Abstract
This study tested the potential of small interfering RNAs (siRNA) targeting human peroxisome proliferator activated receptor gamma (PPARγ) to repress the adipogenic effect of alcohol on human bone marrow-derived mesenchymal cells (hBMSCs). hBMSCs were cultured from hip replacement surgery patients (n = 10). PPARγ-siRNA was transiently transfected into hBMSCs cultured in ostogenic media containing 50 mM alcohol by using a liposome-based strategy. Oil red O staining was used to test the development of differentiated adipocytes, and Alizarin red staining was used to test mineral deposition. Marker genes of adipogenesis (PPARγ2 and aP2) and osteogenesis (Osf2/Cbfa1) were examined through real time RT-PCR and Western blot, respectively. Collagen type I, alkaline phosphatase and osteocalcin protein synthesis of cultures were also assayed. Data were presented as mean ± SD. Differences between the means of the treatment groups were determined with ANOVA. PPARγ-siRNA transfection resulted in significantly lower adipocyte number, increased matrix mineralisation, repressed adipogenic gene markers, up-regulated osteogenic gene marker and bone matrix protein synthesis in the PPARγ-siRNA group compared to controls (P < 0.05). PPARγ-siRNA is a useful strategy to inhibit the adipogenic effect and the osteogenic repression of alcohol on hBMSCs. This may be a novel therapeutic intervention for osteopenic disorders in alcoholism and other conditions.
Introduction
The habitual consumption of significant quantities of alcohol is recognised as a major factor for osteopaenia and increased fracture incidence in alcoholics [1–3]. The current consensus of clinical and experimental studies of alcohol-induced bone loss is that bone remodelling is disrupted because new bone formation is suppressed while only relatively small changes (increase or decrease) occur in bone resorption [4].
In adults, the osteoblast, which is responsible for bone formation, is derived from multipotential mesenchymal stem cells (MSC) in bone marrow. The capacity of these MSCs to differentiate into osteoblasts has a critical role in the cellular processes involved in the maintenance of the adult human skeleton.
The MSCs have been proven to differentiate into multiple lineages and generate progenitors committed to one or more cell lines [5–8]. Among these multiple lineages, those of adipogenesis and osteogenesis are the most closely related. An inverse relationship has been demonstrated between osteogenesis and adipogenesis [9], and there was evidence of transdifferentiation of these cells, suggesting a large degree of plasticity between osteoblasts and adipocytes [10, 11]. Several studies have provided substantial evidence that osteoblasts and adipocytes share a common progenitor: multipotential MSCs in bone marrow [5, 12, 13]. Accumulated evidence of the differentiation switching of these two cell lineages suggests the possible reciprocal relationship between the two differentiation pathways [9, 11, 14].
Because the relationship between adipogenesis and osteogenesis is reciprocal and the adipocytic and osteogenic cells share a common lineage, it is possible that inhibition of adipogenesis may provide an approach to prevent or treat osteoporosis or other bone diseases; thus, the signal transduction pathways implicated in these processes are evaluated as potential targets for therapeutic intervention of osteopaenic disorders.
Several’s studies [15–17] have provided evidence that alcohol shifts lineage commitment and differentiation of MSCs from osteogenesis to adipogenesis by upregulation of gene expression for peroxisome proliferator-activated receptor gamma (PPARγ); however, the molecular mechanism underlying the reciprocal relationship is not yet well understood.
Peroxisome proliferator-activated receptor gamma (PPARγ) is a nuclear receptor and regarded as a master regulator of adipogenesis and metabolic homeostasis [18]. It was demonstrated that PPARγ plays requisite and sufficient roles in the regulation of adipocyte differentiation in PPARγ knockout animals [19, 20]. Recent work of Akune et al. [21] has shown that PPARγ deficiency promotes embryonic stem cell osteoblastogenesis but failed to undergo adipogenesis.
RNA interference (RNAi) is an evolutionary conserved post-transcriptional mechanism of gene silencing induced by sequence-specific double-stranded RNA [22]. Over the last few years, RNA interference (RNAi) has been used in functional genomics and offers innovative approaches in the development of novel therapeutics for certain human diseases [23, 24].
Hosono et al. have reported using an adenovirus-vector-mediated RNAi system specific to PPARγ to suppress adipogenesis in a murine preadipocyte cell line (3T3-L1) [25]. Suppressing PPARγ function through the TAK1/TAB1/NIK cascade leads to cellular differentiation towards osteoblasts rather than adipocytes in multipotent MSCs [26], which suggests that PPARγ acts as a gatekeeper of multipotency in MSCs. Xu et al. [27] used a simpler liposome-mediated PPARγ-siRNA strategy to inhibit adipocyte differentiation in human preadipocytes and human fetal-femur-derived MSCs; however, they did not test the RNAi effect on osteogensis while adipogenesis was inhibited.
Given the emerging potential of RNAi to target PPARγ, RNAi may provide an alternative approach to prevent the adipogenic effect of alcohol on human bone marrow-derived MSCs which may reciprocally enhance the osteogenesis of the multipotent cells. In this study, we cultured human bone marrow-derived MSCs in specific conditions containing osteogenic inducers and alcohol as described by Gong and Wezeman [15]. We examined whether the adipogenic effect of alcohol on human bone marrow-derived MSCs could be inhibited by using the RNAi technique targeting the human PPARγ gene and whether the inhibition effect on adipogenesis would result in promotion of osteogenesis.
