Abstract
VEGF was first described as vascular permeability factor, a potent inducer of vascular leakage. Genetic evidence indicates that VEGF-stimulated endothelial proliferation in vitro and angiogenesis in vivo depend on heparan sulfate, but a requirement for heparan sulfate in vascular hyperpermeability has not been explored. Here we show that altering endothelial cell heparan sulfate biosynthesis in vivo decreases hyperpermeability induced by both VEGF165 and VEGF121. Because VEGF121 does not bind heparan sulfate, the requirement for heparan sulfate suggested that it interacted with VEGF receptors rather than the ligand. By applying proximity ligation assays to primary brain endothelial cells, we show a direct interaction in situ between heparan sulfate and the VEGF receptor, VEGFR2. Furthermore, the number of heparan sulfate-VEGFR2 complexes increased in response to both VEGF165 and VEGF121. Genetic or heparin lyase-mediated alteration of endothelial heparan sulfate attenuated phosphorylation of VEGFR2 in response to VEGF165 and VEGF121, suggesting that the functional VEGF receptor complex contains heparan sulfate. Pharmacological blockade of heparan sulfate-protein interactions inhibited hyperpermeability in vivo, suggesting heparan sulfate as a potential target for treating hyperpermeability associated with ischemic disease.
Keywords: Cell Surface Receptor, Endothelium, Growth factors, Heparan Sulfate, Signal Transduction, Tumor, Hyperpermeability, Proteoglycan, VEGF, VEGFR2
Introduction
The endothelium lining blood vessels forms an essential selective barrier that regulates the passage of plasma macromolecules between blood and tissues. The barrier function of the endothelium is well maintained under normal physiological conditions, but ischemia and inflammation can lead to its breakdown and vascular hyperpermeability (1, 2). The primary molecular event that initiates hyperpermeability involves binding of VEGF to VEGF receptor complexes, resulting in VEGF receptor phosphorylation and activation of Src, endothelial NOS (eNOS),2 and other downstream signal transduction proteins (3). Expression of VEGF is rapidly induced under ischemic conditions by activation of the transcription factor, hypoxia-inducible factor 1 α (4, 5). Many types of tumors constitutively express VEGF as well, and the enhanced vascular leakage in tumors increases the risk of metastasis and raises tumor interstitial pressure while reducing the efficacy of chemotherapeutic drug delivery (6–11). Thus, understanding the mechanism by which VEGF interacts with its receptors might provide insights into new therapeutic strategies for the treatment of ischemic disease or cancer.
Three subgroups of cell surface proteins bind VEGF. The first group is composed of the receptor tyrosine kinases, VEGF receptor-1 (VEGFR1), VEGFR2, and VEGFR3. The second group includes neuropilin-1 (Nrp1) and Nrp2, which lack intrinsic kinase activity. The last group consists of one or more heparan sulfate proteoglycans (HSPGs), which consist of a core protein and one or more covalently attached heparan sulfate chains. Various combinations of these receptors form the functional signaling complexes in endothelial cells lining the blood and lymphatic vasculature. In general, VEGFR2 plays the central role in mitogen-activated signaling in blood endothelial cells. Nrp1 and one or more HSPGs are thought to act as co-receptors to promote more efficient binding of the ligand to VEGFR2 and enhance the stability of the receptor complex (3).
All VEGF-A isoforms (VEGF121, VEGF145, VEGF165, VEGF189, VEGF206) except VEGF121 also bind to heparan sulfate or to heparin, a highly sulfated form of heparan sulfate (12, 13). Heparin augments the interaction of VEGF165 with VEGFR2 (14) and binds to VEGFR1, Nrp1, and Nrp2 (15–17). Because heparin/heparan sulfate can interact with both ligands and receptors, it might serve as a template to facilitate approximation of the components, similar to its role in the assembly of FGF-FGF receptor complexes (18, 19) and antithrombin-thrombin complexes (20, 21). Evidence for the co-receptor activity of heparan sulfate proteoglycans in VEGF signaling has been provided in the context of tumor angiogenesis and vasculogenesis (22–24), where heparan sulfate potentiates the duration and magnitude of VEGF signaling and regulates the localization and quantity of VEGF-VEGFR2 complexes in the tumor vasculature. VEGF121 does not bind heparan sulfate except at low pH (24, 25), but heparan sulfate apparently can regulate the interaction of VEGF121 to VEGFR1 (26), possibly by binding independently to the receptor. In contrast, binding of VEGF121 to VEGFR2 does not appear to be modulated by heparan sulfate or heparin (27).
In the present study, we investigated the role of heparan sulfate in the VEGF-induced vascular hyperpermeability, which is mediated by VEGFR2. We found that mice deficient in endothelial heparan sulfate had reduced dermal and tumor vascular permeability, providing the first genetic evidence that heparan sulfate is directly involved in vascular leakage in vivo. Surprisingly, the deficiency of heparan sulfate not only affected hyperpermeability induced by VEGF165 but also equally affected responses mediated by VEGF121. Mechanistically, we identified a direct interaction between heparan sulfate and VEGFR2 in situ, strongly suggesting that heparan sulfate regulates the assembly of functional VEGF receptor complexes. Blockade of heparan sulfate by the heparin-neutralizing agent, surfen (28), significantly attenuated VEGF-induced dermal vascular hyperpermeability, thus providing pharmacological evidence that altering heparan sulfate-VEGF receptor interactions might prove beneficial for treatment of disorders associated with ischemia, such as stroke, myocardial infarction, or cancer.
EXPERIMENTAL PROCEDURES
Reagents
Mouse VEGF164 (catalogue number 450-32) and VEGF121 (catalogue number 100-20A) from PeproTech were used for the Miles assay (32) and in vitro signaling studies. Human VEGF165 (catalogue number 100-44) and VEGF121 (catalogue number 100-116) from Shenandoah Biotechnology were also used in some experiments. Histamine was from Calbiochem, and protamine sulfate and Evan's Blue were from Sigma. Surfen (bis-2-methyl-4-amino-quinolyl-6-carbamide) was obtained from the Open Chemical Repository in the Developmental Therapeutic Program at the NCI, National Institutes of Health (NSC12155). Antibodies to phospho-Tyr-1175-VEGFR2 (catalogue number 2478), VEGFR2 (catalogue number 2497), phospho-Ser-1177-eNOS (catalogue number 9571), and eNOS (catalogue number 9572) were from Cell Signaling. Monoclonal antibody to heparan sulfate (H1890) from U. S. Biological and goat polyclonal antibody to mouse VEGFR2 (catalogue number AF644) from R&D systems were used for proximity ligation assays. Polyclonal antibodies against VEGFR2 from Santa Cruz Biotechnology (catalogue number sc-48161) were used for immunoprecipitation. FITC-conjugated antibodies to endothelial markers CD31 (catalogue number 11-0311), CD34 (catalogue number 11-0341), and phycoerythrin-conjugated antibodies to CD105 (catalogue number 12-1051) and CD166 (catalogue number 12-1661) were from eBioscience. Heparin lyases I, II, and III were generous gifts from Dr. Jian Liu (University of North Carolina at Chapel Hill).
