Summary
Lymphopoiesis generates mature B, T, and NK lymphocytes from hematopoietic stem cells via a series of increasingly restricted developmental intermediates. The transcriptional networks that regulate these fate choices are composed of both common and lineage-specific components, which combine to create a cellular context that informs the developmental response to external signals. E proteins are an important factor during lymphopoiesis, and E2A in particular is required for normal T and B-cell development. Although the other E proteins, HEB and E2-2, are expressed during lymphopoiesis and can compensate for some of E2A's activity, E2A proteins have non-redundant functions during early T-cell development and at multiple checkpoints throughout B lymphopoiesis. More recently, a role for E2A has been demonstrated in the generation of lymphoid-primed multipotent progenitors and has shown to favor their specification towards lymphoid over myeloid lineages. This review summarizes both our current understanding of the wide-ranging functions of E proteins during the development of adaptive lymphocytes and the novel functions of E2A in orchestrating a lymphoid-biased cellular context in early multipotent progenitors.
Keywords: hematopoietic progenitor cells, lineage commitment, transcription factors, cell differentiation, gene regulation
Introduction
Lymphopoiesis is the process by which B and T lymphocytes as well as natural killer (NK) cells, lymphoid tissue inducing (LTi) cells, and possibly subsets of dendritic cells (DCs) are generated from hematopoietic stem cells (HSCs). The gene expression programs that characterize a given fate are driven by specific combinations of lineage-restricted and ubiquitous transcription factors. Some of these lineage-specifying factors function solely to induce lineage associated genes, whereas others commit cells to a lineage by prevent activation of genes associated with alternative fates. Recent findings also suggest that down-regulation of stem cell-associated transcription factors just prior to lineage commitment may extinguish the ability of ubiquitous or common transcriptional activators to access genes for alternative lineages (1, 2). Our understanding of how transcriptional regulatory networks stabilize lineage-associated gene programs and how specific combinations of transcription factors within these networks synergize or antagonize each others' activities is evolving. The E protein transcription factors are key components of the transcriptional networks that promote specification and commitment to the adaptive T and B-lymphoid fates, and they also directly regulate essential features of these lineages such as the rearrangement and expression of the immunoglobulin (Ig) and T-cell receptor (TCR) loci. E proteins are believed to antagonize innate lymphocyte fates, particularly NK cells and LTi cells, and the Id inhibitors of E proteins are necessary for the development of these lineages (3, 4). The mechanisms by which E proteins antagonize innate lymphocyte development remain controversial but, in analogy with their role in adaptive lymphocytes, may involve the induction of transcriptional networks inappropriate for innate lineages. While E proteins regulate transcriptional networks in adaptive lymphocytes, we propose that at earlier stages of lymphopoiesis, in the common progenitors of both adaptive and innate lymphocytes, E proteins play a different role: collaborating with other factors to establish the context for specification and commitment to lymphoid fates (Fig. 1). In this review, we examine both the evidence supporting this hypothesis and the functions of E proteins in later adaptive lymphocyte development.
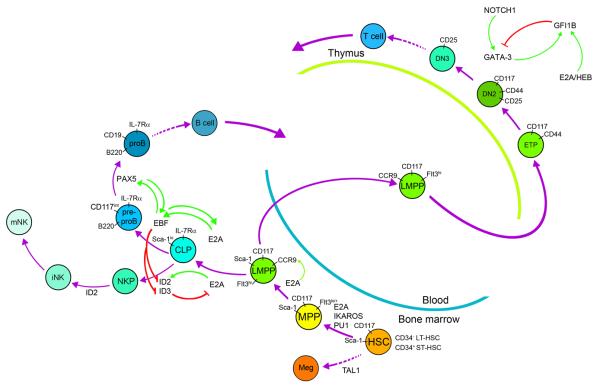
Fig. 1. Mechanism of lymphocyte specification and commitment.
E2A collaborates with multiple factors to promote lymphoid priming, which is the low abundance expression of multiple lymphocyte-associated genes. The complement of genes induced in individual LMPPs may differ depending on the concentration of each of the transcriptional regulators leading to a heterogenous population. Heterogeneous lymphoid priming leads to the expression of multiple receptors and signaling molecules that allows variation in the response to extrinsic signals. These signaling pathways (IL-7R or Notch signaling) can induce transcription factors that collaborate with E proteins to induce novel transcriptional regulatory networks required for B or T lymphopoiesis. Commitment to the adaptive or innate lymphoid pathways involves activation of repression of essential regulators of the alterative fate.
E proteins and their interacting partners
E proteins
In mammals, there are 4 class I helix-loop-helix proteins (HLH), or E proteins, encoded by 3 genes: E2A (Tcf3), Heb (Tcf12), and E2-2 (Tcf4) (reviewed in 5). The structure of E proteins has been reviewed extensively, and primary references can be found in the review by Massari and Murre (5). E proteins are orthologues of the Drosophila protein DAUGHTERLESS, which functions in a wide range of cell fate decisions in flies, including sex determination, selection of sensory organ progenitors, and selection of the R7 photoreceptor. The first mammalian E proteins to be identified, E12 and E47, were cloned on the basis of their ability to bind an ‘E box’ sequence (κE2) in the immunoglobulin κ (Igκ) light chain enhancer. E12 and E47 differ only in their bHLH domain and derive via alternate exon usage from the E2A gene. HEB and E2-2 were cloned shortly thereafter based on their ability to bind to similar E box sequences. The E proteins bind DNA as a dimer and protein-protein interactions are mediated by the HLH domain. Immediately upstream of the HLH domain is a basic region that is required for DNA binding and may play a role in nuclear localization. All of the E proteins share a high degree of sequence similarity within their bHLH domains as well as in two upstream transcriptional activation domains (ADs). In the E2A proteins, AD1 and AD2 collaborate in activating transcription. However, if both domains are deleted, the remaining N-terminal portion retains some transcriptional activation activity (6). E12 and E47 are primarily transcription activators that function, in part, by recruiting the co-activator protein p300/CBP, which in turn recruits histone acetyltransferases and RNA polymerase II to the promoter or enhancers of target genes (5, 7, 8). However, there is evidence that E2A proteins also repress some genes in pre-B cells, although the mechanism underlying this repression remains to be resolved (9). Interestingly, while E47 readily forms homodimers, an inhibitory domain in E12 may restrict its function to heterodimers (10). Since E47 lacks this inhibitory domain, the proportion of E47 homo- to heterodimers is most likely solely dependent on the quantity of potential binding partners, although the redox state may also alter dimerization and DNA-binding potential under physiological conditions (11). In keeping with this, in B-lymphocyte cell lines, where E47 protein is expressed at higher concentrations than the other E proteins, E47 homodimers are the dominant E box-binding complexes detected by electrophoretic mobility shift assays (EMSA) (5). In contrast, in T lymphocytes where HEB and E47 are both highly expressed, heterodimers bind to the E-box sequence (12, 13).
Alternative splice variants have not been reported for E2-2 and HEB, although northern blot analysis revealed multiple mRNA species reactive with Heb cDNA. However, in contrast to the E2A gene, there is evidence that alternative promoter usage generates two forms of the HEB protein, which were reported to have different functions (14). The shorter form of HEB has only one activation domain and, by analogy with E2A, would be predicted to function in transcriptional activation (6). However, this protein also contains a novel sequence that is not found in other E proteins, termed an Alt domain (14). While it is tempting to speculate that Alt confers a novel function on HEB, such as transcriptional repression, this remains to be formally investigated. An alternative promoter for the E2-2 protein, resulting in a form of E2-2 that lacks AD1 but has an Alt domain, is also predicted based on DNA sequence comparison but thus far has not been ascribed functional significance (15).
Tissue-specific bHLH proteins
As E proteins are broadly expressed and bind a common E box element, one aspect of their function that is not completely understood is how they mediate so many different developmental outcomes. One means by which specificity can be lent to E proteins is via dimerization with the more tissue-restricted class II bHLH proteins, which, in the context of hematopoiesis, refer to the highly-related stem cell leukemia (SCL)/T-lymphocyte acute lymphoblastic leukemia 1 (TAL1), and lymphoblastic leukemia 1 (LYL1) proteins. Both TAL1 and LYL1 were cloned from translocations associated with T-cell acute lymphoblastic leukemia (T-ALL) and, like the E proteins, possess an HLH domain and adjacent basic region (reviewed in 16). Class II HLH proteins homodimerize poorly and generally require heterodimerization with E proteins to allow DNA binding (5). Binding site selection assays reveal that the E-box sequence preferred by TAL1-E protein heterodimers differs from those preferred by E protein homodimers (17, 18). This alteration of E protein DNA binding specificity may explain the ability of TAL1 and LYL1 to interfere with E2A-dependent transcription of target genes and provides one mechanism by which TAL1 or LYL1 over expression might contribute to T-cell leukemogenesis (reviewed in 19).
Id proteins
Class IV HLH proteins are comprised of the mammalian Inhibitor of DNA binding (ID1-4) proteins and Drosophila EXTRAMACROCHAETAE (emc), which lack the basic domain required for DNA-binding but retain the HLH domain and the ability to dimerize with E proteins (5). Id proteins inhibit E protein activity by directly preventing their DNA-binding but do not interact appreciably with the hematopoietic class II HLH proteins (20-22). Despite some evidence that ID2 can interact with the retinoblastoma (RB) tumor suppressor and that ID2 and ID3 interact with Ets family members, the most well characterized role of ID proteins is in the regulation of E protein activity (23).
E proteins and hematopoietic stem cells
Like all hematopoietic cells, lymphocytes are derived from hematopoietic stem cells (HSCs) that reside in the bone marrow within the lineage marker (Lin)−Sca-1+CD117 (Kit)+ (LSK) population that also contains the CD34− self-renewing long-term HSCs (LT-HSCs) and their short-term reconstituting CD34+ progeny (ST-HSCs) (24). For many immunologists, E2A is thought of as a specific factor for B-cell development and mice lacking E2A do have a dramatic defect in B lymphopoiesis (25, 26). However, a role for E proteins, particularly E2A, at earlier stages of hematopoiesis has not been thoroughly investigated, even though E protein interacting proteins are essential for HSC formation and function. Specifically, the class II HLH proteins TAL1 and LYL1 have well characterized functions in HSCs, implicating E proteins as their obligate dimerization partners. However, it remains to be determined whether a specific E protein is critical for any TAL1 or LYL1 functions and how this specificity arises.
Tal1-deficient embryos die by day 10.5 of gestation, due to a failure of primitive hematopoiesis (27, 28). Blastocyst chimeras confirm that Tal1−/− cells are also severely impaired in definitive hematopoiesis but can contribute to non-hematopoietic tissues. Although critical during the development of HSCs from mesoderm, following hematopoietic specification Tal1 is largely dispensable for the maintenance and function of definitive HSCs but remains essential for erythroid and megakaryocyte lineages (reviewed in 29). Interestingly, primitive hematopoiesis can be rescued with a wide range of TAL1 mutants, even those lacking the basic domain and thus DNA-binding capacity, so long as their dimerization function is intact (30). While these findings may be interpreted as evidence that sequestering E proteins is the primary function of TAL1 during primitive hematopoiesis, there is evidence that TAL1-E protein heterodimers can be recruited to DNA through interactions with other transcription factors such as Gata1 (31). In contrast, the basic region of TAL1 is required for definitive hematopoiesis as well as for development of mature erythrocytes, myelocytes, and megakaryocytes, clearly implicating TAL1-mediated transcription in these processes (30).
In contrast to Tal1−/− mice, Lyl1−/− mice are viable but have reduced numbers of the more primitive LSKCD34− HSCs and a defect in the generation of B cells, while myeloid lineages are largely spared (32). Lyl1 expression during embryonic development, as determined by a LacZ knock-in model, largely overlaps with that of Tal1 (33). Despite the high degree of similarity and overlapping expression patterns, endogenous LYL1 does not rescue Tal1−/− embryos from lethality (27, 28), suggesting that LYL1 protein may not be present during primitive hematopoiesis. It was recently shown that LYL1 and TAL1 do compensate for each other to some extent in adult HSCs, since mice deficient in both Tal1 and Lyl1 show a more severe HSC deficiency in that their HSCs fail to repopulate lethally-irradiated recipients and are highly apoptotic (34).