Methods and materials
Materials
Dexamethasone, β-glycerophosphate, Percoll, L-ascorbic acid 2-phosphate and foetal bovine serum (FBS) were purchased from Sigma (St. Louis, MO). Dulbecco's minimum essential medium (DMEM), Dulbecco’s phosphate-buffered saline (D-PBS), and antibiotics/antimycotics were purchased from GIBCO (Grand Island, NY). Opti-MEM I Reduced Serum Medium and Lipofectamine™ RNAiMAX were purchased from Invitrogen Life Technologies (UK). Specific and negative siRNAs were purchased from Ambion. Antihuman aP2, PPARγ and Osf2/Cbfa1 monoclonal antibody were from R&D Systems (USA). Staining solutions and all other biochemical reagents were obtained from Sigma-Aldrich (UK) unless otherwise stated.
Cell preparation and culture methods
Human MSCs were harvested from cancellous bony fragments of patients undergoing routine total hip replacement surgery (age range 39–56 years; n = 10) after informed consent. All patients showed no evidence of concurrent illness and were not receiving any medications that could affect bone metabolism. None of the patients were alcoholics.
Primary bone marrow-derived MSCs were purified by a modification of Percoll density gradient centrifugation as in previously described methods [5]. When MSC cultures became nearly confluent, cells were trypsinised and plated in 75 cm2 flasks or 6-well plates for protein analyses or RNA analyses and histochemical analysis, respectively, after siRNA transfection. Cells were used for siRNA transfection/adipogenic induction/control at passage two or three only.
Cell surface markers were evaluated by flow cytometry (Cytomics FC-500, Beckman Coulter, Fullerton, CA, USA) using the monoclonal antibodies conjugated with fluorescein isothiocyanate (FITC) or phycoerythrine (PE): CD34-FITC, CD105-PE, CD166-PE.
Transfection of PPARγ-siRNA
FAM™ Labeled Negative Control siRNA #1 (a fluorescent random siRNA) was used to monitor and optimise siRNA transfection in hMSC by using Lipofectamine™ RNAiMAX as a carrier. Negative siRNA has no significant homology to any known human gene sequences (Ambion manufacturers held data).
siRNA transfection was performed on MSCs after cells were seeded and grown to approximately 50% confluence. Cells were transfected with Silencer™ validated siRNAs targeting PPARγ (validated siRNAs ID 5821, cat. #51323; Ambion, Inc.) using Lipofectamine™ RNAiMAX transfection agent according to the manufacturers' instructions. Briefly, the Lipofectamine™ RNAiMAX was diluted and incubated in non-serum DMEM for 15 minutes and mixed with siRNA diluted in non-serum DMEM for an additional 15 min to form the transfection complexes. Transfection complexes were added onto the cells of which culture medium was replaced with non-serum DMEM prior to transfection. Lipofectamine at 0.2% (v/v) and PPARγ-siRNA (60 nM) were used in a total transfection volume of 2.4 ml per well or 8 ml per flask. Six hours after adding the transfection complex, cell culture medium was changed into DMEM containing 10% FBS.
Where appropriate, Lipofectamine and negative siRNA (validated Silencer™ negative control #1 siRNA, cat. #4611; Ambion, Inc.) were established in parallel to demonstrate nonspecific effects of gene silencing and to confirm the transfection procedure, respectively.
Treatment groups and adipogenic induction
When treatment started (day 0), MSCs at passage two or three were cultured in the following six conditions: (1) Normal cell control group: medium with no additions except for 10% FBS; (2) Osteogenic control group: medium to which osteogenic inducers (10 mMβ-glycerophosphate, 50 μg/ml ascorbic acid, and10-7 M dexamethasone) were added; (3) Adipogenic control group [16]: medium containing 10% FBS, 50 mM alcohol, and osteogenic inducers; (4) Lipofectamine control group: medium containing 10% FBS, 50 mM alcohol, the osteogenic inducers and transfection reagents Lipofectamine; (5) Negative siRNA control group: medium containing 10% FBS, 50 mM alcohol, the osteogenic inducers and MSCs transfected with negative PPARγ-siRNA; (6) PPARγ-siRNA group: medium containing 10% FBS, 50 mM alcohol, the osteogenic inducers and MSCs transfected with PPARγ-siRNA.
We chose the concentration of alcohol (50 mM) because previous studies have reported that the half-maximal effective concentration of alcohol to inhibit osteoblast proliferation in vitro is approximate 50 mM, a level within the physiological range observed in actively imbibing alcoholics [28].
Due to the volatile nature of alcohol, its concentration was maintained by placing a reservoir containing the same concentration of alcohol in a chamber during culture. The reservoir was replenished daily [29].
Oil red O staining and quantitation
For morphological assessment of adipogenesis, the accumulation of intracellular lipid droplets was visualised by being stained with Oil red O. The cells were fixed in 4% neutral buffered formalin for ten minutes followed by washing with 3% isopropanol, and then incubated with a newly filtered Oil red O staining solution for one hour at room temperature. After staining, the cells were rinsed with double-distilled H2O. Cells were considered as lipid-positive when droplets were stained red under a light microscope. Lipid-positive cells were counted under nine different microscopic fields (×200 magnification), with three fields per chamber slide to get the average number of lipid-positive cells for each group.
Alizarin red staining
To detect mineral deposition, alizarin red staining was performed on MSCs after cultured in chamber slides for 28 days. Briefly, cells were fixed in 4% formaldehyde in PBS for 45 minutes at 4°C. After being washed with distilled water, cells were exposed to alizarin red (2% aqueous, Sigma) for ten minutes at room temperature and then washed again with distilled water. Finally, cells were observed and photographed under phase contrast microscopy.
