Abstract
Objective
The arylamine N-acetyltransferase 2 (NAT2) slow acetylation phenotype is an established risk factor for urinary bladder cancer. We previously reported on this risk association using NAT2 phenotypic categories inferred from NAT2 haplotypes based on 7 single nucleotide polymorphisms (SNPs) in a study in Spain. In a subsequent genome-wide scan, we have identified a single common tag SNP (rs1495741) located in the 3′ end of NAT2 that is also associated with bladder cancer risk. The aim of this report is to evaluate the agreement between the common tag SNP and the 7-SNP NAT2 inferred phenotype.
Methods
The agreement between the 7-SNP NAT2 inferred phenotype and the tag SNP, rs1495741, was initially assessed in 2,174 subjects from the Spanish Bladder Cancer Study (SBCS), and confirmed in a subset of subjects from the Main and Vermont component the New England Bladder Cancer Study (NEBCS). We also investigated the association of rs1495741 genotypes with NAT2 catalytic activity in cryopreserved hepatocytes from 154 individuals of European background.
Results
We observed very strong agreement between rs1495741 and the 7-SNP inferred NAT2 phenotype: sensitivity and specificity for the NAT2 slow phenotype was 99% and 95%, respectively. Our findings were replicated in an independent population from the United States. Estimates for the association between NAT2 slow phenotype and bladder cancer risk in the SBCS and its interaction with cigarette smoking were comparable for the 7-SNP inferred NAT2 phenotype and rs1495741. In addition, rs1495741 genotypes were strongly related to NAT2 activity measured in hepatocytes (P<0.0001).
Conclusion
A novel NAT2 tag SNP (rs1495741) predicts with high accuracy the 7- SNP inferred NAT2 phenotype, and thus can be used as a sole marker in pharmacogenetic or epidemiological studies of populations of European background. These findings illustrate the utility of tag SNPs, often employed in genome-wide association studies (GWAS), to identify novel phenotypic markers. Further studies are required to determine the functional implications of this novel SNP and the structure and evolution of the haplotype on which it resides.
INTRODUCTION
Differences in the acetylation activity of the arylamine N-acetyltransferase 2 (NAT2) gene were first discovered in 1950’s as modifiers of the neurotoxic effects of the anti-tuberculosis drug isoniazid1. Polymorphisms in the NAT2 gene have been extensively studied as modifiers of drug efficacy/toxicity and cancer risk because of its key role of the isoenzyme in the metabolism of arylamine and hydrazine drugs and carcinogens.2
The underlying genetic basis of differences in NAT2 enzyme activity has been explained by a combination of single nucleotide polymorphisms (SNPs) in the coding region (exon 2) of the NAT2 gene.3 Extensive effort has been devoted to the characterization of the functional effects of these SNPs and their haplotypes on NAT2 activity. 3–7 These observations have established a solid foundation for inferring the NAT2 rapid, intermediate and slow phenotypic classification based on the co-dominant expression of rapid and slow acetylator NAT2 haplotypes.4, 5 Many epidemiological studies of cancer risk have used genotyping to infer NAT2 phenotypic classification, since directly measuring NAT2 activity in humans would require complex and cumbersome phenotypic assays, such as measuring urinary metabolites after caffeine administration.8 A panel of 7 NAT2 SNPs has been recommended to accurately assign NAT2 haplotypes and inferred phenotype in populations of European background.9 These include four SNPs known to reduce NAT2 activity by different mechanisms (rs1801279 (R64Q), rs1801280 (I114T), rs1799930 (R197Q) and rs1799931 (G286E)), and three SNPs that do not cause changes in enzyme activity (rs1041983 (Y94Y), rs1799929 (L161L) and rs1208 (K268R)), 3, 5–7 but are needed to assign the NAT2 haplotypes and inferred phenotypic categories with high accuracy.9, 10
One of the most consistent associations with cancer risk has been for an increased risk of urothelial cell carcinoma of the bladder in NAT2 slow compared to rapid/intermediate acetylators.11, 12 Specifically, meta-analyses have shown that the NAT2 slow acetylation phenotype (mostly inferred from NAT2 haplotypes based on 3 to 7 NAT2 SNPs) is associated with an increased bladder cancer risk in populations of European background: summary odds ratio (OR) of 1.35 (95% CI 1.22–1.55) P=2×10−8, based on 4,403 cases and similar numbers of controls from 24 studies.12 In addition, the NAT2 association might be limited to cigarette smokers, which is consistent with the role of NAT2 in detoxification of aromatic amines in tobacco smoke, known bladder carcinogens. 11, 12 In this report, we evaluate the agreement between rs1495741, a tag SNP ~14kb 3′ of NAT2 discovered in a bladder cancer genome-wide association study (GWAS)13, with the NAT2 inferred phenotype11.
