Abstract
Hydroxyl radical protein footprinting coupled to mass spectrometry has been developed over the last decade and has matured to a powerful method for analyzing protein structure and dynamics. It has been successfully applied in the analysis of protein structure, protein folding, protein dynamics, and protein-protein and protein-DNA interactions. Using synchrotron radiolysis, exposures of proteins to a “white” x-ray beam for milliseconds provide sufficient oxidative modifications to surface amino acid side chains that can be easily detected and quantified by mass spectrometry. Thus, conformational changes in proteins or protein complexes can be examined using a time-resolved approach, which would be a valuable method for the study of macromolecular dynamics. In this review, we describe a new application of hydroxyl radical protein footprinting to probe the time evolution of the calcium-dependent conformational changes of gelsolin on the millisecond timescale. The data suggest a cooperative transition as multiple sites in different molecular sub-domains have similar rates of conformational change. These findings demonstrate that time-resolved protein footprinting is suitable for studies of protein dynamics that occur over periods ranging from milliseconds to seconds. In this review we also show how the structural resolution and sensitivity of the technology can be improved as well. The hydroxyl radical varies in its reactivity to different side chains by over two orders of magnitude, thus oxidation of amino acid side chains of lower reactivity are more rarely observed in such experiments. Here we demonstrate that selected reaction monitoring (SRM)-based method can be utilized for quantification of oxidized species, improving the signal to noise ratio. This expansion of the set of oxidized residues of lower reactivity will improve the overall structural resolution of the technique. This approach is also suggested as a basis for developing hypothesis driven structural mass spectrometry experiments.
General methods to study macromolecular structure and dynamics
The multidomain proteins that interact in large complexes play an important role in every cellular process (1, 2). Furthermore, the regulation of cell function is delicately balanced by the relative affinities of the various protein partners in such macromolecular assemblies, these interactions are regulated by the binding of ligands, nucleic acids, other proteins, metal ions, and by the structural effects of posttranslational modifications. Understanding of the detailed interactions of these multi-component assemblies is essential to understanding their functions. X-ray crystallography and nuclear magnetic resonance (NMR) are the methods of choice to reveal structural details and dynamics of macromolecules at the atomic level. However, both approaches have limitations, most notably for the analysis of structure and dynamics of proteins in conformational states of interest and are especially challenging for large complexes and membrane proteins. Alternative methods are needed in cases where it is not possible to determine high-resolution structure for a conformational state of interest or to confirm the structure of models. Due to significant improvement in sensitivity, speed and mass accuracy of mass spectrometry instruments over the past decade, structural mass spectrometry based technologies are increasingly being used to address these important questions. These approaches include hydrogen-deuterium exchange (3, 4, 5), chemical cross-linking (6,7) and covalent labeling (8, 9).
In hydrogen-deuterium exchange (HDX) amide protons in the backbone of a protein undergo exchange with the deuterium atoms from the deuterium water solutions mixed with protein. The amide hydrogens at the surface of protein that are not involved in stable secondary structure exchange very rapidly with deuterium atoms, while amide hydrogens that are buried or H-bonded have much slower exchange rates (10). Thus, the rate of hydrogen exchange is a function of protein structure and solvent accessibility that can be sensitively and accurately detected and quantified by mass spectrometry. To attenuate hydrogen back-exchange, which is typically observed during the analysis, low pH and low temperature must be utilized. However, only proteases with activity at acid pH can be used for generation of the peptides to be analyzed.
Chemical cross-linking methods utilize chemical reagents to form a covalent bond between functional groups of adjacent protein molecules. These covalent bonds can be formed within a protein (intramolecular cross-linking) and between different components of protein complexes (intermolecular cross-linking). The identification of intramolecular cross-linked sites that impose distance constraints on the location of two amino acid side chains by mass spectrometry can provide information on the three-dimensional structure of proteins. The identification of individual complexed proteins and their sites of interaction provides valuable information that can be used to define the orientation of interacting proteins in the protein complexes. However, the combinatorial nature of the cross-linking chemistry, the low abundance of specific interpeptide cross-links, and the typically complicated tandem MS fragmentation of cross-linked peptides are the primary challenges that must be overcome.
In covalent labeling approaches the structure and dynamics of proteins in various conformational states are probed by reacting surface accessible amino acid side chains with reagents that form covalent bonds. In these experiments either non-specific or amino acid-specific covalent labels are utilized. There are a number of amino acid-specific reagents that are currently available to map protein structure. These include: vicinal dicarbonyl compounds that modify arginine residues, carbodiimides to modify carboxyl groups in proteins, and various organic acid anhydrides that modify lysine residue (9). Iodoacetamide and its derivatives are extensively used to modify cysteines (9). However, the drawbacks of using these chemical reagents in protein footprinting experiments include preference of these reagents to a specific position within the protein, limiting the number of sites that can be probed. Approaches that use non-specific (or at least less specific) covalent labels generally rely on protein reactions with hydroxyl radicals (11–13) and provide structural resolution at the single amino acid level. The rate of modification of amino acid side chains in proteins depends on the reactivity of their side chains to hydroxyl radical attack and their solvent accessibility. The mass increases of peptides in hydroxyl radical footprinting and as in HDX methods are subsequently evaluated using protease cleavage and LC-MS analysis. In contrast to the deuterium-exchange approach, the highly reactive hydroxyl radicals react with surface accessible amino acid side chains (as opposed to the backbone) resulting in stable covalent modifications that facilitate downstream analysis. Furthermore, hydroxyl radicals have solvent properties (similar size and mobility) similar to water molecules (14), thus the extent of oxidation for any reactive amino acid depends directly on solvent accessibility. In addition, the site and extent of oxidation can be successfully measured by mass spectrometry for most amino acids providing structural information about macromolecular assemblies with the single amino acid resolution (11–13). An advantage of all three structural mass spectrometry techniques includes the ability to probe protein structure and dynamics at physiologically relevant protein concentrations for a wide range of solution conditions in the presence of multiple ligands (15–17). In this review we primarily focus on the studies that rely on synchrotron radiation to produce hydroxyl radicals using beamline X28C at National Synchrotron Light Source and illustrate some new approaches to probe time-resolved dynamics of conformational change as well as increase the sensitivity of detection using the power of selected reaction monitoring methods of mass spectrometry.