Real-time quantitative RT-PCR
Total RNA was prepared from cultures by using the Trizol reagent (Invitrogen) according to the manufacturer’s instructions for messenger RNA (mRNA) expression of adipogenic markers (PPARγ2 and aP2) on day six and osteogenic maker Osf2/Cbfa1 on day ten. Quantitative RT-PCR analysis was performed with the QuantiTect SYBR Green RT-PCR Kit (Qiagen, Valencia, CA) using the GeneAmp 5700 sequence detection system (Applied Biosystems) according to the manufacturer's instructions. β-actin was used as a normalisation control. Samples were assayed in six duplicates, and the values were normalised to the relative amounts of β-actin.
Western blot analysis
Cell lysates containing 20 –25 μg of protein from six- or ten-day cultures for adipogenic (PPAR-γ2 and aP2) and osteogenic markers (Osf2/Cbfa1) were separated by SDS-polyacrylamide gel electrophoresis and transferred to PVDF membranes (Millipore Corp., Bedford, MA), respectively. The membranes were blocked for one hour in defatted milk (10% in Tris-buffered saline with TWEEN-20 [TBST] buffer) incubated with indicated primary antibodies (1:1000) and then incubated in horseradish peroxidase-conjugated secondary antibody (1:1,000). After washing, the blots were developed by using the SuperSignal West Picochemiluminescent detection system (Pierce, Rockford, IL) and exposed to X-ray film to detect the protein band. Relative densitometric units were determined using the analysis software Diversity Database.
Alkaline phosphatase, collagen-I and osteocalcin assay
On day ten, alkaline phosphatase activity was assayed by measuring the formation of p-nitrophenol from p-nitrophenyl phosphate with the ALP Optimised clorimetric test kit according to the manufacturer’s instructions. C-terminal propeptide of type I collagen was measured by enzyme-linked immunoadsorbent assay with a Metra CICP EIA kit (Quidel, San Diego, CA). For assessing osteocalcin, cultures were added into 10-7 M 1,25-dihydroxycholecalciferol during the last 24 hours. A Metra Osteocalcin EIA kit (Quidel) was used for osteocalcin production measurement in the culture media from the culture of day ten. The results of alkaline phosphatase activity, type I procollagen and osteocalcin production were standardised to total protein as measured with Bio-Rad total protein assay reagents (Bio-Rad, Hercules, CA).
Statistical analysis
Data are presented as mean ± SD. Differences between the means of the treatment groups were determined with ANOVA. When a significant difference was determined, the Bonferroni post hoc analysis was used, and a P value less than 0.05 was considered significant.
Results
Identification of MSCs
The expression of the surface markers in the MSCs were determined by flow cytometry using the monoclonal antibodies CD34-FITC, CD105-PE, and CD166-PE in the undifferentiated states. The MSCs expressed the CD105 and CD166, markers of MSC, but did not express the CD34, hematopoietic cell markers. The cells prepared by Percoll separation were composed of 98.5% ± 2.03% of CD34 negative cells, 93.9% ± 1.78% of CD105 positive cells and 92.0% ± 2.31% of CD166 positive cells.
siRNA transfection of human mesenchymal stem cells
Fluorescence was predominantly noted in nuclei of MSCs. Highest transfection efficacy (95%) was obtained when the concentration of fluorescent siRNA was 60 nM in 0.17% (v/v). Lipofectamine™ RNAiMAX transfection agent had no detectable effects on cell proliferation or differentiation. No significant differences between cells transfected with the siRNA and control transfections were observed in duration of cell cultures reaching confluence (P > 0.05).
Inhibition of alcohol-induced adipogenesis in cultured MSCs transfected with PPARγ-siRNA
Adipocyte development and quantification
Accumulation of lipid droplets was observed in all the groups except for the normal cell control group after 24 days. Adipocyte quantification showed that the average lipid-positive cell number in osteogenic controls and cell cultures transfected with PPARγ-siRNA were 3.87 ± 1.64 and 5.12 ± 1.46, respectively, which were significantly lower than the adipogenic control group (64.0% lower, P = 0.000 and 52.4% lower, P = 0.000, respectively). The number was slightly higher in the PPARγ-siRNA group than in the osteogenic control group; however, the difference was not statistically significant. No significant differences in cell cultures among the Lipofectamine™ RNAiMAX control group, the negative siRNA control group and the adipogenic control group were observed (P = 0.835) (Fig. 1).
Fig. 1.
Oil red O staining of hBMSCs transfected with PPRAγ-siRNA or controls followed by treatment for 24 days. a Cell control. b Osteogenic control. c Adipogenic control. d Lipofectamine™ RNAiMAX control. e Negative siRNA control. f PPARγ-siRNA (magnification ×200, scale bars = 20 μm). g Data presented as mean±standard deviation of 9 fields, 3 fields per slide. *P < 0.01 compared to the adipogenic control group; **P < 0.01 compared to the normal cell control group
Gene expression and protein synthesis for adipogenic markers
Real time RT-PCR analysis showed that compared to the osteogenic control group, the PPARγ2 and aP2 mRNA of adipogenic, negative SiRNA and Lipofectamine control groups were significantly higher (P < 0.001). The cells treated with PPARγ-siRNA exhibited 73.8% and 67.8% reductions in endogenous PPARγ2 and aP2 mRNAs, respectively, compared with those in the adipogenic control group (P < 0.001). Otherwise, cells in the osteogenic control group showed slight but significant increase of PPARγ2 and aP2 mRNA expression compared to the normal cell control group (P < 0.001), which may be due to the effect of dexamethasone on adipogenic differentiation of MSCs in the osteogenic inducer [16].
