Abstract
Among mutations in human Runx1/AML1 transcription factors, the t(8;21)(q22;q22) genomic translocation that creates an AML1-ETO fusion protein is implicated in etiology of the acute myeloid leukemia. To identify genes and components associated with this oncogene we used Drosophila as a genetic model. Expression of AML1-ETO caused an expansion of hematopoietic precursors in Drosophila, which expressed high levels of reactive oxygen species (ROS). Mutations in functional domains of the fusion protein suppress the proliferative phenotype. In a genetic screen, we found that inactivation of EcRB1 or activation of Foxo and superoxide dismutase-2 (SOD2) suppress the AML1-ETO–induced phenotype by reducing ROS expression in the precursor cells. Our studies indicate that ROS is a signaling factor promoting maintenance of normal as well as the aberrant myeloid precursors and suggests the importance of antioxidant enzymes and their regulators as targets for further study in the context of leukemia.
Introduction
RUNX genes encode highly conserved Runt domain transcription factors (AML/RUNX proteins), which play critical roles during blood cell development in metazoan species.1–4 Mutations that disrupt the activity of AML1/Runx1 lead to hematopoietic disorders including myelodysplastic syndrome and acute myeloid leukemia (AML).2,5 AML is the most frequent and debilitating form of leukemia with approximately a quarter million new cases diagnosed each year worldwide. The t(8;21) (q22;q22) chromosomal translocation that creates the AML1-ETO fusion product is present in 12%-15% of all cases of AML patients. The M2 subtype of AML carrying the AML1-ETO fusion protein is characterized by excessive production of granulocyte precursors.6
Under normal conditions, AML1/RUNX1 (also known as core binding factor-α, CBFα) in association with CBFβ binds the enhancer core sequence, TGT/CGGT, to activate tissue-specific expression of a number of hematopoietic genes.4 ETO (also known as MTG8) functions as a transcriptional repressor by recruiting a protein complex consisting of histone deacetylases (HDACs), the nuclear receptor co-receptor (N-CoR), mSin3A and the silencing mediator of retinoid and thyroid hormone receptor (SMRT).7 ETO function has been implicated in the Notch pathway,8 and it interacts with SON and TCF4.9,10 The (8,21)(q22;q22) chromosomal aberration produces a potent oncogenic fusion protein consisting of most of the Runt domain of AML1 and nearly the complete sequence of ETO.11 The general view among investigators in the field is that the AML1-ETO fusion product acts as a constitutive repressor form of AML1.12,13 It has also been shown that this fusion protein has other activities in addition to interfering with AML1 function.14,15 For example, AML1-ETO knock-in mouse embryos contain highly proliferative multilineage hematopoietic progenitors that are not found when either AML1 or CBFβ is disrupted.12,14 Evidently, the full range of AML1-ETO target genes as well as components involved in the pathway, and their relevance to AML pathogenesis is not completely understood.
To further our understanding of AML pathogenesis, additional animal models that allow in vivo genetic analysis are needed to complement studies in the existing mouse models. To this end, a Drosophila model is valuable due to the availability of facile genetic screens, a nearly complete collection of mutants, and advanced genetic technologies. The Drosophila hematopoietic system bears similarity to the mammalian myeloid system, and the molecular machinery that regulates differentiation and proliferation of blood cells tends to be evolutionary conserved.16,17 In Drosophila, the hematopoietic tissue originates from 2 independent embryonic anlagen. A cardiogenic mesoderm differentiates into blood cells or hemocytes in the hematopoietic organ called the lymph gland, while a head mesoderm anlage gives rise to circulating blood cells, known also as embryonic hemocytes.18,19 For both hematopoietic waves, multipotential hematopoietic progenitors are maintained and then differentiate subsequently into mature hemocyte cell types, called plasmatocytes, crystal cells, and lamellocytes.20–24 The Drosophila transcription factor Lozenge (Lz), which has homology to AML1/Runx1, is required for specification of crystal cells.19 Other Runx orthologs in Drosophila are expressed in the hematopoietic system, but their function has not yet been determined. In this study we analyzed the effect of human AML1-ETO fusion protein when expressed in the hemocyte precursors of the Drosophila embryonic blood compartment and identified genetic modifiers of the phenotype as potential effectors of the fusion protein.
We found that this oncogene induces expansion of hematopoietic precursors, which are characterized by high level of expression of reactive oxygen species (ROS). Among identified mutations in the modifier genetic screen, we found that nuclear receptors, including EcR-B1, are required for abnormal proliferation caused by AML1-ETO. Our studies revealed that negative regulation of ROS production by inactivation of EcR-B1 or activation of Foxo and its direct target, superoxide dismutase-2 (SOD2), substantially suppress generation of the aberrant hemocyte precursors induced by AML1-ETO. Our in vivo studies reveal tumor suppressor activity of Foxo and SOD2 in the context of AML1-ETO–induced pathology that warrants analysis in the context of the human disorder.
Methods
Genetics
UAS-RasV12, w1118, EcRq50st, EcRm554fc, Kr2, E2f 07 172, wnt4EMS23, UAS-SOD2, UAS-Foxo, UAS-Cat, UAS-EcR-ADN, UAS-EcR-AdsRNA, UAS-EcR-B1DN, UAS-EcR-ADN, UAS-Akt1, UAS-PtendsRNA, UAS-SOD2dsRNA, Drosophila deficiency kit and numerous alleles used in the genetic screen were obtained from the Bloomington Drosophila Stock Center. UAS-dsRNA lines of EcR-B1, err, Hnf4, and Hr96 were obtained from the NIG-FLY stock collection, and lines of E75C, E78B, DHR39, DHR4, DHR3, Akt1, and ND75 were obtained from the VDRC stock center. Dfz2 and fz alleles were received from R. Nusse, and S0429-38, S1364-03, S1364-03, and S0279-04 alleles were from Szeged stock center. For the modifier genetic screen, mutant P-element alleles were combined with CyO,Kr-Gal4,UAS-GFP or Tm6,Tb balancers to distinguish mutant chromosomes. For the screen the following fly stocks were generated: w; hmlΔ-Gal4, UAS-2xeGFP; UAS-AML1-ETO/Tm6,Tb. Crosses were maintained on standard Drosophila cornmeal/sucrose/yeast medium at 25° to 29°C. Larvae were imaged with a Zeiss SteREO Lumar.V12 Fluorescence Stereomicroscope (12× magnification, a Zeiss NeoLumar S 0.8 × FWD 80-mm objective lenses) and a Zeiss AxioCam HRC digital camera.
