Abstract
Background
Streptomycin (SM) is a broad spectrum antibiotic and is an important component of any anti-tuberculosis therapy regimen. Several mechanisms have been proposed to explain the emergence of resistance but still our knowledge is inadequate. Proteins form a very complex network and drugs are countered by their modification/efflux or over expression/modification of targets. As proteins manifest most of the biological processes, these are attractive targets for developing drugs, immunodiagnostics or therapeutics. The aim of present study was to analyze and compare the protein profile of whole cell extracts from Mycobacterium tuberculosis clinical isolates susceptible and resistant to SM.
Results
Two-dimensional gel electrophoresis (2DE) and matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry was employed for analyzing the protein profiles. Homology and in silico characterization for identified proteins was assessed using BLAST, InterProScan and KEGG database searches. Computational studies on the possible interactions between SM and identified proteins were carried out by a battery of online servers and softwares, namely, CLUSTALW (KEGG), I-TASSER, VMD, PatchDock and FireDock. On comparing 2DE patterns, nine proteins were found consistently overexpressed in SM resistant isolates and were identified as Rv0350, Rv0440, Rv1240, Rv3075c, Rv2971, Rv3028c, Rv2145c, Rv2031c and Rv0569. In silico docking analysis showed significant interactions of SM with essential (Rv0350, Rv0440 and Rv2971) and non essential (Rv1240, Rv3075c and Rv2031c) genes.
Conclusions
The computational results suggest high protein binding affinity of SM and suggested many possible interactions between identified proteins and the drug. Bioinformatic analysis proves attributive for analysis of diversity of proteins identified by whole proteome analysis. In-depth study of the these proteins will give an insight into probable sites of drug action other than established primary sites and hence may help in search of novel chemotherapeutic agents at these new sites as inhibitors.
Background
Tuberculosis is one of the most challenging infectious diseases. Globally, 9.2 million new cases and 1.7 million deaths occur due to this disease [1]. Its impact on public health is further aggravated by co-infection with human immunodeficiency virus, emergence of multi-drug resistant strains and reactivation of the dormant bacteria. Attempt for primary prevention using Bacillus Calmette Guerin (BCG) and other integral vaccines have generally been disappointing though some subunit vaccines are under trial [2]. The excessive emergence of drug resistant tuberculosis has stimulated interest in the understanding of the underlying mechanisms of drug resistance in Mycobacterium tuberculosis and significant progress has been made in this field [3]. Streptomycin (SM) is first line anti-tuberculosis drug and preferred for treatment of relapses. It inhibits protein synthesis in susceptible bacteria by interacting with steps of translation. Modification of the primary target of the drug by mutations in the genes encoding either the 16 S rRNA or S12 ribosomal proteins primarily affects the activity of SM and is clinically significant in Mycobacterium species [4,5]. However, nearly one third of resistant isolates of M. tuberculosis do not have these mutations suggesting the involvement of some other mechanism(s) responsible for resistance [6-8]. Role of efflux pumps in SM resistance have been demonstrated by inhibitor assays [9]. Recently an acetyltransferase gene Rv0262c [10] and mutation within gidB gene [11] have also been reported to confer low level SM resistance in M. tuberculosis. Still there is a scope for unraveling more underlying mechanisms for SM resistance.
Two-dimension gel electrophoresis (2DE) along with mass spectrometry is a powerful and direct tool to study differential protein expression in cells. Identification and characterization of mycobacterial proteins as drug targets, diagnostics and vaccine candidates have been popular research objectives, but comparative proteome profiling of drug susceptible and resistant isolates remain unexplored in relation to SM. Therefore, the aim of the present study was to compare the protein profiles of cell extracts from M. tuberculosis isolates sensitive and resistant to SM.
Results
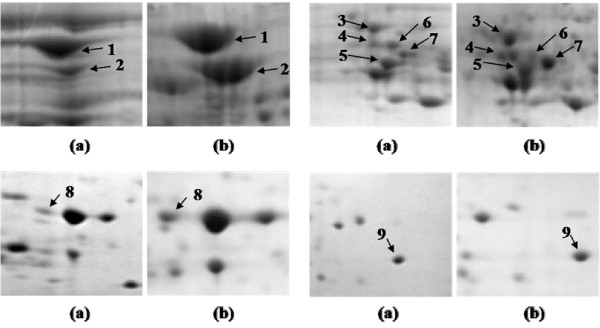
This study was focused on the proteins overexpressed in SM resistant M. tuberculosis clinical isolates. Protein profiles were compared by 2DE run in triplicates for each isolate. 2DE patterns for SM susceptible and SM resistant isolates are shown as Additional File 1 and Additional File 2 respectively. Spots appearing consistently overexpressed among resistant isolates were further processed. We found nine protein spots to be overexpressed in SM resistant isolates in comparison to sensitive isolates (Figure 1). Magnified regions of these overexpressed proteins are shown in Figure 2. These protein spots were further identified as DnaK, 60 kDa chaperonin2, Malate dehydrogenase, Probable oxidoreductase, Electron transfer flavoprotein subunit alpha, Antigen 84, 14 kDa antigen and two hypothetical proteins by matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (Table 1) and identity also confirmed by MS/MS (Table 2). The level of over expression has been represented as densitometric ratio in Table 1. All the spots except one exhibited at least two fold over expression. However, Antigen 84 overexpressed maximally more than four fold.
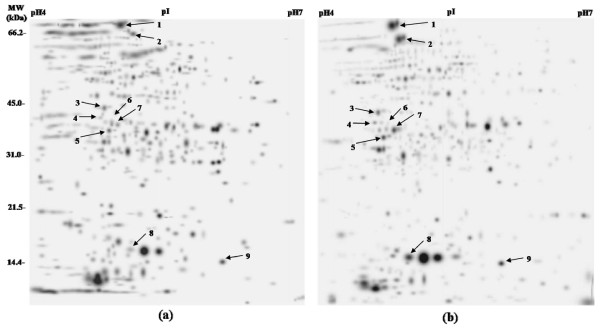
Figure 1.
Composite images of 2 D gels of proteins extracted from M. tuberculosis clinical isolates: (a) SM susceptible, (b) SM resistant. Spots indicated by arrow were excised and analyzed by MS.
Figure 2.
Magnified regions of 2 D gels showing the overexpressed proteins (a) sensitive isolate; (b) SM resistant isolate.
Table 1.
