Abstract
Metal binding by apo-manganese superoxide dismutase (apo-MnSOD) is essential for functional maturation of the enzyme. Previous studies have demonstrated that metal binding by apo-MnSOD is conformationally gated, requiring protein reorganization for the metal to bind. We have now solved the X-ray crystal structure of apo-MnSOD at 1.9 Å resolution. The organization of active site residues is independent of the presence of the metal cofactor, demonstrating that protein itself templates the unusual metal coordination geometry. Electrophoretic analysis of mixtures of apo- and (Mn2)-MnSOD, dye-conjugated protein, or C-terminal Strep-tag II fusion protein reveals a dynamic subunit exchange process associated with cooperative metal binding by the two subunits of the dimeric protein. In contrast, (S126C) (SS) apo-MnSOD, which contains an inter-subunit covalent disulfide crosslink, exhibits anticooperative metal binding. The protein concentration dependence of metal uptake kinetics implies that protein dissociation is involved in metal binding by the wild type apo-protein, although other processes may also contribute to gating metal uptake. Protein concentration dependent small-zone size exclusion chromatography is consistent with apo-MnSOD dimer dissociation at low protein concentration (KD = 1×10−6 M). Studies on metal uptake by apo-MnSOD in Escherichia coli cells show that the protein exhibits similar behavior in vivo and in vitro.
Keywords: superoxide; dismutase, manganese; protein interactions; metal binding; electrophoretic mobility shift
1. Introduction
Superoxide dismutase (SOD) is an important component of the cellular defense against oxidative stress, protecting cells from damage by toxic superoxide free radical formed as an unavoidable by-product of aerobic metabolism [1,2]. All known SODs are metalloenzymes containing a redox-active metal ion (Cu, Fe, Mn or Ni) in their active site [3–6]. Acquisition of the metal cofactor is a crucial step in the maturation of the SOD metalloproteins, and is required for expression of their catalytic activity. Recent studies have shed light on the mechanism of metal cofactor insertion into Cu,Zn SOD. In yeast, a copper chaperone (CCS) has been shown to be required for intracellular delivery of copper to the precursor SOD apo-protein [7,8]. In contrast, no metallochaperone has yet been identified for metal insertion into the yeast mitochondrial MnSOD, and in vitro studies have shown that E. coli apo-MnSOD is capable of picking up metal on its own without assistance from other proteins [9–11].
The metal binding site in the homodimeric E. coli MnSOD is buried in the interior of the protein, near the subunit interface (Fig. 1) [5]. At present very little is known about how E. coli MnSOD acquires its metal cofactor inside the cell. Early studies showed that metal binding by MnSOD in vivo is relatively unselective, and Mn2-, Fe2- and mixed (Mn,Fe)- forms of the protein are all present [13–15]. In spite of this nonspecific metal binding behavior, only the manganese-containing form is active, reflecting a strict catalytic specificity for the Mn ion [16].
Figure 1.

Organization of the E. coli Mn2-MnSOD holo-protein. The homodimeric protein is shown with the metal ion rendered as a cyan sphere, and surrounding amino acid residues comprising the metal environment (His26, His81, Gln146, Asp167, His171) shown in space-filling view (purple). Individual subunits are color-coded (red and green), and one subunit surface is rendered transparent to reveal the internal architecture. (A) View perpendicular to molecular 2-fold axis. Inset expands the view of the metal environment. (B) View parallel to molecular 2-fold axis. (Based on PDB ID: 1VEW; visualized using CHIMERA [12]).
Nevertheless, the nonspecific metal binding behavior of apo-MnSOD has made it possible to monitor the metal uptake process in vitro using a continuous fluorimetric metal binding assay based on the efficient quenching of intrinsic protein tryptophan luminescence by cobalt ions [10]. The metal binding reaction exhibits two distinct kinetic phases: a fast phase that represents a fraction of the protein that is capable of rapidly binding the metal ion from solution (the “open” state), and a slow phase that reflects the relatively slow conversion from the stable “closed” state of the protein to the “open” state. The proportion of open state increases with increasing pH and temperature, indicating that the two species of protein in the sample are interconverting in a dynamic equilibrium. Previous studies aimed at defining the structural determinants involved in the conversion of closed to open form in conformationally gated metal binding demonstrated that, while covalently constraining the domain interface within each subunit has no effect, covalent disulfide constraints on the subunit interface significantly perturbs metal uptake by apo-MnSOD, but does not prevent the process [11]. Mutagenesis of residues in the active site environment (near the subunit interface) appears to favor formation of the open state. Together, these observations have led to a mechanistic proposal for metal uptake by apo-MnSOD involving reorientation of residues on the subunit interface to create a metal entry channel leading to the buried metal binding site.
We have now solved the structure of E. coli apo-MnSOD crystallized under conditions favoring the “open” form of the protein, for comparison with the structure of fully metallated Mn2-MnSOD. We have also extended the investigations of subunit interactions in apo-MnSOD using a combination of approaches (including electrophoretic resolution of the different metalloforms present in MnSOD mixtures and small-zone size exclusion chromatography (SEC)), revealing dynamic aspects of the protein structure, and providing insight into the physical processes underlying metal binding by apo-MnSOD. The role of subunit separation in the metal uptake reaction has been investigated through analysis of the protein concentration dependence of metal uptake kinetics, and production of apo-MnSOD in E. coli cells has allowed the in vivo and in vitro metal uptake behavior to be compared.
2. Materials and Methods
2.1 Biochemical reagents
All reagents were from commercial sources and used without purification.
2.2 Culture media
Terrific Broth (TB) (12 g/L tryptone, 24 g/L yeast extract, 2.31 g/L potassium phosphate monobasic, 12.54 g/L potassium phosphate dibasic) and Luria-Bertani medium (LB) (5 g/L NaCl, 5 g/L yeast extract, 10 g/L tryptone) were supplemented with antibiotics as required for selection (carbenicillin, 125 mg/L). Modified MOPS minimal medium (40 mM MOPS, 4 mM Tricine, pH 7.2, containing 50 mM sodium chloride, 1.32 mM potassium phosphate, 9.5 mM ammonium chloride, 0.28 mM potassium sulfate, 0.5 mM magnesium sulfate, 0.5 μM calcium chloride, and micronutrients, but without iron or manganese) was prepared as previously described [17].
2.3 Biological materials
Ultracompetent Escherichia coli XL2-Blue cells were from Stratagene (La Jolla, CA). Electrocompetent E. coli cells of other strains were routinely prepared by standard procedures [18] and electrotransformed using an Eppendorf 2510 electroporator with 1 mm cuvette (18 kV/cm). The E. coli ΔsodA knockout strain was constructed by phage lambda Red recombineering methods from E. coli BW25113 [11,19,20].
2.4 Expression Plasmids
The arabinose-inducible pBAD2sodA vector containing the E. coli sodA structural gene under the tight, strong PBAD promoter [21] was routinely used for homologous expression of recombinant E. coli MnSOD, as previously described [11]. The Quik-Change Multi mutagenesis procedure (Stratagene, La Jolla, CA) was used to prepare the pBAD-sodA (Q21C) mutational variant, using pBAD2sodA as template and the mutagenic primer 5′P- CCGCA CTTCG ATAAG TGCAC CATGG AAATC CACCA CACC-3′. The pBAD-sodACStrep vector for expression of C-terminal Strep-tag II [22] MnSOD fusion protein was constructed by amplification of the sodA coding sequence from the pQGBsodA template [23,24] using a forward primer containing an Nde I restriction site (5′-CGACA TATGA GCTAT ACCCT GCCAT CCCTG CCG-3′) and a reverse primer containing nucleotides encoding the Strep-tag II peptide (Trp-Ser-His-Pro-Gln-Phe-Glu-Lys)[22] and a Hind III restriction site (5′-GTCAA GCTTA TTATT TTTCG AACTG CGGGT GGCTC CAAGC GCTTT TTTTC GCCGC AAAAC GTGCC-3′). The sodA-CStrep PCR product was triply digested with a mixture of Nde I, Hind III, and Dra I and ligated into an Nde I/Hind III digested pBAD2 vector to produce the pBAD-sodA-CStrep expression vector. The sequences of the inserts were verified by direct sequence analysis (Molecular Biology Core Sequencing Facility, Oregon Health and Science University). Expression vectors were electrotransformed into electrocompetent E. coli BW25113 ΔsodA cells for protein production.
2.5 Recombinant protein expression
E. coli BW25113 ΔsodA | pBAD2sodA and E. coli BW25113 ΔsodA | pBAD2sodA (Q21C) expression strains were inoculated into 1 L of Terrific Broth (TB) supplemented with 2 g/L glucose and 125 mg/L carbenicillin and grown to saturation at 37°C with shaking. Cells were collected and resuspended in 1 L TB (minus phosphate) supplemented with 5 mM MnCl2, 2 g/L glycerol and 0.5 g/L L-(+)-arabinose. Cells were harvested after incubation at 37°C for 24 h with shaking. The E. coli BW25113 ΔsodA | pBAD2sodA-CStrep expression strain was grown in modified MOPS minimal medium. E. coli MnSOD (E170A) and (S126C) variants were prepared as previously described [11,25].
2.6 Protein purification
E. coli MnSOD was purified by a modification of previously described methods [11]. The cell-free extract from 15 g of cells was loaded onto an ion exchange column (DE-52, 5×45 cm) and the flow-through SOD fractions were further purified by CM-52 chromatography or chromatofocussing (PBE-94 Polybuffer Exchanger developed with Polybuffer-74 ampholyte).
2.7 Preparation of apo-protein
Apo-MnSOD was prepared by denaturation of the purified metal-containing protein in 3.5 M Guanidinium HCl (GdnHCl) (pH 3.5) containing 10 mM EDTA and renaturation by dialysis against 20 mM Tris/EDTA (pH 7.6) as previously described [9,10]. EDTA and 2-mercaptoethanol (2-ME) were removed by gel filtration (BioGel P-6). Apo-MnSOD isolated from cells grown under metal-limitation in MOPS minimal medium was purified from cell extracts by DE-52 and CM-52 ion exchange chromatography. The apo-protein product was stored at −80°C.