Expression of transgenic constructs
To produce upstream activation sequence (UAS) transgenic flies, cDNA of AML1-B, ETO, Lz, AML1-ETO and its mutant forms25,26 were transferred from pBluescript into the pUAST vector.27 To make transgenic flies, w1118 embryos were injected with pUAST constructs by BestGene, and transformed progeny were recovered by standard protocol.27 AML1-ETO mutant proteins remain stable during ectopic expression experiments.25,26 We used hmlΔ-Gal4 to express proteins in circulating hemocytes.28 For controls, larvae from crosses of hmlΔ-Gal4 and w1118 were used. Two independent insertions of each UAS-AML1-ETO mutant as well as UAS-dsRNA constructs were analyzed in each experiment.
Immunocytochemistry, hemocyte counts, ROS staining, and phagocytosis analysis
Antibodies against EcRB1 (Mab AD4.4),29 Runx1/AML1, P1 (anti-Nimrod), L1, peroxidasin (Pxn), and prophenoloxidase (PPO) were used.22 Nuclear DNA was visualized with the ToPro-3 dye (Invitrogen). Cy3-labeled anti–mouse and anti–rabbit secondary antibodies were obtained from Jackson ImmunoResearch Laboratories. Hemocyte immunostaining was performed as previously described,21 with use of Cy3-labeled secondary antibodies. To determine total hemocyte number in different mutant backgrounds, the circulating hemocytes from single larva were collected in 20 μL phosphate-buffered saline, while the lymph gland remained untouched. Cells were counted using a hemocytometer and calculated as number of cells per single larva. Cell numbers of 10 independent samples for each genotype were analyzed to ensure reproducibility. Expression of ROS was monitored with the fluorescent dye dihydroethidium (Invitrogen) as described previously.30 Phagocytosis in circulating hemocytes was monitored ex vivo with use of FluoSpheres (blue fluorescent 0.1-μm carboxylate-modified microsphere beads; Invitrogen). Briefly, hemocytes from single larvae were isolated in 30 mL Schneider medium into 14-well glass slides (Fisher Scientific) containing FluoSpheres (1:1000) and incubated for 15-20 minutes. Glass wells were gently washed once and processed for ROS staining. Cells were fixed with 4% formaldehyde for 5 minutes and mounted with Vectashield (Vector Laboratories) and imaged using a Bio-Rad Radiance 2000 confocal microscope (40× lenses and digital magnifications) or a Zeiss Axioskop 2 plus Fluorescence Stereomicroscope (40× lenses and digital magnification) with a Zeiss AxioCam HRC digital camera.
Results
AML1-ETO induces proliferation of hemocyte precursors in Drosophila larvae
To express human AML1 and AML1-ETO in Drosophila blood cells, we used the Gal4/UAS binary misexpression system.27 Transgenic fly strains were made that allow transcription of these transgenes under control of the UAS. To express these human proteins in Drosophila blood cells, we used transgenic flies that produce Gal4 transcription factor upon activation of the hemolectin (hml) promoter (hmlΔ-Gal4) or peroxidasin (pxn) promoter (pxn-Gal4; not shown). These drivers are activated during late embryonic and all larval stages in a majority of circulating hemocytes as well as hemocyte precursors.28,31 We found that AML1-ETO causes increased proliferation of circulating hemocytes, raising their numbers to 10 times the wild-type level (Figure 1A). In contrast, neither AML1-B or its Drosophila homolog Lz, or ETO had any similar overt effects on the proliferation of circulating hemocytes (Figure 1A). However some proliferative activity of AML1-B could be mediated because of its transcriptional repression features such as the presence of binding sites for Groucho, or due to interference with endogenous CBFβ. The hyperproliferation caused by AML1-ETO is primarily associated with a stimulating effect of this oncogene on the immature hemocyte precursors. We found a significant increase in the number of hemocyte precursors in larvae with AML1-ETO expression versus wild-type ones (Figure 1B-C). These immature precursors were identified as cells that do not express any differentiation marker, including nimrod (P1), Pxn, PPO, and L1 (Figure 1B). Noticeably, hemocytes from AML1-ETO mutant background tend to be relatively small and round in shape.
Figure 1.
AML1-ETO induces generation of ROS+, Wg+ hemocyte precursors, and increased proliferation of hemocytes in larval circulation. (A) AML1-ETO, AML1, ETO, and Lz were expressed under control of the hemocyte specific driver hmlΔ-Gal4, using Gal4/UAS system. Hemocytes from 3rd instar larva were extracted and counted for each genotype. hmlΔ-Gal4,UAS-GFP heterozygous were used as wild-type control (WT). AML1-ETO causes a robust increase in the number of hemocytes in larval circulation compared with wild-type and overexpression of the other proteins. (B) Hemocyte precursors, identified as cells that do not express maturation markers: P1, Pxn, PPO, and L1 (all green) are rarely seen in wild-type (rare examples shown by arrow), but are significantly elevated upon AML1-ETO expression (arrows). (C) Quantitation of the data in panel B (n = 10, P < .001). (D) Wg is highly expressed in rare precursor cells that are hml negative (arrowhead) in circulation of wild-type larvae (WT). Number of Wg+ hemocytes is significantly increased in AML1-ETO mutant background. (E) Quantitation of the data in panel D (n = 10, P < .001). (F) ROS is expressed in hmllow(1) and rarely at low levels in some hml+(2) hemocytes in circulation of wild-type larva (WT). Number of cells expressing high level of ROS is increased in mutant background (AML1-ETO). (G) Quantitation of the data in panel F (n = 10, P < .001). (H) ROS+ cells (indicated by arrowhead) in WT and AML1-ETO backgrounds fail to phagocytose FluoSpheres (FS, blue) microparticles, suggesting they are precursors. Hemocyte markers, ROS, ToPro-3, and microspheres dyes color-coded in left panels. Scale bars, 5 μm.
Wingless signaling is required for maintenance of the stem cell population in the hematopoietic organ called the lymph gland.32 In circulation of wild-type larvae, the number of such precursors is relatively low, and they also express Wg (Figure 1D). It is also a feature of these cells to express relatively low level of hml. We found that AML1-ETO causes a significant increase in the number of Wg+ hemocyte precursors, the majority of which tend to express low level of hml (Figure 1D-E). This is unlike the lymph gland precursors of the medullary zone that do not express hml. As another marker of stem cells, ROS are expressed at significantly higher levels in the hemocyte precursors of the larval hematopoietic organ, the lymph gland.30 Our analysis reveals that ROS is also differentially expressed in precursor cells in circulation; high ROS is seen in a small number of hml− cells found in wild-type and is not detectable in mature hml+ hemocytes (Figure 1F). Lack of phagocytic activity additionally indicates that the ROS+ hemocytes represent immature precursor populations in circulation (Figure 1H). Importantly, the number of ROS+ hemocyte precursors is significantly elevated in larvae expressing AML1-ETO in hemocytes (Figure 1F-H).
