Abstract
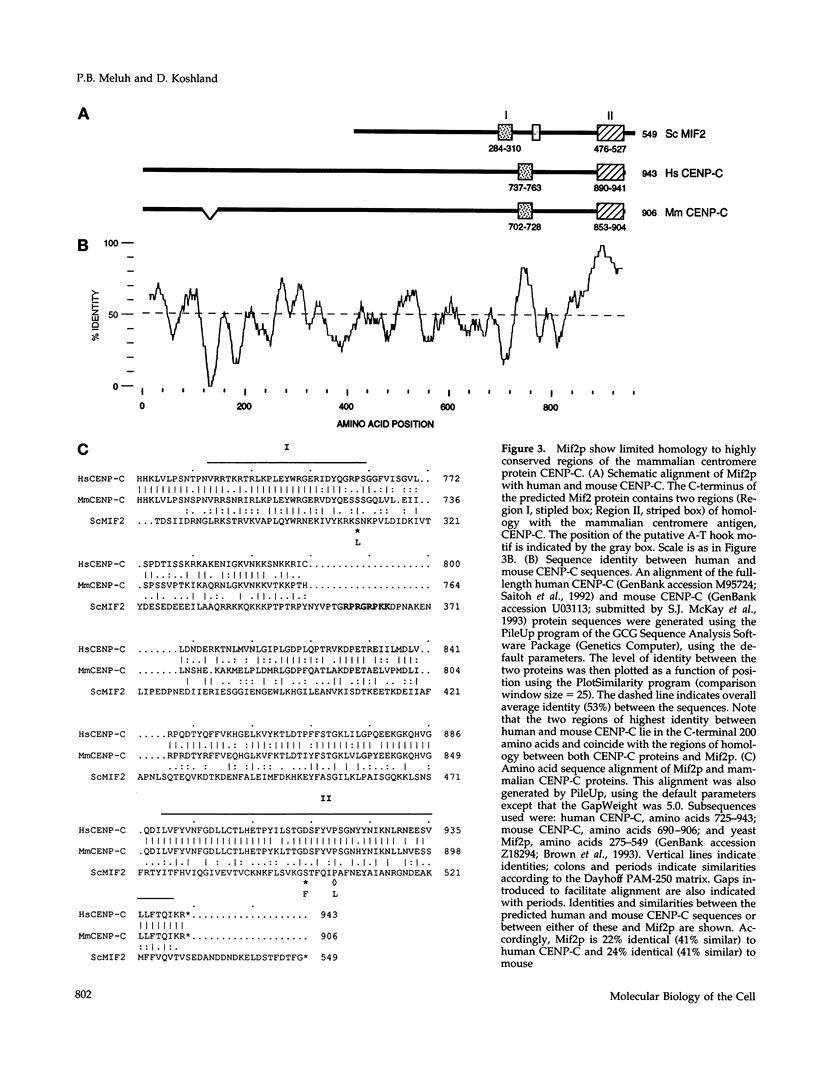
The MIF2 gene of Saccharomyces cerevisiae has been implicated in mitosis. Here we provide genetic evidence that MIF2 encodes a centromere protein. Specifically, we found that mutations in MIF2 stabilize dicentric minichromosomes and confer high instability (i.e., a synthetic acentric phenotype) to chromosomes that bear a cis-acting mutation in element I of the yeast centromeric DNA (CDEI). Similarly, we observed synthetic phenotypes between mutations in MIF2 and trans-acting mutations in three known yeast centromere protein genes-CEP1/CBF1/CPF1, NDC10/CBF2, and CEP3/CBF3B. In addition, the mif2 temperature-sensitive phenotype can be partially rescued by increased dosage of CEP1. Synthetic lethal interactions between a cep1 null mutation and mutations in either NDC10 or CEP3 were also detected. Taken together, these data suggest that the Mif2 protein interacts with Cep1p at the centromere and that the yeast centromere indeed exists as a higher order protein-DNA complex. The Mif2 and Cep1 proteins contain motifs of known transcription factors, suggesting that assembly of the yeast centromere is analogous to that of eukaryotic enhancers and origins of replication. We also show that the predicted Mif2 protein shares two short regions of homology with the mammalian centromere Ag CENP-C and that two temperature-sensitive mutations in MIF2 lie within these regions. These results provide evidence for structural conservation between yeast and mammalian centromeres.
