Abstract
Background
Multi-allelic microsatellite markers have become the markers of choice for the determination of genetic structure in plants. Synteny across cereals has allowed the cross-species and cross-genera transferability of SSR markers, which constitute a valuable and cost-effective tool for the genetic analysis and marker-assisted introgression of wild related species. Hordeum chilense is one of the wild relatives with a high potential for cereal breeding, due to its high crossability (both interspecies and intergenera) and polymorphism for adaptation traits. In order to analyze the genetic structure and ecogeographical adaptation of this wild species, it is necessary to increase the number of polymorphic markers currently available for the species. In this work, the possibility of using syntenic wheat SSRs as a new source of markers for this purpose has been explored.
Results
From the 98 wheat EST-SSR markers tested for transferability and polymorphism in the wild barley genome, 53 primer pairs (54.0%) gave cross-species transferability and 20 primer pairs (20.4%) showed polymorphism. The latter were used for further analysis in the H. chilense germplasm. The H. chilense-Triticum aestivum addition lines were used to test the chromosomal location of the new polymorphic microsatellite markers. The genetic structure and diversity was investigated in a collection of 94 H. chilense accessions, using a set of 49 SSR markers distributed across the seven chromosomes. Microsatellite markers showed a total of 351 alleles over all loci. The number of alleles per locus ranged from two to 27, with a mean of 7.2 alleles per locus and a mean Polymorphic Information Content (PIC) of 0.5.
Conclusions
According to the results, the germplasm can be divided into two groups, with morphological and ecophysiological characteristics being key determinants of the population structure. Geographic and ecological structuring was also revealed in the analyzed germplasm. A significant correlation between geographical and genetic distance was detected in the Central Chilean region for the first time in the species. In addition, significant ecological influence in genetic distance has been detected for one of the population structure groups (group II) in the Central Chilean region. Finally, the association of the SSR markers with ecogeographical variables was investigated and one marker was found significantly associated with precipitation. These findings have a potential application in cereal breeding.
Background
Wild species usually exhibit large genetic variability, which serves as a resource for adaptability to changing environments. On the contrary, cultivated plants are usually more limited in number and display less genetic variability, as a result of the genetic bottlenecks occurring at domestication, translocation and transition from landraces to modern breeding [1]. A consequence of such genetic erosion is genetic uniformity, which may result in the loss of relevant traits, such as resistance to biotic and abiotic stresses.
Thus, wild species related to cultivated crops represent interesting sources of genetic variation, through the introgression of new and better performing alleles. The large genetic variability present in the wild cereals is an invaluable resource for cereal crop improvement. Hordeum chilense Roemer et Schultes, a native South American diploid wild barley (2n = 2x = 14), offers a high potential for cereal breeding among the species of the genus Hordeum, because of its high crossability with other members of the Triticeae tribe and its agronomically interesting characteristics. Crosses between wheat and H. chilense lead to fertile amphiploids named tritordeums. They represent the basic genetic material for introducing genetic variability from H. chilense into wheat breeding programs [2] and for transferring useful genes from H. chilense to wheat. The analysis of the germplasm genetic structure is the basis of management, research and utilization of such germplasm [3], since it is critical to identify and correctly interpret the associations between functional and molecular diversity [4,5]. H. chilense has been found in a wide range of environments, and shows high genetic as well as phenotypic diversity [6]. The analysis of the structure of such high variations is important for breeding purposes, especially to identify genes or genomic regions involved in environmental adaptation and showing high diversity. The genetic structure of populations has been widely documented in most of the studies investigating the diversity of elite crop germplasm, especially in self-pollinating cereals [7-9]. Molecular markers and development of statistical techniques to analyze such data have been the subject of recent intensive studies [10-21], allowing the analysis of the genetic structure in several species and eliminating many of the problems linked with spurious associations. However, although significant efforts to increase the availability of genomic tools, such as molecular markers for cereal crops, have been undertaken in the last years, these developments were not made for wild species with scarce direct agronomic interest, like H. chilense. To solve this problem, the transferability of wheat and barley microsatellite markers (or Simple Sequence Repeats; SSR) to wild related species was evaluated [22,23]. Comparative genomic analyses have indicated a good conservation of coding regions across genomes of different grass species, suggesting that this part of the genome can be used to develop transferable molecular markers [24-27]. The development of high throughput sequencing technologies in recent years has allowed the generation of large Expressed Sequence Tag (EST) datasets in a number of plant species, including cereals, which can be systematically searched for SSR [28]. For example, Yu et al. [29] tested EST-SSR primers originating from hexaploid wheat and rice ESTs on four cereal crops (wheat, rice, barley and maize) and found that 62% of the primer pairs produced Polymerase Chain Reaction (PCR) amplicons on at least two species. Similarly, Zhang et al. [30] reported the transferability of 116 wheat EST-SSRs on 168 accessions, representing 18 grass species. The transferability among the Triticeae ranged from 73.7% for Aegilops longissima to 100% for wheat subspecies (Triticum compactum), but was also good for less related species such as rye (72.8%) or maize (40.4%). In barley, Varshney et al. [31] reported that 78.2% of the SSR markers used (165) showed amplification in wheat, followed by 75.2% in rye and 42.4% in rice. Finally, Gupta et al. [32] reported that 55.12% of wheat EST-SSRs were transferable to barley. Recently, it was shown that the barley EST-SSRs represent a promising source of molecular markers to screen the H. chilense genome [33].
In addition of their high degree of transferability across species, it was recently demonstrated that EST-SSRs are useful for genetic variability studies. For example, Gupta et al. [32] assessed the genetic diversity of EST-SSRs in a collection of 52 elite exotic wheat genotypes. Their results indicate that EST-SSRs are more useful for diversity analyses than genomic microsatellites (g-SSRs). Yang et al. [34] also used EST-SSRs to measure the genetic diversity among three hexaploid wheat populations. They concluded that EST-SSR markers are ideal markers for assessing genetic diversity in wheat. In addition, Balfourier et al. [35] used 38 g-SSRs or 44 EST-SSRs to analyze the structure and the diversity of a collection of 372 wheat varieties and obtained identical results with both types of markers. Recently, Hübner et al. [21] studied the genetic analysis of a new collection of the wild barley H. spontaneum with a set of 42 EST-SSRs, revealing that wild barley populations can be divided into seven major genetically differentiated clusters, as well as the evidence of temperature and precipitation as environmental cues that have shaped the genetic makeup of wild barley. Pan et al. [36] investigated the genetic diversity among 15 wild emmer wheat (T. dicoccum) populations using 25 EST-SSRs, detecting a considerable amount of genetic variation, partly related to ecological factors.
The goals of the current study were to determine: (i) the transferability level of wheat EST-SSR markers and their usefulness for H. chilense; (ii) the genetic diversity of the H. chilense species, using a wide set of available microsatellite markers; (iii) the genetic structure of a natural collection of 94 H. chilense accessions; and (iv) the possible influence of spatial, morphological and environmental factors in the observed structure.
Results
Transferability and polymorphism of wheat EST-SSRs
The transferability of the 98 wheat EST-SSRs was evaluated on a set of eight accessions of H. chilense. A SSR was considered as transferable when the PCR amplification of a band of the expected size and SSR pattern was observed. The primers that showed null alleles in some samples were tested at least twice, in order to avoid false negatives (eg., non-amplification due to PCR failure).
Among the 98 wheat EST-SSRs, 53 (54%) showed cross-species transferability. The percentage of transferred markers was about 50% on each chromosome, thereby indicating a uniform distribution across the genome. Among the 53 transferable SSRs, 20 PCR primer pairs (20.4%) showed polymorphism in the accessions studied and were used for further analysis in the H. chilense germplasm. Between two and 10 alleles per primer pair were observed and only 11.7% of the total alleles had the same size as the allele found in Triticum aestivum cv. 'Chinese Spring', confirming the good potential of H. chilense for wheat and barley genetic diversity improvement. These markers were first assigned to H. chilense chromosomes using the available set of wheat-H. chilense addition lines ([37]). Fifteen of the 20 polymorphic primer pairs were located on the same linkage group as in wheat. One of them was located on chromosome 7 D in wheat (CFE135), while it amplified a product on chromosome 1Hch in H. chilense. Four markers showed the same PCR amplicon sizes in both species, and thus their locations could not be confirmed.