We also noticed an increase in the number of lamellocytes (L1+ cells) and crystal cells (Lz+, PPO+) as well as an occurrence of melanotic tumors (30%-40%) in larvae with expressing AML1-ETO (not shown). It is likely that the increased proliferation of the hemocyte precursors contributes to the formation of tumors, as has been seen when different signaling regulators, such as fly JAK kinase (HopTum−l) and HP1 protein are mutated.16,33 These tumors can be recognized by the innate immune system, which melanizes them as part of the cellular immune response.
Interaction of AML1-ETO with DNA, CBFβ, and components of the transcription repressor complex is crucial for hemocyte proliferation
In a mammalian cell, AML1-ETO acts as a constitutive repressor of AML1 target genes by recruiting a transcriptional repressor complex through its ETO domains12,13 (Figure 2A). It has been shown that interaction of the oncoprotein with proteins of the transcriptional repressor complex as well as with target DNA and CBFβ are essential for leukemia progression.25,34 In order for the Drosophila hematopoietic model to be useful as a system to study AML1-ETO, it is important to determine whether the essential characteristics of the protein as a transcriptional regulator are maintained. The R174Q mutation in AML1 abrogates its DNA binding,34 and the Y113A/T161A mutation diminishes its interaction with CBFβ.26 To assess the functional relevance of these domains to the observed phenotype in Drosophila, we expressed each of the mutated UAS-AML1-ETO constructs in hemocytes. We also confirmed that these mutant proteins are stable and localized in nuclei of hemocytes (supplemental Figure 1, available on the Blood Web site; see the Supplemental Materials link at the top of the online article). Unlike the wild-type oncogene, both constructs, encoding proteins compromised for binding to CBFβ or DNA, were significantly deficient in inducing overproliferation (Figure 2A-B). A construct that disrupts both the CBFβ- and DNA-binding (Y113A/T161A/ R174Q, YTR mutant) also failed to induce the overproliferation phenotype (Figure 2A-B). In addition, we also found that coexpressing an excess of Drosophila CBFβ proteins Brother or Big Brother (using UAS-Bro or UAS-Bgb) significantly suppresses AML1-ETO–induced proliferation probably due to depletion or sequestering of AML1-ETO away from the DNA (supplemental Figure 2).
Figure 2.
Functional domains of AML1-ETO are critical for its activity in Drosophila blood cells. (A) Schematic representation of the AML1-ETO domains and mutations. Interaction sites of AML1-ETO with DNA, CBFβ (light blue lines), and HDAC1-3, Sin3A, N-CoR/SMRT, ETO, and SON (dark blue lines) are indicated below the protein scheme. (B) UAS constructs encoding wild-type AML1-ETO and its mutant variants (indicated on panel) were expressed under control of hmlΔ-Gal4. hmlΔ-Gal4,UAS-GFP heterozygous were used as wild-type control (WT control). In contrast to intact AML1-ETO, mutant forms with altered DNA-binding (R174Q), CBFβ-binding [Y113A/T161A (YT)], or both DNA- and CBFβ-binding [R174Q/Y113A/T161A (YTR)] domains failed to induce proliferation of hemocytes (n = 10, P < .001). Truncated AML1-ETO(NHR3X) and (NHR2X) proteins were less active in inducing hemocyte proliferation than intact protein (n = 10, P < .001). The AML1-ETO(NHR2X542,m7) mutant with disrupted HHR, NHR3, and MYND domains was unable to induce hemocyte proliferation.
Truncated forms of AML1-ETO with both NHR3 and MYND domains deleted (NHR2X542) or the MYND domain deleted (NHR3X663)25 also result in a 50% reduction of hemocyte proliferation, compared with full-length protein (Figure 2A-B). Removal of the HHR (dimerization mutant m7), NHR3 and MYND domains (NHR2X542, m7) suppresses proliferation to near wild-type level (Figure 2A-B). These results strongly suggest that the proliferative response in Drosophila blood precursor cells requires the interaction of AML1-ETO with CBFβ and the target DNA sequence as well as with the transcriptional repressors through its HHR, NHR3, and MYND domains.
A genetic screen for modifiers of AML1-ETO phenotype
To identify functionally important genes that interact with the AML1-ETO oncogenic pathway, we designed a genetic screen, which allowed identification of single copy second site suppressor and enhancer alleles (Figure 3A). Fly stocks containing green fluorescent protein (GFP)–marked circulating hemocytes (hmlΔ-Gal4, UAS-GFP) expressing AML1-ETO were crossed with lethal mutations, placed over marked balancers, and progeny larvae were analyzed under a fluorescent microscope (Figure 3A). Screening a panel(231) of genomic deficiencies on Drosophila autosomes revealed 28 deficiencies that enhance AML1-ETO phenotype and 8 that suppress the phenotype, validating the concept that single-copy mutations can be identified. Testing available mutant alleles within some of the identified genomic intervals as well as an additional screen of approximately 1500 autosomal P-element insertional mutations revealed that alleles of Protein kinase-C 53E (Pkc53E), wnt4, frizzled (fz), Dfz2, Ryanodine receptor 44F (Rya-R44F), dissatisfaction (dsf), Another transcription unit (Atu), Bgb and the uncharacterized gene lethal-(2)06 496 (l(2)06 496) significantly enhance the phenotype, while mutations in Ecdysone receptor (EcR), sallimus (sls), Sin3A, Nurf38, Kruppel (Kr), e2f, male sex lethal-3 (msl-3), couch potato (cpo), and the uncharacterized genes CG5033 and S0279-04 decreased the AML1-ETO–induced phenotype (Table 1). As controls, heterozygous larvae of the identified mutations show wild-type hemocyte counts, indicating that the observed genetic interactions are specifically associated with the proliferative effect of AML1-ETO. Among the identified mutants, we can roughly classify at least 2 major groups of genes; one is associated with steroid receptor signaling (EcR, Pkc53E, sls, Sin3A, Nurf38) and the other related to Wnt4/Fz and calcium signaling (Pkc53E, wnt4, fz, Dfz2, Rya-R44F). Herein we have primarily focused on investigating the AML1-ETO interaction with the ecdysone signaling pathway.
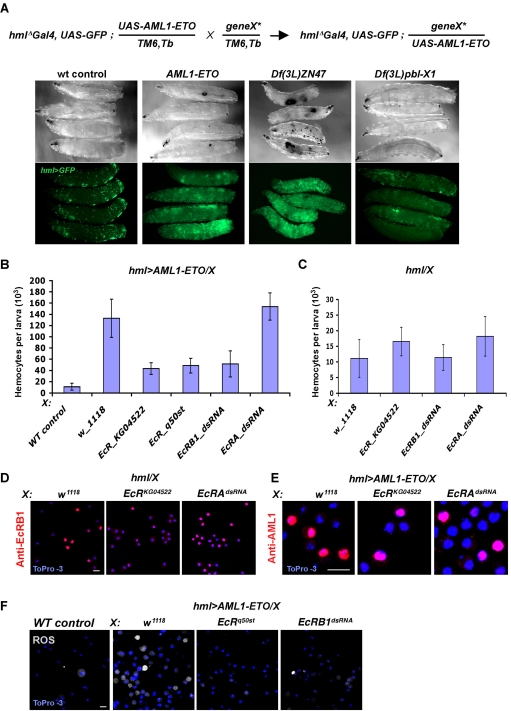
Figure 3.