Mutant Mouse Strains
Ndst1f/fTie2Cre+ mice were generated and maintained as described previously (29). Vegfr2+/− mice were purchased from The Jackson Laboratory. Ext1f/+Tie2Cre+ mice were generated by crossing Ext1f/f mice (30) to Tie2Cre transgenic mice (31). All animals were fully backcrossed on C57Bl/6 background.
Miles Assay
For the Miles assay (32), mice (6–10 weeks of age) were injected intravenously with 200 μl of 0.5% Evan's Blue, and 15 min later 200 ng of VEGF in 20 μl of PBS with 0.1% BSA was injected intradermally into shaved dorsal skin. An equal volume of PBS-BSA was injected into a separate dorsal skin patch in each mouse as a negative control. Histamine (1 μg) in PBS-BSA was used as positive control. Mice were sacrificed 40 min after VEGF injection, and a 6-mm diameter skin biopsy was obtained at the injection site using a dermal punch (Acuderm, Inc.). The sample was then extracted with 1 ml of formamide at 55 °C for 24 h. Evan's Blue was quantified by measuring absorbance at 605 nm and converted to mass by using a standard curve.
Protamine inhibition was measured by intradermal injection (20 μl) of a mixture of 12.5 μg of protamine and 200 ng of VEGF in PBS-BSA. When surfen was tested, 18 μl of 60 μm surfen (in PBS-BSA and 0.2% DMSO) was mixed with 2 μl of PBS-BSA containing 200 ng of VEGF. As a positive control, 18 μl of PBS-BSA with 0.2% DMSO was mixed with 2 μl of PBS-BSA containing 200 ng of VEGF.
LLC Tumor Vascular Permeability Model
Lewis lung carcinoma cells (4 × 105 cells, LL/2 line from ATCC) were inoculated subcutaneously into the hindquarters of 12–14-week old mice. Mice were used for experiments when tumors first became visible, 9–14 days after tumor inoculation. Subcutaneous Lewis lung carcinoma cells (LLC) tumors with an average volume of 50–100 mm3 (∼50 mg of wet weight) were processed and stained with anti-CD31 mAb, and microvascular density was measured as described previously (23). These small, early tumors were vascularized to the same extent in all animals regardless of genotype. Mice were injected intravenously with 200 μl of 0.5% Evan's Blue and sacrificed 1 h later. Tumors were carefully removed without any attached skin, weighed, and extracted in 0.5 ml of formamide solution at 55 °C for 24 h. Extracted dye was quantitated as described above in the Miles assay. For the assessment of VEGF expression, tumors were homogenized in 1 ml of radioimmune precipitation buffer (100 mm Tris, 150 mm NaCl, 1% Triton X-100, 0.1% SDS, pH 7.4) containing a protease inhibitor mixture, and 20 μg of tumor homogenate was resolved by SDS-PAGE and immunoblotted with a rabbit polyclonal antibody to VEGF (Abcam, catalogue number ab46154).
Isolation of Primary Mouse Microvascular Endothelial Cells
Murine brain microvascular endothelial cells (mBMEC) were isolated from cerebral cortex essentially as described (33), except that cells were selected with 5 μg/ml puromycin for 4 days in DMEM medium (Invitrogen) supplemented with 20% FBS (Atlanta Biologicals), 50 μg/ml endothelial growth supplement (BTI, Inc.), 50 μg/ml heparin, nonessential amino acids, penicillin, and streptomycin. Cells generally reached confluence in 7 days and were used without passage. Murine lung microvascular cells were isolated and cultured as described previously (23). Disaccharide analysis of heparan sulfate derived from the cells was determined by liquid chromatography/mass spectrometry as described (34).
VEGF Binding Assay
Murine lung endothelial cells were treated with (or without) 10 milliunits/ml heparan lyase I for 20 min at room temperature. Endothelial cells (2 × 105) were incubated with VEGF164 and VEGF121 (40 ng) for 1 h at 4 °C. After rinsing, the cells were incubated with 1 μg/ml biotinylated rabbit anti-hVEGF (catalogue number 500-P10Bt, PeproTech) for 1 h at 4 °C. Cells were stained with streptavidin-phycoerythrin (eBioscience) at a dilution of 1:400 and analyzed by flow cytometry.
Immunoblotting
mBMEC were serum-starved in DMEM for 5 h prior to treatment with VEGF. In some experiments, mBMEC were incubated with a mixture of heparin lyases I (2 milliunits/ml), II (2 milliunits/ml), and III (5 milliunits/ml) in serum-free DMEM at 37 °C for 15 min. Cells were then stimulated with 20 ng/ml mVEGF164 or 50 ng/ml hVEGF121 at 37 °C for 2 or 15 min. Cells were lysed in radioimmune precipitation buffer (Cell Signaling) with a protease and phosphatase inhibitor mixture (Pierce). Samples (10 μg) were resolved on 4–12% Bis-Tris NuPAGE gel and blotted onto polyvinylidene difluoride membrane. After incubation with primary and secondary antibodies, reactive bands were visualized by SuperSignal West Femto (for anti-phospho-Tyr-1175-VEGFR2 and anti-phospho-Ser-1177-eNOS mAbs) or West Pico (for all other antibodies) chemiluminescent substrate (Pierce).
Proximity Ligation Assay
mBMEC were treated with heparan lyases as described above in growth medium lacking serum. Cells were then stimulated with 20 ng/ml mVEGF164 or 50 ng/ml hVEGF121 at 37 °C for 2, 5, and 15 min followed by fixation in 2% paraformaldehyde and blocking with 5% donkey serum in PBS. To study heparan sulfate/VEGFR2 interaction, cells were incubated overnight at 4 °C with 1 μg/ml anti-heparan sulfate (mAb 10E4), a mouse monoclonal antibody that recognizes N-sulfated glucosamine units, and 2 μg/ml affinity-purified goat polyclonal antibody against mouse VEGFR2. Proximity ligation was performed using a Duolink assay kit (Olink) according to the protocol provided by the manufacturer, except that the oligonucleotide-conjugated anti-mouse and anti-goat probes were used at 1:10 dilution. All images were taken with a DeltaVision deconvolution microscopy system with a 20× objective, deconvolved with SoftWoRx software, and analyzed with NIH ImageJ software. Five images were taken from each sample well (0.2 cm2). Each experiment represents a fresh preparation of mBMEC.