Details of overexpressed proteins identified by MALDI-TOF mass spectrometry in SM resistant M. tuberculosis clinical isolates
| Spot No.† | Accession No. | Proteins identified | Mascot score | Nominal mass (Da) | pI | No. of peptides matched | Sequence coverage (%) | ORF No. | Densitometric ratio of protein expression between sensitive vs. resistant isolates |
|---|---|---|---|---|---|---|---|---|---|
| 1. | P0A5B9 (DNAK_MYCTU) | Chaperone protein dnaK | 165 | 66790 | 4.85 | 22 | 38 | Rv0350 | 1: 1.48 |
| 2. | P0A520 (CH602_MYCTU) | 60 kDa chaperonin 2 | 85 | 56692 | 4.85 | 12 | 27 | Rv0440 | 1: 3.17 |
| 3. | P0A5J6 (MDH_MYCTU) | Malate dehydrogenase | 103 | 34358 | 4.65 | 11 | 51 | Rv1240 | 1: 2.45 |
| 4. | P95083 (P95083_MYCTU) | Hypothetical protein | 98 | 33194 | 4.73 | 9 | 28 | Rv3075c | 1: 3.69 |
| 5. | P95124 (Y2971_MYCTU) | Probable oxidoreductase | 97 | 30516 | 4.70 | 11 | 47 | Rv2971 | 1: 3.10 |
| 6. | O53275 (ETFA_MYCTU) | Electron transfer flavoprotein subunit alpha (α-ETF) | 118 | 31672 | 4.71 | 11 | 42 | Rv3028c | 1: 2.97 |
| 7. | P0A5N2 (AG84_MYCTU) | Antigen 84 | 132 | 28260 | 4.80 | 9 | 44 | Rv2145c | 1: 4.27 |
| 8. | P0A5B7 (ACR_MYCTU) | 14 kDa Antigen (16 kDa Antigen, HSP16.3) | 117 | 16217 | 5.00 | 9 | 70 | Rv2031c | 1: 2.69 |
| 9. | O53766 (Y0569_MYCTU) | Hypothetical protein | 80 | 9517 | 5.70 | 5 | 54 | Rv0569 | 1: 2.43 |
†Spot number of protein marked in Figure 1 & Figure 2.
Table 2.
MALDI-TOF/TOF (MS/MS) analysis of all nine overexpressd proteins in SM resistant M. tuberculosis clinical isolates
| Spot No. | Peak Mass (Da) | Protein Identified | Nominal mass | Mascot Score | pI | Sequence of peptide | ORF No. |
|---|---|---|---|---|---|---|---|
| 1 | 1062.611 | Chaperone protein dnaK | 66790 | 33 | 4.85 | RTTPSIVAFARN | Rv0350 |
| 1226.711 | Chaperone protein dnaK | 66790 | 34 | 4.85 | KDAGQIAGLNVLRI | Rv0350 | |
| 1567.931 | Chaperone protein dnaK | 66790 | 61 | 4.85 | KLLGSFELTGIPPAPRG | Rv0350 | |
| 1645.962 | Chaperone protein dnaK | 66790 | 40 | 4.85 | RIVNEPTAAALAYGLDKG | Rv0350 | |
| 2613.426 | Chaperone protein dnaK | 66790 | 60 | 4.85 | RSETFTTADDNQPSVQIQVYQGERE | Rv0350 | |
| 2 | 940.555 | 60 kDa Chaperonin 2 | 56692 | 58 | 4.85 | KDLLPLLEKV | Rv0440 |
| 1503.710 | 60 kDa Chaperonin 2 | 56692 | 45 | 4.85 | KGYISGYFVTDPERQ | Rv0440 | |
| 1658.900 | 60 kDa Chaperonin 2 | 56692 | 89 | 4.85 | KQIAFNSGLEPGVVAEKV | Rv0440 | |
| 1790.868 | 60 kDa Chaperonin 2 | 56692 | 20 | 4.85 | KDETTIVEGAGDTDAIAGRV | Rv0440 | |
| 2075.110 | 60 kDa Chaperonin 2 | 56692 | 65 | 4.85 | KTDDVAGDGTTTATVLAQALVRE | Rv0440 | |
| 2203.240 | 60 kDa Chaperonin 2 | 56692 | 114 | 4.85 | KKTDDVAGDGTTTATVLAQALVRE | Rv0440 | |
| 2317.275 | 60 kDa Chaperonin 2 | 56692 | 62 | 4.85 | KVVVTKDETTIVEGAGDTDAIAGRV | Rv0440 | |
| 3 | 1693.989 | Malate Dehydrogenase | 34358 | 52 | 4.65 | RLASGSLLGPDRPIELRL | Rv1240 |
| 1722.957 | Malate Dehydrogenase | 34358 | 38 | 4.65 | KVAVTGAAGQIGYSLLFRL | Rv1240 | |
| 1879.944 | Malate Dehydrogenase | 34358 | 56 | 4.65 | KGGNWTIVSGLEIDEFSRG | Rv1240 | |
| 2423.360 | Malate Dehydrogenase | 34358 | 52 | 4.65 | RVGVTGNPANTNALIAMTNAPDIPRE | Rv1240 | |
| 4 | 995.545 | Hypothetical Protein Rv3075c | 33194 | 30 | 4.73 | RLAFGIGDFRR | Rv3075c |
| 1016.503 | Hypothetical Protein Rv3075c | 33194 | 26 | 4.73 | KEFFAEFARD | Rv3075c | |
| 1322.619 | Hypothetical Protein Rv3075c | 33194 | 63 | 4.73 | RWFGDGNADWVRI | Rv3075c | |
| 1491.919 | Hypothetical Protein Rv3075c | 33194 | 32 | 4.73 | RLPNVPIVALVETARG | Rv3075c | |
| 1583.780 | Hypothetical Protein Rv3075c | 33194 | 64 | 4.73 | RDTGFGEDPATLAYARS | Rv3075c | |
| 1648.036 | Hypothetical Protein Rv3075c | 33194 | 45 | 4.73 | KRLPNVPIVALVETARG | Rv3075c | |
| 5 | 897.627 | Probable Oxidoreductase | 30516 | 15 | 4.70 | KTPAQVLLRW | Rv2971 |
| 1105.658 | Probable Oxidoreductase | 30516 | 27 | 4.70 | KLATPDQGFTRS | Rv2971 | |
| 1368.911 | Probable Oxidoreductase | 30516 | 72 | 4.70 | RWNLQLGNAVVVRS | Rv2971 | |
| 1382.701 | Probable Oxidoreductase | 30516 | 12 | 4.70 | RWNLQLGNAVVVRS | Rv2971 | |
| 6 | 1069.609 | Electron transfer flavoprotein subunit alpha | 31672 | 43 | 4.71 | KVAPQLTEAIKA | Rv3028c |
| 1341.748 | Electron transfer flavoprotein subunit alpha | 31672 | 56 | 4.71 | RIGSGLLVDVVDVRE | Rv3028c | |
| 1577.902 | Electron transfer flavoprotein subunit alpha | 31672 | 113 | 4.71 | MAEVLVLVEHAEGALKK | Rv3028c | |
| 1706.093 | Electron transfer flavoprotein subunit alpha | 31672 | 65 | 4.71 | MAEVLVLVEHAEGALKKV | Rv3028c | |
| 1973.063 | Electron transfer flavoprotein subunit alpha | 31672 | 69 | 4.71 | RAAVDSGYYPGQFQVGQTGKT | Rv3028c | |
| 2024.322 | Electron transfer flavoprotein subunit alpha | 31672 | 67 | 4.71 | KTVSPQLYIALGISGAIQHRA | Rv3028c | |
| 2692.697 | Electron transfer flavoprotein subunit alpha | 31672 | 14 | 4.71 | KNGLVLVIDGQLWTITEFQHVKPGKG | Rv3028c | |
| 7 | 1088.702 | Antigen 84 | 28260 | 22 | 4.80 | RLIEENSDLRQ | Rv2145c |
| 1171.748 | Antigen 84 | 28260 | 32 | 4.80 | RANAEQILGEARH | Rv2145c | |
| 1807.228 | Antigen 84 | 28260 | 38 | 4.80 | RLKTYLESQLEELGQRG | Rv2145c | |
| 1817.241 | Antigen 84 | 28260 | 33 | 4.80 | RVLSLAQDTADRLTNTAKA | Rv2145c | |
| 8 | 885.507 | 14 kDa antigen | 16217 | 21 | 5.00 | MATTLPVQRH | Rv2031c |
| 1162.563 | 14 kDa antigen | 16217 | 47 | 5.00 | RSEFAYGSFVRT | Rv2031c | |
| 1715.053 | 14 kDa antigen | 16217 | 28 | 5.00 | KGILTVSVAVSEGKPTEKH | Rv2031c | |
| 1869.098 | 14 kDa antigen | 16217 | 46 | 5.00 | RAELPGVDPDKDVDIMVRD | Rv2031c | |
| 9 | 929.516 | Hypothetical protein Rv0569 | 9517 | 22 | 5.70 | KVGDWLVIKG | Rv0569 |
| 1109.493 | Hypothetical protein Rv0569 | 9517 | 12 | 5.70 | KGATIDQPDHRG | Rv0569 | |
| 1163.530 | Hypothetical protein Rv0569 | 9517 | 4 | 5.70 | RSSDGSPPYVVRW | Rv0569 | |
| 1269.646 | Hypothetical protein Rv0569 | 9517 | 17 | 5.70 | RFGAVQSAILHARG | Rv0569 |
Results of computational analysis of all nine overexpressed proteins using different softwares and servers are as follows.
