Abstract
Protein interaction reporter (PIR) technology can enable identification of in vivo protein interactions with the use of specialized chemical cross-linkers, liquid chromatography, and high-resolution mass spectrometry. PIR-cross-linkers contain labile bonds that are specifically fragmented under low energy collision or photodissociation conditions in the mass spectrometer source, thus releasing cross-linked peptides. Successful analysis of PIR-cross-linked proteins requires the use of expected mathematical relationships between cross-linked complexes released peptides after fragmentation of the labile PIR bonds. Presented here is a next-generation software tool, BLinks, for use in the analysis and identification of PIR-cross-linked proteins. BLinks is an advancement beyond our previous efforts by incorporation of chromatographic profiles that must match between cross-linked complexes and released peptides to enable estimation of p values to help filter true relationships from complex datasets. Additionally, BLinks was used to incorporate Mascot database searching results from subsequent MS/MS analysis of the released peptides to facilitate identification of cross-linked proteins. BLinks was used in the analysis of human serum albumin, and 46 inter-peptide relationships were found spanning thirty proximal residues with a 2.2% false discovery rate. BLinks was also used to track peptides involved in multiple, co-eluting relationships that make accurate identification of protein interactions difficult. An additional 10 inter-peptide relationships were identified despite poor correlation using the profiling tools provided with BLinks. Additionally, BLinks can be used to globally map all inter-peptide relationships from the data analysis and customize subsequent analysis to target specific peptides of interest, thus making it a useful tool for both discovery of protein interactions and mapping protein topology.
Introduction
Protein-protein interactions have been studied using many different technologies that include the yeast two-hybrid system1, tagged protein co-immuniprecipitation2,3, protein microarrays4,5, and most recently chemical cross-linking combined with mass spectrometry.6 Protein interaction reporters (PIRs) are a novel type of chemical cross-linker that are useful for identifying protein-protein interactions, particularly for proteins within their native environment.7,8 PIRs are membrane-permeable and capable of cross-linking proteins in vivo across the exposed lysine residues of interacting domains. Cross-linked proteins are captured by affinity purification with a tag included in the PIR technology, enzymatically digested to peptides, and analyzed by reversed-phase liquid chromatography (RPLC) with a Fourier-transform mass analyzer. Key to the design of the PIRs are labile bonds that release the cross-linked peptides specifically within the ion source after they are separated chromatographically. The released peptide ions can then be identified by tandem mass spectrometry (MS/MS) techniques.
The controllable cleavage of PIR labile bonds allows linked peptides to be observed as intact structures or as individual peptide ions. Cleavage is performed with low energy collisions in the ion source that results in PIR bond dissociation while leaving the peptide bonds intact. By alternating scans without and with in-source collisional activation, inter-cross-linked peptides can be observed linked together and individually. It is possible to infer inter-cross-link interactions by accurate mass from the mathematical relationship of two released peptide masses to their intact mass in the preceding scan. A previous study9 showed the feasibility of identifying inter-peptide relationships using computational methods. The employed software called X-links used the mathematical relationships between the ions of any two consecutive spectra to enable cross-linked peptide relationship identification. Additionally, X-links provided a set of visual tools to aid the user in the evaluation of the results, including a chromatographic histogram of the ions in a relationship, and a list of candidate peptide sequences obtained by accurate mass from a tryptic peptide database for the organism of study.
Despite the availability of existing computational tools, the analysis of PIR-linked proteins in complex biological samples is tedious. Anderson et al. showed that as the number of ions in the fragmented scans increases, so does the rate of false discovery9. Currently, inter-cross-linked relationships are counted on a scan-by-scan basis so that the number of possible relationships to be validated in a complex sample is compounded by redundancy. Furthermore, peptide sequence identification is difficult when using accurate mass with a large protein database from multi-cellular organisms.