Statistics
All data were analyzed by unpaired one-tailed Student's t test to assess statistical significance, except that the paired test was used for protamine and surfen inhibition assays. p values <0.05 were considered statistically significant.
RESULTS
Ndst1-deficient Mice Show Reduced Dermal Vascular Hyperpermeability in Response to VEGF165 and VEGF121
To investigate whether heparan sulfate is involved in VEGF-induced vascular hyperpermeability, we tested the response to VEGF in vivo using mutant mice with altered endothelial heparan sulfate induced by the conditional inactivation of Ndst1 (Ndst1f/fTie2Cre+). Ndst1 encodes the enzyme N-deacetylase-N-sulfotransferase-1, which is partially responsible for the N-deacetylation and subsequent N-sulfation of subsets of N-acetylglucosamine residues in heparan sulfate. Offspring of genotype Ndst1f/fTie2Cre + show ∼50% decrease in overall sulfation of heparan sulfate due to coupling of N-sulfation to other downstream modifications (O-sulfation and uronic acid epimerization) of the chain (23, 29). Because endothelial cells also express a second Ndst isozyme (Ndst2), incomplete reduction of sulfation of endothelial heparan sulfate occurs in Ndst1-deficient mice (29). Compounding these two mutations, however, results in loss of viability (24, 35).
Dermal vascular hyperpermeability was measured by the leakage of Evan's Blue in response to local intradermal injection of VEGF (modified Miles assay) (32). As shown in Fig. 1A, mutant mice showed significantly reduced leakage of Evan's Blue into sites injected with VEGF165 when compared with wild-type littermates (1.5 ± 0.2 in Ndst1f/fTie2Cre+ versus 2.1 ± 0.2 μg in Ndst1f/fTie2Cre−). After subtraction of the basal level of permeability (i.e. when mice were injected intradermally with PBS-BSA instead of VEGF), we calculated that the extent of leakage was reduced 2-fold in the mutant. In contrast, mutant and wild-type mice showed no difference in leakage induced by histamine, an inflammatory factor that does not depend on heparan sulfate (Fig. 1A). We also assayed leakage in Ext1f/+Tie2Cre + mice, which lack one copy of the copolymerase Ext1 responsible for the formation of the backbone of heparan sulfate. This mutation results in shortened heparan sulfate chains (∼50% reduction in length) (36). Altering chain length, however, did not affect VEGF165-induced vascular hyperpermeability (Fig. 1B), emphasizing the importance of sulfation of the heparan sulfate in the hyperpermeability response.
FIGURE 1.
Dermal vascular hyperpermeability induced by VEGF is reduced in Ndst1-deficient mice. A, dermal vascular permeability was measured by Miles assay, in which Evan's Blue was injected intravenously. The amount of dye that leaked into the injection site was measured by absorbance after extraction. Ndst1f/fTie2Cre− (WT) and Ndst1f/fTie2Cre+ (labeled as Ndst1 −/−) mice were injected with 200 ng of VEGF165 (n = 20) and VEGF121 (n = 10). Histamine (1 μg, n = 12) was injected as positive control, and PBS with 0.1% BSA (n = 20) was injected as negative control. Error bars indicate S.E. B, dermal vascular hyperpermeability of Ext1f/+Tie2Cre+ (labeled as Ext1+/−) and Ext1f/+Tie2Cre− (WT) mice was measured as in A. n = 6–9. C, binding of VEGF165 and VEGF121 to primary lung endothelial cells measured by flow cytometry. VEGF binding to untreated cells is shown as a black line, binding to heparin lyase I-treated cells is shown as a gray line, and the control sample, incubated only with biotinylated antibody and streptavidin-phycoerythrin, is shown as the filled gray histogram. % of Max, percentage of maximum.
As a control, we examined the response to VEGF121, a splice isoform that does not bind to heparin or heparan sulfate (12, 13, 24). As shown in Fig. 1C, VEGF121 bound poorly to isolated endothelial cells when compared with VEGF165. (Under these conditions, binding of VEGF165 was primarily mediated by heparan sulfate because treatment of the cells with the degradative enzyme, heparin lyase I, reduced binding by >90%, whereas binding of VEGF121 was not affected.) Surprisingly, Ndst1-deficient mice also showed reduced hyperpermeability in response to VEGF121 (Fig. 1A; 1.3 ± 0.2 in Ndst1f/fTie2Cre+ versus 2.0 ± 0.1 μg in Ndst1f/fTie2Cre−). Thus, the ability of VEGF to induce hyperpermeability in this system depends less on the interaction of heparan sulfate with the ligand than on its interaction with one or more signaling receptors activated by VEGF. Note that the binding of VEGF121 to endothelial cells was about 2.5-fold lower than the binding of VEGF165 to heparin lyase-treated cells. Experiments with mAb 10E4, which reacts with native heparan sulfate, showed complete removal of the epitope by heparin lyase digestion (data not shown). Thus, the difference in binding of the two VEGF isoforms might reflect a lower affinity of VEGF121 for VEGF receptors when compared with VEGF165 as reported previously (37).
Ndst1-deficient Mice Show Reduced Tumor Vascular Permeability
To test whether endothelial heparan sulfate plays a similar role in regulating tumor vascular hyperpermeability, we injected LLC subcutaneously into Ndst1f/fTie2Cre+mice and wild-type littermates (Ndst1f/fTie2Cre−). When tumors were visually discernable, mice were injected with Evan's Blue, and the extent of leakage was measured. As shown in Fig. 2A, less dye extravasation occurred in tumors grown in Ndst1-deficient mice when compared with tumors grown in wild-type mice (21.3 ± 2.1 versus 30.7 ± 2.3 ng/mg of wet tumor mass in mutant and wild-type mice, respectively). In previous studies, we showed decreased LLC tumor growth and vascular density in Ndst1-deficient mice due to deficient sprouting of endothelial cells (23). However, that study examined much larger tumors (volume = 150–400 mm3) taken 15 days after tumor injection. In the experiments reported here, care was taken to ensure that the tumors used had comparable mass (48 ± 4 versus 50 ± 5 mg of wet weight for mutant and wild-type mice, respectively, corresponding to 50–100 mm3, Fig. 2B) and similar tumor microvascular density (Fig. 2C). Western blotting showed comparable levels of VEGF expression in tissue extracts prepared from mutant and wild-type tumors (Fig. 2D). Analysis of tumor sections for infiltration of leukocytes, which might contribute to the tumor vascular hyperpermeability, also showed no differences (Fig. 2E). The results of the skin assays and tumor studies provide the first in vivo evidence that endothelial heparan sulfate regulates VEGF-induced vascular permeability.
