Abstract
Cilia are important sensory organelles, which are thought to be essential regulators of numerous signaling pathways. In Caenorhabditis elegans, defects in sensory cilium formation result in a small-body phenotype, suggesting the role of sensory cilia in body size determination. Previous analyses suggest that lack of normal cilia causes the small-body phenotype through the activation of a signaling pathway which consists of the EGL-4 cGMP-dependent protein kinase and the GCY-12 receptor-type guanylyl cyclase. By genetic suppressor screening of the small-body phenotype of a cilium defective mutant, we identified a chb-3 gene. Genetic analyses placed chb-3 in the same pathway as egl-4 and gcy-12 and upstream of egl-4. chb-3 encodes a novel protein, with a zf-MYND motif and ankyrin repeats, that is highly conserved from worm to human. In chb-3 mutants, GCY-12 guanylyl cyclase visualized by tagged GFP (GCY-12::GFP) fails to localize to sensory cilia properly and accumulates in cell bodies. Our analyses suggest that decreased GCY-12 levels in the cilia of chb-3 mutants may cause the suppression of the small-body phenotype of a cilium defective mutant. By observing the transport of GCY-12::GFP particles along the dendrites to the cilia in sensory neurons, we found that the velocities and the frequencies of the particle movement are decreased in chb-3 mutant animals. How membrane proteins are trafficked to cilia has been the focus of extensive studies in vertebrates and invertebrates, although only a few of the relevant proteins have been identified. Our study defines a new regulator, CHB-3, in the trafficking process and also shows the importance of ciliary targeting of the signaling molecule, GCY-12, in sensory-dependent body size regulation in C. elegans. Given that CHB-3 is highly conserved in mammal, a similar system may be used in the trafficking of signaling proteins to the cilia of other species.
Author Summary
Caenorhabditis elegans is a 1–2 mm long nematode. Its body size is controlled by sensory inputs; some mutants with defects in sensory perception grow into small size (20%–30% decrease in body volume), although the animals seem to feed normally. The EGL-4 cGMP-dependent protein kinase and the GCY-12 guanylyl cyclase act in sensory neurons to control body size downstream of sensory inputs. GCY-12 is localized to cilia, antenna-like cellular structures of sensory neurons. In the cilia, a number of signaling molecules are localized. Dysfunction of cilia is known to cause several human disorders such as Bardet-Biedl syndrome, illustrating the importance of these organelles. In this study, we identified a novel protein, CHB-3, involved in sensory-dependent body size regulation. Our analyses suggest that CHB-3 protein regulates the trafficking of GCY-12 from the cell bodies to the cilia. Without CHB-3 protein, GCY-12 fails to localize to cilia and body size is not controlled properly. Thus, the cilia are a special place for sensory information processing in body size regulation. Our analyses identified CHB-3 as a novel trafficking regulator of ciliary protein(s).
Introduction
Cilia are highly conserved microtubule-based hair-like organelles, which are specialized subcellular compartments where a number of transmembrane receptors and channels are localized [1]. Through these molecules in cilia, environmental signals are transmitted to the cytoplasm and nucleus. For example, the outer segments of photoreceptor cone and rod cells in vertebrates are developmentally derived from cilia and the first steps of photosensation occur within the cilium. In addition to such classic sensory cilia, most vertebrate cells have a non-motile primary cilium, which also has a sensory function [2]. Dysfunction of the primary cilia causes a plethora of developmental disorders (ciliopathies) [3], illustrating the importance that these organelles play in signal transduction.
C. elegans is an excellent model system with which to understand how cilia form and function. Many mutants with defects in generation of normal sensory cilia have been isolated and the analyses of these mutants and the responsible genes have revealed the mechanism of cilium generation [4]. Interestingly, most of these mutants which lack normal sensory cilia exhibit a small-body phenotype, suggesting that sensory cilia are involved in the regulation of body size in C. elegans [5]. We have previously shown that the small body size of cilium-defective mutants is not due to an inability to locate food and that body size is regulated by a subset of amphid sensory neurons [6]. In order to identify the molecular mechanisms underlying sensory regulation of body size, we conducted a genetic screen for suppressors of the small body size of a cilium-defective mutant, che-2. These suppressors are termed Chb mutants for suppressor of che-2 small body size. One of the Chb mutants, egl-4, exhibits a large-body phenotype and the body size is not decreased when a che-2 mutation is introduced, indicating that cilium defects cause the small body size through egl-4 function [6]. The egl-4 gene encodes a cGMP-dependent protein kinase (PKG) [6]–[8]. EGL-4 appears to regulate body size by functioning in sensory neurons, because EGL-4 expression in several sensory neurons is sufficient for growth to a normal body size [6].
By genetic analyses, the DBL-1/TGF-β signaling pathway, which regulates hypodermal ploidy and consequently affects cell growth and body size [9], was shown to act downstream of EGL-4 [6], [7]. Indeed, che-2 and egl-4 mutants exhibit low and high ploidy of the hypodermal cell, respectively [10]. Because EGL-4 PKG can regulate gene expression by antagonizing a histone deacetylase [11], it is possible that EGL-4 kinase, under the control of sensory inputs, regulates body size by transcriptional regulation of such humoral factors as DBL-1/TGF-β.
Hence, lack of normal sensory cilia affects a signal transduction pathway of EGL-4 kinase and ultimately changes body size of C. elegans. However, the mechanisms how the EGL-4 activity is controlled through the cilia remain to be established. In this study, we establish that CHB-3 is a new upstream component of EGL-4 kinase that regulates body size. CHB-3 is a zf-MYND motif protein conserved from C. elegans to humans. We show that CHB-3 mediates the cilium localization of a guanylyl cyclase, GCY-12, and consequently regulates EGL-4 cGMP-dependent protein kinase activity. In a chb-3 mutant, the frequency and the velocity of the dendritic transport of GCY-12 to cilia seem to be decreased, compared to those of wild-type animals. Our findings suggest that appropriate control of body size requires that certain signaling components, such as GCY-12, are localized to cilia (or their basement regions) to activate their proper targets (such as EGL-4), highlighting the indispensable role of sensory cilia as signaling centers.
Results
chb-3 suppresses the small-body phenotype in the che-2 mutant
chb-3(eg52) was previously isolated as a mutant which suppresses the small body size of che-2, but not the cilium structural defect of che-2, in a screen that also identified egl-4 [6]. As shown in Figure 1A and 1B, the chb-3(eg52) mutation caused an increase in body size of che-2(e1033). Compared with wild-type animals (WT, N2 strain), the chb-3(eg52) single mutant were longer and larger overall (Figure 2). Suppression of the che-2 small-body phenotype by chb-3, however, is unlikely a result of simple additive effects of the mutation, because a che-2 transgene did not increase the size of the chb-3(eg52);che-2(e1033) mutant (chb-3(eg52);che-2(e1033);Ex[che-2] in Figure 1A). Moreover, the chb-3(eg52) single mutant and the chb-3(eg52);che-2(e1033) double mutant were similar in body size (Figure 1B). We defined the group of Chb mutants displaying body sizes that were not affected by the che-2 mutation as class I mutants, whereas the other Chb mutants were designated as class II mutants [6]. On the whole, the class I Chb genes are required for animals to make the body size small when animals do not perceive sensory inputs (e.g., cilium-defective mutants). The epistasis of chb-3 to che-2 indicates that chb-3 is a class I Chb mutant, suggesting a role for the chb-3 gene in the sensory processing-mediated regulation of body size.
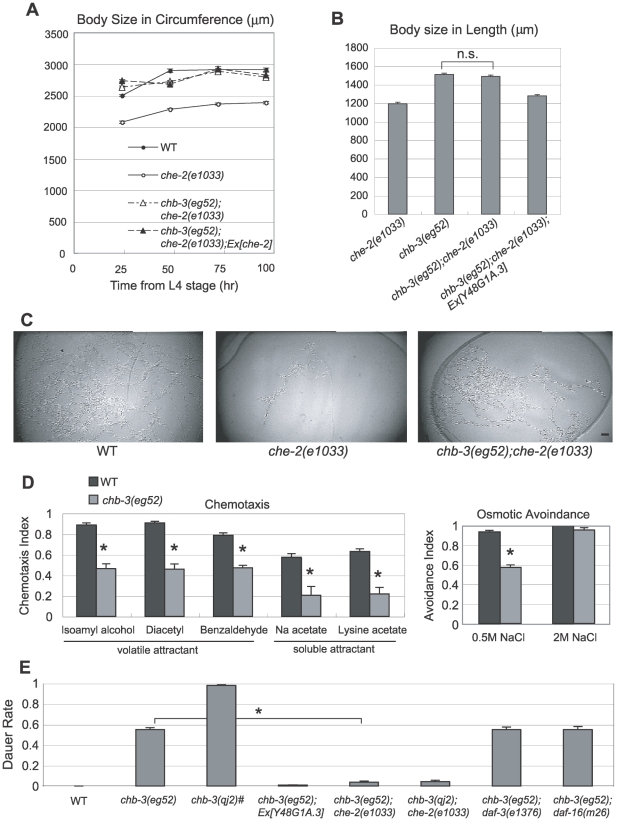
Figure 1. chb-3 is a che-2 small-body-size suppressor.
(A) Time course of changes in body size indicated by the length of the perimeter of the lateral image of the animals. Body size was measured at the indicated time points from the L4 stage. Error bars indicate the standard error of the mean (s.e.m). (B) Body size indicated by the length of the animals 48 hours after the L4 stage. “n.s.” indicates that the difference is not significant (p>0.05, t test). (C) Typical tracks left by a single animal on an E. coli lawn during an 18-hours period. Scale bar, 10 mm. (D) Chemotaxis and osmotic avoidance assays. Each data point represents an average of at least 3 independent assays. Error bars indicate s.e.m. The mark (*) indicates the significant difference (p<0.01, t test). (E) The rate of dauer formation at 20°C. The Daf-c phenotype of chb-3 was suppressed by che-2(e1033) but not by daf-3(e1376) or daf-16(m26). Rescue of the chb-3(eg52) Daf-c phenotype by the Y48G1A.3 transgene (Ex[Y48G1A.3/myo-3::gfp]) is also shown. Each data point represents an average of at least 5 independent assays. Error bars indicate s.e.m. The mark (*) indicates the significant difference from chb-3(eg52) (p<0.01, t test). #The dauer formation rate of chb-3(qj2) was determined by gathering eggs from the chb-3(qj2); Ex[tax-4p::CHB-3::GFP] rescued line, and observing hatched animals that lost the Ex array.
