Abstract
We use classical molecular dynamics and sixteen combinations of force fields and water models to simulate a protein crystal observed by room-temperature X-ray diffraction. The high resolution of the diffraction data (0.96Å) and the simplicity of the crystallization solution (nearly pure water) makes it possible to attribute any inconsistencies between the crystal structure and our simulations to artifacts of the models rather than inadequate representation of the crystal environment or uncertainty in the experiment. All simulations were extended for 100ns of production dynamics, permitting some long-timescale artifacts of each model to emerge. The most noticeable effect of these artifacts is a model-dependent drift in the unit cell dimensions, which can become as large as 5% in certain force fields; the underlying cause is the replacement of native crystallographic contacts with non-native ones, which can occur with heterogeneity (loss of crystallographic symmetry) in simulations with some force fields. We find that the AMBER FF99SB force field maintains a lattice structure nearest that seen in the X-ray data, and produces the most realistic atomic fluctuations (by comparison to crystallographic B-factors) of all the models tested. We find that the choice of water model has a minor effect in comparison to the choice of protein model. We also identify a number of artifacts that occur throughout all of the simulations: excessive formation of hydrogen bonds or salt bridges between polar groups and loss of hydrophobic interactions. This study is intended as a foundation for future work that will identify individual parameters in each molecular model that can be modified to improve their representations of protein structure and thermodynamics.
Keywords: molecular dynamics, protein structure, crystal lattice, force field
1 Introduction
Molecular dynamics (MD) simulations of proteins are routinely performed as adjuncts to biochemical studies,1 in lead finding and analysis for drug design, 2 and as a lens for examining protein folding and function.3,4,5 The success of molecular models in these applications indicates the great potential for computation in biophysical problems, but these successes must be taken in the context of many other results in which the calculated chemical properties were not converged or were only qualitatively correct.6,7,8,9 Improvements in computing hardware continue to extend the range of molecular simulations, increasing statistical certainty in the calculated properties to produce commensurate increases in researchers’ ability to eliminate approximations and discriminate between model parameters.
Rigorous validation of the molecular models (force fields) has been performed for amino acid side chain analogs, using distributed computing to obtain converged solvation energies with a number of popular force fields and water models. 10,11 Solvation energies of biological molecules are a crucial property to treat correctly because they are integral to the energetics of ligand binding and biomolecular recognition. Furthermore, the ability of particular models to predict trends in solvation energy for most of the 20 natural amino acids is an indication that the models are producing the correct energies because they are physically reasonable, not merely over-fitted. Details of the ensemble of conformations adopted by individual amino acids in solution can also be verified by NMR experiments; 12 an accurate reproduction of the conformations of many amino acids packed in close proximity is needed to simulate proteins.
Another strategy for testing detailed structural characteristics of many amino acids interacting in close proximity is to simulate protein crystals, which are up to 70% polypeptide by volume. 13,15,14,16 Although the complex structure of each protein component in the crystal lattice presents a challenge of adequately sampling its relevant conformations, protein crystal simulations offer some advantage in that multiple independent copies of the unique asymmetric unit (usually a protein monomer) may be simulated simultaneously in the unit cell, without the burden of simulating large amounts of water (as must be included in typical studies of solvated systems to minimize finite size artifacts). For the purpose of validating protein models, crystal simulations also have the advantage that many incorrect models which lead to erroneous conformational states will be quickly identifiable. For example, one pioneering study by Krieger and colleagues 17 made incremental changes to the AMBER force field using root mean squared deviations in α-carbon positions to guide the optimization. However, validation studies must still undertake a great deal of sampling to ensure that the model does not merely represent the native state as a trap in the potential energy surface which would be traded for even more favorable conformations as the system evolves.
High-resolution X-ray structures of proteins are potentially useful because it is straightforward to assign deviations in the mean positions of atoms to errors in the force field. However, the simulation protocols themselves must be carefully designed to recreate the conditions of the experiment. X-ray structures of protein crystals present images of biomolecules in a complex solvent environment; there are probably non-aqueous components to the solvent, and the solvent itself is typically only ordered enough to be observed in the first solvation shell around the protein, if at all. Modeling the non-aqueous components of the solvent can be particularly challenging, and the relative abundances of each species near the surfaces of lattice proteins are not certain, even though the crystallization solutions themselves have known compositions. 13 Compounding the uncertainty about the experimental conditions, many X-ray diffraction studies make use of cryogenic conditions (flash freezing in liquid nitrogen) to improve the resolution of the data; this technique can also lead to minor alterations in the conformations of side-chains on protein surfaces, 18 and there is currently no adequate way to simulate biomolecules at temperatures in the range of 100K. For these reasons, X-ray structures must be chosen selectively for use as protein force field validation benchmarks.
Toxin protein II from the scorpion Androctonus australis Hector is an excellent system for testing how different combinations of molecular force fields and water models reproduce the known experimental structure of a protein. The toxin is chemically stable for months in a solution of 0.2M ammonium acetate at pH 6.8, and forms crystals in the orthorhombic P212121 space group by slow evaporation at 4°C.19 X-ray diffraction of these crystals has yielded resolutions as good as 0.96Å at 287K (PDB accession codes 1AHO20 and 1PTX21). Aside from its exceptionally high resolution, the 1AHO structure has a number of other helpful attributes. The asymmetric unit of the crystal is a single copy of the small, 64-residue protein; the unit cell consists of only about 6000 atoms when fully solvated. The ammonium acetate present in the crystallization solution has been modeled successfully before, 13 and a neutral pH of 6.8 requires no special considerations such as addition of hydronium ions or inclusion of protonated acidic residues, which are not commonly modeled in molecular simulations.
In this article, we present simulations of the Toxin II protein lattice using all pairwise combinations of four different molecular force fields and four different water models. In addition to analysis of the integrity of individual proteins, we make direct comparisons between the atomic fluctuations in our simulations and those observed in the X-ray structure, and dissect interactions across seven protein:protein interfaces in the crystal lattice to rate the performance of each model. We find that the integrity of the protein lattice is best maintained by the AMBER FF99SB force field, 22 and that both AMBER FF99SB and CHARMM22 with CMAP corrections 24,23 obtain several correct results in terms of the conformations of individual proteins within the lattice. We include a detailed online supplement to offer additional pictorial guides to the protein lattice as well as detailed information on the analysis of interfaces, in the interest of improving the parameterizations for simulating crystallized and solvated proteins.
2 Methods
2.1 Choice of protein force fields and water models
Four popular molecular force fields were evaluated: the AMBER FF99SB forcefield with improvements to protein backbone dihedral parameters suggested by Hornak and colleagues, 22 the AMBER FF03 force field developed by Duan and colleagues, 25 the CHARMM22 all-atom force field with CMAP corrections for protein backbone dihedral potentials 24,23, and the OPLS all-atom force field. 26 All of these force fields explicitly represent all hydrogen atoms, are non-polarizable, and contain only pairwise-additive terms for non-bonded interactions. All of the force fields use torsional potentials fitted to potential energy surfaces obtained from quantum mechanical calculations. Atomic charge parameters on each force field were fitted to reproduce either electrostatic potential surfaces obtained from quantum mechanical calculations (FF99SB and FF03), experimental properties of condensed liquids (OPLS), or a combination of experimental properties and interaction energies obtained from quantum mechanical calculations (CHARMM22).
All force fields used in the present study specify the TIP3P water model27 as the preferred water model for simulations of biomolecules in aqueous solutions. The CHARMM force fields were parameterized using a modified TIP3P water model (with Lennard-Jones terms added to hydrogen atoms), making it the only force field to contain its own description of water. However, the TIP3P water model was not designed for use with long-ranged electrostatics, the standard in modern simulations; investigators also apply different water models to any particular force field. We therefore tested all four molecular force fields with the TIP3P (or modified TIP3P in the case of CHARMM22), SPC-Extended (SPC/E),28 TIP3P-Ewald,29 and TIP4P-200530 water models in separate simulations. All of these water models are rigid, non-polarizable, and (with the exception of the original TIP3P model) compatible with long-ranged electrostatics.
2.2 Construction of the simulation cell
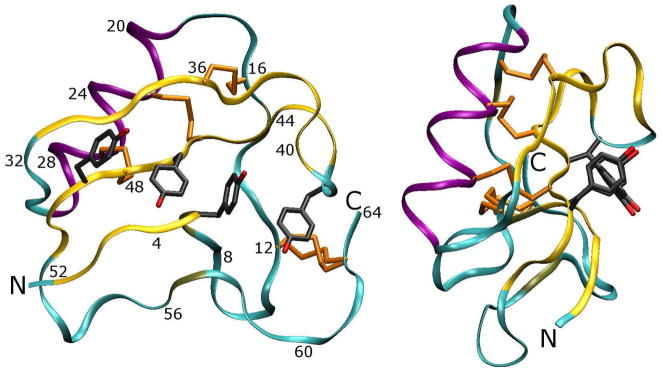
The unit cell was reconstructed by applying the symmetry operations of the P212121 space group to the protein and all water molecules identified in the 1AHO PDB file. 20 The major structural features of the crystallographic asymmetric unit (a 64-residue protein monomer) are presented in Figure 1: the monomer is primarily stabilized by four disulfide bridges as well as a series of four tyrosine residues arranged to optimize T -stacking interactions between aromatic rings. The monomer constitutes one asymmetric unit of the lattice; four symmetry-related copies of the monomer make up one unit cell. Figures 2, 3, and 4 illustrate the place of the monomer among its fourteen nearest neighbors.
Figure 1. Map of the scorpion toxin protein (one asymmetric unit of the crystal lattice).