FIGURE 2.
Tumor vascular permeability is reduced in Ndst1-deficient mice. A, LLC (4 × 105) were injected into the hindquarters of Ndst1f/fTie2Cre− (WT) and Ndst1f/fTie2Cre+ (labeled as Ndst1 −/−) mice. Mice with comparably sized tumors were injected i.v. with 1 mg of Evan's Blue (EB) and sacrificed 1 h later. The amount of dye extravasation into the tumor was measured spectrophotometrically and normalized to the amount tumor mass that was analyzed (wet weight). B, mass (wet weight) of the tumors used in the experiment shown in panel A. C, multiple sections from tumors (n = 4) were stained with antibody to CD31. The density of tumor microvasculature was estimated by counting the number of CD31+ microvessels per 400× microscopic field. D, VEGF expression in tumors was assessed by Western blotting after separation of 20 μg of tumor lysate by SDS-PAGE. Each lane represents a separate tumor. E, sections from tumors (n = 4) were stained with antibody to CD45. The number of CD45+ cells per 400× microscopic field was determined. Error bars indicate S.E. HPF, high power field.
Loss of Endothelial Heparan Sulfate Attenuates VEGFR2 Phosphorylation in Response to VEGF165 and VEGF121
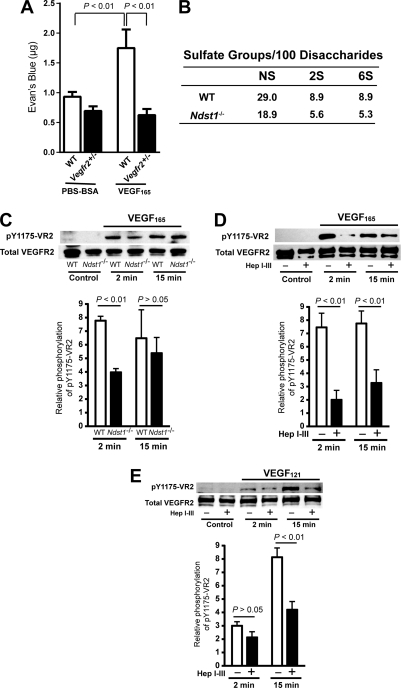
VEGFR2 is the primary VEGF receptor expressed in blood endothelial cells and is thought to be essential for mediating the hyperpermeability response induced by VEGF (38, 39). Consistent with these observations, we found that dermal permeability of Vefgr2+/− mice was abrogated in the Miles assay (Fig. 3A). VEGFR2 expression was readily detectable by Western blotting in isolated neonatal dermal endothelial cells (data not shown). The complete loss of response in the heterozygous mice was surprising, a finding that might suggest that a critical threshold of receptors is needed for hyperpermeability response.
FIGURE 3.
Loss of heparan sulfate attenuates VEGFR2 phosphorylation in response to both VEGF165 and VEGF121. A, dermal vascular hyperpermeability induced by VEGF165 was blocked in Vegfr2+/− mice. Wild-type or Vegfr2+/− mice were injected intradermally with 200 ng of VEGF165 (n = 6) or PBS with 0.1% BSA (n = 6) as negative control. B, chemical analysis of heparan sulfate derived from the mBMEC. Heparan sulfate was isolated by anion-exchange chromatography and digested with heparin lyases I, II, and III, and the resulting disaccharides were analyzed by liquid chromatography/mass spectrometry (34). The number of sulfate groups at each position was determined and expressed per 100 disaccharides. Ndst1−/−, Ndst1f/fTie2Cre+. C, immunoblot analysis of VEGFR2 phosphorylation in mBMEC isolated from mutant and wild-type mice after stimulation with 20 ng/ml VEGF165 for 2 and 15 min (n = 3). D and E, wild-type mBMEC were treated with heparan lyases I, II, and III (Hep I–III) for 15 min and then stimulated with 20 ng/ml VEGF165 (n = 4) or 50 ng/ml VEGF121 (n = 3). Representative immunoblots are shown. The bar graphs represent the relative signal intensity after correction for the background and normalization to the loading control. The average values ± S.E. were determined from the indicated number of replicates. pY1175, phospho-Tyr-1175.
To establish that the alteration in vascular leakage in Ndst1-deficient mice was caused by alterations in VEGFR2 signaling, we isolated primary mBMEC, which, in contrast to dermal endothelial cells, can be readily obtained from adult animals. Cells derived from Ndst1-deficient and wild-type animals showed reactivity for characteristic markers of mBMEC (CD31, CD34, CD105, and CD166, supplemental Fig. S1). The level of N-sulfation, 2-O-sulfation, and 6-O-sulfation of heparan sulfate derived from mutant mBMEC was reduced when compared with the wild type (Fig. 3B), and the extent of reduction was similar to that observed previously in lung microvascular endothelial cells (29).
Treatment of mBMEC with VEGF165 induced rapid and robust phosphorylation of VEGFR2 in wild-type cells, based on reactivity to a phospho-specific antibody that recognizes phospho-Tyr-1175 (Fig. 3C). After normalizing the band intensity to the total amount of VEGFR2 in each extract, we found that phosphorylation of VEGFR2 in Ndst1-deficient cells was reduced ∼50% at 2 min, with attenuation of the effect 15 min after stimulation. Removal of endothelial heparan sulfate by treatment with heparin lyases caused a more substantial reduction of VEGFR2 phosphorylation (73% at 2 min and 58% at 15 min, Fig. 3D). Thus, the incomplete reduction of VEGFR2 phosphorylation in Ndst1 mutant cells was likely due to partial reduction in N-sulfation. VEGF121 activation also displayed a 60% reduction in VEGFR2 phosphorylation in heparin lyase-treated cells when compared with mock-treated controls (Fig. 3E).
eNOS is an important regulator of VEGF-induced hyperpermeability that lies downstream of VEGFR2 and its activation is associated with autophosphorylation of VEGFR2 at Tyr-1175 (40–42). Heparan lyase treatment of mBMEC attenuated eNOS phosphorylation at Ser-1177 by VEGF165 and VEGF121 (Fig. 4). Src kinase is another downstream regulator of VEGF-induced hyperpermeability. Blockade of Src has been shown to reduce tissue injury following cerebral ischemia and myocardial infarction (4, 5). However, attempts to measure Src activation (phosphorylation at Tyr-418) in cultured mBMEC did not meet with success due to the high basal level of phosphorylation in these cells (data not shown).