Genetic variability analysis
To perform the genetic variability analysis, 21 barley EST-SSRs and 8 gSSRs previously identified as useful for the genetic analysis in H. chilense ([33,38]) were added to the 20 polymorphic wheat EST-SSRs transferred in this work. A total of 351 alleles were detected over the whole sample of 94 accessions for the 49 SSR loci. Among the 351 alleles, 162 originated from the 21 barley EST-SSRs, 94 from the 20 wheat EST-SSRs and 95 from the eight g-SSRs. The number of alleles per locus ranged from two (for GBM1411, GBM1323, GPW7425, CFE10 and CFE23) to 27 (for GBM1464), with a mean of 7.2 alleles and Polymorphic Information Content (PIC) of 0.5 per locus (Table 1). The highest (0.91) and lowest (0.04) PIC values were observed for GBM1464 and GPW7213, respectively. Generally, wheat EST-SSRs exhibited lower PIC values and fewer number of alleles than barley EST-SSRs. Barley EST-SSRs detected almost twice more alleles and higher PIC values than wheat EST-SSRs.
Table 1.
Summary of the genetic parameters shown by 49 SSR markers used for the characterization of the H. chilense accessions.
| Marker ID | Chromosome location | Allele No. | PIC |
|---|---|---|---|
| GPW7577 | 1B | 5 | 0.140 |
| CFE023 | 1B | 2 | 0.331 |
| GPW7296 | 1D | 3 | 0.189 |
| GBM1029 | 1H | 7 | 0.606 |
| GBM1002 | 1H | 5 | 0.574 |
| GBM1411 | 1H | 2 | 0.369 |
| GBMS14 | 1H | 14 | 0.832 |
| CFE175 | 2A | 3 | 0.177 |
| GPW7438 | 2B | 6 | 0.153 |
| CFE068 | 2D | 7 | 0.679 |
| GBM1047 | 2H | 4 | 0.396 |
| GBM1036 | 2H | 7 | 0.515 |
| GBM1462 | 2H | 6 | 0.673 |
| GBMS233 | 2H | 13 | 0.736 |
| GPW7213 | 3A | 3 | 0.042 |
| GPW7335 | 3B | 8 | 0.581 |
| GWM1047 | 3D | 19 | 0.883 |
| GPW7553 | 3D | 8 | 0.700 |
| GPW7663 | 3D | 6 | 0.144 |
| GBM1069 | 3H | 6 | 0.605 |
| GBMS198 | 3H | 8 | 0.633 |
| CFE188 | 4B | 3 | 0.193 |
| GWM1302 | 4D | 10 | 0.800 |
| GBM1055 | 4H | 8 | 0.627 |
| GBM1067 | 4H | 9 | 0.485 |
| GBM1020 | 4H | 6 | 0.638 |
| GBM1465 | 4H | 5 | 0.222 |
| GBM1323 | 4H | 2 | 0.361 |
| GBM1350 | 4H | 19 | 0.906 |
| GBMS214 | 4H | 3 | 0.402 |
| CFE037 | 5A | 4 | 0.093 |
| GPW7425 | 5B | 2 | 0.078 |
| CFE239 | 5B, 5D | 3 | 0.072 |
| GBM1064 | 5H | 5 | 0.603 |
| GBMS154 | 5H | 10 | 0.674 |
| GPW7455 | 6A | 6 | 0.511 |
| CFE002 | 6A, 6B, 6D | 8 | 0.497 |
| CFE080 | 6A, 6B, 6D | 2 | 0.172 |
| GWM1103 | 6D | 18 | 0.873 |
| GBM1008 | 6H | 13 | 0.744 |
| GBM1076 | 6H | 6 | 0.701 |
| GBM1400 | 6H | 3 | 0.557 |
| CFE010 | 7A, 7B, 7D | 2 | 0.198 |
| CFE100 | 7A, 7B, 7D | 10 | 0.820 |
| CFE135 | 7D | 3 | 0.390 |
| GBM1060 | 7H | 13 | 0.838 |
| GBM1058 | 7H | 3 | 0.217 |
| GBM1464 | 7H | 27 | 0.913 |
| GBM1432 | 7H | 6 | 0.724 |
| Mean | 7.163 | 0.4951 | |
Additionally, several rare or specific alleles were found among the analyzed germplasm. Out of the 351 alleles, 184 were found at a frequency lower than 5%, and were therefore considered as rare. Clustering showed that the germplasm can be separated into two groups (see below) with a total of 66 specific alleles found in group I (39 being rare) and 134 specific alleles found in group II (82 being rare). The gene diversity, polymorphic information content (PIC), and the number of specific alleles was lower in group I than in group II (Table 2).
Table 2.
Number of accessions and specific alleles, gene diversity and its standard deviation (SD), calculated for 49 microsatellite loci typed for the H. chilense germplasm, per genetic structure group.
| Structure groups | No. of accessions | No. of specific alleles | Gene diversity | Gene diversity SD |
|---|---|---|---|---|
| Group I | 41 | 66 | 0.3811 | 0.0428 |
| Group II | 53 | 134 | 0.4874 | 0.0387 |
Analysis of genetic structure and differentiation among inferred groups
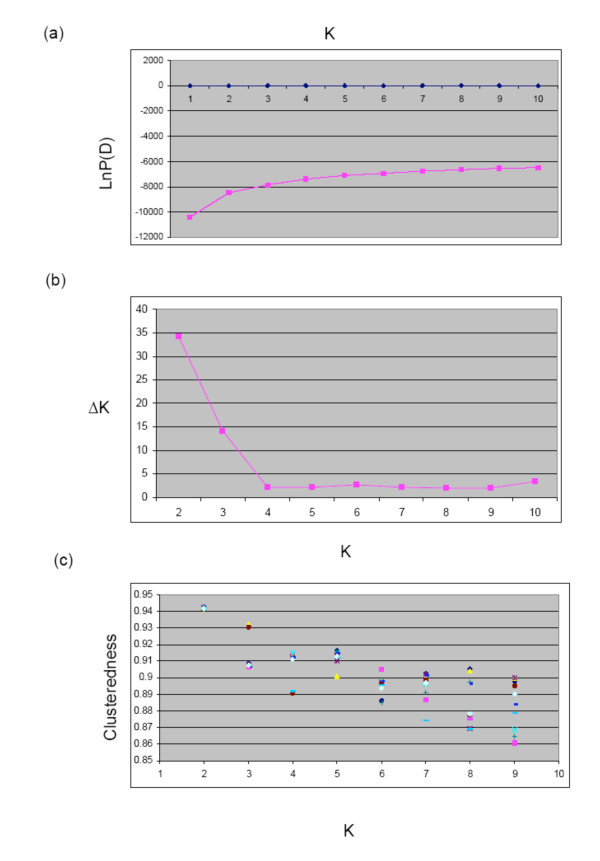
The genotyping data obtained from the 49 SSRs were used to analyze the genetic structure of the germplasm, using the Bayesian clustering model implemented in the Structure software. The natural logarithm of the probability of the data, proportional to the posterior probability of K, showed no clear peak between 1 and 10 for K, and therefore it was difficult to determine the true number of populations (K) (Figure 1a). We applied the rate of change in the Napierian logarithm probability relative to standard deviation (ΔK), as described by Evanno et al. [39]. The results showed the highest peak at 2 (Figure 1b), which was confirmed by the clusteredness index [40], showing the highest median level at K = 2 (Figure 1c). Moreover, using the Geneland software, we observed that the posterior distributions of the estimated number of populations (K) across 10 replicates displayed a clear mode at K = 2 in all of them (additional file 1), corroborating again the previous data. Thus, these results suggest that the analyzed H. chilense germplasm can be divided into two genetically distinct groups.
Figure 1.
Estimation of the most probable number of clusters (K), based on 20 independent runs and K ranging from 1 to 10. (a) Evolution of the natural logarithm probability of the data against K; (b) magnitude of ΔK for each K value; and (c) clusteredness analysis of the H. chilense accessions.
To find the key determinants to the inferred structure of these two groups, we investigated the geographical proximity, as well as the morphological, agronomical and ecological characteristics in the accessions belonging to each group. Figure 2a shows the distribution of each accession into the two populations (identified by the Structure software, and designated as group I and group II, hereafter) and the three clusters (according to morphological and agronomical data as classified by Vaz Patto et al. [6]). Geographic origins divided in 8 zones (Figure 2b) and ecological regions (Figure 2c) according to the classification established by DiCastri and Hajek [41] at K = 2 populations, are also shown.
Figure 2.
Membership of H. chilense accessions. Model-based populations when K = 2, and in the predefined groups according to: (a) agro-morphological data; (b) geographic origins; and (c) ecological regions.