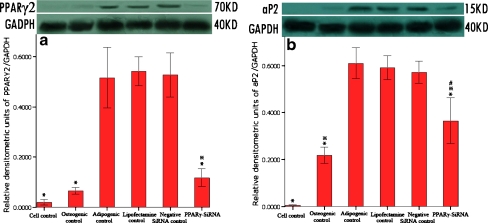
Furthermore, Western blot analysis verified that protein levels of aP2 corresponding with PPARγ2 were significantly lower in the PPARγ-siRNA group compared with the adipogenic control group (P < 0.001). The protein level of PPARγ2 in the osteogenic control group was slightly higher, however, without statistical significance. The aP2 protein level in the osteogenic control group was significantly increased compared to the normal cell control group (P = 0.000), although lower than the PPARγ-siRNA group (P = 0.000) (Fig. 2).
Fig. 2.
Effect of PPARγ-siRNA on protein levels of PPARγ2 and aP2. The data are expressed as the relative protein levels of PPARγ2 and aP2 divided by corrected GDPH value for the same sample. *P < 0.01 versus the adipogenic control group; **P < 0.01 compared to the normal cell control group; ***P = 0.000 compared to the osteogenic control group; n = 4
Osteogenesis was enhanced in MSCs transfected with PPARγ-siRNA
Mineral deposition
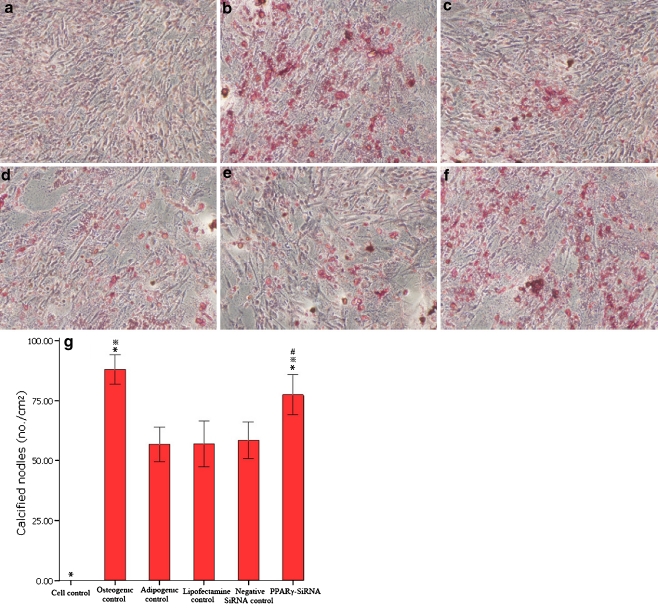
Alizarin red staining showed that throughout the 28-day culture period, the number of calcification nodules in the adipogenic control group was 56.75 ± 4.57 per square centimeter, which was significantly decreased by 35.51% compared with the osteogenic control group (P = 0.000). The cells transfected with PPARγ-siRNA presented 36.6% increased calcification nodules compared with the adipogenic control group (P = 0.000), though slightly less than the nodules in osteogenic controls (11.9% less, P = 0.004). No significant differences among the Lipofectamine™ RNAiMAX control group, the negative siRNA control group and the adipogenic control group were observed (P = 0.877). Due to deficiency of osteogenic inducers in cultures [15], no calcification nodule was found in the normal cell control group (Fig. 3).
Fig. 3.
Alizarin red staining identification of mineralisation in cell cultures for 28 days. a Cell control. b Osteogenic control. c Adipogenic control. d Lipofectamine™ RNAiMAX control. e Negative siRNA control. f PPARγ-siRNA (magnification 100×, scale bars = 20 μm). g Data presented as mean±standard deviation of calcification nodules per square centimeter. *P < 0.01 versus adipogenic control group; **P < 0.01 compared to normal cell control group; ***P = 0.004 compared to osteogenic control group
Gene expression and protein synthesis for osteogenic markers
Quantitative analysis of the mRNA levels by real-time RT-PCR method revealed that Osf2/Cbfa1, the marker gene for osteogenesis in the PPARγ-siRNA group, was up-regulated by 17.4% compared to the adipogenic control group (P = 0.000), and almost reached the gene level of cells cultured in pure osteogenic medium without alcohol (P = 0.302).
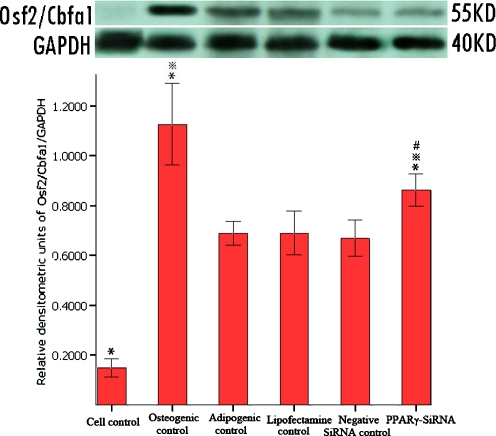
Subsequently, Western blot analysis confirmed that the protein level of Osf2/Cbfa1 corresponding with mRNA level was significantly up-regulated in MSCs transfected with PPARγ-siRNA compared with adipogenic controls (25.0% higher, P = 0.000), although still lower than osteogenic controls (P = 0.000). As other indices observed previously, no significant differences were found among the Lipofectamine™ RNAiMAX, negative siRNA and adipogenic control groups (P = 0.75) (Fig. 4).
Fig. 4.
