Abstract
The title enamino ketone, C12H15NO, a derivative of 4-(phenylamino)pent-3-en-2-one, presents a roughly planar [greatest displacement of an atom from the pentenone plane is 0.033 (2) Å] pentenone backbone, enhanced by an intramolecular N—H⋯O hydrogen bond; the asymmetry in C—C distances in the group suggests the presence of unsaturated bonds. The overall geometry in the free ligand differs significantly from that in other reported compounds, in which it is coordinated to rhodium; this is reflected in the bond distances [the N⋯O distance is significantly increased (0.2 Å) upon coordination to the metal] and the dihedral angle between the benzene ring and the pentenone backbone [49.53 (5)°]. All of the methyl goups are rotationally disordered over two orientations of equal occupancy.
Related literature
For synthetic background, see: Shaheen et al. (2006 ▶). For applications of enaminoketones in liquid crystals, see: Pyżuk et al. (1993 ▶), in fluorescence, see: Xia et al. (2008 ▶), in complexes of medical interest, see: Tan et al. (2008 ▶); Chen & Rhodes (1996 ▶), in catalysis, see: Nair et al. (2002 ▶); Van Aswegen et al. (1991 ▶); Steyn et al. (1992 ▶, 1997 ▶); Otto et al. (1998 ▶); Roodt & Steyn (2000 ▶); Brink et al. (2010 ▶). For the structures of related ligand systems, see: Damoense et al. (1994 ▶); Venter et al. (2009a
▶,b
▶). 
Experimental
Crystal data
C12H15NO
M r = 189.25
Monoclinic,

a = 7.5674 (7) Å
b = 11.5075 (9) Å
c = 12.0996 (11) Å
β = 92.154 (5)°
V = 1052.91 (16) Å3
Z = 4
Mo Kα radiation
μ = 0.08 mm−1
T = 100 K
0.55 × 0.23 × 0.12 mm
Data collection
Bruker X8 APEXII 4K Kappa CCD diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 2004 ▶) T min = 0.960, T max = 0.991
10022 measured reflections
2308 independent reflections
1854 reflections with I > 2σ(I)
R int = 0.034
Refinement
R[F 2 > 2σ(F 2)] = 0.053
wR(F 2) = 0.150
S = 1.07
2308 reflections
129 parameters
H-atom parameters constrained
Δρmax = 0.45 e Å−3
Δρmin = −0.35 e Å−3
Data collection: APEX2 (Bruker, 2005 ▶); cell refinement: SAINT-Plus (Bruker, 2004 ▶); data reduction: SAINT-Plus; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: DIAMOND (Brandenburg & Putz, 2005 ▶); software used to prepare material for publication: WinGX (Farrugia, 1999 ▶).
Supplementary Material
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536810021045/bg2348sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536810021045/bg2348Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| N11—H11⋯O12 | 0.91 | 1.90 | 2.6345 (19) | 136 |
Table 2. Comparative geometrical parameters (Å, °) for free and coordinated N,O-bidendate (N,O-bid) compounds.
| Parameters | (I) | (II) | (III) | (IV) |
|---|---|---|---|---|
| C111—N11 | 1.422 (2) | 1.521 (4)/1.463 (3) | 1.440 (4) | - |
| C2—N11 | 1.345 (2) | 1.320 (4) | 1.319 (4) | 1.303 (6) |
| C4—O12 | 1.257 (2) | 1.290 (3) | 1.291 (4) | 1.281 (6) |
| C2—C3 | 1.383 (3) | 1.410 (4) | 1.423 (4) | 1.396 (7) |
| C3—C4 | 1.420 (2) | 1.365 (3) | 1.382 (3) | 1.388 (9) |
| O12⋯N11 | 2.635 (2) | 2.885 (3) | 2.886 (3) | 2.826 (6) |
| N11—C2—C4—O12 | −0.5 (1) | 4.1 (2) | −2.6 (2) | 1.2 (4) |
| Dihedral angle | 49.53 (5) | 87.47 (4)/89.36 (8) | 85.58 (8) | - |
Notes: (I) This work; (II) N,O-bid = 4-(2,3-dimethyl phenylamino)pent-3-en-2-onato (Venter et al., 2009a ▶); (III) N,O-bid = 4-(2,6-dimethyl phenylamino)pent-3-en-2-onato (Venter et al., 2009b ▶). (IV) N,O-bid = 4-amino-pent-3-en-2-onato (Damoense et al., 1994 ▶). The dihedral angle is defined as the torsion angle between the N–C–C–C–O plane and the benzene ring. A positive angle denotes a clockwise rotation.