BLAST Analysis
Blastp analysis was carried out for two hypothetical proteins and two proteins with unknown functions. Hypothetical protein (Rv3075c) was found highly conserved (99 ± 0.05% identical) in few species of mycobacteria and in others it acts as citrate lyase subunit beta-like protein (CitE) or HpcH/HpaI aldolase. Other one hypothetical protein (Rv0569) was also found highly conserved in mycobacteria and other microbes and significant function could not be assigned except in few microbes as signal-transduction protein (Frankia and Nocardioides sps.) and DNA-binding protein (Streptomyces coelicolor &S. lividans). Probable oxidoreductase (Rv2971) appeared highly conserved in some species of mycobacteria, some as 2, 5-diketo-D-gluconic acid reductase A (M. avium) or morphine 6-dehydrogenase (M. smegmatis). Probable electron transfer flavoprotein alpha-subunit (fixb) was also highly conserved in mycobacteria and other microbes.
Phylogenetic analysis
CLUSTALW analysis was performed for all overexpressed proteins. Rv0350, Rv0440 and Rv1240 were highly conserved among all organisms and Rv0350 had 51% homology with human heat shock 70 kDa protein, Rv0440 showed 45% homology with human heat shock 60 kDa protein-1 and Rv1240 exhibited 48.93% homology with human malate dehydrogenase. Rv3075c had 20.19% homology with CitE of human, 86.31% with M. marinum & 85.66% with M. ulcerans, Rv2971 revealed 34% homology with human aldo-keto reductase and Rv3028c had 39% homology with human electron transfer flavoprotein. Rv2145c had 18.34% homology with ankyrin repeat domain 24 protein of human. Rv2031c had 29.86% homology with HSP20/alpha crystallin family protein of M. avium and 20% homology with 18 kDa antigen of M. leprae. However it showed 17% homology with human outer dense fiber of sperm tails protein. The hypothetical protein (Rv0569) did not show homology with any human protein.
InterProScan analysis
InterProScan analysis of Rv0350 showed nine signature motifs [SPRINT: PR00301] (Figure 3a) and motifs 1, 5 and 6 were most conserved [INTERPRO: IPR018181]. One peptide binding domain [SUPERFAMILY: SSF100920] and two actin like ATPas domains were also present in it [SUPERFAMILY: SSF53067]. Rv0440 showed five conserved motifs [SPRINT: PR00298], one conserved site (from residues 403-414) for 60 kDa Chaperonin [INTERPRO: IPR018370] (Figure 3b) and GroEL apical domain like region from residues 182-374 [SUPERFAMILY: SSF52029]. Rv1240 revealed one characteristic motif which provides a signature for L-lactate/malate dehydrogenase [PFAM: PF00056], one active site from residues 156-168 [PROSITE: PS00068] (Figure 3c) and NAD (P)-binding Rossmann fold domains [SUPERFAMILY: SSF51735]. Rv3075c showed characteristic HpcH-HpaI motifs [PFAM: PF03328], phosphoenolpyruvate/pyruvate domain from residues 22 to 239 [SUPERFAMILY: SSF51621] and citrate lyase beta subunit domain from residues 1-288 which provides a signature for lyase/aldolase activity [UNIPROT: PIRSF015582]. Rv2971 showed five Aldo/keto reductase subgroups 43-67, 99-117, 131-148, 165-194 and 202-226 [SPRINT: PR00069] out of which one is conserved site of aldo/keto reductase from residues 131-148 [PROSITE: PS00062] (Figure 3e). Rv3028c showed alpha/beta-subunit motif from residues 4-125 [PFAM: PF01012], one alpha subunit from residues 197-227 [PFAM: PF00766] and one conserved site found from residues 257-283 in alpha subunit at C-terminal side [PROSITE: PS00696] (Figure 4a). Rv2145c showed divIVA motif from residues 3-61 [PFAM: PF05103] and DivIVA domain from residues 3-39 [CMR: TIGR03544] (Figure 4b). Rv2031c confirmed its relation with Hsp20/alpha crystallin family [PFAM: PF00011], heat shock hsp20 proteins family profile [PROSITE: PS01031] and HSP20-like chaperone [SUPERFAMILY: SSF49764]. Amino acid residues were almost common for showing its relation with all three families. Signature for chaperone proteins and HSP20 family motif is also present in the same region from residues 22-142 [PANTHER: PTHR11527]. Rv0569 showed domain of unknown function DUF1918 from residues 1-58 [PFAM: PF08940] (Figure 4c). None of the nine proteins had signal peptide or transmembrane domains neither they had any site for post translational modifications.
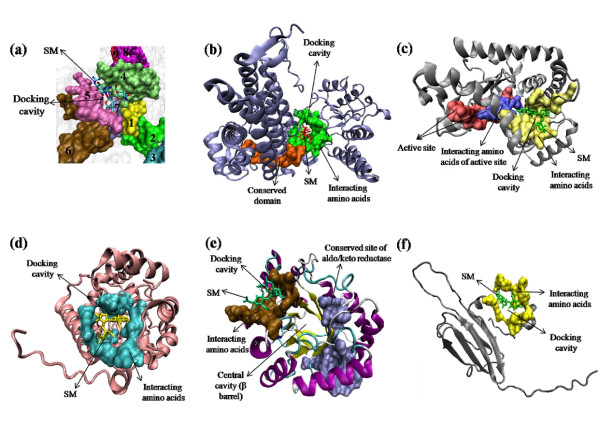
Figure 3.