2.8 In vivo metal uptake by apo-MnSOD
E. coli BW25113 ΔsodA cells containing the pBAD2sodA expression vector were used for the production of apo-MnSOD. The cells were grown in modified MOPS minimal medium [17] for both growth phase (supplemented with 0.2% glucose, 100 nM iron sulfate and 125 μg/mL carbenicillin) and induction phase (supplemented with 0.2% glycerol, 30 mM L-(+)-arabinose and 125 μg/mL carbenicillin). Cells from a fresh agar plate were inoculated into 50 mL of growth phase medium to O.D.600 nm = 0.05 and grown overnight at 35°C. The cells were collected by centrifugation (5000 rpm, 10 min) and resuspended in 24 mL of induction medium containing L-(+)-arabinose. 2 mL aliquots of the cell suspension were used for protein expression and metal uptake studies. After 4 hours induction at 35°C, chloramphenicol (Cm) was added to the medium (to 50 μg/mL) to arrest de novo protein biosynthesis. After 10 min at 35°C, manganese chloride was added and incubation was continued. The cells were collected by centrifugation and washed twice with 1 mL of 100 mM potassium phosphate buffer (pH 7). The cell pellet was resuspended in 0.3 mL of B-PER reagent (Pierce Biotechnology, Rockford, IL) containing 1 mM EDTA, and the soluble protein was extracted from the cells according to the instructions provided by the manufacturer. The cell-free extract was loaded onto a DE-52 ion exchange chromatography column (0.75×3.5 cm) equilibrated with native PAGE sample buffer (62.5 mM Tris-HCl, pH 6.8, containing 1 mM EDTA). The column flow-through peak fraction (monitored by 280 nm absorption and gel electrophoresis) was used for further analysis by electrophoresis. Holo- and apo-MnSOD were resolved on precast 12% Tris-HCl Ready-Gels (Bio Rad Laboratories, Hercules, CA) with 1 mM EDTA added to the running buffer. The gel chamber was chilled in ice and run for approximately 2 hours (20 mA/gel). The gel was prefixed with a 50% methanol/7% acetic acid solution and stained with GelCode Blue stain reagent (Pierce Biotechnology, Rockford, IL). For superoxide dismutase activity assays, the B-PER cell-free extract was used directly, without further processing.
2.9 Dye conjugation
Purified apo-MnSOD (Q21C) (2 mg) in 1 mL of 10 mM HEPES pH 7.5 containing 150 mM NaCl and 1 mM EDTA was reacted with 0.5 mg Alexa-Fluor 594 C5-maleimide or Alexa-Fluor 647 C2-maleimide (Invitrogen, Carlsbad, CA) dissolved in 30 μL of anhydrous N,N-dimethyl formamide. The reaction mixture was stirred under argon at 35°C for 1 hour, and the labeled protein was separated from excess free dye by gel filtration over Sephadex G-25 (1.5×49 cm) equilibrated with the reaction buffer. The protein-containing fractions were concentrated by YM-10 ultrafiltration (Millipore, Billerica, MA) and desalted over Bio-Gel P-6. The degree of labeling was estimated by measuring the absorption at 280 nm and 588 nm (for Alex Fluor 594) or 650 nm (for Alexa Fluor 647) (Equation 1):
| (1) |
where AAF is the absorption of the dye conjugate at the stated wavelength; εAF is the extinction coefficient for the dye at that wavelength (εAF 594 (588 nm) = 7.3×104 M−1cm−1; εAF 647 (650 nm) = 2.39×105 M−1cm−1); cAF reflects the relative contribution of the Alexa Fluor dye to absorption at 280 nm (0.56 for Alexa Fluor 594 and 0.03 For Alexa Fluor 647); and εMnSOD is the molar extinction coefficient for E. coli MnSOD at 280 nm (εMnSOD 280 nm =8.66×104 M−1cm−1)[26].
2.10 Protein crystallization
Crystallization trials for apo-MnSOD were set up as previously described for holo-MnSOD [5], except that 1 mM EDTA was included in the crystallization buffer and the cryo-protection solution. Crystals were grown at room temperature using the sitting drop method. The drops consisted of 4 μL of protein (17 mg/mL in 1 mM EDTA) and 4 μL of reservoir solution (20% PEG 6000, 0.1 M Bicine pH 8.0, 1 mM EDTA). Drops were surrounded with 0.9 mL of reservoir solution. Colorless crystals of a rod-shaped or cubic morphology grew, under the same conditions, over 1–2 weeks, and were designated forms I & II. Selected crystals were transferred to cryo-protectant solution (25% PEG 6000, 15% MPD, 1 mM EDTA in 0.1 M Bicine pH 8.0) for structure analysis.
2.11 Structure determination
X-ray diffraction data were collected at 100K from the crystals. Form I diffracted X-rays to 2.0Å resolution, measured on a Rigaku R-Axis IV image plate detector, using Cu-Kα radiation from a Rigaku (RU-200) rotating anode generator. Form II diffracted X-rays to 1.9Å resolution at the Advanced Light Source synchrotron beam line 4.2.2, measured on a NOIR-1 CCD detector. The diffraction patterns of all crystals revealed secondary lattices, likely due to crystal cracking or satellite crystals, but these were weak enough that they could be ignored. In spite of the obvious morphological differences between the two crystal forms, preliminary structures determined from them were essentially identical. Form II yielded higher resolution diffraction, so a dataset of 360 × 0.5° oscillation images was used in refinement. Diffraction images were indexed and integrated in Denzo and the resulting data were scaled and merged in Scalepack [27]. A summary of the data processing statistics is shown in Table 1. The crystals belong to spacegroup C2221 and contain 2 MnSOD dimers in the asymmetric unit (a.s.u.).
Table 1.
Summary of diffraction data and refinement statistics
| Parameter | apo MnSOD |
|---|---|
| Space group | C2221 |
| Unit cell dimensions (Å) | a = 94.9, b = 106.6, c = 178.0 |
| Resolution range (Å)a | 44.0–1.90 (1.93–1.90) |
| I/σI | 35.8 (2.8) |
| Rmerge (%)b | 6.9 (54.6) |
| Redundancy | 7.2 (5.4) |
| Completeness (%) | 99.9 (99.4) |
| Rcryst/Rfree | 0.192/0.236 |
| RMSD bond lengths (Å)/RMSD bond angles (°) | 0.006/1.03 |
| Number of atoms in a.s.u. | 7202 |
| Protein | 6562 |
| Water | 640 |
| Avg. B-factor (Å2) | 47.5 |
Numbers in parentheses are values in highest resolution shell
Rmerge = Σhkl Σi |Ii(hkl) − <I(hkl)>|/Σhkl Σi Ii(hkl), where Ii(hkl) is the ith observation of a symmetry equivalent of reflection hkl.
The prior holo-MnSOD structure [5] (PDB ID: 1VEW) was nearly isomorphous with the current apo-MnSOD form and could be used as a phasing model. Cross-validation reflections were not identified in the holo-MnSOD database entry, so a new test set of 3569 reflections (5%) were selected randomly for refinement of the apo form. A model consisting of the protein atoms from the Mn-bound structure was optimized against the apo-MnSOD data, starting with rigid group refinement of each chain (4 chains total from 2 homo-dimers). Simulated annealing refinement followed to mitigate bias in the electron density maps from the starting model. Additionally, averaged kick mFo-DFc electron density maps [28] calculated with active site residues His26, His81, Asp167 and His171 omitted showed clear electron density, verifying that the maps and final structure were not biased. Refinement of coordinates, individual B-factors and TLS parameters were performed in Phenix [29], and manual adjustments to the model were performed using Coot [30]. Water molecules were first added in Phenix, and additional waters were identified in Coot. The model was refined to a final Rwork/Rfree of 0.192/0.236 (Table 1). The quality of the model was assessed using Procheck [31]. Alignment of the apo- and Mn-bound structures of MnSOD was performed by secondary structure matching (SSM) calculations within Coot.
2.12 Typhoon imaging
Fluorescence imaging of dye-labeled proteins resolved on native PAGE gels was performed using a Typhoon TRIO+ Variable Mode Imager (Amersham Biosciences). For detection of Alex Fluor 594 dye conjugate, the green laser (532 nm) excitation was used and the emission filter was set to 610 nm with 30 nm bandpass; for detection of Alex Fluor 647 dye conjugate, the red laser (633 nm) excitation was used and the emission filter was set to 670 nm with 30 nm bandpass. For detection of FRET, green laser (532 nm) excitation was used with the emission filter set to 670 nm (30 nm bandpass). Scans were analyzed using ImageQuant software (Amersham Biosciences).
2.13 Size exclusion chromatography
Protein concentration-dependent dimer dissociation of apo- and holo-MnSOD was determined by Sephacryl 100-HR (1.5×99 cm) small-zone size exclusion chromatography at 45°C in a thermostated column with temperature controlled by a circulating water bath. The column was equilibrated in 20 mM MOPS buffer (pH 7.6) containing 1 mM EDTA. Variable amounts of protein (35–700 μg) were loaded and the elution profile was monitored by optical density at 280 nm or intrinsic protein luminescence (λEX = 280 nm; λEM = 333 nm). Experimental elution profiles were simulated using the program SCIMMS [32,33]. Blue dextran, bovine serum albumin, Mn2-MnSOD and Fe2-MnSOD (E170A) (1 mg each) were used as molecular weight standards.