Two views of the protein are presented. The protein backbone is shown in ribbon trace, with residue numberings to aid some points of the Results and Discussion in the main text. The protein is stabilized by at least two major features: first, a set of disulfide bridges links β-sheets and random coil regions of the protein, and second, a series of four tyrosine residues is arranged in a stable π-stacking arrangement. The Cys12–Cys63 disulfide bridge is observed in two conformations in the X-ray structure.
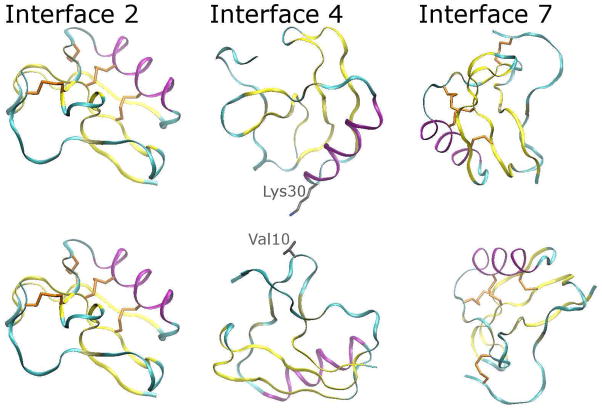
Figure 2. Position of the protein monomer in the 1AHO lattice (part I).
One protein monomer constitutes one asymmetric unit of the crystal lattice; four symmetry-related monomers make up one unit cell. This figure shows the orientation of one asymmetric unit relative to some of its neighbors (note that the lattice contacts are equivalent for any particular asymmetric unit). Each monomer is situated amongst fourteen near neighbors, defining a set of seven unique interactions, which we have numbered arbitrarily. The three interactions with the smallest extent of contact between monomers (2, 4, and 7) are shown above; side-chains of residues in each monomer containing heavy atoms that reside within 5Å of one another in the X-ray structure are detailed in stick form. The coloring scheme and convention of detailing the four disulfide bridges, which can be used as landmarks in the protein, is carried over from Figure 1. The evolution of all seven interfaces will be discussed throughout this article.
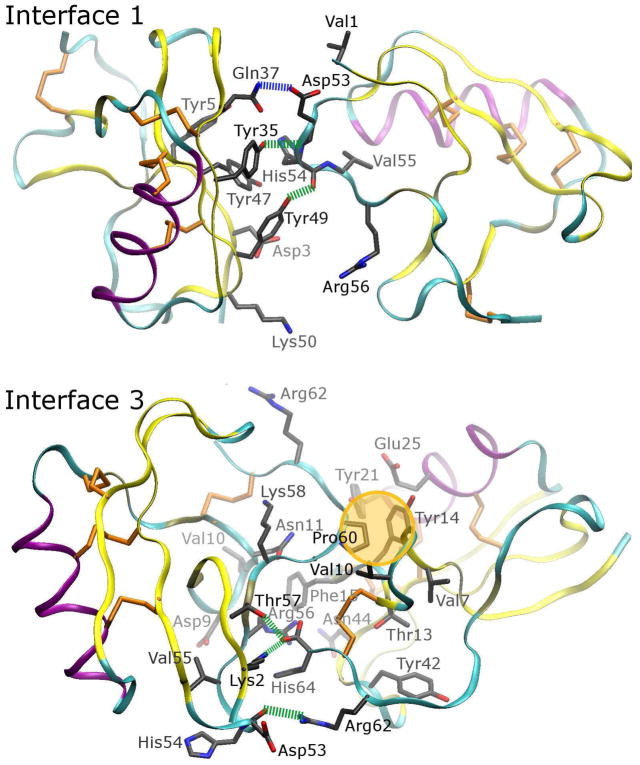
Figure 3. Position of the protein monomer in the 1AHO lattice (part II).
Interfaces 1 and 3 between monomers are extensive, with specific hydrophobic contacts (highlighted with a gold bubble), hydrogen bonds or salt bridges involving backbone atoms (indicated by green dashed lines), and hydrogen bonds between side chains on separate monomers (indicated by blue dashed lines). The color scheme for each of these specific features is designed to match the flags in Figure 12, and Figures S14 to S16 of the Supporting Information, where the evolution of these specific contacts during each of the simulations is discussed in more detail. Overall, interface 3 may be the most significant lattice interaction.
Figure 4. Position of the protein monomer in the 1AHO lattice (part III).
Interfaces 5 and 6 show numerous residues in close proximity and two more specific contacts, both of them involving Arg62 in interface 6. The π-stacking interaction between this arginine and Tyr35 of the adjacent monomer (the tyrosine is not part of the T -stacking arrangement illustrated in Figure 1), coupled with excellent geometry for a hydrogen bond between one of the arginine’s η nitrogens and the Tyr49 hydroxyl group (Tyr49 is part of the T -stack) suggests that it is a particularly strong point of contact between the monomers.
The 1AHO structure itself was refined from the same X-ray diffraction data set as the previously published 1PTX structure, 21 as a proof-of-concept application of the SnB program 31 for ab initio structure determination and refinement from X-ray diffraction data. The backbones of the two structures are superimposable to less than 0.1 Å RMSD, and the positions of side chains, including the Cys12:Cys63 disulfide bridge observed to occupy two conformations in both structures, are virtually identical. A few atoms, such as distal atoms of the Lys30 and Lys50 side chains as well as part of the Asp9 carboxylate group, were unobserved by the 1AHO structure determination process but reconstructed with the TLEAP module of AMBER10.32 The 1AHO structure’s 0.96 Å resolution was not necessarily a reason to choose it over the 1PTX structure’s 1.3 Å resolution, but the 1AHO structure’s identification of a larger number of water molecules, owing to the SnB program’s aggressive fitting of solvent molecules into the electron density of the X-ray diffraction data, provided a reason to choose that structure over 1PTX. Of the 129 water molecules in the 1AHO structure (see Supporting Information Figure S1), 42 have incomplete occupancies; the 1PTX structure contains 106 distinct water molecules, nine of them delocalized over two or more sites. As in previous studies, all crystallographic water molecules were included in the initial structures for all simulations. 13 To address the problem of uncertainty in the positions of numerous water molecules in the present simulations, waters were added at all partially occupied sites with the expectation that additional waters would be necessary to completely hydrate the unit cell, and that the first few picoseconds of dynamics, if not the energy minimization process, would settle any moderate steric clashes at the observed water positions.
Periodic boundary conditions are a valid approximation for simulations of crystal lattices. However, it is important to realize that no two unit cells within a crystal are instantaneously identical: the proteins within the lattice are subject to thermal fluctuations, and the lattice itself may contain more significant defects which can be thought of as “rigid body” movements of entire symmetry-related lattice subunits (asymmetric units). In addition, the periodicity of the simulated system can introduce artificial long-range ordering because adjacent copies of the system cannot move independently; periodic boundary conditions may therefore still impose an artificial constraint on the system. In order to suppress any artifacts from the periodicity of our model, we constructed the simulation cell from twelve individual unit cells in a 2×2×3 arrangement, a rectilinear box initially measuring 91.8 Å×81.4 Å×90.3 Å containing a total of 48 protein monomers.
The crystals were grown in a 0.2M ammonium acetate solution by slow evaporation. To mimic these conditions and neutralize the system charge, 84 acetate and 36 ammonium ions were added, based on the volume of the unit cell not taken up by protein atoms. These ions were modeled using the parameters of Saigal and Pranata 33 when in simulations with the FF99 and FF03 protein force fields, using OPLS-AA parameters 26 for the relevant small molecules in simulations with the OPLS protein force field, and using parameters and charges taken from model compounds included with the CHARMM22 force field.34 The ions were added to the lattice interstices (regions of space not occupied by proteins within the crystal lattice) using the “AddToBox” program described in a previous article. 13 As before, we placed the ions first, requiring that the AddToBox program leave at least 4.0 Å between the ions and any protein or crystallographic water molecules. We also required that the AddToBox program place new ions at least 4.0 Å from any other ions previously placed, to ensure that the ions would be evenly distributed throughout the simulation box from the outset of the simulations. We then added various numbers of water molecules, iteratively refining the amount and restarting each simulation several times until the system volume reached an equilibrium value within 0.3% of the experimental value under constant pressure dynamics as described below. The iterative solvation and equilibration process required that each simulation be restarted after as much as 10ns of constant pressure dynamics to ensure that the amount of added solvent was properly maintaining the crystal unit cell volume. After fully solvating the lattices with each model, the total number of particles ranged from 72000 to 83000 depending on the protein force field and water model, as listed in Table 1.
Table 1. Compositions of the protein crystal system for four protein force fields and four water models.
Force fields and water models are described and referenced in the Methods. A total of 46176 atoms were included in the protein component of each system (48 toxin monomers in 12 independent representations of the crystallographic unit cell, with all hydrogens represented explicitly). Additionally, 84 acetate and 36 ammonium ions were added to neutralize the overall charge of the system and represent the 0.2M ammonium acetate concentration present in the crystallization solution (768 atoms were present in the ammonium acetate ions). Most systems contained roughly 73,000 particles, but systems using TIP4P-2005 contained up to 83,000 due to massless “extra points” on the water molecules.
| Protein Force Field and Number of Watersa | ||||
|---|---|---|---|---|
| Water Model | FF99SB | FF03 | CHARMM22 | OPLS |
| SPC/E | 9000 | 8904 | 8345 | 8357 |
| TIP3P | 8898 | 8850 | 8484 | 8182 |
| TIP3P-Ewb | 8970 | 8862 | 8238 | 8238 |
| TIP4P-2005 | 9012 | 8916 | 8360 | 8360 |
Includes 6192 water molecules present (or at least partially occupied) in the 1AHO structure
Model “F” presented in the work by Price et al. 29 was used in simulations with the FF99SB and FF03 force fields, but the related model “B” was used with the CHARMM22 and OPLS force fields due to NAMD’s lack of a homogeneity approximation for long-ranged van-der Waals interactions.