MATERIALS AND METHODS
Spanish Bladder Cancer Study (SBCS)
The Spanish Bladder Cancer Study is a hospital-based case-control study conducted in 18 hospitals from five different regions in Spain.11 DNA for genotype assays was extracted from leukocytes or buccal cell samples by Puregene® DNA Isolation Kit (Gentra Systems, Minneapolis, MN) or phenol-chloroform, respectively. This report includes study subjects who had adequate DNA quantity and quality for scanning on the Illumina Infinium®1M array. Informed consent was obtained in accordance with the National Cancer Institute and local Institutional Review Boards.
New England Bladder Cancer Study (NEBCS, Main and Vermont)
The New England Bladder Cancer Study is a population-based case-control study conducted in the states of Maine, Vermont and New Hampshire14. DNA was extracted from buccal cells by Puregene® DNA Isolation Kit (Gentra Systems, Minneapolis, MN) or phenol-chloroform. This report includes non-Hispanic study subjects of European descent, residents of Maine and Vermont (NEBCS (ME,VT)) who provided buccal cell samples and had adequate DNA quantity and quality to be scanned with the Infinium® Human 610-Quad array. Informed consent was obtained in accordance with the National Cancer Institute and local Institutional Review Boards.
Cryopreserved human hepatocytes
Cryopreserved human hepatocytes were received from Celsis In Vitro Technologies (Baltimore, MD) and stored in liquid nitrogen until use (see http://www.celsis.com/ivt/characterization-tables/ for details of the source of cells). Hepatocytes were thawed, centrifuged according to manufacturer’s instructions and as described previously15, and pelleted cells were washed in 1X phosphate buffered saline.
Genotyping
Seven NAT2 SNPs haplotypes, and inferred phenotypic classification were determined as previously described (http://www.louisville.edu/medschool/pharmacology/NAT.html). 11, 16 The seven SNPs included four SNPs known to reduce NAT2 activity by different mechanisms (rs1801279 (R64Q), rs1801280 (I114T), rs1799930 (R197Q) and rs1799931 (G286E), and three SNPs that do not cause changes in enzyme activity (rs1041983 (Y94Y), rs1799929 (L161L) and rs1208 (K268R) but are needed to assign the NAT2 haplotypes and inferred phenotypic categories with high accuracy. A description and methods for each specific genotype assay used in SBCS samples can be found at http://snp500cancer.nci.nih.gov/. Individuals possessing two of the NAT2 alleles associated with rapid acetylation activity (NAT2*4, NAT2*11A, NAT2*12A, NAT2*12B, NAT2*12C, NAT2*13) were classified as rapid acetylators; individuals possessing one of these alleles and one allele associated with slow acetylation (NAT2*5, NAT2*6, NAT2*7, and NAT2*14) were classified as intermediate acetylators, and those individuals that possessed two slow acetylation alleles were classified as slow acetylators. SNP rs1495741 was genotyped on the Illumina Infinium®1M array for the SBCS and the Illumina Infinium®610K array for the NEBCS (ME,VT)) as described (Rothman N, Garcia-Closas M et al., in prep).