In contrast to the severe consequences to hematopoiesis of deleting Tal1 or Lyl1, deletion of E2A results in more subtle alterations of the HSC compartment. In the absence of E2A, HSCs and their downstream progeny within the LSK compartment show increased bromodeoxyuridine (BrdU) incorporation, suggesting that there are more cells in cycle than in wildtype (WT) controls (35-37). This increased proliferative capacity was linked to decreased expression of mRNA encoding the cell cycle inhibitor p21 (35, 36). Competitive bone marrow transplantation assays revealed that E2A−/− HSCs are compromised slightly in their ability to undergo self-renewal (36). In addition, under stress conditions, E2A−/− HSCs become exhausted and fail to sustain hematopoiesis (35). However, under steady-state conditions the requirement for E2A appears to be minimal, suggesting either that HEB or E2-2 are more important for HSCs, or that all E proteins function redundantly within HSCs.
E proteins and development of lymphoid-primed multipotent progenitors
Lymphoid-primed multipotent progenitors (LMPPs) represent the first lineage restriction point along the route from HSC to committed lymphocyte. LMPPs are a subset of LSKCD34+ cells that have lost self-renewal capacity, and most megakaryocyte/erythrocyte (MegE) differentiation potential, but retain the ability to give rise to all lymphocyte subsets as well as granulocyte/macrophage progenitors (GMPs) (38, 39). LMPPs can give rise to common lymphoid progenitors (CLPs) (40), which produce B lymphocytes and NK cells in the bone marrow and are likely among the bone marrow precursors that function as thymus seeding progenitors (TSPs) (41, 42). A growing list of markers demonstrates the heterogeneity of the LMPP population. In particular, a subset of cells expresses the chemokine receptor CCR9 (43, 44), which is required for efficient homing of TSPs to the thymus (45). A partially overlapping subset of LMPPs has low expression of vascular cell adhesion molecule-1 (VCAM-1)/CD106 (44, 46), which characterizes more primitive LSKs that retain MegE potential. In addition, a small population of LMPPs express GFP from a Rag1 reporter gene, indicating that they overlap with early lymphoid progenitors (ELPs), which are more lymphoid-biased than the bulk LMPP population (47). LMPPs with a CD90−CD62L+ phenotype show heightened T-cell differentiation potential in vitro and in vivo (48-50). While this heterogeneity may be a consequence of a developmental hierarchy within the LMPP subset an alternative interpretation is that the LMPP population contains cells with distinct gene expression profiles that may reflect differential biases in their developmental potential.
The two critical attributes that distinguish LMPPs from MPPs are (i) the loss of MegE potential and (ii) the initiation of ‘lymphoid priming’, a process in which genes associated with multiple distinct lymphoid lineages are co-expressed at low abundance (39, 51). E2A is necessary for the development of LMPPs and proper lymphoid priming, but it is not required to extinguish MegE potential (37). Instead, the loss of MegE potential coincides with the loss of thrombopoietic receptor Mpl expression and may also be associated with restricted expression of Gata1, a transcription factor essential for both megakaryocyte and erythroid differentiation (52, 53). Similarly, decreased expression of TAL1 in LMPPs may decrease MegE potential and liberate E proteins to function in homodimeric complexes that can contribute to lymphoid gene expression. Consistent with that hypothesis, the expression of E47 protein does not change during differentiation of HSCs into LMPPs, indicating that the essential function of E2A in lymphoid priming does not require increased E2A protein expression (37).
While E2A is not required for the restriction of MegE potential, it does play a role in the myeloid versus lymphoid decision of LMPPs. The few LMPPs present in E2A−/− mice show enhanced myeloid cloning efficiency compared to WT LMPPs, and E2A−/− MPPs are also skewed toward macrophage differentiation (37). This finding is consistent with a study by Cochrane and colleagues in which inhibition of E protein function via ectopic expression of ID1 favored myelopoiesis at the expense of B cell development (54). However, while ID1 expression is associated with myeloid rather than lymphoid differentiation, Id1−/−LMPPs did not show a significant alteration in lineage potential. Therefore, these studies confirm that E proteins promote lymphopoiesis from LMPPs at the expense of myelopoiesis, but indicate that Id1 is unlikely to be the major physiological determinant of E protein function at this stage of development. Interestingly, progenitors from mice transgenic for an inducible version of a hybrid E47, containing TAL1's ID-impervious bHLH domain, preferentially gave rise to B lymphocytes over myeloid cells (54). This finding was interpreted to support the hypothesis that E proteins promote lymphoid differentiation from LMPPs but raises questions about the DNA binding specificity of the E protein complexes involved since this hybrid E47 should bind DNA sequences preferred by TAL1-E protein heterodimers. Nonetheless, most evidence indicates that E2A functions within LMPPs to promote lymphoid rather than myeloid differentiation. Indeed, the fact that MPPs from E2A−/− bone marrow are also skewed toward myeloid differentiation raises the possibility that the failure to accumulate E2A−/−LMPPs is a consequence of the propensity of these cells to precociously adopt the myeloid fate without forming a stable LMPP intermediate.
In addition to their increased propensity to give rise to myeloid cells, E2A−/− LMPPs demonstrated a striking dysregulation of lymphoid-associated genes that includes both known E2A targets such as Rag1, Il7ra, Dntt, Igh-6, and Notch1, and genes whose regulation was not previously shown to involve E2A, such as Ccr9 and Ets1 (37). Surprisingly however, genes encoding transcription factors known to promote lymphopoiesis early in hematopoiesis, such as Sfpi1 (PU.1), Ikzf1 (IKAROS), and Gfi1 were not dependent on E2A for their expression. Analysis of the DNA sequences flanking the transcriptional start site for E2A-dependent genes revealed an enrichment of binding sites for IKAROS and PU.1, suggesting that while E2A does not regulate their expression, it collaborates with these factors to initiate lymphoid priming (37). These observations suggest that, unlike commitment to the B-lymphocyte fate, the emergence and function of LMPPs may not depend on E2A to induce an entirely novel transcriptional regulatory network. Rather, transcription factors already expressed in LSKs and MPPs may cooperate with E2A to drive lymphoid priming and further differentiation to the CLP stage (Fig. 1). Lymphoid priming allows cells within the LMPP and CLP compartment to express a variety of cell surface receptors, including IL7R, NOTCH1, CCR9 (and likely CCR7), and CD69L, which endows them with the ability to respond to their microenvironment, and influences the set of genes regulated by the basal cast of transcriptional regulators inherited from earlier stages. Indeed, individual CLPs or ETPs may have different probabilities of responding to an environmental cue with a given outcome based on small fluctuations in transcription factor expression that may become amplified during their development from LMPPs.
While it is likely that E2A cooperates with Ikaros and PU.1 to promote lymphoid priming, E2A-deficiency has more severe consequences for the development of LMPPs than deficiency of either of the other proteins. As both IKAROS and PU.1 control the expression of Flt3, it is difficult to unambiguously identify LMPPs in Sfpi1−/− or Ikzf1−/− mice (55-57), but Yoshida et al. (56) cleverly overcame this problem, using an Ikaros promoter-GFP reporter mouse that displays increased GFP expression concomitant with the development of LMPPs. When crossed to mice carrying a null allele of Ikaros, GFPhi LSKs are clearly present and these cells have appropriate myeloid potential without generating megakaryocytes, indicating that LMPPs are present but lack expression of FLT3. However, these Ikaros−/− LMPPs show altered lymphoid gene expression and fail to properly generate B or T lymphocytes in vitro. The set of IKAROS dependent genes in LMPPs shows significant but not complete overlap with those regulated by E2A, consistent with these two transcription factors collaborating in gene regulation (58). GFI1 has also been implicated in the generation of LMPPs and appears to function in part by limiting the expression of PU.1, which is both essential for lymphoid differentiation and can block it if expressed at high concentrations (59-61). Therefore, GFI1 may be necessary to allow appropriate cooperative interactions between E2A, IKAROS, and PU.1 to promote lymphoid priming, while limiting PU.1-driven myelopoiesis. Our current understanding of how extracellular signals and the expressed cohort of transcription factors, in collaboration or antagonism with E2A, lead to the transcriptional regulatory networks that dictate specification and commitment to a given lymphocyte fate is the focus of the remaining sections of this review.
E proteins and B-cell development
Specification of the B-cell fate
Specification refers to the process of turning on a lineage-associated gene expression program and, for the B-cell fate, begins in the CLP population. The critical event driving CLPs toward the B-cell fate is the induction of the B-cell-associated transcription factor EBF1, whose mRNA and target gene expression have been detected in a subset of CLPs (62-64). E12 is able to induce Ebf1 expression in a macrophage cell line (65), and E2A binding sites have been identified in the Ebf1α promoter, which is active at the CLP stage (66, 67). Therefore, E2A proteins appear to be regulators of the initial expression of the Ebf1 gene. Moreover, the requirement for E2A in early B-cell development can be bypassed by forced expression of EBF1 in E2A−/− progenitors, which rescues the expression of B-cell-associated genes and IgH V(D)J recombination (68, 69). However, these data should not be interpreted as evidence that E proteins are superfluous in pro-B cells after the induction of EBF1. As discussed below and reviewed elsewhere, E proteins play many critical roles during B-cell development; however, the E protein activity provided by HEB and E2-2 is sufficient for survival and expression of most E protein target genes in pro-B cells, provided that EBF1 is expressed appropriately.
Ebf1 is not among the lymphoid genes whose expression is primed by E2A in LMPPs, rather, the induction of Ebf1 occurs in a subset of CLPs that also show increased expression of E2A protein compared to the bulk of CLPs and LMPPs (37, 63, 64). This increase in E2A protein levels is dependent on EBF1, suggesting that it is a consequence of and not the cause of EBF1 induction (71). This finding raises the question of why Ebf1 induction is delayed until the CLP stage if E2A protein levels remain constant from the HSC compartment. The answer likely lies in the requirement for a change in cellular context that occurs between the LMPP and CLP stages, which is, in turn, dependent on signaling through the IL7R.
IL7R surface expression is not observed until the CLP stage even though Il7ra is among the lymphoid genes transcribed in LMPPs (51, 72). IL7R signaling has been implicated in the expression of Ebf1 in CLPs, although the mechanism responsible remains controversial (73-76). One possible scenario is that the chromatin context at the Ebf1 gene is altered as a consequence of IL7R signals, allowing E2A and other regulators of Ebf1 expression, such as PU.1, to gain access and promote its transcription. Such a change in chromatin context is likely mediated by the STAT5 transcription factors, which may be activated upon expression of IL7R at the CLP stage, even before the initial transcription of the Ebf1 gene is detected. This hypothesis is supported by the observation that Ebf1 expression is extremely low in the CLPs of Il7−/− and Il7ra−/− mice but can be restored by retroviral expression of a constitutively active STAT5 (74, 75). However, Ebf1 is expressed in the pro-B cells of mice in which the Stat5a and Stat5b genes were deleted using Rag1-Cre, suggesting that if STAT5 opens the Ebf1 gene locus, it must act prior to the function of Rag1-Cre and via a hit-and-run mechanism (77). Alternative hypotheses can also explain these data; for example, in mature T lymphocytes, deletion of Stat5a and Stat5b allows the compensatory use of alternate STAT transcription factors by IL7R, such as STAT6 (78). Thus, the connection between IL7R signaling and histone modifications at the Ebf1 gene, and the role of Stat5, remain poorly defined. In embryonic stem cells and Lin− bone marrow cells, the Ebf1 gene is in a poised state containing histone modifications associated with both gene activation (H3K4me3) and repression (H3K27me3) (79). H3K27me3 is maintained by the polycomb repressor complex 1 (PRC1), and deletion of Bmi-1, encoding a component of PRC1, in hematopoietic cells, leads to the loss of H3K27me3 at the Ebf1 promoters and premature Ebf1 mRNA expression (79). Therefore, the regulation of PRC1 function may alter the context of the Ebf1 gene to allow for its E2A-dependent induction at the CLP stage.