Effect of PPARγ-siRNA on protein levels of Osf2/Cbfa1. The image on the top is the Western blot analyses, below which the data are expressed as the relative protein levels of Osf2/Cbfa1 divided by corrected GDPH value for the same sample. *P < 0.01 versus adipogenic control; **P < 0.01 versus normal cell control; ***P = 0.000 versus osteogenic control; n = 4
Alkaline phosphatase activity, type I collagen, and osteocalcin production assay
To further assess the enhancement of osteogenesis induced by PPARγ-siRNA in the MSCs, alkaline phosphatase activity, type I collagen, and osteocalcin production were assayed after ten days of treatment. Alkaline phosphatase activity, type I collagen production and osteocalcin production were all inhibited in cells cultured in medium containing alcohol compared to osteogenic controls (P < 0.05). Cells transfected with PPARγ-siRNA presented 28.6% up-regulation in alkaline phosphatase activity, 58.8% up-regulation in type I collagen production and 33.7% in osteocalcin production compared to the adipogenic control group, respectively (P < 0.001), even though Alkaline phosphatase activity and osteocalcin production of these cells were slightly but statistically lower than osteogenic controls (11.7% lower, P = 0.011; 7.1% lower, P = 0.046, respectively). No significant differences were observed among the Lipofectamine™ RNAiMAX, negative siRNA and adipogenic control groups (P > 0.05).
Discussion
Chronic consumption of excessive alcohol eventually not only increases incidence of osteoporosis and fractures, but also delays fracture healing compared with nonalcoholics [4].
The increase in marrow adipogenesis associated with osteopenia in chronic alcoholics compared with nonalcoholics is well known clinically when medullary cavities of long bones are opened during orthopaedic surgical procedures [30]. These observations in humans can also be recreated in experimental animal models [17, 31].
The clinical and experimental studies of alcohol-induced bone loss suggest that the changes in the cell dynamics of the marrow tissue may lead to abnormal bone remodelling and result in alcohol-induced bone loss. It is well known that the capacity of MSCs to differentiate into osteoblasts has a critical role in bone remodelling of the adult human [32, 33].
Marrow adipocytes share a common bone MSC pool with bone-forming osteoblasts; thus, increase of adipogenesis will result in reduction of osteogenesis. Some studies of gene silencing and overexpression have provided insight into critical pathways that determine the fate of these multipotential bone marrow stem cells. One of these pathways—that of the nuclear hormone receptor peroxisome proliferator activated receptor-γ (PPARγ), the master regulator in the process of adipocyte differentiation— when activated, promotes adipogenesis and inhibits osteogenesis [34–36].
Alcohol may disturb bone remodelling, which involves both a suppression of osteogenic competence and a promotion of adipogenesis of the MSCs in the marrow. Recently, Gong and Wezeman demonstrated the osteogenic inhibitory effect and adipogenic effect of alcohol on human bone-derived MSCs [15, 16]. They found that the adipogenic effect of alcohol is through upregulation of PPARγ2 at the point of lineage commitment as well as through enhancement of lipid transport and storage through increased aP2 synthesis. Alcohol inhibited MSCs osteogenic differentiation by reducing collagen type I synthesis and alkaline phosphatase activity. In addition, collagen type I gene expression was down-regulated by alcohol.
Our study indicates the potential of specific siRNA to inhibit PPARγ mRNA and reduce the adipogenic effect of alcohol on human bone marrow-derived MSCs, which leads to a concomitant increase in osteogenesis by using ex vivo cell culture systems.
Although adipogenic cells or stem cells are thought typically to be resistant to siRNA transfection [25, 37], our study confirms that primary human bone marrow-derived MSCs can be efficiently transfected using a liposomal-based strategy. Liposomal reagent control and siRNA control play a necessary role in verifying and specifying specific siRNAs. Negative siRNA and Lipofectamine reagent revealing no inhibitory effect in this study indicates a specific inhibitory effect of PPARγ-siRNAs on adipogenesis in human bone marrow-derived MSCs.
In our study, adipogenesis was evaluated morphologically with Oil red O staining. The adipogenesis observed in osteogenic controls was due to cultures containing osteogenic inducers added with dexamethasone [15]. Moreover, the adipogenic feature of osteogenic inducers could be enhanced by alcohol as described previously [16]. The adipocyte quantification result suggests that the adipogenic effect of alcohol was significantly inhibited by PPARγ-siRNA through specifically down-regulating PPARγ gene expression. This down-regulation was confirmed by both real time RT-PCR and Western blot analysis which showed that cells treated with PPARγ-siRNA exhibited significant reduction in both endogenous PPARγ mRNA and protein product compared to controls. PPARγ, especially PPARγ2 as EBF-1 (early B-cell factor), is a critical transcription factor that induces expression of multiple adipogenic markers and leads to terminal adipogenic differentiation of cells [38]. The correlation between PPARγ gene expression down-regulation and adipogenesis inhabitation observed in this study is consistent with the results of Xu et al. [27].
aP2, one of the well-known downstream genes regulated by PPARγ, serves as a lipid shuttle that dissolves hydrophobic fatty acids and delivers them to the appropriate metabolic system for use [39]. Its expression is induced at a very late stage during adipogenesis differentiation and is routinely considered as a lineage selective marker. In our study, aP2 was inhibited in MSCs transfected with PPARγ-siRNA at both the mRNA and protein levels, which suggested the inhibition of the adipogenic effect of alcohol through PPARγ-siRNA transfection. Moreover, in our study, aP2 followed as the up-regulation of PPARγ gene expression was promoted at both mRNA and protein levels in cultures containing osteogenic inducers and alcohol. This, however, seems at odds with a previous report by Wezeman and Gong [16], considering that as promoting PPARγ gene expression, aP2 mRNA was not regulated at the mRNA level, although the protein level of aP2 was significantly increased by alcohol. This discrepancy can perhaps be explained by differences of mRNA analysis techniques between ours and previous studies. Wezeman and Gong chose Northern blot which was semiquantitative to detect the changes at mRNA level. However, in our study, we chose real time RT-PCR to detect the more subtle changes of mRNA expression.