Acknowledgments
Financial assistance from the University of the Free State is gratefully acknowledged. We also express our gratitude towards SASOL, the UFS Materials and Nanosciences Academic Cluster and the South African National Research Foundation (SA-NRF/THRIP) for financial support of this project. Part of this material is based on work supported by the SA-NRF/THRIP under grant No. GUN 2068915. Opinions, findings, conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the SA-NRF.
supplementary crystallographic information
Comment
A well-known system in organometallic chemistry is the β-diketone compound AcacH (acetylacetone; or when coordinated acetylacetonato, acac-). A multitude of derivatives have been synthesized to date, with enaminoketones being one type. Since enaminoketones contain nitrogen and oxygen atoms as well as an unsaturated C—C bond, these electron-rich compounds are of interest in various fields including liquid crystals [Pyżuk et al. (1993)], fluorescence studies [Xia et al. (2008)] as well as formation of complexes of medical interest [Tan et al. (2008); Chen & Rhodes (1996)]. It also has significant application possibilities in catalysis [Nair et al. (2002); Van Aswegen et al. (1991); Steyn et al. (1992; 1997); Otto et al. (1998); Roodt & Steyn (2000); Brink et al. (2010)].
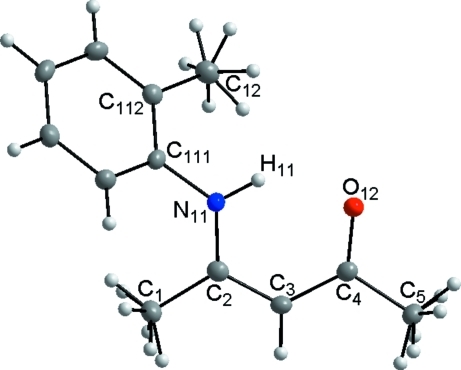
The title enaminoketone is a derivative of 4-(phenylamino)pent-3-en-2-one [PhonyH; Shaheen et al. (2006)]. Fig. 1 shows a view of the molecule. The C2–C3 distance of 1.383 (3) Å, versus the C3–C4 distance of 1.420 (2)Å indicates an unsaturated bond in the pentenone backbone, which is otherwise planar, probably helped by an intramolecular N1—H1···O1 bond (N1—H1: 0.91Å; H1···O1: 1.90Å; N1···O1:2.635 (2)Å, N1—H1···O: 136.4°). In general terms, the geometry in the free ligand differs significantly from that in other reported compounds where it is coordinated to rhodium [Table1; Venter et al. (2009a; 2009b); Damoense et al. (1994)]; fon instance, the N1···O1 distance is greatly ncreased (~0.2 Å) upon coordination to the metal, as it is the dihedral angle between the phenyl ring and the pentenone backbone. All the methyl goups appear rotationally disordered in two sites of similar occupation.
Experimental
A solution of acetylacetone (11.07 g, 0.1106 mol), 2-Me-aniline (10.73 g, 0.1008 mol) and 2 drops of H2SO4(conc.) in 150 ml benzene was refluxed for 6 hours in a DeanStark trap, filtered and left to crystallize. Crystals suitable for X-Ray diffraction were obtained in 17.86 g (94.32 %) yield. This compound is stable in air and light over a period of several months.