3D models of overexpressed proteins showing docking with SM. Residues constituting interacting site, active site and conserved site are represented as space filled models with rest of the structure represented by cartoon structures. (a) Rv0350: Nine motifs are marked by numerals, docking cavity and SM are indicated by arrows and motifs 4, 5 & 7 are interacting with SM. (b) Rv0440: Red coloured SM, green coloured cavity and orange coloured conserved domain has been marked. Conserved domain is in the close vicinity of interacting site. (c) Rv1240: SM (green) is interacting with active site residues (blue) and other residues in the close vicinity (yellow) in the cavity. (d) Rv3075c: SM (yellow) interacting clearly with the central cavity residues (blue) of the globular protein. (e) Rv2971: SM (green) binding at the opposite side (brown) from conserved site of aldo/keto reductase (light blue) in the protein. Central cavity is present in the middle of complete β-barrel. (f) Rv2031c: SM (green) interacting with the outer part (yellow) of the protein in place of conserved HSP20-like chaperone domain.
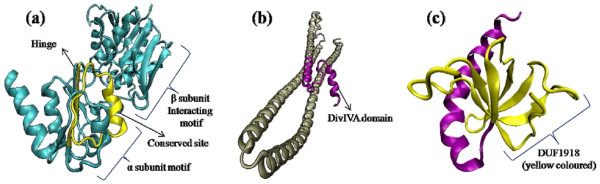
Figure 4.
3D models of proteins found non-interacting with SM represented by cartoon structures. (a) Rv3028c: Protein contains one α-subunit motif and one β-subunit interacting motif and one conserved site in the α-subunit motif. (b) Rv2145c: Whole protein consists of only α-helix and contains DivIVA domain indicating its role in cell division/cell shape. (c) Rv0569: Domain of unknown function (DUF1918) is present in the β-barrel of protein.
3D modeling and docking
All selected 3D models showed less than 2% discrepancy from Ramachandran plot. These models were further explored for in silico docking studies to identify the binding of SM. Parameters used for selection of 3D models and their docking with SM are represented in Table 3. There were possible active cavities observed on the surface of the 3D structures. Molecular docking of SM with secondary structures of Rv0350, Rv0440, Rv1240 & Rv3075c proteins showed successful binding (Figure 3) into the central cavity of the protein and the drug molecule fits well in the cavity. However, with Rv0350 and Rv3075c, drug showed binding at the conserved active site whereas in Rv1240, SM is interacting with four residues of conserved active site and for Rv0440 drug showed binding in the close vicinity of the conserved site. In docked complex of Rv2971 and Rv2031c, binding of drug was far from the conserved/active site (Figure 3). Remaining three proteins Rv3028c, Rv2145c & Rv0569 did not show significant binding of the drug and their 3D models are shown in Figure 4.
Table 3.
3D modeling and docking parameters used for bioinformatic analysis
| Identified protein's ORF number | TM-score | RMSD value (Å) | Global energy | Attractive Vander-wall force | Repulsive Vander-wall force | ACE | Interacting amino acid residues | Remarks |
|---|---|---|---|---|---|---|---|---|
| Rv0350 | 0.75 ± 0.10 | 7.2 ± 4.2 | -45.45, | -24.49, | 5.13 | -9.21 | 11, 70, 71, 119, 121, 122, 126, 175, 177, 193-195 and 197 | Interacting amino acids belong to or are in close proximity of signature motifs 1, 4 and 5 |
| Rv0440 | 0.44 ± 0.08 | 5.4 ± 3.4 | -42.74, | -25.89, | 5.64 | -5.26 | 117, 172-175, 192, 206, 216, 212, 320, 322, 324, 326, 327, 329, 391, 395, 398, 399 and 402 | Interacting residues are in the close vicinity of the one conserved site |
| Rv1240 | 0.94 ± 0.05 | 3.2 ± 2.2 | -51.86, | -22.83, | 5.69 | -15.08 | 156-168 | Amino acid residues found interacting with SM are from the active site for function |
| Rv3075c | 0.49 ± 0.15 | 10.5 ± 4.6 | -44.95, | -21.94, | 5.20 | -10.62 | 31, 57, 159-162, 165, 198, 200, 224, 226, 278, 281, 282, 285 and 286 | amino acid residues found interacting with SM are situated at the central cavity of the molecule |
| Rv2971 | 0.93 ± 0.06 | 2.9 ± 2.1 | -41.57, | -18.69, | 6.30 | -14.35 | 30, 31, 195-200, 237-340 and 243 | amino acid residues found interacting with SM are situated far from the conserved site (residue 131 to 148) |
| Rv3028c | 0.99 ± 0.04 | 2.4 ± 1.8 | -39.04, | -21.43, | 8.55 | -10.03 | poor docking | - |
| Rv2145c | 0.30 ± 0.09 | 15.4 ± 3.4 | -35.93 | -22.04 | 22.52 | -13.70 | poor docking | - |
| Rv2031c | 0.82 ± 0.09 | 3.2 ± 2.3 | -41.10, | -26.91, | 19.30 | -14.57 | 8, 10, 14, 18, 21, 22, 25, 28, 32 and 34 | SM interacts with region exclusively conserved in genus Mycobacterium, no active site found |
| Rv0569 | 0.83 ± 0.08 | 2.1 ± 1.7 | -30.57, | -17.34, | 7.52 | -12.89 | poor docking | - |
Discussion
The 2DE profile of SM resistant M. tuberculosis clinical isolates exhibited many proteins that were overexpressed. Protein spots 1 and 2 are chaperone protein 70 kDa DnaK (Rv0350) and 60 kDa Chaperonin-2/GroEL-2 (Rv0440) respectively. These essential genes prevent misfolding and promote the refolding and proper assembly of unfolded/misfolded polypeptides generated under stress conditions. Rv0350 have characteristic peptide binding domain and ATPase domain, which indicate its role in active protein refolding and proper assembly. The most conserved motifs are interacting with docked SM which indicates that the drug may inhibit/modulate the functioning of this protein and might influence SM toxicity. In prokaryotes DnaK is known to negatively regulate/auto-regulate expression of many HSP proteins by promoting degradation of σ32 heat shock transcriptional regulator [12] and σ32 regulates over expression of other HSPs [13]. Thus blocking of DnaK activity not only positively regulates its expression but may also change HSP profile of the cell. Rv0350 has also been found overexpressed in SM induced culture of SM mono-resistant clinical isolate of M. tuberculosis [14]. Rv0440 (GroEL) is the second copy of 60 kDa Chaperonin in M.tuberculosis genome. This gene is situated away from classic GroEL-Cpn10 operon and suggests a specialised regulation in M.tuberculosis. SM showed binding in the close vicinity of the conserved site on the apical domain, which is a peptide binding domain [15] and therefore it may be predicted that it could be a possible inhibitor/modulator. Duplicate copies of this gene suggest its importance as an energy independent chaperonin in slow metabolism efficient genome of M. tuberculosis [16]. Over expression of this gene could be to compensate inhibited/modulated molecules or to neutralise drug by binding. Further, spot 8 corresponding to Rv2031c encodes 14 kDa antigen (HSP16.3/HSPX) which is a member of the small heat-shock protein family of chaperones. It has been shown to be induced under oxygen-deficient conditions [17]. Its role in maintenance of long term viability during latent, asymptomatic infections and in replication during initial infection has also been proposed. Docking studies revealed that the SM binds to a demarcated cavity comprising of 10 amino acids though the interacting amino acids do not belong to any known activity region but these are highly conserved among mycobacteria. These leads can be exploited for better understanding of function of these domains by employing inhibition studies.