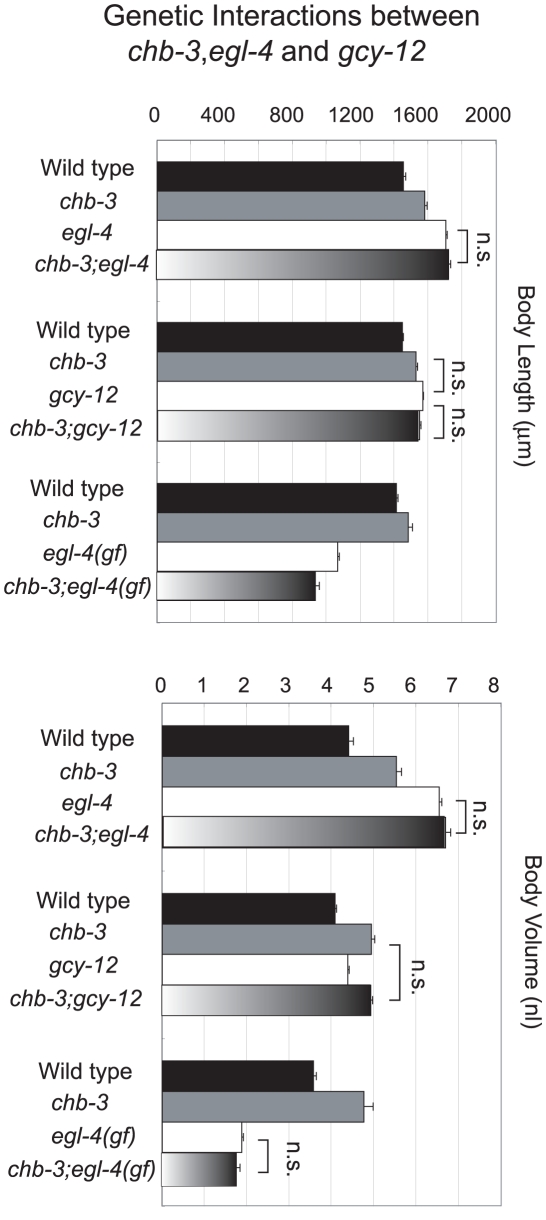
Figure 2. Genetic interactions among chb-3, egl-4, and gcy-12.
Body size, indicated using the length (µm) and volume (nl), of animals 48 hours after the L4 stage. Error bars indicate s.e.m. “n.s.” indicates that the difference is not significant (p>0.05, t test).
We have previously shown that all class I Chb mutants isolated to date, such as egl-4, suppress the characteristic locomotory defect of che-2 mutants [6]. Normally, wild-type animals alternate between two locomotory states: roaming and dwelling. Roaming is characterized by a relatively higher speed of movement with less turning, whereas dwelling is characterized by a relatively slower speed of movement with more turning. che-2 mutants exhibit a more confined tracking pattern owing to increased time dwelling and decreased time roaming. Like egl-4, chb-3 suppressed this locomotory defect of che-2. The chb-3(eg52);che-2(e1033) double mutant showed a wider tracking pattern compared with che-2 (Figure 1C). This result is consistent with a role for chb-3 downstream of sensory inputs.
chb-3 animals exhibit defects in sensory-mediated behaviors
To analyze the role of chb-3 in sensory perception, we examined other sensory behaviors of the chb-3(eg52) mutant. We found that chb-3(eg52) exhibits defects in chemotaxis behavior to a variety of attractants and in osmotic avoidance behavior (Figure 1D). For instance, compared with wild-type animals, the chb-3(eg52) mutant showed muted responses to either volatile (isoamyl alcohol, benzaldehyde, diacetyl) or soluble (Na, lysine) attractants. Aversive responses to high osmolarity solutions (0.5 M NaCl) were also less pronounced in chb-3(eg52) animals, although the response returned to a wild-type level if the osmolarity was high enough (2 M NaCl).
Another Chb mutant, egl-4, also shows defects in sensory behaviors [12]. These sensory defects suggest an important role for these molecules in sensory signal transduction and are consistent with the hypothesis that EGL-4 and CHB-3 are involved in sensory information processing underlying body size control of C. elegans. However, the phenotypes of chb-3 and egl-4 mutant animals are not completely identical; chb-3(eg52) showed reduced responses not only to isoamyl alcohol and diacetyl but also to benzaldehyde, whereas egl-4 mutants responded normally to benzaldehyde [12].
chb-3 belongs to the Daf-c group of mutants with defects in cGMP signaling
We noted that chb-3(eg52) also exhibits a dauer-constituitive (Daf-c) phenotype. C. elegans develops into a diapause larvae called dauer if the worm experiences harsh environmental conditions such as a low food supply, a high population density or high temperature during early development. Daf-c mutants form dauers independent of the environmental conditions. About 50% of chb-3(eg52) animals formed dauers under conditions in which wild-type animals never formed dauers (Figure 1E).
Previous studies revealed that Daf-c mutants can be classified into three groups, thereby defining three branches of a pathway that act in parallel to regulate dauer formation [13]–[15]. Group I Daf-c mutants including daf-11 have defects in a cGMP-signaling pathway. For example, daf-11 encodes a receptor-type guanylyl cyclase [16]. The group I Daf-c mutants were strongly suppressed by cilium-defective mutations such as che-2, suggesting that certain sensory inputs are required for dauer formation by the group I mutants [14]. The group II Daf-c mutants have been shown to carry mutations in genes encoding components of the DAF-7/TGF-β signaling pathway [17]–[19]. Their Daf-c phenotype is suppressed by mutations in the daf-3 gene, which encodes a downstream Smad transcription factor in the TGF-β signaling pathway [14], [20]. Another group of Daf-c mutants is caused by defects in insulin signaling. The Daf-c phenotype of this group is known to be suppressed by mutations in the daf-16 gene, which encodes a FOXO transcription factor acting downstream of insulin [21].
To determine the Daf-c group in which chb-3 belongs, double mutant analyses were performed. The Daf-c phenotype of chb-3(eg52) was strongly suppressed by che-2(e1033), but not by daf-3(e1376) or daf-16(m26) (Figure 1E). In addition, one of the group I Daf-c mutants, daf-11(m47), did not exhibit a higher dauer rate when the chb-3(eg52) mutation was introduced (data not shown). These analyses suggest that chb-3 is a group I Daf-c gene, which may be involved in cGMP signaling.
Genetic analysis of the relationship among egl-4, chb-3, and gcy-12
We previously showed that the cGMP-dependent protein kinase EGL-4 regulates body size by acting downstream of sensory inputs in C. elegans [6]. Recently, we identified a receptor type guanylyl cyclase gene, gcy-12, as another class I Chb gene (manuscript in preparation). gcy-12 mutations, like those in egl-4, cause an increase in body length and volume compared with wild-type animals. Importantly, the gcy-12;egl-4 double mutant does not become larger than the egl-4 single mutant, indicating that EGL-4 and GCY-12 act in a linear pathway for body size regulation. Therefore, we hypothesized that GCY-12 produces cGMP, which activates the EGL-4 cGMP-dependent protein kinase and regulates body size in C. elegans. Because chb-3 appears to play a role in a cGMP signaling pathway for dauer regulation, we asked if CHB-3 acts in the same pathway as EGL-4 and GCY-12. The chb-3 mutation does not show synergy with either egl-4 or gcy-12 for body size; the chb-3(eg52);egl-4(ky185) and chb-3(eg52);gcy-12(ks100) double mutants were as large as the single mutants (Figure 2). Although we cannot exclude the possibility that the synergic effect was not observed by a physical constraint in the double mutants, these results suggest that CHB-3, EGL-4, and GCY-12 may function in the same regulatory pathway of body size control. To determine whether CHB-3 acts downstream or upstream of EGL-4, we used an egl-4 (gf) allele. egl-4 (ad450gf) is a gain-of-function mutant of egl-4, which seems to produce a constitutively activated EGL-4 kinase, and exhibits opposite phenotypes of egl-4(lf) including a small-body phenotype [22]. The chb-3(eg52) mutation did not increase body size in the egl-4(ad450gf) background. The chb-3(eg52); egl-4(ad450gf) double mutant animals exhibited the same body volume as egl-4(ad450gf) with the slightly shortened body length, indicating that activated EGL-4 does not require functional CHB-3 when producing a small-body phenotype (Figure 2). Therefore, CHB-3 likely acts upstream of EGL-4, such as a regulator of kinase activity.
chb-3 encodes a novel protein with a zf-MYND motif
By SNP mapping and germline transformation experiments, we identified a corresponding gene for the chb-3(eg52) mutation, Y48G1A.3 (Figure 3A, also see Materials and Methods). A genomic DNA fragment containing a single predicted gene (Y48G1A.3) restored the small body and the confined tracking pattern which are characteristic of the che-2 single mutant, when introduced into the chb-3(eg52);che-2(e1033) double mutant (Figure 1B and data not shown). The same fragment also rescued the Daf-c phenotype and the chemotaxis defect to diacetyl, when introduced into the chb-3(eg52) mutant (Figure 1E and data not shown). BLAST searches and conserved domain analysis (CDART) were performed with the predicted protein sequence of 388 amino acids. We found highly homologous proteins (∼30% identity/∼50% similarity throughout the protein) in species from flies to humans (Figure 3C), although none have defined functions. CDART revealed that CHB-3/Y48G1A.3 contains 4 consecutive ankyrin repeats in the N-terminal portion (amino acids 9-127). Ankyrin repeats are known to mediate protein-protein interactions in a diverse range of protein families. CHB-3/Y48G1A.3 also contains a motif termed a MYND type Zn-finger (zf-MYND) in the region close to the C terminus (amino acids 321-357). The zf-MYND motif is found in several different types of protein, including transcription factors and scaffolding proteins, and is thought to also mediate protein-protein interactions [23], [24].
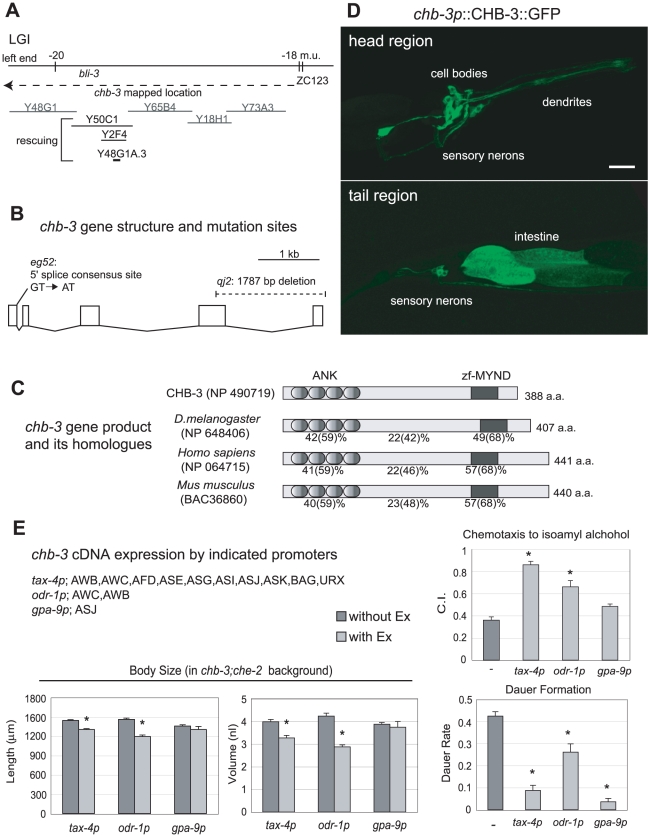
Figure 3. chb-3 encodes a novel protein with a zf-MYND motif.
(A) Genetic and physical map of the chb-3 locus. Y50C1, Y2F4 (C. elegans genomic YAC clone), and a Y48G1A.3 PCR fragment (6.5 kb) which contained the Y48G1A.3 predicted gene and 1 kb of upstream sequence, rescued the Daf-c phenotype of chb-3(eg52). The other clones are all YAC clones that failed to rescue the chb-3 phenotypes. (B) Gene structure and mutation sites of chb-3/Y48G1A.3. Exons are boxed. (C) chb-3 encodes a novel protein with four consecutive ankyrin repeats (ANK) close to the N terminus and a MYND-type Zn-finger motif (zf-MYND) close to the C terminus. Homologues from D. melanogaster (NP 648406), Homo sapiens (NP 064715), and Mus musculus (BAC36860) are included in the alignment. The amino-acid identities (and similarities in parentheses) to CHB-3 are shown for each ANK and zf-MYND domain and the region between the domains. (D) GFP expression from chb-3p::CHB-3::GFP in the head and tail regions of a wild-type animal (young adult stage). Projection of confocal microscopic sectioning images. Anterior is to the right. The scale bar represents 20 µm. (E) Phenotype rescue by chb-3 expressed under the control of the indicated sensory promoters. Sensory neurons in which each promoter drives expression are also shown. To examine rescuing of the Chb phenotype (suppression of che-2-small-body size), each construct was introduced into chb-3(eg52);che-2(e1033) animals as an extrachromosomal (Ex) array. Decrease of the size with the Ex array means rescuing of the Chb phenotype. To examine rescuing of the chemotaxis defect and Daf-c phenotype, each construct was introduced into chb-3(eg52) animals as an Ex array. Chemotaxis assays were performed with isoamyl alcohol (1/100 dilution). Animals with (light gray bars) and without (dark gray bars) the Ex array were compared. The mark (*) indicates the significant difference from control (p<0.01, t test).