Refinement
The methyl and aromatic H atoms were placed in geometrically idealized positions and constrained to ride on their parent atoms, with C—H = 0.95 and 0.98Å and Uiso(H) = 1.5Ueq(C) and 1.2Ueq(C), respectively. The methyl groups were generated to fit the difference electron density and the groups were then refined as rigid rotors. The highest residual electron-density peak is 0.47Å from H1F.
Figures
Fig. 1.
Molecular structure of the title compound. Displacement ellipsoids are drawn at the 50% probability displacement level.
Crystal data
| C12H15NO | F(000) = 408 |
| Mr = 189.25 | Dx = 1.194 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 2889 reflections |
| a = 7.5674 (7) Å | θ = 2.7–28.2° |
| b = 11.5075 (9) Å | µ = 0.08 mm−1 |
| c = 12.0996 (11) Å | T = 100 K |
| β = 92.154 (5)° | Plate, colourless |
| V = 1052.91 (16) Å3 | 0.55 × 0.23 × 0.12 mm |
| Z = 4 |
Data collection
| Bruker X8 APEXII 4K Kappa CCD diffractometer | 2308 independent reflections |
| Radiation source: fine-focus sealed tube | 1854 reflections with I > 2σ(I) |
| graphite | Rint = 0.034 |
| ω and φ scans | θmax = 27°, θmin = 2.4° |
| Absorption correction: multi-scan (SADABS; Bruker, 2004) | h = −9→9 |
| Tmin = 0.960, Tmax = 0.991 | k = −14→12 |
| 10022 measured reflections | l = −15→15 |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.053 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.150 | H-atom parameters constrained |
| S = 1.07 | w = 1/[σ2(Fo2) + (0.0654P)2 + 0.9506P] where P = (Fo2 + 2Fc2)/3 |
| 2308 reflections | (Δ/σ)max < 0.001 |
| 129 parameters | Δρmax = 0.45 e Å−3 |
| 0 restraints | Δρmin = −0.35 e Å−3 |
Special details
| Experimental. The intensity data was collected on a Bruker X8 ApexII 4 K Kappa CCD diffractometer using an exposure time of 60 seconds/frame. A total of 688 frames were collected with a frame width of 0.5° covering up to θ = 28.24° with 99.1% completeness accomplished. |
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | Occ. (<1) | |
| C1 | 0.4288 (3) | 0.30725 (17) | 0.73241 (16) | 0.0212 (4) | |
| H1A | 0.4068 | 0.3907 | 0.723 | 0.032* | 0.5 |
| H1B | 0.3886 | 0.2659 | 0.6653 | 0.032* | 0.5 |
| H1C | 0.5557 | 0.2939 | 0.7458 | 0.032* | 0.5 |
| H1D | 0.4939 | 0.243 | 0.6998 | 0.032* | 0.5 |
| H1E | 0.5121 | 0.3678 | 0.7575 | 0.032* | 0.5 |
| H1F | 0.345 | 0.3397 | 0.6769 | 0.032* | 0.5 |
| C2 | 0.3300 (2) | 0.26340 (16) | 0.82877 (15) | 0.0172 (4) | |
| C3 | 0.3410 (2) | 0.14730 (15) | 0.85770 (15) | 0.0174 (4) | |