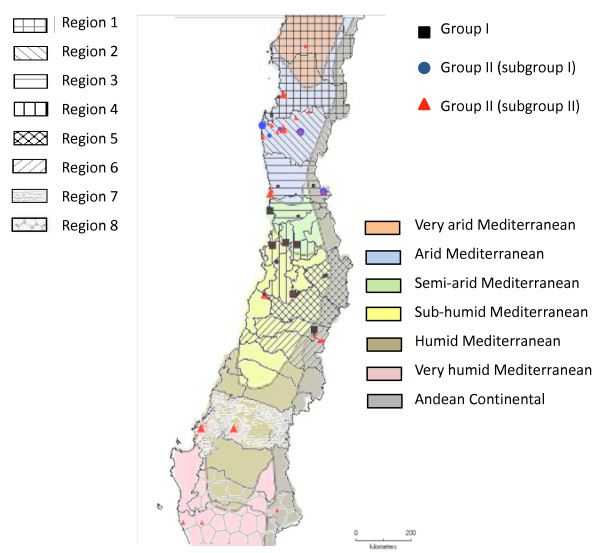
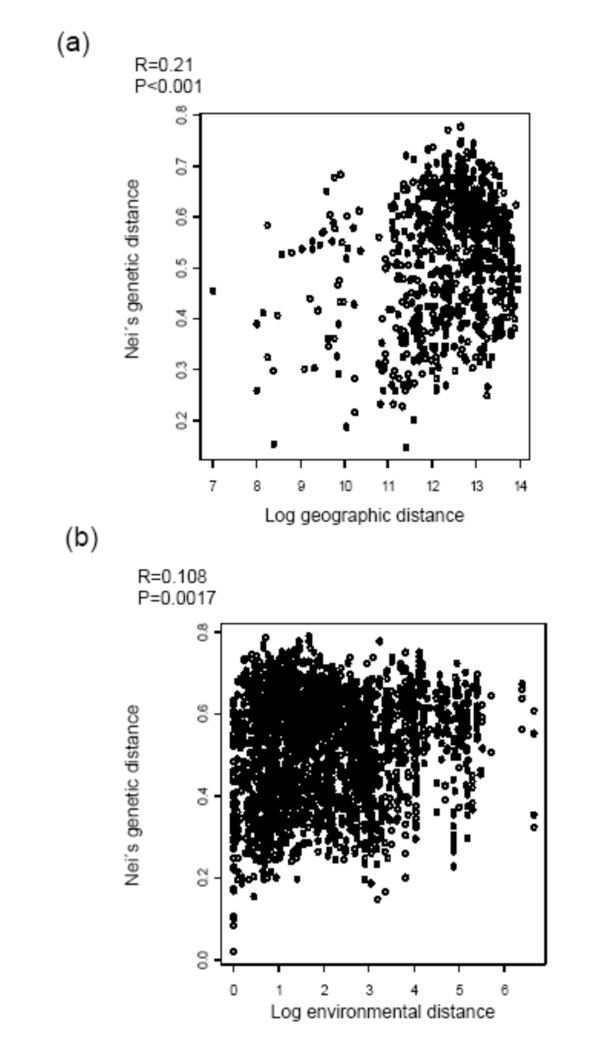
The genetic structure analysis, according to the geographical origins of the accessions (Figure 2b) revealed that, in some regions, the accessions were grouped according to the geographical location. The zones 2, 4, 5 and 7 showed a uniform structure, while the rest of the zones were more or less admixed. Uniform structure was considered when more than 80% of the accessions in one group had more than 80% of membership in this group. The geographical origin of the accessions and their membership to the inferred groups are represented in Figure 3. We calculated the correlation coefficient (r) between the geographic and the genetic distance matrices using the Mantel test [42]. We observed an r value of 0.21 (Figure 4a), revealing a low but significant correlation (p < 0.001). The correlation was then analyzed separately for both inferred groups (I and II), and the results showed that group I had a uniform distribution in the Center of the country, yet group II expanded across the North, Center and South of the country. Therefore, group II was analyzed separately for the North, the Center and the South regions. We found a significant correlation between geographical proximity and genetic distance for group I and for group II Central. Thus, our results demonstrate a geographical influence in population structure in the Central Chilean region for both structure groups (additional file 2). Such influence was not detected for the group II accessions, either at the North or South regions (the latter is probably due to the scarcity of accessions).
Figure 3.
Geographical and environmental distribution of the H. chilense accessions, according to their classification in two populations(group I and group II). Symbol size represents more than one sample at the same location.
Figure 4.
Mantel test showing the relationship of genetic, geographic and environmental distances. (4a) Relationship between Nei's genetic distance and Napierian logarithm of geographic distances; and (4b) relationship between Nei's genetic distance and environmental distance.
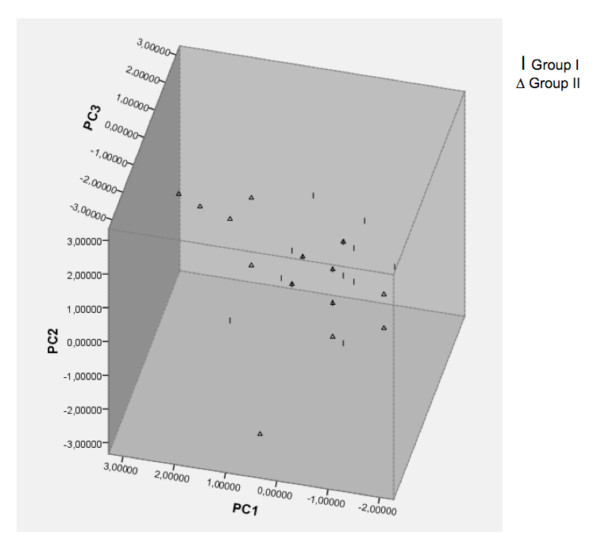
To investigate the impact of ecological characteristics in the inferred structure (K = 2), the accessions were grouped according to bioclimatic parameters. The results revealed admixed populations, except for some provinces that showed a more uniform structure (like humid Mediterranean, very arid Mediterranean, very humid Mediterranean, and sub-humid Mediterranean; see Figure 2c). Comparative analyses between ecogeographical data (latitude, longitude, altitude, mean temperature of coldest and warmest month, and rainfall of driest and wettest month) and genetic data (Nei's genetic distance matrix) revealed a weak but significant correlation (r = 0.108; p = 0.0017) (Figure 4b). When analyzing the influence of the same ecogeographical data in both structured groups separately, the group I did not show any ecogeographical influence, whereas the group II Central exhibited a significant ecogeographical influence (additional file 3). In addition, the ecogeographical data were used to separate the H. chilense accessions through Principal Component (PC) analysis. The first component (PC1, Figure 5 and Table 3) was explained by variation in latitude and rainfall, accounting for 42.9% of the variation. The second component (PC2) accounted for 35.4% of the variation, being explained by variation in temperature and longitude. PC3 accounted for 12.3% of the variation, and was explained by the variation in elevation.
Figure 5.
Principal Component Analysis based on ecogeographic values. The samples shapes are displayed according to the groups defined by the Structure software.
Table 3.
Principal component analysis based on ecogeographical data of H. chilense accessions.
| Variable | Principal Component | ||
|---|---|---|---|
| PC1 | PC2 | PC3 | |
| Elevation | 0.126 | -0.586 | 0.755 |
| Warmest month | -0.445 | 0.779 | 0.389 |
| Coldest month | -0.568 | 0.763 | 0.253 |
| Latitude | 0.906 | 0.327 | 0.171 |
| Longitude | 0.213 | 0.865 | -0.147 |
| Wettest month | 0.896 | 0.181 | 0.148 |
| Driest month | 0.893 | 0.236 | -0.038 |
| Variance (%) | 42.9 | 35.4 | 12.3 |
The unrooted Neighbor-Joining (NJ) tree (Figure 6) distinguished two groups of accessions, corresponding to the structure grouping. Neither geographical nor ecological evidence was detected in the grouping. Results of distance and Bayesian cluster analyses evidenced the presence of a structured genetic diversity among the groups. The Analysis of MOlecular VAriance (AMOVA) of the two inferred groups by the Structure software revealed a 33.16% of the genetic variation among groups, with the remaining 66.84% due to differences within groups. The genetic variances within and among groups were significant (FST = 0.331, p < 0.001; being FST the variance among subpopulations relative to the total variance), supporting the presence of a genetic structure.
Figure 6.
Unrooted neighbor-joining tree based on the Nei's genetic distances obtained using 49 microsatellites. The sample shapes indicate the two groups, inferred by the Structure software.