FIGURE 4.
Loss of heparan sulfate attenuates eNOS phosphorylation in response to both VEGF165 and VEGF121. Immunoblot analysis of eNOS phosphorylation in mBMEC cells after treatment with heparan lyases I, II, and III (Hep I–III) for 15 min and stimulation with 20 ng/ml VEGF165 or 50 ng/ml VEGF121 was performed. Representative immunoblots are shown. The bar graphs represent the normalized relative signal intensity. n = 3. Error bars indicate S.E. pS1177, phospho-Ser-1177.
Pharmacological Inhibition of Heparan Sulfate Inhibits Vascular Hyperpermeability
The above findings suggested that antagonizing the action of endothelial heparan sulfate might provide an effective method to reduce vascular hyperpermeability. To explore this possibility, we first examined the effect of protamine, a highly positively charged protein commonly used to neutralize anticoagulant heparin (43). The addition of protamine to mBMEC caused a dose-dependent suppression of VEGFR2 phosphorylation in response to VEGF165 (Fig. 5A, left). Inhibition was achieved at a concentration as low as 0.1 μm, and with near complete inhibition occurring at 1 μm. Protamine also inhibited VEGF121-induced VEGFR2 phosphorylation, although inhibition required higher concentration (Fig. 5A, right). We also tested the effect of surfen, a recently described low molecular mass (372 Da) heparan sulfate antagonist (average molecular mass of protamine = 5,000 Da) (28). Surfen showed a dose-dependent inhibition of VEGFR2 phosphorylation induced by VEGF165 in mBMEC (Fig. 5B, left), with complete inhibition occurring at 10 μm. Like protamine, surfen was able to inhibit VEGF121-induced VEGFR2 phosphorylation but required higher concentration for unknown reasons (Fig. 5B, right).
FIGURE 5.
Protamine and surfen inhibit VEGFR2 phosphorylation in vitro and dermal vascular permeability in vivo. A, immunoblot analysis of VEGFR2 phosphorylation in mBMEC stimulated with VEGF165 (50 ng/ml) or VEGF121 (100 ng/ml) for 5 min. Cells were treated with protamine at the indicated concentration prior to application of VEGF. The immunoblots are representative of three similar experiments. B, immunoblot analysis of VEGFR2 phosphorylation was performed as in A, except that cells were treated with surfen. C, mice were injected intradermally with 200 ng of VEGF165 alone (filled squares) or with 12.5 μg of protamine (open squares), and the amount of Evan's Blue leakage was measured; n = 9. D, mice were injected intradermally with 200 ng of VEGF165 alone (filled squares) or with 54 μm surfen (open squares); n = 9. E, mice were injected with 200 ng of VEGF121 alone (filled squares) or with 54 μm surfen (open squares); n = 11. Experimental and control treatments of the same mouse in C–E are linked with a line. PBS with 0.1% BSA was used as negative control, which showed an average leakage of 0.9 ± 0.1 μg (C), 0.8 ± 0.1 μg (D), and 0.7 ± 0.1 μg (E). pY1175, phospho-Tyr-1175.
We next tested the ability of protamine and surfen to block VEGF-induced leakage in the skin. Protamine attenuated VEGF165-induced Evan's Blue modestly (22% reduction, Fig. 5C). The lack of potency of protamine in this assay might have been caused by poor tissue penetration as we observed a highly focal inhibition of dye leakage at a site injected with a mixture of protamine and VEGF. Consistent with this idea, intradermal injection of surfen reversed VEGF-induced leakage more effectively than protamine (57%; 1.9 ± 0.2 μg of Evan's Blue with VEGF165 versus 1.2 ± 0.2 μg with VEGF165 plus surfen, Fig. 5D). A similar effect of surfen was also observed when VEGF121 was tested (50% reduction; 1.3 ± 0.1 μg of Evan's Blue versus 1 ± 0.1 μg with surfen (Fig. 5E). Thus, pharmacological inhibition of heparan sulfate-protein interactions in wild-type animals mimics the effect of altering the biosynthesis of heparan sulfate genetically.
VEGF165 and VEGF121 Induce a Transient Increase of Heparan Sulfate-VEGFR2 Complexes
To study the mechanism by which heparan sulfate facilitates VEGF-induced VEGFR2 signaling and hyperpermeability, we examined the formation of heparan sulfate-VEGFR2 complexes in situ by proximity ligation assay in isolated mBMEC using antibodies to heparan sulfate (mAb 10E4) and the extracellular domain of VEGFR2. The specificity of the two antibodies was verified by performing proximity ligation using mBMEC treated with heparan lyases and by using mBMEC isolated from Vegfr2+/− mice (supplemental Fig. S2). Interestingly, heparan sulfate-VEGFR2 complexes were readily detected under basal condition in the absence of VEGF, which may reflect the range of interaction assessed by the assay (up to 30 nm) (44) (Fig. 6A). The apparent number of complexes underwent a rapid increase in response to both VEGF121 and VEGF165 stimulation (70 and 60%, respectively). The response was rapid (2 min was the earliest feasible time point), and then the number of complexes decreased to the basal level by 15 min (Fig. 6B). The formation of new complexes depended on heparan sulfate based on the lack of induction of complexes by either VEGF121 or VEGF165 in mBMEC isolated from Ndst1-deficient mice (Fig. 6C). A 47% decrease in the number of heparan sulfate-VEGFR2 complexes was noted under basal conditions (compare basal conditions for mutant and wild-type cells in Fig. 6C), which was expected because the mutant heparan sulfate contains fewer mAb 10E4 epitopes (N-sulfated glucosamine residues) (29). Although one might argue that the induction of complexes of VEGFR2 with heparan sulfate by VEGF165 reflects a bridging activity of the ligand, VEGF121 does not bind to heparan sulfate yet induces a transient increase in receptor complexes as well (Fig. 1C).
FIGURE 6.