Presented here is an algorithm and software tool, BLinks, which is used to facilitate identification of inter-cross-link relationships by providing and then extending the capabilities of X-links for the analysis of complex biological samples. Blinks is used to identify inter-cross-link species by mass relationships and chromatographically, not just on a scan-by-scan basis. This step dramatically reduces the number of inter-cross-linked peptides that need to be validated. In addition, because relationships are tracked chromatographically, it is possible to evaluate them statistically, and provide an automated test for validation to reduce the number of false discoveries. Finally, BLinks is used to incorporate peptide sequence information from Mascot database searches of MS/MS spectra.
Identification of inter-cross-linked relationships can be used to infer protein-protein interactions when the peptides arise from different proteins. However, the information is still useful if the two interacting peptides come from the same protein. Because of the difficulty in obtaining accurate crystal structures for many proteins, chemical cross-linking technologies, including PIR technology, can be used to identify proximal lysine residues within a protein in vivo. Such information is useful in determining the protein folding topology of both proteins and protein complexes10.
Materials & Methods
Cross-linker
The BRink PIR cross-linker was synthesized in-house using an Aapptec Endeavor 90 peptide synthesizer and FMOC chemistry. Biotinylated lysine was first coupled to glycine. A second lysine was then coupled to provide a branch point for the coupling of two Rink groups. Succinyl anhydride was then coupled to each Rink group and the cross-linker stored at −80°C. Prior to use the BRink carboxylate groups were activated by forming N-hydroxysuccinimide (NHS) esters using the TFA-NHS synthesis route.11
Sample preparation
PIR cross-linking was performed on 1.0 mg/mL human serum albumin (HSA, Sigma-Aldrich) in 20 mM HEPES buffer using BRink. BRink was added to the HSA solution for a final cross-linker concentration of 1 mM, and allowed to react for 30 minutes at room temperature. The sample was reduced with dithiolthreitol (DTT) and alkylated with iodoacetamide (IAA). Removal of excess cross-linker was performed using filter aided sample preparation (FASP)12, and the HSA collected in PBS by reversing the flow of buffer through the filter. PIR-labeled HSA was then captured on monomeric avidin beads (Thermo Fisher Scientific). Digestion with trypsin was done directly on the beads for 2 hours at 37 °C. A second avidin capture step was performed following digestion. The cross-linked peptides were then eluted from the beads using 70% acetonitrile and 0.5% TFA buffer. The peptides were concentrated using a speed-vacuum and resuspended in water with 0.1% formic acid. The peptides were then analyzed by RPLC on an LTQ-FTICR mass analyzer (Thermo Fisher Scientific).
Data Acquisition
The PIR-cross-linked HSA digest was loaded from the autosampler onto a 75 micron inner diameter fused-silica capillary column packed with 20 cm of Magic Beads (Michrom Bioresources, Inc.). The column was mounted on an in-house constructed nanospray source and high pressure liquid chromatography (HPLC) was performed using a Waters NanoAcquity system. A binary mobile phase gradient was used to elute the peptides. Mobile phase component A consisted of water with 0.1% formic acid. Mobile phase component B contained acetonitrile and 0.1% formic acid. The gradient program consisted of four steps: 1) Peptide elution from 5% to 15% solvent B for 10 minutes, 2) Peptide elution from 15% to 40% B for 120 minutes, 3) Column wash at 80% r B for 20 minutes, and 4) Column re-equilibration at 5% B for 30 minutes.
Ion analysis was performed using a LTQ-FT Ultra hybrid mass spectrometer (Thermo Fisher Scientific). Two methods were used to acquire the data for BLinks software analysis. The first method consisted of two alternating scans that allowed acquisition of spectra in the ICR cell at 25,000 resolution (at 400 m/z) either with ion source collision induced dissociation (ISCID) at 80 volts or without ISCID. During an ISCID scan, an offset of the specified 80 volts is applied to ion optics downstream from the skimmer (i.e. lenses, multipoles, and ion trap) to accelerate ions for fragmentation. The offset is applied by subtraction of the offset value from the tuned value for each individual component. Data acquisition with ISCID at 80 volts is referred to as a “high energy” scan, and without ISCID is referred to as a “low energy” scan. The second, follow-up method used a six scan cycle with ISCID at 80 volts for all scans. The first scan was acquired in the ICR cell, followed by 5 MS/MS scans in the LTQ. Additionally, a third method was used for validation of inter-cross-links identified with BLinks. The validation method contained a single scan in the ICR cell that was acquired without ISCID followed by two CID MS/MS events in the ICR that targeted inter-cross-linked peptides using a mass and time target list.