Genomic DNA from hepatocytes was isolated from pelleted cells prepared as described above using the QIAamp DNA Mini Kit (Qiagen) according to manufacturer’s instructions. NAT2 SNPs, haplotypes and deduced phenotypic classification were inferred as described.4, 11, 16 rs1495741 was determined by TaqMan assay (http://snp500cancer.nci.nih.gov/
NAT2 activity assays in cryopreserved human hepatocytes
Hepatocytes were lysed in 20 mM sodium phosphate (pH 7.4), 1 mM EDTA, 1 mM dithiothreitol, 100 μM phenylmethanesulfonyl fluoride, and 1 μg/ml aprotin and 1 μM pepstatin A by three rounds of freeze (−70°C)/thawing (37°C). Lysates were centrifuged at 15,000 × g for 20 min and the resulting supernatant assayed for the level of NAT2 activity using the NAT2-selective substrate sulfamethazine (SMZ) as described previously17. Briefly, reactions containing lysates (< 2 mg protein/ml which translates into < 0.2 mg protein/reaction), 300 μM SMZ, and 1 mM acetyl coenzyme A were incubated at 37°C for 10 min, ensuring that there was linearity over the reaction period. Reactions were terminated by the addition of 1/10 volume 1 M acetic acid. The reaction tubes were centrifuged to precipitate protein and supernatant was injected onto a Lichrospher 100 RP-18 (125 mm × 4 mm; 5 μm) reverse phase column. Reactants and products were separated by high performance liquid chromatography (HPLC) (Beckman, Fullerton, CA). N-acetyl-SMZ was quantitated by absorbance at 260 nm. Variability per lysate was assessed by conducting reactions in triplicate.
Statistical methods
We constructed 3×3 contingency tables to visualize the agreement between the three rs1495741 genotypes and the three 7-SNP NAT2 inferred phenotype categories. We also estimated sensitivity and specificity as measures of agreement between rs1495741 and the 7-SNP NAT2 inferred phenotype, using the latter as the gold standard and after combining the AG/GG genotypes and the intermediate/rapid NAT2 phenotype categories. We estimated odds ratios (OR) and 95% confidence intervals (CI) from logistic regression models to test for associations between bladder cancer risk and NAT2 genotypes, haplotypes and phenotypes, as well as interactions with cigarette smoking, as previously described11.
NAT2 activity according to the rs1495741 genotype was assessed graphically and differences in mean activity by genotype categories were analyzed using one-way analysis of variance (ANOVA). We then fit a linear regression of NAT2 activity against the rs1495741 genotype (coded as an ordered variable with values 0,1,2 for AA, AG and GG genotypes, respectively) to test for a linear dose-response between the two variables.
RESULTS AND DISCUSSION
In a case-control study of urinary bladder in the Spanish Bladder Cancer Study (SBCS), we inferred the NAT2 acetylator phenotype based on 7 SNPs in exon 2 of NAT2 (7-SNP inferred phenotype).11 Samples from 2,174 subjects (1,097 cases and 1,077 controls) in this study were also genotyped for a set of one million SNPs (Illumina Infinium® 1M array), as part of a bladder cancer GWAS13. This study indentified a strong association for bladder cancer risk for rs1495741, shown in Table 1: per-allele OR (95%CI)= 1.34 (1.15–1.55); P=1.40 × 10−4. This common SNP, located approximately 14Kb in the 3′ end of the NAT2 gene has major allele A and minor allele G (frequency of 0.24 in controls). rs1495741 is located within a large LD block including the 7 previously identified SNPs (S_Figure 1 http://links.lww.com/FPC/A193). Although rs1495741 was highly correlated with NAT2 haplotypes derived from the 7 SNPs, it only showed a modest pairwise correlation with the individual 7 SNPs, four pairs of which were highly correlated (r2 ≥0.85; S_Table 1 http://links.lww.com/FPC/A194).
Table 1.