2.3 Molecular dynamics simulations
After constructing the simulation cell, energy minimization and restrained equilibration were carried out as described previously 35. All force calculations for this work were performed with a 9 Å cutoff on real-space interactions and Smooth Particle-Mesh Ewald electrostatics on a 96 × 96 × 96 mesh. The Ewald reciprocal space force calculation was performed at every step of dynamics; no multiple-time-stepping scheme was used. Production dynamics were performed with a 2 fs time step, using an Andersen thermostat to maintain the system temperature at 287 K by random velocity reassignments every 2 ps. Restrained equilibration began in the constant-volume ensemble but then transitioned into the constant-pressure ensemble as described in our previous studies. 35 In this work, volume rescaling was anisotropic based on the components of the virial tensor perpendicular to each face of the simulation cell. Constant pressure was maintained at 1 bar by a Berendsen barostat with a relaxation time constant of 1 ps−1.
Simulations using the FF99SB and FF03 force fields were conducted slightly differently than those with the CHARMM22 and OPLS force fields: tests of both AMBER force fields were conducted with the PMEMD program in the AMBER1032 software package, wheras tests of the CHARMM22 and OPLS force fields were carried out with NAMD v2.6 36 with extensions (now available in NAMD v2.7) for the use of TIP4P water. Several variants of Ewald-compatible TIP3P models were developed by Price and colleagues29: model F was paired with the FF99SB and FF03 force fields because the AMBER software package supports a homogeneity approximation for long-ranged van-der Waals corrections to the virial, but model B (identical to model F except for a change to the Lennard-Jones parameters to compensate for a lack of any long-ranged van-der Waals correction) was paired with the CHARMM22 and OPLS force fields for calculations in the NAMD software package (at the time these simulations were begun NAMD did not support such a correction). The homogeneity approximation would have no effect on the dynamics of a simulation conducted at constant volume, and the contributions to the virial are small in the context of a condensed phase system. Also, an additional 10ns of equilibration dynamics, using a Langevin thermostat37 and Nosè-Hoover Langevin piston barostat, 36 were performed for the CHARMM and OPLS force fields before beginning 100ns of production dynamics, but because most analyses were restricted to the final few ns of the 100 ns “production” dynamics this additional equilibration does not mean that the configurations in any set of simulations were significantly more evolved than another. Finally, the disulfide bridge between Cys12 and Cys63, shown in two conformations with a 55%/45% distribution in the 1AHO structure, was modeled with two of the four asymmetric units having either conformation in every unit cell to iniate the FF99SB and FF03 simulations, but modeled solely in the major conformation for simulations with the CHARMM22 and OPLS force fields, reflecting the preferences of different investigators managing each set of simulations. The conformation of the disulfide bridge was observed to flip in some protein monomers in all simulations (data not shown); although only 10 to 20 flips (and even fewer cases in which a single disulfide bridge flipped more than once) were observed in each simulation, the existence of such flips is one indication that the initial states of the bridges did not adversely affect the outcome of each simulation.
3 Results
3.1 Hydration requirements depend on the choice of force field
The first comparison that can be made between the various force fields and water models is the number of water molecules required to maintain the volume of each simulation cell near the experimental value. These numbers are listed in Table 1. Simulations with the FF99SB and FF03 required 5–10% more water molecules than simulations with CHARMM22 or OPLS, but for any particular force field, the overall number of water molecules required to solvate the unit cell was more consistent. Most force fields required slightly fewer molecules of TIP3P water than of the other water models studied; this could be expected as TIP3P underestimates the density of real water by ~1% when used with long-range electrostatics. 38 The CHARMM/TIP3P simulation required more water molecules than other CHARMM simulations, but as stated in the Methods the TIP3P water used with CHARMM is a modified version of the original TIP3P model. 24 While we tried to place water in each simulation to hold its equilibrium volume very near the experimental value, we also considered the possibility that the structural results were sensitive to minor variations in the total numbers of waters. To address this concern, we conducted an additional 100ns simulation using the FF99SB force field paired with the TIP4P-2005 water model and 72 more water molecules than the simulation presented in these results (a volume increase of 0.3%). Several of the analyses presented below were repeated on this control. As shown in Figure S2 of the Supporting Information, we could not detect any effect from this volume perturbation on the results we sought.
3.2 Model-dependent deformation of the unit cell
A striking result of all 16 crystal simulations was the fact that maintaining a constant pressure via anisotropic unit cell rescaling altered the unit cell dimensions far in excess of the experimental error, even while the overall volume of the system remained at a value well within the experimental error as shown in Table 2. As mentioned in the Methods, refinements of the 1AHO structure and the 1PTX structure21 both started from the same data set, but took different approaches. The unit cell dimensions of 1AHO were reported as 45.90 × 40.70 × 30.10 Å, while those of 1PTX were reported as 45.94 × 40.68 × 29.93 Å. Based on these figures, we estimated the experimental error of each unit cell dimension to be ±0.1Å. Assuming the errors in each unit cell dimension to be independent, the total experimental error in the unit cell volume can be estimated as 0.8%. In a previous communication, 39 we estimated the experimental error in the volume of a much larger unit cell to be 0.5%, based on a broader sample of isomorphous crystals, but as shown in Figure 5 either value would lead to the same qualitative result when comparing the experimental and simulated errors in the unit cell dimensions.
Table 2. Deviation of the total volume in each simulation cell over 100ns of dynamics.
Total volumes of all simulations were close to the experimental values and consistent throughout each simulation. As shown in the Supporting Information, the simulation results appear to be insensitive to minor perturbations of the total volume.
| Volume Deviation from Experiment (%) | ||||
|---|---|---|---|---|
| Water Model | FF99SB | FF03 | CHARMM22 | OPLS |
| SPC/E | 0.13 ± 0.05 | 0.00 ± 0.05 | −0.33 ± 0.07 | 0.09 ± 0.06 |
| TIP3P | 0.13 ± 0.04 | 0.26 ± 0.04 | 0.20 ± 0.09 | −0.11 ± 0.05 |
| TIP3P-Ew | 0.18 ± 0.05 | 0.33 ± 0.05 | −0.19 ± 0.12 | −0.06 ± 0.08 |
| TIP4P-2005 | 0.09 ± 0.08 | −0.28 ± 0.07 | −0.13 ± 0.10 | 0.05 ± 0.07 |
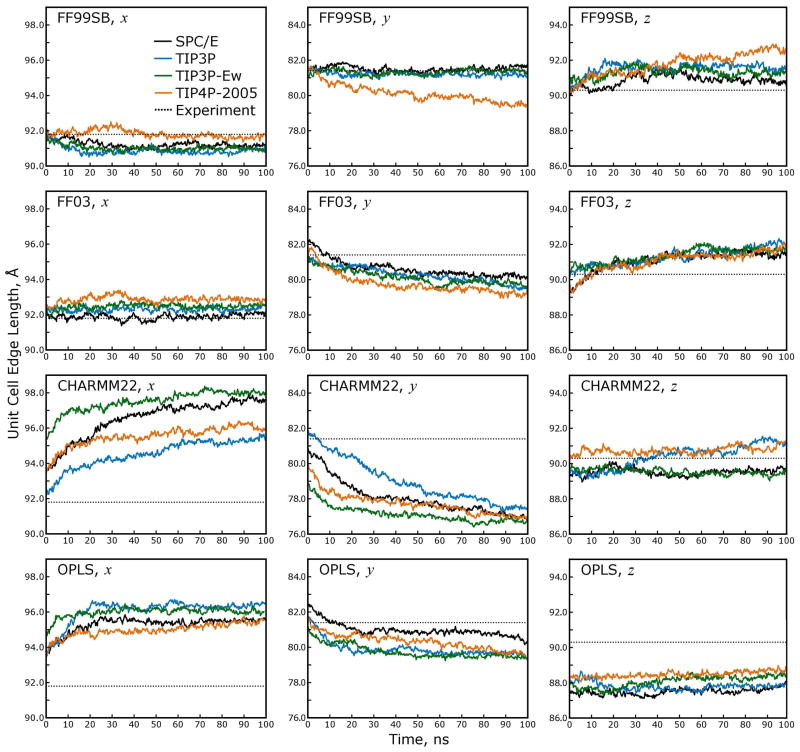
Figure 5. The unit cell dimensions are not maintained by all force fields during constant-pressure dynamics with anisotropic rescaling.
The ability of a force field to maintain the unit cell aspect ratios gives an early indication of whether the model is behaving correctly. As shown in the panels above, the unit cell volume is closely maintained near the experimental value throughout all simulations, but the individual unit cell dimensions drift away from their experimental values, particularly in simulations with the CHARMM22 and OPLS force fields. While the protein force field has the most influence over the evolution of the unit cell dimensions during each simulation, the water model also can have an effect; the three-point water models all give similar results, though some slight differences are apparent, when applied with the FF99SB or FF03 force fields. In contrast, the four-point TIP4P-2005 water model tends to give the poorest results when paired with the FF99SB or FF03 force fields, and no generalizations can be made about pairings of water models with the CHARMM22 or OPLS force fields. The inset legend in the upper right panel applies to all panels.