2.14 Metal uptake assay
Fluorimetric metal uptake measurements were performed as previously described [10,11] using a Cary Eclipse spectrofluorimeter (Varian, Walnut Creek, CA) equipped with a Cary temperature controller and a Peltier cell holder. Gel mobility was also used to monitor the progress of the metal uptake reaction. For time-dependent studies, a slight excess (1.25 equivalents) or substoichiometric (0.4 equivalents) amount of ice cold manganese chloride or cobalt chloride was added to the apo-MnSOD protein (0.1 mM) in 20 mM MOPS buffer pH 7.8 in an ice bath. After removing a zero time reference sample, the mixture was transferred to a preheated thin-walled PCR tube in a 37°C water bath. Aliquot samples were transferred to ice cold thin-walled PCR tubes to stop the reaction at the specified times. Stoichiometric metal uptake was measured by addition of aliquots of solution of the metal ion to apo-protein in 40 mM MOPS buffer pH 7.8 and incubation of the sample in thin-walled PCR tube for 30 min in a 37°C water bath.
2.15 Subunit exchange
Equal amounts of Mn2-MnSOD and apo-MnSOD (80 μM each) in ice cold 20 mM MOPS pH 7.8 (with or without EDTA, 6.25 mM) were incubated at 45°C for 0–3 hours, and then chilled in an ice bath to quench the reaction. Holo-, apo-, and half-metallated MnSOD protein species were resolved by gel mobility shift using native PAGE. A reaction mixture containing equal amounts of Mn2-MnSOD and apo-MnSOD incubated at 37°C for 15 hours was used as a standard for the three species. Subunit exchange experiments using dye-labeled Alexa Fluor 594 and Alexa Fluor 647 conjugated protein (40 μM each) was performed at 37°C for 0–1 hour. Subunit exchange between wt Mn2-MnSOD and C-terminal Strep-tag II Mn2-MnSOD fusion protein was performed at 37°C for 15 hours. In the latter case, the protein species were resolved by native gel electrophoresis run for 4 hours with the gel chamber cooled in an ice bath.
2.16 Image analysis
Scanned images of electrophoresis gels were quantitated by gel densitometry using the program ImageJ [34,35]. Data from the densitometric processing was utilized to evaluate kinetic rate constants and equilibrium parameters using the programs Kaleidograph (Synergy Software, Reading, PA) and Scientist (Micromath, Saint Louis, MO).
2.17 Evaluation of equilibrium parameters and binding cooperativity from gel electrophoretic mobility shift data
The densitometric data generated by electrophoretic resolution of species (apo-, half-apo-and holo-protein) populated in the course of the stoichiometric metal binding titration (see above, section 2.14) were normalized within each lane to the sum of densities over all three species. The normalized speciation data was then analyzed using the program Scientist to evaluate binding constants. The reaction associated with stoichiometric metal binding may be written (Equation 2):
| (2) |
where P represents the uncomplexed apo-protein, PM is the half-apo form and PM2 the holo-protein, and K1 and K2 are association constants for the formation of half-apo and holo-protein complexes, respectively (Equations 3&4):
| (3) |
| (4) |
The cooperativity of metal binding is reflected in the relative values of the association constants K1 and K2. The fraction of each stoichiometric species resolved in the gels may be represented by Equations 5–7[36]:
| (5) |
| (6) |
| (7) |
with Z = (1 + K1[M] + K2[M]2) and Θi being defined as the mole fraction of protein with exactly i metal ions bound. Each of these equations contains the independent variable [M], corresponding to the concentration of free uncomplexed metal ion in the sample. Under stoichiometric conditions ([P] ≫ K), [M] is vanishingly small, and in particular cannot be reliably estimated from the total metal ion concentration ([M]0) and the concentrations of the complexed species (Equation 8):
| (8) |
However, in the course of the titration, [M] increases (approximately logarithmically) from 0 to a concentration approaching [P] as equivalency is reached. The titration progress parameter ([M]0/[P]0) can be transformed to a variable in the space of the free metal ion [M] by exponentiation (10(([M]0/[P]0)q1−q2)), with parameters q1 and q2 representing the spread and shift of the range of [M], respectively. The individual speciation curves for the three components are then globally fit to equations 5–7 for the fractional populations, extracting four parameters (K1, K2, q1 and q2).
2.18 Protein characterization
The concentration of purified SOD was determined by optical absorption measurements, using the published molar extinction coefficient (ε280 nm =8.66×104 M−1cm−1) [26], or by the method of Lowry et al. [37]. Superoxide dismutase activity was measured with the xanthine oxidase/cytochrome c inhibition assay [38]. Metal analyses were performed using a Varian Instruments SpectrAA Model 20B atomic absorption spectrometer equipped with a GTA 96 graphite furnace. Protein homogeneity was routinely evaluated using discontinuous SDS-PAGE in 12% Tris-HCl Ready-Gel (Bio-Rad Laboratories, Hercules, CA) with 5% 2-ME included in the sample buffer.
3. Results and Discussion
3.1 X-ray crystal structure of E. coli apo-MnSOD
The structure of apo-MnSOD has been determined to 1.9Å resolution, permitting a comparison with the holo-protein structure [5]. Despite the expectation that the apo form of the protein would reveal a more open conformation, the apo and Mn-bound forms of MnSOD are surprisingly similar. After separately aligning each monomer from the two structures by the secondary structure matching algorithm [39], the root mean squared deviation (RMSD) for Cα atoms is 0.33Å for the first monomer and 0.34Å for the second monomer. The maximum- likelihood based coordinate error estimate of the Apo-MnSOD model is 0.28Å, corresponding to a random component of ~0.4Å in an all-atom difference calculation. Therefore, most of the differences between the apo- and Mn-bound structures are not statistically significant. The largest differences are near the N- and C-termini and within two surface-exposed regions of the protein. One region contains residues Ser44 through Glu47, and the other contains residues Gly135 and Asp136. The latter region is involved in crystal packing contacts, the likely cause of local Cα deviations of 0.76–0.99Å.
Within the dimer, there is a rotation of 2.2° of one subunit relative to the other and a displacement of 1.1Å. The change is small, but statistically significant, and indicates a degree of flexibility at the subunit interface. A much larger difference was observed in the relative orientation of the two apo-MnSOD dimers in the crystallographic asymmetric unit. Comparing the crystal packing of the proteins in the Mn-bound and apo-MnSOD structures, one dimer is rotated by 22° and translated by 26Å relative to the other. Although this difference is probably due to crystal packing and is unlikely to be physiologically relevant, it provides further verification that the apo crystals are distinguishable from the metallated crystals.
The similarity of apo- and holo-enzymes is particularly striking in the region of the Mn-binding site. The RMSDs of atoms in the five side-chains comprising the extended coordination environment of the metal ion (His26, His81, Gln146, Asp167, His171) range between 0.19Å and 0.35Å and therefore are not significantly different beyond the estimated coordinate error of the model. Electron density maps calculated from the apo-MnSOD data did not show any evidence of Mn (or any other metal ion) in the active site, although a water molecule (modeled as a hydroxide ion in the holo-MnSOD structure) was clearly present (Fig. 2). In order to confirm that the crystals were devoid of Mn, a full spectrum X-ray fluorescence scan of the crystal was performed. No Mn emission peak was observed at 5898.8eV, confirming that the crystals were of the apo-form. The distances between the active site water and the coordinating residues (2.5–2.85Å from Gln146 NE2, Asp167 OD2, and His171 NE2) further suggest a hydrogen-bonding network involving these residues, as opposed to the metal coordination that orients the buried solvent in the earlier holo-structure [5].
Figure 2.

Structure of the metal binding site in apo-MnSOD. The active site of apo-MnSOD is shown with four of the active site residues displayed as sticks and labeled (active site residue Asp167 is omitted for clarity). The active site water molecule is shown as a light blue sphere, and is within hydrogen-bonding distance from Gln146, Asp167, and His171. Manganese, superposed from the holo-MnSOD structure (PDB ID: 1VEW), is shown as a magenta sphere for reference, but was not present in the apo crystals. A 2mFo−DFc electron density map is shown as a blue mesh at a contour level of 1.0σ surrounding the atoms in the active site. (Based on PDB ID: 3OT7. The image was generated using PyMOL [40].)
Neither the apo- nor Mn-bound SOD protein subunits contain an open channel connecting the Mn-binding site to the surrounding solvent that could serve as an entry point for metal binding, although the active site is situated quite close to the dimer interface. A total of 875Å2 of the surface on each monomer is buried at the dimer interface. Interestingly, His171, a key residue in coordinating Mn binding, forms part of the dimer interface in both the holo- [5] and the apo-protein. In the apo-MnSOD structure, His171 from the first subunit contributes 40Å2, while the corresponding residue in the opposing subunit contributes 55Å2 of the surface area at the dimer interface. This important contribution of His171 to the dimer interface in both apo- and Mn-bound forms of MnSOD might account for its conserved structure. The position of His171 is constrained, in part, by interactions with Glu170 of the adjacent dimeric subunit. In the dissociated monomer, this constraint would be removed, potentially allowing His171 to adopt alternative conformations. In fact, modeling an alternate rotomer conformation for the side-chain of His171 opens a channel that could provide access to the active site [11]. Such a change may be facilitated in the monomeric form of MnSOD
3.2 Electrophoretic mobility shift analysis of metal uptake by apo-MnSOD
The E. coli MnSOD polypeptide contains a nearly equal number of acidic (Glu,Asp: 25 residues) and basic (Lys,Arg: 23 residues) amino acids, and consequently has only a small net charge under standard discontinuous native protein electrophoresis conditions (Tris-HCl pH 8.3). Binding a metal cation significantly alters the net protein charge and as a result, different metalloforms of E. coli MnSOD exhibit distinct electrophoretic mobilities (Fig. 3), allowing the progress of metal uptake by wt apo-MnSOD to be monitored. Time-dependent metal uptake by apo-MnSOD at 37°C in the presence of a slight excess of metal ion (1.25 equivalents Mn2+) shows that all apo-protein is converted to holo-protein within 30 minutes (Fig. 3A, 3–10). Interestingly, no half-metallated species was detected during the entire time-course. The pseudo first order rate constants for disappearance of apo-protein (k = 0.21±0.02 min−1) and appearance of holo-protein (k = 0.17±0.01 min−1) evaluated from the gel data (Figure 3B) are the same, consistent with rapid, sequential binding of two metal ions in the limit of strong positive cooperativity.