Commitment to the B-cell fate
Commitment to a particular lineage requires repressing lineage inappropriate genes to prevent differentiation down alternative pathways. The first cells fully committed to the B-cell lineage express both B220 and CD19 on their surface, along with CD43, HSA and the receptor c-kit (CD117). These cells are absent in E2A−/− and Ebf1−/− mice and most B-cell associated genes and IgH rearrangements are undetectable in the bone marrow or fetal liver of these mice (26, 71, 80). Mice heterozygous for the E2A and Ebf1 mutations show a significant reduction of committed pro-B cells (defined as B220+CD43+) and have reduced expression of many B-cell genes (81). Among the genes regulated by E2A and EBF1 is Pax5, which encodes a transcription factor that is essential for development beyond the pro-B-cell stage. Although pro-B cells develop in Pax-5−/− mice, there is substantial evidence that they are not committed to the B-cell lineage (82, 83). For instance, Pax-5−/− pro-B lymphocytes expanded in vitro in IL-7 express many lineage inappropriate genes, which can be repressed by ectopic expression of Pax-5 (84). Remarkably, Pax5−/− pro-B cells retain the ability to differentiate into alternative lineages, including myeloid cells, when IL-7 is removed from the culture (82). Moreover, these IL-7-expanded Pax-5−/− pro-B cells, and to some degree primary Pax-5−/− pro-B cells, can function as HSCs and can reconstitute all hematopoietic lineages when transplanted into lethally irradiated mice. The requirement for Pax-5 to maintain commitment to the B-cell lineage persists throughout B-cell development, since even mature B cells lose their B-cell characteristics and express lineage inappropriate genes following Pax-5 deletion (85). These mature cells can regress to a more immature developmental state, and even switch to the T-cell lineage, but most frequently seed the development of B-cell neoplasms. Taken together, these studies clearly demonstrate the essential role of Pax5 in B-cell lineage commitment and E2A as an essential regulator of Pax5 expression.
It has only recently been appreciated that Pax-5 does not act alone in repressing alternative lineage genes in B lymphocytes. Pax-5, like E2A, is involved in regulating the expression of Ebf1 via a second Ebf1 promoter, designated Ebf1β. Therefore, in the absence of PAX5, Ebf1 is only transcribed from the Ebf1α promoter and does not reach the concentrations found in WT pro-B cells (67). EBF1, in turn, has been implicated in the repression of non-B-lineage genes (86, 87), although a direct test of Ebf1's role in maintaining B-cell commitment awaits the development of a conditional allele for Ebf1 that allows its deletion after B-cell commitment. At the present time, studies are limited to an analysis of EBF1 function in cells or cell lines prior to B-cell lineage commitment (57, 87). Nonetheless, EBF1 is capable of repressing the Id2 and Id3 genes (86). This observation is particularly intriguing, because E2A proteins have been implicated in the induction of Id2 as well as Ebf1 (37, 88). Therefore, when E2A protein levels increase in CLPs both EBF1 and ID2 should increase, but these proteins have antagonistic effects on E2A function and B-cell development. The ability of EBF1 to repress Id2 and Id3 suggests that EBF1 augments E2A function through both positive feedback and repression of E protein inhibitors.
No other targets of EBF1 repression have been identified, and the mechanism of repression remains to be determined. Genome-wide association studies revealed that EBF1 does bind to genes that are repressed during B-cell development, but the relevance of most of these genes remains to be determined (89). In contrast, PAX5 repressed genes include essential regulators of other hematopoietic fates, such as the T-cell-associated genes Notch1, Ptcra, and the macrophage colony stimulating factor receptor gene Csfr1 (84). Nonetheless, genome-wide association assays confirmed that EBF1 binding at cis-regulatory sequences is frequently associated with E-box binding sites (in 13.5% of cases), consistent with the fact that EBF1 and E2A proteins cooperatively regulate multiple B-cell-associated genes (89). In addition, genome-wide binding data for E2A, comparing Ebf1-deficient and sufficient cells, suggest that EBF1 may alter the chromatin context at cis-regulatory sites to allow binding by E2A, since the number of E2A bound sequences was nearly 3-fold higher in Ebf1-sufficient cells (90). EBF1 was shown to bind to some of its B-cell targets in a T-cell line and promote H3K4 dimethylation, a modification which may facilitate binding of E2A or other regulators of B-cell genes (89).
E proteins in pro-B lymphocytes
E47 homodimers are the major E-box binding proteins in pro-B lymphocytes (5). However, it is the dose of E proteins rather than the specific E protein that appears to be important for B-cell development. This conclusion is based on an analysis of the B-cell phenotype in mice carrying combined mutations in the E2A, Heb, or E2-2 genes (91). Deletion of Heb or E2-2 has a minor effect on B-cell development, but when combined with heterozygosity of E2A, their deletion results in a more striking effect on B-cell generation. Similarly, the dose of mutant forms of E12 required to rescue B-cell development from E2A−/− MPPs is proportional to the transcriptional activation capacity of the mutants (6). The notion that the dose of E proteins is more critical than the identity of the E protein is also supported by experiments in which HEB was able to promote B-cell development when expressed in place of E2A from the E2A gene (70). Our data support the notion that high concentrations of E proteins are required for the initial induction of Ebf1 but that lower levels are sufficient after EBF1 is expressed (68). This finding is likely a reflection of the multiple changes that occur in cells after expression of EBF1 and the fact that EBF1, E2A and PAX5, as well as other factors such as ETS1 and FOXO1, function cooperatively to regulate a large portion of the B-cell transcriptome (90). Lower concentrations of E protein may be sufficient for these cooperative interactions, whereas higher concentrations of E proteins may be necessary for the initial induction of Ebf1. The altered chromatin context and cooperative interactions may explain why deletion of E2A leads to only a modest decline in Ebf1 in pro-B cells or Abelson transformed B-cell lines (69, 92).
In addition to its role in initiating B-cell lineage specification and commitment, E2A coordinates these differentiation events with IL-7-induced proliferation. In the absence of E2A, in the case where EBF1 is expressed ectopically in E2A−/− MPPs, HEB, and E2-2 are sufficient to maintain pro-B-cell survival but do not allow expansion in response to IL-7 (68). This deficit is linked to an inability to up-regulate the IL7R signaling dependent gene N-myc. E2A is not required in pro-B cells for Il7ra expression, surface IL7R, or initial IL7R signalling events, because STAT5 activation is observed in E2A−/− EBF1-expressing pro-B cells. Therefore, E2A is likely directly involved in regulating the N-myc gene. Inhibition of all E protein activity via ectopic expression of ID3 has much more severe consequences than deleting E2A in pro-B cells, inducing catastrophic growth arrest and caspase-2 dependent apoptosis (69, 92, 93). There are multiple mRNA changes upon ID3 expression that contribute to grow arrest, including decreased expression of N-myc and c-myc and decreased expression of genes encoding proteins involved in ribosome function (93). Therefore, E proteins are essential for the viability of pro-B cells and link their survival to the differentiation program.
E2A proteins were identified as regulators of Ig enhancers, and E proteins are essential for IgH expression in pro-B cells, although E2A itself is not absolutely required because E2A−/− pro-B cells rescued with EBF1 undergo VH(DH)JH rearrangement and express IgH at WT levels (68). In addition, IgH is among the E2A-dependent genes primed in LMPPs (37). Nonetheless, EBF1 and PAX5 clearly have important roles in controlling IgH expression and rearrangement. Rag1 is the gene most affected in pro-B cells expressing ID3, indicating that E protein activity is critical for V(D)J recombination (93). In pre-B cells, E2A is indispensable for Igκ light chain expression and rearrangement (94). Removal of the E2A binding sites in the Igκ intronic enhancer impairs recombination (68, 95) and expression of ID3 or deletion of E2A in pro-B cells or Abelson pre-B-cell lines severely impairs Igκ gene transcription (93). Expression of either E12 or E47, in combination with the Rag genes, is sufficient to induce Igκ locus recombination in non-lymphoid cells indicating that E proteins are sufficient to promote transcription of Igk in this context (96, 97). In pro-B cells, IL7R signaling is known to prevent the activation and rearrangement of Igk and this has been shown to be mediated by STAT5, through its ability to impede the binding of E2A to the Igκ intronic enhancer (98). These data demonstrate that E2A proteins are major regulators of Igκ expression and rearrangement.
E proteins and T-cell development
In contrast to B lymphocytes, T precursors require the thymic environment to develop. In the adult, blood-borne TSPs enter the thymus at the cortical-medullary junction (CMJ), persist for about 10 days, then migrate outwards to the subcapsular zone and back to the medulla over the course of another 5 days (99). The most immature thymocytes lack mature lineage markers and both CD4 and CD8 major histocompatibility complex (MHC) coreceptors and are called ‘double negative’ (DN) cells (100). DN cells are further subdivided by expression of the adhesion molecule CD44, CD117, and CD25, the α chain of the interleukin-2 (IL-2) receptor. The immediate progeny of TSPs are CD44+CD117+CD25− early thymic progenitors (ETPs), which also retain myeloid, NK, and DC potential. ETPs are heterogeneous, with a CCR9+ and Flt3+ subset that resembles the putative TSP and posses more B-cell potential and greater proliferative capacity than the CCR9− and Flt3− subset (43, 101). DN2 stage cells, characterized by expression of CD25 and CD117, lack B-cell potential but can produce NK and DCs (2). The last non-T potentials are lost in the transition to the CD117−CD44−CD25+ DN3 stage during which thymocytes become further specified to the T-cell lineage as evidenced by expression of Ptcra, Rag1, and Rag2 (2). The RAGs, in conjunction with constitutively-expressed DNA repair machinery, rearrange the Tcrb, Tcrg, and Tcrd loci in DN2/3 cells, allowing for further development based on the production of a functional pre-TCR containing preTα and TCRβ or a TCRγδ receptor.
E proteins and T-lymphocyte specification
Multiple transcription factors have been implicated in specification of the T-lymphocyte gene expression program, including NOTCH1, RUNX1, GATA3, IKAROS, GFI1, LEF1/TCF1, and E2A. E2A proteins are required for normal thymocyte development. In their absence, the thymus is about 20% smaller than in a WT mouse and the mice develop an aggressive T-cell lymphoma by 3-6 months of age (13, 102). E2A was reported to be up-regulated in DN2 cells in parallel with the inception of T-lymphocyte specification and commitment (103, 104). However, these early studies examined E2A expression in DN thymocytes without depleting Lin+ cells, and therefore the DN1 population was heterogeneous and not highly enriched for ETPs. Using new staining strategies that allow a better resolution of the ETP and DN2 subsets, we found that E2A is expressed at similar levels in ETPs, DN2 and DN3 cells and is up-regulated in a subset of DN4 cells (Fig. 2). These findings are largely consistent with recent mRNA expression analysis for E2A in Lin− thymocytes (2). In addition, we found that E2A−/− mice have severely reduced numbers of both ETPs and DN2 cells (Xu and Kee, unpublished data), a finding that is consistent with the reduced number of LMPPs and CLPs in E2A−/− mice, and their failure to express the thymus-homing chemokine receptor CCR9 (37). Flt3l−/− mice, which also have a reduced number of LMPPs, and Ccr9−/− mice, have similar defects in the generation of ETPs and DN2 cells, indicating that LMPP/CLP numbers and homing potential directly determine ETP and DN2 cell numbers (24, 42).
Fig. 2. E47 is expressed at similar levels in all DN thymocyte subsets.