Chemistry of hydroxyl radical protein footprinting
Hydroxyl radicals can be generated by multiple methods including: transition metal-dependent chemical generation from peroxide, photolysis of peroxide, and radiolysis of water. Among the chemical methods, the most commonly used is Fenton chemistry (18), where hydroxyl radicals are generated by reaction between metal-bound ethylene diamine tetraacetic acid and hydrogen peroxide. Fenton chemistry is an inexpensive approach. However, it requires relatively long reaction times and uses chemical additives whose interactions with macromolecules may bias the observed local reactivity. More recent “rapid” Fenton methods, developed for nucleic acids footprinting, have decreased the exposure time considerably (19). On the other hand, techniques including radiolysis with x-rays, γ-rays, or electron beams (20–24) and UV-photodissociation of dilute hydrogen peroxide (25–28) generate hydroxyl radicals isotropically in biological solutions. Furthermore, the laser-induced dissociation of hydrogen peroxide and high flux X-rays from synchrotron sources generate stable oxidation products of macromolecular assemblies on microsecond to millisecond timescales. These last two approaches for hydroxyl radical generation have significant advantages compared to Fenton chemistry and γ-irradiation methods. Fist, rapid generation of hydroxyl radicals (whose concentration decays rapidly), can help ensure that oxidation events occur prior to any changes in macromolecular structure. Second, the time-resolved kinetic studies of protein dynamics are possible by monitoring the changes in oxidation extent as a function of time (29, 30).
The synchrotron bending magnet beamline can deliver between 1014 and 1015 photons/sec of a continuous spectrum x-ray beam (white beam) in the range of 3–30 keV. Hydroxyl radicals are generated at sub-micromolar steady state concentrations as the solution volume is irradiated with synchrotron light according to the following overall chemistry:
 |
(1) |
Under aerobic conditions, superoxide anions and hydroperoxy radicals are also generated according to:
 |
(2) |
These hydroxyl radicals react withamino acid side chains on the macromolecular surface on microsecond to millisecond timescales, depending on the sample concentration and scavenging effects of buffer or other components (31, 32). The relative reactivity of the amino acid side chains with respect to hydroxyl radicals under aerobic conditions have been established to be: Cys > Met > Trp > Tyr > Phe > Cystine > His > Leu ~ Ile > Arg ~ Lys ~ Val > Ser ~ Thr ~ Pro > Gln ~ Glu > Asp ~ Asn > Ala > Gly (21, 22, 33). The mechanism of amino acid side chain modifications largely depends on side chain chemistry. However, the predominate modifications of amino acid side chains are generated from incorporation of hydroxyl (or oxo) groups resulting in the mass shift of +16 (and +14) atomic mass units (amu). A number of residues also have other characteristic modifications including −30 amu mass shift for Asp and Glu, −43 amu for Arg, +32 and +48 amu mass shift for Cys, and −22, −10 and +5 amu mass shift for His. Overall, the published set of modified amino acid residues in proteins that contribute to structural analysis currently numbers 15 out of 20 (33).
LC-MS-based quantification of peptide oxidation in hydroxyl radical footprinting
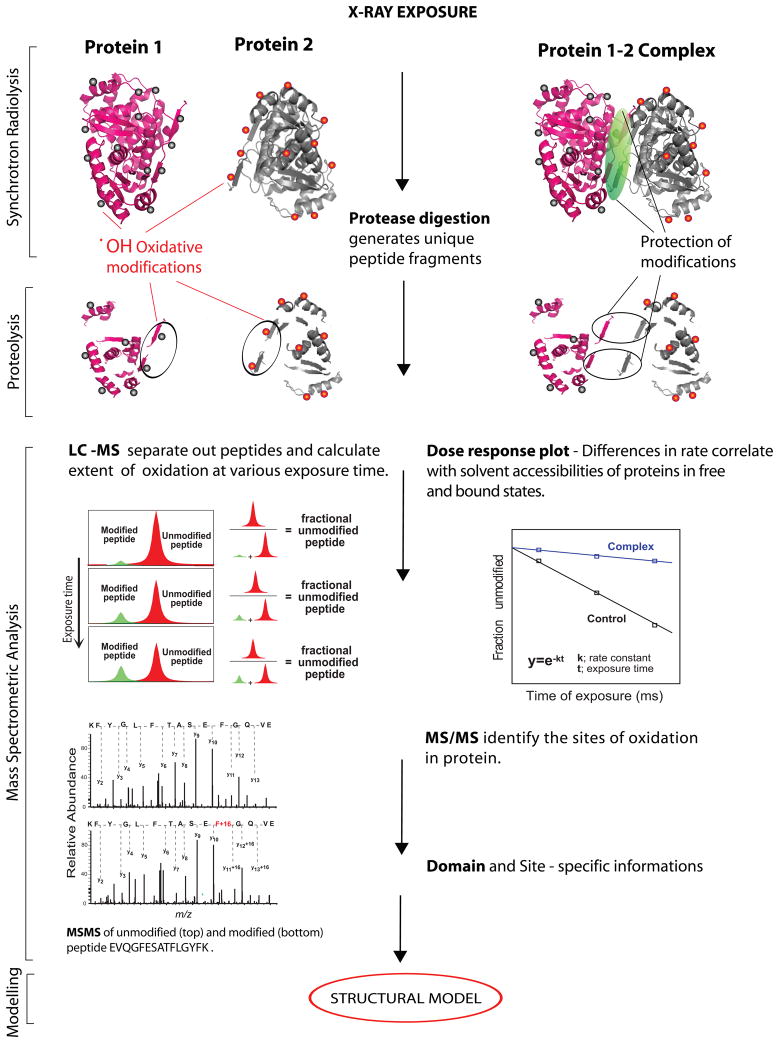
The experimental set up for the typical hydroxyl radical mediated footprinting experiment at X28C is illustrated in Figure 1. After exposure, proteins and their complexes are subjected to mass spectrometry analysis for quantification of peptide oxidation subsequent to protein digestion. Because the radiolytic modifications are covalent and stable, the range of experimental conditions such as different buffers or proteases, or reducing agents to digest heavily disulfide-bonded proteins, can be easily employed. Unmodified and modified proteolytic peptides in these footprinting experiments are typically separated by reverse-phase liquid chromatographic (LC) analysis and quantified using extracted ion current values derived from total ion current chromatogram (Figure 1). As the oxygen adducts result in the formation of more polar peptides, the modified peptides have a retention times different from that of unmodified counterparts in LC analysis. Typically, the peptides with oxygen (+14 or +16) adducts elute prior to the unmodified products (34). However, due to the different chemistry, peptides with modified Arg and His residues elute after unmodified peptides (35). Following MS analysis of the digest, the fraction of unmodified peptide is calculated using the ratio of the peak area under the ion signal of the unmodified peptide to the peak areas of total unmodified and modified ions derived from the selected ion current chromatogram (Figure 1). The rate of modification is calculated from the dose-response curve plotted for each peptide as a function of exposure time using a pseudo-first-order function. The identity of unmodified peptides and the modification sites of the oxidized counterparts are determined by tandem MS analysis. We utilized ProtMapMS software developed by our group (35) for the comprehensive automatic data analysis from hydroxyl radical footprinting experiments. Finally, domain- and site-specific information derived from MS-based hydroxyl radical footprinting analysis can be utilized for structural modeling of macromolecular assemblies.