Spot 3 (Rv1240) encodes malate dehydrogenase, which is involved in the conversion of malate to oxaloacetate. Role of this protein in M. tuberculosis drug resistance remains to be elucidated. SM interacts with the amino acids of conserved active site of Rv1240, which suggests that the drug can affect the activity of this enzyme. Spot 5 (Rv2971) belonging to oxidoreductase of aldo/keto reductase family is probably involved in cellular metabolism. Previous studies also found that this protein was differentially expressed between BCG and H37Rv and was regarded as a candidate antigen for development of novel vaccine [18,19]. Furthermore, this protein has also been reported to be differentially expressed between isoniazid (INH) susceptible and resistant strains [20]. Residues found interacting with SM were situated far from conserved site and therefore it is suggested that this protein might be playing a secondary role in imparting resistance.
Spot 6 (Rv3028c) is an electron transfer flavoprotein alpha subunit. Bioinformatic analysis indicated that alpha subunit provides a signature for the electron transfer flavoprotein family. The electron transfer flavoprotein serves as a specific electron acceptor for other dehydrogenases. It transfers the electrons to the main respiratory chain via ETF-ubiquinone oxidoreductase (ETF dehydrogenase) [21] and participates in the oxidation of fatty acids [22]. The 3D structure of Rv3028c adopts a typical bi-lobed structure with four alpha helices each packing against the hydrophobic beta sheet comprising of 5 plates in alpha subunit motif and 7 plates in beta subunit interacting motif. No significant in silico interactions with SM were found. Further studies regarding substrate specificity & interacting proteins might reveal a relation with SM resistance. Spot 7 (Rv2145c) encodes antigen 84 and also corresponds to wag31 which was originally identified as antigen of pathogenic mycobacteria that is recognized by serum from tuberculosis patients [23]. Studies indicate that this gene is a homologue of the cell shape/cell division protein DivIVA and one of the substrates of PknA and PknB [24]. Orthologs of Rv2145c have also been found as immunogenic, cell division initiation protein or secreted antigen Wag31 in some microbes. Rv3028c & Rv2145c were also found overexpressed in INH resistant M. tuberculosis isolates [20]. Its 3D structure showed that the protein has a bi-lobbed ribbon structure comprising mostly of alpha helices & the conserved site of the protein had no evident interaction with SM.
Two protein spots 4 (Rv3075c) and 9 (Rv0569) which encode hypothetical protein could not be assigned any function. Phylogenetic analysis of these two proteins revealed existence of their homologous sequences in other mycobacterial species with known as well as unknown functions and reflected dynamic and interesting scenarios of evolutionary importance. Rv3075c, besides showing homology with hypothetical protein, also showed homology with CitE of some mycobacterial species. While the bacterial citrate lyase is a heterotrimeric complex with three subunits, the M. tuberculosis genome does not contain α and γ subunits of this complex, implying that M. tuberculosis CitE act differently from other bacterial CitE proteins [25]. These data hint that the biochemical function of the M. tuberculosis and human CitE may differ from other bacterial CitE proteins, and that M. tuberculosis CitE may be critical for pathogenesis, encompassing part of a novel pathway for fatty acid biosynthesis or anaerobic energy metabolism [26].
SM docking with Rv3075c revealed that the drug binds to the active site of the protein and interacts with 16 residues, out of which, one residue (162, Asp) has been reported as a part of active site in E. coli [27,28]. Thus it is suggested that the SM is binding at the active site of Rv3075c. It is assumed that the drug might interfere with protein function. blastp analysis of the other hypothetical protein (Rv0569) did not show any significant identity (exhibited homology with hypothetical proteins). Phylogenetic and blastp analysis exhibited that Rv0569 might work as signal transduction protein or DNA binding protein but no function has been assigned. 3D model of this protein showed semi-lunar beta barrel with a very compact structure and an alpha helix protruding out instead of packing against hydrophobic beta plates. Rv0569 showed no significant binding with SM. It is quite likely that being a possible signaling protein it might modulate the expression of other proteins.
Conclusions
To conclude, this study has employed proteomic approach, which is a direct method, to identify proteins from resistant M. tuberculosis isolates compared to sensitive isolates. Nine protein spots were consistently overexpressed in SM resistant isolates. We expect that these proteins might be contributing in conferring resistant phenotype to the isolates. Further these proteins were subjected to advanced bioinformatics analysis to generate an understanding of the subtle relation of SM with the overexpressed proteins. Homology searches and InterProScan generated insights to the possible functions and essential domains of the proteins. Rv0350, Rv0440, Rv1240, Rv3075c, Rv2971 and Rv2031c showed significant interaction in silico with SM thus their over expression in the resistant isolates could be compensating the inhibited/modulated molecules. Other proteins which are overexpressed but do not exhibit good binding with drug might be indirectly associated with SM. The elucidated mechanisms and associations may be further exploited to develop newer therapeutic agents derived from SM.
Methods
Mycobacterial growth and drug susceptibility testing
Six SM resistant and three sensitive (to five first line drugs) M. tuberculosis clinical isolates were obtained from Mycobacterial Repository Centre of our Institute. Susceptibility testing was performed by conventional LJ proportion method [29]. Bacteria were grown in Sauton's liquid medium at 37°C for four weeks (late log phase).
Preparation of mycobacterial cell extract
Mycobacterial cell extract was prepared according to modified protocol of Brodie et al., [30]. Cells were washed three times with normal saline and then suspended in sonication buffer (50 mM Tris-HCl containing 10 mM MgCl2, 0.1% sodium azide, 1 mM PMSF and 1 mM EGTA; pH 7.4) at a concentration of 1 g wet cell mass per 5 ml and then broken by intermittent sonication for 15 min at 4°C using sonicator (Sonics & Materials Inc, Newtown, CT, USA). The homogenate was centrifuged at 12,000 g for 20 min at 4°C. Pellets were discarded and supernatant was stored at -70°C until used.
Protein precipitation with SDS-TCA-acetone
Cell extracts were treated with 1% SDS and then subjected to trichloro acetic acid (TCA)-acetone precipitation procedure [31]. 10% TCA was added to the cell extract, the mixture was incubated at -20°C overnight and then precipitated protein was collected by centrifugation (18,000 g, 4°C, 15 min). It was again washed twice with 100% ice cold acetone and allowed to air dry. The protein pellet was suspended in appropriate volume of two-dimensional rehydration buffer (BIO-RAD, Hercules, CA, USA). Protein concentration was estimated using the Bradford assay [32].