| H3 | 0.4142 | 0.0982 | 0.8159 | 0.021* | |
| C4 | 0.2494 (2) | 0.09702 (15) | 0.94616 (14) | 0.0162 (4) | |
| C5 | 0.2647 (3) | −0.03183 (16) | 0.96633 (16) | 0.0217 (4) | |
| H5A | 0.1945 | −0.0532 | 1.0296 | 0.033* | 0.5 |
| H5B | 0.3889 | −0.0521 | 0.982 | 0.033* | 0.5 |
| H5C | 0.2208 | −0.0739 | 0.9005 | 0.033* | 0.5 |
| H5D | 0.3417 | −0.0663 | 0.9118 | 0.033* | 0.5 |
| H5E | 0.1472 | −0.0674 | 0.9594 | 0.033* | 0.5 |
| H5F | 0.3153 | −0.0456 | 1.0409 | 0.033* | 0.5 |
| C12 | 0.2730 (3) | 0.49042 (17) | 1.07167 (15) | 0.0218 (4) | |
| H12A | 0.2961 | 0.4067 | 1.0685 | 0.033* | 0.5 |
| H12B | 0.1797 | 0.5055 | 1.1238 | 0.033* | 0.5 |
| H12C | 0.3812 | 0.5311 | 1.0964 | 0.033* | 0.5 |
| H12D | 0.2752 | 0.5555 | 1.1239 | 0.033* | 0.5 |
| H12E | 0.3917 | 0.4567 | 1.0687 | 0.033* | 0.5 |
| H12F | 0.1901 | 0.4311 | 1.0961 | 0.033* | 0.5 |
| O12 | 0.15430 (17) | 0.15514 (11) | 1.00878 (10) | 0.0195 (3) | |
| C111 | 0.1994 (2) | 0.45704 (15) | 0.86788 (15) | 0.0160 (4) | |
| C112 | 0.2147 (2) | 0.53330 (16) | 0.95849 (15) | 0.0172 (4) | |
| C113 | 0.1737 (2) | 0.64979 (16) | 0.94050 (16) | 0.0203 (4) | |
| H113 | 0.1821 | 0.7026 | 1.0008 | 0.024* | |
| C114 | 0.1207 (2) | 0.69078 (16) | 0.83647 (16) | 0.0208 (4) | |
| H114 | 0.0932 | 0.7707 | 0.8262 | 0.025* | |
| C115 | 0.1081 (2) | 0.61469 (16) | 0.74784 (15) | 0.0192 (4) | |
| H115 | 0.0738 | 0.6426 | 0.6763 | 0.023* | |
| C116 | 0.1456 (2) | 0.49751 (16) | 0.76367 (15) | 0.0180 (4) | |
| H116 | 0.1344 | 0.445 | 0.7032 | 0.022* | |
| N11 | 0.2320 (2) | 0.33691 (13) | 0.88747 (13) | 0.0176 (3) | |
| H11 | 0.1801 | 0.3060 | 0.9471 | 0.021* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| C1 | 0.0225 (9) | 0.0206 (9) | 0.0210 (9) | 0.0019 (7) | 0.0057 (7) | 0.0032 (7) |
| C2 | 0.0167 (8) | 0.0190 (9) | 0.0159 (9) | 0.0001 (7) | 0.0001 (7) | −0.0007 (7) |
| C3 | 0.0192 (9) | 0.0163 (9) | 0.0168 (9) | 0.0028 (7) | 0.0029 (7) | −0.0015 (7) |
| C4 | 0.0176 (8) | 0.0157 (9) | 0.0152 (8) | 0.0008 (7) | −0.0013 (7) | −0.0010 (7) |
| C5 | 0.0254 (10) | 0.0152 (9) | 0.0250 (10) | 0.0014 (7) | 0.0062 (8) | 0.0005 (7) |
| C12 | 0.0221 (9) | 0.0244 (10) | 0.0188 (9) | −0.0010 (8) | 0.0000 (7) | −0.0015 (7) |
| O12 | 0.0254 (7) | 0.0157 (6) | 0.0176 (7) | 0.0015 (5) | 0.0051 (5) | −0.0001 (5) |
| C111 | 0.0158 (8) | 0.0122 (8) | 0.0202 (9) | −0.0008 (6) | 0.0032 (7) | 0.0003 (7) |
| C112 | 0.0164 (8) | 0.0177 (9) | 0.0177 (9) | −0.0018 (7) | 0.0023 (7) | −0.0002 (7) |
| C113 | 0.0222 (9) | 0.0170 (9) | 0.0220 (9) | −0.0018 (7) | 0.0033 (7) | −0.0059 (7) |