Figure 1.
A schematic representation of MS-based hydroxyl radical footprinting experiments. Proteins and their complex are exposed to x-ray beam for a different time intervals. Oxidized proteins are subjected to proteolysis and mass spectrometry analysis. LC-MS is employed to separate out peptides and quantitatively measure the extent of modification. Dose-response profiles are plotted for each modified peptides within proteins sequences. Modification rates derived from free proteins and their complex are calculated and compared. The specific oxidation sites are determined by MS/MS analysis.
Time-resolved hydroxyl radical protein footprinting - new approach to study conformational dynamics of protein macromolecule assemblies
Understanding conformational dynamics in macromolecules and their complexes during processes such as ligand binding, folding, and macromolecular assembly is an important step towards providing detailed molecular descriptions of biological systems. An ideal method would analyze the conformational dynamics on timescales comparable with the conformational transition, monitor multiple sites within the molecule simultaneously, be compatible with a wide range of solution conditions for various sizes and classes of macromolecules and their macromolecular complexes, and utilize small amounts of material. These were the principles upon which time-resolved hydroxyl radical footprinting for nucleic acids was developed. In this paper we show how these concepts can be brought to bear to develop similar protein footprinting studies (30).
There are several biophysical approaches including fluorescence, NMR, and x-ray crystallography that can provide conformational dynamics information for proteins. Fluorescence spectroscopy has enviable time resolution, requires small sample quantities, provides solution phase information, and can probe interactions at both the solvent interface and the interior of a macromolecule. However, a fluorescent probe, including intrinsic fluorophores like Tryptophan, must be separately and repetitively incorporated in specific regions of the structure to probe multiple regions of the macromolecule and provide site-specific conformational information. This requires multiple forms of the protein to be generated along with multiple experiments. The desired fluorescent probes can be difficult to incorporate in the macromolecule of interest and may be limited by the overlap of the fluorescence spectra (30).
Other dynamics techniques such as NMR mediated deuterium exchange quench studies of protein folding (36) have yielded major insights into conformational dynamics of macromolecules with high structural resolution. However, the major disadvantages for these powerful exchange methods are limitation of the size of the proteins or complexes that can be analyzed and the need for quenching methods to “freeze” the exchange.
Time-resolved x-ray crystallography methods have advanced considerably in recent years (37). This approach has been primarily successful in the examination of structural intermediates that have been generated by flash photolysis and examined in real-time or trapped with cryogenic conditions (38, 39). Major limitations of these methods are the amounts of material needed, the difficulty in crystallizing large complexes, and substrate diffusion and dynamics of the protein may be inhibited by the crystal lattice (37, 40).
Time-resolved synchrotron footprinting provides a method that probes the solvent accessible surface of macromolecules and their biologically relevant complexes (30, 41). The formation of stable modifications provides an ability to utilize a wide range of gel and/or mass spectrometry techniques to identify the sites and quantify the extent of modification and/or cleavage at the amount of material three orders of magnitude less than needed for NMR. Time-resolved footprinting method was first introduced by Sclavi et al. (42) to study folding of nucleic acids. In principle, the rapid mixing of nucleic acids with solutions that contains metal ions prior synchrotron X-ray exposure allows nucleic acid conformational changes and ligand binding reactions to be followed on a timescales of millisecond to minutes with single nucleotide resolution. In these experiments the irradiation of nucleic acids with synchrotron light results in the cleavage in the DNA or RNA strand in a sequence-independent reaction (18). The change in hydroxyl radical protection for every nucleotide in a nucleic acid chain is monitored using SDS polyacrylamide gel electrophoresis and due to the high resolution of nucleic acid gel methods, single nucleotide structural resolution is achieved. Introduction of this method created a powerful approach for conducting comprehensive thermodynamic and kinetic studies of nucleic acids (42–44). Similar approaches can be utilized for time-resolved kinetic studies of protein assemblies by monitoring the transitions undergone by proteins on the timescales of milliseconds to seconds. In contrast with nucleic acid footprinting, where cleavage of the macromolecule is required, synchrotron exposure of proteins for milliseconds provides sufficient oxidized product to be analyzed by mass spectrometry. If conformational changes in the protein or protein complexes appear on the milliseconds or seconds timescale, resulting in the change in solvent accessibility of a side chain probe, a time-resolved protein footprinting can be a valuable approach to understanding the mechanism of conformational change.
Time-resolved footprinting: experimental design to study evolution of Ca2+-dependent gelsolin activation
In this section we show the dynamic changes in gelsolin conformation upon Ca2+ activation employing time-resolved protein footprinting. Gelsolin is a Ca2+-dependent actin regulatory protein that severs actin filaments and caps the fast-growing barbed end with high affinity. This ability to remodel the actin filaments results in gelsolin playing a key role in cell motility and differentiation processes (45). Gelsolin protein contains six repeats of a 15 kDa domain with overall similar structural organization that are separated into an N-terminal half (S1–S3) and a C-terminal half (S4–S6) of the molecule (Figure 2). The helical tail of S6 (Figure 2) interacts in a noncovalent manner with F-actin binding helix of S2, masking its actin-binding sites in the absence of Ca2+ (45). It was postulated that in the presence of Ca2+, the S6 helical tail is unlatched from S2, and the interface between S4 and S6 β-sheets where a Ca2+-binding site is located are disrupted in order to providing additional binding sites to actin filaments (46). We used a time-resolved protein footprinting approach to observe the time evolution of the calcium dependent conformational changes seen for gelsolin. The rapid mixing of gelsolin with buffer containing Ca2+ and then exposure of the solution to millisecond doses of radiation after specific delay times were used to define potential structural-kinetic intermediates in the activation process.