Mutational analysis of chb-3(eg52)
To identify the chb-3(eg52) mutation, we sequenced the entire coding region of Y48G1A.3, revealing a mutation in the 5′ splice consensus site of the 1st intron (GT to AT; Figure 3B). 5′ splice consensus is highly conserved in C. elegans, and this change may disrupt appropriate removal of the 1st intron from chb-3 mRNA. By 5′ RACE analysis using total RNA prepared from chb-3(eg52) mutant animals, we confirmed that the 81-bp 1st intron, which is removed in mRNA isolated from wild-type animals, remained in chb-3(eg52) mRNA. This extra 81-bp sequence does not contain an in-frame stop codon, suggesting that the CHB-3 protein in chb-3(eg52) mutant animals contains 27 extra amino acids inserted into the middle of the 2nd ankyrin repeat. The chb-3(eg52) mutation is fully recessive and the mutant phenotypes can be rescued by introducing the wild-type chb-3 gene, suggesting that chb-3(eg52) is a loss-of-function mutation. chb-3(eg52), however, does not appear to be a null mutation. Therefore, we obtained a deletion mutation caused by Tc1 transposon excision in the chb-3 gene. This deletion allele, qj2, lacks approximately 300 bp of coding sequence, including the region encoding the zf-MYND domain (Figure 3B). chb-3(qj2) showed a similar but more severe phenotype than eg52. As shown in Figure 1E, qj2 exhibits a highly penetrant Daf-c phenotype, which can be suppressed by che-2. qj2 also suppresses the che-2 small-body phenotype as does eg52 (data not shown). Because of the strong Daf-c phenotype, we could not analyze qj2 for other phenotypes, such as chemotaxis behavior. The common phenotypes (Daf-c, Chb, and gcy-12 localization pattern (see below)) and the molecular lesions in eg52 and qj2, however, indicate that both eg52 and qj2 are loss-of-function mutants and that qj2 is likely a null allele.
chb-3 function is critical in a subset of sensory neurons
We created a GFP reporter construct by using the chb-3 promoter (2.3-kb upstream region) to express chb-3 cDNA fused with gfp at the 3′ terminus (chb-3p::CHB-3::GFP). GFP expression was observed in many sensory neurons, including the amphid and phasmid neurons (Figure 3D). We also observed relatively weak expression in the intestine. GFP-fused CHB-3 appeared cytoplasmic and excluded from the nucleus (Figure 3D). To identify the tissues in which chb-3 expression is required for normal function, chb-3 cDNA was expressed in various sets of sensory neurons using several sensory neuron promoters (tax-4p, odr-1p, and gpa-9p) (Figure 3E) [25]–[28]. In these constructs, a GFP reporter gene was inserted into the 3′ end of chb-3 cDNA for monitoring cellular expression and sub-cellular localization of CHB-3. Animals expressing these constructs were examined for dauer formation, chemotaxis, and suppression of the che-2 small-body phenotype (Figure 3E). The tax-4 promoter construct (tax-4p::CHB-3::GFP) rescued all the defects examined. In a chb-3;che-2 background, tax-4p::CHB-3::GFP restored the che-2-like small-body phenotype (Figure 3E). In a chb-3 background, tax-4p::CHB-3::GFP also restored chemotaxis to diacetyl and isoamyl alcohol, and decreased the rate of dauer formation (Figure 3E and data not shown). The tax-4 promoter drives expression in 10 pairs of sensory neurons, suggesting an important role of chb-3 in these sensory neurons (Figure 3E). CHB-3 expression was also limited to fewer sensory neurons using the odr-1 and gpa-9 promoters. In terms of the body size phenotypes, odr-1p expression showed rescuing activity (equivalent to tax-4p), suggesting that the AWB and AWC sensory neurons were involved in this regulatory process. For chemotaxis to isoamyl alcohol, none of these constructs rescued as strongly as was observed with tax-4p, and we were not able to identify a smaller subset of neurons in which CHB-3 specifically contributes to chemotaxis behavior. For the regulation of dauer formation, chb-3 expression in the ASJ neuron is important, because the gpa-9p construct strongly rescued this phenotype. These results suggest that CHB-3 acts in a subset of sensory neurons to regulate the sensory processing required for multiple developmental and behavioral functions in C. elegans.
The GCY-12 localization pattern is altered in chb-3 mutants
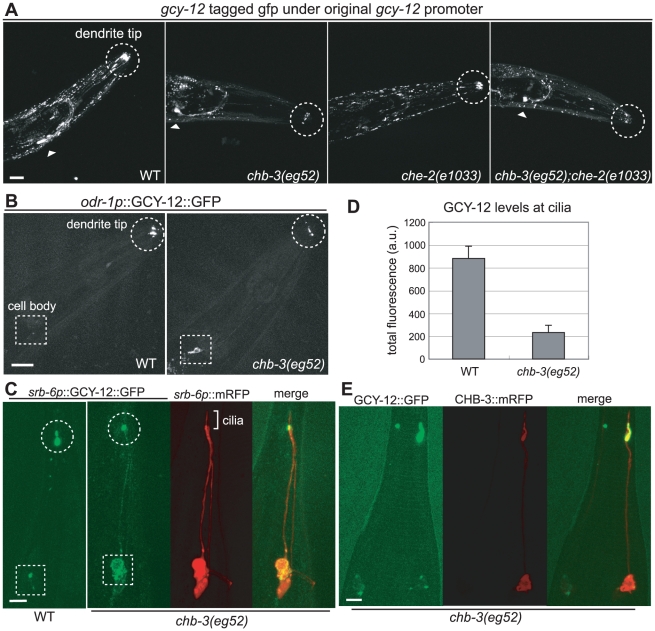
We observed GCY-12 expression and found that the GCY-12 localization pattern was changed in the chb-3 mutant background. We fused a genomic fragment, which contains a 3-kb upstream region and the entire gcy-12 coding region, with gfp at its 3′ end in frame (GCY-12::GFP). When the GCY-12::GFP construct was introduced into wild-type animals, GFP was strongly localized to the dendrite tip of sensory neurons, where cilia exist (Figure 4A). Punctuate GFP signals were also observed in head muscle tissue (Figure 4A). This expression profile likely reflected the actual expression pattern, because the construct retained the ability to rescue the gcy-12 body size phenotype (data not shown). Next, we examined the gcy-12 expression pattern in chb-3 mutants. We transferred the extrachromosomal transgene, Ex[GCY-12::GFP], by mating from wild-type animals into a chb-3 background, and found that the GCY-12 level in the dendrite tip was decreased in chb-3 mutants (Figure 4A for eg52; Figure S1 for qj2). To observe the detailed localization pattern, we expressed gcy-12 cDNA tagged with gfp in a limited number of sensory neurons; in AWC and AWB sensory neurons by using the odr-1 promoter (odr-1p::GCY-12::GFP) and in PHA and PHB sensory neurons by using the srb-6 promoter (srb-6p::GCY-12::GFP) [29]. AWC and AWB are the sensory neurons in which CHB-3 plays a vital role in body size regulation. PHA and PHB are neurons in the tail phasmid sensory organ with relatively short dendrites and morphologically simple cilia. In both sets of sensory neurons, GCY-12 at the dendrite tip was decreased in chb-3(eg52) (Figure 4B and 4C). In contrast, we found more GCY-12 accumulated in the cell bodies in many chb-3(eg52) animals (Figure 4B and 4C). We quantified the ciliary GFP levels in wild-type and chb-3(eg52) animals by 3D analysis with confocal microscopic images. The ciliary GFP level in chb-3 animals expressing odr-1p::GCY-12::GFP was 26% of that of wild-type animals (Figure 4D). Based on the expression of srb-6p::mRFP, chb-3(eg52) retained normal PHA and PHB cilia, and GCY-12::GFP was localized to the bases of cilia which is called transition zone (Figure 4C).
Figure 4. chb-3 is required for proper localization of the GCY-12 guanylyl cyclase to cilia.
(A) Expression of GFP-tagged, full-length GCY-12 under the control of the native promoter (GCY-12::GFP). Expression of the ttx-3p::mRFP injection marker in the AIY neurons is also shown (arrowheads). (B) Expression of GFP-tagged, full-length GCY-12 under the control of the odr-1 promoter (odr-1p::GCY-12::GFP). (C) Expression of GFP-tagged, full-length GCY-12 under the control of the srb-6 promoter (srb-6p::GCY-12::GFP). srb-6p::mRFP was also expressed to visualize the morphology of the cilia in chb-3(eg52). (D) Ciliary GFP levels in animals expressing odr-1p::GCY-12::GFP were quantified using the confocal 3D analysis. Numbers of examined animals; wild-type (16), chb-3 (18). Error bars indicate s.e.m. (E) Decreased GCY-12 levels in the cilia of chb-3(eg52) were rescued cell autonomously. The tail region of a chb-3(eg52) animal carrying two extrachromosomal transgenes, Ex[srb-6p::GCY-12::GFP] and Ex[srb-6p::CHB-3::mRFP], was shown. In this animal, Ex[srb-6p::CHB-3::mRFP] showed mosaic expression only in the right phasmid neuron, and only the right phasmid ciliary level of GCY-12::GFP was increased. (A, B, C, E) Projection of confocal microscopic sectioning images. Dendrite tips and cell bodies are marked with circles and squares, respectively. Head (A, B) and tail (C, E) regions of young adults are shown. Scale bars represent 20 µm (A, B) and 5 µm (C, E).
Decreased GCY-12 levels in the cilia of chb-3(eg52) animals were rescued cell autonomously. By expressing CHB-3 protein tagged with mRFP by using the srb-6 promoter (srb-6p::CHB-3::mRFP) and the odr-1 promoter (odr-1p::CHB-3::mRFP), high GCY-12 levels at the cilia were restored in PHA/PHB and in AWC/AWB neurons, respectively (Figure 4E and Figure S2). Due to the mosaic expression of the extrachromosomal transgene, we observed an animal in which CHB-3 tagged with mRFP was expressed in a phasmid neuron in one side but not in a neuron in the bilaterally opposite side. In this case, high GCY-12 level was restored only in the cilia of the CHB-3 expressing neuron (Figure 4E, the neuron in the right side), but not in the cilia of the other neuron. These results indicate that CHB-3 functions to localize GCY-12 to the cilia cell autonomously.
Decreased GCY-12 levels at the cilia is not likely due to a decrease in gcy-12 transcription, because the gcy-12 3-kb promoter fusion with gfp gave nearly the same expression level in wild-type and chb-3(eg52) animals (Figure S3). We also asked if ciliary membrane molecules other than GCY-12 require CHB-3 for their localization to cilia. We examined the expression of GFP-tagged constructs of a cyclic nucleotide-gated channel (CNG) TAX-2, and the G protein coupled receptors (GPCRs) ODR-10 (expressed under the control of its original promoter or an AWB-specific (str-1p) promoter) and SRD-1 (kindly gifted by Cori Bargmann). All of the examined proteins showed normal cilium localization in chb-3(eg52) animals, indicating that CHB-3 is not required for their cilium localization (Figure S4 and data not shown). The expression of GFP-tagged constructs also showed no apparent morphological defect of the examined cilia of chb-3(eg52) (Figure S4).