| C114 | 0.0214 (9) | 0.0129 (8) | 0.0284 (10) | 0.0004 (7) | 0.0047 (8) | 0.0012 (7) |
| C115 | 0.0189 (9) | 0.0190 (9) | 0.0198 (9) | −0.0001 (7) | 0.0024 (7) | 0.0031 (7) |
| C116 | 0.0200 (9) | 0.0163 (9) | 0.0178 (9) | −0.0020 (7) | 0.0020 (7) | −0.0018 (7) |
| N11 | 0.0232 (8) | 0.0132 (7) | 0.0168 (8) | 0.0009 (6) | 0.0053 (6) | 0.0012 (6) |
Geometric parameters (Å, °)
| C1—C2 | 1.496 (2) | C12—C112 | 1.506 (3) |
| C1—H1A | 0.98 | C12—H12A | 0.98 |
| C1—H1B | 0.98 | C12—H12B | 0.98 |
| C1—H1C | 0.98 | C12—H12C | 0.98 |
| C1—H1D | 0.98 | C12—H12D | 0.98 |
| C1—H1E | 0.98 | C12—H12E | 0.98 |
| C1—H1F | 0.98 | C12—H12F | 0.98 |
| C2—N11 | 1.345 (2) | C111—C116 | 1.391 (3) |
| C2—C3 | 1.383 (3) | C111—C112 | 1.406 (3) |
| C3—C4 | 1.420 (2) | C111—N11 | 1.422 (2) |
| C3—H3 | 0.95 | C112—C113 | 1.391 (3) |
| C4—O12 | 1.257 (2) | C113—C114 | 1.389 (3) |
| C4—C5 | 1.506 (2) | C113—H113 | 0.95 |
| C5—H5A | 0.98 | C114—C115 | 1.385 (3) |
| C5—H5B | 0.98 | C114—H114 | 0.95 |
| C5—H5C | 0.98 | C115—C116 | 1.390 (3) |
| C5—H5D | 0.98 | C115—H115 | 0.95 |
| C5—H5E | 0.98 | C116—H116 | 0.95 |
| C5—H5F | 0.98 | N11—H11 | 0.9071 |
| C2—C1—H1A | 109.5 | H5B—C5—H5F | 56.3 |
| C2—C1—H1B | 109.5 | H5C—C5—H5F | 141.1 |
| H1A—C1—H1B | 109.5 | H5D—C5—H5F | 109.5 |
| C2—C1—H1C | 109.5 | H5E—C5—H5F | 109.5 |
| H1A—C1—H1C | 109.5 | C112—C12—H12A | 109.5 |
| H1B—C1—H1C | 109.5 | C112—C12—H12B | 109.5 |
| C2—C1—H1D | 109.5 | H12A—C12—H12B | 109.5 |
| H1A—C1—H1D | 141.1 | C112—C12—H12C | 109.5 |
| H1B—C1—H1D | 56.3 | H12A—C12—H12C | 109.5 |
| H1C—C1—H1D | 56.3 | H12B—C12—H12C | 109.5 |
| C2—C1—H1E | 109.5 | C112—C12—H12D | 109.5 |
| H1A—C1—H1E | 56.3 | H12A—C12—H12D | 141.1 |
| H1B—C1—H1E | 141.1 | H12B—C12—H12D | 56.3 |
| H1C—C1—H1E | 56.3 | H12C—C12—H12D | 56.3 |
| H1D—C1—H1E | 109.5 | C112—C12—H12E | 109.5 |
| C2—C1—H1F | 109.5 | H12A—C12—H12E | 56.3 |
| H1A—C1—H1F | 56.3 | H12B—C12—H12E | 141.1 |
| H1B—C1—H1F | 56.3 | H12C—C12—H12E | 56.3 |
| H1C—C1—H1F | 141.1 | H12D—C12—H12E | 109.5 |
| H1D—C1—H1F | 109.5 | C112—C12—H12F | 109.5 |
| H1E—C1—H1F | 109.5 | H12A—C12—H12F | 56.3 |
| N11—C2—C3 | 120.21 (16) | H12B—C12—H12F | 56.3 |
| N11—C2—C1 | 120.06 (16) | H12C—C12—H12F | 141.1 |
| C3—C2—C1 | 119.73 (16) | H12D—C12—H12F | 109.5 |
| C2—C3—C4 | 123.91 (16) | H12E—C12—H12F | 109.5 |
| C2—C3—H3 | 118 | C116—C111—C112 | 120.72 (16) |
| C4—C3—H3 | 118 | C116—C111—N11 | 121.30 (16) |
| O12—C4—C3 | 122.97 (16) | C112—C111—N11 | 117.91 (16) |