Figure 2.

Ribbon representation of the subunit structure of horse plasma gelsolin highlighting the six homologues domains such as S1 (blue), S2 (magenta), S3 (green), S4 (cyan), S5 (yellow), S6 (red), the C-terminal tail (orange) and the subdomain linkers (grey).
In these experiments, we utilized a commercial Kin-Tek stopped flow device. A “push-pause” timing sequence for the motor powering the mixer was applied. For example, a set of time-resolved footprinting experiments was carried out in which 20 μl of the 4 μM of protein solution (in 10 mM cacodylic acid sodium salt trihydrate with 0.5 mM EGTA, pH 7.0) in one syringe was rapidly mixed with 20 μl of 10 mM CaCl2 solution (in 10 mM cacodylic acid sodium salt trihydrate, 0.5 mM EGTA, pH 7.0) in the other syringe providing a final concentration of 2 μM of protein and 5 mM of Ca2+. At these final concentrations of Ca2+, we have previously shown that both the S2/S6 and S4/S6 interactions are disrupted as footprinting probe sites within these sub-domains show increased oxidation as a function of Ca2+(16). The solution of activated gelsolin was passed through the X-ray beam and the sample collected. The exposure time depends on the flow rate of the sample solution through the beam and the beam size. In all experiments, a constant X-ray exposure time of 50 ms was employed while the reaction time between gelsolin sample and the Ca2+ buffer or ligand was varied from 10 milliseconds to 3 seconds. During and after the mixing time (dead time < 5 ms), calcium-induced conformational changes are expected to occur. The X-ray beam was used to provide a snapshot of solvent exposure at the specific delay times after mixing gelsolin with Ca2+. The dynamics of Ca2+ activation was studied upon different final Ca2+ concentrations such as 0.5 μM, 10 μM, 100 μM and 5 mM chosen based on our previous equilibrium studies (16). Two control points including Ca2+-free samples and samples containing 5 mM Ca2+ gelsolin were also exposed.
Following tryptic digestion of the exposed protein samples, the specific peptides 49–72 within S1 of gelsolin, 431–454 within S4, and 722–748 within S6, containing the sites of interest, were analyzed for time-dependent changes in their modifications using mass spectrometry (16). The amount of modified fraction for each of these samples was quantified using our published approach and normalized to the endpoints represented by 0 and 5 mM Ca+2 as previously described (47). Eleven reaction time points ranging from 10 ms to 3 seconds delay were used to construct a progress curves for these peptides to monitor the Ca2+-induced rearrangement of the gelsolin sub-units.
Time-resolved footprinting revealed Ca2+-dependent dynamic rearrangements of gelsolin sub-domains
The first series of time-resolved footprinting experiments were carried out where 4 μM of gelsolin was mixed with 1 μM of Ca2+ in the ratio 1:1 and the samples exposed with delay time ranging from 10 ms to 3 sec. Progress curves for peptides 49–72, 722–748 and 431–454 were obtained by plotting the oxidized fraction normalized to 1 for each peptide (representing the endpoint of modification) as a function of the reaction time. At the final concentration of 0.5 μM Ca2+ there were no changes in the extent of the oxidation for any peptides compared to the zero Ca2+ control points (data not shown). This concentration was below the threshold for observing any conformational changes based on our previous studies. As the final Ca2+ concentration was increased to 10 μM, peptides 49–72 and 722–748 showed changes in the extent of modification (progress curves in Figure 3A) with increase of the delay time with rates of 4.1 s−1 and 3.5 s−1 respectively (Figure 3A). These two rates are essentially equal showing that these conformational changes are apparently concerted. In contrast, the peptide composed of residues 431–454 did not experience any changes in the extent of the oxidation for 10 μM of Ca+2 concentration, as this concentration of ion was below the binding affinity of the site monitored by these residues (Figure 3A). Specifically, in our equilibrium studies we showed that protein regions 49–72 (S1) and 722–748 (S6 and S6 tail helix) were sensitive to sub-micromolar Ca2+-binding events that saturate at approximately 10 μM (16), while region 431–454, which is adjacent to the salt-link that couples the S4/S6 beta-sheet, shows no changes at these low concentrations of calcium. Overall, the data clearly identifies homogenous conformational changes associated with the specific peptide probes. The data are well fit by single exponential functions (represented by the solid curves) with only modest deviations from the fits and saturation regions of the curve well matched to the independently derived endpoints. No burst phases were evident and overall the conformational dynamics were well characterized. Moreover, gelsolin segments including 371–386, 571–588 and 600–621 did not exhibit any changes in the extent of oxidation upon different Ca2+ concentration (data not shown), which is fully consistent with their lack of response to Ca+2 dependent changes in equilibrium studies (16).
Figure 3.

Progress curve at 10 μM (A), 100 μM (B) and 5 mM (C) Ca2+ for peptides comprised of residues 49–72, 722–748 and 431–454 within gelsolin protein after exposure for 50 ms.
As the Ca2+ concentration was increased to 100 μM, where the S4/S6 site can be occupied, dynamic changes in the extent of the modification were observed for all three protein regions with the rates increasing slightly to ~7 s−1(Figure 3B). At 5 mM of Ca2+ we observed progress curves for all three peptides at 49–72, 431–454 and 722–748 similar to those at 100 μM final Ca2+(Figure 3C). Overall the rates are relatively insensitive to the final Ca+2 concentration, at least within the measurement errors. For example, as the final Ca+2 concentration was increased 500-fold from 10 μM to 5 mM the rate at which the conformation change took place was essentially unchanged for peptides 49–72 and 722–748. Below the apparent Kd of binding for peptide 431–454 there was no conformational change at all, but the increase in Ca+2 concentration of 50-fold (100 μM to 5 mM) had a negligible effect on the rate. This lack of rate effect as a function of changes in ligand concentration suggests that binding and the associated conformational changes are tightly linked in the presence of saturating calcium ion. These results demonstrate that in the presence of 100 μM or higher Ca2+ the release of the “S6 latch” and the disruption of the S4/S6 β-sheet structure represent a concerted if not cooperative process (16).