(A) Total thymocytes from C57Bl/6 mice were stained with a lineage cocktail including antibodies to CD8, CD3, TCRβ, TCRγ, CD11b, CD11c, Gr1, NK1.1, and B220 as well as CD44, CD25 and E47 or control IgG. CD44 and CD25 expression are shown on Lineage negative thymocytes. B) Histogram showing E47 staining in ETPs (Lin−CD44+CD35−), DN2s (Lin−CD44+CD25+), DN3s (Lin−CD44−CD25+), and DN4s (Lin−CD44−CD25−). Isotype control staining is shown in gray.
In contrast to E2A protein levels, HEB and HEBALT are upregulated at the DN2 and DN3 stages and this induction may be uniquely dependent on signals derived from the thymus microenvironment (2, 14). In extracts of total thymocytes, E-box binding complexes are composed predominantly of an E2A:HEB heterodimer (12, 13). The incomplete arrest in T-cell development in E2A−/− mice has been attributed to partial redundancy with HEB, and this conclusion is consistent with the observation that targeting a DNA-binding defective mutant of HEB, that can dimerize with and inhibit E2A, into the Heb locus, causes a severe reduction in the total number of thymocytes (12). However, in contrast to E2A−/− thymocytes, Heb−/− thymocyte numbers are not reduced until after the DN stage of development, when immature CD8+ cells normally undergo proliferation (105), indicating that HEB is not absolutely necessary for differentiation through the DN stages of T-cell development. HEB deficiency also results in thymocyte numbers that are 10%-20% of WT and a loss in the distinction between the thymic cortex and the medulla (105). DN thymocytes and γδ T-cell numbers are unaffected by the absence of HEB, in keeping with a defect that arises after the DN3 stage (105). There is also evidence that E2-2 contributes to T-cell development, and in mice expressing ID1 as an Lck-driven transgene there is massive apoptosis of thymocytes that is more severe than in the absence of either E2A or HEB (106, 107). Taken together, these findings demonstrate that E proteins, primarily E2A homodimers and E2A-HEB heterodimers, play essential roles in T-cell development during lineage specification and commitment. However, the regulatory networks in which E2A functions remain to be fully revealed.
E2A and Notch1
One of the critical specification and commitment factors for early T-cell development is NOTCH1, a transmembrane receptor that also functions as a transcription factor. Mammals have four Notch receptors (12), and five ligands: three Delta-like ligands (Dll1, 3 and 4), and two Serrate-like molecules called JAGGED 1 and JAGGED 2. NOTCH1 and Dll4 are the Notch receptor-ligand pair essential for T-cell development (108). Upon binding of Notch1 to its ligand, the intracellular portion of Notch (ICN) is released by proteolytic cleavage and translocates to the nucleus (reviewed in 109). In the nucleus, ICN binds the constitutively DNA-bound recombination signal binding protein for immunoglobulin kappa J region (RBPJk) or CBF1/suppressor of hairless/Lag-1 (CSL). ICN then forms a complex with CSL, which is stabilized by mastermind-like (MAML) proteins, and recruits coactivators such as p300 and the histone acetyl transferases PCAF and GCN5 to activate target gene transcription. In the context of T-cell development, the critical targets that mediate many of ICN's functions have not been identified, although ICN clearly regulates some essential T-cell-associated genes, including Ptcra, Cd25, and Hes1, an important regulator of DN3 cell proliferation (110).
The importance of Notch signaling in T-cell development has been demonstrated though both gain- and loss-of-function experiments. In the absence of NOTCH1, the ETP population is absent, although the bone marrow LMPP population is intact (111). Progenitors lacking NOTCH1 or CSL, or expressing negative regulators of Notch signals generate an increased number of B lymphocytes in the thymus (112). Extrinsic factors that block progenitors from receiving Notch signals, such as γ-secretase protease inhibitors (113), or other progenitors with a competitive advantage in Notch binding (114), also result in accentuated intrathymic B-cell development. At least a subset of these progenitors has been demonstrated to arise from cells that were destined for the T-cell fate (115). In contrast to Notch deficiency, ectopic NOTCH1 activation enforces the T-cell fate in vitro and in the bone marrow, at the expense of B-cell development (116-122), although constitutive Notch signals frequently result in aberrant development that is arrested at the CD4+CD8+ stage and a high frequency of leukemic transformation involving activation of the proto-oncogene c-myc.
There have been many indications that E proteins and the Notch signaling pathway interact during T-cell development. In LMPPs, E2A is required for proper expression of Notch1, suggesting that any ETPs that develop in E2A−/− mice would be defective in their ability to generate T cells. In vitro, E2A−/− hematopoietic progenitor cell lines and primary progenitor cells cultured in fetal thymic organ culture (FTOC) do fail to generate DN3 cells, and this deficiency can be overcome by ectopic expression of ICN (123). These findings demonstrate the power of ICN to promote T-cell development and are consistent with the hypothesis that Notch1 is a critical target of E2A whose decreased expression prevents T-cell development in vitro. However, analysis of Notch1 mRNA expression in E2A−/− DN2 and DN3 cells revealed that it is present at near WT levels (Xu and Kee, unpublished data). The molecular basis for this increase in Notch1 mRNA is not known; however, Notch1 is a target of the ICN/CSL/MAML complex and therefore the low level of NOTCH1 protein on the surface of E2A−/− TSPs may be sufficient to initiate the Notch signaling pathway leading to up-regulation of Notch1 mRNA. Alternatively, HEB/HEBALT, alone or in combination with other transcription factors, may be sufficient for Notch1 expression in the thymus. That Notch signaling is functional in E2A−/− DN2/DN3 cells seems likely given that they express WT levels of the Notch signaling target gene Deltex1 and can generate some mature T cells (Xu and Kee, unpublished data). However, another group reported that Deltex1 is increased in the DN1-DN2 transitional cells in the DN compartment of ID1 transgenic mice and that it may function to inhibit Notch signaling (124). Importantly, these transitional cells are not apparent in the E2A−/− thymus, indicating that the defects in this population may be unrelated to the functions of E2A in T-cell development.
Notch signaling has been proposed to inhibit E2A function under some conditions. In cells with high levels of activated mitogen-activated protein kinases, such as pro-B cells, Notch signaling promotes the degradation of E2A (125-127). However, during T lymphopoiesis, NOTCH1 and E2A have been shown to collaborate both directly (123, 128, 129) and indirectly via Deltex1 (130, 131). Transduction of E2A−/− progenitors with tamoxifen-inducible E47 (E47ER) directly upregulates targets of the Notch signaling pathway, including Hes-1, and Notch1, and expression of these genes is reduced in E2A−/− hematopoietic progenitor cell lines (123). Further, Notch and E2A synergize to upregulate Ptcra and CD7, and to enhance Tcrb rearrangement (123, 128, 132, 133). However, ectopic ICN1 restores in vitro CD4+CD8+ DP development from E2A−/− fetal thymocytes, suggesting that sufficiently strong Notch signals can overcome the requirement for E2A activity, although not excluding that there is still some compensation from HEB and/or E2-2 (123). The accumulating evidence supports a collaborative role for E2A and Notch signaling in T-cell specification.
T-cell commitment
NOTCH1 is the only protein that has been shown to be sufficient for specification and ultimately commitment to the T-lymphocyte fate, at least when activated in hematopoietic progenitors (134). Nonetheless, activation of NOTCH1 upon entry into the thymus does not immediately commit cells to T cell differentiation. Rather, a slow loss of lineage potentials occurs that is likely dependent on progressive changes in transcription factor expression or function, or accessibility to DNA (2). Unlike Pax5, however, Notch1 is not active at all stages of T-cell development and inhibiting Notch signaling after full commitment to the T-cell lineage does not cause lineage instability or expression of non-T-cell gene programs. Indeed, signaling through the pre-TCR results in a down-regulation of Notch1 mRNA as a consequence of ID3 induction and inhibition of E protein function (135). Therefore, Notch signaling establishes an appropriate context for T-cell lineage commitment but is not required to maintain this program in committed cells. Analysis of gene expression in T-cell progenitors revealed that commitment to the T-cell lineage is associated with a loss of stem cell-associated transcription factor genes such as Tal1, Gata2, and Sfpi1, rather than a dramatic induction of T-cell-associated gene expression (1, 136). This observation suggests that stem cell transcription factors may be maintaining access to alternative differentiation programs and that extinguishing their expression is necessary to prevent access to these programs by transcription factors that function in T-cell differentiation. Interestingly, one transcription factor that does increase during the commitment process is the transcriptional repressor BCL11b and recent reports demonstrate that deletion of Bcl11b in thymocytes leads to activation of NK cell gene expression and lineage conversion (137-139). Therefore, BCL11b plays a role in T=cell lineage commitment akin to that of PAX5 in B lymphocytes. However, the regulatory networks linking BCL11b to NOTCH1 and E proteins are currently not known.
A role for E2A in commitment to the T-cell fate has not been thoroughly investigated. There are more DN3 cells in E2A−/− mice than predicted based on the number of DN2 cells, suggesting that T-cell differentiation is recovering by this stage and that the development of committed cells is not affected by E2A deficiency (Xu and Kee, unpublished data). However, there are multiple observations that suggest a requirement for E2A in the generation of DN3 cells. First, unlike E2A−/− mice, Flt3L−/− and Ccr9−/− mice, which both have reduced ETP and DN2 cells populations, completely recover WT levels of DN3 cells (42, 140). Second, E2A−/− DN3 cells are hyper-proliferative, indicating that the actual number of DN2 cells progressing to the DN3 stage may be lower than evident by the steady state number of DN3 cells (141). Third, E2A−/− hematopoietic progenitors give rise to more NK cells than T cells when cultured in vitro or in FTOC (123). One possible explanation for reduced ability of E2A−/− DN2 cells to suppress NK cell differentiation could be a subtle difference in NOTCH1 expression levels. However, given the dynamic relationship between ID proteins and commitment factors for the B-cell fate, it seems likely that a similar dynamic exists between T-cell commitment factors and ID proteins in DN2 cells. BCL11b is a clear candidate for this antagonistic factor and Id2 is increased after deletion of Bcl11b in T cells (137-139); however, published data do not clearly support a role for E2A in the induction of BCL11b (136). Therefore, the mechanism by which E proteins suppress the NK cell fate remain to be determined.
E2A and Gfi1/1b
To identify E2A target genes in T lymphocytes, two groups have compared mRNA expression profiles in E2A−/− T-cell lymphomas retrovirally transduced with a bicistronic retrovirus encoding GFP alone or E47 and GFP (142, 143). Restoring either E47 or E12 expression to E2A−/− T lymphomas results in growth arrest and apoptosis (144, 145), leading to the hypothesis that E2A proteins remain capable of inducing their target genes in these lymphomas. In one of these retroviral screens, the transcriptional repressor growth factor independence 1b (GFI1b) was identified as a direct E47 target (142). In the other, Gfi1 was shown to be regulated downstream of E47 (143). In both screens, Gata3 mRNA decreased after expression of E47. GFI1 and GFI1b are of particular interest as potential E47 targets because they are paralogues of the Drosophila gene sens, which is a known target of E proteins during fly sensory organ development (146). Transgenic expression of GFI1b under the control of the lck promoter had already been demonstrated to reduce DP numbers (147), suggesting that it might mediate E47 induced growth arrest and apoptosis in E2A−/− lymphomas. Comparison of E2A−/− lymphoma lines transduced with GFI1b or control vector revealed that GFI1b does inhibit their proliferation and survival (142). GFI1b repressed GATA3 as well as GFI1, which was previously known to be repressed by GFI1b (148), and GATA3 was shown to be essential for the proliferation and survival of the lymphomas (142). Interestingly, in these cells GATA3 promoted the expression of GFI1b, suggesting an autoregulatory loop in which GATA3 might fine tune its own expression to control survival or proliferation. However, if E2A and GATA3 cooperatively regulate Gfi1b, Gfi1/1b may be less responsive to GATA3 in E2A−/− thymocytes and the GFI1/1b-mediated feedback inhibition of Gata3 may be defective, causing GATA3 to promote aberrant proliferation and survival of E2A−/− thymocytes. This model is consistent with the tendency of CD2-Gata3 transgenic mice, like E2A−/− mice, to develop an immature T-cell lymphoma.
