M, et al. Drosophila p53 is a structural and functional homolog of the tumor suppressor p53. Cell. 2000;101:91–101. doi: 10.1016/S0092-8674(00)80626-1. [DOI] [PubMed] [Google Scholar]
- Orentreich N, Matias JR, DeFelice A, Zimmerman JA. Low methionine ingestion by rats extends life span. J Nutr. 1993;123:269–274. doi: 10.1093/jn/123.2.269. [DOI] [PubMed] [Google Scholar]
- Pan KZ, Palter JE, Rogers AN, Olsen A, Chen D, et al. Inhibition of mRNA translation extends lifespan in Caenorhabditis elegans. Aging Cell. 2007;6:111–119. doi: 10.1111/j.1474-9726.2006.00266.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Punzo C, Kornacker K, Cepko CL. Stimulation of the insulin/mTOR pathway delays cone death in a mouse model of retinitis pigmentosa. Nat Neurosci. 2009;12:44–52. doi: 10.1038/nn.2234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogina B, Helfand SL. Sir2 mediates longevity in the fly through a pathway related to calorie restriction. Proc Natl Acad Sci U S A. 2004;101:15998–16003. doi: 10.1073/pnas.0404184101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogina B, Helfand SL, Frankel S. Longevity regulation by Drosophila Rpd3 deacetylase and caloric restriction. Science. 2002;298:1745. doi: 10.1126/science.1078986. [DOI] [PubMed] [Google Scholar]
- Skorupa DA, Dervisefendic A, Zwiener J, Pletcher SD. Dietary composition specifies consumption, obesity, and lifespan in Drosophila melanogaster. Aging Cell. 2008 doi: 10.1111/j.1474-9726.2008.00400.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sogame N, Kim M, Abrams JM. Drosophila p53 preserves genomic stability by regulating cell death. Proc Natl Acad Sci U S A. 2003;100:4696–4701. doi: 10.1073/pnas.0736384100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatar M, Bartke A, Antebi A. The endocrine regulation of aging by insulin-like signals. Science. 2003;299:1346–1351. doi: 10.1126/science.1081447. [DOI] [PubMed] [Google Scholar]
- Thomas G, Hall MN. TOR signalling and control of cell growth. Curr Opin Cell Biol. 1997;9:782–787. doi: 10.1016/s0955-0674(97)80078-6. [DOI] [PubMed] [Google Scholar]
- Troen AM, French EE, Roberts JF, Selhub J, Ordovas JM, et al. Lifespan modification by glucose and methionine in Drosophila melanogaster fed a chemically defined diet. Age (Dordr) 2007;29:29–39. doi: 10.1007/s11357-006-9018-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Horst A, Tertoolen LG, de Vries-Smits LM, Frye RA, Medema RH, et al. FOXO4 is acetylated upon peroxide stress and deacetylated by the longevity protein hSir2(SIRT1) J Biol Chem. 2004;279:28873–28879. doi: 10.1074/jbc.M401138200. [DOI] [PubMed] [Google Scholar]
- Vaziri H, Dessain SK, Ng Eaton E, Imai SI, Frye RA, et al. hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell. 2001;107:149–159. doi: 10.1016/s0092-8674(01)00527-x. [DOI] [PubMed] [Google Scholar]
- Wang MC, Bohmann D, Jasper H. JNK extends life span and limits growth by antagonizing cellular and organism-wide responses to insulin signaling. Cell. 2005;121:115–125. doi: 10.1016/j.cell.2005.02.030. [DOI] [PubMed] [Google Scholar]
- Zhang H, Liu J, Li CR, Momen B, Kohanski RA, et al. Deletion of Drosophila insulin-like peptides causes growth defects and metabolic abnormalities. Proc Natl Acad Sci U S A. 2009;106:19617–19622. doi: 10.1073/pnas.0905083106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zid BM, Rogers AN, Katewa SD, Vargas MA, Kolipinski MC, et al. 4E-BP extends lifespan upon dietary restriction by enhancing mitochondrial activity in Drosophila. Cell. 2009;139:149–160. doi: 10.1016/j.cell.2009.07.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.