Figure 3.
Time dependence of in vitro metal uptake by apo-MnSOD. MnSOD standards prepared by incubating equal amounts of Mn2-MnSOD and apo-MnSOD, 37°C, 15 h: a, Mn2-MnSOD, b, Mn1-MnSOD, and c Apo-MnSOD. (A) Metal uptake was initiated by addition of 1.25 equivalents of MnCl2 to apo-MnSOD as described in the Materials and Methods (Section 2.14). (1) MnSOD standards; (2) equal amounts of Mn2-MnSOD and apo-MnSOD mixed immediately before loading on the gel; (3–10) MnCl2 was added to apo-MnSOD and the metal uptake reaction was stopped at the indicated time points: 0, 1, 3, 5, 7, 10, 15, and 30 min, respectively. (B) Kinetic timecourses for metal uptake based on densitometric analysis of electrophoretic mobility shift data in (A). Normalized intensities for apo-MnSOD (●) and Mn2-MnSOD (■) were fitted to first order kinetic equations using nonlinear regression methods. Theoretical lines represent best-fit simulations with k1= 0.18 ± 0.01 min−1 (apo-MnSOD) and k1=0.2±0.02 min−1 (Mn2-MnSOD). (C) Metal uptake was initiated by addition of 0.4 equivalents of MnCl2 to apo-MnSOD as described in the Materials and Methods (Section 2.14). (1) MnSOD standards; (2) equal amounts of Mn2-MnSOD and Apo-MnSOD; (3–9) incubation time: 0, 1, 5, 10, 15, 30 min and 15 h, respectively.
3.2.1 Metal binding by wt apo-MnSOD
When slightly less than 0.5 equivalent of Mn2+ was added to the apo-protein (Fig. 3C), a small amount of half-metallated dimeric species was observed, but a statistical distribution of all three species was achieved only after overnight incubation of the mixture at 37°C (Fig. 3C, 9). Attempts to purify the half-metallated protein from the reaction mixture by CM-52 ion exchange chromatography were unsuccessful, and only holo-MnSOD (with a distorted elution profile, data not shown) and apo-protein were found. These results indicate that the metal binding reaction of wt apo-MnSOD is highly cooperative and symmetric between the subunits of the dimer.
3.2.2 Subunit exchange
The appearance of half-metallated MnSOD following prolonged incubation of apo-MnSOD with substoichiometric Mn2+ at 37°C could be the result of subunit exchange following metal uptake, or metal mobilization between holo- and apo-protein. In order to distinguish between these two possibilities, a 1:1 mixture of holo- and apo-MnSOD was incubated at 45°C in the presence or absence of EDTA in the reaction buffer (Fig. 4A). Under these conditions, a statistical distribution of the three metalloforms (holo-, half-metallated, and apo-MnSOD, 1:2:1) was reached within 2 hours. The presence of EDTA does not affect the formation of all three metalloforms either at 45°C for 3 hours (Fig. 4A, 11), or overnight at 37°C (Fig. 4A, 10), demonstrating that the formation of the half-metallated species is the result of the exchange of subunits between dimers rather than dissociation and rebinding of the metal ion. Rate constants for the subunit exchange reaction (Equation 9):
| (9) |
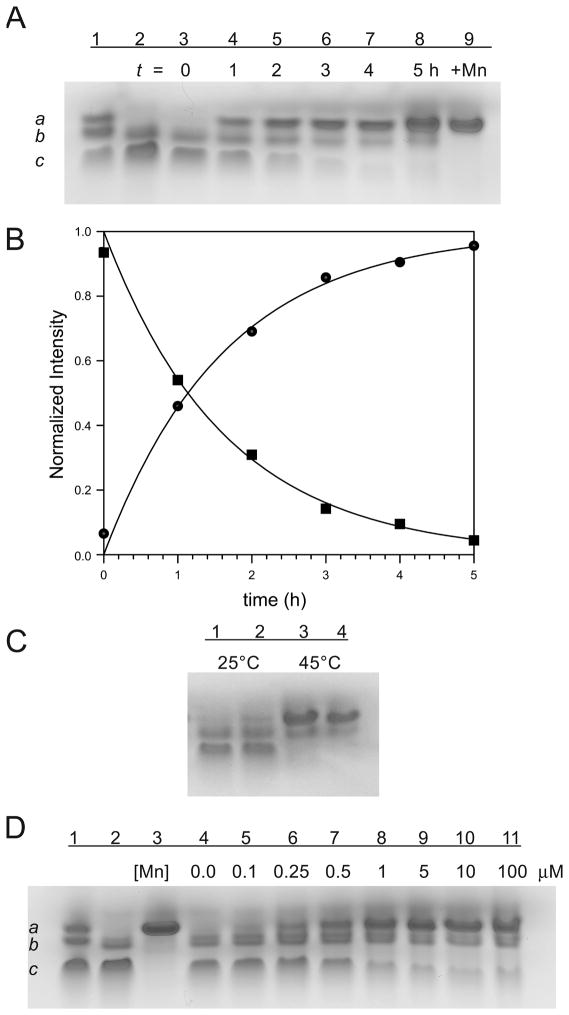
have been estimated by quantitative analysis of the gel data (Fig. 4B) (k1= 1.4 h−1; k2=1.5 h−1). The relatively slow kinetics observed for the subunit exchange process suggests that dissociation of the holo-protein may be rate limiting. Subunit exchange was found to be strongly temperature-dependent (over the range 0–45°C, Fig. 4C, 1–4) but only weakly pH-dependent over the range pH 5.5–8.8 (Fig. 4C, 5–10). Subunit exchange is also inhibited by elevated ionic strength (higher concentrations of sodium chloride or buffers) (Fig. 4D).
Figure 4.
Native PAGE analysis of subunit exchange between Mn2-MnSOD and Apo-MnSOD. MnSOD standards: a, Mn2-MnSOD, b, Mn1-MnSOD, and c Apo-MnSOD. (A) Time dependence of subunit exchange at 45°C. (1) MnSOD standards; (2) Mn2-MnSOD and Apo-MnSOD mixture, initial state (t = 0 min); (3–9) Mn2-MnSOD and Apo-MnSOD mixture incubated at 45°C for: 10 min, 0.5, 1, 1.5, 2, 2.5 3 h, respectively; (10 & 11) reaction in the presence of EDTA: (10) 37°C, 15 h; (11) 45°C, 3 h. (B) Kinetic timecourses for subunit exchange based on densitometric analysis of electrophoretic mobility shift data in (A). Normalized intensities for apo-MnSOD (■), Mn1-MnSOD (●) and Mn2-MnSOD (▲) were globally fit to three differential equations (d[P]/dt = d[PM2]/dt=−k1[P][PM2]+(0.5)k2[PM]2; d[PM2]/dt=(2)k1[P][PM2] −k2[PM2]2) using nonlinear regression methods. Theoretical lines representing best-fit simulations with k1= 1.4 h−1 and k2=1.5 h−1 are shown. (C) Temperature and pH dependence of subunit exchange. (1–4): incubated at 0°C, 37°C, 41°C, and 45°C (20 mM MOPS pH 7.8) for 1 h; (5–10): pH 5.5 (MES), 6.2 (MES), 7.0 (MOPS), 7.8 (MOPS), 8.2 (HEPES), and 8.8 (TAPS) at 45°C for 1 h. (D) Inhibition of subunit exchange by sodium chloride. (1) MnSOD standards; (2–9): 0, 20, 40, 60, 80, 120, 160, and 200 mM sodium chloride added to the reaction mixture (20 mM MOPS pH 7.8) and incubated at 45°C for 1 h.
Subunit exchange during metal uptake was investigated by addition of MnCl2 to a stoichiometric mixture of C-terminal Strep-tag II fusion and untagged wt apo-MnSOD at 37°C (Fig. 5A). The kinetics of metal uptake, monitored fluorimetrically, are the same for C-Strep and wt apo-MnSOD (data not shown). After incubation for 30 min, the mixture was analyzed by native PAGE which showed the formation of a statistical distribution of product metalloforms including C-Strep Mn2-MnSOD, a species with intermediate mobility interpreted as a heterodimeric metallated form containing both C-Strep and wt MnSOD subunits, and wt Mn2-MnSOD (1:2:1) (Fig. 5A, 3). The species with intermediate mobility was also observed in the products resulting from overnight incubation of fully metallated C-Strep Mn2-MnSOD and wt Mn2-MnSOD at 37°C (Fig. 5A, 7), but not at shorter times (30 min, 37°C) (Fig. 5A, 6), suggesting that rapid scrambling of subunits is associated with the metal uptake process.
Figure 5.