Association between markers and ecogeographical factors
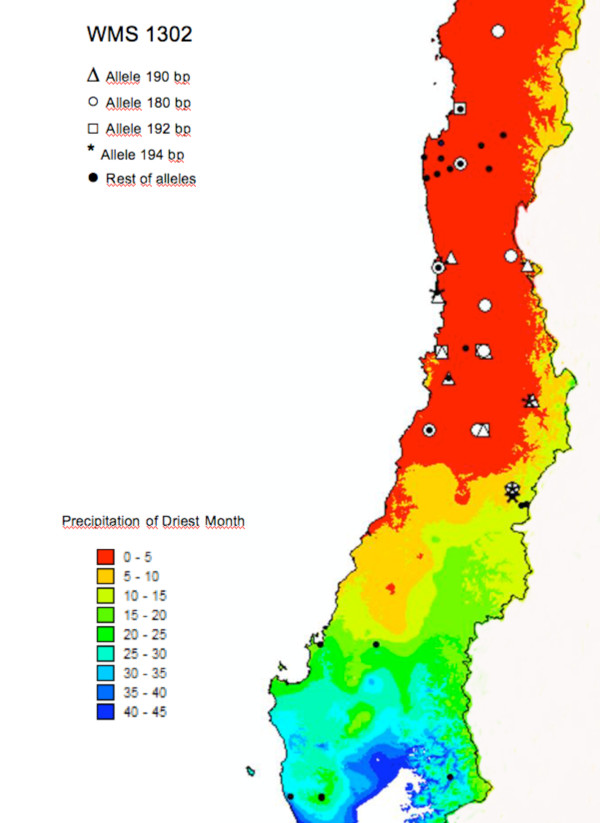
We identified 12 outlier loci that detected high or low variability with respect to the expected neutrality. Among those, 11 loci (GBM1350, GBM1064, GBM1008, GBM1060, GBM1464, GWM1047, GBMS14, GWM1302, CFE135, GPW7335, GPW7663 and GPW7577) are candidates for balanced selection, while the locus CFE135 is a candidate for being subjected to positive selection. The 12 markers were assayed for their association with ecogeographical data. The marker GWM1302 exhibited four different alleles (188, 190, 192 and 194 bp) among 10 alleles that were associated with low rainfall (Figure 7). The other markers did not show any significant association with any of the ecogeographical traits.
Figure 7.
Distribution of alleles associated with rainfall. Distribution of alleles 188 (○), 190 (Δ), 192 (□), 194 (*) of the GWM1302 locus associated with rainfall. Non-associated alleles are also shown (●).
Discussion
Transferability and polymorphism of wheat EST-SSR
The transferability of EST-SSRs across related species has been demonstrated in several species and genera [24,26,29-32,43-45]. Recently, we reported on the usefulness of barley EST-SSRs for genetic analysis in H. chilense [33]. In the present work, we show that more than half (54%) of the assayed wheat EST-SSRs can be transferred to H. chilense, which is lower than the transferability of the barley EST-SSRs (66%). This is likely due to the fact that wheat is evolutively more distant from H. chilense than barley. This result is relatively consistent with the findings of Zhang et al. [30] and Gupta et al. [46], who reported higher transferability of wheat EST-SSRs to barley than to more evolutively distant species, such as maize, rice, sorghum, lolium (ryegrass), oats and purple false brome (Brachypodium).
Despite of the evolutive distance, the overall conservation of the wheat EST-SSRs linkage groups in H. chilense was high, indicating a good level of synteny between these two species. Only one disagreement was observed, that may be due to chromosomal rearrangements, which may be frequent during speciation [47]. Thus, the wheat EST-SSRs markers transferred to H. chilense have an added value as intergeneric syntenic markers, in addition to their direct application to analyze gene diversity. Since numerous additional wheat EST-SSRs are available in the public databases (eg., GrainGenes http://wheat.pw.usda.gov/GG2/index.shtml; [48]), this number could be further increased.
Among the 53 wheat EST-SSRs showing good transferability to H. chilense, about 40% exhibited polymorphism between at least two accessions, which represents 20% of the initial set of wheat EST-SSRs. This was lower compared to what was previously observed between wheat and barley (60% of the transferable EST-SSRs; [30]). This is also lower compared to what was found using barley EST-SSRs (36%), because barley is more closely related to H. chilense than wheat. Thus, the markers transferred from wheat to H. chilense are only those that are more conserved, and therefore likely to show lower polymorphism. Similarly, the number of alleles detected in H. chilense with wheat EST-SSRs was lower compared to the number obtained when using barley EST-SSRs (94 vs 162, respectively). On the other hand, due to their wheat origin, they are not suitable for direct in-tube detection methods [49]. Nevertheless these drawbacks are largely overcome by the fact that they provide valuable anchors for synteny inference [50]. Therefore, we conclude that both wheat and barley represent a good source of markers for genetic diversity and structure studies of H. chilense germplasm collections. In addition, they represent an invaluable tool for the introgression of H. chilense alleles to other cereal species.
Genetic variability analysis
The microsatellite markers revealed a total of 351 alleles across all the 49 loci. The high level of genetic diversity detected could be an adaptive strategy in response to a heterogeneous environment. According to the marker's origin, the SSRs identified in EST databases detected a lower number of alleles and PIC than those obtained through general genomic libraries, including non-transcribed regions. Such is an expected result, due to the more conserved nature of EST-SSRs. The number of alleles detected and PIC found when using EST-SSRs from wheat was lower than with barley EST-SSR and non-transcribed genomic microsatellites, as expected, due to the evolutionary distance between species. Despite these facts, our results show a sufficient level of variation when using EST-SSRs (both from barley and wheat origins) to carry out genetic structure and future association mapping analysis. Therefore, this is one more case where the EST-SSR markers provide an opportunity to examine the functional diversity of germplasm collections, as shown by Eujayl et al. 2002 [51].
The group I identified by Structure was fixed to one allele in eight loci. Seven of these loci could be assumed as being non-neutral, due to their origin from EST databases. The higher number of fixed alleles in non-neutral SSRs could be explained by temporal variation of external factors, generating selection pressures that maintain variation within populations [52]. This may also be due to the fact that not all nucleotide bases on a transcribed DNA are of selective nature. In fact, the third base of the mRNA triplets is less specific (wobble hypothesis, [53]). Furthermore, some amino acids may share similar chemical properties (eg., nonpolar or polar, including acidic or basic), thus being less prone to generate a phenotypic change. A high number of specific alleles were identified, which could be an indication of the relatively high rate of mutation at SSR loci [54], or to a germplasm with a rich genetic diversity and a divergent population structure. The gene diversity is significantly lower in group I than in group II (Table 2). Therefore, the group II is genetically more diverse, corresponding to accessions present in a wider ecogeographical range.
The Spearman correlation showed that genetic diversity is influenced negatively by altitude and positively by temperature. Accessions in group II showed a higher number of alleles. Besides, they were found mainly in places with low altitude and higher temperature than accessions in group I. Therefore, accessions stressed by cold showed less genetic variation.
In this genetic variability analysis of H. chilense germplasm using wheat and barley gSSRs and EST-SSRs, we have defined two main germplasm groups (group I and group II). A previous analysis based on AFLP markers and a Principal-CoOrdinate analysis ([6]) divided the same germplasm into three clusters. The group II defined in this work contains two subgroups, corresponding to the clusters II and III described by Vaz Patto et al. ([6]).
Genetic structure in H. chilense
The analysis of the genetic structure using both Bayesian approaches (Structure and Geneland software) and genetic distance approaches (cluster analysis) of a set of 94 H. chilense accessions, using 49 microsatellite markers, revealed two genetically differentiated groups.
The 'admixture model' implemented by Structure gave a better fit to the species ecophysiological clusters, as defined by Vaz Patto et al. [6], and it was chosen for further association analysis. Therefore, the morphological and agronomic characteristics, which determined the ecophysiological clusters, were key determinants of the population structure of the H. chilense germplasm. Thus, the two inferred groups are mainly in accordance with the agro-morphological clusters described by Vaz Patto et al. [6], as the group I corresponds to cluster I, while the group II (with the exception of three lines) includes clusters II and III (see Figure 2a). According to the geographical origins and the ecological distribution, the inferred genetic structure showed both uniform and admixed populations (Figure 2b and 2c). The accessions included in the geographical zones 1 and 2 showed a uniform genetic structure. They were found in the driest places of these zones, corresponding with Mediterranean arid environments. The accessions grouped in the geographical regions 7 and 8 revealed also a uniform genetic structure, and were found in the wettest places, corresponding to and Mediterranean humid and very humid environments. This points to an influence of the rainfall in shaping the population structure.