A genetic screen identified ecdysone receptors as modifiers of AML1-ETO induced blood disorder. (A) Schema of modifier genetic screen: flies of synthesized genetic background containing hmlΔ-Gal4,UAS-GFP;UAS-AML1-ETO/TM6,Tb were crossed with mutant alleles. Larvae from these crosses were scored under direct (top panels) and fluorescent (bottom panels) illumination (Zeiss SteREO Lumar.V12 Stereomicroscope, 12× magnification); from left to right: normal density of GFP+ hemocytes in wild-type (WT) larvae of hmlΔ-Gal4,UAS-GFP background. Numbers of GFP+ hemocytes and black melanotic tumors are robustly elevated in hmlΔ-Gal4,UAS-GFP;UAS-AML1-ETO larvae (AML1-ETO). Examples of enhancer and suppressor modifiers of AML1-ETO phenotype: single copy deficiencies [Df(3L)ZN47] or [Df(3L)pbl-X1] cause a dramatic increase and decrease, respectively, in the number of circulating hemocytes and melanotic tumors in hmlΔ-Gal4, UAS-GFP; UAS-AML1-ETO larvae. These deficiencies include hundreds of genes, a majority of which remain yet uncharacterized. Analysis of available mutant alleles of genes belonging to these genomic intervals has not yet identified single loci that can modify the phenotype. (B) EcR-B1 is required for induction of hemocyte proliferation by AML1-ETO. EcR-B1 and EcR-A were inactivated in hemocytes of hmlΔ-Gal4,UAS-GFP;UAS-AML1-ETO background (hml > AML1-ETO). AML1-ETO induced proliferation of hemocytes was dramatically suppressed by EcR hemizygosity (alleles KG04522, q50st) and dsRNA of EcR-B1. In contrast,the oncogene induced proliferation of hemocytes was very mildly increased by dsRNA allele of EcR-A. hmlΔ-Gal4,UAS-GFP heterozygous were used as wild-type control (WT control). (C) As controls for (B), hemizygosity for EcR-B1 or inactivation of either EcR-B1 or EcR-A with corresponding dsRNAs in normal hemocytes [hmlΔ-Gal4,UAS-GFP(hml)] does not affect proliferation of otherwise hemocytes. (D) Inactivation of EcR-A causes increase in circulating hemocytes expressing EcR-B1. EcR-B1 is highly expressed in nuclei of a population of circulating hemocytes in wild-type larvae (hml/w1118). Expression of EcR-B1 is significantly reduced in EcR KG04522 heterozygous (hml/EcRKG04522) animals, while the number of cells expressing high level of EcR-B1 is significantly increased upon dsRNA-mediated inactivation of EcR-A (hml/EcR-AdsRNA). (E) Nuclear localization of AML1-ETO in hemocytes is not affected by deficiency of EcR-B1 (EcR KG04522) or EcR-A (EcR-AdsRNA) in AML1-ETO mutant background. (G) Number of ROS+ hemocyte precursors is significantly reduced upon inactivation of EcR-B1 in AML1-ETO mutant larvae (hml > AML1-ETO). Abbreviations of genotypes indicated on top of each panel, cell markers are color coded. Scale bars, 5 μm.
Table 1.
Genes identified in AML1-ETO modifier genetic screen
| Names of genes | Names of alleles | Molecular function | Effect on AML1-ETO phenotype |
|---|---|---|---|
| Suppressors | Suppression of phenotype | ||
| EcR | EcRKG04522, EcRQ50st | Nuclear receptor | Strong |
| msl-3 | msl-3d01070, msl-31 | Chromatin and RNA binding | Moderate |
| cpo | cpoBG02810 | mRNA binding | Moderate |
| Kr | Kr2 | Transcriptional repressor | Strong |
| E2f | E2f07172 | Transcription factor | Moderate |
| CG5033 | CG5033EY04217 | Ribonucleoprotein binding | Moderate |
| sls | slsKG00981 | Myosin light chain kinase | Strong |
| Sin3A | Sin3A08269, RNAi | Transcriptional corepressor | Moderate |
| Nurf38 | Nurf38k16102, RNAi | Chromatin remodeling | Strong |
| S0279-04 | Unknown | Moderate | |
| Enhancers | Enhancement of phenotype | ||
| wnt4 | wnt4EMS23, wnt4C1 | Ligand of Frizzled receptors | Strong |
| fz | fzJB | Frizzled receptor | Strong |
| Dfz2 | Df(3R)469-2, Dfz2DN | Frizzled receptor | Strong/moderate |
| Rya-R44F | Rya-R44FEY12439 | Calcium ion transport receptor | Moderate |
| Atu | Atus1938 | Transcription cofactor | Strong |
| Pkc53E | Pkc53EEY14093 | Protein kinase C activity | Moderate |
| Bgb | BgbD | CBFβ homolog | Moderate |
| Dsf | dsfMB03062 | Nuclear receptor | Moderate |
| l(2)06496 | l(2)06496k14618 | Component of microtubule | Strong |
| S0429-38 | Unknown | Strong | |
| S1364-03 | Unknown | Strong | |
| S1364-03 | Unknown | Strong |
Ecdysone nuclear receptors are involved in AML1-ETO activity in hemocytes
The Ecdysone receptor (EcR) locus encodes 3 isoforms of the receptor, which differ in their N-terminal domains. EcR-A and EcR-B1 are differentially expressed during larval development, where they often function in a nonredundant manner.29,35 Analysis of the EcR alleles revealed that hemizygous EcR-B1 mutations are responsible for suppression of the AML1-ETO phenotype, while alleles that primarily affect the EcR-A isoform tend to enhance the AML1-ETO phenotype (Figure 3B and supplemental Figure 3). In addition, dsRNA and dominant negative alleles of EcR isoform B1 or A cause a strong suppression and mild enhancement (P = .095) of the phenotype, respectively (Figure 3B and supplemental Figure 3). EcR-B1 mutations cause a 3-fold reduction in hemocyte number compared with the AML1-ETO parental background (P < .001; Figure 3B). Thus the 2 isoforms have different and likely opposing interaction in the context of their involvement in the AML1-ETO pathway. We also found that inactivation of the EcR-A isoform up-regulates the expression of EcR-B1 in normal hemocytes, suggesting that the phenotype observed upon loss of EcR-A could, in fact, be indirectly mediated through the up-regulation of EcR-B1 (Figure 3D). Importantly, these interactions are specific for AML1-ETO pathway because inactivation of these receptors did not affect proliferation of wild-type hemocytes (Figure 3C and supplemental Figure 3). Also, mutant alleles of EcRB1 do not alter hopTum−l blood phenotype (supplemental Figure 4). Larvae overexpressing AML1-ETO in hemocytes develop with a normal time cycle indicating that Ecdysone pulses in these larvae are intact. EcR-B1 or EcR-A mutations do not affect the nuclear localization of AML1-ETO in hemocytes (Figure 3E). Importantly, we noticed that EcR-B1 deficiency causes a significant reduction in the number of ROS+ precursors in the AML1-ETO mutant (Figure 3F). The exact relationship between AML1-ETO and EcR remains unclear and requires detailed biochemical analysis beyond the scope of this in vivo genetic study.