Overall, these time-resolved footprinting data are consistent with a three state Ca2+-induced activation process as a function of Ca2+ concentration (16, 48) in which state 1 corresponds to the “Ca2+ -free” form; state 2 involves partial unlatching of the structure mediated by occupancy of high-affinity Ca2+- binding sites; while state 3 is the Ca2+-saturated fully activated gelsolin form. The transition between state 2 and 3 is mediated by occupancy of additional Ca2+ ions in lower-affinity binding sites (16). However, when high enough Ca2+ (final concentration) is used, the kinetic activation process is 2-state, independent of Ca2+ concentration, and entirely concerted. These data illustrate how our mass-spectrometry based quantification approach is very effective for biophysical analysis and that hydroxyl radical mediated protein footprinting is a very effective technique to monitor time-resolved dynamics of proteins.
SRM-based approach for quantitation of tandem MS ion data to increase resolution in protein footprinting
As hydroxyl radical protein footprinting can monitor changes in the oxidation extent of the solvent accessible and reactive amino acid side chains, increasing the number of probes in the protein is essential for achieving the highest possible structural resolution for the technique. The modification products produced as the result of the reaction between the sulfur-containing and aromatic amino acid residues and hydroxyl radicals can be easily detected in MS experiments. However, less reactive residues such as Gly, Ala, Asp, Asn, Arg, Glu and Gln result in less abundant oxidized species, this coupled with poor ionization efficiency for many of the “nonstandard” oxidation products (e.g. for Arg, Asp, Glu), can make detection of the oxidation of these amino acid side chains challenging, in fact these products are not routinely detectible by mass spectrometry (21, 22, 49). Ser and Thr, also have reaction products that are difficult to detect, presumably due to low ionization efficiency of the products. Overall, about half of the 20 naturally occurring amino acids are routinely used as probes in protein footprinting experiments across many laboratories and the oxidation products from these amino acids are relatively easily detected by mass spectrometry. In some cases, we have observed oxidation for other less reactive residues including Ala, Arg and Glu (50,51), increasing the published set of oxidized probes in proteins that contribute to structural analysis (33).
Up to now, a data dependent acquisition approach (DDA) for mass spectrometry is the method commonly used by many laboratories for hydroxyl radical protein footprinting experiments. In DDA the mass spectrometer is typically set up such that the first survey scan monitors a wide range of ion masses from 350 to 1600 m/z, then the second measurement is a tandem MS experiment of the five to eight most intense peptide ions derived from the first scan event. A limitation of this approach contributes to the difficulty in detecting the low abundant or “non-standard” products described above that may not be included the set of most intense masses “switched on” in a DDA analysis.
To address this problem, we propose the use of selected reaction monitoring (SRM)-based method in order to quantify the oxidation extent of selected peptides that contain residues with low reactivity. This approach relies on the existing knowledge that has been developed with respect to the oxidation products from peptides of known sequence (16, 35). Hypothesized product peptides can be predicted based on amino acid reactivity, solvent accessibility, and knowledge of radiolytic chemistry for various amino acid side chains (33). Because the proteolytic peptides and the typical oxidized products have very similar structures and a molecular mass that is shifted by +16 or +32 amu (corresponding to the addition of one or two oxygen atoms, respectively), the ionization and detection efficiencies of these peptides are relatively comparable. However, due to the oxygen adducts that result in the formation of more polar peptides, the modified peptides are typically eluted from the reverse phase column one-half to several minutes prior the unmodified peptides. For example, methionine-containing peptides that are oxidized by one and two oxygen are eluted from the column a few minutes prior to the unmodified peptide, while oxidized phenylalanine-, tyrosine- and tryptophan-containing peptides are eluted from the column typically one-half to one minute prior to their unoxidized peptide (11,17). In contrast, peptides containing histidine and arginine amino acids that suffer ring-open and loss of guanidino group, respectively, are eluted two to five minutes after the unmodified peptide. Thus, the retention times of the oxidized products ions will be different from those of the unoxidized ions, and ionization efficiency may be different as well. This is different from the standard SRM technique where the retention time for mass shifted and unshifted ions is identical. However, this is not a problem in the proposed experiments due to the existing extensive knowledge about oxidation products and their elution times and the fact that we first employ a DDA analysis in all footprinting experiments that provides an accurate identification of the unmodified peptide and clearly defines its retention time as an anchor for the experimental approach. Furthermore, selection of peptides for the SRM-based experiments in hydroxyl radical footprinting include the following criteria: a peptide mass preferably in the 1000–2500 range provides a range of fragment ions in the tandem MS spectra to choose for SRM monitoring, the unmodified peptides have to be “well ionized”, and a solvent accessible amino acid residue of interest must be present in the peptide. In the SRM-based experiment, we set up the mass spectrometer in such a way that the “parent” ions, representing the unoxidized and oxidized peptides separately identified in the chromatogram, are isolated with the isolation width of 1.6–2 amu, and then subjected to tandem fragmentation. Two to three desired fragments are monitored by mass spectrometry for each “parent ion”. The peak areas for these detected tandem fragment ions from both the unmodified and modified peptides are calculated and summed, respectively. All fragment ions representing the unmodified peptide have the same retention time, while all fragment ions derived from the modified peptide(s) also have identical elution time but varying from that for the unmodified counterpart by a fixed amount. Thus, monitoring a few fragment ions for modified and unmodified peptides is recommended to assure that the correct ions are interrogated. Furthermore, the fraction of unmodified peptide (extent of oxidation) of interest is calculated from the ratio of the summed peak areas of the tandem fragments for unmodified peptide to the sum total of all ion fragments (sum of the peak areas for unmodified and modified ion fragments) for each exposure time.
To demonstrate the advantage of this method for the analysis of low abundant oxidative products in comparison with conventional DDA approach, Angiotensin II peptide with sequence DRVYIHPF was exposed and analyzed. The retention time and m/z value of fragment ions for this peptide that gives the highest detection sensitivity by tandem MS analysis was determined for the unoxidized peptide using DDA. The same product ions were assumed to give the highest detection sensitivity for the oxidized peptides. Oxidized products for Arg and His residues within Angiotensin II were monitored using DDA and SRM analysis to compare the sensitivity of both techniques. All experiments were carried out at four exposure time points for each sample in three replicates. The extent of oxidation was calculated for Arg and His residues from DDA and SRM analysis for each exposure time as described above. Dose-response curves are presented in Figure 4 as fraction of unmodified peptide versus x-ray exposure time. The modification product of −43 amu on Arg residue and the modification products of +16, +5, −6, −10 and −22 amu on His residue were monitored throughout all experiments. While, the dose-response curves derived from DDA and SRM analysis overall reflect the first-order process, the rate constants for modification of Arg and His residues derived from the plots with DDA experiments were calculated to be slightly greater than that from the SRM experiments. We suspect this is the case as the numerator ion current values in the DDA analysis contain more noise component with respect to the signal compared to the case of the SRM experiment, e.g. the DDA experiment overstates that actual signal and reports a larger extent of oxidation. Supporting this notion, the SRM-based data analysis of three replicates was observed to be more precise (with an r squared value of approximately 0.99 and a ~1% reported error in the rate constant) than that from DDA analysis (with an r squared value of approximately 0.96 and ~3% reported error). These observations suggests that the signal to noise level in the SRM-based method is greater than in DDA approach and implies an overall increase in the sensitivity for SRM method.