Native PAGE analysis of subunit exchange during metal uptake by apo-MnSOD. (A) C-terminal Strep-tag II (C-Strep) MnSOD apo-protein. (1) wt apo-MnSOD; (2) C-Strep apo-MnSOD; (3) products of metal uptake reaction (37°C for 30 min) for a mixture of C-Strep and wt apo-MnSOD; (4) products of metal uptake reaction for wt apo-MnSOD; (5) products of metal uptake reaction for C-Strep apo-MnSOD; (6) mixture of Mn2-C-Strep MnSOD and Mn2-MnSOD incubated at 37°C for 0.5 h; (7) as (6) but incubated 15 h; (8) Mn2-C-Strep MnSOD and Mn2-MnSOD standards. The gel was run at 20 A/gel for 4 h at 0°C. (B) Alexa Fluor 594 and Alexa Fluor 647 labeled Q21C apo-MnSOD. (1) Alexa Fluor 594 apo-MnSOD (Q21C); (2) Alexa Fluor 647 apo-MnSOD (Q21C); (3) Alexa Fluor 594 Mn2-MnSOD (Q21C) after metal uptake reaction (37°C, 10 min); (4) Alexa Fluor 647 Mn2-MnSOD (Q21C) after metal uptake reaction (37°C, 10 min); (5) products of metal uptake reaction (37°C, 10 min) for equal mixture of Alexa Fluor 594 apo-MnSOD (Q21C) and Alexa Fluor 647 apo-MnSOD (Q21C); (6–8) an equal mixture of Alexa Fluor 594 Mn2-MnSOD (Q21C) (3) and Alexa Fluor 647 Mn2-MnSOD (Q21C) (4) incubated at: (6) 0°C, 3 h; (7) 37°C, 10 min; (8) 37°C, 1 h. The gel was run at 20 A/gel for 1.75 h at 0°C. (C) Fluorescence imaging of Alexa Fluor labeled MnSOD (Q21C) performed as described in the Materials and Methods (Section 2.12). (1) Alexa Fluor 594 Mn2-MnSOD (Q21C), (2) Alexa Fluor 647 Mn2-MnSOD (Q21C), (3) an equal mixture of Alexa Fluor 594 Mn2-MnSOD (Q21C) (1) and Alexa Fluor 647 Mn2-MnSOD (Q21C) (2) incubated at 37°C for 1 h. (a) Coomassie Blue stained native PAGE. (b) Native PAGE scanned with Typhoon TRIO+ fluorescence imager, 532 nm excitation, 647 nm emission. (c) 633 nm excitation, 670 nm emission. (d) 532 nm excitation, 670 nm emission. (e) Scan superposition showing FRET for the intermediate mobility species.
Complete subunit exchange during metal uptake was also demonstrated by using fluorescent dye probes to label the protein. The Q21C variant MnSOD, prepared by site-directed mutagenesis, was covalently conjugated with dye maleimides (Alexa Fluor 594 and Alexa Fluor 647) at the unique Cys residue in each subunit, with estimated labeling efficiencies of 99% (for Alexa Fluor 594) and 76% (for Alexa Fluor 647). The rate of metal uptake was approximately 3× faster for the dye-conjugated apo-protein than for the unlabeled apo-protein at 37°C. The different metalloforms of each of the dye conjugates can be resolved by native PAGE, and addition of MnCl2 to a mixture of the dye-labeled apo-proteins results in formation of both homodimeric holo-proteins and a species having intermediate mobility within 10 min at 37°C (Fig. 5B, 5). Simply incubating the two dye-labeled homodimeric holo-proteins, prepared individually, at 37°C for 1 hour also produces the same mixture (Fig. 5B, 8), but only a small amount of the intermediate mobility species was observed after 10 min (Fig. 5B, 7), again suggesting that relatively rapid scrambling of subunits occurs during metal uptake. The identity of the intermediate mobility species as a heterodimeric form containing both dye-labeled subunits was confirmed by detection of fluorescence resonance energy transfer (FRET), using fluorescent imaging (Fig. 5C). The Förster radius for the Alex Fluor 594/647 dye pair (85Å) is large compared to the separation of the dye molecules in the dimeric protein based on the distance between corresponding CD atoms in the Gln21 side chains in the two subunits in the crystal structure for the E. coli MnSOD holo-protein (PDB ID: 1VEW) (11.6 Å), yielding a predicted FRET efficiency of >95%. The homodimeric species containing a single covalently attached dye molecule can be selectively monitored at their respective excitation and emission wavelengths (Fig. 5C, panels b&c). Selective excitation of the donor dye (Alexa Fluor 594) and detection at the emission wavelength of the acceptor dye (Alexa Fluor 647) leads to a clear enhancement of a FRET signal associated with the intermediate mobility band on the native PAGE gel (Fig. 5C, panel e). No subunit exchange was observed in mixtures of either Alexa Fluor labeled apo-proteins or C-Strep/wt apo-MnSOD (data not shown), suggesting that under native PAGE conditions the apo-protein may migrate as a monomeric species.
3.2.3 Metal binding by disulfide crosslinked apo-MnSOD (S126C)(SS)
Subunit exchange during metal uptake may be a consequence of destabilization of the dimeric protein by the exergonic metal complex formation, or it may indicate that protein dissociation precedes metal binding, as illustrated in Scheme 1. Earlier work [11] demonstrated that covalently crosslinking subunits by disulfide bonds does not prevent metal uptake in the apo-protein. We have further investigated the effects of covalent constraints by monitoring the progress of metal uptake by (S126C)(SS) disulfide crosslinked apo-protein using gel electrophoretic mobility shift analysis (EMSA) (Fig. 6). These experiments reveal asymmetric metal binding by the two subunits of the crosslinked dimer, contrasting with the strongly cooperative, symmetric metal binding of the wt apo-protein. Both metal stoichiometry-dependent (0.3–2.9 equiv. MnCl2 in terms of dimeric protein) (Fig. 6A) and time-dependent (0–30 min) (Fig. 6C) metal uptake reactions show initial formation of the half-metallated species, which appears to be fully populated before the appearance of the holo-protein product. However, no half-metallated form was detected when fully metallated (S126C)(SS) MnSOD was incubated with the apo-protein overnight at 37°C, again indicating that the metal binding is essentially irreversible and the metal ion does not dissociate from the protein after it is bound. As expected, half-metallated (S126C)(SS) MnSOD prepared by addition of approximately 0.5 equivalents of MnCl2 to the apo-protein exhibited about half the SOD activity of the fully metallated (S126C)(SS) product (2400±72 U/mg for protein containing 0.41±0.01 Mn/active site vs. 4652±20 U/mg for protein containing 0.81±0.03 Mn/active site), indicating that the two sites function independently in catalysis.
Scheme 1.
Figure 6.
Metal uptake by disulfide-crosslinked apo-MnSOD (S126C). (A) Native PAGE electrophoretic mobility shift results for stoichiometric metal titration of apo-MnSOD (S126C)(SS). (1) apo-MnSOD (S126C)(SS); (2–8), products formed by addition of MnCl2 to apo-MnSOD (S126C)(SS): 0.3, 0.6, 0.8 1.2, 1.7, 2.3, 2.9 equivalents MnCl2 (20 mM MOPS pH 7.8) based on dimeric protein. (B) Quantitative analysis of species formed during stoichiometric titration of apo-MnSOD (S126C)(SS) by MnCl2. Normalized densitometric intensities for apo-MnSOD (S126C)(SS) (P) (■), Mn1-MnSOD (S126C)(SS) (PM) (●) and Mn2-MnSOD (S126C)(SS) (PM2) (▲) from (A) were globally fit to Equations 5–8 (Methods, section 2.17) using nonlinear regression methods. The solid lines are simulations based on the best-fit parameter values (K1=1.6×107 M−1; K2=4.2×1012 M−2; q1=1.5; q2=8). (C) Time-dependent metal uptake. (1) Mn2-, Mn1-, and apo-MnSOD (S126C)(SS) standards; (2–9) incubation of apo-MnSOD (S126C)(SS) with 1.4 equivalents of MnCl2 at 37°C for 0, 1, 3, 5, 7, 10, 15 or 30 min (20 mM MOPS pH 7.8). (D) Quantitative analysis of kinetic intermediates formed during reaction of apo-MnSOD (S126C)(SS) with MnCl2. Normalized densitometric intensities for apo-MnSOD (S126C)(SS) (P) (■), Mn1-MnSOD (S126C)(SS) (PM) (●) and Mn2-MnSOD (S126C)(SS) (PM2) (▲) from (C) were globally fit to three differential equations (d[P]/dt=−k1[P]; d[PM]/dt=k1[P] −k2[PM]; d[PM2]/dt=k2[PM]) using nonlinear regression methods. Theoretical lines representing best-fit simulations with k1= 2.2 min−1 and k2=0.1 min−1 are shown.
Metal binding by MnSOD (S126C)(SS) apo-protein was investigated quantitatively by analyzing the densitometric data for the progress of the stoichiometric titration, as described in the Materials and Methods (2.17). Normalized gel densitometry data from Fig. 6A is plotted vs. the titration progress parameter ([M]0/[P]0) in Fig. 6B. Extensive conversion to the half-apo species (PM), reflected in the peak in [PM]/[P]0 (Θ1) near the mid-point of the titration ([M]0/[P]0= 1), is characteristic of anti-cooperative metal binding by the two subunits of the dimer. The relative magnitude of the peak is related to the ratio of equilibrium constants for the two complexes (Equation 10):
| (10) |
The degree of cooperativity for the metal binding process can be evaluated quantitatively using the parameter α (α = 4K2/(K1)2) to assess interaction between the two sites [41], with cooperativity associated with α > 1 and anti-cooperativity associated with α < 1. The factor of 4 in the definition of α is a statistical weighting factor, and the expression is written in terms of overall association constants rather than stepwise constants. Alternatively, we may define a parameter w (Equation 11):
| (11) |
such that w = 0 corresponds to non-cooperative behavior of two independent, identical binding sites, w = +1 corresponds to fully cooperative behavior, and w = −1 corresponds to the limit of strong anti-cooperativity. Fitting the experimental data to the speciation equations (Equations 5–7, section 2.17)(Fig. 6B) yields estimates of the binding parameters. While the constants (K1,K2) cannot be individually determined by this analysis, the relative values (reflected in the cooperativity parameters α and w) are well defined. Based on the estimates for K1 and K2 obtained by global fitting of the progress curves (Fig. 6B), α = 0.07 and w = −0.9 for metal binding by MnSOD (S126C)(SS) apo-protein, in the limit of strong anti-cooperativity. Thus, metal binding appears to be mechanistically altered by the disulfide crosslink, which prevents complete separation of subunits but does not block metal uptake (Scheme 2). The presence of the crosslink leads to anti-cooperative, asymmetric metal binding by the subunits of the homodimeric protein, with the disulfide crosslink acting as a symmetry-breaking perturbation, removing the intrinsic degeneracy of the symmetric protein dimer.