CHB-3 regulates the dendritic transport of GCY-12::GFP
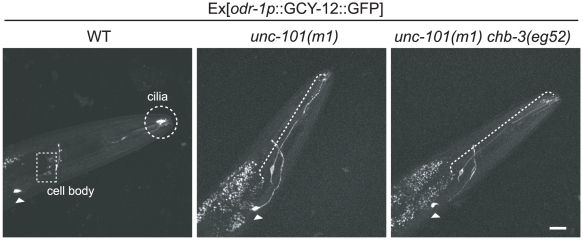
We attempted to analyze further the molecular function of CHB-3. unc-101 is a gene encoding a protein related to the µ1 subunit of the AP-1 clathrin-coated vesicle adaptor complex, which is found in the trans-Golgi network (TGN) [30]. In C. elegans, unc-101 is required for the localization of multiple ciliary membrane proteins, including several GPCRs and a receptor-type guanylyl cyclase, ODR-1 [31]. First, we examined the odr-1p::GCY-12::GFP expression pattern in a unc-101 mutant. In unc-101(m1) null animals, GFP was uniformly distributed over the entire cell surface (Figure 5), indicating that GCY-12 as well as other ciliary membrane proteins is dependent on the somatodendritic sorting by UNC-101. Next, we examined the odr-1p::GCY-12::GFP expression pattern in the chb-3(eg52) unc-101(m1) double mutant, and found that the GCY-12 localization pattern was indistinguishable from that observed in the unc-101 background; GFP was uniformly distributed over the entire cell surface in the double mutant (Figure 5). This suggests that CHB-3 acts after the somatodendritic sorting step in which UNC-101 is involved.
Figure 5. The UNC-101 clathrin adaptor AP-1 complex subunit is required for GCY-12 localization to cilia.
Expression of odr-1p::GCY-12::GFP is indicated with broken lines. Expression of ttx-3p::mRFP (injection marker) in an AIY neuron is also shown (arrowhead). Projection of confocal microscopic sectioning images of head region in young adult. Scale bar represents 20 µm.
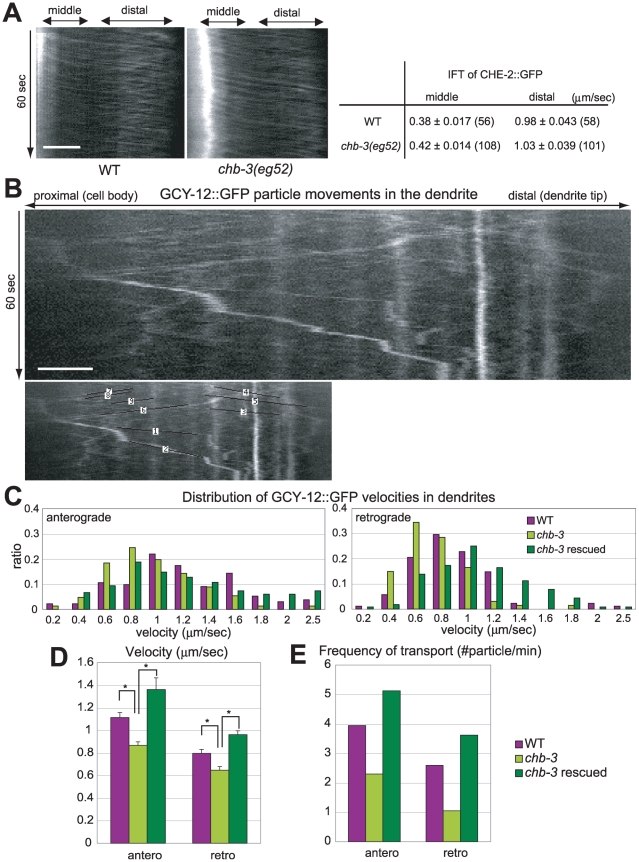
It is possible that CHB-3 may function in GCY-12 loading into the cilia and/or GCY-12 trafficking along cilia. Ciliary components, including several membrane proteins, are known to be transported from the base of the cilium to the tip of the cilium by the mechanism called the intraflagellar transport (IFT) [4], [32]. We examined if the chb-3 mutation causes a defect in IFT and consequently affects the level of GCY-12 retained in the cilia. By using time-lapse fluorescence microscopy, we observed the movement of one of the IFT components CHE-2 tagged with GFP (CHE-2::GFP) in the cilia [33], [34]. CHE-2::GFP was properly localized to the cilia in chb-3(eg52) as was observed in wild-type animals. In the cilia of both wild-type and chb-3(eg52) animals, CHE-2::GFP moved relatively slowly (∼0.4 µm/sec) along the middle segments and was accelerated to ∼1 µm/sec along the distal segments in an anterograde direction (Figure 6A). Previously, it was reported that IFT along the middle and distal segments of cilia utilize distinct anterograde motor systems and exhibit different rates of IFT movements [35]. Our result is consistent with that and suggests that the chb-3 mutant retains the normal IFT machinery. We did not detect the intraflagellar transport of GCY-12::GFP, most of which was accumulated at the bases of cilia, transition zone.
Figure 6. chb-3 is required for proper GCY-12::GFP transport along the dendrites to cilia.
(A) left; Kymographs depicting CHE-2::GFP particle motility within the cilia. right; The velocities of CHE-2::GFP particles along the middle segments and the distal segments of cilia are shown as the average ± s.e.m. (number of particles). (B) A kymograph depicting GCY-12::GFP particle motility along the dendrite of a phasmid sensory neuron in a wild-type animal. Corresponding lines, with which the velocity and the frequency were analyzed, are also shown. The movie is available upon requests. (A, B) The horizontal and vertical axes represent distance and time, respectively. Horizontal scale bar; 2 µm. (C) Distribution of GCY-12::GFP particle velocities in dendrites. (D) Average velocities of GCY-12::GFP particles in anterograde and retrograde directions in the dendrites with s.e.m. Asterisk indicates the significant difference at p<0.001 (t test). (E) Frequencies of GCY-12::GFP particles in anterograde and retrograde directions in the dendrites. (C, D, E) Wild-type, chb-3(eg52) and chb-3(eg52);Ex[srb-6p::CHB-3::mRFP](chb-3 rescued) animals were carrying Ex[srb-6p::GCY-12::GFP] transgene, and their dendrites of the phasmid neurons were observed. Total time of observation, total numbers of particles in anterograde direction and in retrograde direction are follows; WT (34 min, 134, 88), chb-3 (64 min, 147, 67), chb-3 rescued (32 min, 164, 116).
Next, we observed the dendritic transport of GCY-12::GFP. Because GCY-12::GFP appeared to be increased in the cell bodies of chb-3 mutant animals, we examined if the chb-3 mutation causes a defect in the dendritic transport of GCY-12 to the cilium base. First, we observed wild-type animals carrying Ex[srb-6p::GCY-12::GFP] transgene, and detected bidirectional movements of GCY-12::GFP particles along the dendrite of the phasmid sensory neurons by time-lapse fluorescence microscopy (Figure 6B). Depending on directionality, GCY-12::GFP particles move at different rates. Anterograde movement averages 1.11 (s.e.m. ±0.044, n = 134) µm/sec and retrograde movement averages 0.80 (s.e.m. ±0.036, n = 88) µm/sec (Figure 6C and 6D). These rates are comparable to those of the ODR-10 G-protein coupled receptor (anterograde: 1.42 µm/sec, retrograde: 0.71 µm/sec [31]) and those of the PKD-2 polycystin 2 channel (anterograde: 1.26 µm/sec, retrograde: 0.81 µm/sec [36]). Since the dendritic transport of these ciliary membrane proteins was observed in different sensory neurons (GCY-12 in PHA and PHB phasmid neurons, ODR-10 in AWB neurons, PKD-2 in CEM and RnB neurons), these results suggest that common dendritic transport machineries are used for membrane protein trafficking to cilia. Then, we analyzed the dendritic transport of GCY-12::GFP in the chb-3(eg52) mutant. We observed chb-3(eg52) carrying Ex[srb-6p::GCY-12::GFP] transgene, and found that GCY-12::GFP particle motility in the dendrites of the phasmid neurons was decreased (Figure 6C–6E and Figure S5). Compared to wild-type animals, the average velocities both in anterograde and in retrograde direction were lower in the chb-3(eg52) mutant (Figure 6D). Furthermore, the numbers of observed particles which move at least 3 µm in a steady rate were decreased both in anterograde and in retrograde direction in chb-3(eg52) (Figure 6E). chb-3(eg52) animals carrying not only Ex[srb-6p::GCY-12::GFP] but also Ex[srb-6p::CHB-3::mRFP] transgene (chb-3 rescued) recovered motility and exhibited even higher velocities and frequencies of GCY-12::GFP particle movement than wild-type animals (Figure 6D, 6E, and Figure S5). These results suggest that CHB-3 regulates the dendritic transport of GCY-12::GFP, and that decreased levels of GCY-12::GFP in the cilia of chb-3 may be caused by the defect in the dendritic transport.
The decrease in ciliary GCY-12 levels may affect the body size
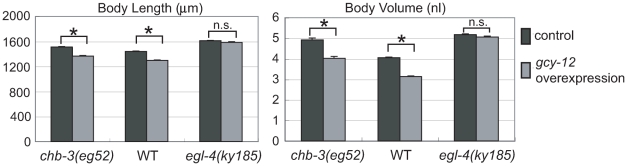
We examined whether decreased levels of GCY-12 in the cilia cause the chb-3 body-size phenotypes. If the decrease in ciliary GCY-12 caused the chb-3 large-body phenotype, overexpression of gcy-12 may rescue the phenotype. Figure 7 shows the effect on body size of gcy-12 overexpression by the original gcy-12 promoter. Overexpressing the gcy-12 gene indeed restored the chb-3(eg52) body size to a level comparable to wild-type animals. gcy-12 overexpression also decreased the body size of wild-type animals, suggesting that gcy-12 was acting in a dose-dependent manner. These GCY-12 overexpression effects were likely through modulation of EGL-4 activity, since overexpression did not affect the body size of the egl-4(ky185) mutant (Figure 7).
Figure 7. The effect of gcy-12 overexpression on body size.
Genomic DNA containing the entire coding region of gcy-12 and its upstream region (3 kb) was introduced into the animals as an extrachromosomal array (Ex). Animals with (gray bars) and without (black) the Ex array were compared. Error bars indicate s.e.m. The mark (*) indicates the significant difference (p<0.005, t test), and “n.s.” indicates that the difference is not significant.
The suppression of the che-2 small-body phenotype by mutations in gcy-12 or egl-4 indicates that the che-2 small-body phenotype is caused by the functions of these molecules. The lack of normal cilia seems to cause inappropriate activation of this signaling pathway, which ultimately makes the body size small. As shown in Figure 4A, we found that GCY-12 was still localized to dendrite tips in the che-2(e1033) mutant. In the che-2(e1033) mutant, the distal and middle segments but not transition zone of the cilia are missing [5]. GCY-12::GFP may be localized to transition zone of the shortened cilia of the che-2 mutant. On the other hand, GCY-12 in the dendrite tip was decreased in the chb-3(eg52);che-2(e1033) double mutant, as was observed for the chb-3 single mutant (Figure 4A). Because mutations in chb-3, as well as in gcy-12 and egl-4, suppress the che-2 small-body phenotype, these localization patterns suggest that GCY-12 needs to be at the cilia (or their bases) for activation of the GCY-12-EGL-4 signaling pathway. The shortened sensory cilia of che-2 mutants may activate GCY-12 to induce the small-body phenotype, then gcy-12 and chb-3 mutations may suppress this phenotype by abolishing and decreasing GCY-12 protein levels in the cilia, respectively. Thus, the ciliary GCY-12 levels may affect body size, and CHB-3 is an important regulator of the localization processes.