Full text
PDF












Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Baker R. E., Fitzgerald-Hayes M., O'Brien T. C. Purification of the yeast centromere binding protein CP1 and a mutational analysis of its binding site. J Biol Chem. 1989 Jun 25;264(18):10843–10850. [PubMed] [Google Scholar]
- Baker R. E., Masison D. C. Isolation of the gene encoding the Saccharomyces cerevisiae centromere-binding protein CP1. Mol Cell Biol. 1990 Jun;10(6):2458–2467. doi: 10.1128/mcb.10.6.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bloom K. S., Carbon J. Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell. 1982 Jun;29(2):305–317. doi: 10.1016/0092-8674(82)90147-7. [DOI] [PubMed] [Google Scholar]
- Boeke J. D. One and two codon insertion mutants of bacteriophage f1. Mol Gen Genet. 1981;181(3):288–291. doi: 10.1007/BF00425599. [DOI] [PubMed] [Google Scholar]
- Bram R. J., Kornberg R. D. Isolation of a Saccharomyces cerevisiae centromere DNA-binding protein, its human homolog, and its possible role as a transcription factor. Mol Cell Biol. 1987 Jan;7(1):403–409. doi: 10.1128/mcb.7.1.403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown M. T., Goetsch L., Hartwell L. H. MIF2 is required for mitotic spindle integrity during anaphase spindle elongation in Saccharomyces cerevisiae. J Cell Biol. 1993 Oct;123(2):387–403. doi: 10.1083/jcb.123.2.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M. J., Davis R. W. Purification of a yeast centromere-binding protein that is able to distinguish single base-pair mutations in its recognition site. Mol Cell Biol. 1989 Jun;9(6):2544–2550. doi: 10.1128/mcb.9.6.2544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M., Davis R. W. Yeast centromere binding protein CBF1, of the helix-loop-helix protein family, is required for chromosome stability and methionine prototrophy. Cell. 1990 May 4;61(3):437–446. doi: 10.1016/0092-8674(90)90525-j. [DOI] [PubMed] [Google Scholar]
- Christianson T. W., Sikorski R. S., Dante M., Shero J. H., Hieter P. Multifunctional yeast high-copy-number shuttle vectors. Gene. 1992 Jan 2;110(1):119–122. doi: 10.1016/0378-1119(92)90454-w. [DOI] [PubMed] [Google Scholar]
- Churchill M. E., Travers A. A. Protein motifs that recognize structural features of DNA. Trends Biochem Sci. 1991 Mar;16(3):92–97. doi: 10.1016/0968-0004(91)90040-3. [DOI] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature. 1980 Oct 9;287(5782):504–509. doi: 10.1038/287504a0. [DOI] [PubMed] [Google Scholar]
- Cumberledge S., Carbon J. Mutational analysis of meiotic and mitotic centromere function in Saccharomyces cerevisiae. Genetics. 1987 Oct;117(2):203–212. doi: 10.1093/genetics/117.2.203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DePamphili M. L. How transcription factors regulate origins of DNA replication in eukaryotic cells. Trends Cell Biol. 1993 May;3(5):161–167. doi: 10.1016/0962-8924(93)90137-p. [DOI] [PubMed] [Google Scholar]
- Densmore L., Payne W. E., Fitzgerald-Hayes M. In vivo genomic footprint of a yeast centromere. Mol Cell Biol. 1991 Jan;11(1):154–165. doi: 10.1128/mcb.11.1.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doheny K. F., Sorger P. K., Hyman A. A., Tugendreich S., Spencer F., Hieter P. Identification of essential components of the S. cerevisiae kinetochore. Cell. 1993 May 21;73(4):761–774. doi: 10.1016/0092-8674(93)90255-O. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dowell S. J., Tsang J. S., Mellor J. The centromere and promoter factor 1 of yeast contains a dimerisation domain located carboxy-terminal to the bHLH domain. Nucleic Acids Res. 1992 Aug 25;20(16):4229–4236. doi: 10.1093/nar/20.16.4229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du W., Thanos D., Maniatis T. Mechanisms of transcriptional synergism between distinct virus-inducible enhancer elements. Cell. 1993 Sep 10;74(5):887–898. doi: 10.1016/0092-8674(93)90468-6. [DOI] [PubMed] [Google Scholar]