Several studies have been carried out to detect the population structure in barley. In most of them, the key factors affecting the genetic structure were growth habit or spike morphology and geographic origin [15,16,19,20,55-57]. In our study, the morphological and agronomic characteristics also have appeared as the main factors to affect the population structure, although we have also shown that geographic locations and ecological patterns of distribution also affect this structure. The analysis of the total germplasm set revealed low but significant associations between geographical and genetic distance, as well as between ecological and genetic distance. By analyzing separately the three main geographical regions of provenance of the species (North, Center and South), a more significant correlation between geographical and genetic distance was detected for the accessions from Central Chile, but no association was found either in Northern or in Southern Chile accessions. This is in agreement with the basal phylogenetic position of H. chilense in South America, established initially in Central Chile from a long-distance continental dispersal from North America, followed by two independent dispersals to the North and to the South [58]. On the other hand, by analyzing both population structure groups separately, a higher correlation between ecological and genetic distance could be detected for group II accessions, but no correlation was found for group I accessions. In fact, group II accessions have shown a better ability to colonize the North and the South Chilean regions (see Figure 3). Thus, the adaptation to geographic and ecological factors may be one of the causes of the genetic structure in the studied germplasm. Thus, the results of our work illustrate the interest to further investigate how morphological characteristics and ecophysiological traits affect the species selection and the population structure. Moreover, the presence of a high level of structure within the H. chilense germplasm should be considered in future association mapping studies. The AMOVA detected higher differences among individuals within-population structure groups than among groups, which is consistent with findings from other studies, indicating that considerable genetic diversity is partitioned within rather than between wild barley populations [59-63]. The proportion of genetic variation within population groups reflected high levels of genetic diversity.
Association of markers with ecogeographical factors
The loci that show unusually low or high levels of genetic differentiation are often assumed to be under natural selection [64]. The accessions that show association of alleles of the locus GWM1302 with low precipitation belong mainly to the group I, and were collected from dry places, thereby suggesting that this environmental factor is involved in a local adaptation after colonization.
Significant correlations between microsatellite markers and ecogeographical factors have been observed in several studies in wild wheat [65] and in wild barley [66], suggesting the impact of natural selection on these markers by creating regional divergence.
Genetic clustering in a principal component analysis revealed that the combination of geographic and ecological data, such as the latitude with rainfall, as the main contributor to the genetic structure of the H. chilense germplasm. The second principal component explained by longitude and temperature significantly contributed to the separation of the two groups. Hübner et al. [21] studied the population structure in Hordeum spontaneum and found a strong correlation of population structure with temperature and precipitation. In our study, the genetic structure of the analyzed germplasm showed a correlation with morphological and ecophysiological characteristics, influenced also to a minor extent by geographic and ecological factors.
Conclusions
Our study shows the utility of barley EST-SSR for the genetic analysis of H. chilense, with a remarkably high level of polymorphism within this species, despite of the evolutionary distance between the wheat and barley genera. The current set of SSR markers available for Hordeum chilense, which includes wheat and barley gSSRs and EST-SSRs, is useful to analyze the genetic structure and ecogeographical adaptation of H. chilense wild barley populations. Both wheat and barley represent a good source of markers for genetic diversity and structure studies of H. chilense germplasm collections. In addition, they represent an invaluable tool for the introgression of H. chilense alleles into other cereal species, and are useful as anchors for the syntenic maps. The analyzed germplasm can be divided into two groups, with morphological and ecophysiological characteristics being key determinants of the population structure. Geographic and ecological structuring was also revealed in the analyzed germplasm. A significant correlation between geographical and genetic distance was detected in the Central Chilean region for the first time in the species. In addition, significant ecological influence in genetic distance has been detected for one of the population structure groups (group II) in the Central Chilean region. Finally, one marker was found significantly associated with precipitation. These findings have a potential application in cereal breeding.
Methods
Plant material and DNA sampling
The DNA collection, isolated by Castillo et al. [38], consisted of 94 samples of H. chilense collected during several expeditions to Chile [67,68]. This germplasm is maintained at the Germplasm Bank of the Institute for Sustainable Agriculture (Prof. A. Martín, IAS, CSIC, Cordoba, Spain). Eight samples of H. chilense DNA were used for selecting transferable and polymorphic microsatellite markers, using Triticum aestivum cv. 'Chinese Spring' DNA as control, to corroborate the pattern of the microsatellites and the fragment sizes. In addition, a DNA set of T. aestivum-H. chilense addition lines developed by Miller [37] were used to determine the chromosomal locations of the polymorphic SSR. The addition lines for chromosomes 4Hch, 5Hch, 6Hch and 7Hch were disomic, whereas the addition lines for chromosomes 1HchS, 2Hch alpha arm, 5Hch, 6HchS arm and 7Hch alpha and beta arm were ditelosomic.
Amplification and transferability of wheat EST-SSRs
A selection of 98 SSRs derived from wheat ESTs [69,70] uniformly distributed across wheat chromosomes were initially screened for their transferability and polymorphism in H. chilense genome, and 20 were selected for germplasm analysis. A set of 21 barley EST-SSRs and eight wheat and barley g-SSRs, previously identified as useful for the genetic analysis in H. chilense [33,68] was added. In total, 49 polymorphic microsatellite markers were applied for fingerprinting H. chilense accessions. The polymerase chain reaction (PCR) amplification and fragment analysis were as previously described [43,71,72], or PCR was carried out using the M13 protocol as described in Nicot et al. [69], with an annealing temperature of 60°C for 30 cycles (30 s at 94°C, 30 s at 60°C, and 30 s at 72°C) and 56°C annealing for eight cycles. Amplification products were visualized using an ABI PRISM 3700 Genetic Analyzer from Life Technologies (Carlsbad, CA, USA). The fragment sizes were calculated using GeneMapper software from the same manufacturer.
Ecogeographical data
The geographic data (altitude, latitude and longitude) of 76 accessions were available [6], and thus they were used to project the data using the DIVA-GIS software http://www.diva-gis.org;. The geographic location of the study area was between 28°15' and 38°42' South latitude and between 70°18' and 73°24' West longitude. The altitude on the sites varied within a wide range, from sea level to high mountains (> 2000 m). Since only one accession is available in some provinces, the accessions were grouped in eight zones along Chile, from North to South, including in some cases various close provinces with similar ecological characteristics. The ecological data like rainfall of wettest and driest month and mean temperature of the warmest and coldest month were obtained for each site using DIVA-GIS. The ecological regions were described following the bioclimatic classification of DiCastri and Hajek [41].
Statistical analysis
The summary statistics including the number of alleles per locus, polymorphism information content (PIC) values and gene diversity were determined using the application PowerMarker version 3.25 [73]. The unrooted neighbor-joining (NJ) tree was constructed using the Nei's index distance [74]. One thousand matrices were obtained by bootstrapping, and the consensus tree was constructed with the program Consense of the Phylip package (version 3.66) [75]. The dendrogram was visualized using the TreeView 1.6.6 software [76]. We performed a Mantel test correlation [42] between Nei's genetic distance and the natural (Napierian) logarithm of the geographic distances, using the library ade4 in the R package (version 2.10.1; R Development Core Team 2008) [78].
A Bayesian model-based analysis for inference of population structure was performed using the program Structure (version 2.2) [78] to estimate the number of groups (K) represented by all sampled individuals and the individual admixture proportions. The Structure software assumes a model in which there are K populations (where K may be unknown), each being characterized by a set of allele frequencies at each locus. Individuals in the sample are probabilistically assigned to a particular population, or associated to two or more populations (if their genotypes indicate that they are admixed). The number of clusters was inferred using 20 independent runs with 100,000 burn-ins and 100,000 iterations after burn-ins, following the admixture ancestry model and correlated allele frequencies, with K ranging from 1 to 10. We have followed the procedure by Evanno et al. [39] to better detect the real number of clusters determined by Structure. Also, the clusteredness index [40] was calculated, which is based on the Q matrix of Structure, being 1 when individuals are assigned completely to a single cluster and 0 when they are equally assigned to all clusters. The individuals can have membership coefficients summing 1 across clusters.
The Distruct 1.1 software [40] was used to graphically represent the estimated population structure, according to geographic proximity, ecological region and agronomical data. Each individual was represented by a thick line, which was partitioned into K colored segments, representing the individual's estimated membership fractions in K clusters.
The genetic structure of the population was also inferred by the Geneland package [79], implemented in the R software. The Geneland software uses geographic coordinates and does not assume admixture, whereas the Structure software does not use geographic coordinates and does assume admixture. We carried out five independent runs using independent allele frequencies with 100,000 iterations, from which each 100th observation was sampled from the Markov chain, with minimum and maximum K being 1 to 10. The run with the highest likelihood was post-processed to obtain the posterior mode of population membership. The genetic differentiation among genetic groups inferred by Structure was estimated by hierarchical analysis of molecular variance (AMOVA), implemented in the Arlequin 3.0 software [80].