The unit cell deformations appear to have reached equilibrium in all simulations, although in the case of CHARMM22 the deformations only stabilized in the last 20ns of the trajectories. The CHARMM22 and OPLS force fields both produce errors in excess of 5% in the individual unit cell dimensions. The choice of water model appears to have an effect, but no model seems to be sufficiently well matched to the CHARMM22 or OPLS force fields to produce the correct unit cell dimensions. In contrast, the closely related FF03 and FF99SB force fields produce errors of 0–2% in the unit cell dimensions. The results for the FF99SB and FF03 force fields again show dependence on the water model, and because the overall errors are so small in the case of FF99SB, the three-point water models stand out as more appropriate than the four-point water model for simulations with this force field.
3.3 Integrity of individual and collective protein structures
Whereas anisotropic deformations of the unit cell provide an important description of the state of the crystal lattice, the root mean square deviations (rmsds) of atomic positions of individual proteins in the lattice provide a preliminary description of the microscopic integrity of the protein structure when simulated with each force field and water model. As shown in Table 3, the backbones of individual proteins remained close to their crystallographic conformations (plots of the average monomer rmsd, as it evolves over the course of each simulation, can be found in Supporting Information Figure S3). However, a small protein with four disulfide bridges would not be expected to show large changes in its backbone conformation. While there is some correlation between the maintenance of the unit cell dimensions and the monomer rmsds in each simulation (FF99SB yields the best rmsds, and OPLS the poorest), monomer rmsds do not effectively discriminate among the various models.
Table 3. Final monomer rmsd measurements after 100ns of dynamics.
Backbone rmsds are given as averages over all 48 protein monomers in each simulation cell; standard deviations in the rmsds obtained for all individual monomers provide error bars.
| Monomer RMSD (Å) | ||||
|---|---|---|---|---|
| Water Model | FF99SB | FF03 | CHARMM22 | OPLS |
| SPC/E | 0.61 ± 0.06 | 0.82 ± 0.14 | 0.85 ± 0.10 | 1.22 ± 0.18 |
| TIP3P | 0.59 ± 0.07 | 0.92 ± 0.11 | 0.91 ± 0.14 | 1.39 ± 0.22 |
| TIP3P-Ew | 0.57 ± 0.04 | 0.86 ± 0.15 | 0.82 ± 0.11 | 1.28 ± 0.28 |
| TIP4P-2005 | 0.66 ± 0.11 | 0.86 ± 0.11 | 0.91 ± 0.10 | 1.17 ± 0.20 |
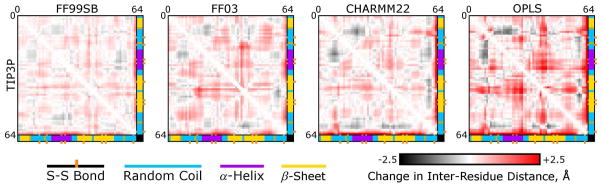
Figure 6 plots deviations in the simulated versus experimental distances between all α-carbons in matrix form, with the protein monomer’s secondary structure elements colored to illustrate how different regions of the monomer move, on average, relative to one another. (As shown by the matrices for all 16 simulations in Figure S5 of the Supporting Information, the choice of water model has little effect on any deformations of the monomer.) Some patterns in the deviations appear to occur in all force fields. In particular, two of the random coil regions in the protein (residues 8 to 12 and residues 52 to 56) seem to be drawn closer to the α-helical region (residues 19 to 29). All force fields correctly reproduced the interatomic distances for backbone atoms within the α-helix. All but the OPLS force field produced very little deviation in the juxtaposition of the helix with the residue 43–51 β-strand; the interaction of these two secondary structures is reinforced by two disulfide bonds, but these bonds take on non-crystallographic torsional configurations when simulated with the OPLS force field. One other pattern appears to be common to all of the matrices in Figure 6: all force fields allow a β-hairpin loop (residues 39–43) to move away from most other parts of the protein; as can be seen in Figure 7, this occurs as the β-hairpin flips outwards, possibly breaking a T -stacking (π-stacking) contact between Tyr42 and Tyr5. In all of the force fields, π-stacking effects are not explicitly included, and can only result from superposition of spherically symmetric potentials centered at each atom site (the interaction of slightly polarized aromatic hydrogens with aromatic carbons); nonetheless, the FF99SB and CHARMM22 force fields maintain the orientatons of Tyr42, Tyr5, Tyr47, and Tyr49 side chains very near the crystallographic values. The FF03 force field occasionally allows Tyr42 to adopt an alternate conformation in which its hydroxyphenyl group is pointed at the phenyl ring of Tyr5. The OPLS force field permits much more severe displacement of the Tyr42 side chain relative to Tyr5, and also displacement of Tyr49 relative to Tyr47. Analysis of the pairwise backbone distances therefore identifies some local structural deviations that distinguish the four force fields, but more generally all four force fields tend to introduce similar errors in the backbone structure of the protein monomers.
Figure 6. Distance deviation matrices for α-carbon atoms in simulations with TIP3P water.
Each panel is a color representation of the matrix of interatomic distances between α-carbons in the protein monomer. (Numbers on the axes of each plot refers to residue numbers, and colored bars on axes refer to secondary structural elements.) Results are presented as differences between the average distances obtained over all 48 symmetry-related monomers over the last 5ns of each simulation and the corresponding distances seen in the X-ray data. While this figure only presents simulations conducted with the TIP3P water model, matrices for all 16 simulations are given in Figure S5 of the Supporting Information.
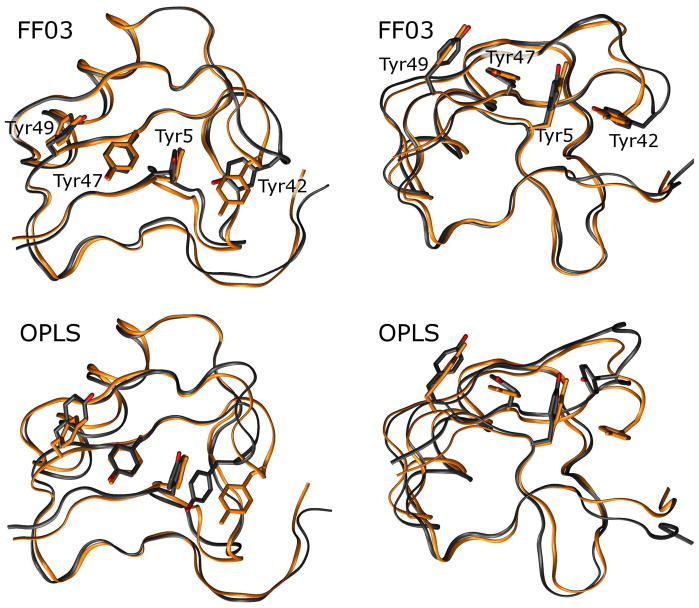
Figure 7. Distortion of the residues 39–43 hairpin can occur with loss of interactions between Tyr42 and Tyr5.
While all force fields seem to allow the residues 39 to 43 β-hairpin turn to become distorted, the FF03 and OPLS force fields allow the associated Tyr42 side chain to adopt non-native conformations in which it breaks its favorable T-stacking interaction with Tyr5. Above, the X-ray structure is shown in orange while a representative monomer from a simulation performed with each force field is shown in black; structures are superimposed by quaternion alignment of backbone atoms. None of the force fields contains explicit terms for the T-stacking effect, though it apparently stabilizes four tyrosine side chains in the X-ray structure. More importantly, the FF99SB and CHARMM force fields maintain the interaction between tyrosine side chains; it is not certain whether the FF99SB and CHARMM force fields have superior pairwise terms to stabilize the nonbonded interactions between side-chains, or if they simply have superior backbone dihedral terms which prevent the side chains from exploring non-native conformations. Future studies with restrained simulations which isolate backbone and side chain degrees of freedom may provide these answers.
When analyzing previous streptavidin crystal simulations, 39 we used the “lattice rmsd,” a measurement of rmsd designed specifically for assessing the arrangement of many proteins within the unit cell. A key feature of this metric was that only crystallographic symmetry operations, not optimal quaternion alignments, were used to superimpose the various asymmetric units onto the experimental reference structure. Before measuring the lattice rmsd, the unit cell found at some time t was also rescaled to be identical to the original unit cell dimensions, and all atomic positions were rescaled proportionately. The lattice rmsd therefore provides complementary information to the measurements of unit cell dimensions and monomer rmsd. Table 4 lists the final lattice rmsd for each force field and water model combination after 100 ns of simulation; plots of the lattice rmsd over the entire simulation are given in Supporting Information Figure S4. Even though unit cell deformation is removed from the calculation, the magnitude of the lattice rmsd in each simulation is strongly correlated with the degree of deformation in the unit cell, confirming that the proteins have shifted relative to their original locations in the lattice.
Table 4. Final lattice rmsd measurements after 100ns of dynamics.