| O12—C4—C5 | 117.93 (16) | C113—C112—C111 | 117.91 (17) |
| C3—C4—C5 | 119.10 (15) | C113—C112—C12 | 120.93 (16) |
| C4—C5—H5A | 109.5 | C111—C112—C12 | 121.16 (16) |
| C4—C5—H5B | 109.5 | C114—C113—C112 | 121.61 (17) |
| H5A—C5—H5B | 109.5 | C114—C113—H113 | 119.2 |
| C4—C5—H5C | 109.5 | C112—C113—H113 | 119.2 |
| H5A—C5—H5C | 109.5 | C115—C114—C113 | 119.74 (17) |
| H5B—C5—H5C | 109.5 | C115—C114—H114 | 120.1 |
| C4—C5—H5D | 109.5 | C113—C114—H114 | 120.1 |
| H5A—C5—H5D | 141.1 | C114—C115—C116 | 119.93 (17) |
| H5B—C5—H5D | 56.3 | C114—C115—H115 | 120 |
| H5C—C5—H5D | 56.3 | C116—C115—H115 | 120 |
| C4—C5—H5E | 109.5 | C115—C116—C111 | 120.08 (17) |
| H5A—C5—H5E | 56.3 | C115—C116—H116 | 120 |
| H5B—C5—H5E | 141.1 | C111—C116—H116 | 120 |
| H5C—C5—H5E | 56.3 | C2—N11—C111 | 128.22 (16) |
| H5D—C5—H5E | 109.5 | C2—N11—H11 | 115.9 |
| C4—C5—H5F | 109.5 | C111—N11—H11 | 115.9 |
| H5A—C5—H5F | 56.3 | ||
| N11—C2—C3—C4 | 1.9 (3) | C112—C113—C114—C115 | −0.1 (3) |
| C1—C2—C3—C4 | −178.62 (17) | C113—C114—C115—C116 | 1.1 (3) |
| C2—C3—C4—O12 | −2.6 (3) | C114—C115—C116—C111 | −1.4 (3) |
| C2—C3—C4—C5 | 177.01 (17) | C112—C111—C116—C115 | 0.7 (3) |
| C116—C111—C112—C113 | 0.3 (3) | N11—C111—C116—C115 | 177.51 (16) |
| N11—C111—C112—C113 | −176.63 (16) | C3—C2—N11—C111 | −177.32 (17) |
| C116—C111—C112—C12 | −179.96 (16) | C1—C2—N11—C111 | 3.2 (3) |
| N11—C111—C112—C12 | 3.1 (3) | C116—C111—N11—C2 | 48.9 (3) |
| C111—C112—C113—C114 | −0.6 (3) | C112—C111—N11—C2 | −134.19 (19) |
| C12—C112—C113—C114 | 179.64 (17) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| N11—H11···O12 | 0.91 | 1.90 | 2.6345 (19) | 136 |
Table 2 Comparative geometrical parameters (Å, °) for free and coordinated N,O-bidendate (N,O-bid) compounds
| Parameters | (I) | (II) | (III) | (IV) |
| N11—C111 | 1.422 (2) | 1.521 (4)/1.463 (3) | 1.440 (4) | - |
| N11—C2 | 1.345 (2) | 1.320 (4) | 1.319 (4) | 1.303 (6) |
| O12—C4 | 1.257 (2) | 1.290 (3) | 1.291 (4) | 1.281 (6) |
| C2—C3 | 1.383 (3) | 1.410 (4) | 1.423 (4) | 1.396 (7) |
| C3—C4 | 1.420 (2) | 1.365 (3) | 1.382 (3) | 1.388 (9) |
| N11···O12 | 2.635 (2) | 2.885 (3) | 2.886 (3) | 2.826 (6) |
| N11—C2—C4—O12 | -0.5 (1) | 4.1 (2) | -2.6 (2) | 1.2 (4) |
| Dihedral angle | 49.53 (5) | 87.47 (4)/89.36 (8) | 85.58 (8) | - |
Notes: (I) This work; (II) N,O-bid = 4-(2,3-dimethyl phenylamino)pent-3-en-2-onato (Venter et al., 2009a); (III) N,O-bid = 4-(2,6-dimethyl phenylamino)pent-3-en-2-onato (Venter et al., 2009b). (IV) N,O-bid = 4-amino-pent-3-en-2-onato (Damoense, et al., 1994).