Because of the reciprocal relationship between osteogenesis and adipogenesis, we postulated that inhibition of the adipogenic effect of alcohol on human BMSC through siRNA targeting PPARγ mRNA would be concomitant with promotion of osteogenesis.
As we observed, Osf2/Cbfa1, the marker gene for osteogenesis, was significantly up-regulated at both mRNA and protein levels in the PPARγ-siRNA group compared to the adipogenic control groups. Osteoblast-specific factor 2/core binding factor a1 (Osf2/Cbfa1), also known as RUNX2, is a transcription factor required for commitment of mesenchymal progenitors to the osteoblast lineage [40, 41]. Recent molecular studies and genetic manipulation of Osf2/Cbfa1 in vivo indicated that the expression of Osf2/Cbfa1 is both necessary and sufficient for MSCs differentiation towards the osteoblast lineage [42, 43].
PPARγ2 can bind Osf2/Cbfa1 and inhibit its transcriptional activity [44]. As PPARγ mRNA was down-regulated by specific siRNA, and as adipogenesis was inhibited, Osf2/Cbfa1 was promoted at both mRNA and protein levels, which stimulated the osteoblast lineage differentiation of MSCs.
In our study, due to up-regulation of Osf2/Cbfa1, the type I collagen, phosphatase activity, and osteocalcin protein, the early, middle and late stage of osteogenic makers, respectively, increased in the PPARγ-siRNA group compared to the adipogenic control. This result confirms the previous findings that the Osf2/Cbfa1 regulatory element can be found in the promoter of all major osteoblast genes, including type I collagen alpha 1 chain, osteopontin, bone sialoprotein and osteocalcin, all of which are involved in the establishment of an osteoblast phenotype [45].
These osteogenic effects of PPARγ-siRNA inhibiting adipogenesis were further tested by alizarin red staining detecting calcification nodules. The ability of the extracellular matrix to undergo cell-mediated mineralisation is partially dependent on the synthesis of type I collagen and alkaline phosphatase [46].
In conclusion, our study confirmed the efficacy of PPARγ-siRNA strategies to reduce the adipogenic effect and the osteogenesis inhibition of alcohol on human bone marrow-derived MSCs. Based on the present and previous evidence, we believe that PPARγ-siRNA may be a novel target for therapeutic intervention of osteopaenic disorders in alcoholism and other osteoporosis conditions.
Acknowledgments
We would like to thank Wang Ying-cheng for his guidance and assistance with the Western blot and real-time PT-PCR studies. We are grateful to Peng Wen-zhen and Li Qiong-ying for assistance with cell culture.
Footnotes
Qiang Huang and Hui Zhang contributed equally to this work.
Contributor Information
Hui Zhang, Email: zhanghui_xx@126.com.
Fu-xing Pei, Email: peifuxing2@163.com.
References
- 1.Turner RT. Skeletal response to alcohol. Alcohol Clin Exp Res. 2000;24:1693–1701. doi: 10.1111/j.1530-0277.2000.tb01971.x. [DOI] [PubMed] [Google Scholar]
- 2.Seeman E. Effects of tobacco and alcohol use on bone. In: Marcus R, editor. Osteoporosis. 2. San Diego: Elsevier; 2001. pp. 771–794. [Google Scholar]
- 3.Clark MK, Sowers MF, Dekordi F, Nichols S. Bone mineral density and fractures among alcohol-dependent women in treatment and in recovery. Osteoporos Int. 2003;14:396–403. doi: 10.1007/s00198-003-1387-2. [DOI] [PubMed] [Google Scholar]
- 4.Chakkalakal DA. Alcohol-induced bone loss and deficient bone repair. Alcohol Clin Exp Res. 2005;29:2077–2090. doi: 10.1097/01.alc.0000192039.21305.55. [DOI] [PubMed] [Google Scholar]
- 5.Pittenger MF, Mackay AM, Beck SC, Jaiswal RK, Douglas R, Mosca JD, Moorman MA, Simonetti DW, Craig S, Marshak DR. Multilineage potential of adult human mesenchymal stem cells. Science. 1999;284:143–147. doi: 10.1126/science.284.5411.143. [DOI] [PubMed] [Google Scholar]
- 6.Declercq H, Vreken N, Maeyer E, Verbeeck R, Schacht E, Ridder L, Cornelissen M. Isolation, proliferation and differentiation of osteoblastic cells to study cell/biomaterial interactions: comparison of different isolation techniques and source. Biomaterials. 2004;25:757–768. doi: 10.1016/S0142-9612(03)00580-5. [DOI] [PubMed] [Google Scholar]
- 7.O'Donoghue K, Fisk NM. Fetal stem cells. Best Pract Res Clin Obstet Gynaecol. 2004;18:853–875. doi: 10.1016/j.bpobgyn.2004.06.010. [DOI] [PubMed] [Google Scholar]
- 8.Oreffo RO, Cooper C, Mason C, Clements M. Mesenchymal stem cells: lineage, plasticity and skeletal therapeutic potential. Stem Cell Rev. 2005;1:169–178. doi: 10.1385/SCR:1:2:169. [DOI] [PubMed] [Google Scholar]