Lattice rmsd measured for all simulations is positively correlated with deformations of the unit cell dimensions (see Figure 5), even though coordinates were rescaled to eliminate such deformations prior to these calculations. The lattice rmsd quantifies translations and rotations of the individual asymmetric units relative to their positions in the X-ray structure, apart from any net deformation of the unit cell which may also have occurred; a complete description of the lattice rmsd is given in the main text. Error bars are not provided as there can be only one measurement of the lattice rmsd per simulation.
| Lattice RMSD (Å) | ||||
|---|---|---|---|---|
| Water Model | FF99SB | FF03 | CHARMM22 | OPLS |
| SPC/E | 1.13 | 1.34 | 1.71 | 2.19 |
| TIP3P | 1.08 | 1.39 | 1.59 | 2.20 |
| TIP3P-Ew | 1.05 | 1.21 | 1.68 | 2.34 |
| TIP4P-2005 | 1.70 | 1.34 | 1.64 | 2.00 |
3.4 Atomic fluctuations in simulations and comparison to experiment
Measurements of atomic positions are a central result of any crystallographic study, but the X-ray results also give indications as to the mobility of atoms in the crystal. The crystallographic “B factors,” sometimes called temperature factors, can be interpreted as mean squared fluctuations of atoms, which can in turn indicate the appropriate qualities of a molecular force field, 40,41 but this characterization is incomplete. More precisely, the isotropic B factors are proportional to the second moment of the distribution of all corresponding copies of a particular atom throughout the entire lattice, including any crystal defects and “lattice disorder,” subtle rigid body movements of proteins averaged over many unit cells. We measured the second moments of distributions in atomic positions over the final 50ns of each simulation and took the square roots to obtain atomic RMS fluctuations F, which can be compared to crystallographic B factors B using the formula:
| (1) |
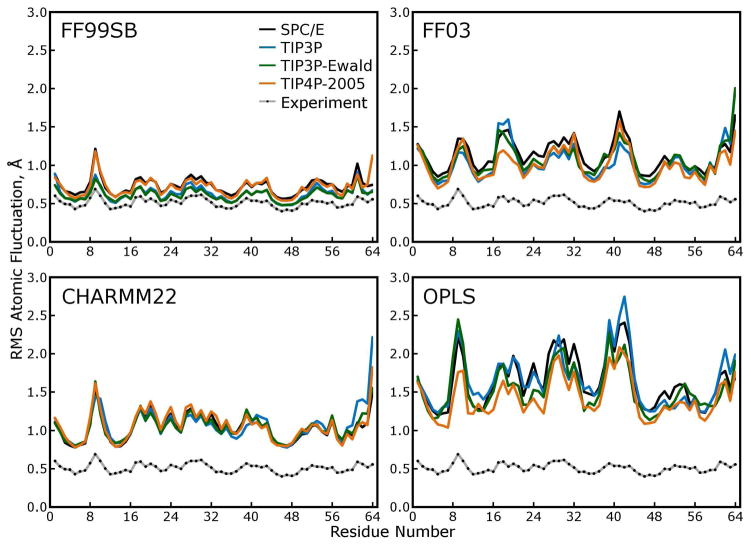
These fluctuations are plotted in Figure 8. While the atomic RMS fluctuations produced by any one force field do not appear to be very sensitive to the water model, across different force fields there was noticeable variability in the magnitudes of the fluctuations and the size of the “baseline” fluctuations of the least variable atomic positions.
Figure 8. Atomic root mean squared (RMS) fluctuations of.
α carbons. These fluctuations were calculated after using the symmetry operations of the P212121 space group to superimpose all 48 asymmetric units (protein monomers) in each simulated lattice and are thus directly comparable to the scaled roots of crystallographic B-factors, as shown in each panel. Larger RMS fluctuations in a simulation rarely indicate tremendous mobility of a particular region of the protein in any of our simulations. Rather, as shown by comparing these results to those of Figure 9, higher RMS fluctuations indicate more lattice disorder, increased rigid-body motions of the individual protein subunits within the lattice.
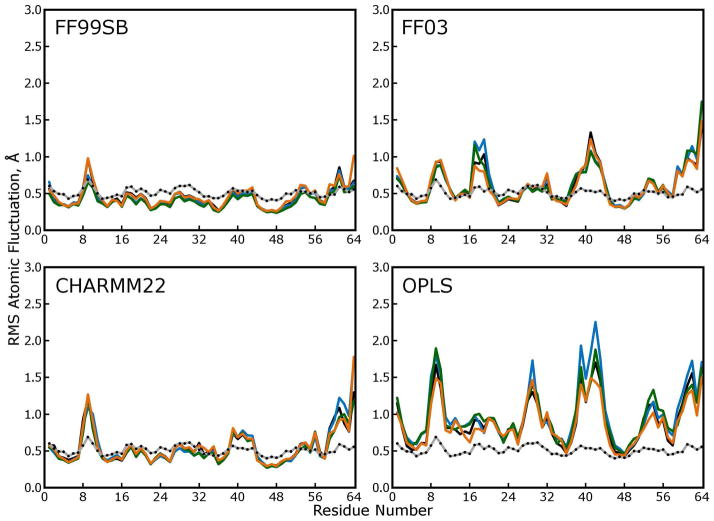
Lattice RMSD measurements confirmed that the proteins in all lattices shifted relative to their experimental positions, but the fluctuations of α carbons about their new mean positions could originate both from flexibility of the protein backbone and rigid body motions of entire asymmetric units within the lattice. To distinguish between these two contributions to the atomic fluctuations, we broke all snapshots of each trajectory into their 48 protein monomers and superimposed all of them using optimal quaternion alignments against the average monomer structure. The atomic fluctuations obtained from this distribution of atomic positions are plotted in Figure 9. When the lattice disorder is removed, the FF99SB and CHARMM22 force fields yield similar overall fluctuations in the protein backbones; the ongoing increase in lattice rmsd evident in Figure S4 of the Supporting Information clearly contributes to the elevated atomic fluctuations for the CHARMM22 force field shown in Figure 8, and the fact that the magnitudes of fluctuations in simulations with the FF99SB force field fall only about 0.3Å if lattice disorder is omitted indicates that any rigid-body movements of proteins in these simulations tend to occur symmetrically across all 48 monomers. In the other three force fields, where the lattice disorder makes a large contribution to the atomic fluctuations, there arises the possibility that whatever contacts are forming and breaking are not consistent across all monomers, such that the lattice is melting.
Figure 9. Atomic fluctuations obtained by quaternion alignments of protein backbones.
In the main text, atomic fluctuations were computed based on the distribution of each atom as obtained in the lattice rmsd calculation (see above), which we believe is the best approximation to the distribution described by crystallographic “B factors” that our simulations can yield. In the plots above, atomic fluctuations were computed based on the distribution of each atom obtained after optimal quaternion alignments of the protein backbones. This method, which is used to compare fluctuations in solution-phase experiments to crystallographic B factors, omits “lattice disorder” from the calculation and therefore lowers the observed fluctuations. Comparison of these plots to Figure 8 shows that fluctuations obtained from simulations with the FF99SB forcefield (which closely reproduce the experimental B factors) contain very little lattice disorder. In contrast, simulations with other force fields produce significant lattice disorder as well as wider variability in protein backbone conformations. Together with the lattice RMSD measurements, these plots suggest that, in simulations with the FF03, CHARMM22, and OPLS force fields, proteins are shifting away from their initial positions in the lattice in many different ways—the original symmetry is being destroyed and the lattice is melting.
3.5 Deformation of the lattice along crystallographic interfaces
The global deformations of lattice dimensions and lattice RMSDs noted above demonstrate that the relative arrangement of subunits within the simulated unit cells shifts from the initial positions. All monomers in the 1AHO crystal structure are initially equivalent; accordingly, each monomer makes the same contacts to its nearest neighbors, as enumerated in Figure 10a. Each monomer contacts fourteen adjacent monomers, defining a set of seven unique interfaces shown in Figures 2, 3, and 4. Relative to an arbitrary definition of axes, the monomers can take on any of four different orientations, enumerated by the symmetry operations of the P 212121 space group; the crystal itself consists of pleated sheets of alternating pairs of these orientations stretching along the yz plane shown in Figure 10b. We examined how the initially homogeneous contacts evolved, and possibly diverged, during each simulation.
Figure 10. Arrangement of monomers in the simulated crystal lattice.
(a) Arbitrary numbering and relative locations of the center of mass vectors between a chosen monomer and its 14 neighboring monomers. Protein coloring runs blue to red from N- to C-terminus. The interactions are broken into seven mutually interacting pairs (each of which shares a color).
(b) Orientation of vector 1 from panel a on each of the 48 monomers in the lattice, superimposed on cartoon representations of the monomers. Monomers are colored based on their absolute orientation relative to the lab frame. (c) As panel b, but with the protein structures omitted.
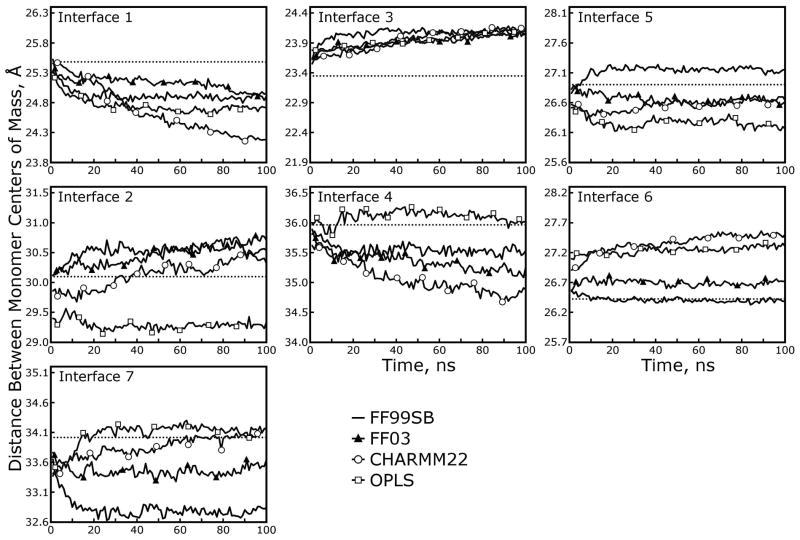
The average distances between monomer centers of mass along the seven principal interactions in the 1AHO structure are plotted in Figure 11. Both increases and decreases in the distances are observed, reflecting loss of crystallographic protein-protein interactions and formation of non-native interactions. Separation of the monomers into four distinct sets based on their absolute spatial orientations did not significantly alter the results (data not shown), indicating that the periodicity imposed by our simulation cell did not affect the evolution of these contacts. Some variations in the centers of mass distances span the four force fields. In general the simulations tighten interfaces 1 and 4 while stretching interfaces 3 and 6. While the metric used here directly measures protein-protein distances, the effects of water models are also apparent: for the CHARMM22 simulations, the trajectory containing TIP3P water generally stayed closest in overall monomer layout to the crystal structure, whereas for OPLS TIP4P appears to perform best, and for FF99SB and FF03 all of the three-point water models produced roughly equivalent accuracy.