Two-dimensional gel electrophoresis (2DE)
Isoelectric focusing (IEF) was carried out using the method of 'in gel rehydration' [33] with slight modifications as described previously [34]. Immobilized pH gradient (IPG) strips of pH 4-7 and length 17 cm (BIO-RAD, Hercules, CA, USA) were rehydrated overnight at 20°C with 500 μg protein which was mixed with rehydration buffer. Strips were then focused on an IEF unit PROTEAN IEF Cell (BIO-RAD, Hercules, CA, USA) at 20°C using the following four step program: a) 0-250 V for 2 h in linear mode; b) 250 V constant for 2 h in rapid mode; c) 250-5000 V for 4 h in linear mode; and d) 5000 V constant until 35 kVh reached. The current limit was set at 50 μA per strip. After IEF, IPG strips were equilibrated for 15 min in equilibration buffer I (6 M urea, 2% SDS, 0.375 M Tris; pH 8.8, 20% glycerol) containing 130 mM dithiothreitol (DTT) followed by equilibration buffer II containing 135 mM iodoacetamide instead of DTT for 15 min.
Proteins were separated in second dimension on 12% SDS-PAGE [35] in a vertical electrophoretic dual gel unit PROTEAN II XI (BIO-RAD, Hercules, CA, USA) at constant voltage of 250 V for 5-6 h and gels were stained with coomassie brilliant blue R250 to visualize proteins. Images of gels were acquired by Chemidoc (BIO-RAD, Segrate [Milan], Italy) using Quantity One software (BIO-RAD, Hercules, CA, USA). 2 D gels were analysed using PDQuest Advanced software (version 8.0) (Bio-Rad, Hercules, CA, USA). Images were analysed using stepwise spot detection and spot matching followed by differential expression analysis. PDQuest employs Student t-test and enumerates spots with differential intensity of significant levels. Resultant composite images for susceptible and resistant isolates were manually checked for artifactual spots, merged spots, and missed spots (Figure 1) and few spots with more isolate specific variability were omitted in the down stream processing. Equal amount of protein was loaded in all gels and experiments were repeated at least three times.
In-gel digestion with trypsin
Method of Shevchenko et al., [36] was followed with slight modifications as described earlier [34]. Protein spots of interest were excised from gels using spot picker 'Investigator ProPic' (Genomic Solutions, Huntingdon, UK) and collected in 96 well PCR plate. Digestion of proteins and spotting of peptides on MALDI-TOF target plate was carried out using protein digester 'Investigator ProPrep' (Genomic Solutions, Huntingdon, UK). The gel plugs were destained and dehydrated by washing three times (~10 min) with 25 mM NH4HCO3-50% acetonitrile (ACN) (1:1). Dried gel plugs were treated with freshly prepared 10 mM DTT in 50 mM NH4HCO3 for 45 min at 56°C. After incubation, the DTT was replaced quickly by the same volume of freshly prepared 55 mM iodoacetamide in 50 mM NH4HCO3 for 30 min and then dehydrated with 100% ACN. The dried gel pieces were incubated for 12 h at 37°C with 25 mM NH4HCO3 containing 0.02 μg/μl of mass spectrometry grade trypsin (Promega, Madison, WI, USA). The resulting peptides were extracted twice from the gel pieces, using peptide extraction buffer (1:1 mixture of 70% ACN and 0.1% trifluoroacetic acid [TFA]).
Mass spectrometry
Mass spectrometry was carried out as described earlier [34]. Digested samples were desalted and concentrated on C-18 ZipTips (Millipore, Billerica, MA, USA) using the manufacturer's protocol. ZipTips were eluted on MTP 384 target plate with 2 μl of α-cyano-4-hydroxycinnamic acid (HCCA) (Sigma-Aldrich, USA) saturated solution dissolved in 50% ACN, 0.2% TFA. Mass spectra of digested proteins were acquired using Autoflex II TOF/TOF 50 (Bruker Daltonik GmbH, Leipzig, Germany) in positive reflectron mode, in the detection range of 500-3000 m/z. External calibration to a spectrum, acquired for a mixture of peptides with masses ranging from 1046 to 2465 Da, was done prior to acquisition. The proteolytic masses obtained were then processed through Flex Analysis v.2.4 programme for peak detection. Submission of peak lists to the UniProtKB/Swiss-Prot database using the Mascot search engine http://www.matrixscience.com to identify the proteins from the annotated M. tuberculosis chromosome (strain H37Rv, EMBL/GenBank/DDBJ entry AL123456) Release 20 (June 2010). The pI and molecular mass of proteins were taken into account for identification of proteins and we did not find any significant difference between the experimental and predicted pI and molecular mass of proteins. Peptide mass tolerance was set to 50 ppm with carbamidomethyl-cystein set as fixed modification, oxidation of methionine as variable modification and 1 missed cleavage site allowed. Few intense peaks from each spectrum were selected for fragmentation by laser-induced dissociation in MALDI-TOF/TOF. The MS⁄MS spectra were calibrated internally to the precursor ion mass and used for sequence specific search at mascot database (Matrix science). In addition, peptide mass fingerprint-based searches were carried out using only the set of peptide masses, in the same database without any constraints for isoelectric point (pI) and molecular mass. The whole procedure was repeated several times to ensure correct protein identification.
Bioinformatic analysis
Protein sequences of all nine overexpressed proteins were retrieved from Tuberculist server http://genolist.pasteur.fr/TubercuList/ hosted by Pasteur Institute, Paris for whole annotated genome of H37Rv. BLAST [37] runs were performed at NCBI server http://blast.ncbi.nlm.nih.gov/ using the default threshold E-value of 10 and inclusion threshold value of 0.005. Motif and domain searches were made on EBI server http://www.ebi.ac.uk/Tools/InterProScan/ employing InterProScan which uses 13 different homology search programs (blastprodom, fprintscan, pfam, pir, panther, tigr, smart, superfamily, gene3 d, scanregexp, profilescan, seg, coils, tm, signalp, GO). Orthologs of proteins from other species of mycobacteria and human were obtained from KEGG http://www.genome.jp/kegg/ by single-directional best-hit option (SBH) and same server was employed for multiple sequence alignments (CLUSTALW) [38] we have used the following combined set of 5 organisms: mtu (M.tuberculosis), mbo (M.bovis), mav (M.avium), mle (M.leprae) and hsa (Homo sapiens). Sequences of H37Rv were submitted for 3-dimentional structure predictions at I-TASSER server http://zhang.bioinformatics.ku.edu/I-TASSER/. Structures were selected on the basis of RMSD values and agreement with Ramachandran Plot using VMD software (University of Illinois). Selected structures were molecularly docked with SM (structure obtained from http://www.drugbank.ca for in-silico interactions studies by submitting the structures to Patch Dock server [39]http://bioinfo3d.cs.tau.ac.il/PatchDock/ which is based on shape complementarity principles and results were refined using FireDock server [40,41]http://bioinfo3d.cs.tau.ac.il/FireDock/ which rearranges the interface side chains and adjusts the relative orientation of the molecules. Interacting amino acid side chains, drug molecule orientation and docking feasibility was based on Fire Dock scores and visualizations with VMD software.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
PS carried out the experiments, participated in the data analysis and drafted the manuscript. BK helped in carrying out mass spectrometric experiments, YG participated in bioinformatic analysis and NS participated in 2DE experiments. VMK and KV helped in the design of the project as well as preparation and critical review of the manuscript. DB conceived and designed the study, interpreted the experiment data and drafted the manuscript. All authors read and approved the final manuscript.