SOD2 activation by Foxo blocks generation of the AML1-ETO–induced ROS+ precursors
As EcR-B1–mediated suppression of AML1-ETO phenotype is associated with a reduction in ROS+ hemocyte precursors, we asked if inactivation of ROS production in AML1-ETO–induced hemocyte precursors would suppress their hyperproliferation. To test this concept we expressed ROS scavenger enzymes SOD2 and Catalase (Cat) in AML1-ETO–expressing hemocytes. SOD2 robustly, and Cat to a lesser extent, suppress the AML1-ETO–induced hemocyte proliferation (Figure 4A). In addition, expression of Foxo, a direct positive regulator of SOD2 and Cat,36 almost completely suppressed the AML1-ETO phenotype (Figure 4A,C). Foxo activation is negatively controlled by the PI3kinase/Akt1 signaling, and dsRNA-mediated inactivation of Akt1 was also found to suppress the phenotype (Figure 4A). Importantly, overexpression of Foxo or SOD2 did not significantly affect proliferation of circulating hemocytes in a wild-type background (Figure 4B). Taken together, our data suggest that AML1-ETO–induced generation of ROS+ hemocyte precursors can be suppressed by reducing the ROS levels.
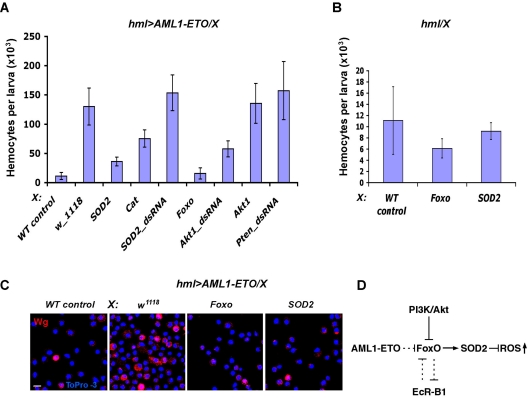
Figure 4.
SOD2 activation by Foxo suppresses generation of AML1-ETO–induced precursors. (A) Activation of SOD2 or Catalase or Foxo, or inactivation of Akt1 causes suppression of AML1-ETO–mediated hemocyte proliferation. Corresponding UAS constructs (indicated on x-axis) were expressed in hemocytes expressing AML1-ETO (hmlΔ-Gal4,UAS-GFP;UAS-AML1-ETO: hml > AML1-ETO). Number of hemocytes in AML1-ETO mutant was significantly reduced by ectopic expression of Foxo, AktdsRNA, SOD2, or Catalase (n = 10, P < .001). Number of hemocytes in AML1-ETO mutant was not significantly affected by overexpression of Akt1, ptendsRNA, SOD2dsRNA. hmlΔ-Gal4,UAS-GFP heterozygous were used as wild-type control (WT control). (B) In a wild-type background, the number of hemocytes is not significantly affected by overexpression of SOD2 or Foxo with hmlΔ-Gal4,UAS-GFP. (C) Activation of SOD2 or Foxo causes significant reduction of Wg+ hemocyte precursors in AML1-ETO mutant. SOD2 or Foxo were expressed in hemocytes expressing AML1-ETO (hml > AML1-ETO). Scale bars, 5 μm. (D) A schematic diagram depicting the relationship between PI3K/Akt and EcR-B1 pathways in negative regulation of FoxO that is required for positive regulation of ROS-inactivating enzymes. We propose that AML1-ETO suppresses FoxO function, leading to an increase of ROS in hemocyte precursors.
Discussion
The AML1-ETO fusion protein is implicated in the etiology of AML in humans. Although this oncogene is one of the most extensively investigated in mouse models, many questions of its function and effective therapy remain to be elucidated. Here we used the advantages of the Drosophila genetic system for an in vivo analysis that allowed us to identify new genes and components that function in the context of AML1-ETO. The genetic model recapitulates basic properties of the fusion protein in that domains for interaction with target promoter DNA and CBFβ as well as various functional motifs within the ETO domain are critical in inducing the expansion of hemocyte precursors in Drosophila. It seems likely that AML1-ETO functions as an antagonistic constitutive repressor form of Drosophila Runx factors.12,37 Despite these conserved features, our analysis did reveal some differences in activity between the Drosophila and mouse systems. The truncated AML1-ETO-NHR2X542 form is less active in inducing the phenotype in Drosophila, while the comparable AML11-ETO9a isoform is highly leukemogenic in mouse models.38 This difference could be attributed to the absence of a SON ortholog in flies, which has been shown to suppress the leukemic activity of the oncogene via an interaction with the MYND domain.10
Drosophila has a low redundancy in gene function, and under conditions of sensitized backgrounds, genes that are not otherwise haploinsufficient in the wild-type fly can be identified as single-copy enhancers and suppressors. This allowed the identification of opposing phenotypes due to EcR-B1 and EcR-A nuclear receptor loss in the context of the AML1-ETO pathway. This result is consistent with the opposite activities of the EcR isoforms in various cell contexts in regulating gene transcription.29,35 Our studies revealed that E78B and E75C nuclear receptors, which are transcriptional targets of ecdysone signaling,39 also significantly suppress the AML1-ETO phenotype (supplemental Figure 5). Interestingly, we also found that functions of Hnf4, err, and Hr96 nuclear receptors are implicated in the progression of the AML1-ETO phenotype as well (supplemental Figure 5). The genetic interaction of the AML1-ETO and nuclear receptors is consistent with the known associations of human vitamin D, androgen and estrogen nuclear receptors, and Runx proteins in modulating their transcriptional activity.40–42 AML1-ETO and nuclear receptor complexes can also interact through corepressor proteins, such as NcoR/SMRT, HDACs, and Sin3A. It is well known that deregulation in retinoid acid signaling or vitamin D signaling contribute to pathogenesis of leukemia in humans.42,43 Moreover recent studies revealed antileukemic activity of corticosteroid compounds in human AML1-ETO–induced leukemia cell lines.44 The human liver X receptor (LXR), an ortholog of EcR, is involved in maturation and function of myeloid dendritic cells.45 Although the hematopoietic function of the other identified nuclear receptors and their human orthologs has not yet been evaluated, our studies suggest they may be involved in development and/or function of myeloid cells.