Discussion
Cilia are important sensory organelles, which are now thought to be essential regulators of numerous signaling pathways. In this paper, we identified chb-3, a new Chb (che-2 small body size suppressor) gene, that is required for body size control through the cilia in C. elegans. Genetic analyses placed chb-3 in the same pathway as egl-4 (a gene encoding a cGMP-dependent protein kinase) and gcy-12 (a gene encoding a receptor-type guanylyl cyclase), and probably upstream of egl-4. chb-3 encodes a highly conserved protein from worms to humans, which contains ankyrin repeats and a zf-MYND motif. We found that CHB-3 acts cell autonomously and mediates GCY-12 localization to cilia through the regulation of the dendritic transport. We also showed that decreased ciliary levels of GCY-12 are a probable cause of the body size change observed in chb-3. These results reveal the role of CHB-3 in ciliary membrane protein trafficking, and highlight the important role of sensory cilia in body size determination.
EGL-4 kinase is regulated positively and negatively at the cilia
Previously, we proposed a model in which sensory perception, acting through modulation of the GCY-12-EGL-4 pathway, regulates body size [6] (manuscript in preparation). Normally, activation of the GCY-12-EGL-4 pathway reduces body size. In wild-type animals, sensory inputs through cilia are likely to negatively regulate the GCY-12-EGL-4 pathway, since the loss of sensory inputs due to che-2 mutations causes inappropriate activation of the GCY-12-EGL-4 pathway and results in a small-body phenotype. In this model, sensory stimuli inhibit, rather than stimulate sensory neurons, as is reported in several sensory signal transduction pathways [37], [38]. Here, we showed that CHB-3 functions in the localization of GCY-12 to cilia and plays an indispensable role in the activation of the GCY-12-EGL-4 pathway. Thus, GCY-12 apparently needs to be present at cilia (or transition zone of cilia) to activate the EGL-4 kinase, although EGL-4 is not restricted to cilia and is rather broadly distributed throughout the entire cytoplasmic region (as far as can be observed with tagged GFP constructs [6]). This is potentially because the signaling complex (which allows for effective signal transduction) may be formed at the cilia, or because GCY-12 may require ligand binding at the cilia to be activated. Hence, the cilium is likely to be a place where the EGL-4 activity is controlled positively (by GCY-12) and negatively (by sensory inputs through unknown molecules being dependent on normal cilia). The levels of EGL-4 kinase activity are a consequence of these opposing controls, which thereby determine the ultimate body size of C. elegans.
CHB-3 acts in a small subset of sensory neurons to regulate of body size
chb-3 is expressed in multiple neurons and the intestine of C. elegans. Rescue experiments with various sensory promoters suggested that CHB-3 functions in a small subset of sensory neurons; it is sufficient to express chb-3 in two pairs of sensory neurons, AWC and AWB, for normal body size regulation. These neurons are included in a set of sensory neurons, EGL-4 expression in which is sufficient for normal body size regulation [6]. However, egl-4 expression in AWC and AWB by the odr-3 promoter is not sufficient to rescue the body size phenotype of the egl-4 mutant [6]. This suggests that EGL-4 may function for body size regulation in other sensory neurons that do not require CHB-3, too. This is consistent with the observation that the egl-4 mutant is larger than the chb-3 mutant and the gcy-12 mutant (Figure 2). EGL-4 may act dependently and also independently of CHB-3 in the regulation of body size.
Other behavioral and developmental roles of CHB-3
chb-3 mutants exhibit not only body-size phenotypes but also defects in chemotaxis and a group I type Daf-c phenotype, which are not shared by egl-4 mutants [12] and gcy-12 mutants (manuscript in preparation). It is possible that CHB-3 mediates the cilium localization of not only GCY-12 but also other signaling molecules that are required for chemotaxis and control of dauer formation. One such candidate is DAF-11, another receptor-type guanylyl cyclase that contributes to dauer regulation and chemotaxis and is localized to sensory cilia [16]. Both chb-3 and daf-11 are group I Daf-c mutants. Our analysis also indicates that the Daf-c phenotype of chb-3 is strongly rescued if chb-3 is expressed in ASJ sensory neurons, where daf-11 acts in dauer control [39]. Strikingly, Jensen VL et al. are reporting that DAF-11 cilium localization is indeed controlled by a protein encoded by daf-25 gene, which turned out to be an identical gene to chb-3/Y48G1A.3 (Jansen VL et al., 2010, in this issue). Thus, based on our and their results, CHB-3/DAF-25 mediates the cilium localization of at least two different receptor-type guanylyl cyclases of C. elegans. Although we observed that several other cilium membrane proteins, including GPCRs and CNG, localized to cilia independent of CHB-3, further analyses may reveal additional molecules that are dependent on.
CHB-3/DAF-25–mediated trafficking
Interestingly, we found that the unc-101(m1) mutation almost completely suppressed the Daf-c phenotype of the chb-3(eg52) mutant (Figure S6). We speculate that the unc-101 mutation may bypass the requirement for CHB-3 by causing the re-localization of signaling components such as DAF-11 throughout the cell surface and consequently elevating the level of these molecules at cilia. If this were the case, unc-101 would also suppress the body size phenotype of chb-3 by distributing GCY-12 throughout the cell surface. We, however, failed to observe this, because the unc-101(m1) mutant itself has a small-body phenotype probably due to a muscle contraction defect.
chb-3 encodes a highly conserved zf-MYND protein that may act in the dendritic transport of ciliary membrane proteins
Recently, cilia have become the focus of intensive studies evaluating their contributions to the transduction of extracellular signals. It is clear that cilia play fundamental roles in several mammalian processes, including Sonic Hedgehog morphogen signaling and left-right axis determination [40]. Ciliary dysfunction also causes a number of human disorders (ciliopathies), including Bardet-Biedl syndrome (BBS) characterized by retinal degeneration, early-onset obesity and cognitive impairment [3]. Proteins mutated in BBS are shown to be involved in the regulation of membrane proteins trafficking into cilia with a small GTPase, Rab8 [41]. Rab8 and several other small GTPases regulate the trafficking of post-TGN vesicles to cilia [41], [42]. However, our understanding of how membrane proteins trafficking from Golgi to the cilium is regulated remains extremely limited [43]. Although only a small number of proteins involved in this process have been identified, the analyses of these proteins suggest that there is an active and highly directed mechanism for placement of membrane proteins to the cilia. IFT20 is a conserved protein which dynamically moves between the Golgi complex and the cilium, and is thought to act in the delivery of ciliary membrane proteins [44]. UNC-101 clathrin AP-1 µ1 subunit in C. elegans acts in the sorting or packaging of ciliary membrane proteins at trans-Golgi network [31]. In C. elegans, ODR-4 membrane protein is thought to act in odorant receptor folding or transport to cilia [45], and CIL-1 PI 5-phosphatase is required to confine PKD-2 polycystin 2 channel to the cilia [46].
Here, we show that CHB-3 may act in the dendritic transport of GCY-12 to the cilia. chb-3 encodes a novel zf-MYND protein, which is highly conserved from C. elegans to humans. The zf-MYND motif, containing two putative zinc fingers, is found in several different types of proteins [47]. Some zf-MYND proteins act as histone deacetylase-dependent transcriptional repressors through binding a silencing mediator complex with their zf-MYND domains [23], [24], [48]–[50]. Another zf-MYND protein, Drosophila nervy, binds a cAMP-dependent protein kinase and the Plexin receptor, and controls neuronal axon guidance by integrating different guidance cues [51], [52]. Thus, the zf-MYND motif appears to mediate specific protein-protein interactions. CHB-3 also contains N-terminal ankyrin repeats, which are generally involved in protein-protein interactions. Two alleles of chb-3, eg52 and qj2, are likely to produce a protein with disturbed ankyrin repeats and a protein lacking the zf-MYND domain, respectively. These alleles exhibit an equivalent defect in GCY-12 localization, indicating that both ankyrin repeats and the zf-MYND motif are important for this function. The homologous proteins of CHB-3 in other species, which also possess these two domains, might be also involved in dendritic transport.
In unc-101 mutants, GCY-12 is evenly distributed over the plasma membranes. This may be because loss of AP-1 µ1 causes GCY-12 to enter a different class of vesicle at the TGN that can fuse anywhere in the plasma membrane. In the chb-3 unc-101 mutant, the GCY-12 localization pattern was indistinguishable from that observed in the unc-101 background, suggesting that chb-3 mutations possibly disrupt steps following the TGN rather than processing occurring before the TGN. We observed that GCY-12::GFP particles move in the dendrites of sensory ciliated neurons. The velocities in anterograde and retrograde directions are comparable to the previously reported velocities of other ciliary membrane proteins in C. elegans. This indicates utilization of common trafficking machineries, although no motor protein such as dynein and kinesin has been identified for ciliary protein transport along dendrites in C. elegans. In the chb-3 mutant, GCY-12::GFP particle motility in the dendrites is decreased; GCY-12::GFP particles move significantly slower and particle movements are observed less frequently. We did not detect dendritic movement of CHB-3::GFP, which was rather evenly distributed in the dendrite. Thus, CHB-3 itself does not appear to move along dendrites. Given that the CHB-3 protein possesses two protein-protein interaction domains, one possible function of CHB-3 is to associate the post-TGN vesicles containing GCY-12::GFP with dendritic transport machineries as a transient adaptor protein. Alternatively, it is possible that CHB-3 itself is a component of the dendritic transport machinery. In either case, CHB-3 seems to act not generally for all ciliary membrane protiens but specifically for a limited number of proteins including GCY-12. Future studies, including the search for proteins interacting with CHB-3 and their homologues in higher organisms, may fully reveal the molecular functions of this protein and mechanisms of ciliary targeting. The search for additional Chb mutations and analyses of their molecular functions may also lead to the identification of other regulators acting in the system.
Our study has identified a new regulator, CHB-3, of the trafficking of membrane proteins to cilia. We also demonstrate the importance of ciliary targeting of a guanylyl cycalse, GCY-12, in sensory-dependent body size regulation. Given that CHB-3 is highly conserved, a similar system might be used for the control of trafficking of signaling proteins in various species.
Materials and Methods
Strains and genetics
C. elegans strain N2 was used as wild-type animals. Worms were grown at 20°C with food, E. coli OP50, using standard methods [53].
The other strains used in this work included chb-3(eg52), chb-3(qj2), gcy-12(ks100), che-2(e1033), egl-4(ky185), daf-3(e1376), daf-16(m26), daf-11(m47), bli-3(e767), and unc-101(m1). Double mutants were generated using standard methods and confirmed by complementation testing or sequencing.
The che-2; Ex[che-2(EcoT14I)/H20::gfp] strain was kindly provided by Isao Katsura [33]. Ex[che-2(EcoT14I)/H20::gfp] (Ex[che-2]) is an extrachromosomal array containing a che-2 gene with its original promoter, and retains a rescuing ability of the che-2 small-body phenotype [6].