Heparan sulfate-VEGFR2 complexes show a transient increase when stimulated by VEGF165 or VEGF121. A, representative images were obtained by proximity ligation assay. Cells were fixed and incubated with antibodies to heparan sulfate (mAb 10E4) and to the extracellular domain of mouse VEGFR2 (goat IgG) followed by proximity ligation assay reagents. Each red dot indicates an interaction between heparan sulfate and VEGFR2. Nuclei are shown in blue. The Control panel represents cells incubated with 10E4 and nonspecific goat IgG. B, mBMEC were stimulated with 50 ng/ml VEGF121 (striped bars) or 20 ng/ml VEGF165 (white bars) for the indicated time. The data are presented as red dots (Duolink signals)/cell. As a negative control, cells were incubated with mAb 10E4 and nonspecific goat IgG, which gave rise to a background of 3.9 ± 0.3 Duolink signals/cell. The error bar represents S.E., n = 4 separate experiments. C, mBMEC derived from Ndst1f/fTie2Cre− (WT) and Ndst1f/fTie2Cre+ (labeled as Ndst1−/−) mice were stimulated with VEGF as indicated and analyzed by proximity ligation assay. The error bars represent S.E., n = 3 separate experiments. All images were taken with a DeltaVision deconvolution microscope (200× magnification), deconvolved with SoftWoRx software and processed with NIH Image J.
DISCUSSION
In this report, we provide two lines of evidence demonstrating that VEGF-induced vascular hyperpermeability depends on endothelial heparan sulfate. First, undersulfation induced by tissue-specific inactivation of the sulfotransferase, Ndst1, attenuated VEGF-induced dermal and tumor vascular hyperpermeability in vivo (Figs. 1 and 2). Secondly, pharmacological antagonism of heparan sulfate-protein interactions diminished VEGF-induced hyperpermeability in the dermis and blocked VEGFR phosphorylation in isolated endothelial cells (Fig. 5). We also showed that VEGF165, as well as VEGF121, which does not bind to heparan sulfate, induced transient association of VEGFR2 with heparan sulfate (Fig. 6).
All VEGF isoforms bind to heparan sulfate with the exception of VEGF121, a splice variant lacking exons 6 and 7, which encode one or more heparin-binding domains (13). The interaction of VEGF165 with VEGF receptors depends on cell surface heparan sulfate because treatment with heparin lyase severely reduces binding (27, 45). In contrast, VEGF121 is thought to act independently of heparan sulfate because cross-linking of VEGF121 to VEGFR2 and phosphorylation of VEGFR2 were unaffected by heparin lyase or exogenous heparin in human umbilical vein endothelial cells (22, 27). The data presented here challenge this view. Heparin lyase treatment of mBMEC prevented VEGF121-induced VEGFR2 phosphorylation and signaling in vitro (Fig. 3E), and genetic alteration of heparan sulfate in vivo reduced VEGF121-induced vascular permeability (Fig. 1). Possibly, differences in the source of cells used in the earlier studies (human umbilical vein endothelial cells) versus microvascular endothelia (mBMEC) used here might explain the discrepancy. Microvascular endothelial cells may be more susceptible to leak from hydrostatic and/or pathologic hyperpermeability stress.
Proximity ligation assays of primary mBMEC provided direct evidence that both VEGF165 and VEGF121 induced an interaction between heparan sulfate and VEGFR2. VEGF165 might accomplish this interaction by acting as bridging molecule, binding to both heparan sulfate and VEGFR2. Consistent with this idea, the heparin-binding domain in VEGF165 is physically distinct from the region that interacts with the receptor and thus can function independently (46). However, the mechanism by which VEGF121 facilitates heparan sulfate-VEGFR2 interaction must be different due to the lack of a heparin-binding domain in the ligand. The extracellular domain of VEGFR2 can bind to heparin with weak affinity (47), perhaps through the sequence QDRKTKKRHC located between the sixth and seventh Ig-like domains (48). Because all VEGF isoforms are secreted as covalent dimers, with the receptor-binding domains located on opposite sides of the dimer interface, VEGF can facilitate dimerization and autophosphorylation of the receptor (49). Thus, VEGF121 might enhance the interaction of heparan sulfate with VEGFR2 by an increase in avidity due to the receptor dimer binding to an extended oligosaccharide sequence in heparan sulfate (50). VEGFR1 also contains a potential heparin-binding domain in its fourth Ig-like extracellular loop, suggesting that a common mode of ligand-induced interaction with heparan sulfate might exist (15, 51).
Heparan sulfate typically occurs covalently linked to a proteoglycan core protein (52). Cells express several membrane and extracellular HSPGs, including glycosylphosphatidylinositol-anchored proteoglycans (glypicans) and transmembrane proteoglycans (e.g. syndecans). Gengrinovitch et al. (53) have suggested glypican-1 as a VEGF165-binding proteoglycan, but endothelial cells express other membrane proteoglycans as well (54). Studies are underway to identify the functional HSPG that interacts with VEGFR2 and VEGF165 and whether the interaction with heparan sulfate depends on the arrangement of sulfate residues in the chain, e.g., by examining how hyperpermeability is affected in tissue-specific mutants altered in uronyl 2-O-sulfation (55) and glucosaminyl 6-O-sulfation (56). Systemic mutants altered in the expression of specific core proteins are also available, which should aid in the identification of the active proteoglycan(s) in this system (57).
Tissue-damaging edema resulting from ischemia is associated with vascular hyperpermeability induced by expression of VEGF (2, 4). Despite the long term beneficial effects of VEGF in promoting angiogenesis within ischemic tissue, the acute phase edema induced by VEGF aggravates tissue injury (8). Indeed, several studies that employed agents that inhibited VEGF receptor phosphorylation (58) and/or phosphorylation at the downstream steps in the signal transduction cascade (4, 5, 39) improved outcome. The current study suggests that targeting of hyperpermeability through heparan sulfate has the potential to be a novel approach to alleviate tissue injury caused by edema and may synergize with other VEGFR2 pathway inhibitors. Although the induction of edema following endothelial barrier disruption represents an important aspect of disease progression, the inflammatory response contributes significantly to the outcome of ischemic tissue injury as well (59, 60). Previous studies of heparan sulfate-deficient mice have suggested that endothelial heparan sulfate plays a role in neutrophil trafficking and chemokine presentation (29). Thus, antagonizing heparan sulfate interactions provides a generally applicable approach for treating tissue injury and the unintended deleterious sequelae of edema and inflammation.
Supplementary Material
Acknowledgments
We thank Danyin Song for general assistance, Erin Foley for help in isolating primary brain endothelial cells, and Yu Yamaguchi for providing Ext1f/f mice.
This work was supported, in whole or in part, by National Institutes of Health Grant HL57345 (to J. D. E.). This work was also supported by Postdoctoral Fellowship 0825274F from the American Heart Association (to D. X.) as well as support from the Veterans Affairs Career Development program and the American Cancer Society (Grant RSG-08-153-01-CS) (to M. M. F.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1 and S2.