Gata3 and T-cell lineage commitment
There is abundant evidence that GATA3 plays a role in both T-lymphocyte lineage specification and commitment (149-151). GATA transcription factors are characterized by two highly conserved C2C2 zinc fingers in the C-terminal domain which cooperatively recognize a T/A(GATA)A/G consensus sequence (152). In mice, complete deficiency in GATA3 results in death before e12.0, as a consequence of noradrenalin deficiency (153). RAG-2−/− blastocyst complementation experiments show that Gata3−/−, Gata3+/− and WT cells contribute similarly to B-cell and myeloid compartments, but Gata3−/− cells fail to contribute to the T-cell lineage (151). Recently studies also showed that GATA3 is required at the earliest stage of T-cell development, since ETPs are already decreased in Gata3−/− mice (154). Gata3 is expressed at very low levels in bone marrow LSKFlt3+ cells but is increased upon entry of TSPs into the thymus or when MPPs are cultured on stomal cells expressing DL1 (136, 155), indicating that Notch signaling controls expression of Gata3 early in development. Gata3 is expressed from two distinct promoters during T-cell development and only the Gata3a promoter is functional during thymopoiesis (156-158). Notch signaling has been shown to regulate expression of the Gata3b promoter in Th1 cells, but there is currently no evidence for a direct effect of the ICN/CSL/MAML complex on the Gata3a promoter (159, 160). Therefore, the mechanisms underlying Gata3 induction during thymopoiesis remain elusive.
While GATA3 is essential for T-cell development, overexpression of GATA3 blocks T-cell differentiation at the DN2 stage, prior to commitment (161, 162). Importantly, in the absence of Notch signaling, GATA3hi DN2 cells are reprogrammed to a mast cell fate through the rapid induction of mast cell genes, including Gata2, and the transient downregulation of Sfpi1. However, Notch signaling prevents the reprogramming of DN2 cells by GATA3 and an anti-proliferative effect of ectopic GATA3 on T-cell progenitors is revealed (161). The ability of increases in GATA3 expression to reprogram DN2 cells into mast cells suggests that there must be tight control of Gata3 expression during T-cell specification and commitment. Our observation that E2A, through induction of GFI1b, can repress Gata3 in T-cell lymphoma lines, suggests that E2A may play a role in preventing GATA3 from blocking T-cell lineage commitment or diverting DN2 cells away from the T-cell pathway. However, GFI1b is not highly expressed in T-cell progenitors and therefore the importance of this pathway in E2A-dependent T-cell development remains to be demonstrated (136, 142). Nonetheless, GFI1 is also regulated by E2A and GATA3 under some conditions and could contribute to limiting Gata3 at these critical stages of T-cell development. Future studies need to be directed at identifying the cis regulatory mechanisms controlling expression of Gata3 and how the transcription factors expressed at the DN2 stage function to promote differentiation and commitment to the T-lymphocyte lineage.
Concluding remarks
The E protein transcription factors are nearly ubiquitously expressed and yet play unique and essential roles at multiple stages of adaptive lymphopoiesis. For a summary of the functions we have described in this review see Fig. 3. Specificity in the functions of E protein transcription factors arise as a consequence of (i) differences in protein dimerization partners, which can alter the concentration of active E proteins, (ii) differences in expression levels, (iii) differences in chromatin context, which can influence which E protein target genes are accessible, and (iv) differences in the complement of cooperating transcription factors. We have argued here that the earliest functions for E proteins in lymphopoiesis are in the development and maintenance of HSCs as a consequence of their dimerization with TAL1 and LYL1, although no single E protein has thus far been shown to be absolutely essential. As HSC differentiate toward the lymphoid fates, E protein homodimers, composed predominately but not exclusively of E2A proteins, collaborate with other transcription factors in MPPs and LMPPs to promote lymphoid-priming and maintain cells on a trajectory to the lymphoid fates. Indeed, without these E2A proteins, LMPPs rapidly adopt a myeloid fate. Within lymphoid specified cells, E2A proteins are necessary to initiate the transcriptional regulatory network that results in B-lymphocyte lineage specification and commitment, and they likely prevent differentiation toward the NK cell fate, although this remains to be demonstrated. However, after commitment to the B-cell lineage lower concentrations of E protein activity are sufficient for the cooperative interactions that maintain the B cell program. The role of E proteins in specification and commitment to the T-lymphocyte lineage are less well defined, but the decreased number of DN3 cells and propensity to develop T-cell lymphoma indicate that E2A plays a critical role in guiding this process. Future studies should be directed at identifying E2A target genes in T lymphocytes and placing E proteins into the transcriptional networks that result in commitment to the T-cell fate. The mechanisms guiding T-cell lineage commitment are likely to have some parallels with B-cell lineage commitment; however, it is already clear that critical differences exist in the architecture of these programs. For example, elevated expression of GATA3, a critical regulator of T-cell lineage specification, arrests T-cell development prior to commitment and divert cells away from the T cell pathway in the absence of activated NOTCH1. E2A proteins likely play a role in harnessing GATA3 expression since targets of E2A in T-cell lymphomas are able to modulate the abundance of Gata3 mRNA. Nonetheless, the myriad of functions of E proteins in the development of adaptive lymphocytes are just beginning to be revealed. Understanding these functions will have important implications for controlling lymphocyte develop and preventing malignant transformation.
Fig. 3. Regulatory Networks in Lymphopoiesis involving E2A.
The model shows precursor progeny relationships in the lymphopoietic system denoted purple arrows. Transcriptional regulation is depicted using green arrows for positive regulation and red bars for negative regulation.
Acknowledgements
Work from our lab was supported by the NIH (R01s CA099978, AI079213, AI078267) and the American Cancer Society. R.F.deP. and B.L.K were supported by the Leukemia and Lymphoma Society.
References
- 1.Yui MA, Feng N, Rothenberg EV. Fine-scale staging of T cell lineage commitment in adult mouse thymus. J Immunol. 2010;185:284–93. doi: 10.4049/jimmunol.1000679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rothenberg EV, Moore JE, Yui MA. Launching the T-cell-lineage developmental programme. Nat Rev Immunol. 2008;8:9–21. doi: 10.1038/nri2232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Boos MD, Yokota Y, Eberl G, Kee BL. Mature natural killer cell and lymphoid tissue-inducing cell development requires Id2-mediated suppression of E protein activity. J Exp Med. 2007;204:1119–30. doi: 10.1084/jem.20061959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yokota Y, et al. Development of peripheral lymphoid organs and natural killer cells depends on the helix-loop-helix inhibitor Id2. Nature. 1999;397:702–6. doi: 10.1038/17812. [DOI] [PubMed] [Google Scholar]
- 5.Massari ME, Murre C. Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Mol Cell Biol. 2000;20:429–40. doi: 10.1128/mcb.20.2.429-440.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bhalla S, et al. differential roles for the E2A activation domains in B lymphocytes and macrophages. J Immunol. 2008;180:1694–703. doi: 10.4049/jimmunol.180.3.1694. [DOI] [PubMed] [Google Scholar]
- 7.Bayly R, et al. Critical role for a single leucine residue in leukemia induction by E2A-PBX1. Mol Cell Biol. 2006;26:6442–52. doi: 10.1128/MCB.02025-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kee BL, Arias J, Montminy MR. Adaptor-mediated recruitment of RNA polymerase II to a signal-dependent activator. J Biol Chem. 1996;271:2373–5. doi: 10.1074/jbc.271.5.2373. [DOI] [PubMed] [Google Scholar]
- 9.Greenbaum S, Lazorchak AS, Zhuang Y. Differential functions for the transcription factor E2A in positive and negative gene regulation in pre-B lymphocytes. J Biol Chem. 2004;279:45028–35. doi: 10.1074/jbc.M400061200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sun XH, Baltimore D. An inhibitory domain of E12 transcription factor prevents DNA binding in E12 homodimers but not in E12 heterodimers. Cell. 1991;64:459–70. doi: 10.1016/0092-8674(91)90653-g. [DOI] [PubMed] [Google Scholar]
- 11.Benezra R. An intermolecular disulfide bond stabilizes E2A homodimers and is required for DNA binding at physiological temperatures. Cell. 1994;79:1057–67. doi: 10.1016/0092-8674(94)90036-1. [DOI] [PubMed] [Google Scholar]
- 12.Barndt RJ, Dai M, Zhuang Y. Functions of E2A-HEB heterodimers in T-cell development revealed by a dominant negative mutation of HEB. Mol Cell Biol. 2000;20:6677–85. doi: 10.1128/mcb.20.18.6677-6685.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bain G, et al. E2A deficiency leads to abnormalities in alphabeta T-cell development and to rapid development of T-cell lymphomas. Mol Cell Biol. 1997;17:4782–91. doi: 10.1128/mcb.17.8.4782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wang D, et al. The basic helix-loop-helix transcription factor HEBAlt is expressed in pro-T cells and enhances the generation of T cell precursors. J Immunol. 2006;177:109–19. doi: 10.4049/jimmunol.177.1.109. [DOI] [PubMed] [Google Scholar]
- 15.Anderson MK. At the crossroads: diverse roles of early thymocyte transcriptional regulators. Immunol Rev. 2006;209:191–211. doi: 10.1111/j.0105-2896.2006.00352.x. [DOI] [PubMed] [Google Scholar]
- 16.Green AR, Begley CG. SCL and related hemopoietic helix-loop-helix transcription factors. Int J Cell Cloning. 1992;10:269–76. doi: 10.1002/stem.5530100504. [DOI] [PubMed] [Google Scholar]
- 17.Miyamoto A, Cui X, Naumovski L, Cleary ML. Helix-loop-helix proteins LYL1 and E2a form heterodimeric complexes with distinctive DNA-binding properties in hematolymphoid cells. Mol Cell Biol. 1996;16:2394–401. doi: 10.1128/mcb.16.5.2394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hsu HL, Cheng JT, Chen Q, Baer R. Enhancer-binding activity of the tal-1 oncoprotein in association with the E47/E12 helix-loop-helix proteins. Mol Cell Biol. 1991;11:3037–42. doi: 10.1128/mcb.11.6.3037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.O'Neil J, Billa M, Oikemus S, Kelliher M. The DNA binding activity of TAL-1 is not required to induce leukemia/lymphoma in mice. Oncogene. 2001;20:3897–905. doi: 10.1038/sj.onc.1204519. [DOI] [PubMed] [Google Scholar]
- 20.Benezra R, Davis RL, Lockshon D, Turner DL, Weintraub H. The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell. 1990;61:49–59. doi: 10.1016/0092-8674(90)90214-y. [DOI] [PubMed] [Google Scholar]