In mouse models, AML1-ETO suppresses myeloid differentiation by increased self-renewal of progenitors.6 Similarly, in Drosophila AML1-ETO alters the differentiation program, inducing the generation of excess hematopoietic precursors. Interestingly, these precursor cells express high level of ROS, and the production of ROS+ precursors is critical for the progression of the AML1-ETO–induced proliferative disorder. Suppression of this phenotype by EcR-B1 loss was attributed to lower numbers of ROS+ precursors. This led to the finding that a reduction of ROS production by SOD2 and Cat enzymes as well as by overexpression of Foxo, which activates expression of these enzymes, substantially suppresses the AML1-ETO–mediated expansion of the precursors. This indicates that ROS plays a central role as a signaling factor in maintaining proliferation of these abnormal hemocyte precursors. In addition, elevated ROS level by the disruption of complex I protein (ND75) of the mitochondrial electron transport chain induces significant proliferation of hemocytes (supplemental Figure 6).30 It has been shown that FoxO-dependent regulation of ROS production is critical for maintenance of hematopoietic stem cells in mammals.36 Murine multipotent progenitors and common myeloid progenitors maintain 100× higher ROS level than hematopoietic stem cells, although the functional significance of high ROS production in myelopoiesis has not yet been evaluated.46 It was recently shown, however, that high ROS levels are critical for promoting human leukemia stem cells, but a certain threshold increase in ROS concentration mediates their terminal differentiation.47,48 In Drosophila, a tight regulation of ROS level is required for maintenance of normal stem cell populations in the lymph gland. These moderate levels of ROS are necessary for the progenitors to be competent to differentiate, and higher ROS levels triggers a differentiation program through the activation of the JNK pathway.30 ROS therefore functions dually for specification of these myeloid precursors as well as oxidative stress sensing.
Elevated levels of ROS associated with active Ras/MAPK and PI3K pathways have also been detected in leukemic cells in humans.49,50 Inactivated FoxO by Flt3-mediated PI3Kinase/Akt signaling was also observed in 30% cases of AML.49 FoxO transcription factors play diverse physiologic roles in different cellular contexts in mammals and Drosophila, including apoptosis, cell-cycle regulation, and oxidative stress resistance.51 Our data indicate that in a context of malignant Drosophila blood precursors, Foxo acts downstream of the PI3Kinase/Akt pathway as a positive regulator of SOD2 and Cat expression. On the other hand, it is well established that FoxO proteins interact with several nuclear receptors leading to changes in the transcriptional activity of both proteins.52 It has been also documented that ecdysone signaling regulates stabilization of Foxo in Drosophila,53 as well as it controls expression of ROS inactivated enzymes including Catalase and MsrA.54,55 The identified suppression of the AML1-ETO phenotype by inactivation of EcR-B1 or activation of Foxo suggests that both genes could act together to regulate expression of the antioxidant enzymes (Figure 4D). Thus, our studies suggest that ROS is a signaling factor promoting maintenance of the myeloid precursor cells in the Drosophila AML1-ETO model and highlights the importance of FoxO, SOD2, and nuclear receptors as targets for further study in the context of the human disease.
Supplementary Material
Acknowledgments
T.H., T.M., S.S., and Q.-M.T. participated as undergraduate students in the UCLA Undergraduate Research Consortium in Functional Genomics, supported by a Howard Hughes Medical Institute Professor's Award to U.B. We thank J. M. Olson, G. B. Call, I. E. Clark, C. Taber, K. Topp, and E. Owusu-Ansah for assistance during the genetic screening and critical comments during these studies. We thank members of the Banerjee laboratory and I. E. Clark for comments on the manuscript. We thank C. J. Evans for assistance in phagocytosis analysis. We acknowledge R. Nusse for fz and dFz2 alleles, J. H. Fessler and I. Ando for antibodies, and J. Canon for UAS-AML1-ETO, UAS-AML1 constructs. Due to space limitations, we apologize that not all studies were referenced. We are grateful to the Bloomington Drosophila stock center, NIG-FLY stock collection, Szeged stock center, and VDRC dsRNA stock center. We thank the Developmental Studies Hybridoma Bank at the University of Iowa for making accessible anti-EcR monoclonal antibodies. We thank the University of California, Los Angeles for providing support and infrastructure for this project.
This work was supported by a National Institutes of Health (NIH) grant (5R01 HL067395) and a seed grant from the Johnson Comprehensive Cancer Center at UCLA to U.B.
Footnotes
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Authorship
Contribution: S.A.S. and U.B. developed the project; S.A.S. conceived, designed, and performed the experiments; S.A.S., T.H., T.M., Q.-M.T., and S.S. performed the genetic screens; M.D.C. and N.A.S. designed and made UAS constructs of mutant forms of AML1-ETO; and S.A.S. and U.B. discussed the results and wrote the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Utpal Banerjee, Department of Molecular, Cell, and Developmental Biology, Molecular Biology Institute, Department of Biological Chemistry, Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, University of California at Los Angeles, 621 Charles E. Young Dr S, Life Sciences Bldg, Rm 2204, Los Angeles, CA 90095; e-mail: banerjee@mbi.ucla.edu.