- 9.Beresford JN, Bennett JH, Devlin C, Leboy PS, Owen ME. Evidence for an inverse relationship between the differentiation of adipocytic and osteogenic cells in rat marrow stromal cell cultures. J Cell Sci. 1992;102:341–351. doi: 10.1242/jcs.102.2.341. [DOI] [PubMed] [Google Scholar]
- 10.Nuttall ME, Patton AJ, Olivera DL, Nadeau DP, Gowen M. Human trabecular bone cells are able to express both osteoblastic and adipogenic phenotype: implications for osteopenic disorders. J Bone Miner Res. 1998;13:371–382. doi: 10.1359/jbmr.1998.13.3.371. [DOI] [PubMed] [Google Scholar]
- 11.Park SR, Oreffo RO, Triffitt JT. Interconversion potential of closed human marrow adipocytes in vitro. Bone. 1999;24:549–554. doi: 10.1016/S8756-3282(99)00084-8. [DOI] [PubMed] [Google Scholar]
- 12.Beresford JN. Osteogenic stem cells and the stromal system of bone and marrow. Clin Orthop. 1999;240:270–280. [PubMed] [Google Scholar]
- 13.Bennett JH, Joyner CJ, Triffitt JT, Owen ME. Adipocytic cells cultured from marrow have osteogenic potential. J Cell Sci. 1991;99:131–139. doi: 10.1242/jcs.99.1.131. [DOI] [PubMed] [Google Scholar]
- 14.Li XH, Zhang JC, Sui SF, Yang MS. Effect of daidzin, genistin, and glycitin on osteogenic and adipogenic differentiation of bone marrow stromal cells and adipocytic transdifferentiation of osteoblasts. Acta Pharmacol Sin. 2005;26:1081–1086. doi: 10.1111/j.1745-7254.2005.00161.x. [DOI] [PubMed] [Google Scholar]
- 15.Gong Z, Wezeman FH. Inhibitory effect of alcohol on osteogenic differentiation in human bone marrow-derived mesenchymal stem cells. Alcohol Clin Exp Res. 2004;28:468–479. doi: 10.1097/01.ALC.0000118315.58404.C1. [DOI] [PubMed] [Google Scholar]
- 16.Wezeman FH, Gong Z. Adipogenic effect of alcohol on human bone marrow-derived mesenchymal stem cells. Alcohol Clin Exp Res. 2004;28:1091–1101. doi: 10.1097/01.ALC.0000130808.49262.F5. [DOI] [PubMed] [Google Scholar]
- 17.Wang Y, Li Y, Mao K, Li J, Cui Q, Wang GJ. Alcohol-induced adipogenesis in bone and marrow: a possible mechanism for osteonecrosis. Clin Orthop Relat Res. 2003;410:213–224. doi: 10.1097/01.blo.0000063602.67412.83. [DOI] [PubMed] [Google Scholar]
- 18.Knouff C, Auwerx J. Peroxisome proliferator-activated receptor-gamma calls for activation in moderation: lessons from genetics and pharmacology. Endocr Rev. 2004;25:899–918. doi: 10.1210/er.2003-0036. [DOI] [PubMed] [Google Scholar]
- 19.Rosen ED, Sarraf P, Troy AE, Bradwin G, Moore K, Milstone DS, Spiegelman BM, Mortensen RM. PPAR gamma is required for the differentiation of adipose tissue in vivo and in vitro. Mol Cell. 1999;4:611–617. doi: 10.1016/S1097-2765(00)80211-7. [DOI] [PubMed] [Google Scholar]
- 20.Barak Y, Nelson MC, Ong ES, Jones YZ, Ruiz-Lozano P, Chien KR, Koder A, Evans RM. PPAR gamma is required for placental, cardiac, and adipose tissue development. Mol Cell. 1999;4:585–595. doi: 10.1016/S1097-2765(00)80209-9. [DOI] [PubMed] [Google Scholar]
- 21.Akune T, Ohba S, Kamekura S, et al. PPARγ insufficiency enhances osteogenesis through osteoblast formation from bone marrow progenitors. J Clin Invest. 2004;113:846–855. doi: 10.1172/JCI19900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–811. doi: 10.1038/35888. [DOI] [PubMed] [Google Scholar]
- 23.Lee NS, Dohjima T, Bauer G, Li H, Li MJ, Ehsani A, Salvaterra P, Rossi J. Expression of small interfering RNAs targeted against HIV-1 rev transcripts in human cells. Nat Biotechnol. 2002;20:500–505. doi: 10.1038/nbt0502-500. [DOI] [PubMed] [Google Scholar]
- 24.Novina CD, Murray MF, Dykxhoorn DM, et al. siRNA-directed inhibition of HIV-1 infection. Nat Med. 2002;8:681–686. doi: 10.1038/nm725. [DOI] [PubMed] [Google Scholar]
- 25.Hosono T, Mizuguchi H, Katayama K, et al. RNA interference of PPARgamma using fiber-modified adenovirus vector efficiently suppresses preadipocyte-to-adipocyte differentiation in 3 T3–L1 cells. Gene. 2005;348:157–165. doi: 10.1016/j.gene.2005.01.005. [DOI] [PubMed] [Google Scholar]
- 26.Suzawa M, Takada I, Yanagisawa J. Cytokines suppress adipogenesis and PPAR-gamma function through the TAK1/TAB1/NIK cascade. Nat Cell Biol. 2003;5:224–230. doi: 10.1038/ncb942. [DOI] [PubMed] [Google Scholar]
- 27.Xu Y, Mirmalek-Sani SH, Yang X, Zhang J, Oreffo RO. The use of small interfering RNAs to inhibit adipocyte differentiation in human preadipocytes and fetal-femur-derived mesenchymal cells. Exp Cell Res. 2006;312:1856–1864. doi: 10.1016/j.yexcr.2006.02.016. [DOI] [PubMed] [Google Scholar]