Scheme 2.
3.3 Protein concentration dependence of metal uptake
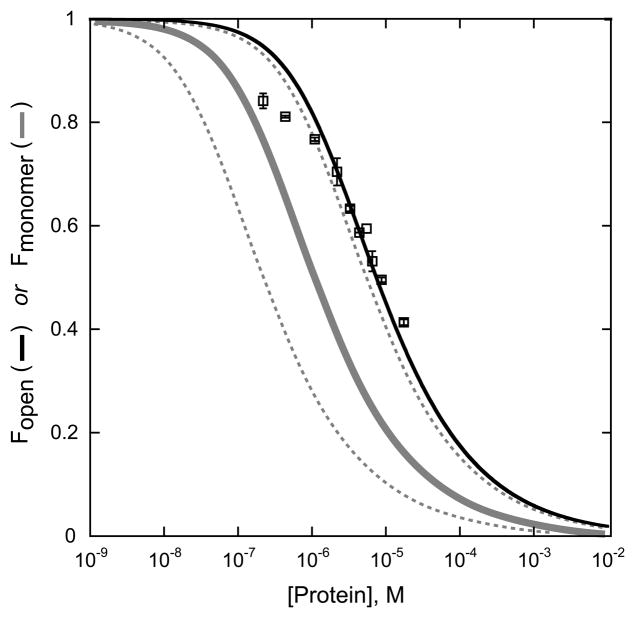
The symmetric metal uptake process observed for wt apo-MnSOD and the exchange of subunits between holo- and apo-protein dimers suggest that dissociation of the dimeric apo-MnSOD plays a significant role in the metal uptake reaction. Previous analysis of metal binding by E. coli apo-MnSOD revealed complex kinetics comprising two exponential phases (fast and slow phases) associated with the metal uptake mechanism. This type of kinetic behavior is consistent with conformational gating of metal uptake, in which a structural reorganization of the protein is required for the metal to bind in the buried active site. The amplitude of the fast phase reflects the fraction of apo-protein in the “open” conformation that is able to bind metal ion readily, and the slow phase reflects the rate of interconversion of the two forms. The open fraction (Fopen) has been shown to increase with increasing temperature and pH. If the conformational change associated with conversion to the open form involves subunit dissociation, the metal uptake kinetics should be sensitive to protein concentration. The dependence of the amplitude of Fopen on total protein concentration is shown in Fig. 7. Fopen clearly decreases with increasing protein concentration, although experimental constraints limit the range of protein concentration that can be investigated. The protein concentration dependence of Fopen can be modeled in terms of a subunit dissociation model for conformational gating, interpreting the open state as the dissociated monomeric species (Fig. 7). The standard protein dimer equilibrium constant KD is expressed in terms of monomer and dimer concentrations (Equation 12)
Figure 7.
Protein concentration dependence of metal uptake kinetics. Metal uptake was monitored fluorimetrically (20 mM MOPS pH 7.6, 45°C) as described in the Material and Methods (Section 2.14). The ratio of fast phase amplitude to total amplitude (Fopen) was plotted against the total protein concentration (subunits) and the results analyzed in terms of equilibrium dissociation of the dimeric protein, as described in the Results (Section 3.3). ( ) theoretical curve for Fopen, KD = (6.6±0.9)×10−6 M. (
) theoretical curve for Fopen, KD = (6.6±0.9)×10−6 M. ( ) theoretical curve for KD(SEC) = 1.1×10−6 M. (
) theoretical curve for KD(SEC) = 1.1×10−6 M. ( )) bounding limits for 5-fold variation of KD(SEC).
)) bounding limits for 5-fold variation of KD(SEC).
| (12) |
where [M] and [D] are the concentrations of monomeric and dimeric species, and KD is the dissociation constant for the dimeric protein. Expressing the dimer concentration in terms of the total protein ([D] = ([P]tot − [M])/2) and rearranging yields a quadratic equation that may be solved for [M] (Equation 13):
| (13) |
Fitting the experimental protein concentration dependence with this expression yields an estimate of the protein dimer dissociation constant, KD = 6.6±0.9 μM. The experimental data appear to deviate from the theoretical prediction at the lowest protein concentrations, approaching a maximum value of Fopen near 0.9. This maximum value is sensitive to temperature and pH, possibly indicating that other processes are involved in conformational gating of metal uptake in addition to protein dissociation.
3.4 Protein concentration dependent SEC
Size exclusion chromatography was used to directly investigate the protein concentration dependence of the E. coli apo-MnSOD quaternary structure. Earlier gel filtration studies [11] have shown that the protein behaves like a dimeric species under metal uptake conditions (45°C, pH 7.5). However, in those studies the protein concentration used for gel filtration (~1 mg/mL) was approximately 20× higher than the concentration used in the fluorimetric metal uptake measurements (50 μg/mL). In order to investigate the protein concentration dependence of the quaternary structure, a series of SEC experiments were performed with varying amounts of apo-MnSOD under metal uptake conditions. At the lower range of protein concentration, protein fluorescence measurements were required to detect the protein elution profile. Over the entire range of protein concentrations (total protein loaded: 35 μg–0.70 mg) the protein eluted as a single band, which broadened and shifted to larger elution volume (Ve) as the amount of protein loaded was decreased (Fig. 8). This behavior is consistent with progressive dissociation of the dimeric apo-protein at low protein concentration together with rapid reassociation, resulting in a fast exchange limit for SEC separation. In contrast, under the same conditions the fully metallated Mn2-MnSOD holo-protein at the lowest concentration (35 μg total protein loaded) only started to show a separation into two peaks (data not shown). Also, at lower temperature (25°C) where metal uptake is very slow, even dilute apo-MnSOD (35 ug protein loaded) elutes as a dimer, again identifying the dimeric structure with the closed form of the protein.
Figure 8.
Protein concentration dependence of size exclusion chromatography. Samples were loaded onto a 1.5×99 cm column of Sephacryl 100-HR equilibrated and run at 45°C in 20 mM MOPS pH 7.6 containing 1 mM EDTA. Elution profiles were monitored by absorption (O.D.280 nm) or fluorescenc e (λEX = 280 nm; λEM = 333 nm) and normalized for analysis. (○—○) Mn2-MnSOD, 1 mg; (□—□) Apo-MnSOD, 700 μg; (△—△) Apo-MnSOD, 350 μg; (◇—◇) Apo-MnSOD, 175 μg; (▽—▽) Apo-MnSOD, 87.5 μg; (■—■) Apo-MnSOD, 35 μg; (●—●) Fe2-MnSOD (E170A), 1 mg. (----) Theoretical elution profiles for all samples generated by SCIMMS simulation program using the following input parameters: diffusion coefficient (percent equilibration/cycle): monomer, 0.03; dimer, 0.015; dispersion coefficient (gaussian spread/translation): monomer, 0.142; dimer, 0.130. This analysis yields an estimate of the protein dimer dissociation constant KD(SEC) = 1.1×10−6 M.
The conditions for these SEC experiments represent the small zone filtration limit, where a stable equilibrium composition is never attained during chromatographic elution. Under these conditions, it is not possible to relate the peak positions to the protein dimer dissociation constant (KD) by simply interpreting peak positions as an average molecular mass [32]. However, the transport processes (diffusion, dispersion, migration) involved in the chromatographic separation may be modeled computationally to estimate the KD. Experimental elution profiles for apo-MnSOD were simulated using the program SCIMMS [32,33], as illustrated in Fig. 8. The limiting dimeric and monomeric species were modeled using the elution profiles of E. coli Mn2-MnSOD holo-protein and E. coli Fe2-MnSOD (E170A), which are known to exist as dimers and monomers, respectively, under these conditions. Based on this analysis, a value of KD = 1×10−6 M has been estimated for equilibrium dissociation of apo-MnSOD under metal uptake conditions. The predicted distribution between monomeric and dimeric forms over the range of protein concentrations used in the kinetic measurements is shown in Fig. 7. The 5-fold difference between the KD estimates based on SEC and metal uptake kinetics may reflect uncertainties inherent in the analysis of the SEC data [32,33], or may reflect destabilization of the dimer in the presence of the metal ion in the reaction mixture.