References
- 1.Canon J, Banerjee U. Runt and Lozenge function in Drosophila development. Semin Cell Dev Biol. 2000;11(5):327–336. doi: 10.1006/scdb.2000.0185. [DOI] [PubMed] [Google Scholar]
- 2.Ito Y. RUNX genes in development and cancer: regulation of viral gene expression and the discovery of RUNX family genes. Adv Cancer Res. 2008;99:33–76. doi: 10.1016/S0065-230X(07)99002-8. [DOI] [PubMed] [Google Scholar]
- 3.Kalev-Zylinska ML, Horsfield JA, Flores MV, et al. Runx1 is required for zebrafish blood and vessel development and expression of a human RUNX1-CBF2T1 transgene advances a model for studies of leukemogenesis. Development. 2002;129(8):2015–2030. doi: 10.1242/dev.129.8.2015. [DOI] [PubMed] [Google Scholar]
- 4.de Bruijn MF, Speck NA. Core-binding factors in hematopoiesis and immune function. Oncogene. 2004;23(24):4238–4248. doi: 10.1038/sj.onc.1207763. [DOI] [PubMed] [Google Scholar]
- 5.Watanabe-Okochi N, Kitaura J, Ono R, et al. AML1 mutations induced MDS and MDS/AML in a mouse BMT model. Blood. 2008;111(8):4297–4308. doi: 10.1182/blood-2007-01-068346. [DOI] [PubMed] [Google Scholar]
- 6.Peterson LF, Zhang DE. The 8;21 translocation in leukemogenesis. Oncogene. 2004;23(24):4255–4262. doi: 10.1038/sj.onc.1207727. [DOI] [PubMed] [Google Scholar]
- 7.Hug BA, Lazar MA. ETO interacting proteins. Oncogene. 2004;23(24):4270–4274. doi: 10.1038/sj.onc.1207674. [DOI] [PubMed] [Google Scholar]
- 8.Wildonger J, Mann RS. Evidence that nervy, the Drosophila homolog of ETO/MTG8, promotes mechanosensory organ development by enhancing Notch signaling. Dev Biol. 2005;286(2):507–520. doi: 10.1016/j.ydbio.2005.08.026. [DOI] [PubMed] [Google Scholar]
- 9.Moore AC, Amann JM, Williams CS, et al. Myeloid translocation gene family members associate with T-cell factors (TCFs) and influence TCF-dependent transcription. Mol Cell Biol. 2008;28(3):977–987. doi: 10.1128/MCB.01242-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ahn EY, Yan M, Malakhova OA, et al. Disruption of the NHR4 domain structure in AML1-ETO abrogates SON binding and promotes leukemogenesis. Proc Natl Acad Sci U S A. 2008;105(44):17103–17108. doi: 10.1073/pnas.0802696105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Miyoshi H, Kozu T, Shimizu K, et al. The t(8;21) translocation in acute myeloid leukemia results in production of an AML1-MTG8 fusion transcript. EMBO J. 1993;12(7):2715–2721. doi: 10.1002/j.1460-2075.1993.tb05933.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yergeau DA, Hetherington CJ, Wang Q, et al. Embryonic lethality and impairment of haematopoiesis in mice heterozygous for an AML1-ETO fusion gene. Nat Genet. 1997;15(3):303–306. doi: 10.1038/ng0397-303. [DOI] [PubMed] [Google Scholar]
- 13.Wildonger J, Mann RS. The t(8;21) translocation converts AML1 into a constitutive transcriptional repressor. Development. 2005;132(10):2263–2272. doi: 10.1242/dev.01824. [DOI] [PubMed] [Google Scholar]
- 14.Okuda T, Cai Z, Yang S, et al. Expression of a knocked-in AML1-ETO leukemia gene inhibits the establishment of normal definitive hematopoiesis and directly generates dysplastic hematopoietic progenitors. Blood. 1998;91(9):3134–3143. [PubMed] [Google Scholar]
- 15.Shimada H, Ichikawa H, Nakamura S, et al. Analysis of genes under the downstream control of the t(8;21) fusion protein AML1-MTG8: overexpression of the TIS11b (ERF-1, cMG1) gene induces myeloid cell proliferation in response to G-CSF. Blood. 2000;96(2):655–663. [PubMed] [Google Scholar]
- 16.Evans CJ, Sinenko SA, Mandal L, et al. Advances in Developmental Biology. Vol 18. Oxford United Kingdom: Elsevier; 2007. Genetic dissection of hematopoiesis using Drosophila as a model system. p. 259. [Google Scholar]
- 17.Martinez-Agosto JA, Mikkola HK, Hartenstein V, Banerjee U. The hematopoietic stem cell and its niche: a comparative view. Genes Dev. 2007;21(23):3044–3060. doi: 10.1101/gad.1602607. [DOI] [PubMed] [Google Scholar]
- 18.Holz A, Bossinger B, Strasser T, Janning W, Klapper R. The two origins of hemocytes in Drosophila. Development. 2003;130(20):4955–4962. doi: 10.1242/dev.00702. [DOI] [PubMed] [Google Scholar]
- 19.Lebestky T, Chang T, Hartenstein V, Banerjee U. Specification of Drosophila hematopoietic lineage by conserved transcription factors. Science. 2000;288(5463):146–149. doi: 10.1126/science.288.5463.146. [DOI] [PubMed] [Google Scholar]
- 20.Markus R, Laurinyecz B, Kurucz E, et al. Sessile hemocytes as a hematopoietic compartment in Drosophila melanogaster. Proc Natl Acad Sci U S A. 2009;106(12):4805–4809. doi: 10.1073/pnas.0801766106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sinenko SA, Kim EK, Wynn R, et al. Yantar, a conserved arginine-rich protein is involved in Drosophila hemocyte development. Dev Biol. 2004;273(1):48–62. doi: 10.1016/j.ydbio.2004.05.022. [DOI] [PubMed] [Google Scholar]
- 22.Jung SH, Evans CJ, Uemura C, Banerjee U. The Drosophila lymph gland as a developmental model of hematopoiesis. Development. 2005;132(11):2521–2533. doi: 10.1242/dev.01837. [DOI] [PubMed] [Google Scholar]
- 23.Lanot R, Zachary D, Holder F, Meister M. Postembryonic hematopoiesis in Drosophila. Dev Biol. 2001;230(2):243–257. doi: 10.1006/dbio.2000.0123. [DOI] [PubMed] [Google Scholar]
- 24.Minakhina S, Steward R. Hematopoietic stem cells in Drosophila. Development. 2010;137(1):27–31. doi: 10.1242/dev.043943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu Y, Chen W, Gaudet J, et al. Structural basis for recognition of SMRT/N-CoR by the MYND domain and its contribution to AML1/ETO's activity. Cancer Cell. 2007;11(6):483–497. doi: 10.1016/j.ccr.2007.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Roudaia L, Cheney MD, Manuylova E, et al. CBFbeta is critical for AML1-ETO and TEL-AML1 activity. Blood. 2009;113(13):3070–3079. doi: 10.1182/blood-2008-03-147207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993;118(2):401–415. doi: 10.1242/dev.118.2.401. [DOI] [PubMed] [Google Scholar]
- 28.Sinenko SA, Mathey-Prevot B. Increased expression of Drosophila tetraspanin, Tsp68C, suppresses the abnormal proliferation of ytr-deficient and Ras/Raf-activated hemocytes. Oncogene. 2004;23(56):9120–9128. doi: 10.1038/sj.onc.1208156. [DOI] [PubMed] [Google Scholar]
- 29.Talbot WS, Swyryd EA, Hogness DS. Drosophila tissues with different metamorphic responses to ecdysone express different ecdysone receptor isoforms. Cell. 1993;73(7):1323–1337. doi: 10.1016/0092-8674(93)90359-x. [DOI] [PubMed] [Google Scholar]