- 28.Klein RF. Alcohol-induced bone disease: impact of ethanol on osteoblast proliferation. Alcohol Clin Exp Res. 1997;21:392–399. doi: 10.1111/j.1530-0277.1997.tb03781.x. [DOI] [PubMed] [Google Scholar]
- 29.Eriksen JL, Druse MJ. Potential involvement of S100B in the protective effects of a serotonin-1A agonist on ethanol treated astrocytes. Dev Brain Res. 2001;128:157–164. doi: 10.1016/S0165-3806(01)00172-9. [DOI] [PubMed] [Google Scholar]
- 30.Meunier P, Aaron J, Edouard C, Vignon G. Osteoporosis and the replacement of cell populations of the marrow by adipose tissue. A quantitative study of 84 iliac bone biopsies. Clin Orthop. 1971;80:147–154. doi: 10.1097/00003086-197110000-00021. [DOI] [PubMed] [Google Scholar]
- 31.Wezeman FH, Gong Z. Bone marrow triglyceride accumulation and hormonal changes during long-term alcohol intake in male and female rats. Alcohol Clin Exp Res. 2001;25:1515–1522. doi: 10.1111/j.1530-0277.2001.tb02155.x. [DOI] [PubMed] [Google Scholar]
- 32.Martin RB, Chow BD, Lucas PA. Bone marrow fat content in relation to bone remodeling and serum chemistry in intact and ovariectomized dogs. Calcif Tissue Int. 1990;46:189–194. doi: 10.1007/BF02555043. [DOI] [PubMed] [Google Scholar]
- 33.Martin RB, Zissimos SL. Relationships between marrow fat and bone turnover in ovariectomized and intact rats. Bone. 1991;12:123–131. doi: 10.1016/8756-3282(91)90011-7. [DOI] [PubMed] [Google Scholar]
- 34.Lecka-Czernik B, Moerman EJ, Grant DF, Lehmann JM, Manolagas SC, Jilka RL. Divergent effects of selective peroxisome proliferator-activated receptor-gamma2 ligands on adipocyte versus osteoblast differentiation. Endocrinology. 2002;143:2376–2384. doi: 10.1210/en.143.6.2376. [DOI] [PubMed] [Google Scholar]
- 35.Rzonca SO, Suva LJ, Gaddy D, Montague DC, Lecka-Czernik B. Bone is a target for the antidiabetic compound rosiglitazone. Endocrinology. 2004;145:401–406. doi: 10.1210/en.2003-0746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rosen ED, Hsu CH, Wang X, Sakai S, Freeman MW, Gonzalez FJ, Spiegelman BM. C/EBPalpha induces adipogenesis through PPARgamma: a unified pathway. Genes Dev. 2002;16:22–26. doi: 10.1101/gad.948702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Oliveira DM, Goodell MA. Transient RNA interference in hematopoietic progenitors with functional consequences. Genesis. 2003;36:203–208. doi: 10.1002/gene.10212. [DOI] [PubMed] [Google Scholar]
- 38.Akerblad P, Mansson R, Lagergren A, et al. Gene expression analysis suggests that EBF-1 and PPARgamma2 induce adipogenesis of NIH-3 T3 cells with similar efficiency and kinetics. Physiol Genomics. 2005;23:206–216. doi: 10.1152/physiolgenomics.00015.2005. [DOI] [PubMed] [Google Scholar]
- 39.Storch J, Thumser EA. The fatty acid transport function of fatty acid-binding proteins. Biochim Biophys Acta. 2000;1486:28–44. doi: 10.1016/s1388-1981(00)00046-9. [DOI] [PubMed] [Google Scholar]
- 40.Otto F, Thornell AP, Crompton T, et al. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell. 1997;89:765–771. doi: 10.1016/S0092-8674(00)80259-7. [DOI] [PubMed] [Google Scholar]
- 41.Xiao ZS, Thomas R, Hinson TK, Quarles LD. Genomic structure and isoform expression of the mouse, rat and human Cbfa1/Osf2 transcription factor. Gene. 1998;214:187–197. doi: 10.1016/S0378-1119(98)00227-3. [DOI] [PubMed] [Google Scholar]
- 42.Karsenty G, Wagner EF. Reaching a genetic and molecular understanding of skeletal development. Dev Cell. 2002;2:389–406. doi: 10.1016/S1534-5807(02)00157-0. [DOI] [PubMed] [Google Scholar]
- 43.Komori T. Regulation of osteoblast differentiation by transcription factors. J Cellular Biochem. 2006;99:1233–1239. doi: 10.1002/jcb.20958. [DOI] [PubMed] [Google Scholar]
- 44.Jeon MJ, Kim JA, Kwon SH, Kim SW, Park KS, Park SW, Kim SY, Shin CS. Activation of peroxisome proliferator-activated receptor-gamma inhibits the Runx2-mediated transcription of osteocalcin in osteoblasts. J Biol Chem. 2003;278:23270–23277. doi: 10.1074/jbc.M211610200. [DOI] [PubMed] [Google Scholar]
- 45.Xiao G, Jiang D, Thomas P, Benson MD, Guan K, Karsenty G, Franceschi RT. MAPK pathways activate and phosphorylate the osteoblast-specific transcription factor, Cbfa1. J Biol Chem. 2000;275:4453–4459. doi: 10.1074/jbc.275.6.4453. [DOI] [PubMed] [Google Scholar]
- 46.Franceschi RT. The developmental control of osteoblast-specific gene expression: role of specific transcription factors and the extracellular matrix environment. Crit Rev Oral Biol Med. 1999;10:40–57. doi: 10.1177/10454411990100010201. [DOI] [PubMed] [Google Scholar]