NAT2 polymorphisms genotyped in the Spanish Bladder Cancer Study (SBCS) and their association with urinary bladder cancer risk
| dbSNPID | Location | aa change | MAF in controls | OR | 95% | CI | P |
|---|---|---|---|---|---|---|---|
| NAT2 SNPs | |||||||
| rs1801279* | 18,301,984 (Ex2+197G>A) | R64Q | 0.001 | - | - | - | |
| rs1041983 | 18,302,075(Ex2+288C>T) | Y94Y | 0.314 | 1.05 | 0.89 | 1.25 | 0.548 |
| rs1801280* | 18,302,134(Ex2+347T>C) | I114T | 0.438 | 1.25 | 1.04 | 1.51 | 0.020 |
| rs1799929 | 18,302,274(Ex2+487C>T) | L161L | 0.429 | 1.24 | 1.03 | 1.49 | 0.023 |
| rs1799930* | 18,302,383(Ex2−580G>A) | R197Q | 0.294 | 1.00 | 0.84 | 1.18 | 0.990 |
| rs1208 | 18,302,596(Ex2−367G>A) | K268R | 0.436 | 1.24 | 1.03 | 1.49 | 0.026 |
| rs1799931* | 18,302,650(Ex2−313G>A) | G286E | 0.023 | 1.07 | 0.72 | 1.60 | 0.725 |
| GWAS SNP | |||||||
| rs1495741 | 18,317,161(14,494bp 3′ of STPA>G) | 0.239 | 1.34 | 1.15 | 1.55 | 1.40E-04 | |
SNPs known to reduce NAT2 activity by different mechanisms. The other NAT2 SNPs (rs1041983, rs1799929 and rs1208) do not cause changes in enzyme activity but are needed to assign the NAT2 haplotypes and inferred phenotypic categories with high accuracy.
Data from the 7 NAT2 SNPs has been previously reported by Garcia-Closas et al. 11.
Data from rs1495741 was included in a bladder cancer GWAS18..
aa: amino acid
MAF: minor allele frequency
OR: association between bladder cancer risk and having at least one variant of the functional SNP. For rs1495741, the OR is calculated under an additive model with the rare allele (rapid) as the reference.
CI: confidence interval
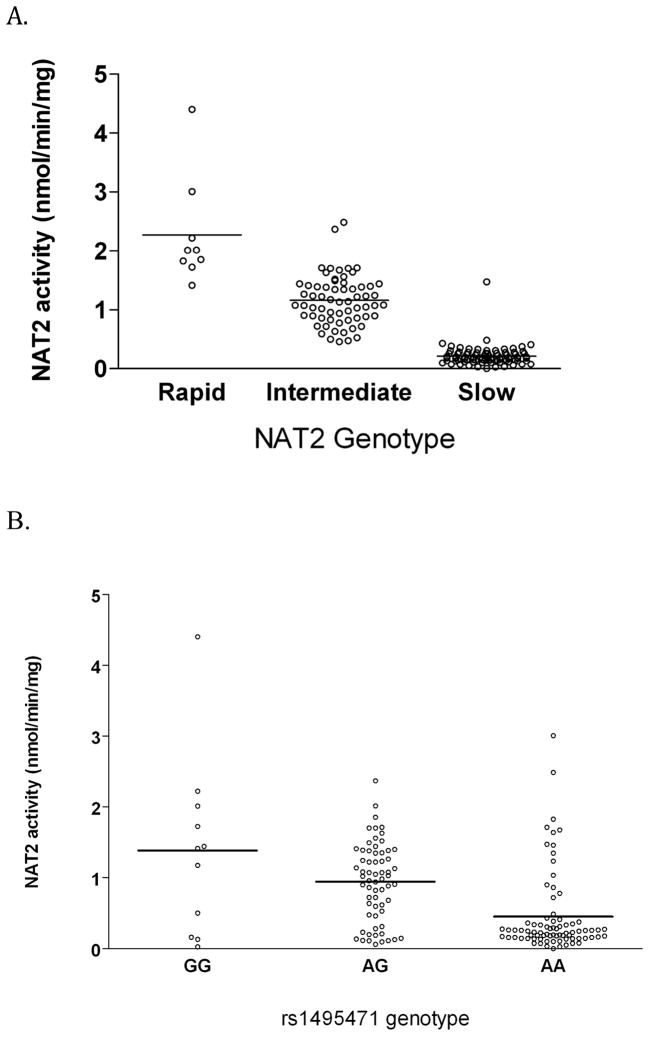
Figure 1.