Body size measurement
L4 crescent stage animals were transferred to 6-cm seeded NGM plates and maintained under well-fed conditions until the body size was measured. For body length and volume, animals were anesthetized with NaN3 48 hours after the L4 stage and measured by using ‘Senchu-gazou-kaiseki-souchi SVK-3A’ (Showa Denki Co., Fukuoka, Japan) as described by Hirose et al. [7]. As another indicator of the body size, we measured the length of perimeter of the lateral image of the animals as previously described [6]. Each data point represents the average volume and length of more than 20 animals. All strains plotted on each graph were measured at the same time.
Locomotory behavioral assay
All seeded plates for the locomotory behavioral assay were freshly spread with bacteria, incubated overnight at 37°C, and allowed to cool to room temperature before use. For the tracking assay; a single well-fed L4 crescent stage animal was transferred to an individual 3.5-cm seeded NGM plate, and allowed to move freely for 17–20 hours at room temperature. The assay plates were not moved during this period to avoid any mechanical stimulation.
Dye-filling assay and other behavioral assays
Dye-filling assay; assays were done as described by Starich et al. using DiI (10 µg/ml) [54].
Osmotic avoidance; assays were performed basically as described by Culotti and Russell [55]. First, rings (1 cm in diameter) were drawn on plates with 50 µl of NaCl solution of indicated concentration. Then, 25 animals were placed inside of the ring. After 30 minutes, the animals inside and outside of the ring were counted. The avoidance index was calculated as the number of animals which stayed inside of the ring/the total number of animals.
Chemotaxis; assays were done as described by Bargmann [56]. One microliter of isoamyl alcohol (1/100 dilution), diacetyl (1/100 dilution) or benzaldehyde (1/100 dilution) for volatile attractants, and agar plugs (7 mm in diameter) soaked with sodium acetate (0.4 M) or lysine acetate (1 M) for soluble attractants, were used.
Dauer formation; eggs were gathered by bleaching well-fed adults, and placed on seeded plates (200–300 eggs/plate). After 50 hours at 20°C, the dauers and non-dauers were counted. The criteria for dauer were a dark intestine, long shape, suppressed pumping, and lethargic movements.
Quantification of GCY-12–tagged GFP in cilia
Young adult animals (48 hours from the L4 stage) harboring Ex[odr-1p::GCY-12::GFP] were examined for fluorescence at the nose tip. An LSM510 confocal microscope (Carl Zeiss) equipped with a 100x objective lens was used to scan GFP fluorescence using a 488-nm excitation beam and a BP 500–550 nm emission filter. Nose tips were optically sliced in 0.7-µm intervals, and the stacked images were processed with “3D for LSM” software (Carl Zeiss) to calculate the total fluorescence at the nose tip (the product of the volume of the fluorescent area and the mean fluorescence). Each data point represents the average from 16-18 animals.
IFT and dendritic transport analyses
Time-lapse imaging analyses were performed basically as described by Signor et al. [57]. Time-lapse images were obtained with Olympus IX71 microscope equipped with 100 x, 1.4 N.A. objective and ORCA-ER digital camera (Hamamatsu) with Uniblitz VMM-D1 shutter unit (Ancient Vassociates). All images were collected at 1 frame/0.5 sec intervals, and at 100–400 msec exposure time for 1 min duration, with adult hermaphrodite anesthetized with 10 mM levamisole. When animal's positions were not completely fixed by levamisole, those were digitally fixed by the custom registration software (written in Matlab). In the software, the algorithm minimizes the mean square difference between the pixel intensities of ROI (region of interest) of two consecutive time slices [58]. Then, movies and kymographs were created by using MetaMorph software. Animals were cultured at 20°C and observed at 22–23°C. The intraflagellar transport of CHE-2::GFP was analyzed with phasmid neurons of animals expressing CHE-2 tagged GFP driven under the che-2 promoter [33]. Our IFT analysis of CHE-2::GFP in wild-type animals yielded velocities which are slightly slower than those in a previous report (anterograde middle segment; 0.7 µm/sec, anterograde distal segment; 1.3 µm/sec [59]). This may be due to the differences in the observing conditions. For the analysis of GCY-12::GFP dendritic transport, the same transgene (Ex[srb-6p::GCY-12::GFP]) was transferred to N2 and chb-3(eg52) by mating for comparison under the same expression level of GCY-12::GFP. As the “rescued chb-3(eg52)” line, Ex[srb-6p::CHB-3::mRFP] transgene was introduced into chb-3(eg52); Ex[srb-6p::GCY-12::GFP], and animals carrying two transgenes were used. The frequencies of GCY-12 transport along dendrites were analyzed based on kymographs, and a path containing at least one straight segment corresponding to a movement longer than 3 µm in a steady rate was counted as 1 event. If a GCY-12::GFP particle changes the speeds or the directions in a path, the longest straight segment was analyzed for the representative speed and direction of the particle.
Molecular biology methods
For the chb-3 cDNA expression analysis, most of the chb-3 cDNA was obtained from the yk280C9 cDNA clone (provided by Yuji Kohara). The 5′ region of the first exon was obtained using 5′ RACE (5′-RACE system, Invitrogen), and fused to the yk280C9 cDNA to create a full-length cDNA. A BamHI site just upstream of the initial ATG codon was artificially created using PCRs. The stop codon was changed to a BalI site using PCRs. The cDNA was inserted in-frame into pPD95.75 (GFP expression vector; a gift from Andy Fire) using the BamHI and BalI sites (CHB-3::GFP vector). Upstream promoter sequences from chb-3, tax-4, odr-1, srb-6 and gpa-9 (2.3 kb for chb-3, and approximately 3 kb for the other) were obtained using PCRs and inserted upstream of the chb-3 coding region of the CHB-3::GFP vector to create chb-3p::CHB-3::GFP, tax-4p::CHB-3::GFP, odr-1p::CHB-3::GFP, srb-6p::CHB-3::GFP, and gpa-9p::CHB-3::GFP, respectively. The activities of these promoters (other than chb-3) have previously been reported [25]-[29] and were confirmed using the GFP expression patterns obtained with each construct. odr-1p::CHB-3::mRFP, srb-6p::CHB-3::mRFP were created by replacing the GFP coding region with mRFP cDNA.
The GCY-12::GFP construct was made by inserting an 8.1-kb genomic fragment, which contained 3 kb of the promoter region and the entire coding region, in-frame into pPD95.77 using the XbaI and BalI sites. The stop codon was changed to a BalI site using PCRs such that GFP was added to the C terminus of GCY-12. The odr-1p::GCY-12::GFP and srb-6p::GCY-12::GFP constructs were made by inserting the odr-1 promoter region (2.4 kb) or the srb-6 promoter region (3.3 kb) with gcy-12 cDNA (obtained from the yk316d1 cDNA clone provided by Yuji Kohara; the most 5′ region was supplemented with a product of 5′ RACE) into pPD95.77.
Generation of transgenic worms
Transgenic strains were generated by standard microinjection methods [60]. Unless otherwise noted, test DNA was injected at 100 ng/µl with a co-injection marker such as myo-3::gfp DNA (33 ng/µl, a GFP construct expressed in body wall muscle), lin-44::gfp (33 ng/µl, a GFP construct expressed in hypodermal cells at the tip of the tail) or ttx-3::mRFP (33 ng/µl, a RFP construct expressed in the AIY neuron). Generally we isolated two to five independent transgenic lines for each injection and confirmed that there were no major differences between them. To compare the expression profiles in different genetic backgrounds, the same extrachromosomal arrays, which were transferred by mating between strains, were used.
For gcy-12 overexpression analysis, a genomic fragment encompassing 3 kb of the promoter region and the entire coding region of gcy-12 was sub-cloned into pPD49.26 (expression vector; a gift from Andy Fire), and injected into N2 animals at 150 ng/µl with a injection marker (33 ng/µl sra-6::gfp).
Mapping and cloning of chb-3
chb-3(eg52) was mapped using the SNP method [61] based on its suppression of the confined tracking phenotype associated with che-2. After mating chb-3(eg52);che-2(e1033); Ex [che-2] and CB4856 animals, we isolated approximately 100 F2 chb-3(eg52);che-2(e1033) animals, which were identified by a cilium defect (indicated by a dye-filling defect) and the wide ranged tracking pattern. Genotype analysis of these lines allowed us to map chb-3(eg52) to the left end of LG I. Further mapping was performed based on the Daf-c phenotype. The dauer phenotype was examined in F3 animals instead of F2 offspring of chb-3(eg52) and CB4856, because the chb-3 dauer phenotype was maternally rescued; chb-3/+ hermaphrodites did not produce dauer progenies. Thus, F2 progenitors were bleached for their F3 eggs. F3 animals were cultured for 50–60 hours under well-fed conditions and ∼300 dauers were isolated as chb-3 homozygotes. Genotype analysis of these dauer lines allowed us to map chb-3 to a ∼820-kb region between the ZC123 marker and the left end of LGI. Six YACs (Y48G1C, Y50C1, Y65B4, Y18H1, Y73A3, and Y2F4) covering this region were introduced separately into chb-3(eg52) animals. Two overlapping YACs (Y50C1 and Y2F4) rescued the Daf-c phenotype. Y2F4, a relatively small YAC (190∼230 kb), contained approximately 30 predicted genes. We PCR-amplified genomic fragments covering each gene, and introduced them into chb-3(eg52) animals. A 6.5-kb PCR fragment, which was amplified using primers 5′-GATGTTTTCATGGGATGTGCAC-3′ and 5′-TCGGAGATCAATTTTGAGGGC-3′, rescued the Daf-c phenotype of chb-3(eg52). This PCR fragment contained one predicted gene, Y48G1A.3 and ∼1 kb of upstream sequence. A full-length cDNA for Y48G1A.3 was obtained by fusing the yk280C9 clone (provided by Yuji Kohara) and a 5′ fragment (obtained using 5′ RACE). This full-length cDNA also rescued each of the chb-3 phenotypes examined (see the Results).
To identify the molecular lesion in chb-3(eg52), the genomic coding sequence was amplified from the eg52 mutant. Three independent PCRs were combined and the products were directly sequenced.
The chb-3(qj2) allele was generated using Tc1 transposon insertion and subsequent excision from the chb-3 coding region basically as described by Zwaal et al. [62]. We used MT3126 mut-2(r459);dpy-19(n1347) animals as the mutator line. The isolated deletion allele (qj2) was back-crossed 7 times, and maintained as trans-hetero animals with bli-3 or as the rescued line with an Ex[tax-4p::CHB-3::GFP] transgene, because the Daf-c phenotype in qj2 homozygotes was severe and few grew into fertile adults. The qj2 allele lacks 1787 bp beginning in the middle of the 4th exon and ending 61 bp after the 5th exon. This deletion allele appears to produce a CHB-3 protein that lacks the 101 C-terminal amino acids.
Analysis of chb-3 transcripts in chb-3(eg52) animals using 5′ RACE
Total RNA was isolated from wild-type and chb-3(eg52) animals using RNeasy (Qiagen) and reverse transcribed using a chb-3 gene-specific primer (5′-AGGCGGCATACGTCCTGTTTTC-3′). After TdT tailing, the 5′-end of the chb-3 cDNA was amplified by using a 5′ RACE system (Invitrogen), an Anchor abridged primer, and a chb-3 gene specific primer (5′-ATAAGGCGGCAAACATGAGTGG-3′). The PCR products were cloned and sequenced.
Supporting Information
The GCY-12 levels in the cilia were decreased in chb-3(eg2) and chb-3(qj2) mutants. chb-3 mutants were examined in a che-2(e1033) mutation background in order to suppress the Daf-c phenotype. Head region of L4 stage animals were shown. Anterior is to the right. The same extrachromosomal array (Ex[GCY-12::GFP]) was transffered by mating from line to line for comparison.