Figure 11. Deformation of the lattice measured according to distances between protein monomers.
As shown in Figure 10, seven unique interfaces may be defined for closely interacting Toxin II monomers in the lattice. The distances between monomer centers of mass are plotted for simulations with all four force fields and the TIP3P water model (a complete presentation of all sixteen simulations can be found in Figures S6 and S7 of the Supporting Information). Dotted lines show the correct value from the X-ray structure. While the FF99SB force field performed the best in terms of maintaining the overall unit cells dimensions, this analysis shows that all of the force fields produce similar behavior in terms of the relative positions of the monomers.
It is also instructive to consider how the observed deformations in subunit-subunit distances differ from those that would occur simply as a result of the (anisotropic) rescaling of the unit cell at each step. We therefore rescaled the crystallographic unit cell and protein coordinates to match the box dimensions observed at timesteps in each of the sixteen simulations, and recomputed the inter-monomer distances from the rescaled crystallographic coordinates. Differences in these intersubunit distances and the actual distances observed in each simulation are plotted in Supporting Information Figures S8 and S9. Several systematic trends are apparent across all water models and force fields in this analysis—interactions 3 and 6 are consistently longer than expected based on symmetric deformation, whereas interactions 1 and 7 are shorter than expected. This uniformity suggests that all force fields share propensities to overestimate the strength of certain interactions and perhaps underestimate the strength of crystallographic native contacts.
3.6 Evolution of individual residue contacts between monomers
The protein-protein interactions contributing to deformation of the initial crystal lattice can be probed in detail through analysis of the specific residue-residue contacts forming intersubunit interactions. Lists of “general” contacts, residues on different monomers having any heavy atoms within 5Å of one another, are presented in Figures S10 to S13 of the Supporting Information. The initial set of “general” contacts present in the X-ray structure is shown alongside color-coded matrices denoting the formation of new contacts or loss of native ones. Several changes in intersubunit interactions occur similarly across all water models and all protein force fields, such as the formation of a large number of new contacts across interfaces 1, 2, and 5, and the loss of a large number of contacts across interface 3. These changes in interactions are, in turn, reflected in the changes in distances between monomer centers of mass noted in the previous section.
More significant changes throughout the simulations become apparent when considering only favorable interactions between residues, such as hydrogen bonds, salt bridges, or hydrophobic packing as defined by proximal apolar carbons. We identified all of these interactions, summarized in Table 5, by a distance criterion of 3.5 Å or less between the relevant heavy atoms, with one exception for a “hydrophobic” contact involving a cation-π interaction between Tyr35 and Arg62, which was considered intact if the distance between the Arg62 Cζ atom and the centroid of the tyrosine ring was less than 4.0Å. The evolution of the contacts listed in Table 5 is presented in Figure 12, and comprehensive lists of all favorable interactions formed or broken over the course of each trajectory are given in Figures S14 to S16 of the Supporting Information. All combinations of protein and water force fields show a variety of new hydrogen bonds forming between monomers, though only a few of these bonds occur consistently and most reflect heterogeneity in the lattice structure at the end of each simulation. Table 6 offers a simple characterization of each force field’s propensity to form salt bridges, hydrogen bonds, and hydrophobic contacts by presenting changes in the total numbers of each type of specific interaction that have formed, across all interfaces, during the course of each simulation. In this analysis, all force fields are qualitatively very similar, with a propensity to create new hydrogen bonds, whether between the polypeptide backbones or various side chain groups. The total number of hydrophobic interactions decreased in nearly all simulations: as shown in Figure S15 of the Supporting Information, a hydrophobic contact between the Tyr14 and Pro60 side chains was consistently broken in all simulations, and the cation-π interaction between Tyr35 and Arg62, preserved in the FF99SB simulations, was often broken in other simulations. Figure S15 of the Supporting Information shows that a few new hydrophobic contacts did form in the place of these native contacts, though with modest occupancies.
Table 5. Specific contacts between protein monomers inferred from visual inspection of the X-ray structure.
| Contact Type | Interface | Atoms of Monomer 1 | Atoms of Monomer 2 |
|---|---|---|---|
| HB, Backbonea | 1 | Tyr49 Oη | His54 O |
| Salt Bridge | 1 | Tyr35 Oη | His54 N |
| HB, Side Chainb | 1 | Gln37 Nε | Asp53 Oδ |
| HB, Backbone | 3 | Lys2 Nζ | His64 OXT |
| HB, Backbone | 3 | Thr57 Oγ1 | His64 OXT |
| HB, Backbone | 3 | Asp53 O | Arg62 Nη |
| Hydrophobic | 3 | Pro60 Cγ, Cδ | Tyr14 aromatic ring |
| Hydrophobicc | 6 | Arg62 guanidinium | Tyr35 aromatic ring |
| HB, Side Chain | 6 | Arg62 Nη | Tyr49 Oη |
Hydrogen bond involving one or more backbone atoms
Hydrogen bond involving only side chain atoms
Actually involves a cation-π interaction between the arginine guanidinium group and the tyrosine aromatic ring, labeled “hydrophobic” for lack of a better category
Figure 12. Evolution of specific contacts identified in the 1AHO crystal structure over the course of 16 simulations.
The occupancy of each specific contact listed in Table 5 at the final frame of each trajectory is shown as a shaded square. Each column represents a separate force field and water model combination. Black squares indicate that a contact has been completely lost, while white squares indicate that a contact has been maintained. The residues involved in each contact are listed on the left side of the table, and the type of contact and interface it spans are indicated by the colored, numbered boxes on the right: green, hydrogen bonds involving only backbone atoms; blue, hydrogen bonds involving side chain atoms; yellow, hydrophobic contacts.
Table 6. Gain or loss of specific contacts of three different types in all simulations.
Hydrogen bonds (or salt bridges) and hydrophobic interactions between separate monomers (essential to lattice contacts) formed and lost during each simulation were tallied and split into three general categories. First column: salt bridges or hydrogen bonds involving two side chain groups. Second column: salt bridges or hydrogen bonds involving at least one backbone group. Third column: hydrophobic interactions, including the cation-π interaction between Arg62 and Tyr35. All specific contacts originally present in the X-ray structure are listed in Table 5.
| Force Field | Water Model | Salt Bridge/Hydrogen Bond (Side Chain) | Salt Bridge/Hydrogen Bond (Backbone) | Hydrophobic |
|---|---|---|---|---|
| FF99SB | SPC/E | 1.6 | 2.1 | −0.4 |
| TIP3P | 2.1 | 1.5 | −0.4 | |
| TIP3P-Ew | 2.1 | 1.7 | −0.3 | |
| TIP4P-2005 | 1.8 | 2.2 | 0.0 | |
| FF03 | SPC/E | 1.2 | 3.5 | −0.6 |
| TIP3P | 1.7 | 3.0 | −0.6 | |
| TIP3P-Ew | 1.5 | 3.2 | −0.5 | |
| TIP4P-2005 | 1.0 | 3.0 | −0.4 | |
| CHARMM | SPC/E | 0.3 | 0.9 | −0.6 |
| TIP3P | 1.0 | 0.7 | −1.3 | |
| TIP3P-Ew | 0.8 | 1.7 | −0.9 | |
| TIP4P-2005 | 0.7 | 0.1 | −1.0 | |
| OPLS | SPC/E | 2.5 | 2.6 | −1.2 |
| TIP3P | 3.9 | 3.1 | −1.3 | |
| TIP3P-Ew | 3.4 | 4.0 | −1.1 | |
| TIP4P-2005 | 2.6 | 2.8 | −1.1 | |
As shown in Figure 12, of the nine specific crystallographic interactions we identified, only two are preserved across greater than 50% of the monomers in all 16 simulations: a hydrogen bond between the Tyr49 hydroxyl group and the His54 backbone oxygen across interface 1, and a salt bridge or hydrogen bond between the Lys2 side chain amino group and another monomer’s C terminus across interface 3. A hydrogen bond between the Gln37 and Asp53 side chains (interface 1) is preserved only in simulations using the CHARMM22 force field, and is preserved best when the simulations are performed with the TIP3P or TIP3P-Ewald water models. All simulations indicate that the hydrophobic interaction between Tyr14 and P60 is replaced by a hydrogen bond between the Phe15 (peptide nitrogen) and Pro60 (peptide oxygen) backbone groups. In the X-ray structure there is a water-mediated hydrogen bonding network, including a crystallographic water with 100% occupancy, connecting these backbone groups, but all simulation force fields show a preference for a protein-protein interaction. In addition, a salt bridge or hydrogen bond between Arg62 and Asp53 and a hydrogen bond between the polypeptide C terminus and Thr57 are generally maintained in all except the CHARMM trajectories. Finally, a hydrogen bond between Tyr49 and Arg62 (interface 6) is maintained in the FF99 trajectories and some FF03 and CHARMM trajectories.