Software Analysis
Low and high energy MS scans were separated into two sets and analyzed with Hardklör13. Hardklör was operated with the default parameters in addition to using a correlation threshold of 0.90, and a maximum charge state of 9. MS/MS spectra were analyzed with Mascot14 (Matrix Sciences) and exported to comma separated values (.csv) file after applying an expect cut-off of 0.05. The low and high energy Hardklör results were imported into BLinks along with each MS/MS results file obtained on the dataset from Mascot.
The Hardklör results imported into BLinks were used to create extracted ion chromatograms (XICs) for all persistent ion signals. Persistent ion signals were defined as isotope distributions observed in at least three or more consecutive spectra with a 10.0 ppm mass tolerance. The monoisotopic mass values for each persistent ion signal were used to identify cross-linked PIR mass relationships. PIR cross-linked relationships were made by summing the masses of co-eluting PIR-fragment ions in the high energy scans, namely the PIR reporter ion mass and one or two released peptide masses, and matching them to the mass of an intact PIR precursor ion in the low energy scans, within 5.0 ppm. The XICs of persistent ion signals involved in PIR mass relationships were then aligned by retention time, and, where retention time overlap in the XICs was found, the signal intensity profiles of XICs were correlated to obtain the Pearson product moment coefficient (r).
To determine the significance of each correlation, r was used to compute Student’s t statistic. For the null case of no correlation, the following equation
is distributed like Student’s t-distribution with N-2 degrees of freedom, where N is the number of data points.15,16 Using this statistic, a p value for each PIR relationship was calculated. An N of at least 20 was used in this study, which, given the duty cycle of the mass spectrometer (two ICR scans at 25,000 resolution), approximated to 15 seconds of chromatographic retention time. Given that all XICs have the same general shape, some correlation scores are observed even for the null case. Furthermore, p values give significance in terms of false positive rate17. For these reasons, the p value is used as a filter for relevant cross-linked peptide relationships prior to subsequent false discovery rate (FDR) calculations.
A false discovery rate was determined by identifying “decoy” cross-linked PIR mass relationships. As previously described, mass relationships are determined by summing the fragmented PIR product peptide masses with the reporter ion mass to match the intact PIR precursor mass. A decoy PIR relationship is obtained when the peptide ion masses are summed to an incorrect reporter ion mass to match an intact precursor ion mass. To create a decoy mass, a +11 dalton mass shift was applied to the reporter ion mass, in a manner similar to decoy strategies used for accurate mass and time (AMT) tag studies.18 Product peptide ion masses that sum together with the decoy reporter ion mass to match a precursor mass are false. These false relationships are correlated as described above and used to assess a FDR for the same p value cutoff as PIR relationships determined using the correct reporter ion mass.
Results
PIR-labeled HSA was digested to peptides and the PIR-linked peptides were enriched by avidin capture as described in the methods. The PIR-labeled HSA peptides were analyzed by μLC using two methods: 1. MS analysis in the ICR while alternating the use of low collision energy in source, and 2. shotgun MS/MS analysis with constant low collision energy in source. The first method acquired high-resolution spectra containing either the intact PIR-linked precursor ions or the released reporter and peptide ions resulting from cleavage of the labile PIR bonds at the low collision energy. The second method was used to obtain MS/MS spectra for the released peptides resulting from PIR cleavage. Spectrum analysis for each method was performed using Hardklör or Mascot, respectively, and imported into BLinks for the identification of cross-linked peptide pairs.