- eNOS
- endothelial nitric-oxide synthase
- HSPG
- heparan sulfate proteoglycan
- VEGFR
- VEGF receptor
- hVEGF
- human VEGF
- mVEGF
- mouse VEGF
- mBMEC
- murine brain microvascular endothelial cells
- LLC
- Lewis lung carcinoma cells
- DMSO
- dimethyl sulfoxide
- Bis-Tris
- 2-(bis(2-hydroxyethyl)amino)-2-(hydroxymethyl)propane-1,3-diol.
REFERENCES
- 1. Mehta D., Malik A. B. (2006) Physiol. Rev. 86, 279–367 [DOI] [PubMed] [Google Scholar]
- 2. Marti H. J., Bernaudin M., Bellail A., Schoch H., Euler M., Petit E., Risau W. (2000) Am. J. Pathol. 156, 965–976 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Olsson A. K., Dimberg A., Kreuger J., Claesson-Welsh L. (2006) Nat. Rev. Mol. Cell Biol. 7, 359–371 [DOI] [PubMed] [Google Scholar]
- 4. Paul R., Zhang Z. G., Eliceiri B. P., Jiang Q., Boccia A. D., Zhang R. L., Chopp M., Cheresh D. A. (2001) Nat. Med. 7, 222–227 [DOI] [PubMed] [Google Scholar]
- 5. Weis S., Shintani S., Weber A., Kirchmair R., Wood M., Cravens A., McSharry H., Iwakura A., Yoon Y. S., Himes N., Burstein D., Doukas J., Soll R., Losordo D., Cheresh D. (2004) J. Clin. Invest. 113, 885–894 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Senger D. R., Van de Water L., Brown L. F., Nagy J. A., Yeo K. T., Yeo T. K., Berse B., Jackman R. W., Dvorak A. M., Dvorak H. F. (1993) Cancer Metastasis Rev. 12, 303–324 [DOI] [PubMed] [Google Scholar]
- 7. Tong R. T., Boucher Y., Kozin S. V., Winkler F., Hicklin D. J., Jain R. K. (2004) Cancer Res. 64, 3731–3736 [DOI] [PubMed] [Google Scholar]
- 8. Weis S. M., Cheresh D. A. (2005) Nature 437, 497–504 [DOI] [PubMed] [Google Scholar]
- 9. Willett C. G., Boucher Y., di Tomaso E., Duda D. G., Munn L. L., Tong R. T., Chung D. C., Sahani D. V., Kalva S. P., Kozin S. V., Mino M., Cohen K. S., Scadden D. T., Hartford A. C., Fischman A. J., Clark J. W., Ryan D. P., Zhu A. X., Blaszkowsky L. S., Chen H. X., Shellito P. C., Lauwers G. Y., Jain R. K. (2004) Nat. Med. 10, 145–147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Criscuoli M. L., Nguyen M., Eliceiri B. P. (2005) Blood 105, 1508–1514 [DOI] [PubMed] [Google Scholar]
- 11. Weis S., Cui J., Barnes L., Cheresh D. (2004) J. Cell Biol. 167, 223–229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Carmeliet P., Ng Y. S., Nuyens D., Theilmeier G., Brusselmans K., Cornelissen I., Ehler E., Kakkar V. V., Stalmans I., Mattot V., Perriard J. C., Dewerchin M., Flameng W., Nagy A., Lupu F., Moons L., Collen D., D'Amore P. A., Shima D. T. (1999) Nat. Med. 5, 495–502 [DOI] [PubMed] [Google Scholar]
- 13. Ruhrberg C. (2003) Bioessays 25, 1052–1060 [DOI] [PubMed] [Google Scholar]
- 14. Tessler S., Rockwell P., Hicklin D., Cohen T., Levi B. Z., Witte L., Lemischka I. R., Neufeld G. (1994) J. Biol. Chem. 269, 12456–12461 [PubMed] [Google Scholar]
- 15. Kendall R. L., Thomas K. A. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 10705–10709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Vander Kooi C. W., Jusino M. A., Perman B., Neau D. B., Bellamy H. D., Leahy D. J. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 6152–6157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Mamluk R., Gechtman Z., Kutcher M. E., Gasiunas N., Gallagher J., Klagsbrun M. (2002) J. Biol. Chem. 277, 24818–24825 [DOI] [PubMed] [Google Scholar]
- 18. Pellegrini L., Burke D. F., von Delft F., Mulloy B., Blundell T. L. (2000) Nature 407, 1029–1034 [DOI] [PubMed] [Google Scholar]
- 19. Schlessinger J., Plotnikov A. N., Ibrahimi O. A., Eliseenkova A. V., Yeh B. K., Yayon A., Linhardt R. J., Mohammadi M. (2000) Mol. Cell 6, 743–750 [DOI] [PubMed] [Google Scholar]
- 20. Dementiev A., Petitou M., Herbert J. M., Gettins P. G. (2004) Nat. Struct. Mol. Biol. 11, 863–867 [DOI] [PubMed] [Google Scholar]
- 21. Li W., Johnson D. J., Esmon C. T., Huntington J. A. (2004) Nat. Struct. Mol. Biol. 11, 857–862 [DOI] [PubMed] [Google Scholar]
- 22. Ashikari-Hada S., Habuchi H., Kariya Y., Kimata K. (2005) J. Biol. Chem. 280, 31508–31515 [DOI] [PubMed] [Google Scholar]
- 23. Fuster M. M., Wang L., Castagnola J., Sikora L., Reddi K., Lee P. H., Radek K. A., Schuksz M., Bishop J. R., Gallo R. L., Sriramarao P., Esko J. D. (2007) J. Cell Biol. 177, 539–549 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Jakobsson L., Kreuger J., Holmborn K., Lundin L., Eriksson I., Kjellén L., Claesson-Welsh L. (2006) Dev. Cell 10, 625–634 [DOI] [PubMed] [Google Scholar]
- 25. Goerges A. L., Nugent M. A. (2003) J. Biol. Chem. 278, 19518–19525 [DOI] [PubMed] [Google Scholar]
- 26. Cohen T., Gitay-Goren H., Sharon R., Shibuya M., Halaban R., Levi B. Z., Neufeld G. (1995) J. Biol. Chem. 270, 11322–11326 [DOI] [PubMed] [Google Scholar]
- 27. Gitay-Goren H., Cohen T., Tessler S., Soker S., Gengrinovitch S., Rockwell P., Klagsbrun M., Levi B. Z., Neufeld G. (1996) J. Biol. Chem. 271, 5519–5523 [DOI] [PubMed] [Google Scholar]
- 28. Schuksz M., Fuster M. M., Brown J. R., Crawford B. E., Ditto D. P., Lawrence R., Glass C. A., Wang L., Tor Y., Esko J. D. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 13075–13080 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Wang L., Fuster M., Sriramarao P., Esko J. D. (2005) Nat. Immunol. 6, 902–910 [DOI] [PubMed] [Google Scholar]