Interface 3 merits special consideration as it has a large number of contacts in the X-ray structure that dissociate in the CHARMM22, OPLS, and FF03 simulations but remain intact during the FF99SB trajectories. The preceding analysis of the deformations in the distances between monomer centers of mass also implicated interface 3 as a key element in unit cell deformations. As can be seen in Figure 3, interface 3 involves interactions between a flexible hairpin linker region of one subunit (residues 7–16) and the mobile C-terminus of an adjacent subunit (residues 55–64). Substantial rearrangements can occur in these secondary structures over the course of the trajectories: Figure 13 shows that the protein’s C terminus remains close to its conformation in the X-ray structure in simulations with the FF99SB force field, but adopts an increasingly large range of conformations in simulations with the FF03, CHARMM22, and OPLS force fields. Despite the variability of this region of the protein, Figure 12 shows that the hydrogen bond between the Lys2 side chain and the C terminus (His64) remains intact even in simulations with the OPLS force field—the flexibility of the side chain apparently compensates for the variability of the C terminus. However, the hydrogen bond between the Thr57 peptide nitrogen and the other monomer’s C terminus is more likely to be broken, particularly in the CHARMM22 and OPLS simulations.
Figure 13. Interactions at interface 3 after 100 ns of simulated dynamics.
In interface 3, the N- and C-termini of interacting monomers may be very important for maintaining the lattice structure, giving rise to two of the specific hydrogen bonding lattice contacts. Above, all instances of interface 3 from the final frame of a representative simulation carried out with each force field are shown in transparent relief, superimposed by quaternion alignment of backbone atoms against the X-ray structure (shown in solid color). The FF99SB force field again stands out in this comparison: all instances of the interface are tightly clustered around the X-ray structure, as opposed to simulations with FF03, CHARMM22, and OPLS force fields which portray the N- and C-termini as increasingly disordered. Despite the disorder, the hydrogen bond between Lys2 and the C-terminus is at least 80% occupied in all simulations, and the hydrogen bond between Thr57 and the C-terminus is better than 50% occupied in all but the CHARMM22 simulations. These results may indicate that better dihedral parameters, particularly on the backbone groups, are needed for long-timescale simulations.
Analysis of the interactions between particular residues that accompany deformations at each interface and the lattice as a whole offers information that could be useful for improving the parameters of each force field, but additional studies will be needed to reach a sufficient level of understanding about which individual parameters must be changed. The peptide C terminus could fluctuate more in simulations with FF03, CHARMM22, and OPLS because the backbone hydrogen bond between Thr27 and the His64 carboxylate is not strong enough, but given the integrity of other backbone hydrogen bonds in the well-conserved α-helical and β-sheet secondary structures throughout all simulations (see Figure 6) and the propensity of all models to form more hydrogen bonds than the X-ray structure contains, we speculate that backbone dihedral parameters may be to blame. Additional simulations that hold various parts of the protein fixed, as we will discuss in more detail at the conclusion of the article, are needed to further narrow the search space for optimizing each force field’s parameter set.
4 Discussion
Of the four molecular force fields studied in this article, the FF99SB model stands out for its ability to maintain the correct unit cell dimensions and reproduce the backbone structure and atomic fluctuations of the Toxin II crystal observed in X-ray diffraction experiments. However, its success comes with many caveats. None of the force fields was able to maintain the correct distances between monomer centers of mass: in fact, all of them showed very similar deformations of the lattice in this respect. The FF99SB model’s ability to maintain five out of the eight specific contacts in the X-ray structure may be integral to its success, but like other models it showed a propensity to form more hydrogen bonds than should really exist, and a tendency to underestimate the strength of hydrophobic contacts. Clearly, it is not necessary for a force field to maintain all native contacts (or avoid the formation of non-native contacts) between protein monomers in order to preserve the protein backbone structure observed in experiments. Some of the native contacts could also have been maintained (by the FF99SB model, or others) for the wrong reasons—the hydrogen bonding interaction of Arg62 with Tyr49, for example, could maintain the juxtaposition of Arg62 and Tyr35 if it were parameterized to be too strong. In reality, the cation-π interaction between Arg62 and Tyr35 is highly favorable and makes its own contribution to the stability of the lattice structure, in the same way that other cation-π interactions are known to stabilize folded proteins.42 In our simulations, the Arg62:Tyr35 interaction is not supported by any explicit terms; other studies have concluded that explicit treatment of electronic polarization is needed to properly account for the strength of cation-π interactions. 43 Finally, the sensitivity of outcomes such as lattice deformation to individual force field parameters are not yet known, and the CHARMM22 force field obtained a number of correct results (particularly, the correlation of atomic fluctuations with experiment) if the macroscopic deformation of the lattice was removed from consideration. The FF99SB force field therefore emerges as the preferred model for simulating the structural details of Toxin II, but further optimization of any of the models may be productive.
While previous studies 10,11 identified significant effects from particular water models, defining some water models as more appropriate for particular force fields on the basis of accurate solvation free energies, we found that the choice of water model has little effect on the structural properties of the monomers and the lattice as a whole. The most significant exception to this generalization is the observation that three-point water models are more appropriate than the TIP4P-2005 model for simulations with the FF99SB force field; in this case, the conservation of structure was high enough that the effects of individual water models rose above the inaccuracies of the force field. The fact that we did not find the choice of water model to be as significant as the force field itself can be partly attributed to the precision available in our simulations, which as we have stated is much less than that available to simulations of solvated small molecules. Our results were insensitive to factors such as implicit polarization or long-range van-der Waals energy contributions, which only affect the computed heat of vaporization or the system pressure tensor and so have little to no effect on the structure despite their influence on thermodynamic quantities. We also note that the solvation enthalpies observed by Hess and co-workers are tightly clustered about particular values that depend much more on the description of the solute than the water model, which is in agreement with our findings. Even though the choice of water model had little impact on the structure, it is notable that different force fields required significantly different numbers of waters to fully hydrate the unit cell. The interaction of proteins and water molecules remains a central challenge in biomolecular modeling, even if all of the available water models behave more similarly than expected.
Some errors oberserved in this study such as unnatural hydrogen bond formation spanning all four force fields corroborate other structural validation studies. In general, the popular non-polarizable molecular force fields exhibit a tendency to over-stabilize α-helices; 44,8 the CHARMM22 force field (with CMAP corrections, as used in this study) has been shown before to bias conformational equilibria of a β-sheet protein towards α-helical structures in folding simulations. 45,9 The source of this bias is not that the force fields were over-fitted to simulate α-helical structures; none of the force fields were trained to stabilize α-helices at all. Instead, the bias may be inherent to non-polarizable models, arising from an implicit hyper-polarization of dipoles within molecules which simply reinforces the orderly hydrogen-bonding interactions that make α-helices stable in reality. This hyper-polarization may also be responsible for the excessive formation of hydrogen bonds between separate protein monomers observed in our simulations of the Toxin II lattice.
The set of simulations presented in this article is insufficient to prescibe particular changes to the force field parameters, but the results do suggest better protocols for performing structure-based validation based on crystallographic data. Global deformation of the lattice is entwined with a sampling problem (because it is such a slow process) and may also distort results such as backbone rmsds, atomic fluctuations, or side chain dihedral conformations obtained for individual monomers. Future validation studies may be able to identify the problematic force field parameters by iteratively restraining different parts of the system. Restraints on all backbone atoms, letting side chains move freely to assess a force field’s accuracy in side-chain packing, would serve as an excellent counterpart to the existing thermodynamic studies on hydration free energies of amino acid analogs. 10,11 Intramolecular retraints that maintain backbone structure and intermolecular restraints that maintain lattice contacts may be useful for identifying particular contacts within proteins or between proteins that contribute to global lattice deformations as well as deviations in the monomer structure.
Supplementary Material
Acknowledgments
This work was supported by NIH grants RR12255 and RR05969; supercomputing time was partially provided by Large Resource Allocation Committee MCA93S028. D.S. Cerutti and P.L. Freddolino gratefully acknowledge Terry P. Lybrand and Klaus Schulten for encouraging preliminary work on this project.