(8.43 MB EPS)
Decreased GCY-12 levels in AWC/AWB cilia were rescued by chb-3 expression in AWC/AWB neurons. Cilia (nose tips) and cell bodies are marked with circles and squares, respectively (Upper panels) A chb-3(eg52) animal carrying the Ex[odr-1p::GCY-12::GFP] extrachromosomal array. (Bottom panels) A chb-3(eg52) animal carrying two extrachromosomal arrays: Ex[odr-1p::GCY-12::GFP] and Ex[odr-1p::CHB-3::mRFP]. Ex[odr-1p::CHB-3::mRFP] directs the expression of mRFP-tagged, full-length CHB-3 in AWC and AWB neurons. CHB-3::mRFP is distributed throughout the cell, but is excluded from the nucleus. The Ex[odr-1p::GCY-12::GFP] array contains ttx-3p::mRFP as an injection marker, which was expressed in the AIY neuron (*). Scale bar represents 20 µm.
(4.90 MB EPS)
The expression patterns of gcy-12 promoter fusion with gfp in wild-type (N2) and chb-3(eg52) animals. The same extrachromosomal array (Ex[gcy-12p::GFP]) array was used for comparison. The anterior regions of adult animals are shown. No differences in the expression level and pattern between N2 and chb-3 are seen.
(2.09 MB EPS)
In chb-3(eg52) animals, ODR-10 GPCR normally localizes to cilia of AWA sensory neuron (top panel) and AWB sensory neuron (bottom panel). For the localizaiton patterns in wild-type animals, see Dwyer et al., 1998 [45]. GFP patterns also show normal cilium morpholgy of these sensory neurons in chb-3(eg52).
(1.52 MB EPS)
Kymographs depicting GCY-12::GFP particle motility along the dendrite of a phasmid sensory neuron in chb-3(eg52) (Upper panels) and chb-3(eg52) rescued (Lower panels) animals are shown. The horizontal and vertical axes represent distance and time, respectively. Corresponding lines, with which the velocity and the frequency were analyzed, are also shown. Movies are available upon requests.
(1.57 MB EPS)
The rate of dauer formation at 20°C. The Daf-c phenotype of chb-3(eg52) is suppressed by unc-101(m1). Error bars indicate s.e.m.
(0.32 MB EPS)
Acknowledgments
We thank the C. elegans Genetics Center for worm strains, the Sanger Center for YAC clones, Andy Fire for the various vectors, Yuji Kohara for the cDNA clones, and Cornelia I. Bargmann for the GFP tagged constructs of TAX-4, SRD-1, and ODR-10. We thank Isao Katsura for comments on the manuscript. We also thank V. L. Jensen and D. L. Riddle for communication of their work before publication.
Footnotes
The authors have declared that no competing interests exist.
This work was supported by Grants-in-Aid for Scientific Research (B) (18370004 to TI), for Young Scientists (B) (15770002 to MF), and on Priority Areas-Molecular Brain Science (17024044 to TI) and Systems Molecular Ethology (20115003 to TI) of the Ministry of Education, Culture, Sports, Science, and Technology of Japan (http://www.mext.go.jp/english/), as well as other grants from The Uehara Foundation (TI)(http://www.ueharazaidan.com), The Naito Foudation (TI) (http://www.naito-f.or.jp), and The Asahi Glass Foundation (TI) (http://www.af-info.or.jp/en/). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Fliegauf M, Benzing T, Omran H. When cilia go bad: cilia defects and ciliopathies. Nat Rev Mol Cell Biol. 2007;8:880–93. doi: 10.1038/nrm2278. [DOI] [PubMed] [Google Scholar]
- 2.Pazour GJ, Witman GB. The vertebrate primary cilium is a sensory organelle. Curr Opin Cell Biol. 2003;15:105–10. doi: 10.1016/s0955-0674(02)00012-1. [DOI] [PubMed] [Google Scholar]
- 3.Beales PL. Lifting the lid on Pandora's box: the Bardet-Biedl syndrome. Curr Opin Genet Dev. 2005;15:315–23. doi: 10.1016/j.gde.2005.04.006. [DOI] [PubMed] [Google Scholar]
- 4.Scholey JM, Anderson KV. Intraflagellar transport and cilium-based signaling. Cell. 2006;125:439–42. doi: 10.1016/j.cell.2006.04.013. [DOI] [PubMed] [Google Scholar]
- 5.Lewis JA, Hodgkin JA. Specific neuroanatomical changes in chemosensory mutants of the nematode Caenorhabditis elegans. J Comp Neurol. 1977;172:489–510. doi: 10.1002/cne.901720306. [DOI] [PubMed] [Google Scholar]
- 6.Fujiwara M, Sengupta P, McIntire SL. Regulation of body size and behavioral state of C. elegans by sensory perception and the EGL-4 cGMP-dependent protein kinase. Neuron. 2002;36:1091–102. doi: 10.1016/s0896-6273(02)01093-0. [DOI] [PubMed] [Google Scholar]
- 7.Hirose T, Nakano Y, Nagamatsu Y, Misumi T, Ohta H, et al. Cyclic GMP-dependent protein kinase EGL-4 controls body size and lifespan in C elegans. Development. 2003;130:1089–99. doi: 10.1242/dev.00330. [DOI] [PubMed] [Google Scholar]
- 8.L'Etoile ND, Coburn CM, Eastham J, Kistler A, Gallegos G, et al. The cyclic GMP-dependent protein kinase EGL-4 regulates olfactory adaptation in C. elegans. Neuron. 2002;36:1079–89. doi: 10.1016/s0896-6273(02)01066-8. [DOI] [PubMed] [Google Scholar]
- 9.Lozano E, Saez AG, Flemming AJ, Cunha A, Leroi AM. Regulation of growth by ploidy in Caenorhabditis elegans. Curr Biol. 2006;16:493–8. doi: 10.1016/j.cub.2006.01.048. [DOI] [PubMed] [Google Scholar]
- 10.Tain LS, Lozano E, Saez AG, Leroi AM. Dietary regulation of hypodermal polyploidization in C. elegans. BMC Dev Biol. 2008;8:28. doi: 10.1186/1471-213X-8-28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.van der Linden AM, Wiener S, You YJ, Kim K, Avery L, et al. The EGL-4 PKG acts with KIN-29 salt-inducible kinase and protein kinase A to regulate chemoreceptor gene expression and sensory behaviors in Caenorhabditis elegans. Genetics. 2008;180:1475–91. doi: 10.1534/genetics.108.094771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Daniels SA, Ailion M, Thomas JH, Sengupta P. egl-4 acts through a transforming growth factor-beta/SMAD pathway in Caenorhabditis elegans to regulate multiple neuronal circuits in response to sensory cues. Genetics. 2000;156:123–41. doi: 10.1093/genetics/156.1.123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Riddle DL, Swanson MM, Albert PS. Interacting genes in nematode dauer larva formation. Nature. 1981;290:668–71. doi: 10.1038/290668a0. [DOI] [PubMed] [Google Scholar]
- 14.Thomas JH, Birnby DA, Vowels JJ. Evidence for parallel processing of sensory information controlling dauer formation in Caenorhabditis elegans. Genetics. 1993;134:1105–17. doi: 10.1093/genetics/134.4.1105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Vowels JJ, Thomas JH. Genetic analysis of chemosensory control of dauer formation in Caenorhabditis elegans. Genetics. 1992;130:105–23. doi: 10.1093/genetics/130.1.105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Birnby DA, Link EM, Vowels JJ, Tian H, Colacurcio PL, et al. A transmembrane guanylyl cyclase (DAF-11) and Hsp90 (DAF-21) regulate a common set of chemosensory behaviors in caenorhabditis elegans. Genetics. 2000;155:85–104. doi: 10.1093/genetics/155.1.85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Georgi LL, Albert PS, Riddle DL. daf-1, a C. elegans gene controlling dauer larva development, encodes a novel receptor protein kinase. Cell. 1990;61:635–45. doi: 10.1016/0092-8674(90)90475-t. [DOI] [PubMed] [Google Scholar]
- 18.Estevez M, Attisano L, Wrana JL, Albert PS, Massague J, et al. The daf-4 gene encodes a bone morphogenetic protein receptor controlling C. elegans dauer larva development. Nature. 1993;365:644–9. doi: 10.1038/365644a0. [DOI] [PubMed] [Google Scholar]
- 19.Ren P, Lim CS, Johnsen R, Albert PS, Pilgrim D, et al. Control of C. elegans larval development by neuronal expression of a TGF-beta homolog. Science. 1996;274:1389–91. doi: 10.1126/science.274.5291.1389. [DOI] [PubMed] [Google Scholar]
- 20.Patterson GI, Koweek A, Wong A, Liu Y, Ruvkun G. The DAF-3 Smad protein antagonizes TGF-beta-related receptor signaling in the Caenorhabditis elegans dauer pathway. Genes Dev. 1997;11:2679–90. doi: 10.1101/gad.11.20.2679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gottlieb S, Ruvkun G. daf-2, daf-16 and daf-23: genetically interacting genes controlling Dauer formation in Caenorhabditis elegans. Genetics. 1994;137:107–20. doi: 10.1093/genetics/137.1.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Raizen DM, Cullison KM, Pack AI, Sundaram MV. A novel gain-of-function mutant of the cyclic GMP-dependent protein kinase egl-4 affects multiple physiological processes in Caenorhabditis elegans. Genetics. 2006;173:177–87. doi: 10.1534/genetics.106.057380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ansieau S, Leutz A. The conserved Mynd domain of BS69 binds cellular and oncoviral proteins through a common PXLXP motif. J Biol Chem. 2002;277:4906–10. doi: 10.1074/jbc.M110078200. [DOI] [PubMed] [Google Scholar]
- 24.Lutterbach B, Sun D, Schuetz J, Hiebert SW. The MYND motif is required for repression of basal transcription from the multidrug resistance 1 promoter by the t(8;21) fusion protein. Mol Cell Biol. 1998;18:3604–11. doi: 10.1128/mcb.18.6.3604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Komatsu H, Mori I, Rhee JS, Akaike N, Ohshima Y. Mutations in a cyclic nucleotide-gated channel lead to abnormal thermosensation and chemosensation in C. elegans. Neuron. 1996;17:707–18. doi: 10.1016/s0896-6273(00)80202-0. [DOI] [PubMed] [Google Scholar]
- 26.L'Etoile ND, Bargmann CI. Olfaction and odor discrimination are mediated by the C. elegans guanylyl cyclase ODR-1. Neuron. 2000;25:575–86. doi: 10.1016/s0896-6273(00)81061-2. [DOI] [PubMed] [Google Scholar]
- 27.Peckol EL, Zallen JA, Yarrow JC, Bargmann CI. Sensory activity affects sensory axon development in C. elegans. Development. 1999;126:1891–902. doi: 10.1242/dev.126.9.1891. [DOI] [PubMed] [Google Scholar]
- 28.Jansen G, Thijssen KL, Werner P, van der Horst M, Hazendonk E, et al. The complete family of genes encoding G proteins of Caenorhabditis elegans. Nat Genet. 1999;21:414–9. doi: 10.1038/7753. [DOI] [PubMed] [Google Scholar]
- 29.Troemel ER, Chou JH, Dwyer ND, Colbert HA, Bargmann CI. Divergent seven transmembrane receptors are candidate chemosensory receptors in C. elegans. Cell. 1995;83:207–18. doi: 10.1016/0092-8674(95)90162-0. [DOI] [PubMed] [Google Scholar]