- 30. Inatani M., Irie F., Plump A. S., Tessier-Lavigne M., Yamaguchi Y. (2003) Science 302, 1044–1046 [DOI] [PubMed] [Google Scholar]
- 31. Constien R., Forde A., Liliensiek B., Gröne H. J., Nawroth P., Hämmerling G., Arnold B. (2001) Genesis 30, 36–44 [DOI] [PubMed] [Google Scholar]
- 32. Miles A. A., Miles E. M. (1952) J. Physiol. 118, 228–257 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Perrière N., Demeuse P., Garcia E., Regina A., Debray M., Andreux J. P., Couvreur P., Scherrmann J. M., Temsamani J., Couraud P. O., Deli M. A., Roux F. (2005) J. Neurochem. 93, 279–289 [DOI] [PubMed] [Google Scholar]
- 34. Lawrence R., Olson S. K., Steele R. E., Wang L., Warrior R., Cummings R. D., Esko J. D. (2008) J. Biol. Chem. 283, 33674–33684 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Holmborn K., Ledin J., Smeds E., Eriksson I., Kusche-Gullberg M., Kjellén L. (2004) J. Biol. Chem. 279, 42355–42358 [DOI] [PubMed] [Google Scholar]
- 36. Lin X., Wei G., Shi Z., Dryer L., Esko J. D., Wells D. E., Matzuk M. M. (2000) Dev. Biol. 224, 299–311 [DOI] [PubMed] [Google Scholar]
- 37. Pan Q., Chathery Y., Wu Y., Rathore N., Tong R. K., Peale F., Bagri A., Tessier-Lavigne M., Koch A. W., Watts R. J. (2007) J. Biol. Chem. 282, 24049–24056 [DOI] [PubMed] [Google Scholar]
- 38. Gille H., Kowalski J., Li B., LeCouter J., Moffat B., Zioncheck T. F., Pelletier N., Ferrara N. (2001) J. Biol. Chem. 276, 3222–3230 [DOI] [PubMed] [Google Scholar]
- 39. Scheppke L., Aguilar E., Gariano R. F., Jacobson R., Hood J., Doukas J., Cao J., Noronha G., Yee S., Weis S., Martin M. B., Soll R., Cheresh D. A., Friedlander M. (2008) J. Clin. Invest. 118, 2337–2346 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Fischer S., Clauss M., Wiesnet M., Renz D., Schaper W., Karliczek G. F. (1999) Am. J. Physiol. 276, C812–C820 [DOI] [PubMed] [Google Scholar]
- 41. Fukumura D., Gohongi T., Kadambi A., Izumi Y., Ang J., Yun C. O., Buerk D. G., Huang P. L., Jain R. K. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 2604–2609 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Fulton D., Gratton J. P., McCabe T. J., Fontana J., Fujio Y., Walsh K., Franke T. F., Papapetropoulos A., Sessa W. C. (1999) Nature 399, 597–601 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Park K. W. (2004) Int. Anesthesiol. Clin. 42, 135–145 [DOI] [PubMed] [Google Scholar]
- 44. Söderberg O., Gullberg M., Jarvius M., Ridderstråle K., Leuchowius K. J., Jarvius J., Wester K., Hydbring P., Bahram F., Larsson L. G., Landegren U. (2006) Nat. Methods 3, 995–1000 [DOI] [PubMed] [Google Scholar]
- 45. Gitay-Goren H., Soker S., Vlodavsky I., Neufeld G. (1992) J. Biol. Chem. 267, 6093–6098 [PubMed] [Google Scholar]
- 46. Holmes K., Roberts O. L., Thomas A. M., Cross M. J. (2007) Cell Signal 19, 2003–2012 [DOI] [PubMed] [Google Scholar]
- 47. Chiang M. K., Flanagan J. G. (1995) Growth Factors 12, 1–10 [DOI] [PubMed] [Google Scholar]
- 48. Dougher A. M., Wasserstrom H., Torley L., Shridaran L., Westdock P., Hileman R. E., Fromm J. R., Anderberg R., Lyman S., Linhardt R. J., Kaplan J., Terman B. I. (1997) Growth Factors 14, 257–268 [DOI] [PubMed] [Google Scholar]
- 49. Robinson C. J., Stringer S. E. (2001) J. Cell Sci. 114, 853–865 [DOI] [PubMed] [Google Scholar]
- 50. Robinson C. J., Mulloy B., Gallagher J. T., Stringer S. E. (2006) J. Biol. Chem. 281, 1731–1740 [DOI] [PubMed] [Google Scholar]
- 51. Park M., Lee S. T. (1999) Biochem. Biophys. Res. Commun. 264, 730–734 [DOI] [PubMed] [Google Scholar]
- 52. Bernfield M., Götte M., Park P. W., Reizes O., Fitzgerald M. L., Lincecum J., Zako M. (1999) Annu. Rev. Biochem. 68, 729–777 [DOI] [PubMed] [Google Scholar]
- 53. Gengrinovitch S., Berman B., David G., Witte L., Neufeld G., Ron D. (1999) J. Biol. Chem. 274, 10816–10822 [DOI] [PubMed] [Google Scholar]
- 54. Qiao D., Meyer K., Mundhenke C., Drew S. A., Friedl A. (2003) J. Biol. Chem. 278, 16045–16053 [DOI] [PubMed] [Google Scholar]
- 55. Stanford K. I., Wang L., Castagnola J., Song D., Bishop J. R., Brown J. R., Lawrence R., Bai X., Habuchi H., Tanaka M., Cardoso W. V., Kimata K., Esko J. D. (2010) J. Biol. Chem. 285, 286–294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Izvolsky K. I., Lu J., Martin G., Albrecht K. H., Cardoso W. V. (2008) Genesis 46, 8–18 [DOI] [PubMed] [Google Scholar]
- 57. Bishop J. R., Schuksz M., Esko J. D. (2007) Nature 446, 1030–1037 [DOI] [PubMed] [Google Scholar]
- 58. van Bruggen N., Thibodeaux H., Palmer J. T., Lee W. P., Fu L., Cairns B., Tumas D., Gerlai R., Williams S. P., van Lookeren Campagne M., Ferrara N. (1999) J. Clin. Invest. 104, 1613–1620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Kleinig T. J., Vink R. (2009) Curr. Opin. Neurol. 22, 294–301 [DOI] [PubMed] [Google Scholar]
- 60. Wang Q., Tang X. N., Yenari M. A. (2007) J. Neuroimmunol. 184, 53–68 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.