- Earnshaw W. C., Rothfield N. Identification of a family of human centromere proteins using autoimmune sera from patients with scleroderma. Chromosoma. 1985;91(3-4):313–321. doi: 10.1007/BF00328227. [DOI] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Clarke L., Carbon J. Nucleotide sequence comparisons and functional analysis of yeast centromere DNAs. Cell. 1982 May;29(1):235–244. doi: 10.1016/0092-8674(82)90108-8. [DOI] [PubMed] [Google Scholar]
- Funk M., Hegemann J. H., Philippsen P. Chromatin digestion with restriction endonucleases reveals 150-160 bp of protected DNA in the centromere of chromosome XIV in Saccharomyces cerevisiae. Mol Gen Genet. 1989 Oct;219(1-2):153–160. doi: 10.1007/BF00261171. [DOI] [PubMed] [Google Scholar]
- Gaudet A., Fitzgerald-Hayes M. Alterations in the adenine-plus-thymine-rich region of CEN3 affect centromere function in Saccharomyces cerevisiae. Mol Cell Biol. 1987 Jan;7(1):68–75. doi: 10.1128/mcb.7.1.68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaudet A., Fitzgerald-Hayes M. Mutations in CEN3 cause aberrant chromosome segregation during meiosis in Saccharomyces cerevisiae. Genetics. 1989 Mar;121(3):477–489. doi: 10.1093/genetics/121.3.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerring S. L., Spencer F., Hieter P. The CHL 1 (CTF 1) gene product of Saccharomyces cerevisiae is important for chromosome transmission and normal cell cycle progression in G2/M. EMBO J. 1990 Dec;9(13):4347–4358. doi: 10.1002/j.1460-2075.1990.tb07884.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goh P. Y., Kilmartin J. V. NDC10: a gene involved in chromosome segregation in Saccharomyces cerevisiae. J Cell Biol. 1993 May;121(3):503–512. doi: 10.1083/jcb.121.3.503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes F., Austin S. Topological scanning of the P1 plasmid partition site. J Mol Biol. 1994 Oct 21;243(2):190–198. doi: 10.1006/jmbi.1994.1646. [DOI] [PubMed] [Google Scholar]
- Hegemann J. H., Fleig U. N. The centromere of budding yeast. Bioessays. 1993 Jul;15(7):451–460. doi: 10.1002/bies.950150704. [DOI] [PubMed] [Google Scholar]
- Hegemann J. H., Shero J. H., Cottarel G., Philippsen P., Hieter P. Mutational analysis of centromere DNA from chromosome VI of Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jun;8(6):2523–2535. doi: 10.1128/mcb.8.6.2523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heintz N. H. Transcription factors and the control of DNA replication. Curr Opin Cell Biol. 1992 Jun;4(3):459–467. doi: 10.1016/0955-0674(92)90012-2. [DOI] [PubMed] [Google Scholar]
- Hieter P., Pridmore D., Hegemann J. H., Thomas M., Davis R. W., Philippsen P. Functional selection and analysis of yeast centromeric DNA. Cell. 1985 Oct;42(3):913–921. doi: 10.1016/0092-8674(85)90287-9. [DOI] [PubMed] [Google Scholar]
- Hoffman C. S., Winston F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene. 1987;57(2-3):267–272. doi: 10.1016/0378-1119(87)90131-4. [DOI] [PubMed] [Google Scholar]
- Hyman A. A., Middleton K., Centola M., Mitchison T. J., Carbon J. Microtubule-motor activity of a yeast centromere-binding protein complex. Nature. 1992 Oct 8;359(6395):533–536. doi: 10.1038/359533a0. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang W. D., Philippsen P. Purification of a protein binding to the CDEI subregion of Saccharomyces cerevisiae centromere DNA. Mol Cell Biol. 1989 Dec;9(12):5585–5593. doi: 10.1128/mcb.9.12.5585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang W., Lechner J., Carbon J. Isolation and characterization of a gene (CBF2) specifying a protein component of the budding yeast kinetochore. J Cell Biol. 1993 May;121(3):513–519. doi: 10.1083/jcb.121.3.513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kent N. A., Tsang J. S., Crowther D. J., Mellor J. Chromatin structure modulation in Saccharomyces cerevisiae by centromere and promoter factor 1. Mol Cell Biol. 1994 Aug;14(8):5229–5241. doi: 10.1128/mcb.14.8.5229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kingsbury J., Koshland D. Centromere-dependent binding of yeast minichromosomes to microtubules in vitro. Cell. 1991 Aug 9;66(3):483–495. doi: 10.1016/0092-8674(81)90012-x. [DOI] [PubMed] [Google Scholar]