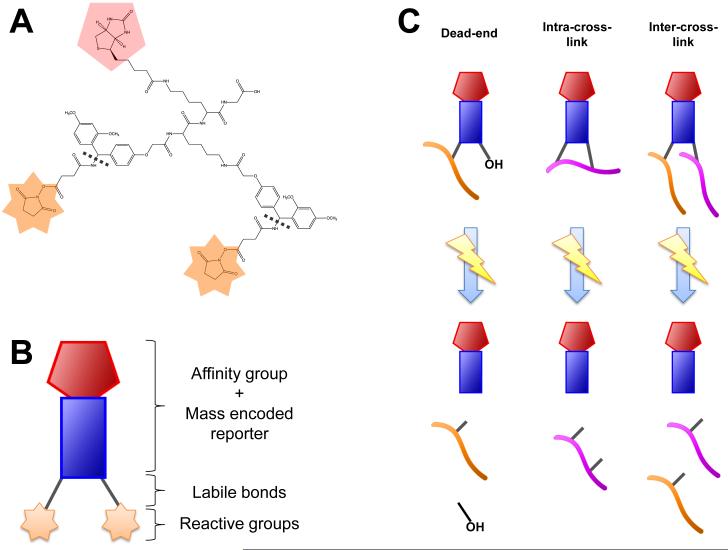
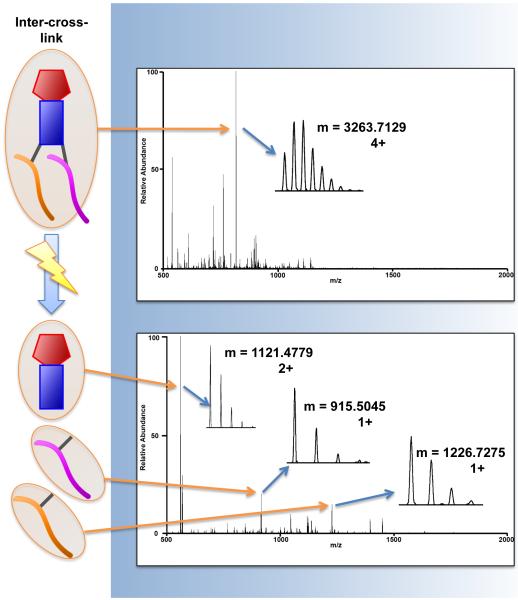
PIR cross-linkers fragment to yield three distinct components when ion source collision energy is increased: A reporter ion containing the PIR backbone and biotin group, and two short arms bound to lysine residues (Figure 1A and 1B). BLinks was used to analyze the ISCID fragmentation scans to identify three types of PIR-linked peptide relationships: dead-end peptides, intra-cross-linked peptides, and inter-cross-linked peptides (Figure 1C). Dead-end peptides are formed when only one short arm of the PIR cross-linker reacts with a lysine. Intra-cross-linked peptides are single peptides containing two lysines bound to each short arm. Inter-cross-linked peptides are two distinct peptides attached to each short arm of the cross-linker. For dead-ends and intra-cross-links, BLinks compares the intact PIR-linked mass to the summed masses of the reporter ion and a single peptide ion. For inter-cross-linked peptides, two peptide ion masses plus the reporter ion mass are summed together to find the intact PIR-linked mass (Figure 2).
Figure 1.
Illustration of PIR cross-linking technology. (A) The chemical structure for the in-house synthesized cross-linker, Brink. Shown in (B) is a cartoon illustration of Brink, highlighting the affinity group and mass encoded tag, the labile bond regions, and the reactive groups. (C) Three general PIR products are formed from the fragmentation of PIR-linked peptides: dead-end, intra-cross-links, and inter-cross-links. Dead-ends and intra-cross-linked relationships are made from the contribution of a single peptide mass and the reporter ion mass. Inter-cross-linked relationships involve two peptide ions and the reporter ion.