- 21.Langlands K, Yin X, Anand G, Prochownik EV. Differential interactions of Id proteins with basic-helix-loop-helix transcription factors. J Biol Chem. 1997;272:19785–93. doi: 10.1074/jbc.272.32.19785. [DOI] [PubMed] [Google Scholar]
- 22.Sun XH, Copeland NG, Jenkins NA, Baltimore D. Id proteins Id1 and Id2 selectively inhibit DNA binding by one class of helix-loop-helix proteins. Mol Cell Biol. 1991;11:5603–11. doi: 10.1128/mcb.11.11.5603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Iavarone A, Garg P, Lasorella A, Hsu J, Israel MA. The helix-loop-helix protein Id-2 enhances cell proliferation and binds to the retinoblastoma protein. Genes Dev. 1994;8:1270–84. doi: 10.1101/gad.8.11.1270. [DOI] [PubMed] [Google Scholar]
- 24.Sitnicka E, Bryder D, Theilgaard-Monch K, Buza-Vidas N, Adolfsson J, Jacobsen SE. Key role of flt3 ligand in regulation of the common lymphoid progenitor but not in maintenance of the hematopoietic stem cell pool. Immunity. 2002;17:463–72. doi: 10.1016/s1074-7613(02)00419-3. [DOI] [PubMed] [Google Scholar]
- 25.Zhuang Y, Soriano P, Weintraub H. The helix-loop-helix gene E2A is required for B cell formation. Cell. 1994;79:875–84. doi: 10.1016/0092-8674(94)90076-0. [DOI] [PubMed] [Google Scholar]
- 26.Bain G, et al. E2A proteins are required for proper B cell development and initiation of immunoglobulin gene rearrangements. Cell. 1994;79:885–92. doi: 10.1016/0092-8674(94)90077-9. [DOI] [PubMed] [Google Scholar]
- 27.Shivdasani RA, Mayer EL, Orkin SH. Absence of blood formation in mice lacking the T-cell leukaemia oncoprotein tal-1/SCL. Nature. 1995;373:432–4. doi: 10.1038/373432a0. [DOI] [PubMed] [Google Scholar]
- 28.Robb L, et al. Absence of yolk sac hematopoiesis from mice with a targeted disruption of the scl gene. Proc Natl Acad Sci U S A. 1995;92:7075–9. doi: 10.1073/pnas.92.15.7075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lecuyer E, et al. The SCL complex regulates c-kit expression in hematopoietic cells through functional interaction with Sp1. Blood. 2002;100:2430–40. doi: 10.1182/blood-2002-02-0568. [DOI] [PubMed] [Google Scholar]
- 30.Porcher C, Liao EC, Fujiwara Y, Zon LI, Orkin SH. Specification of hematopoietic and vascular development by the bHLH transcription factor SCL without direct DNA binding. Development. 1999;126:4603–15. doi: 10.1242/dev.126.20.4603. [DOI] [PubMed] [Google Scholar]
- 31.Lahlil R, Lecuyer E, Herblot S, Hoang T. SCL assembles a multifactorial complex that determines glycophorin A expression. Mol Cell Biol. 2004;24:1439–52. doi: 10.1128/MCB.24.4.1439-1452.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Capron C, et al. The SCL relative LYL-1 is required for fetal and adult hematopoietic stem cell function and B-cell differentiation. Blood. 2006;107:4678–86. doi: 10.1182/blood-2005-08-3145. [DOI] [PubMed] [Google Scholar]
- 33.Giroux S, et al. lyl-1 and tal-1/scl, two genes encoding closely related bHLH transcription factors, display highly overlapping expression patterns during cardiovascular and hematopoietic ontogeny. Gene Expr Patterns. 2007;7:215–26. doi: 10.1016/j.modgep.2006.10.004. [DOI] [PubMed] [Google Scholar]
- 34.Souroullas GP, Salmon JM, Sablitzky F, Curtis DJ, Goodell MA. Adult hematopoietic stem and progenitor cells require either Lyl1 or Scl for survival. Cell Stem Cell. 2009;4:180–6. doi: 10.1016/j.stem.2009.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yang Q, et al. E47 controls the developmental integrity and cell cycle quiescence of multipotential hematopoietic progenitors. J Immunol. 2008;181:5885–94. doi: 10.4049/jimmunol.181.9.5885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Semerad CL, Mercer EM, Inlay MA, Weissman IL, Murre C. E2A proteins maintain the hematopoietic stem cell pool and promote the maturation of myelolymphoid and myeloerythroid progenitors. Proc Natl Acad Sci U S A. 2009;106:1930–5. doi: 10.1073/pnas.0808866106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Dias S, Mansson R, Gurbuxani S, Sigvardsson M, Kee BL. E2A proteins promote development of lymphoid-primed multipotent progenitors. Immunity. 2008;29:217–27. doi: 10.1016/j.immuni.2008.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Forsberg EC, Serwold T, Kogan S, Weissman IL, Passegue E. New evidence supporting megakaryocyte-erythrocyte potential of flk2/flt3+ multipotent hematopoietic progenitors. Cell. 2006;126:415–26. doi: 10.1016/j.cell.2006.06.037. [DOI] [PubMed] [Google Scholar]
- 39.Adolfsson J, et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell. 2005;121:295–306. doi: 10.1016/j.cell.2005.02.013. [DOI] [PubMed] [Google Scholar]
- 40.Kondo M, Weissman IL, Akashi K. Identification of clonogenic common lymphoid progenitors in mouse bone marrow. Cell. 1997;91:661–72. doi: 10.1016/s0092-8674(00)80453-5. [DOI] [PubMed] [Google Scholar]
- 41.Bhandoola A, von Boehmer H, Petrie HT, Zuniga-Pflucker JC. Commitment and developmental potential of extrathymic and intrathymic T cell precursors: plenty to choose from. Immunity. 2007;26:678–89. doi: 10.1016/j.immuni.2007.05.009. [DOI] [PubMed] [Google Scholar]
- 42.Schwarz BA, Sambandam A, Maillard I, Harman BC, Love PE, Bhandoola A. Selective thymus settling regulated by cytokine and chemokine receptors. J Immunol. 2007;178:2008–17. doi: 10.4049/jimmunol.178.4.2008. [DOI] [PubMed] [Google Scholar]
- 43.Benz C, Bleul CC. A multipotent precursor in the thymus maps to the branching point of the T versus B lineage decision. J Exp Med. 2005;202:21–31. doi: 10.1084/jem.20050146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lai AY, Kondo M. Identification of a bone marrow precursor of the earliest thymocytes in adult mouse. Proc Natl Acad Sci U S A. 2007;104:6311–6. doi: 10.1073/pnas.0609608104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zlotoff DA, Sambandam A, Logan TD, Bell JJ, Schwarz BA, Bhandoola A. CCR7 and CCR9 together recruit hematopoietic progenitors to the adult thymus. Blood. 2010;115:1897–905. doi: 10.1182/blood-2009-08-237784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lai AY, Kondo M. Asymmetrical lymphoid and myeloid lineage commitment in multipotent hematopoietic progenitors. J Exp Med. 2006;203:1867–73. doi: 10.1084/jem.20060697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Igarashi H, Gregory SC, Yokota T, Sakaguchi N, Kincade PW. Transcription from the RAG1 locus marks the earliest lymphocyte progenitors in bone marrow. Immunity. 2002;17:117–30. doi: 10.1016/s1074-7613(02)00366-7. [DOI] [PubMed] [Google Scholar]
- 48.Perry SS, Welner RS, Kouro T, Kincade PW, Sun XH. Primitive lymphoid progenitors in bone marrow with T lineage reconstituting potential. J Immunol. 2006;177:2880–7. doi: 10.4049/jimmunol.177.5.2880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Perry SS, Pierce LJ, Slayton WB, Spangrude GJ. Characterization of thymic progenitors in adult mouse bone marrow. J Immunol. 2003;170:1877–86. doi: 10.4049/jimmunol.170.4.1877. [DOI] [PubMed] [Google Scholar]
- 50.Perry SS, Wang H, Pierce LJ, Yang AM, Tsai S, Spangrude GJ. L-selectin defines a bone marrow analog to the thymic early T-lineage progenitor. Blood. 2004;103:2990–6. doi: 10.1182/blood-2003-09-3030. [DOI] [PubMed] [Google Scholar]
- 51.Mansson R, et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity. 2007;26:407–19. doi: 10.1016/j.immuni.2007.02.013. [DOI] [PubMed] [Google Scholar]
- 52.Arinobu Y, et al. Reciprocal activation of GATA-1 and PU.1 marks initial specification of hematopoietic stem cells into myeloerythroid and myelolymphoid lineages. Cell Stem Cell. 2007;1:416–27. doi: 10.1016/j.stem.2007.07.004. [DOI] [PubMed] [Google Scholar]
- 53.Luc S, et al. Down-regulation of Mpl marks the transition to lymphoid-primed multipotent progenitors with gradual loss of granulocyte-monocyte potential. Blood. 2008;111:3424–34. doi: 10.1182/blood-2007-08-108324. [DOI] [PubMed] [Google Scholar]
- 54.Cochrane SW, Zhao Y, Welner RS, Sun XH. Balance between Id and E proteins regulates myeloid-versus-lymphoid lineage decisions. Blood. 2009;113:1016–26. doi: 10.1182/blood-2008-06-164996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Carotta S, et al. The transcription factor PU.1 controls dendritic cell development and Flt3 cytokine receptor expression in a dose-dependent manner. Immunity. 2010;32:628–41. doi: 10.1016/j.immuni.2010.05.005. [DOI] [PubMed] [Google Scholar]
- 56.Yoshida T, Ng SY, Zuniga-Pflucker JC, Georgopoulos K. Early hematopoietic lineage restrictions directed by Ikaros. Nat Immunol. 2006;7:382–91. doi: 10.1038/ni1314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Medina KL, et al. Assembling a gene regulatory network for specification of the B cell fate. Dev Cell. 2004;7:607–17. doi: 10.1016/j.devcel.2004.08.006. [DOI] [PubMed] [Google Scholar]
- 58.Ng SY, Yoshida T, Zhang J, Georgopoulos K. Genome-wide lineage-specific transcriptional networks underscore Ikaros-dependent lymphoid priming in hematopoietic stem cells. Immunity. 2009;30:493–507. doi: 10.1016/j.immuni.2009.01.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.DeKoter RP, Singh H. Regulation of B lymphocyte and macrophage development by graded expression of PU.1. Science. 2000;288:1439–41. doi: 10.1126/science.288.5470.1439. [DOI] [PubMed] [Google Scholar]
- 60.Spooner CJ, Cheng JX, Pujadas E, Laslo P, Singh H. A recurrent network involving the transcription factors PU.1 and Gfi1 orchestrates innate and adaptive immune cell fates. Immunity. 2009;31:576–86. doi: 10.1016/j.immuni.2009.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Zeng H, Yucel R, Kosan C, Klein-Hitpass L, Moroy T. Transcription factor Gfi1 regulates self-renewal and engraftment of hematopoietic stem cells. EMBO J. 2004;23:4116–25. doi: 10.1038/sj.emboj.7600419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Inlay MA, et al. Ly6d marks the earliest stage of B-cell specification and identifies the branchpoint between B-cell and T-cell development. Genes Dev. 2009;23:2376–81. doi: 10.1101/gad.1836009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Mansson R, et al. Single-cell analysis of the common lymphoid progenitor compartment reveals functional and molecular heterogeneity. Blood. 2010;115:2601–9. doi: 10.1182/blood-2009-08-236398. [DOI] [PubMed] [Google Scholar]
- 64.Zandi S, Mansson R, Tsapogas P, Zetterblad J, Bryder D, Sigvardsson M. EBF1 is essential for B-lineage priming and establishment of a transcription factor network in common lymphoid progenitors. J Immunol. 2008;181:3364–72. doi: 10.4049/jimmunol.181.5.3364. [DOI] [PubMed] [Google Scholar]
- 65.Kee BL, Murre C. Induction of early B cell factor (EBF) and multiple B lineage genes by the basic helix-loop-helix transcription factor E12. J Exp Med. 1998;188:699–713. doi: 10.1084/jem.188.4.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Smith EM, Gisler R, Sigvardsson M. Cloning and characterization of a promoter flanking the early B cell factor (EBF) gene indicates roles for E-proteins and autoregulation in the control of EBF expression. J Immunol. 2002;169:261–70. doi: 10.4049/jimmunol.169.1.261. [DOI] [PubMed] [Google Scholar]