Supplementary Material
2DE patterns of three M. tuberculosis clinical isolates. a, b & c are sensitive to all first line drugs. 500 μg of proteins were first separated on 17 cm IPG strips of pH 4-7 by IEF and then by 12% SDS-PAGE in second dimension. Proteins were stained by coomassie brilliant blue. Regions showing low expressed proteins are highlighted by circles and squares.
2DE patterns of six M. tuberculosis clinical isolates. a, b, c, d, e & f are resistant to SM. Regions showing overexpressed proteins are highlighted by circles and squares.
Contributor Information
Prashant Sharma, Email: prashantanu@gmail.com.
Bhavnesh Kumar, Email: bhavnesh_micro@yahoo.co.in.
Yash Gupta, Email: yashgupta@lycos.com.
Neelja Singhal, Email: neelja30@gmail.com.
Vishwa Mohan Katoch, Email: vishwamohan_katoch@yahoo.co.in.
Krishnamurthy Venkatesan, Email: venkatesan_52@rediffmail.com.
Deepa Bisht, Email: abd1109@rediffmail.com.
Acknowledgements
The research was supported by grant from Department of Biotechnology, New Delhi (BT/PR7872/Med/14/1154/2006). PS, BK & YG are SRFs (ICMR, New Delhi), NS is SRF (CSIR-UGC, New Delhi). We thank Mr. Ajeet Pratap Singh for assistance.
References
- World Health Organization. Global tuberculosis control: epidemiology, strategy, financing. WHO Report. 2009. http://www.who.int/tb/publications/global_report/2009/pdf/full_report.pdf
- Franco H, Wowk PF, Silva CL, Trombone APF, Coelho-Castelo AAM, Oliver C, Jamur MC, Moretto EL, Bonato VLD. A DNA vaccine against tuberculosis based on the 65 kDa heat-shock protein differentially activates human macrophages and dendritic cells. Genet Vaccines Ther. 2008;6:3. doi: 10.1186/1479-0556-6-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Telenti A. In: Molecular Genetics of Mycobacteria. Hatfull GF, Jacobs WR, editor. Washington DC: ASM Press; 2000. Genetics of drug resistance in Mycobacterium tuberculosis; pp. 235–254. [Google Scholar]
- Honore N, Cole ST. Streptomycin resistance in mycobacteria. Antimicrob Agents Chemother. 1994;38(2):238–242. doi: 10.1128/aac.38.2.238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulger M, Aslan G, Emekdas G, Tezcan S, Serin MS. Investigation of rpsL and rrs gene region mutations in streptomycin resistant Mycobacterium tuberculosis complex isolates. Mikrobiyol Bul. 2009;43(1):115–120. [PubMed] [Google Scholar]
- Cole ST, Telenti A. Drug resistance in Mycobacterium tuberculosis. Europ Resp J Suppl. 1995;20:701–713. [PubMed] [Google Scholar]
- Morris S, Bai GH, Suffys P, Portillo-Gomez L, Fairchok M, Rouse D. Molecular mechanisms of multiple drug resistance in clinical isolates of Mycobacterium tuberculosis. J Infect Dis. 1995;171(4):954–960. doi: 10.1093/infdis/171.4.954. [DOI] [PubMed] [Google Scholar]
- Katsukawa C, Tamaru A, Miyata Y, Abe C, Makino M, Suzuki Y. Characterisation of the rpsL and rrs genes of streptomycin-resistant clinical isolates of Mycobacterium tuberculosis in Japan. J Appl Microbiol. 1997;83(5):634–640. doi: 10.1046/j.1365-2672.1997.00279.x. [DOI] [PubMed] [Google Scholar]
- Spies FS, da Silva PE, Ribeiro MO, Rossetti ML, Zaha A. Identification of mutations related to streptomycin resistance in clinical isolates of Mycobacterium tuberculosis and possible involvement of efflux mechanism. Antimicrob Agents Chemother. 2008;52(8):2947–2949. doi: 10.1128/AAC.01570-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hegde SS, Javid-Majd F, Blanchard JS. Overexpression and mechanistic analysis of chromosomally encoded aminoglycoside 2'-N-acetyltransferase [AAC(2')-Ic] from Mycobacterium tuberculosis. J Biol Chem. 2001;276(49):45876–45881. doi: 10.1074/jbc.M108810200. [DOI] [PubMed] [Google Scholar]
- Okamoto S, Tamaru A, Nakajima C, Nishimura K, Tanaka Y, Tokuyama S, Suzuki Y, Ochi K. Loss of a conserved 7-methylguanosine modification in 16 S rRNA confers low-level streptomycin resistance in bacteria. Mol Microbiol. 2007;63(4):1096–1106. doi: 10.1111/j.1365-2958.2006.05585.x. [DOI] [PubMed] [Google Scholar]
- Liberek K, Galitski TP, Zylicz M, Georgopoulos C. The DnaK chaperone modulates the heat shock response of Escherichia coli by binding to the σ32 transcription factor. Proc Natl Acad Sci USA. 1992;89:3516–3520. doi: 10.1073/pnas.89.8.3516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grossman AD, Straus DB, Walter WA, Gross CA. σ32 synthesis can regulate the synthesis of heat shock proteins in Escherichia coli. Genes Dev. 1987;1:179–184. doi: 10.1101/gad.1.2.179. [DOI] [PubMed] [Google Scholar]
- Sharma P, Kumar B, Singhal N, Katoch VM, Venkatesan K, Chauhan DS, Bisht D. Streptomycin induced protein expression analysis in Mycobacterium tuberculosis by two-dimensional gel electrophoresis and mass spectrometry. Indian J Med Res. 2010;132:400–408. [PubMed] [Google Scholar]
- Bukau B, Horwich AL. The Hsp70 and Hsp60 chaperone machines. Cell. 1998;92(3):351–366. doi: 10.1016/S0092-8674(00)80928-9. [DOI] [PubMed] [Google Scholar]
- Qamra R, Srinivas V, Mande SC. Mycobacterium tuberculosis GroEL homologues unusually exist as lower oligomers and retain the ability to suppress aggregation of substrate proteins. J Mol Biol. 2004;342:605–617. doi: 10.1016/j.jmb.2004.07.066. [DOI] [PubMed] [Google Scholar]
- Tabira Y, Ohara N, Kitaura H, Matsumoto S, Naito M, Yamada T. The 16-kDa α-crystallin-like protein of Mycobacterium bovis BCG is produced under conditions of oxygen deficiency and is associated with ribosomes. Res Microbiol. 1998;149(4):255–264. doi: 10.1016/S0923-2508(98)80301-X. [DOI] [PubMed] [Google Scholar]
- Mollenkopf HJ, Grode L, Mattow J, Stein M, Mann P, Knapp B, Ulmer J, Kaufmann SHE. Application of mycobacterial proteomics to vaccine design: Improved protection by Mycobacterium bovis BCG Prime-Rv3407 DNA boost vaccination against tuberculosis. Infect Immun. 2004;72(11):6471–6479. doi: 10.1128/IAI.72.11.6471-6479.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt F, Donahoe S, Hagens K, Mattow J, Schaible UE, Kaufmann SHE, Aebersold R, Jungblut PR. Complementary analysis of the Mycobacterium tuberculosis proteome by two-dimensional electrophoresis and isotope-coded affinity tag technology. Mol Cell Proteomics. 2004;3:24–42. doi: 10.1074/mcp.M300074-MCP200. [DOI] [PubMed] [Google Scholar]
- Jiang X, Zhang W, Gao F, Huang Y, Lv C, Wang H. Comparison of the proteome of isoniazid-resistant and susceptible strains of Mycobacterium tuberculosis. Microb Drug Resis. 2006;12(4):231–238. doi: 10.1089/mdr.2006.12.231. [DOI] [PubMed] [Google Scholar]
- Protein knowledgebase (UniProtKB) Electron transfer flavoprotein subunit alpha. UniProt. 2010. http://www.uniprot.org/uniprot/O53275 August 10, Version 66.