Figure 2.
Illustration of PIR fragmentation and data acquisition using in-source collision induced dissociation (ISCID). Use of ISCID is alternated between each spectrum acquisition, generating mass spectra with either intact PIR precursor ions, or fragmented PIR product ions. Inter-cross-linked relationships are made from the summation two peptide ion masses and the reporter ion mass in the product ion scans to produce the intact precursor ion mass observed in the previous scan event.
Because PIR analyses typically consist of thousands of spectra, performing PIR analysis on a scan-by-scan basis produces thousands of redundant cross-linked relationships. To reduce the redundancy of the PIR relationships reported, BLinks was developed and used to trace ion signals chromatographically and produce a single entry for each persistent ion signal. Persistent ion signals were defined as isotope distributions observed in at least three or more consecutive spectra, within a 10.0 ppm mass tolerance, and allowing for a single gap. The extracted ion chromatograms (XICs) for each persistent ion signal were then used to identify PIR mass relationships. In cases where multiple charge states were observed for either the complexes or the released peptides, only XICs for the most intense charge states were analyzed with Blinks to further reduce redundancy. Use of the XICs from the most intense charge state gave better correlation values than the alternative approach of summing signal intensities across charge states. This was because the lower intensity charge states were often not detectible from the noise over the same retention times as the more intense charge states. Thus, summing the XICs produced an abnormal spike in intensity at the apex of elution instead of a normally distributed signal profile. Depending on the ionization properties of the different PIR fragment ions, this spiking effect could be mild or pronounced. For inter-cross-linked relationships, this spiking effect could produce a poor correlation if it was pronounced for one PIR fragment, but not the other. For this experiment, the HSA ion signals were divided into a set containing the persistent intact PIR-linked ions and a set containing the persistent ISCID PIR peptide fragments. BLinks was then used to identify persistent ion signals in each set and PIR relationships were made by summing persistent ion masses from the fragmented set and comparing them to persistent ion masses from the intact set.
By reducing all the ions identified to a set of persistent ion signals, the redundancy in the results was reduced with BLinks. For the HSA sample analyzed, 298 dead-ends, 207 intra- and 606 inter-cross-linked peptide relationships were identified with BLinks at 5ppm mass accuracy. The reduction in redundancy is an essential step in obtaining tractable results and provides additional utility over the use of PIR-analysis software, X-links, where 19,809 dead-end, 13,735 intra- and 20,277 inter-cross-linked peptide relationships are made at 5ppm mass accuracy using the same dataset, due to redundancy that results from multiple scans and charge states. For all previously reported PIR interaction data, complex manual verification based on co-eluting appearance of cross-linked and released peptides was performed, followed by repeated MS/MS validation8. BLinks represents a computational approach to achieve similar verification in an automated fashion.
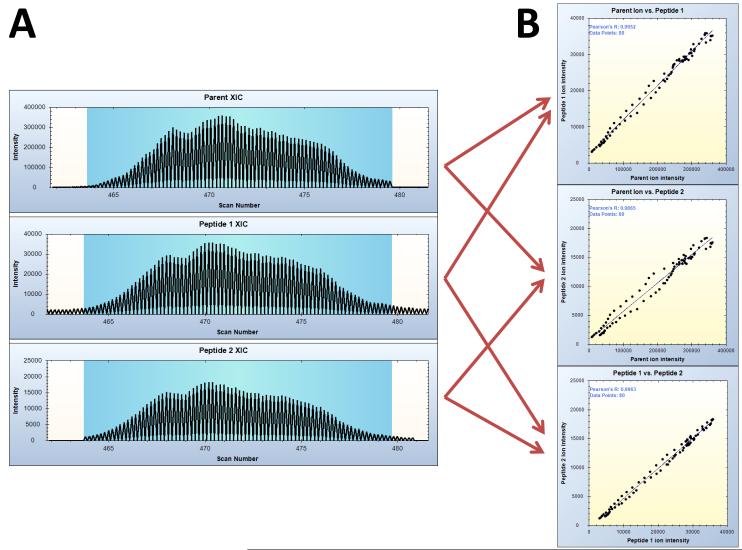
BLinks was used to correlate and obtain a p value for each cross-linked PIR relationship from the HSA sample, as described in the methods. Whereas use of X-links requires manual inspection of the ion XICs for each relationship, BLinks is used to automate the correlation of the XICs of the precursor ion to the fragment ions for each relationship. For dead-end and intra-cross-links, a single r value is obtained. For intra-cross-linked peptides, three correlations are performed: 1) the parent ion to the first fragment ion, 2) the parent ion to the second fragment ion, and 3) the two fragment ions to each other (Figure 3). For each XIC, correlations are performed only for the data points shared in all components of the relationship. The p values for each correlation are calculated as described in the methods.