We used the Lositan software [81] to identify outlier loci that had excessively high or low Fst compared to neutral expectations. The basic rationale is that (i) loci influenced by directional (also called adaptive or positive) selection will show a larger genetic differentiation than neutral loci; and that (ii) loci that have been subject to balancing (also called negative or purifying) selection will show a lower genetic differentiation. Thus, the methods generally consist of identifying loci that present Fst coefficients that are "significantly" different from those expected under neutral theory (they are called outlier loci). To avoid false positives caused by population structure, the Fst was calculated for the inferred structure groups (the significance level chosen was 0.001, which corresponds to a statistical significance level of 0.05), applying a Bonferroni standard correction. The association of alleles of outlier loci with ecogeographical factors was assayed by linear regression analyses, using the SPSS package version 17.0.0 from SPSS (Chicago, IL, USA). Alleles with frequencies below 5% were excluded. Alleles of each locus were introduced as dependent variables in the model and ecogeographical factors were the independent variables. Significance was calculated for the model, which included only one allele, with the significance threshold set at 0.05, using a Bonferroni correction, as already mentioned.
Values of environmental variables were first standardized and the Euclidean distance between the samples was computed using SPSS. The correlation between genetic distance and environmental distance in the collection was calculated by the Mantel test. Also, the Principal Component Analysis (PCA) was computed from environmental values, and the samples were plotted in genetic structure grouping. The Spearman rank correlation was used to assess differences in mean number of alleles and ecogeographic variables among the inferred groups.
Authors' contributions
AC carried out the experiments and drafted the manuscript. CF, PS, GD and PH were involved in designing and planning the work, interpreting the results and critically editing the manuscript. PH, CF and PS conceived the study. PH coordinated the study and helped to draft the manuscript. All authors have read and approved the final manuscript.
Supplementary Material
Estimated number of populations from Geneland analysis. (a) Posterior density distribution of the number of clusters estimated from analysis in five replicates; and (b) genetic assignment of H. chilense individuals.
Mantel test showing the relationship between genetic distance and geographic distance. (B1) Relationship between Nei's genetic distance and Napierian logarithm of geographic distances for group I accessions; and (B2) relationship between Nei's genetic distance and Napierian logarithm of geographic distances for group II-Center accessions.
Mantel test showing the relationship between genetic distance and environmental distance for group II accessions. Plot of genetic distance vs. Ln (environmental distance).
Contributor Information
Almudena Castillo, Email: almudena.castillo@ias.csic.es.
Gabriel Dorado, Email: bb1dopeg@uco.es.
Catherine Feuillet, Email: catherine.feuillet@clermont.inra.fr.
Pierre Sourdille, Email: pierre.sourdille@clermont.inra.fr.
Pilar Hernandez, Email: phernandez@ias.csic.es.
Acknowledgements
Financial support from the Spanish Ministry of Science and Innovation (MICCIN-FEDER grant AGL2010-17316) is acknowledged. We gratefully acknowledge Dr Marion Röder for providing GWM primer sequences. The authors are grateful for the careful sampling and maintenance of the Hordeum chilense germplasm (see materials and methods), and for the data made publicly available by the DIVA-GIS project. This work is dedicated to the memory of Dr. Juan Ballesteros.
References
- Tanksley SD, McCouch SR. Seed banks and molecular maps: unlocking genetic potential from the wild. Science. 1997;277:1063–1066. doi: 10.1126/science.277.5329.1063. [DOI] [PubMed] [Google Scholar]
- Martín A, Martínez C, Rubiales D, Ballesteros J. In: Triticale: Today and Tomorrow. Guedes-Pinto H, Darvey N, editor. Carnide V: Kluwer Academic Publishers; 1996. Tritordeum: triticale's new brother cereal; pp. 57–72. [Google Scholar]
- Waples R. Evolutionary significant units and the conservation of biological diversity under the endangered species. Acta Am Fish Soc Symp. 1995;17:8–27. [Google Scholar]
- Pritchard JK, Rosenberg NA. Use of unlinked genetic markers to detect population stratification in association studies. Am J Hum Genet. 1999;65(1):220–228. doi: 10.1086/302449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buckler ES, Thornsberry JM. Plant molecular diversity and applications to genomics. Current Opinion in Plant Biology. 2002;5:107–111. doi: 10.1016/S1369-5266(02)00238-8. [DOI] [PubMed] [Google Scholar]
- Vaz Patto MC, Aardse A, Buntjer J, Rubiales D, Martín A, Niks RE. Morphology and AFLP markers suggest three Hordeum chilense ecotypes that differ in avoidance to rust fungi. Canadian Journal of Botany. 2001;79(2):204–213. doi: 10.1139/cjb-79-2-204. [DOI] [Google Scholar]
- Melchinger AE, Graner A, Singh M, Messmer MM. Relationships among european barley germplasm: Genetic diversity among winter and spring cultivars revealed by RFLPs. Crop Sci. 1994;34(5):1191–1199. doi: 10.2135/cropsci1994.0011183X003400050009x. [DOI] [Google Scholar]
- Barrett BA, Kidwell KK. AFLP-Based genetic diversity assessment among wheat cultivars from the Pacific Northwest. Crop Sci. 1998;38(5):1261–1271. doi: 10.2135/cropsci1998.0011183X003800050025x. [DOI] [Google Scholar]
- Huang XQ, Börner A, Röder MS, Ganal MW. Assessing genetic diversity of wheat (Triticum aestivum L.) germplasm using microsatellite markers. Theoretical and Applied Genetics. 2002;105(5):699–707. doi: 10.1007/s00122-002-0959-4. [DOI] [PubMed] [Google Scholar]
- Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ES. Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet. 2001;28(3):286–289. doi: 10.1038/90135. [DOI] [PubMed] [Google Scholar]
- Garris AJ, McCouch SR, Kresovich S. Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.) Genetics. 2003;165(2):759–769. doi: 10.1093/genetics/165.2.759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraakman ATW, Niks RE, Van den Berg PMMM, Stam P, Van Eeuwijk FA. Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics. 2004;168(1):435–446. doi: 10.1534/genetics.104.026831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nevo E, Beharav A, Meyer RC, Hackett CA, Forster BP, Russell JR, Powell W. Genomic microsatellite adaptive divergence of wild barley by microclimatic stress in 'Evolution Canyon', Israel. Biological Journal of the Linnean Society. 2005;84(2):205–224. doi: 10.1111/j.1095-8312.2005.00425.x. [DOI] [Google Scholar]
- Breseghello F, Sorrells ME. Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics. 2006;172(2):1165–1177. doi: 10.1534/genetics.105.044586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malysheva-Otto L, Ganal M, Röder M. Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.) BMC Genetics. 2006;7(1):6. doi: 10.1186/1471-2156-7-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rostoks N, Ramsay L, MacKenzie K, Cardle L, Bhat PR, Roose ML, Svensson JT, Stein N, Varshney RK, Marshall DF. et al. Recent history of artificial outcrossing facilitates whole-genome association mapping in elite inbred crop varieties. Proceedings of the National Academy of Sciences. 2006;103(49):18656–18661. doi: 10.1073/pnas.0606133103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chao S, Zhang W, Dubcovsky J, Sorrells M. Evaluation of genetic diversity and genome-wide linkage disequilibrium among U.S. wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci. 2007;47(3):1018–1030. doi: 10.2135/cropsci2006.06.0434. [DOI] [Google Scholar]