- Koshland D., Kent J. C., Hartwell L. H. Genetic analysis of the mitotic transmission of minichromosomes. Cell. 1985 Feb;40(2):393–403. doi: 10.1016/0092-8674(85)90153-9. [DOI] [PubMed] [Google Scholar]
- Koshland D., Rutledge L., Fitzgerald-Hayes M., Hartwell L. H. A genetic analysis of dicentric minichromosomes in Saccharomyces cerevisiae. Cell. 1987 Mar 13;48(5):801–812. doi: 10.1016/0092-8674(87)90077-8. [DOI] [PubMed] [Google Scholar]
- Lechner J. A zinc finger protein, essential for chromosome segregation, constitutes a putative DNA binding subunit of the Saccharomyces cerevisiae kinetochore complex, Cbf3. EMBO J. 1994 Nov 1;13(21):5203–5211. doi: 10.1002/j.1460-2075.1994.tb06851.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lechner J., Carbon J. A 240 kd multisubunit protein complex, CBF3, is a major component of the budding yeast centromere. Cell. 1991 Feb 22;64(4):717–725. doi: 10.1016/0092-8674(91)90501-o. [DOI] [PubMed] [Google Scholar]
- Mann C., Davis R. W. Instability of dicentric plasmids in yeast. Proc Natl Acad Sci U S A. 1983 Jan;80(1):228–232. doi: 10.1073/pnas.80.1.228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masison D. C., O'Connell K. F., Baker R. E. Mutational analysis of the Saccharomyces cerevisiae general regulatory factor CP1. Nucleic Acids Res. 1993 Aug 25;21(17):4133–4141. doi: 10.1093/nar/21.17.4133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGrew J., Diehl B., Fitzgerald-Hayes M. Single base-pair mutations in centromere element III cause aberrant chromosome segregation in Saccharomyces cerevisiae. Mol Cell Biol. 1986 Feb;6(2):530–538. doi: 10.1128/mcb.6.2.530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meeks-Wagner D., Wood J. S., Garvik B., Hartwell L. H. Isolation of two genes that affect mitotic chromosome transmission in S. cerevisiae. Cell. 1986 Jan 17;44(1):53–63. doi: 10.1016/0092-8674(86)90484-8. [DOI] [PubMed] [Google Scholar]
- Mellor J., Jiang W., Funk M., Rathjen J., Barnes C. A., Hinz T., Hegemann J. H., Philippsen P. CPF1, a yeast protein which functions in centromeres and promoters. EMBO J. 1990 Dec;9(12):4017–4026. doi: 10.1002/j.1460-2075.1990.tb07623.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meluh P. B., Rose M. D. KAR3, a kinesin-related gene required for yeast nuclear fusion. Cell. 1990 Mar 23;60(6):1029–1041. doi: 10.1016/0092-8674(90)90351-e. [DOI] [PubMed] [Google Scholar]
- Middleton K., Carbon J. KAR3-encoded kinesin is a minus-end-directed motor that functions with centromere binding proteins (CBF3) on an in vitro yeast kinetochore. Proc Natl Acad Sci U S A. 1994 Jul 19;91(15):7212–7216. doi: 10.1073/pnas.91.15.7212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchison T. J. Microtubule dynamics and kinetochore function in mitosis. Annu Rev Cell Biol. 1988;4:527–549. doi: 10.1146/annurev.cb.04.110188.002523. [DOI] [PubMed] [Google Scholar]
- Murphy M. R., Fowlkes D. M., Fitzgerald-Hayes M. Analysis of centromere function in Saccharomyces cerevisiae using synthetic centromere mutants. Chromosoma. 1991 Dec;101(3):189–197. doi: 10.1007/BF00355368. [DOI] [PubMed] [Google Scholar]