- Covert BA, Spencer JS, Orme IM, Belisle JT. The application of proteomics in defining the T cell antigens of Mycobacterium tuberculosis. Proteomics. 2001;1(4):574–586. doi: 10.1002/1615-9861(200104)1:4<574::AID-PROT574>3.0.CO;2-8. [DOI] [PubMed] [Google Scholar]
- Sassetti CM, Boyd DH, Rubin EJ. Genes required for mycobacterial growth defined by high density mutagenesis. Mol Microbiol. 2003;48(1):77–84. doi: 10.1046/j.1365-2958.2003.03425.x. [DOI] [PubMed] [Google Scholar]
- Kang C, Abbott DW, Park ST, Dascher CC, Cantley LC, Husson RN. The Mycobacterium tuberculosis serine/threonine kinases PknA and PknB: substrate identification and regulation of cell shape. Genes Dev. 2005;19:1692–1704. doi: 10.1101/gad.1311105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soderberg C, Lind P. A novel mammalian homologue of a bacterial citrate-metabolizing enzyme. Ann NY Acad Sci. 2002;967:476–481. doi: 10.1111/j.1749-6632.2002.tb04305.x. [DOI] [PubMed] [Google Scholar]
- Goulding CW, Bowers PM, Segelke B, Lekin T, Kim CY, Terwilliger TC, Eisenberg D. The structure and computational analysis of Mycobacterium tuberculosis protein CitE suggest a novel enzymatic function. J Mol Biol. 2007;365(2):275–283. doi: 10.1016/j.jmb.2006.09.086. [DOI] [PubMed] [Google Scholar]
- Izard T, Blackwelly NC. Crystal structures of the metal dependent 2-dehydro-3-deoxy-galactarate aldolase suggest a novel reaction mechanism. EMBO J. 2000;19(15):3849–3856. doi: 10.1093/emboj/19.15.3849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Namboori S, Mhatre N, Sujatha S, Srinivasan N, Pandit SB. Enhanced functional and structural domain assignments using remote similarity detection procedures for proteins encoded in the genome of Mycobacterium tuberculosis H37Rv. J Biosci. 2004;29(3):245–259. doi: 10.1007/BF02702607. [DOI] [PubMed] [Google Scholar]
- Canetti G, Froman S, Grosset J, Hauduroy P, Langerova M, Mahler HT, Meissner G, Mitchison DA, Sula L. Mycobacteria: laboratory methods for testing drug sensitivity and resistance. Bull Wld Hlth Org. 1963;29:565–578. [PMC free article] [PubMed] [Google Scholar]
- Brodie AF, Kalra VK, Lee SH, Cohen NS. Properties of energy-transducing systems in different types of membrane preparations from Mycobacterium phlei--preparation, resolution, and reconstitution. Methods Enzymol. 1979;55:175–200. doi: 10.1016/0076-6879(79)55024-1. full_text. [DOI] [PubMed] [Google Scholar]
- Bisht D, Singhal N, Sharma P, Venkatesan K. An improved sample preparation method for analyzing mycobacterial proteins in two-dimensional gels. Biochemistry (Moscow) 2007;72(6):672–674. doi: 10.1134/S0006297907060119. [DOI] [PubMed] [Google Scholar]
- Bradford MM. A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye-binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Gorg A, Obermaier C, Boguth G, Harde A, Scheibe B, Wildgruber R, Weiss W. The current state of two-dimensional electrophoresis with immobilized pH gradients. Electrophoresis. 2000;21(6):1037–1053. doi: 10.1002/(SICI)1522-2683(20000401)21:6<1037::AID-ELPS1037>3.0.CO;2-V. [DOI] [PubMed] [Google Scholar]
- Kumar G, Sharma P, Rathore G, Bisht D, Sengupta U. Proteomic analysis of outer membrane proteins of Edwardsiella tarda. J Appl Microbiol. 2010;108(6):2214–2221. doi: 10.1111/j.1365-2672.2009.04627.x. [DOI] [PubMed] [Google Scholar]
- Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Shevchenko A, Wilm M, Vorm O, Mann M. Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal Chem. 1996;68(5):850–858. doi: 10.1021/ac950914h. [DOI] [PubMed] [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Pearson WR, Lipman DJ. Improved tools for biological sequence comparison. Proc Natl Acad Sci USA. 1988;85(8):2444–2448. doi: 10.1073/pnas.85.8.2444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 2005. pp. W363–367. [DOI] [PMC free article] [PubMed]
- Andrusier N, Nussinov R, Wolfson HJ. FireDock: fast interaction refinement in molecular docking. Proteins. 2007;69(1):139–159. doi: 10.1002/prot.21495. [DOI] [PubMed] [Google Scholar]
- Mashiach E, Schneidman-Duhovny D, Andrusier N, Nussinov R, Wolfson HJ. FireDock: a Web server for fast interaction refinement in molecular docking. Nucleic Acids Res. 2008. pp. W229–232. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
2DE patterns of three M. tuberculosis clinical isolates. a, b & c are sensitive to all first line drugs. 500 μg of proteins were first separated on 17 cm IPG strips of pH 4-7 by IEF and then by 12% SDS-PAGE in second dimension. Proteins were stained by coomassie brilliant blue. Regions showing low expressed proteins are highlighted by circles and squares.
2DE patterns of six M. tuberculosis clinical isolates. a, b, c, d, e & f are resistant to SM. Regions showing overexpressed proteins are highlighted by circles and squares.