Figure 4.
Dose-response curves for the oxidation of Angiotensin II (DRVYIHPF) peptide as a function of exposure time. A, dose-response curve for the Arg oxidation derived from DDA analysis. B, dose-response curve for the His oxidation derived from DDA analysis. C, dose-response curve for the Arg oxidation derived from SRM analysis. B, dose-response curve for the His oxidation derived from SRM analysis. Fragment ion b6 was monitored in all SRM experiments.
To more precisely define the sensitivity differences, we monitored two low abundant oxidized products including +5 and −6 amu modifications on His residue by DDA and SRM experiments. In order to quantify the specific sites of oxidation (the oxidized His residue in this case), two SRM transitions (in m/z) such as 526.2→789.4 and 520.8→778.3 were monitored. The doubly charged ions with m/z of 526.2 and 520.8 correspond to the “parent” ions of the oxidized peptides that have mass shifts of +5 and −6 amu, respectively, while singly charged ions with m/z of 789.4 and 778.3 represent b6–fragment ions that have mass shifts of +5 and −6 amu, respectively, corresponding to the oxidation of His residue (Figure 5E, F). These are compared to the b6–fragment ion intensity for the transition 523.8→ 784.5 for unmodified ion that has no mass shift (Figure 5D). For comparison, the extracted ion current values corresponding to the doubly charged “parent” ions with m/z of 523.8, 526.2 and 520.8 were monitored using the conventional DDA method (Figure 5A, B, C). The integrated areas of these unmodified and modified “parent” ions and unmodified and modified fragment ions were obtained from selected ion chromatograms in DDA and SRM spectra, respectively along with the signal to noise ratio (Figure 5). For the +5-His modification no significant extracted ion current value could be seen in Figure 5B while the SRM approach had a clearly defined extracted ion intensity associated with the modified species. For the −6 amu modification, although a clear signal could be seen in the DDA experiment, the signal to noise ratio was dramatically improved using the SRM approach.
Figure 5.
DDA vs SRM analysis of Angiotensin II peptide exposed for 40 ms. (A) DDA of doubly protonated ion with m/z= 523.6 oxidized on His; (B) DDA of doubly protonated ion with m/z= 526.2 that oxidized on His; (C) DDA of doubly protonated ion with m/z= 520.8 that oxidized on His; (D) SRM analysis of transition 523.6→784.5; (E) SRM of transition 526.2→789.4; (F) SRM of transition 520.8→778.3. Fragment ion b6 was monitored in all SRM experiments.
Overall, the sensitivity of the detection and most importantly the quantification of the oxidized products was increased ~100-fold using the SRM approach. No oxidized product of +5 amu was observed by DDA, while SRM method yields quantifiable oxidized products that were eluted approximately 5 min after unmodified peptide. Note that the unoxidized and oxidized peptides have distinct m/z values for the “parent” ions selected for fragmentation, and the retention time for both peptides will be different but predictable as described above.
In summary, in the typical protein footprinting analysis oxidation rate for peptides containing sulfur and aromatic residues, as well as some aliphatic residues are easily determined by DDA mass spectrometry analysis. Peptides that potentially involved in binding interfaces and contain solvent accessible amino acids with low reactivity such as glutamine, serine, threonine, lysine, arginine and alanine can be subjected to detailed examination using the SRM experiment in a targeted fashion. Thus, this SRM-based footprinting analysis can overall increase the structural resolution of hydroxyl radical footprinting.
Acknowledgments
We would like thank Sayan Gupta for assistance at the X28C beamline at National Synchrotron Light Source and Paul Janmey for providing gelsolin. This work was supported in part by grants from the NIH, National Institute for Biomedical Imaging and Bioengineering, P30-EB-09998 and R01-EB-09688. The NSLS is supported by the Department of Energy.