3.5 In vivo metal uptake by apo-MnSOD
The in vivo metal uptake behavior of E. coli apo-MnSOD has been investigated as a test of the biological relevance of the in vitro results. An engineered sodA-knockout strain of E. coli (BW25113 ΔsodA) lacking endogenous MnSOD was transformed with the pBAD2sodA expression vector, which permits tightly regulated high-level homologous expression of recombinant MnSOD protein under control of the PBAD arabinose-sensitive promoter [11,21]. When the strain was cultivated in chemically defined growth medium lacking manganese but supplemented with a small amount of iron (100 nM), then transferred to medium lacking both iron and manganese for induction, the cells produced significant amounts of MnSOD protein, mainly as apo-protein but with small amounts of half-metallated MnSOD (Fig. 9A, 2). Further decreasing the amount of iron in the growth medium reduced the total amount of recombinant protein but did not eliminate the minor species. Metal uptake by apo-MnSOD inside the E. coli cells was monitored by native PAGE following addition of MnCl2 to the cells after arresting de novo protein biosynthesis by addition of chloramphenicol (to 50 μg/mL). The time-dependent metal uptake is shown in Fig. 9A. At 35°C, the majority of the intracellular apo-MnSOD is converted to Mn2-MnSOD within 3 hours. A zero-time control was used to test whether metal uptake occurred during sample processing. The lack of conversion of the zero-time control sample (Fig. 9A, 3) demonstrates that the observed metal uptake occurred inside the E. coli cells, and not during the subsequent processing steps. Quantitative analysis of the kinetics of metal uptake in vivo (Fig. 9B) yields a pseudo first order rate constant k=0.01 min−1 at 35°C, approximately 8-fold lower than previously observed for in vitro metal binding at this temperature (k=0.08 min−1) [11]. The slower kinetics of metal uptake in vivo may be the result of rate-limiting metal transport, or macromolecular crowding, since in vitro metal uptake was performed at 1.1 μM apoprotein, nearly 100-fold lower than the apo-MnSOD concentration present in vivo in our experiments (see below). In vivo metal uptake was temperature-dependent. At 45°C all of the apo-MnSOD was metallated within 1 hour (Fig. 9C, 3&4), while at 25°C only a small fraction was converted to the metallated form even after 3 hours (Fig. 9C, 1&2). The dependence of in vivo metal uptake on exogenous manganese concentration at 35°C is shown in Fig. 9D and in Table 2. At the lower concentrations of manganese, almost all of the metal added to the medium was incorporated into the apo-MnSOD, as estimated by the SOD activity measured in the cell extracts (Table 2). The same results are observed at 45°C (data not shown). Scaled-up cultivation of metal-limited E. coli allowed isolation of 17 mg of purified apo-MnSOD from 500 mL of induction medium, in a cell mass (3.9 g) concentrated from 2 L of growth medium. This represents an intracellular apo-MnSOD concentration of approximately 100 μM. The apo-MnSOD isolated directly from the cells exhibited the same metal uptake kinetic behavior in the in vitro fluorimetric metal uptake assay as previously observed for apo-MnSOD prepared by denaturation and refolding from holo-MnSOD (data not shown). The holoenzyme prepared by reconstituting this apo-MnSOD with manganese has the same high specific activity (7235±254 U/mg; 1.00±0.04 Mn/active site) as Mn2-MnSOD isolated directly from E. coli [15].
Figure 9.
In vivo metal uptake by apo-MnSOD. (A) Native PAGE electrophoretic mobility shift results for time-dependent Mn incorporation into apo-protein in E. coli after addition of MnCl2 to the induction medium. Reaction conditions as described in the Materials and Methods (Section 2.8). (1) top to bottom: Mn2-MnSOD, Fe2-MnSOD, Apo-MnSOD standards; (2) induction control with no chloramphenicol or MnCl2 added, cells collected after 4 hours induction; (3) cells collected immediately after addition of MnCl2 to 100 μM; (4–8) incubation at 35°C following addition of MnCl2; (4) 1 h; (5) 2 h; (6) 3 h; (7) 4 h; (8) 5 h; (9) 100 μM MnCl2 was added during induction and cells were collected after 4 h induction. (B) Quantitative analysis of in vivo metal uptake kinetics. Normalized densitometric intensities for apo-MnSOD (P) (■) and Mn2-MnSOD (PM2) (●) from (A) were globally fit to two differential equations (d[P]/dt=−k1[P]; d[PM2]/dt=k1[P]) using nonlinear regression methods. Theoretical lines representing best-fit simulations with k1= 0.6 h−1 are shown. (C) Temperature dependence of Mn uptake by apo-MnSOD following addition of Mn to the induction medium. (1) 25°C 1 h; (2) 25°C 3h; (3) 45°C 1 h; (4) 45°C 3 h. (D) Manganese concentration dependence of in vivo metal uptake by apo-MnSOD. (1) Mn2-MnSOD, Fe2-MnSOD, Apo-MnSOD standards; (2) 4 h induction with no MnCl2 added; (3) induction in presence of 100 μM MnCl2; (4–11) MnCl2 added 10 min after arresting de novo protein biosynthesis with chloramphenicol, and incubation continued at 35°C for 4 h: (4) 0 μM MnCl2; (5) 0.1 μM MnCl2; (6) 0.25 μM MnCl2; (7) 0.5 μM MnCl2; (8) 1 μM MnCl2; (9) 5 μM MnCl2; (10) 10 μM MnCl2; (11) 100 μM MnCl2.
Table 2.
Superoxide dismutase in vivo metallation
| sample | Culture conditionsa | SOD Activity (U/mL)d | [MnSOD] (μM active sites)e | ||
|---|---|---|---|---|---|
| Inductionb | Post-Inductionc | ||||
| Mn | Cm | [Mn], (μM) | |||
| 1 | − | − | 0.0 | 5.3±0.4 | – |
| 2 | + | − | 0.0 | 486.6±1.7 | – |
| 3 | − | + | 0.0 | 4.7±0.1 | – |
| 4 | − | + | 0.1 | 22.1±0.5 | 0.10±0.002 |
| 5 | − | + | 0.25 | 45.2±0.5 | 0.24±0.003 |
| 6 | − | + | 0.5 | 93.8±0.5 | 0.53±0.003 |
| 7 | − | + | 1 | 149.2±7.2 | 0.86±0.04 |
| 8 | − | + | 5 | 188.6±1.6 | 1.09±0.01 |
| 9 | − | + | 10 | 201.7±2.9 | 1.17±0.02 |
| 10 | − | + | 100 | 196.1±5.5 | 1.14±0.03 |
E. coli BW25113 ΔsodA | pBAD2sodA was grown in modified MOPS medium containing 0.2% glucose as described in the Materials and Methods (Section 2.8).
Incubated at 35°C for 4 hours in 2 mL modified MOPS medium containing 0.2% glycerol and 0.05% L-(+)-arabinose as described in the Materials and Methods (Section 2.8).
Incubated at 35°C for 4 hours following addition of MnCl2.
SOD activity in 1 mL of medium measured using the xanthine oxidase/cytochrome c inhibition assay as described in the Materials and Methods (Section 2.16).
Calculated from experimental SOD activity after subtraction of background activity (average of Samples 1&3), based on a specific activity of 7300 U/mg for fully metallated Mn2-MnSOD [15].
4. Conclusions
The structure of E. coli MnSOD appears to be essentially independent of the presence or absence of the metal cofactor (Fig. 2). This clearly demonstrates that the MnSOD protein templates metal coordination, and that the unusual trigonal bipyramidal geometry of the Mn complex is intrinsically defined by the protein structure, not by interactions with the metallocofactor. This “entatic state” [42] control of the metal ion environment is likely to be important in facilitating rapid electron transfer events during the enzymatic dismutation cycle. The constrained metal coordination geometry may be expected to lower the reorganization barrier for oxidation-reduction reactions at this site. The shell of amino acid side chains that surround and coordinate the metal ion occupy virtually the same positions (0.19–0.35Å RMSD over all atoms) in apo-MnSOD as in the holo-protein complex, and share the same side-chain conformations in both structures. As a result, access to the metal binding site is severely restricted in the conformational state represented by this structure, as there is no channel to the protein surface large enough to allow entry of a metal ion to the buried metal binding site. The structure thus appears to correspond to the “closed” conformational state, in spite of crystallization under conditions that would tend to favor stabilization of the “open” form of the apo-protein. It also implies that protein dynamics must be a part of metal binding.
Electrophoretic analysis of mixtures of MnSOD metalloforms has detected a previously unrecognized subunit exchange process, driven by dissociation of the dimeric protein. Complete subunit exchange is reached after prolonged incubation of a mixture of holo- and apo-protein at elevated temperature (Figs. 3 & 4), or rapidly in the course of metal binding (Fig. 5). The relatively rapid subunit exchange observed during the metal uptake reaction may indicate that the dimeric structure is destabilized by the exergonic metal uptake reaction, or it may reflect dissociation of the dimeric apo-proteins. The protein concentration dependence of metal binding kinetics is consistent with a metal binding mechanism involving subunit dissociation (KD = 6×10−6 M) (Fig. 7), although other factors may also contribute to conformational gating of metal uptake. Small-zone SEC analysis of the stability of the apo-MnSOD dimer yields a similar estimate of KD(SEC) = 1×10−6 M (Fig. 8). This connection between the physical process of subunit dissociation and conformational gating of metal uptake underscores the importance of the subunit interface region on metal binding by the apo-protein. The observation of subunit exchange during metal uptake, and the highly cooperative, symmetric character of the metal uptake reaction are also consistent with a role for subunit dissociation in metal binding by wt E. coli apo-MnSOD. Subunit separation may be expected to facilitate reorientation of active site residues that appears to be required to create an access channel allowing entry of the metal ion into the buried binding site [11]. A covalent crosslink between the two subunits, which prevents subunit dissociation, does not block metal uptake, showing that complete dissociation of the dimeric protein is not required for metal binding to occur, as previously observed. However, the presence of the crosslink significantly perturbs the metal binding process, resulting in highly anti-cooperative metal uptake between the two subunits of the dimer. Thus, although the cross-linked protein is capable of binding metal ions, the mechanism of metal uptake appears to be fundamentally altered. While these results confirm earlier studies showing that subunit dissociation is not absolutely required for metal binding by apo-MnSOD [11], they also demonstrate that subunit dissociation is an essential feature of the binding process for wt apo-protein.
Purified apo-MnSOD produced in vivo is indistinguishable from that prepared from the metallated protein in vitro by denaturation, chelation and refolding. The folded, soluble apo-MnSOD produced in vivo under metal-limited growth conditions can also pick up its metal cofactor inside the E. coli cell when the cells are incubated with exogenous metal ions under conditions similar to those characteristic of in vitro metal uptake of the apo-protein (Fig. 9). At low concentration of manganese (0.1–0.5 μM) virtually all of the metal added to the extracellular medium was incorporated into Mn2-MnSOD, demonstrating the remarkable efficiency of E. coli in concentrating the dilute exogenous metal ion. The similarity of behavior of apo-MnSOD inside and out of the cell demonstrates the relevance of the in vitro studies to metal binding in vivo. In vitro experiments linking metal uptake with MnSOD dimer dissociation suggest that metal delivery occurs to the monomeric protein in vivo, as well.
Acknowledgments
The authors would like to thank Jay Nix at the Advanced Light Source beam line 4.2.2. for crystallographic data collection and X-ray fluorescence scans of apo-MnSOD crystals. Support for this work from the National Institutes of Health (R01 GM042680 to J.W.W. and R01 GM077543 to M.S.C.) and the American Heart Association (10 POST 2600203 to T.F.L.) is gratefully acknowledged.