Scatter plots of SMZ N-acetyltransferase activities in cryopreserved human hepatocytes from individuals of European background. Each dot represents SMZ N-acetyltransferase activity in each of the 154 human hepatocyte samples, stratified by categories of NAT2 acetylator phenotype inferred from 7 NAT2 SNPs (Panel A), and by genotypes of the NAT2 tag SNP (rs1495741) identified in a genome-wide scan (Panel B). Lines illustrate mean activity in each genotype.
ANOVA P-value for differences in SMZ NAT2 mean activity by genotype categories is P<0.0001 in both Panel A and B.
Panel A: The slope from a linear regression of SMZ NAT2 activity against NAT2 deduced phenotype (coded as 2=Rapid, 1=Intermediate, 0=Slow) is 0.745 P<0.0001.
Panel B: The slope from a linear regression of SMZ NAT2 activity against the rs1495741 genotype (coded as 0=AA, 1=AG and 2=GG) is 0.465, P<0.0001.
Relative risk estimates for the association with bladder cancer in the SBCS were very similar when we used data for the NAT2 7-SNP inferred phenotype11 and or the rs1495741 genotypes13 (Table 2). These risk estimates were also similar to those previously reported in large meta-analyses, which included SBCS data.11, 12 Furthermore, estimates of the interaction between the NAT2 slow vs rapid/intermediate inferred phenotype and smoking status (ever vs never smoking) were also comparable when we used the NAT2 7-SNP inferred variable (ORint (95%CI)=1.693 (1.04–2.75) P=0.034) or the GG vs GA/AA rs1495741 genotype (ORint=1.698 (1.04–2.78) P=0.035).
Table 2.
Association between urinary bladder cancer risk with NAT2 inferred phenotype and rs1495741 in 1097 cases and 1077 controls from the Spanish Bladder Cancer Study (SBCS)
| Controls | Cases | |||||||
|---|---|---|---|---|---|---|---|---|
| N | Freq. | N | Freq. | OR | 95% CI | P | ||
| NAT2 Inferred Phenotype* | ||||||||
| Rapid | 64 | 0.06 | 53 | 0.05 | 1.00 | |||
| Intermediate | 409 | 0.38 | 336 | 0.31 | 0.95 | 0.63 | 1.43 | 0.819 |
| Slow | 604 | 0.56 | 708 | 0.65 | 1.39 | 0.93 | 2.06 | 0.104 |
| Slow vs rapid/intermediate | 1.45 | 1.21 | 1.73 | 5.58E-05 | ||||
| rs1495741** | ||||||||
| GG | 61 | 0.06 | 48 | 0.04 | 1.00 | |||
| AG | 396 | 0.37 | 324 | 0.30 | 1.01 | 0.66 | 1.54 | 0.968 |
| AA | 620 | 0.58 | 725 | 0.66 | 1.47 | 0.97 | 2.21 | 0.066 |
| AA vs GG/AG | 1.46 | 1.22 | 1.75 | 4.50E-05 | ||||
NAT2 phenotype inferred from haplotypes determined by 7 NAT2 SNPs: rs1801279 (R64Q), rs1041983 (Y94Y), rs1801280 (I114T), rs1799929 (L161L), rs1799930 (R197Q), rs1208 (K268R), rs1799931 (G286E). This data was included in a previous report from the SBCS 11. Previously published numbers differ slightly because analyses in Table 2 are restricted to subjects with genome-wide scan data.
Data on rs1495741 from the SBCS has been included in a previously report from a bladder cancer GWAS13
Haplotype analyses based on the 8 SNPs (rs1495741 and the 7 NAT2 SNPs) identified one rapid NAT2 allele (NAT2*4), five slow NAT2 alleles (NAT2*5B, *6A, *7B, *5C, 5A, from the most to the least common), and a new haplotype that was NAT2*4 with the A allele of rs1495741 (S_Table 2 http://links.lww.com/FPC/A195). All of the acetylator haplotypes previously recognized as slow (NAT2*5C, *5A, *5B, *7B, *6A) shared the A allele of rs1495741, whereas the rapid haplotype NAT2*4 occurred both with the G allele (0.227 of controls) and, less frequently, with the A allele (0.010 of controls).