- 30.Owusu-Ansah E, Banerjee U. Reactive oxygen species prime Drosophila haematopoietic progenitors for differentiation. Nature. 2009;461(7263):537–541. doi: 10.1038/nature08313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Stramer B, Wood W, Galko MJ, et al. Live imaging of wound inflammation in Drosophila embryos reveals key roles for small GTPases during in vivo cell migration. J Cell Biol. 2005;168(4):567–573. doi: 10.1083/jcb.200405120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sinenko SA, Mandal L, Martinez-Agosto JA, Banerjee U. Dual role of wingless signaling in stem-like hematopoietic precursor maintenance in Drosophila. Dev Cell. 2009;16(5):756–763. doi: 10.1016/j.devcel.2009.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Minakhina S, Steward R. Melanotic mutants in Drosophila: pathways and phenotypes. Genetics. 2006;174(1):253–263. doi: 10.1534/genetics.106.061978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yan M, Ahn EY, Hiebert SW, Zhang DE. RUNX1/AML1 DNA binding domain and ETO/MTG8 NHR2 dimerization domain are critical to AML1-ETO9a leukemogenesis. Blood. 2009;113(4):883–886. doi: 10.1182/blood-2008-04-153742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mouillet JF, Henrich VC, Lezzi M, Vogtli M. Differential control of gene activity by isoforms A, B1 and B2 of the Drosophila ecdysone receptor. Eur J Biochem. 2001;268(6):1811–1819. [PubMed] [Google Scholar]
- 36.Tothova Z, Kollipara R, Huntly BJ, et al. FoxOs are critical mediators of hematopoietic stem cell resistance to physiologic oxidative stress. Cell. 2007;128(2):325–339. doi: 10.1016/j.cell.2007.01.003. [DOI] [PubMed] [Google Scholar]
- 37.Osman D, Gobert V, Ponthan F, Heidenreich O, Haenlin M, Waltzer L. A Drosophila model identifies calpains as modulators of the human leukemogenic fusion protein AML1-ETO. Proc Natl Acad Sci U S A. 2009;106(29):12043–12048. doi: 10.1073/pnas.0902449106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yan M, Burel SA, Peterson LF, et al. Deletion of an AML1-ETO C-terminal NcoR/SMRT-interacting region strongly induces leukemia development. Proc Natl Acad Sci U S A. 2004;101(49):17186–17191. doi: 10.1073/pnas.0406702101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.King-Jones K, Thummel CS. Nuclear receptors—a perspective from Drosophila. Nat Rev Genet. 2005;6(4):311–323. doi: 10.1038/nrg1581. [DOI] [PubMed] [Google Scholar]
- 40.Khalid O, Baniwal SK, Purcell DJ, et al. Modulation of Runx2 activity by estrogen receptor-alpha: implications for osteoporosis and breast cancer. Endocrinology. 2008;149(12):5984–5995. doi: 10.1210/en.2008-0680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Baniwal SK, Khalid O, Sir D, Buchanan G, Coetzee GA, Frenkel B. Repression of Runx2 by androgen receptor (AR) in osteoblasts and prostate cancer cells: AR binds Runx2 and abrogates its binding to DNA. Mol Endocrinol. :2009. doi: 10.1210/me.2008-0470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Puccetti E, Obradovic D, Beissert T, et al. AML-associated translocation products block vitamin D(3)-induced differentiation by sequestering the vitamin D(3) receptor. Cancer Res. 2002;62(23):7050–7058. [PubMed] [Google Scholar]
- 43.Drumea K, Yang ZF, Rosmarin A. Retinoic acid signaling in myelopoiesis. Curr Opin Hematol. 2008;15(1):37–41. doi: 10.1097/MOH.0b013e3282f20a9c. [DOI] [PubMed] [Google Scholar]
- 44.Corsello SM, Roti G, Ross KN, et al. Identification of AML1-ETO modulators by chemical genomics. Blood. 2009;113(24):6193–6205. doi: 10.1182/blood-2008-07-166090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Bensinger SJ, Tontonoz P. Integration of metabolism and inflammation by lipid-activated nuclear receptors. Nature. 2008;454(7203):470–477. doi: 10.1038/nature07202. [DOI] [PubMed] [Google Scholar]
- 46.Tothova Z, Gilliland DG. FoxO transcription factors and stem cell homeostasis: insights from the hematopoietic system. Cell Stem Cell. 2007;1(2):140–152. doi: 10.1016/j.stem.2007.07.017. [DOI] [PubMed] [Google Scholar]
- 47.Abdel-Wahab O, Levine RL. Metabolism and the leukemic stem cell. J Exp Med. 2010;207(4):677–680. doi: 10.1084/jem.20100523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Callens C, Coulon S, Naudin J, et al. Targeting iron homeostasis induces cellular differentiation and synergizes with differentiating agents in acute myeloid leukemia. J Exp Med. 2010;207(4):731–750. doi: 10.1084/jem.20091488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Brandts CH, Sargin B, Rode M, et al. Constitutive activation of Akt by Flt3 internal tandem duplications is necessary for increased survival, proliferation, and myeloid transformation. Cancer Res. 2005;65(21):9643–9650. doi: 10.1158/0008-5472.CAN-05-0422. [DOI] [PubMed] [Google Scholar]
- 50.Zhu QS, Xia L, Mills GB, Lowell CA, Touw IP, Corey SJ. G-CSF induced reactive oxygen species involves Lyn-PI3-kinase-Akt and contributes to myeloid cell growth. Blood. 2006;107(5):1847–1856. doi: 10.1182/blood-2005-04-1612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Arden KC. FOXO animal models reveal a variety of diverse roles for FOXO transcription factors. Oncogene. 2008;27(16):2345–2350. doi: 10.1038/onc.2008.27. [DOI] [PubMed] [Google Scholar]
- 52.van der Vos KE, Coffer PJ. FOXO-binding partners: it takes two to tango. Oncogene. 2008;27(16):2289–2299. doi: 10.1038/onc.2008.22. [DOI] [PubMed] [Google Scholar]
- 53.Colombani J, Bianchini L, Layalle S, et al. Antagonistic actions of ecdysone and insulins determine final size in Drosophila. Science. 2005;310(5748):667–670. doi: 10.1126/science.1119432. [DOI] [PubMed] [Google Scholar]
- 54.Roesijadi G, Rezvankhah S, Binninger DM, Weissbach H. Ecdysone induction of MsrA protects against oxidative stress in Drosophila. Biochem Biophys Res Commun. 2007;354(2):511–516. doi: 10.1016/j.bbrc.2007.01.005. [DOI] [PubMed] [Google Scholar]
- 55.Radyuk SN, Klichko VI, Orr WC. Catalase expression in Drosophila melanogaster is responsive to ecdysone and exhibits both transcriptional and post-transcriptional regulation. Arch Insect Biochem Physiol. 2000;45(2):79–93. doi: 10.1002/1520-6327(200010)45:2<79::AID-ARCH4>3.0.CO;2-Y. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.