References
- 1.Sali A, Glaeser R, Earnest T, Baumeister W. From words to literature in structural proteomics. Nature. 2003;422:216. doi: 10.1038/nature01513. [DOI] [PubMed] [Google Scholar]
- 2.Russell RB, Albert F, Aloy P, Davis FP, Korkin D, Pichaud M, Topf M, Sali A. A structural perspective on protein-protein interactions. Curr Opin Struc Biol. 2004;14:313. doi: 10.1016/j.sbi.2004.04.006. [DOI] [PubMed] [Google Scholar]
- 3.Woods VL, Jr, Hamuro Y. High resolution, high-throughput amide deuterium exchange-mass spectrometry (DXMS) determination of protein binding site structure and dynamics: utility in pharmaceutical design. J Cell Biochem Suppl. 2001;37:89. doi: 10.1002/jcb.10069. [DOI] [PubMed] [Google Scholar]
- 4.King D, Bergmann C, Orlando R, Benen JA, Kester HC, Visser J. Use of amide exchange mass spectrometry to study conformational changes within the endopolygalacturonase II-homogalacturonan-polygalacturonase inhibiting protein system. Biochemistry. 2002;41:10225. doi: 10.1021/bi020119f. [DOI] [PubMed] [Google Scholar]
- 5.Englander JJ, Del Mar C, Li W, Englander SW, Kim JS, Stranz DD, Hamuro Y, Woods VL., Jr Protein structure change studied by hydrogen-deuterium exchange, functional labeling, and mass spectrometry. Proc Natl Acad Sci U S A. 2003;100:7057. doi: 10.1073/pnas.1232301100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sinz A. Chemical cross-linking and mass spectrometry for mapping three-dimensional structures of proteins and protein complexes. J Mass Spectrom. 2003;38:1225. doi: 10.1002/jms.559. [DOI] [PubMed] [Google Scholar]
- 7.Novak P, Giannakopulos AE. Chemical cross-linking and mass spectrometry as structure determination tools. Eur J Mass Spectrom (Chichester, Eng) 2007;13:105. doi: 10.1255/ejms.868. [DOI] [PubMed] [Google Scholar]
- 8.Maleknia SD, Ralston CY, Brenowitz MD, Downard KM, Chance MR. Determination of macromolecular folding and structure by synchrotron x-ray radiolysis techniques. Anal Biochem. 2001;289:103. doi: 10.1006/abio.2000.4910. [DOI] [PubMed] [Google Scholar]
- 9.Mendoza LM, Vachet RW. Probing protein structure by amino acid-specific covalent labeling and mass spectrometry. Mass Spectrom Rev. 2009;28:785. doi: 10.1002/mas.20203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Busenlehner LS, Armstroneng RN. Insights into enzyme structure and dynamics elucidated by amide H/D exchange mass spectrometry. Arch Biochem Biophys. 2004;433:34. doi: 10.1016/j.abb.2004.09.002. [DOI] [PubMed] [Google Scholar]
- 11.Maleknia SD, Brenowitz M, Chance MR. Millisecond radiolytic modification of peptides by synchrotron X-rays identified by mass spectrometry. Anal Chem. 1999;71:3965. doi: 10.1021/ac990500e. [DOI] [PubMed] [Google Scholar]
- 12.Kiselar JG, Maleknia SD, Sullivan M, Downard KM, Chance MR. Hydroxyl radical probe of protein surfaces using synchrotron X-ray radiolysis and mass spectrometry. Int J Radiat Biol. 2002;78:101. doi: 10.1080/09553000110094805. [DOI] [PubMed] [Google Scholar]
- 13.Guan JQ, Vorobiev S, Almo SC, Chance MR. Mapping the G-actin binding surface of cofilin using synchrotron protein footprinting. Biochemistry. 2002;41:5765. doi: 10.1021/bi0121104. [DOI] [PubMed] [Google Scholar]
- 14.Buxton GV, Greenstock CL, Helman WP, Ross AB. Critical Reveview of rate constant for reactions of hydrated electrons, hydrogen atoms and hydroxyl radicals ((•OH/•O− in Aqueous Solution. J Phys Chem Ref Data. 1988;17:513. [Google Scholar]
- 15.Chance MR. Unfolding of apomyoglobin examined by synchrotron footprinting. Biochem Biophys Res Commun. 2001;287:614. doi: 10.1006/bbrc.2001.5628. [DOI] [PubMed] [Google Scholar]
- 16.Kiselar JG, Janmey PA, Almo SC, Chance MR. Visualizing the Ca2+-dependent activation of gelsolin by using synchrotron footprinting. Proc Natl Acad Sci U S A. 2003;100:3942. doi: 10.1073/pnas.0736004100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kiselar JG, Mahaffy R, Pollard TD, Almo SC, Chance MR. Visualizing Arp2/3 complex activation mediated by binding of ATP and WASp using structural mass spectrometry. Proc Natl Acad Sci U S A. 2007;104:1552. doi: 10.1073/pnas.0605380104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tullius TD, Dombroski BA. Hydroxyl radical “footprinting”: high-resolution information about DNA-protein contacts and application to lambda repressor and Cro protein. Proc Natl Acad Sci U S A. 1986;83:5469. doi: 10.1073/pnas.83.15.5469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Shcherbakova I, Mitra S, Beer RH, Brenowitz M. Fast Fenton footprinting: a laboratory-based method for the time-resolved analysis of DNA, RNA and proteins. Nucleic Acids Res. 2006;34:e48. doi: 10.1093/nar/gkl055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Greiner DP, Hughes KA, Meares CF. Radiolytic protein surface mapping. Biochem Biophys Res Commun. 1996;225:1006. doi: 10.1006/bbrc.1996.1285. [DOI] [PubMed] [Google Scholar]
- 21.Xu G, Takamoto K, Chance MR. Radiolytic modification of basic amino acid residues in peptides: probes for examining protein-protein interactions. Anal Chem. 2003;75:6995. doi: 10.1021/ac035104h. [DOI] [PubMed] [Google Scholar]
- 22.Xu G, Chance MR. Radiolytic modification of sulfur-containing amino acid residues in model peptides: fundamental studies for protein footprinting. Anal Chem. 2005;77:2437. doi: 10.1021/ac0484629. [DOI] [PubMed] [Google Scholar]
- 23.Takamoto K, Chance MR. Radiolytic protein footprinting with mass spectrometry to probe the structure of macromolecular complexes. Annu Rev Biophys Biomol Struct. 2006;35:251. doi: 10.1146/annurev.biophys.35.040405.102050. [DOI] [PubMed] [Google Scholar]
- 24.Sharp JS, Tomer KB. Analysis of the oxidative damage-induced conformational changes of apo- and holocalmodulin by dose-dependent protein oxidative surface mapping. Biophys J. 2007;92:1682. doi: 10.1529/biophysj.106.099093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sharp JS, Becker JM, Hettich RL. Analysis of protein solvent accessible surfaces by photochemical oxidation and mass spectrometry. Anal Chem. 2004;76:672. doi: 10.1021/ac0302004. [DOI] [PubMed] [Google Scholar]