- 30.Lee J, Jongeward GD, Sternberg PW. unc-101, a gene required for many aspects of Caenorhabditis elegans development and behavior, encodes a clathrin-associated protein. Genes Dev. 1994;8:60–73. doi: 10.1101/gad.8.1.60. [DOI] [PubMed] [Google Scholar]
- 31.Dwyer ND, Adler CE, Crump JG, L'Etoile ND, Bargmann CI. Polarized dendritic transport and the AP-1 mu1 clathrin adaptor UNC-101 localize odorant receptors to olfactory cilia. Neuron. 2001;31:277–87. doi: 10.1016/s0896-6273(01)00361-0. [DOI] [PubMed] [Google Scholar]
- 32.Qin H, Burnette DT, Bae YK, Forscher P, Barr MM, et al. Intraflagellar transport is required for the vectorial movement of TRPV channels in the ciliary membrane. Curr Biol. 2005;15:1695–9. doi: 10.1016/j.cub.2005.08.047. [DOI] [PubMed] [Google Scholar]
- 33.Fujiwara M, Ishihara T, Katsura I. A novel WD40 protein, CHE-2, acts cell-autonomously in the formation of C. elegans sensory cilia. Development. 1999;126:4839–48. doi: 10.1242/dev.126.21.4839. [DOI] [PubMed] [Google Scholar]
- 34.Qin H, Rosenbaum JL, Barr MM. An autosomal recessive polycystic kidney disease gene homolog is involved in intraflagellar transport in C. elegans ciliated sensory neurons. Curr Biol. 2001;11:457–61. doi: 10.1016/s0960-9822(01)00122-1. [DOI] [PubMed] [Google Scholar]
- 35.Snow JJ, Ou G, Gunnarson AL, Walker MR, Zhou HM, et al. Two anterograde intraflagellar transport motors cooperate to build sensory cilia on C. elegans neurons. Nat Cell Biol. 2004;6:1109–13. doi: 10.1038/ncb1186. [DOI] [PubMed] [Google Scholar]
- 36.Bae YK, Qin H, Knobel KM, Hu J, Rosenbaum JL, et al. General and cell-type specific mechanisms target TRPP2/PKD-2 to cilia. Development. 2006;133:3859–70. doi: 10.1242/dev.02555. [DOI] [PubMed] [Google Scholar]
- 37.Bargmann CI, Horvitz HR. Control of larval development by chemosensory neurons in Caenorhabditis elegans. Science. 1991;251:1243–6. doi: 10.1126/science.2006412. [DOI] [PubMed] [Google Scholar]
- 38.Chalasani SH, Chronis N, Tsunozaki M, Gray JM, Ramot D, et al. Dissecting a circuit for olfactory behaviour in Caenorhabditis elegans. Nature. 2007;450:63–70. doi: 10.1038/nature06292. [DOI] [PubMed] [Google Scholar]
- 39.Murakami M, Koga M, Ohshima Y. DAF-7/TGF-beta expression required for the normal larval development in C. elegans is controlled by a presumed guanylyl cyclase DAF-11. Mech Dev. 2001;109:27–35. doi: 10.1016/s0925-4773(01)00507-x. [DOI] [PubMed] [Google Scholar]
- 40.Singla V, Reiter JF. The primary cilium as the cell's antenna: signaling at a sensory organelle. Science. 2006;313:629–33. doi: 10.1126/science.1124534. [DOI] [PubMed] [Google Scholar]
- 41.Nachury MV, Loktev AV, Zhang Q, Westlake CJ, Peranen J, et al. A core complex of BBS proteins cooperates with the GTPase Rab8 to promote ciliary membrane biogenesis. Cell. 2007;129:1201–13. doi: 10.1016/j.cell.2007.03.053. [DOI] [PubMed] [Google Scholar]
- 42.Mazelova J, Astuto-Gribble L, Inoue H, Tam BM, Schonteich E, et al. Ciliary targeting motif VxPx directs assembly of a trafficking module through Arf4. Embo J. 2009;28:183–92. doi: 10.1038/emboj.2008.267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Leroux MR. Taking vesicular transport to the cilium. Cell. 2007;129:1041–3. doi: 10.1016/j.cell.2007.05.049. [DOI] [PubMed] [Google Scholar]
- 44.Follit JA, Tuft RA, Fogarty KE, Pazour GJ. The intraflagellar transport protein IFT20 is associated with the Golgi complex and is required for cilia assembly. Mol Biol Cell. 2006;17:3781–92. doi: 10.1091/mbc.E06-02-0133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Dwyer ND, Troemel ER, Sengupta P, Bargmann CI. Odorant receptor localization to olfactory cilia is mediated by ODR-4, a novel membrane-associated protein. Cell. 1998;93:455–66. doi: 10.1016/s0092-8674(00)81173-3. [DOI] [PubMed] [Google Scholar]
- 46.Bae YK, Kim E, L'Hernault S W, Barr MM. The CIL-1 PI 5-phosphatase localizes TRP Polycystins to cilia and activates sperm in C. elegans. Curr Biol. 2009;19:1599–607. doi: 10.1016/j.cub.2009.08.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gross CT, McGinnis W. DEAF-1, a novel protein that binds an essential region in a Deformed response element. Embo J. 1996;15:1961–70. [PMC free article] [PubMed] [Google Scholar]
- 48.Lutterbach B, Westendorf JJ, Linggi B, Patten A, Moniwa M, et al. ETO, a target of t(8;21) in acute leukemia, interacts with the N-CoR and mSin3 corepressors. Mol Cell Biol. 1998;18:7176–84. doi: 10.1128/mcb.18.12.7176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gottlieb PD, Pierce SA, Sims RJ, Yamagishi H, Weihe EK, et al. Bop encodes a muscle-restricted protein containing MYND and SET domains and is essential for cardiac differentiation and morphogenesis. Nat Genet. 2002;31:25–32. doi: 10.1038/ng866. [DOI] [PubMed] [Google Scholar]
- 50.Liu Y, Chen W, Gaudet J, Cheney MD, Roudaia L, et al. Structural basis for recognition of SMRT/N-CoR by the MYND domain and its contribution to AML1/ETO's activity. Cancer Cell. 2007;11:483–97. doi: 10.1016/j.ccr.2007.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Fukuyama T, Sueoka E, Sugio Y, Otsuka T, Niho Y, et al. MTG8 proto-oncoprotein interacts with the regulatory subunit of type II cyclic AMP-dependent protein kinase in lymphocytes. Oncogene. 2001;20:6225–32. doi: 10.1038/sj.onc.1204794. [DOI] [PubMed] [Google Scholar]
- 52.Terman JR, Kolodkin AL. Nervy links protein kinase a to plexin-mediated semaphorin repulsion. Science. 2004;303:1204–7. doi: 10.1126/science.1092121. [DOI] [PubMed] [Google Scholar]
- 53.Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Starich TA, Herman RK, Kari CK, Yeh WH, Schackwitz WS, et al. Mutations affecting the chemosensory neurons of Caenorhabditis elegans. Genetics. 1995;139:171–88. doi: 10.1093/genetics/139.1.171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Culotti JG, Russell RL. Osmotic avoidance defective mutants of the nematode Caenorhabditis elegans. Genetics. 1978;90:243–56. doi: 10.1093/genetics/90.2.243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Bargmann CI, Hartwieg E, Horvitz HR. Odorant-selective genes and neurons mediate olfaction in C. elegans. Cell. 1993;74:515–27. doi: 10.1016/0092-8674(93)80053-h. [DOI] [PubMed] [Google Scholar]
- 57.Signor D, Rose LS, Scholey JM. Analysis of the roles of kinesin and dynein motors in microtubule-based transport in the Caenorhabditis elegans nervous system. Methods. 2000;22:317–25. doi: 10.1006/meth.2000.1084. [DOI] [PubMed] [Google Scholar]
- 58.Yu D, Ponomarev A, Davis RL. Altered representation of the spatial code for odors after olfactory classical conditioning; memory trace formation by synaptic recruitment. Neuron. 2004;42:437–49. doi: 10.1016/s0896-6273(04)00217-x. [DOI] [PubMed] [Google Scholar]
- 59.Ou G, Blacque OE, Snow JJ, Leroux MR, Scholey JM. Functional coordination of intraflagellar transport motors. Nature. 2005;436:583–7. doi: 10.1038/nature03818. [DOI] [PubMed] [Google Scholar]
- 60.Mello CC, Kramer JM, Stinchcomb D, Ambros V. Efficient gene transfer in C.elegans: extrachromosomal maintenance and integration of transforming sequences. Embo J. 1991;10:3959–70. doi: 10.1002/j.1460-2075.1991.tb04966.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Wicks SR, Yeh RT, Gish WR, Waterston RH, Plasterk RH. Rapid gene mapping in Caenorhabditis elegans using a high density polymorphism map. Nat Genet. 2001;28:160–4. doi: 10.1038/88878. [DOI] [PubMed] [Google Scholar]
- 62.Zwaal RR, Broeks A, van Meurs J, Groenen JT, Plasterk RH. Target-selected gene inactivation in Caenorhabditis elegans by using a frozen transposon insertion mutant bank. Proc Natl Acad Sci U S A. 1993;90:7431–5. doi: 10.1073/pnas.90.16.7431. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The GCY-12 levels in the cilia were decreased in chb-3(eg2) and chb-3(qj2) mutants. chb-3 mutants were examined in a che-2(e1033) mutation background in order to suppress the Daf-c phenotype. Head region of L4 stage animals were shown. Anterior is to the right. The same extrachromosomal array (Ex[GCY-12::GFP]) was transffered by mating from line to line for comparison.
(8.43 MB EPS)
Decreased GCY-12 levels in AWC/AWB cilia were rescued by chb-3 expression in AWC/AWB neurons. Cilia (nose tips) and cell bodies are marked with circles and squares, respectively (Upper panels) A chb-3(eg52) animal carrying the Ex[odr-1p::GCY-12::GFP] extrachromosomal array. (Bottom panels) A chb-3(eg52) animal carrying two extrachromosomal arrays: Ex[odr-1p::GCY-12::GFP] and Ex[odr-1p::CHB-3::mRFP]. Ex[odr-1p::CHB-3::mRFP] directs the expression of mRFP-tagged, full-length CHB-3 in AWC and AWB neurons. CHB-3::mRFP is distributed throughout the cell, but is excluded from the nucleus. The Ex[odr-1p::GCY-12::GFP] array contains ttx-3p::mRFP as an injection marker, which was expressed in the AIY neuron (*). Scale bar represents 20 µm.
(4.90 MB EPS)
The expression patterns of gcy-12 promoter fusion with gfp in wild-type (N2) and chb-3(eg52) animals. The same extrachromosomal array (Ex[gcy-12p::GFP]) array was used for comparison. The anterior regions of adult animals are shown. No differences in the expression level and pattern between N2 and chb-3 are seen.
(2.09 MB EPS)
In chb-3(eg52) animals, ODR-10 GPCR normally localizes to cilia of AWA sensory neuron (top panel) and AWB sensory neuron (bottom panel). For the localizaiton patterns in wild-type animals, see Dwyer et al., 1998 [45]. GFP patterns also show normal cilium morpholgy of these sensory neurons in chb-3(eg52).
(1.52 MB EPS)
Kymographs depicting GCY-12::GFP particle motility along the dendrite of a phasmid sensory neuron in chb-3(eg52) (Upper panels) and chb-3(eg52) rescued (Lower panels) animals are shown. The horizontal and vertical axes represent distance and time, respectively. Corresponding lines, with which the velocity and the frequency were analyzed, are also shown. Movies are available upon requests.
(1.57 MB EPS)
The rate of dauer formation at 20°C. The Daf-c phenotype of chb-3(eg52) is suppressed by unc-101(m1). Error bars indicate s.e.m.
(0.32 MB EPS)