- Ng R., Carbon J. Mutational and in vitro protein-binding studies on centromere DNA from Saccharomyces cerevisiae. Mol Cell Biol. 1987 Dec;7(12):4522–4534. doi: 10.1128/mcb.7.12.4522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niedenthal R. K., Sen-Gupta M., Wilmen A., Hegemann J. H. Cpf1 protein induced bending of yeast centromere DNA element I. Nucleic Acids Res. 1993 Oct 11;21(20):4726–4733. doi: 10.1093/nar/21.20.4726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niedenthal R., Stoll R., Hegemann J. H. In vivo characterization of the Saccharomyces cerevisiae centromere DNA element I, a binding site for the helix-loop-helix protein CPF1. Mol Cell Biol. 1991 Jul;11(7):3545–3553. doi: 10.1128/mcb.11.7.3545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oertel W., Mayer M. Structure and mitotic stability of minichromosomes originating in yeast cells transformed with tandem dimers of CEN11 plasmids. Mol Gen Genet. 1984;195(1-2):300–307. doi: 10.1007/BF00332763. [DOI] [PubMed] [Google Scholar]
- Panzeri L., Landonio L., Stotz A., Philippsen P. Role of conserved sequence elements in yeast centromere DNA. EMBO J. 1985 Jul;4(7):1867–1874. doi: 10.1002/j.1460-2075.1985.tb03862.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Saitoh H., Tomkiel J., Cooke C. A., Ratrie H., 3rd, Maurer M., Rothfield N. F., Earnshaw W. C. CENP-C, an autoantigen in scleroderma, is a component of the human inner kinetochore plate. Cell. 1992 Jul 10;70(1):115–125. doi: 10.1016/0092-8674(92)90538-n. [DOI] [PubMed] [Google Scholar]
- Saunders M., Fitzgerald-Hayes M., Bloom K. Chromatin structure of altered yeast centromeres. Proc Natl Acad Sci U S A. 1988 Jan;85(1):175–179. doi: 10.1073/pnas.85.1.175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schleif R. DNA looping. Annu Rev Biochem. 1992;61:199–223. doi: 10.1146/annurev.bi.61.070192.001215. [DOI] [PubMed] [Google Scholar]
- Schulman I., Bloom K. S. Centromeres: an integrated protein/DNA complex required for chromosome movement. Annu Rev Cell Biol. 1991;7:311–336. doi: 10.1146/annurev.cb.07.110191.001523. [DOI] [PubMed] [Google Scholar]
- Shore D., Langowski J., Baldwin R. L. DNA flexibility studied by covalent closure of short fragments into circles. Proc Natl Acad Sci U S A. 1981 Aug;78(8):4833–4837. doi: 10.1073/pnas.78.8.4833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sorger P. K., Severin F. F., Hyman A. A. Factors required for the binding of reassembled yeast kinetochores to microtubules in vitro. J Cell Biol. 1994 Nov;127(4):995–1008. doi: 10.1083/jcb.127.4.995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strunnikov A. V., Kingsbury J., Koshland D. CEP3 encodes a centromere protein of Saccharomyces cerevisiae. J Cell Biol. 1995 Mar;128(5):749–760. doi: 10.1083/jcb.128.5.749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thanos D., Maniatis T. The high mobility group protein HMG I(Y) is required for NF-kappa B-dependent virus induction of the human IFN-beta gene. Cell. 1992 Nov 27;71(5):777–789. doi: 10.1016/0092-8674(92)90554-p. [DOI] [PubMed] [Google Scholar]
- Thomas D., Jacquemin I., Surdin-Kerjan Y. MET4, a leucine zipper protein, and centromere-binding factor 1 are both required for transcriptional activation of sulfur metabolism in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Apr;12(4):1719–1727. doi: 10.1128/mcb.12.4.1719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tjian R., Maniatis T. Transcriptional activation: a complex puzzle with few easy pieces. Cell. 1994 Apr 8;77(1):5–8. doi: 10.1016/0092-8674(94)90227-5. [DOI] [PubMed] [Google Scholar]
- Tomkiel J., Cooke C. A., Saitoh H., Bernat R. L., Earnshaw W. C. CENP-C is required for maintaining proper kinetochore size and for a timely transition to anaphase. J Cell Biol. 1994 May;125(3):531–545. doi: 10.1083/jcb.125.3.531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao Z., McGrew J. T., Schroeder A. J., Fitzgerald-Hayes M. CSE1 and CSE2, two new genes required for accurate mitotic chromosome segregation in Saccharomyces cerevisiae. Mol Cell Biol. 1993 Aug;13(8):4691–4702. doi: 10.1128/mcb.13.8.4691. [DOI] [PMC free article] [PubMed] [Google Scholar]