References
- 1.Muley L, Baum B, Smolinski M, Freindorf M, Heine A, Klebe D, Hangauer DG. Enhancement of hydrophobic interactions and hydrogen bond strength by cooperativity: synthesis, modeling, and molecular dynamics simulations of a congeneric series of thrombin inhibitors. J Med Chem [Online] 2010;53:2126–2135. doi: 10.1021/jm9016416. [DOI] [PubMed] [Google Scholar]
- 2.Williams SL, McCammon JA. Conformational dynamics of the flexible catalytic loop in mycobacterium tuberculosis 1-deoxy-D-xylulose 5-phosphate reductoisomerase. Chem Biol Drug Des [Online] 2009;73:26–38. doi: 10.1111/j.1747-0285.2008.00749.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hu X, Wang H, Ke H, Kuhlman B. Computer-based redesign of a β sandwich protein suggests that extensive negative design is not required for de novo β sheet design. Structure [Online] 2008;16:1799–1805. doi: 10.1016/j.str.2008.09.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Adcock SA, McCammon JA. Molecular dynamics: survey of methods for simulating the activity of proteins. Chem Rev [Online] 2006;106:1589–1615. doi: 10.1021/cr040426m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Karplus M, Kuriyan J. Molecular dynamics and protein function. Proc Natl Acad Sci USA [Online] 2005;102:6679–6685. doi: 10.1073/pnas.0408930102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Allen TW, Baştuğ T, Kuyucak S, Chung SH. Gramicidin A channel as a test ground for molecular dynamics force fields. Biophys J [Online] 2003;84:2159–2168. doi: 10.1016/S0006-3495(03)75022-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Paschek D, Nymeyer H, García AE. Replica exchange simulation of reversible folding/unfolding of the Trp-cage miniprotein in explicit solvent: on the structure and possible role of internal water. J Struct Biol [Online] 2007;157:524–533. doi: 10.1016/j.jsb.2006.10.031. [DOI] [PubMed] [Google Scholar]
- 8.Best RB, Hummer G. Optimized molecular dynamics force fields applied to the helix-coil transition of polypeptides. J Phys Chem B [Online] 2009;113:9004–9015. doi: 10.1021/jp901540t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Freddolino PL, Park S, Roux B, Schulten K. Force field bias in protein folding simulations. Biophys J [Online] 2009;96:3772–3780. doi: 10.1016/j.bpj.2009.02.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hess B, van der Vegt NFA. Hydration thermodynamic properties of amino acid analogues: a systematic comparison of biomolecular force fields and water models. J Phys Chem B [Online] 2006;110:17616–17626. doi: 10.1021/jp0641029. [DOI] [PubMed] [Google Scholar]
- 11.Shirts MR, Pande VS. Solvation free energies of amino acid side chain analogs for common molecular mechanics water models. J Chem Phys [Online] 2005;122:134508. doi: 10.1063/1.1877132. [DOI] [PubMed] [Google Scholar]
- 12.Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE. Improved side-chain torsion potentials for the AMBER 399SB protein force field. Proteins [Online] 2010;78:1950–1958. doi: 10.1002/prot.22711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cerutti DS, Le Trong I, Stenkamp RE, Lybrand TP. Simulations of a protein crystal: explicit treatment of crystallization conditions links theory and experiment in the streptavidin-biotin complex. Biochemistry [Online] 2008;47:12065–12077. doi: 10.1021/bi800894u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Meinhold L, Merzel F, Smith JC. Lattice dynamics of a protein crystal. Phys Rev Lett [Online] 2007;99:138101. doi: 10.1103/PhysRevLett.99.138101. [DOI] [PubMed] [Google Scholar]
- 15.Meinhold L, Smith JC. Correlated dynamics determining X-ray diffuse scattering from a crystalline protein revealed by molecular dynamics simulation. Phys Rev Lett [Online] 2005;95:218103. doi: 10.1103/PhysRevLett.95.218103. [DOI] [PubMed] [Google Scholar]
- 16.Ceccarelli M, Marchi M. Simulation of a protein crystal at constant pressure. J Phys Chem B [Online] 1997;101:2105–2108. [Google Scholar]
- 17.Krieger E, Darden TA, Nabuurs SB, Finkelstein A, Vriend G. Making optimal use of empirical energy functions: force-field parameterization in crystal space. Proteins [Online] 2004;57:678–683. doi: 10.1002/prot.20251. [DOI] [PubMed] [Google Scholar]
- 18.Halle B. Biomolecular cryocyrstallography: Structural changes during flash cooling. Proc Natl Acad Sci US A [Online] 2004;101:4793–4798. doi: 10.1073/pnas.0308315101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fontecilla-Camps JC, Habersetzer-Rochat C, Rochat H. Orthorhombic crystals and three-dimensional structure of the potent toxin II from the scorpion Androctonus australis Hector. Proc Natl Acad Sci US A [Online] 1988;85:7443–7447. doi: 10.1073/pnas.85.20.7443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Smith GD, Blessing RH, Ealick SE, Fontecilla-Camps JC, Hauptman HA, Housset D, Langs DA, Miller R. Ab initio structure determination and refinement of a scorpion protein toxin. Acta Crystallogr D [Online] 1997;53:551–557. doi: 10.1107/S0907444997005386. [DOI] [PubMed] [Google Scholar]
- 21.Housset D, Habersetzer-Rochat C, Astier J, Fontecilla-Camps JC. Crystal structure of toxin II from the scorpion Androctonus australis Hector refined at 1.3Å resolution. J Mol Biol [Online] 1994;238:88–103. doi: 10.1006/jmbi.1994.1270. [DOI] [PubMed] [Google Scholar]
- 22.Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins [Online] 2006;65:712–725. doi: 10.1002/prot.21123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.MacKerrel AD, Feig M, Brooks CL., III Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comp Chem [Online] 2004;25:1400–1415. doi: 10.1002/jcc.20065. [DOI] [PubMed] [Google Scholar]
- 24.MacKerell AD, Jr, Bashford D, Bellott M, Dunbrack RL, Jr, Evanseck J, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher IWE, Roux B, Schlenkrich M, Smith J, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B [Online] 1998;102:3586–3616. doi: 10.1021/jp973084f. [DOI] [PubMed] [Google Scholar]
- 25.Duan Y, Wu C, Chowdhury S, Lee MC, Xiong G, Zhang W, Yang R, Cieplak P, Luo R, Lee T, Caldwell J, Wang J, Kollman P. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J Comput Chem [Online] 2003;24:1999–2012. doi: 10.1002/jcc.10349. [DOI] [PubMed] [Google Scholar]
- 26.Jorgensen WL, Maxwell DS, Tirado-Rives J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc [Online] 1996;118:11225–11236. [Google Scholar]
- 27.Jorgensen WL, Chandrasekhar D, Madura JD, Impey RW, Klein ML. Comparison of simple potential functions for simulating liquid water. J Chem Phys [Online] 1983;79:926–935. [Google Scholar]
- 28.Berendsen HJC, Grigera JR, Straatsma TP. The missing term in effective pair potentials. J Phys Chem [Online] 1987;91:6269–6271. [Google Scholar]
- 29.Price DJ, Brooks CL., III A modified TIP3P water potential for simulation with Ewald summation. J Chem Phys [Online] 2004;121:10096–10103. doi: 10.1063/1.1808117. [DOI] [PubMed] [Google Scholar]
- 30.Abascal JLF, Vega C. A general purpose model for the condensed phases of water: TIP4P/2005. J Chem Phys [Online] 2005;123:234505. doi: 10.1063/1.2121687. [DOI] [PubMed] [Google Scholar]
- 31.Miller R, Gallo SM, Khalak HG, Weeks CM. SnB: crystal structure determination via Shake-and-Bake. J Appl Cryst [Online] 1994;27:613–621. [Google Scholar]
- 32.Case DA, Cheatham TE, III, Darden TA, Gohlke H, Luo R, Merz KM, Jr, Onufriev A, Simmerling C, Wang B, Woods RJ. The AMBER biomolecular simulation programs. J Comput Chem [Online] 2005;26:1668–1688. doi: 10.1002/jcc.20290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Saigal S, Pranata J. Monte Carlo simulations of guanidinium acetate and methylammonium acetate ion pairs in water. Bioorg Chem[Online] 1997;25:11–21. [Google Scholar]
- 34.Nygaard TP, Rovira C, Peters GH, Jensen MØ. Ammonium Recruitment and Ammonia Transport by E. coli Ammonia Channel AmtB. Biophys J [Online] 2006;91:4401–4412. doi: 10.1529/biophysj.106.089714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cerutti DS, Le Trong I, Stenkamp RE, Lybrand TP. Dynamics of the streptavidin-biotin complex in solution and in its crystal lattice: distinct behavior revealed by molecular simulations. J Phys Chem B [Online] 2009;113:6971–6985. doi: 10.1021/jp9010372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kalé L, Schulten K. Scalable molecular dynamics with NAMD. J Comp Chem [Online] 2005;26:1781–1802. doi: 10.1002/jcc.20289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Izaguirre JA, Catarello DP, Wozniak JM, Skeel RD. Langevin stabilization of molecular dynamics. J Chem Phys [Online] 2001;114:2090–2098. [Google Scholar]
- 38.Wu Y, Tepper HL, Voth GA. Flexible simple point-charge water model with improved liquid-state properties. J Chem Phys [Online] 2006;124:024503. doi: 10.1063/1.2136877. [DOI] [PubMed] [Google Scholar]
- 39.Cerutti DS, Duke RE, Freddolino PL, Fan H, Lybrand TP. A vulnerability in several popular molecular dynamics packages concerning Langevin and Andersen dynamics. J Chem Theory Comput [Online] 2008;4:1669–1680. doi: 10.1021/ct8002173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Brooks BR, Janežič D, Karplus M. Harmonic analysis of large systems. I. Methodology. J Comput Chem [Online] 1995;16:1522–1542. [Google Scholar]
- 41.Janežič D, Brooks BR. Harmonic analysis of large systems. II. Comparison of different protein models. J Comput Chem [Online] 1995;16:1543–1553. [Google Scholar]
- 42.Gallivan JP, Dougherty DA. Cation-π interactions in structural biology. Proc Natl Acad Sci US A [Online] 1999;96:9459–9464. doi: 10.1073/pnas.96.17.9459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Aschi M, Mazza F, Di Nola A. Cation-π interactions between ammonium ion and aromatic rings: an energy decomposition study. J Mol Struc-Theochem [Online] 2002;587:177–188. [Google Scholar]
- 44.Best RB, Buchete N, Hummer G. Are current molecular dynamics force fields too helical? Biophys J [Online] 2008;95:L07–09. doi: 10.1529/biophysj.108.132696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Freddolino PL, Liu F, Gruebele M, Schulten K. Ten-microsecond MD simulation of a fast-folding WW domain. Biophys J [Online] 2008;94:L75–77. doi: 10.1529/biophysj.108.131565. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.