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: BG2348).
References
- Brandenburg, K. & Putz, H. (2005). DIAMOND Crystal Impact GbR, Bonn, Germany.
- Brink, A., Visser, H. G., Steyl, G. & Roodt, A. (2010). Dalton Trans 39, 5572–5578. [DOI] [PubMed]
- Bruker (2004). SAINT-Plus and SADABS Bruker AXS Inc., Madison, Wisconsin, USA.
- Bruker (2005). APEX2 Bruker AXS Inc., Madison, Wisconsin, USA.
- Chen, H. & Rhodes, J. (1996). J. Mol. Med.74, 497–504. [DOI] [PubMed]
- Damoense, L. J., Purcell, W., Roodt, A. & Leipoldt, J. G. (1994). Rhodium Express, 5, 10–13.
- Farrugia, L. J. (1999). J. Appl. Cryst.32, 837–838.
- Nair, V. A., Suni, M. M. & Sreekumar, K. (2002). Proc. Indian Acad. Sci. (Chem. Sci.), 114, 481–486.
- Otto, S., Roodt, A., Swarts, J. C. & Erasmus, J. C. (1998). Polyhedron, 17, 2447–2453.
- Pyżuk, W., Krówczynsk, A. & Górecka, E. (1993). Mol. Cryst. Liq. Cryst.237, 75–84.
- Roodt, A. & Steyn, G. J. J. (2000). Recent Research Developments in Inorganic Chemistry, Vol. 2, pp. 1–23. Trivandrum: Transworld Research Network.
- Shaheen, F., Marchio, L., Badshah, A. & Khosa, M. K. (2006). Acta Cryst.E62, o873–o874.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Steyn, G. J. J., Roodt, A. & Leipoldt, J. G. (1992). Inorg. Chem 31, 3477–3481.
- Steyn, G. J. J., Roodt, A., Poletaeva, I. A. & Varshavsky, Y. S. (1997). J. Organomet. Chem.536/7, 197–205.
- Tan, H. Y., Loke, W. K., Tan, Y. T. & Nguyen, N.-T. (2008). Lab. Chip, 8, 885–891. [DOI] [PubMed]
- Van Aswegen, K. G., Leipoldt, J. G., Potgieter, I. M., Roodt, A. & Van Zyl, G. J. (1991). Transition Met. Chem 16, 369–371.
- Venter, G. J. S., Steyl, G. & Roodt, A. (2009a). Acta Cryst. E65, m1321–m1322. [DOI] [PMC free article] [PubMed]
- Venter, G. J. S., Steyl, G. & Roodt, A. (2009b). Acta Cryst. E65, m1606–m1607. [DOI] [PMC free article] [PubMed]
- Xia, M., Wu, B. & Xiang, G. (2008). J. Fluorine Chem.129, 402–408.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536810021045/bg2348sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536810021045/bg2348Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report