- Cockram J, White J, Leigh F, Lea V, Chiapparino E, Laurie D, Mackay I, Powell W, O'Sullivan D. Association mapping of partitioning loci in barley. BMC Genetics. 2008;9(1):16. doi: 10.1186/1471-2156-9-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yahiaoui S, Igartua E, Moralejo M, Ramsay L, Molina-Cano J, Ciudad F, Lasa J, Gracia M, Casas A. Patterns of genetic and eco-geographical diversity in Spanish barleys. Theoretical and Applied Genetics. 2008;116(2):271–282. doi: 10.1007/s00122-007-0665-3. [DOI] [PubMed] [Google Scholar]
- Comadran J, Thomas W, van Eeuwijk F, Ceccarelli S, Grando S, Stanca A, Pecchioni N, Akar T, Al-Yassin A, Benbelkacem A. et al. Patterns of genetic diversity and linkage disequilibrium in a highly structured Hordeum vulgare association-mapping population for the Mediterranean basin. TAG Theoretical and Applied Genetics. 2009;119(1):175–187. doi: 10.1007/s00122-009-1027-0. [DOI] [PubMed] [Google Scholar]
- Hübner S, Höffken M, Oren E, Haseneyer G, Stein N, Graner A, Schmid K, Fridman E. Strong correlation of wild barley (Hordeum spontaneum) population structure with temperature and precipitation variation. Molecular Ecology. 2009;18:1523–1536. doi: 10.1111/j.1365-294X.2009.04106.x. [DOI] [PubMed] [Google Scholar]
- Sourdille P, Tavand M, Charmet G, Bernard M. Transferability of wheat microsatellites to diploid Triticeae species carrying the A, B and D genomes. Theoretical and Applied Genetics. 2001;103(2-3):346–352. doi: 10.1007/s00122-001-0542-4. [DOI] [Google Scholar]
- Hernández P, Laurie DA, Martin A, Snape JW. Utility of barley and wheat simple sequence repeat (SSR) markers for genetic analysis of Hordeum chilense and tritordeum. Theoretical and Applied Genetics. 2002;104(4):735–739. doi: 10.1007/s001220100674. [DOI] [PubMed] [Google Scholar]
- Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ. Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Science. 2001;160:1115–1123. doi: 10.1016/S0168-9452(01)00365-X. [DOI] [PubMed] [Google Scholar]
- Taylor C, Madsen K, Borg S, Moller MG, Boelt B, Holm PB. The development of sequence-tagged sites (STSs) in Lolium perenne L.: the application of primer sets derived from other genera. Theoretical and Applied Genetics. 2001;103:648–658. doi: 10.1007/PL00002920. [DOI] [Google Scholar]
- Decroocq V, Favé MG, Hagen L, Bordenave L, Decroocq S. Development and transferability of apricot and grape EST microsatellite markers across taxa. Theoretical and Applied Genetics. 2003;106(5):912–922. doi: 10.1007/s00122-002-1158-z. [DOI] [PubMed] [Google Scholar]
- Gao L, Tang J, Li H, Jia J. Analysis of microsatellites in major crops assessed by computational and experimental approaches. Molecular Breeding. 2003;12(3):245–261. doi: 10.1023/A:1026346121217. [DOI] [Google Scholar]
- Varshney RK, Thiel T, Stein N, Langridge P, Graner A. In silico analysis on frequency and distribution of microsatellites in ESTs of some cereal species. Cellular & Molecular Biology Letters. 2002;7(2A):537–546. [PubMed] [Google Scholar]
- Yu JK, La Rota M, Kantety R, Sorrells M. EST derived SSR markers for comparative mapping in wheat and rice. Molecular Genetics and Genomics. 2004;271:742–751. doi: 10.1007/s00438-004-1027-3. [DOI] [PubMed] [Google Scholar]
- Zhang LY, Bernard M, Ravel C, Balfourier F, Leroy P, Feuillet C, Sourdille P. Wheat EST-SSRs for tracing chromosome segments from a wide range of grass species. Plant Breeding. 2007;126(3):251–258. doi: 10.1111/j.1439-0523.2007.01293.x. [DOI] [Google Scholar]
- Varshney RK, Sigmund R, Borner A, Korzun V, Stein N, Sorrells ME, Langridge P, Graner A. Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice. Plant Science. 2005;168(1):195–202. doi: 10.1016/j.plantsci.2004.08.001. [DOI] [Google Scholar]
- Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS. Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Molecular Genetics and Genomics. 2003;270(4):315–323. doi: 10.1007/s00438-003-0921-4. [DOI] [PubMed] [Google Scholar]
- Castillo A, Budak H, Varshney RK, Dorado G, Graner A, Hernández P. Transferability and polymorphism of barley EST-SSR markers used for phylogenetic analysis in Hordeum chilense. BMC Plant Biology. 2008;8(1):97. doi: 10.1186/1471-2229-8-97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X, Liu P, Han Z, Ni Z, Sun Q. Genetic diversity revealed by genomic-SSR and EST-SSR markers among common wheat, spelt and compactum. Progress in Natural Science. 2005;15(1):24–33. doi: 10.1080/10020070512331341730. [DOI] [Google Scholar]
- Balfourier F, Roussel V, Strelchenko P, Exbrayat-Vinson F, Sourdille P, Boutet G, Koenig J, Ravel C, Mitrofanova O, Beckert M. et al. A worldwide bread wheat core collection arrayed in a 384-well plate. TAG Theoretical and Applied Genetics. 2007;114(7):1265–1275. doi: 10.1007/s00122-007-0517-1. [DOI] [PubMed] [Google Scholar]
- Pan D, Yu-Ming W, Guo-Yue C, Wei L, Ji-Rui W, Eviatar N, You-Liang Z. EST-SSR diversity correlated with ecological and genetic factors of wild emmer wheat in Israel. Hereditas. 2009;146(1):1–10. doi: 10.1111/j.1601-5223.2009.02098.x. [DOI] [PubMed] [Google Scholar]
- Miller TE, Reader SM, Chapman V. In: Proc Int Symp Eucarpia on Induced variability in plant breeding. Broertjes C, editor. Pudoc, Wageningen; 1982. The addition of Hordeum chilense chromosomes to wheat; pp. 79–81. [Google Scholar]
- Castillo A, Budak H, Martín AC, Dorado G, Börner A, Röder M, Hernandez P. Interspecies and intergenus transferability of barley and wheat D-genome microsatellite markers. Annals of Applied Biology. 2010;156:347–356. doi: 10.1111/j.1744-7348.2010.00391.x. [DOI] [Google Scholar]
- Evanno G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology. 2005;14(8):2611–2620. doi: 10.1111/j.1365-294X.2005.02553.x. [DOI] [PubMed] [Google Scholar]
- Rosenberg NA. Distruct: a program for the graphical display of population structure. Molecular Ecology Notes. 2004;4:137–138. doi: 10.1046/j.1471-8286.2003.00566.x. [DOI] [Google Scholar]
- Di Castri F, Hajek E. Bioclimatología de Chile. Ediciones Universidad de Chile. 1976. p. 129.
- Mantel N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967;27(2_Part_1):209–220. [PubMed] [Google Scholar]
- Thiel T, Michalek W, Varshney RK, Graner A. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.) TAG Theoretical and Applied Genetics. 2003;106(3):411–422. doi: 10.1007/s00122-002-1031-0. [DOI] [PubMed] [Google Scholar]
- Zhang LY, Ravel C, Bernard M, Balfourier F, Leroy P, Feuillet C, Sourdille P. Transferable bread wheat EST-SSRs can be useful for phylogenetic studies among the Triticeae species. Theoretical and Applied Genetics. 2006;113(3):407–418. doi: 10.1007/s00122-006-0304-4. [DOI] [PubMed] [Google Scholar]
- Chabane K, Varshney R, Graner A, Valkoun J. Generation and exploitation of EST-derived SSR markers for assaying molecular diversity in durum wheat populations. Genetic Resources and Crop Evolution. 2008;55(6):869–881. doi: 10.1007/s10722-007-9292-8. [DOI] [Google Scholar]
- Gupta PK, Mir RR, Mohan A, Kumar J. Wheat genomics: present status and future prospects. International Journal of Plant Genomics. 2008. [DOI] [PMC free article] [PubMed]
- Salse J, Bolot S, Throude M, Jouffe V, Piegu B, Quraishi UM, Calcagno T, Cooke R, Delseny M, Feuillet C. Identification and characterization of shared duplications between rice and wheat provide new insight into grass genome evolution. Plant Cell. 2008;20(1):11–24. doi: 10.1105/tpc.107.056309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sorrells ME, La Rota M, Bermudez-Kandianis CE, Greene RA, Kantety R, Munkvold JD, Mahmoud A, Ma X, Gustafson PJ. et al. Comparative DNA sequence analysis of wheat and rice genomes. Genome Research. 2003;13(8):1818–1827. doi: 10.1101/gr.1113003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernández P, Dorado G, Cabrera A, Laurie DA, Snape JW, Martin A. Rapid verification of wheat-Hordeum introgressions by direct staining of SCAR, STS, and SSR amplicons. Genome. 2002;45(1):198–203. doi: 10.1139/g01-087. [DOI] [PubMed] [Google Scholar]
- Hernández P, Dorado G, Prieto P, Gimenez MJ, Ramirez MC, Laurie DA, Snape JW, Martin A. A core genetic map of Hordeum chilense and comparisons with maps of barley (Hordeum vulgare) and wheat (Triticum aestivum) Theoretical and Applied Genetics. 2001;102(8):1259–1264. doi: 10.1007/s001220000514. [DOI] [Google Scholar]