- 67.Roessler S, et al. Distinct promoters mediate the regulation of Ebf1 gene expression by interleukin-7 and Pax5. Mol Cell Biol. 2007;27:579–94. doi: 10.1128/MCB.01192-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Seet CS, Brumbaugh RL, Kee BL. Early B cell factor promotes B lymphopoiesis with reduced interleukin 7 responsiveness in the absence of E2A. J Exp Med. 2004;199:1689–700. doi: 10.1084/jem.20032202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kwon K, et al. Instructive role of the transcription factor E2A in early B lymphopoiesis and germinal center B cell development. Immunity. 2008;28:751–62. doi: 10.1016/j.immuni.2008.04.014. [DOI] [PubMed] [Google Scholar]
- 70.Zhuang Y, Barndt RJ, Pan L, Kelley R, Dai M. Functional replacement of the mouse E2A gene with a human HEB cDNA. Mol Cell Biol. 1998;18:3340–9. doi: 10.1128/mcb.18.6.3340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zhuang Y, Jackson A, Pan L, Shen K, Dai M. Regulation of E2A gene expression in B-lymphocyte development. Mol Immunol. 2004;40:1165–77. doi: 10.1016/j.molimm.2003.11.031. [DOI] [PubMed] [Google Scholar]
- 72.Schlenner SM, et al. Fate mapping reveals separate origins of T cells and myeloid lineages in the thymus. Immunity. 2010;32:426–36. doi: 10.1016/j.immuni.2010.03.005. [DOI] [PubMed] [Google Scholar]
- 73.Kikuchi K, Kasai H, Watanabe A, Lai AY, Kondo M. IL-7 specifies B cell fate at the common lymphoid progenitor to pre-proB transition stage by maintaining early B cell factor expression. J Immunol. 2008;181:383–92. doi: 10.4049/jimmunol.181.1.383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kikuchi K, Lai AY, Hsu CL, Kondo M. IL-7 receptor signaling is necessary for stage transition in adult B cell development through up-regulation of EBF. J Exp Med. 2005;201:1197–203. doi: 10.1084/jem.20050158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Dias S, Silva H, Jr., Cumano A, Vieira P. Interleukin-7 is necessary to maintain the B cell potential in common lymphoid progenitors. J Exp Med. 2005;201:971–9. doi: 10.1084/jem.20042393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Malin S, McManus S, Busslinger M. STAT5 in B cell development and leukemia. Curr Opin Immunol. 2010;22:168–76. doi: 10.1016/j.coi.2010.02.004. [DOI] [PubMed] [Google Scholar]
- 77.Malin S, et al. Role of STAT5 in controlling cell survival and immunoglobulin gene recombination during pro-B cell development. Nat Immunol. 2010;11:171–9. doi: 10.1038/ni.1827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Park JH, et al. Signaling by intrathymic cytokines, not T cell antigen receptors, specifies CD8 lineage choice and promotes the differentiation of cytotoxic-lineage T cells. Nat Immunol. 2010;11:257–64. doi: 10.1038/ni.1840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Oguro H, et al. Poised lineage specification in multipotential hematopoietic stem and progenitor cells by the polycomb protein Bmi1. Cell Stem Cell. 2010;6:279–86. doi: 10.1016/j.stem.2010.01.005. [DOI] [PubMed] [Google Scholar]
- 80.Lin H, Grosschedl R. Failure of B-cell differentiation in mice lacking the transcription factor EBF. Nature. 1995;376:263–7. doi: 10.1038/376263a0. [DOI] [PubMed] [Google Scholar]
- 81.O'Riordan M, Grosschedl R. Coordinate regulation of B cell differentiation by the transcription factors EBF and E2A. Immunity. 1999;11:21–31. doi: 10.1016/s1074-7613(00)80078-3. [DOI] [PubMed] [Google Scholar]
- 82.Nutt SL, Heavey B, Rolink AG, Busslinger M. Commitment to the B-lymphoid lineage depends on the transcription factor Pax5. Nature. 1999;401:556–62. doi: 10.1038/44076. [DOI] [PubMed] [Google Scholar]
- 83.Rolink AG, Nutt SL, Melchers F, Busslinger M. Long-term in vivo reconstitution of T-cell development by Pax5-deficient B-cell progenitors. Nature. 1999;401:603–6. doi: 10.1038/44164. [DOI] [PubMed] [Google Scholar]
- 84.Nutt SL, Morrison AM, Dorfler P, Rolink A, Busslinger M. Identification of BSAP (Pax-5) target genes in early B-cell development by loss- and gain-of-function experiments. EMBO J. 1998;17:2319–33. doi: 10.1093/emboj/17.8.2319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Cobaleda C, Jochum W, Busslinger M. Conversion of mature B cells into T cells by dedifferentiation to uncommitted progenitors. Nature. 2007;449:473–7. doi: 10.1038/nature06159. [DOI] [PubMed] [Google Scholar]
- 86.Thal MA, et al. Ebf1-mediated down-regulation of Id2 and Id3 is essential for specification of the B cell lineage. Proc Natl Acad Sci U S A. 2009;106:552–7. doi: 10.1073/pnas.0802550106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Pongubala JM, et al. Transcription factor EBF restricts alternative lineage options and promotes B cell fate commitment independently of Pax5. Nat Immunol. 2008;9:203–15. doi: 10.1038/ni1555. [DOI] [PubMed] [Google Scholar]
- 88.Kee BL. E and ID proteins branch out. Nat Rev Immunol. 2009;9:175–84. doi: 10.1038/nri2507. [DOI] [PubMed] [Google Scholar]
- 89.Treiber T, et al. Early B cell factor 1 regulates B cell gene networks by activation, repression, and transcription- independent poising of chromatin. Immunity. 2010;32:714–25. doi: 10.1016/j.immuni.2010.04.013. [DOI] [PubMed] [Google Scholar]
- 90.Lin YC, et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat Immunol. 2010;11:635–43. doi: 10.1038/ni.1891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Zhuang Y, Cheng P, Weintraub H. B-lymphocyte development is regulated by the combined dosage of three basic helix-loop-helix genes, E2A, E2-2, and HEB. Mol Cell Biol. 1996;16:2898–905. doi: 10.1128/mcb.16.6.2898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Lazorchak AS, Wojciechowski J, Dai M, Zhuang Y. E2A promotes the survival of precursor and mature B lymphocytes. J Immunol. 2006;177:2495–504. doi: 10.4049/jimmunol.177.4.2495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Kee BL. Id3 induces growth arrest and caspase-2-dependent apoptosis in B lymphocyte progenitors. J Immunol. 2005;175:4518–27. doi: 10.4049/jimmunol.175.7.4518. [DOI] [PubMed] [Google Scholar]
- 94.Greenbaum S, Zhuang Y. Identification of E2A target genes in B lymphocyte development by using a gene tagging-based chromatin immunoprecipitation system. Proc Natl Acad Sci U S A. 2002;99:15030–5. doi: 10.1073/pnas.232299999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Schlissel MS. Regulation of activation and recombination of the murine Igkappa locus. Immunol Rev. 2004;200:215–23. doi: 10.1111/j.0105-2896.2004.00157.x. [DOI] [PubMed] [Google Scholar]
- 96.Romanow WJ, et al. E2A and EBF act in synergy with the V(D)J recombinase to generate a diverse immunoglobulin repertoire in nonlymphoid cells. Mol Cell. 2000;5:343–53. doi: 10.1016/s1097-2765(00)80429-3. [DOI] [PubMed] [Google Scholar]
- 97.Goebel P, Janney N, Valenzuela JR, Romanow WJ, Murre C, Feeney AJ. Localized gene-specific induction of accessibility to V(D)J recombination induced by E2A and early B cell factor in nonlymphoid cells. J Exp Med. 2001;194:645–56. doi: 10.1084/jem.194.5.645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Mandal M, et al. Ras orchestrates exit from the cell cycle and light-chain recombination during early B cell development. Nat Immunol. 2009;10:1110–7. doi: 10.1038/ni.1785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Petrie HT. Cell migration and the control of post-natal T-cell lymphopoiesis in the thymus. Nat Rev Immunol. 2003;3:859–66. doi: 10.1038/nri1223. [DOI] [PubMed] [Google Scholar]
- 100.Scollay R, Bartlett P, Shortman K. T cell development in the adult murine thymus: changes in the expression of the surface antigens Ly2, L3T4 and B2A2 during development from early precursor cells to emigrants. Immunol Rev. 1984;82:79–103. doi: 10.1111/j.1600-065x.1984.tb01118.x. [DOI] [PubMed] [Google Scholar]
- 101.Sambandam A, et al. Notch signaling controls the generation and differentiation of early T lineage progenitors. Nat Immunol. 2005;6:663–70. doi: 10.1038/ni1216. [DOI] [PubMed] [Google Scholar]
- 102.Yan W, Young AZ, Soares VC, Kelley R, Benezra R, Zhuang Y. High incidence of T-cell tumors in E2A-null mice and E2A/Id1 double-knockout mice. Mol Cell Biol. 1997;17:7317–27. doi: 10.1128/mcb.17.12.7317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Engel I, Johns C, Bain G, Rivera RR, Murre C. Early thymocyte development is regulated by modulation of E2A protein activity. Journal of Experimental Medicine. 2001;194:733–45. doi: 10.1084/jem.194.6.733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Pan L, Hanrahan J, Li J, Hale LP, Zhuang Y. An analysis of T cell intrinsic roles of E2A by conditional gene disruption in the thymus. J Immunol. 2002;168:3923–32. doi: 10.4049/jimmunol.168.8.3923. [DOI] [PubMed] [Google Scholar]
- 105.Barndt R, Dai MF, Zhuang Y. A novel role for HEB downstream or parallel to the pre-TCR signaling pathway during alpha beta thymopoiesis. J Immunol. 1999;163:3331–43. [PubMed] [Google Scholar]
- 106.Yang Y, Wang HC, Sun XH. Id1 induces apoptosis through inhibition of RORgammat expression. BMC Immunol. 2008;9:20. doi: 10.1186/1471-2172-9-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kim D, Peng XC, Sun XH. Massive apoptosis of thymocytes in T-cell-deficient Id1 transgenic mice. Mol Cell Biol. 1999;19:8240–53. doi: 10.1128/mcb.19.12.8240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Koch U, et al. Delta-like 4 is the essential, nonredundant ligand for Notch1 during thymic T cell lineage commitment. J Exp Med. 2008;205:2515–23. doi: 10.1084/jem.20080829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Pear WS, Radtke F. Notch signaling in lymphopoiesis. Semin Immunol. 2003;15:69–79. doi: 10.1016/s1044-5323(03)00003-4. [DOI] [PubMed] [Google Scholar]
- 110.Tomita K, Hattori M, Nakamura E, Nakanishi S, Minato N, Kageyama R. The bHLH gene Hes1 is essential for expansion of early T cell precursors. Genes Dev. 1999;13:1203–10. doi: 10.1101/gad.13.9.1203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Tan JB, Visan I, Yuan JS, Guidos CJ. Requirement for Notch1 signals at sequential early stages of intrathymic T cell development. Nat Immunol. 2005;6:671–9. doi: 10.1038/ni1217. [DOI] [PubMed] [Google Scholar]
- 112.Han H, et al. Inducible gene knockout of transcription factor recombination signal binding protein-j reveals it's essential role in T versus B lineage decision. International Immunity. 2002;14:637–45. doi: 10.1093/intimm/dxf030. [DOI] [PubMed] [Google Scholar]
- 113.Doerfler P, Shearman MS, Perlmutter RM. Presenilin-dependent gamma-secretase activity modulates thymocyte development. Proc Natl Acad Sci U S A. 2001;98:9312–7. doi: 10.1073/pnas.161102498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Koch U, et al. Subversion of the T/B lineage decision in the thymus by lunatic fringe-mediated inhibition of Notch-1. Immunity. 2001;15:225–36. doi: 10.1016/s1074-7613(01)00189-3. [DOI] [PubMed] [Google Scholar]
- 115.Feyerabend TB, et al. Deletion of Notch1 converts pro-T cells to dendritic cells and promotes thymic B cells by cell-extrinsic and cell-intrinsic mechanisms. Immunity. 2009;30:67–79. doi: 10.1016/j.immuni.2008.10.016. [DOI] [PubMed] [Google Scholar]