- 26.Sharp JS, Guo JT, Uchiki T, Xu Y, Dealwis C, Hettich RL. Photochemical surface mapping of C14S-Sml1p for constrained computational modeling of protein structure. Anal Biochem. 2005;340:201. doi: 10.1016/j.ab.2005.02.005. [DOI] [PubMed] [Google Scholar]
- 27.Aye TT, Low TY, Sze SK. Nanosecond laser-induced photochemical oxidation method for protein surface mapping with mass spectrometry. Anal Chem. 2005;77:5814. doi: 10.1021/ac050353m. [DOI] [PubMed] [Google Scholar]
- 28.Hambly DM, Gross ML. Laser flash photolysis of hydrogen peroxide to oxidize protein solvent-accessible residues on the microsecond timescale. J Am Soc Mass Spectrom. 2005;16:2057. doi: 10.1016/j.jasms.2005.09.008. [DOI] [PubMed] [Google Scholar]
- 29.Stocks BB, Konermann L, BB Time-dependent changes in side-chain solvent accessibility during cytochrome c folding probed by pulsed oxidative labeling and mass spectrometry. J Mol Biol. 2010;398:362. doi: 10.1016/j.jmb.2010.03.015. [DOI] [PubMed] [Google Scholar]
- 30.Chance MR, Sclavi B, Woodson SA, Brenowitz M. Examining the conformational dynamics of macromolecules with time-resolved synchrotron X-ray ‘footprinting’. Structure. 1997;5:865. doi: 10.1016/s0969-2126(97)00241-4. [DOI] [PubMed] [Google Scholar]
- 31.Gupta S, Sullivan M, Toomey J, Kiselar J, Chance MR. The Beamline X28C of the Center for Synchrotron Biosciences: a national resource for biomolecular structure and dynamics experiments using synchrotron footprinting. J Synchrotron Radiat. 2007;14:233. doi: 10.1107/S0909049507013118. [DOI] [PubMed] [Google Scholar]
- 32.Sclavi B, Woodson SA, Sullivan M, Chance MR, Brenowitz M. Following the folding of RNA with time-resolved synchrotron X-ray footprinting. Methods in Enzymology. 1998;295:379. doi: 10.1016/s0076-6879(98)95050-9. [DOI] [PubMed] [Google Scholar]
- 33.Xu G, Chance MR. Radiolytic modification of acidic amino acid residues in peptides: probes for examining protein-protein interactions. Anal Chem. 2004:1213. doi: 10.1021/ac035422g. [DOI] [PubMed] [Google Scholar]
- 34.Kiselar JG, Janmey PA, Almo SC, Chance MR. Structural analysis of gelsolin using synchrotron protein footprinting. Mol Cell Proteomics. 2003;2:1120. doi: 10.1074/mcp.M300068-MCP200. [DOI] [PubMed] [Google Scholar]
- 35.Kaur P, Kiselar JG, Chance MR. Integrated algorithms for high-throughput examination of covalently labeled biomolecules by structural mass spectrometry. Anal Chem. 2009;81:8141. doi: 10.1021/ac9013644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bai Y, Englander SW. Future directions in folding: the multi-state nature of protein structure. Proteins. 1996;24:145. doi: 10.1002/(SICI)1097-0134(199602)24:2<145::AID-PROT1>3.0.CO;2-I. [DOI] [PubMed] [Google Scholar]
- 37.Moffat K. Time-resolved crystallography. Acta Crystallogr A. 1998;54(Pt 6 Pt 1):833. doi: 10.1107/s0108767398010605. [DOI] [PubMed] [Google Scholar]
- 38.Schlichting I, Berendzen J, Phillips GN, Jr, Sweet RM. Crystal structure of photolysed carbonmonoxy-myoglobin. Nature. 1994;371:808. doi: 10.1038/371808a0. [DOI] [PubMed] [Google Scholar]
- 39.Teng TY, Srajer V, Moffat K. Initial trajectory of carbon monoxide after photodissociation from myoglobin at cryogenic temperatures. Biochemistry. 1997;36:12087. doi: 10.1021/bi971140x. [DOI] [PubMed] [Google Scholar]
- 40.Moffat K. Laue diffraction. Methods Enzymol. 1997;277:433. doi: 10.1016/s0076-6879(97)77024-1. [DOI] [PubMed] [Google Scholar]
- 41.Sclavi B, Sullivan M, Chance MR, Brenowitz M, Woodson SA. RNA folding at millisecond intervals by synchrotron hydroxyl radical footprinting. Science. 1998;279:1940. doi: 10.1126/science.279.5358.1940. [DOI] [PubMed] [Google Scholar]
- 42.Sclavi B, Woodson S, Sullivan M, Chance MR, Brenowitz M. Time-resolved synchrotron X-ray “footprinting”, a new approach to the study of nucleic acid structure and function: application to protein-DNA interactions and RNA folding. J Mol Biol. 1997;266:144. doi: 10.1006/jmbi.1996.0775. [DOI] [PubMed] [Google Scholar]
- 43.Brenowitz M, Chance MR, Dhavan G, Takamoto K. Probing the structural dynamics of nucleic acids by quantitative time-resolved and equilibrium hydroxyl radical ‘footprinting’. Current Opinion in Structural Biology. 2002;12:648. doi: 10.1016/s0959-440x(02)00366-4. [DOI] [PubMed] [Google Scholar]
- 44.Adilakshmi T, Bellur DL, Woodson SA. Concurrent nucleation of 16S folding and induced fit in 30S ribosome assembly. Nature. 2008;455:1268. doi: 10.1038/nature07298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Burtnick LD, Robinson RC, Choe S. Structure and function of gelsolin. Results Probl Cell Differ. 2001;32:201. doi: 10.1007/978-3-540-46560-7_14. [DOI] [PubMed] [Google Scholar]
- 46.Pope BJ, Gooch JT, Weeds AG. Probing the effects of calcium on gelsolin. Biochemistry. 1997;36:15848. doi: 10.1021/bi972192p. [DOI] [PubMed] [Google Scholar]
- 47.Ralston CY, Sclavi B, Sullivan M, Deras ML, Woodson SA, Chance MR, Brenowitz M. Time-resolved synchrotron X-ray footprinting and its application to RNA folding. Methods Enzymol. 2000;317:353. doi: 10.1016/s0076-6879(00)17024-7. [DOI] [PubMed] [Google Scholar]
- 48.Ashish, Paine MS, Perryman PB, Yang L, Yin HL, Krueger JK. Global structure changes associated with Ca2+ activation of full-length human plasma gelsolin. J Biol Chem. 2007;282:25884. doi: 10.1074/jbc.M702446200. [DOI] [PubMed] [Google Scholar]
- 49.Xu G, Chance MR. Radiolytic modification and reactivity of amino acid residues serving as structural probes for protein footprinting. Anal Chem. 2005;77:4549. doi: 10.1021/ac050299+. [DOI] [PubMed] [Google Scholar]
- 50.Xu G, Liu R, Zak O, Aisen P, Chance MR. Structural allostery and binding of the transferrin*receptor complex. Mol Cell Proteomics. 2005;4:1959. doi: 10.1074/mcp.M500095-MCP200. [DOI] [PubMed] [Google Scholar]
- 51.Bohon J, Jennings L, Phillips C, Licht S, Chance M. Synchrotron Protein Footprinting Supports Substrate Translocation by ClpA via ATP-Induced Movements of the D2 Loop. Structure. 2008;16:1157. doi: 10.1016/j.str.2008.04.016. [DOI] [PMC free article] [PubMed] [Google Scholar]