As expected by the consistency of risk associations for the NAT2 7-SNP inferred phenotype11 and rs1495741 identified in the GWAS13, the two variables showed strong agreement in samples from the SBCS (Table 3 Panel A). There was no misclassification for the extreme categories, i.e. misclassified individuals were always NAT2 intermediate phenotype or NAT2 heterozygous for rs1495741, and the sensitivity and specificity for slow vs intermediate/rapid NAT2 inferred phenotype was 99.4% and 95.2%, respectively. We used data from a US study to replicate these findings in an independent population, observing similarly high levels of agreement in a sample of 1,272 subjects of European background with complete genotype information (Table 3, Panel B). The high levels of agreement in two independent populations of European background from Spain and the United States, indicate that rs1495741 can be substituted for the panel of 7 NAT2 SNPs, as an accurate marker of the NAT2 phenotype in molecular epidemiology studies. The accuracy of rs1495741 in predicting the NAT2 phenotype in other ethnic groups remains to be evaluated.
Table 3.
Agreement between NAT2 phenotype inferred from 7 SNPs in NAT2* with rs1495741** identified in a genome-wide scan in the Spanish Bladder Cancer Study (Panel A) and the New England Bladder Cancer Study (ME, VT) (Panel B)
| A | rs1495741 | ||||
|---|---|---|---|---|---|
| GG | AG | AA | Total | ||
| NAT2 | Rapid | 109 | 8 | 0 | 117 |
| Intermediate | 0 | 704 | 41 | 745 | |
| Slow | 0 | 8 | 1,304 | 1312 | |
| Total | 109 | 720 | 1345 | 2174 | |
| Sensitivity = 99.4% 95%CI (98.8%–99.7%) | |||||
| Specificity = 95.2% 95%CI (93.6%–96.6%) | |||||
| B | rs1495741 | ||||
|---|---|---|---|---|---|
| GG | AG | AA | Total | ||
| NAT2 | Rapid | 54 | 8 | 0 | 62 |
| Intermediate | 1 | 428 | 21 | 450 | |
| Slow | 0 | 4 | 756 | 760 | |
| Total | 55 | 440 | 777 | 1272 | |
| Sensitivity = 99.5% (98.7%–99.9%) | |||||
| Specificity = 95.9% (93.8%–97.4%) | |||||
Data on the NAT2 inferred phenotype was included in a previous report from the SBCS 11, and is being included in a separate report from the NEBCS19.
Data on rs1495741 from the SBCS and NEBVS (ME,VT) has been included in a previously report from a bladder cancer GWAS13
Sensitivity is defined as the probability of being rs1495741 AA for subjects with the NAT2 slow 7-SNP inferred phenotype.
Specificity is defined as the probability of being rs1495741=GG or AG for subjects with the NAT2 rapid/intermediate 7-SNP inferred phenotype.
To evaluate the relationship between the NAT2 tag SNP and NAT2 hepatic activity, we determined the rs1495741 genotypes and NAT2 catalytic activity in crypopreserved hepatocytes from 154 individuals of European background. The distribution of NAT2 activity measurements in hepatocytes was strongly associated with the NAT2 7-SNP inferred phenotypes (Figure 1, Panel A), as previously reported17, and with the rs1495741 genotypes (Figure 1, Panel B). The NAT2 activity measurements are tighter for the three NAT2 7-SNP inferred phenotype categories than for the three rs1495741 genotype categories, which is consistent with the slight misclassification we observed between these two variables in the SBCS and NEBCS (ME,VT). rs1495741 genotypes showed similar levels of NAT2 catalytic activity in cryopreserved hepatocytes from nine individuals with the homozygous NAT2*4/*4 rapid acetylator genotype (data not shown). Additional functional characterization of rs1495741 on NAT2 activity is required to evaluate whether it impinges on NAT2 function independently or through its association with previously identified NAT2 haplotypes. It should also be noted that the NAT2 catalytic activity measured in vitro is not a perfect measure of the true phenotype, and it can be influenced by many extraneous factors such as the source and quality of the hepatocytes and assay conditions, as well as by NAT2 protein instability. These sources of variation could underestimate the relationship between genotypes and NAT2 activity.