ABBREVIATIONS
- SOD
superoxide dismutase
- SEC
size exclusion chromatography
- FRET
fluorescence resonance energy transfer
- PAGE
polyacrylamide gel electrophoresis
- Cm
chloramphenicol
- wt
wild type
- RMSD
root mean square deviation
- GdnHCl
guanidinium hydrochloride
- 2-mercaptoethanol
2-ME
- a.s.u.
asymmetric unit
- EMSA
electrophoretic mobility shift analysis
Footnotes
Accession Numbers
The structure factors and coordinates of Escherichia coli apo-MnSOD have been deposited to the RCSB Protein Data Bank with the PDB ID code 3OT7.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.McCord JM. Oxygen-derived free radicals. New Horiz. 1993;1:70–76. [PubMed] [Google Scholar]
- 2.Fridovich I. Superoxide radical and superoxide dismutases. Annu Rev Biochem. 1995;64:97–112. doi: 10.1146/annurev.bi.64.070195.000525. [DOI] [PubMed] [Google Scholar]
- 3.Tainer JA, Getzoff ED, Beem KM, Richardson JS, Richardson DC. Determination and analysis of the 2 Å-structure of copper, zinc superoxide dismutase. J Mol Biol. 1982;160:181–217. doi: 10.1016/0022-2836(82)90174-7. [DOI] [PubMed] [Google Scholar]
- 4.Lah MS, Dixon MM, Pattridge KA, Stallings WC, Fee JA, Ludwig ML. Structure-function in Escherichia coli iron superoxide dismutase: comparisons with the manganese enzyme from Thermus thermophilus. Biochemistry. 1995;34:1646–1660. doi: 10.1021/bi00005a021. [DOI] [PubMed] [Google Scholar]
- 5.Edwards RA, Baker HM, Whittaker MM, Whittaker JW, Jameson GB, Baker EN. Crystal structure of Escherichia coli manganese superoxide dismutase at 2.1 Å resolution. J Biol Inorg Chem. 1998;3:161–171. [Google Scholar]
- 6.Wuerges J, Lee JW, Yim YI, Yim HS, Kang SO, Djinovic Carugo K. Crystal structure of nickel-containing superoxide dismutase reveals another type of active site. Proc Natl Acad Sci USA. 2004;101:8569–8574. doi: 10.1073/pnas.0308514101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rosenzweig AC. Metallochaperones: bind and deliver. Chem Biol. 2002;9:673–677. doi: 10.1016/s1074-5521(02)00156-4. [DOI] [PubMed] [Google Scholar]
- 8.Culotta VC, Yang M, O’Halloran TV. Activation of superoxide dismutases: putting the metal to the pedal. Biochim Biophys Acta. 2006;1763:747–758. doi: 10.1016/j.bbamcr.2006.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mizuno K, Whittaker MM, Bächinger HP, Whittaker JW. Calorimetric studies on the tight binding metal interactions of Escherichia coli manganese superoxide dismutase. J Biol Chem. 2004;279:27339–27344. doi: 10.1074/jbc.M400813200. [DOI] [PubMed] [Google Scholar]
- 10.Whittaker MM, Mizuno K, Bächinger HP, Whittaker JW. Kinetic analysis of the metal binding mechanism of Escherichia coli manganese superoxide dismutase. Biophys J. 2006;90:598–607. doi: 10.1529/biophysj.105.071308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Whittaker MM, Whittaker JW. Conformationally gated metal uptake by apo- manganese superoxide dismutase. Biochemistry. 2008;47:11625–11636. doi: 10.1021/bi8015636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. UCSF Chimera - A Visualization System for Exploratory Research and Analysis. J Comput Chem. 2004;25:1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- 13.Ose DE, Fridovich I. Manganese-containing superoxide dismutase from Escherichia coli: reversible resolution and metal replacements. Arch Biochem Biophys. 1979;194:360–364. doi: 10.1016/0003-9861(79)90628-3. [DOI] [PubMed] [Google Scholar]
- 14.Beyer WF, Jr, Fridovich I. In vivo competition between iron and manganese for occupancy of the active site region of the manganese-superoxide dismutase of Escherichia coli. J Biol Chem. 1991;266:303–308. [PubMed] [Google Scholar]
- 15.Whittaker MM, Whittaker JW. Mutagenesis of a proton linkage pathway in Escherichia coli manganese superoxide dismutase. Biochemistry. 1997;36:8923–31. doi: 10.1021/bi9704212. [DOI] [PubMed] [Google Scholar]
- 16.Whittaker JW. The irony of manganese superoxide dismutase. Biochem Soc Trans. 2003;31:1318–1321. doi: 10.1042/bst0311318. [DOI] [PubMed] [Google Scholar]
- 17.Neidhardt FC, Bloch PL, Smith DF. Culture medium for enterobacteria. J Bacteriology. 1974;119:736–747. doi: 10.1128/jb.119.3.736-747.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Miller EM, Nickoloff JA. Escherichia coli electrotransformation. Methods Mol Biol. 1995;47:105–113. doi: 10.1385/0-89603-310-4:105. [DOI] [PubMed] [Google Scholar]
- 19.Datsenko KA, Wanner BL. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA. 2000;97:6640–6645. doi: 10.1073/pnas.120163297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu P, Jenkins NA, Copeland NG. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res. 2003;13:476–484. doi: 10.1101/gr.749203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Guzman LM, Belin D, Carson MJ, Beckwith J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose pBAD promoter. J Bacteriol. 1995;177:4121–4130. doi: 10.1128/jb.177.14.4121-4130.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Schmidt TGM, Skerra A. The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nature Protocols. 2007;2:1528–1535. doi: 10.1038/nprot.2007.209. [DOI] [PubMed] [Google Scholar]
- 23.Whittaker MM, Whittaker JW. Recombinant superoxide dismutase from a hyperthermophilic archaeon, Pyrobaculum aerophilum. J Biol Inorg Chem. 2000;5:402–408. [PubMed] [Google Scholar]
- 24.Gao B, Flores SC, Bose SK, McCord JM. A novel Escherichia coli vector for oxygen-inducible high level expression of foreign genes. Gene. 1996;176:269–270. doi: 10.1016/0378-1119(96)00246-6. [DOI] [PubMed] [Google Scholar]
- 25.Whittaker MM, Whittaker JW. A glutamate bridge is essential for dimer stability and metal selectivity in manganese superoxide dismutase. J Biol Chem. 1998;273:22188–22193. doi: 10.1074/jbc.273.35.22188. [DOI] [PubMed] [Google Scholar]
- 26.Beyer WF, Jr, Reynolds JA, Fridovich I. Differences between the manganese- and the iron-containing superoxide dismutases of Escherichia coli detected through sedimentation equilibrium, hydrodynamic, and spectroscopic studies. Biochemistry. 1989;28:4403–4409. doi: 10.1021/bi00436a042. [DOI] [PubMed] [Google Scholar]
- 27.Otwinowski Z, Minor W. Processing of X-Ray Diffraction Data Collected in Oscillation Mode. Methods in Enzymology. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 28.Praaenikar J, Afonine PV, Guncar G, Adams PD, Turk D. Averaged kick maps: less noise, more signal... and probably less bias. Acta Crystallogr D Biol Crystallogr. 2009;65:921–931. doi: 10.1107/S0907444909021933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Adams PD, Grosse-Kunstleve RW, Hung LW, Ioerger TR, McCoy AJ, Moriarty NW, Read RJ, Sacchettini JC, Sauter NK, Terwilliger TC. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D Biol Crystallogr. 2002;58:1948–1954. doi: 10.1107/s0907444902016657. [DOI] [PubMed] [Google Scholar]
- 30.Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 31.Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst. 1993;26:283–291. [Google Scholar]
- 32.Stevens FJ, Schiffer M. Computer simulation of protein self-association during small-zone filtration. Biochem J. 1981;195:213–219. doi: 10.1042/bj1950213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wilton R, Myatt EA, Stevens FJ. Analysis of protein-protein interactions by simulation of small-zone gel filtration chromatography. Methods Molec Biol. 2004;261:137–154. doi: 10.1385/1-59259-762-9:137. [DOI] [PubMed] [Google Scholar]
- 34.W.S. Rasband. ImageJ. U.S. National Institutes of Health; Bethesda, Maryland, USA: pp. 1997–2009. http://rsb.info.nih.gov/ij/ [Google Scholar]
- 35.Abramoff MD, Magelhaes PJ, Ram SJ. Image processing with ImageJ. Biophotonics Int. 2004;11:36–42. [Google Scholar]
- 36.Senear DF, Brenowitz M. Determination of binding constants for cooperative site-specific protein-DNA interactions using the gel mobility shift assay. J Biol Chem. 1991;266:13661–13671. [PubMed] [Google Scholar]
- 37.Lowry OH, Rosebrough NJ, Farr AL, Randall RJ. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951;193:265–275. [PubMed] [Google Scholar]
- 38.McCord JM, Fridovich I. Superoxide dismutase. An enzymic function for erythrocuprein (hemocuprein) J Biol Chem. 1969;244:6049–6055. [PubMed] [Google Scholar]
- 39.Krissinel E, Henrick K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr D Biol Crystallogr. 2004;60:2256–2268. doi: 10.1107/S0907444904026460. [DOI] [PubMed] [Google Scholar]
- 40.The PyMOL Molecular Graphics System. San Carlos, CA, USA: DeLano Scientific; [Google Scholar]
- 41.Conners KA. Binding Constants: The Measurement of Molecular Complex Stability. Wiley Interscience; New York: 1987. [Google Scholar]
- 42.Frey PA, Hegeman AD. Enzymatic Reaction Mechanisms. Oxford University Press; Oxford: 2006. [Google Scholar]