- Eujayl I, Sorrells ME, Baum M, Wolters P, Powell W. Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor Appl Genet. 2002;104(2-3):399–407. doi: 10.1007/s001220100738. [DOI] [PubMed] [Google Scholar]
- Lazrek F, Roussel V, Ronfort J, Cardinet G, Chardon F, Aouani M, Huguet T. The use of neutral and non-neutral SSRs to analyse the genetic structure of a Tunisian collection of Medicago truncatula lines and to reveal associations with eco-environmental variables. Genetica. 2008;135:391–402. doi: 10.1007/s10709-008-9285-3. [DOI] [PubMed] [Google Scholar]
- Crick FHC. Codon-anticodon pairing: The wobble hypothesis. Journal of Molecular Biology. 1966;19(2):548–555. doi: 10.1016/S0022-2836(66)80022-0. [DOI] [PubMed] [Google Scholar]
- Matus IA, Hayes PM. Genetic diversity in three groups of barley germplasm assessed by simple sequence repeats. Genome. 2002;45:1095–1106. doi: 10.1139/g02-071. [DOI] [PubMed] [Google Scholar]
- Russell JR, Ellis RP, Thomas WTB, Waugh R, Provan J, Booth A, Fuller J, Lawrence P, Young G, Powell W. A retrospective analysis of spring barley germplasm development from 'foundation genotypes' to currently successful cultivars. Molecular Breeding. 2000;6(6):553–568. doi: 10.1023/A:1011372312962. [DOI] [Google Scholar]
- Kilian B, Özkan H, Kohl J, von Haeseler A, Barale F, Deusch O, Brandolini A, Yucel C, Martin W, Salamini F. Haplotype structure at seven barley genes: relevance to gene pool bottlenecks, phylogeny of ear type and site of barley domestication. Molecular Genetics and Genomics. 2006;276(3):230–241. doi: 10.1007/s00438-006-0136-6. [DOI] [PubMed] [Google Scholar]
- Zhang L, Marchand S, Tinker N, Belzile F. Population structure and linkage disequilibrium in barley assessed by DArT markers. TAG Theoretical and Applied Genetics. 2009;119(1):43–52. doi: 10.1007/s00122-009-1015-4. [DOI] [PubMed] [Google Scholar]
- Blattner FR. Multiple intercontinental dispersals shaped the distribution area of Hordeum (Poaceae) New Phytologist. 2006;169(3):603–614. doi: 10.1111/j.1469-8137.2005.01610.x. [DOI] [PubMed] [Google Scholar]
- Nevo E, Brown AHD, Zohary D. Genetic diversity in the wild progenitor of barley in Israel. Cellular and Molecular Life Sciences. 1979;35(8):1027–1029. doi: 10.1007/BF01949920. [DOI] [Google Scholar]
- Nevo E, Zohary D, Beiles A, Kaplan D, Storch N. Genetic diversity and environmental associations of wild barley, Hordeum spontaneum, in Turkey. Genetica. 1986;68(3):203–213. doi: 10.1007/BF02424444. [DOI] [Google Scholar]
- Dawson IK, Chalmers KJ, Waugh R, Powell W. Detection and analysis of genetic variation in Hordeum spontaneum populations from Israel using RAPD markers. Molecular Ecology. 1993;2(3):151–159. doi: 10.1111/j.1365-294X.1993.tb00104.x. [DOI] [PubMed] [Google Scholar]
- Baum BR, Nevo E, Johnson DA, Beiles A. Genetic diversity in wild barley (Hordeum spontaneum C. Koch) in the Near East: a molecular analysis using Random Amplified Polymorphic DNA (RAPD) markers. Genetic Resources and Crop Evolution. 1997;44:147–157. doi: 10.1023/A:1008655023906. [DOI] [Google Scholar]
- Turpeinen T, Vanhala T, Nevo E, Nissilä E. AFLP genetic polymorphism in wild barley (Hordeum spontaneum) populations in Israel. TAG Theoretical and Applied Genetics. 2003;106(7):1333–1339. doi: 10.1007/s00122-002-1151-6. [DOI] [PubMed] [Google Scholar]
- Beaumont MA, Nichols RA. Evaluating loci for use in the genetic analysis of population structure. Proceedings: Biological Sciences. 1996;263(1377):1619–1626. doi: 10.1098/rspb.1996.0237. [DOI] [Google Scholar]
- Li Y, Fahima T, Korol AB, Peng J, R MS, Kirzhner V, Beiles A, Nevo E. Microsatellite diversity correlated with ecological-edaphic and genetic factors in three microsites of wild emmer wheat in North Israel. Mol Biol Evol. 2000;17(6):851–862. doi: 10.1093/oxfordjournals.molbev.a026365. [DOI] [PubMed] [Google Scholar]
- Ivandic V, Hackett CA, Nevo E, Keith R, Thomas WTB, Forster BP. Analysis of simple sequence repeats (SSRs) in wild barley from the Fertile Crescent: associations with ecology, geography and flowering time. Plant Molecular Biology. 2002;48(5):511–527. doi: 10.1023/A:1014875800036. [DOI] [PubMed] [Google Scholar]
- Tobes N, Ballesteros J, Martínez C, Lovazzano G, Contreras D, Cosio F, Gastó J, Martín L. Collection mission of H. chilense Roem. et Schult. in Chile and Argentina. Genetic Resources and Crop Evolution. 1995;42(3):211–216. doi: 10.1007/BF02431254. [DOI] [Google Scholar]
- Giménez MJ, Cosío F, Martínez C, Silva F, Zuleta A, Martín LM. Collecting Hordeum chilense Roem. et Schult. germplasm in desert and steppe dominions of Chile. Plant Genet Resour Newsl. 1997;109:17–19. [Google Scholar]
- Nicot N, Chiquet V, Gandon B, Amilhat L, Legeai F, Leroy P, Bernard M, Sourdille P. Study of simple sequence repeat (SSR) markers from wheat expressed sequence tags (ESTs) TAG Theoretical and Applied Genetics. 2004;109(4):800–805. doi: 10.1007/s00122-004-1685-x. [DOI] [PubMed] [Google Scholar]
- Zhang LY, Bernard M, Leroy P, Feuillet C, Sourdille P. High transferability of bread wheat EST-derived SSRs to other cereals. TAG Theoretical and Applied Genetics. 2005;111(4):677–687. doi: 10.1007/s00122-005-2041-5. [DOI] [PubMed] [Google Scholar]
- Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW. A microsatellite map of wheat. Genetics. 1998;149(4):2007–2023. doi: 10.1093/genetics/149.4.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li JZ, Sjakste TG, Roder MS, Ganal MW. Development and genetic mapping of 127 new microsatellite markers in barley. Theoretical and Applied Genetics. 2003;107(6):1021–1027. doi: 10.1007/s00122-003-1345-6. [DOI] [PubMed] [Google Scholar]
- Liu K, Muse SV. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics. 2005;21(9):2128–2129. doi: 10.1093/bioinformatics/bti282. [DOI] [PubMed] [Google Scholar]
- Nei M, Li H. Mathematical model for studying genetic variation in terms restriction endonucleases. Proceedings of the National Academy of Sciences. 1979;76:5269–5273. doi: 10.1073/pnas.76.10.5269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Felsenstein J PHYLIP (Phylogeny Inference Package) version 3.66 Distributed by the author. Department of Genome Sciences. University of Washington, Seattle 2006. 16364968
- Page RDM. Tree View: An application to display phylogenetic trees on personal computers. Computer Applications Biosciences. 1996;12(4):357–358. doi: 10.1093/bioinformatics/12.4.357. [DOI] [PubMed] [Google Scholar]
- Dray S, Dufour AB. The ade4 package: implementing the duality diagram for ecologists. Journal of statistical software. 2007;4:1–20. [Google Scholar]
- Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155(2):945–959. doi: 10.1093/genetics/155.2.945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guillot G, Mortier F, Estoup A. Geneland: a computer package for landscape genetics. Molecular Ecology Notes. 2005;5(3):712–715. doi: 10.1111/j.1471-8286.2005.01031.x. [DOI] [Google Scholar]
- Excoffier L, Laval G, Schneider S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online. 2005;1:47–50. [PMC free article] [PubMed] [Google Scholar]
- Antao T, Lopes A, Lopes R, Beja-Pereira A, Luikart G. LOSITAN: A workbench to detect molecular adaptation based on a Fst-outlier method. BMC Bioinformatics. 2008;9(1):323. doi: 10.1186/1471-2105-9-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Estimated number of populations from Geneland analysis. (a) Posterior density distribution of the number of clusters estimated from analysis in five replicates; and (b) genetic assignment of H. chilense individuals.
Mantel test showing the relationship between genetic distance and geographic distance. (B1) Relationship between Nei's genetic distance and Napierian logarithm of geographic distances for group I accessions; and (B2) relationship between Nei's genetic distance and Napierian logarithm of geographic distances for group II-Center accessions.
Mantel test showing the relationship between genetic distance and environmental distance for group II accessions. Plot of genetic distance vs. Ln (environmental distance).