CONCLUSION
We have identified a SNP in a bladder cancer GWAS (rs1495741) located ~14kb 3′ of NAT2 that shows almost perfect agreement with the NAT2 phenotype inferred from 7 previously identified SNPs in exon 2 of NAT2. These findings demonstrate the power of GWAS to identify novel markers for disease risk and other phenotypes, including markers of complex functional haplotypes such those known to occur in NAT2. Furthermore, our results indicate that a single SNP, rs1495741 can be analyzed and thus, reduce the complexity of assessing the NAT2 phenotype in epidemiological studies.
Supplementary Material
Acknowledgments
The SBCS thanks Robert C Saal (Westat, Rockville, MD, USA), Leslie Carroll, and Jane Wang (both Information Management Services, Silver Spring, MD, USA) for their support in study and data management; Drs. Adonina Tardón (Universidad de Oviedo, Oviedo, Spain), Consol Serra (Consorci Hospitalari Parc Taulí, Sabadell, Spain), Alfredo Carrato (Hospital General de Elche, Elche, Spain), Reina García-Closas (Unidad de Investigación, Hospital Universitario de Canarias, LaLaguna, Spain), Josep Lloreta (Department of Pathology, Hospital del Mar-IMAS, Barcelona, Spain), and Montserrat Torà, Gemma Castaño-Vinyals and Maria Sala (all from Institut Municipal d’Investigació Mèdica, Barcelona, Spain) for their work in data collection and study coordination. The NEBCS (ME, VT) thanks Anna McIntosh, Paul Hurwitz, Patricia Clark and Vanessa Olivo (Westat, Rockville, MD) for their support in study and data management, and Anne Taylor and Mary McAdams (Information Management Services, Silver Spring, MD) for their programming help. We would like to acknowledge Dr Michael Jones (Maine Medical Center), Sue Ledoux and Dawn Nicolaides (Maine Cancer Registry), Dr Masatoshi Kida (University of Vermont), William Apao and Carolyn Greene (Vermont Cancer Registry) for their contributions during the fieldwork and data collection phases. We also thank all fieldwork staff, interviewers, and data abstractors for their dedicated work, and our study participants for agreeing to be part of the studies.
Funding: The SBCS and NEBCS (ME,VT) were supported by Intramural Research Program of the National Institute of Health, National Cancer Institute, Division of Cancer Epidemiology and Genetics (contract numbers N02-CP-11015 for SBCS and N02-CP-01037 for NEBCS). The SBCS was also funded by grants FIS/Spain 00/0745, Fundació Marató TV3, Red Temática Investigación Cooperativa en Cáncer (RTICC), G03/174, and CA34627. The work on NAT2 catalytic activity in human hepatocytes was partially supported by United States Public Health Service grant R01-CA034627 from the United States National Cancer Institute, National Institutes of Health, Department of Health and Human Services.