- 116.Hozumi K, Abe N, Chiba S, Hirai H, Habu S. Active form of Notch members can enforce T lymphopoiesis on lymphoid progenitors in the monolayer culture specific for B cell development. J Immunol. 2003;170:4973–9. doi: 10.4049/jimmunol.170.10.4973. [DOI] [PubMed] [Google Scholar]
- 117.De Smedt M, et al. Active form of Notch imposes T cell fate in human progenitor cells. J Immunol. 2002;169:3021–9. doi: 10.4049/jimmunol.169.6.3021. [DOI] [PubMed] [Google Scholar]
- 118.Garcia-Peydro M, de Yebenes VG, Toribio ML. Notch1 and IL-7 receptor interplay maintains proliferation of human thymic progenitors while suppressing non-T cell fates. J Immunol. 2006;177:3711–20. doi: 10.4049/jimmunol.177.6.3711. [DOI] [PubMed] [Google Scholar]
- 119.Pui JC, et al. Notch1 expression in early lymphopoiesis influences B versus T lineage determination. Immunity. 1999;11:299–308. doi: 10.1016/s1074-7613(00)80105-3. [DOI] [PubMed] [Google Scholar]
- 120.Dorsch M, et al. Ectopic expression of Delta4 impairs hematopoietic development and leads to lymphoproliferative disease. Blood. 2002;100:2046–55. [PubMed] [Google Scholar]
- 121.Yan XQ, et al. A novel Notch ligand, Dll4, induces T-cell leukemia/lymphoma when overexpressed in mice by retroviral-mediated gene transfer. Blood. 2001;98:3793–9. doi: 10.1182/blood.v98.13.3793. [DOI] [PubMed] [Google Scholar]
- 122.de La Coste A, et al. In vivo and in absence of a thymus, the enforced expression of the Notch ligands delta-1 or delta-4 promotes T cell development with specific unique effects. J Immunol. 2005;174:2730–7. doi: 10.4049/jimmunol.174.5.2730. [DOI] [PubMed] [Google Scholar]
- 123.Ikawa T, Kawamoto H, Goldrath AW, Murre C. E proteins and Notch signaling cooperate to promote T cell lineage specification and commitment. J Exp Med. 2006;203:1329–42. doi: 10.1084/jem.20060268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Wang HC, Perry SS, Sun XH. Id1 attenuates Notch signaling and impairs T-cell commitment by elevating Deltex1 expression. Mol Cell Biol. 2009;29:4640–52. doi: 10.1128/MCB.00119-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Ordentlich P, et al. Notch inhibition of E47 supports the existence of a novel signaling pathway. Mol Cell Biol. 1998;18:2230–9. doi: 10.1128/mcb.18.4.2230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Huang Z, Nie L, Xu M, Sun XH. Notch-induced E2A degradation requires CHIP and Hsc70 as novel facilitators of ubiquitination. Mol Cell Biol. 2004;24:8951–62. doi: 10.1128/MCB.24.20.8951-8962.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Nie L, Xu M, Vladimirova A, Sun XH. Notch-induced E2A ubiquitination and degradation are controlled by MAP kinase activities. Embo J. 2003;22:5780–92. doi: 10.1093/emboj/cdg567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Agata Y, et al. Regulation of T cell receptor beta gene rearrangements and allelic exclusion by the helix-loop-helix protein, E47. Immunity. 2007;27:871–84. doi: 10.1016/j.immuni.2007.11.015. [DOI] [PubMed] [Google Scholar]
- 129.Tanigaki K, Honjo T. Regulation of lymphocyte development by Notch signaling. Nat Immunol. 2007;8:451–6. doi: 10.1038/ni1453. [DOI] [PubMed] [Google Scholar]
- 130.Matsuno K, et al. Human deltex is a conserved regulator of Notch signalling. Nat Genet. 1998;19:74–8. doi: 10.1038/ng0598-74. [DOI] [PubMed] [Google Scholar]
- 131.Kishi N, et al. Murine homologs of deltex define a novel gene family involved in vertebrate Notch signaling and neurogenesis. Int J Dev Neurosci. 2001;19:21–35. doi: 10.1016/s0736-5748(00)00071-x. [DOI] [PubMed] [Google Scholar]
- 132.Wolfer A, Wilson A, Nemir M, MacDonald HR, Radtke F. Inactivation of Notch1 impairs VDJbeta rearrangement and allows pre-TCR-independent survival of early alpha beta Lineage Thymocytes. Immunity. 2002;16:869–79. doi: 10.1016/s1074-7613(02)00330-8. [DOI] [PubMed] [Google Scholar]
- 133.Reizis B, Leder P. Direct induction of T lymphocyte-specific gene expression by the mammalian Notch signaling pathway. Genes Dev. 2002;16:295–300. doi: 10.1101/gad.960702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.de Pooter R, Zuniga-Pflucker JC. T-cell potential and development in vitro: the OP9-DL1 approach. Curr Opin Immunol. 2007;19:163–8. doi: 10.1016/j.coi.2007.02.011. [DOI] [PubMed] [Google Scholar]
- 135.Yashiro-Ohtani Y, et al. Pre-TCR signaling inactivates Notch1 transcription by antagonizing E2A. Genes Dev. 2009;23:1665–76. doi: 10.1101/gad.1793709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.David-Fung ES, et al. Transcription factor expression dynamics of early T-lymphocyte specification and commitment. Dev Biol. 2009;325:444–67. doi: 10.1016/j.ydbio.2008.10.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Ikawa T, et al. An essential developmental checkpoint for production of the T cell lineage. Science. 2010;329:93–6. doi: 10.1126/science.1188995. [DOI] [PubMed] [Google Scholar]
- 138.Li L, Leid M, Rothenberg EV. An early T cell lineage commitment checkpoint dependent on the transcription factor Bcl11b. Science. 2010;329:89–93. doi: 10.1126/science.1188989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Li P, et al. Reprogramming of T cells to natural killer-like cells upon Bcl11b deletion. Science. 2010;329:85–9. doi: 10.1126/science.1188063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Sitnicka E, et al. Critical role of FLT3 ligand in IL-7 receptor independent T lymphopoiesis and regulation of lymphoid-primed multipotent progenitors. Blood. 2007;110:2955–64. doi: 10.1182/blood-2006-10-054726. [DOI] [PubMed] [Google Scholar]
- 141.Engel I, Murre C. E2A proteins enforce a proliferation checkpoint in developing thymocytes. Embo J. 2004;23:202–11. doi: 10.1038/sj.emboj.7600017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Xu W, Kee BL. Growth factor independent 1B (Gfi1b) is an E2A target gene that modulates Gata3 in T-cell lymphomas. Blood. 2007;109:4406–14. doi: 10.1182/blood-2006-08-043331. [DOI] [PubMed] [Google Scholar]
- 143.Schwartz R, Engel I, Fallahi-Sichani M, Petrie HT, Murre C. Gene expression patterns define novel roles for E47 in cell cycle progression, cytokine-mediated signaling, and T lineage development. Proc Natl Acad Sci U S A. 2006;103:9976–81. doi: 10.1073/pnas.0603728103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Engel I, Murre C. Ectopic expression of E47 or E12 promotes the death of E2A-deficient lymphomas. Proc Natl Acad Sci U S A. 1999;96:996–1001. doi: 10.1073/pnas.96.3.996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Park ST, Nolan GP, Sun XH. Growth inhibition and apoptosis due to restoration of E2A activity in T cell acute lymphoblastic leukemia cells. J Exp Med. 1999;189:501–8. doi: 10.1084/jem.189.3.501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Jafar-Nejad H, et al. Senseless acts as a binary switch during sensory organ precursor selection. Genes Dev. 2003;17:2966–78. doi: 10.1101/gad.1122403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Doan LL, et al. Growth factor independence-1B expression leads to defects in T cell activation, IL-7 receptor alpha expression, and T cell lineage commitment. J Immunol. 2003;170:2356–66. doi: 10.4049/jimmunol.170.5.2356. [DOI] [PubMed] [Google Scholar]
- 148.Vassen L, Fiolka K, Mahlmann S, Moroy T. Direct transcriptional repression of the genes encoding the zinc-finger proteins Gfi1b and Gfi1 by Gfi1b. Nucleic Acids Res. 2005;33:987–98. doi: 10.1093/nar/gki243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Anderson MK, et al. Evolutionary origins of lymphocytes: ensembles of T cell and B cell transcriptional regulators in a cartilaginous fish. J Immunol. 2004;172:5851–60. doi: 10.4049/jimmunol.172.10.5851. [DOI] [PubMed] [Google Scholar]
- 150.Ko LJ, Yamamoto M, Leonard MW, George KM, Ting P, Engel JD. Murine and human T-lymphocyte GATA-3 factors mediate transcription through a cis-regulatory element within the human T-cell receptor delta gene enhancer. Mol Cell Biol. 1991;11:2778–84. doi: 10.1128/mcb.11.5.2778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Ting CN, Olson MC, Barton KP, Leiden JM. Transcription factor GATA-3 is required for development of the T-cell lineage. Nature. 1996;384:474–8. doi: 10.1038/384474a0. [DOI] [PubMed] [Google Scholar]
- 152.Rothenberg EV, Scripture-Adams DD. Competition and collaboration: GATA-3, PU.1, and Notch signaling in early T-cell fate determination. Semin Immunol. 2008;20:236–46. doi: 10.1016/j.smim.2008.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Lim KC, Lakshmanan G, Crawford SE, Gu Y, Grosveld F, Engel JD. Gata3 loss leads to embryonic lethality due to noradrenaline deficiency of the sympathetic nervous system. Nat Genet. 2000;25:209–12. doi: 10.1038/76080. [DOI] [PubMed] [Google Scholar]
- 154.Hosoya T, et al. GATA-3 is required for early T lineage progenitor development. J Exp Med. 2009;206:2987–3000. doi: 10.1084/jem.20090934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Taghon TN, David ES, Zuniga-Pflucker JC, Rothenberg EV. Delayed, asynchronous, and reversible T-lineage specification induced by Notch/Delta signaling. Genes Dev. 2005;19:965–78. doi: 10.1101/gad.1298305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Asnagli H, Afkarian M, Murphy KM. Cutting edge: Identification of an alternative GATA-3 promoter directing tissue-specific gene expression in mouse and human. J Immunol. 2002;168:4268–71. doi: 10.4049/jimmunol.168.9.4268. [DOI] [PubMed] [Google Scholar]
- 157.Nawijn MC, et al. Enforced expression of GATA-3 during T cell development inhibits maturation of CD8 single-positive cells and induces thymic lymphoma in transgenic mice. J Immunol. 2001;167:715–23. doi: 10.4049/jimmunol.167.2.715. [DOI] [PubMed] [Google Scholar]
- 158.Hendriks RW, Nawijn MC, Engel JD, van Doorninck H, Grosveld F, Karis A. Expression of the transcription factor GATA-3 is required for the development of the earliest T cell progenitors and correlates with stages of cellular proliferation in the thymus. Eur J Immunol. 1999;29:1912–8. doi: 10.1002/(SICI)1521-4141(199906)29:06<1912::AID-IMMU1912>3.0.CO;2-D. [DOI] [PubMed] [Google Scholar]
- 159.Fang TC, Yashiro-Ohtani Y, Del Bianco C, Knoblock DM, Blacklow SC, Pear WS. Notch directly regulates Gata3 expression during T helper 2 cell differentiation. Immunity. 2007;27:100–10. doi: 10.1016/j.immuni.2007.04.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Amsen D, et al. Direct regulation of Gata3 expression determines the T helper differentiation potential of Notch. Immunity. 2007;27:89–99. doi: 10.1016/j.immuni.2007.05.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Taghon T, Yui MA, Rothenberg EV. Mast cell lineage diversion of T lineage precursors by the essential T cell transcription factor GATA-3. Nat Immunol. 2007;8:845–55. doi: 10.1038/ni1486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Anderson MK, Hernandez-Hoyos G, Dionne CJ, Arias AM, Chen D, Rothenberg EV. Definition of regulatory network elements for T cell development by perturbation analysis with PU.1 and GATA-3. Dev Biol. 2002;246:103–21. doi: 10.1006/dbio.2002.0674. [DOI] [PubMed] [Google Scholar]