References
- 1.Hughes HB, Biehl JP, Jones AP, Schmidt LH. Metabolism of isoniazid in man as related to the occurrence of peripheral neuritis. Am Rev Tuberc. 1954;70:266–73. doi: 10.1164/art.1954.70.2.266. [DOI] [PubMed] [Google Scholar]
- 2.Hein DW, Doll MA, Fretland AJ, et al. Molecular genetics and epidemiology of the NAT1 and NAT2 acetylation polymorphisms. Cancer Epidemiol Biomarkers Prev. 2000;9:29–42. [PubMed] [Google Scholar]
- 3.Walraven JM, Zang Y, Trent JO, Hein DW. Structure/function evaluations of single nucleotide polymorphisms in human N-acetyltransferase2. Curr Drug Metab. 2008;9:471–86. doi: 10.2174/138920008784892065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Hein DW. N-acetyltransferase 2 genetic polymorphism: effects of carcinogen and haplotype on urinary bladder cancer risk. Oncogene. 2006;25:1649–58. doi: 10.1038/sj.onc.1209374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zang Y, Doll MA, Zhao S, States JC, Hein DW. Functional characterization of single-nucleotide polymorphisms and haplotypes of human N-acetyltransferase 2. Carcinogenesis. 2007;28:1665–71. doi: 10.1093/carcin/bgm085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hein DW, Ferguson RJ, Doll MA, Rustan TD, Gray K. Molecular genetics of human polymorphic N-acetyltransferase: enzymatic analysis of 15 recombinant wild-type, mutant, and chimeric NAT2 allozymes. Hum Mol Genet. 1994;3:729–34. doi: 10.1093/hmg/3.5.729. [DOI] [PubMed] [Google Scholar]
- 7.Fretland AJ, Leff MA, Doll MA, Hein DW. Functional characterization of human N-acetyltransferase 2 (NAT2) single nucleotide polymorphisms. Pharmacogenetics. 2001;11:207–15. doi: 10.1097/00008571-200104000-00004. [DOI] [PubMed] [Google Scholar]
- 8.Butler MA, Lang NP, Young JF, et al. Determination of CYP1A2 and NAT2 phenotypes in human populations by analysis of caffeine urinary metabolites. Pharmacogenetics. 1992;2:116–27. doi: 10.1097/00008571-199206000-00003. [DOI] [PubMed] [Google Scholar]
- 9.Deitz AC, Rothman N, Rebbeck TR, et al. Impact of misclassification in genotype-exposure interaction studies: example of N-acetyltransferase 2 (NAT2), smoking, and bladder cancer. Cancer Epidemiol Biomarkers Prev. 2004;13:1543–6. [PubMed] [Google Scholar]
- 10.Hein DW. N-acetyltransferase SNPs: emerging concepts serve as a paradigm for understanding complexities of personalized medicine. Expert Opin Drug Metab Toxicol. 2009;5:353–66. doi: 10.1517/17425250902877698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Garcia-Closas M, Malats N, Silverman D, et al. NAT2 slow acetylation, GSTM1 null genotype, and risk of bladder cancer: results from the Spanish Bladder Cancer Study and meta-analyses. Lancet. 2005;366:649–59. doi: 10.1016/S0140-6736(05)67137-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rothman N, Garcia-Closas M, Hein DW. Commentary: Reflections on G. M. Lower and colleagues’ 1979 study associating slow acetylator phenotype with urinary bladder cancer: meta-analysis, historical refinements of the hypothesis, and lessons learned. International journal of epidemiology. 2007;36:23–8. doi: 10.1093/ije/dym026. [DOI] [PubMed] [Google Scholar]
- 13.Rothman N, Garcia-Closas M, Chatterjee N, et al. A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. doi: 10.1038/ng.687. Under review. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Baris D, Karagas MR, Verrill C, et al. A case-control study of smoking and bladder cancer risk: emergent patterns over time. J Natl Cancer Inst. 2009;101:1553–61. doi: 10.1093/jnci/djp361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Martin RC, Li Y, Liu Q, et al. Manganese superoxide dismutase V16A single-nucleotide polymorphism in the mitochondrial targeting sequence is associated with reduced enzymatic activity in cryopreserved human hepatocytes. DNA Cell Biol. 2009;28:3–7. doi: 10.1089/dna.2008.0788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Doll MA, Hein DW. Comprehensive human NAT2 genotype method using single nucleotidepolymorphism-specific polymerase chain reaction primers and fluorogenic probes. Anal Biochem. 2001;288:106–8. doi: 10.1006/abio.2000.4892. [DOI] [PubMed] [Google Scholar]
- 17.Doll MA, Zang Y, Moeller T, Hein DW. Co-dominant expression of N-acetylation and O-acetylation activities catalyzed by N-acetyltransferase 2 in human hepatocytes. J Pharmacol Exp Ther. doi: 10.1124/jpet.110.168567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rothman N, Garcia-Closas M, Chatterjee N, et al. A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. doi: 10.1038/ng.687. Under review 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Moore L, Baris D, Figueroa J, et al. GSTM1 null and NAT2 slow acetylation genotypes, smoking intensity, and bladder cancer risk: results from the New England bladder cancer case-control study and meta-analyses. doi: 10.1093/carcin/bgq223. Submitted 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.