Figure 3.
Extracted ion chromatogram correlation for inter-crosslinked peptides. The chromatogram intensities shared between the intact PIR-linked ion and the two short arms (indicated in the blue boxes of A) are used to produce three correlation scores (B) relating 1) the intact PIR-linked ion to the first short arm, 2) the intact PIR-linked ion to the second short arm, and 3) the two short arms to each other.
For the HSA sample, cross-linked PIR relationships were analyzed with BLinks that had a mass tolerance of 5 ppm and at least 20 data points in the XIC correlations. 104 unique inter-cross-linked peptide relationships were identified. The p value for each relationship was used as score to filter out relevant PIR relationships. Forty-six of the 104 inter-cross-linked relationships had a p value less than 0.05 (Table 1). An estimate of the false discovery rate (FDR) was made using a decoy reporter ion mass shifted by +11 Daltons, as described in the methods. Only a single inter-cross-linked peptide relationship at p<0.05 was made using the decoy reporter ion mass (2.2% FDR). Mascot search results from the MS/MS spectra of product ions were used to assign peptide sequences. The FDR is computed for the PIR cross-linking relationship, and is not based on peptide sequence identification, and thus any sequence identifications from the Mascot results do not influence the FDR calculations. These Mascot sequence identifications are incorporated within the Blinks analysis to infer protein relationships between inter-cross-links. Forty-one of the 46 inter-cross-linked relationships had at least one peptide identified with a modified residue. Seven of the peptides that were not identified by MS/MS were too short for the database search algorithm, but could be identified by accurate mass and manual inspection of the MS/MS spectra. With manual peptide identification, both peptides could be identified for 42 of the 46 inter-cross-linked relationships. Of the four remaining inter-cross-linked relationships, two had single peptide sequence identifications. Distances between the ε-amines of inter-cross-links were computed using the crystal structure for HSA (Figure 4). Most cross-linked distances were between 15-25 angstroms, which was within the computed maximum cross-linker distance of approximately 43 angstroms.
Table 1.
Inter-cross-linked peptide relationships for HSA
| Parent Neutral Mass (Da) |
Mass Accuracy (ppm) |
Peptide #1 Sequence a | Peptide #1 Neutral Mass (Da) |
Peptide #2 Sequence a | Peptide #2 Neutral Mass (Da) |
Figure Key |
|---|---|---|---|---|---|---|
| 2343.2042 | −2.5158 | R.QIKK.Q | 614.3774 | K.HKPK.A | 607.3458 | S1 |
| 2674.3313 | −2.5589 | K.ATKEQLK.A | 915.5047 | R.YTKK.V | 637.3447 | S2 |
| 2773.3644 | −2.466 | K.ATKEQLK.A | 915.5047 | R.YTKK.V | 736.3777 | S3 |
| 2893.3304 | −0.9445 | K.VGSKCCK.H | 936.4175 | K.HPEAKR.M | 835.4333 | S4 |
| 3035.4762 | −3.957 | R.AFKAWAVAR.L | 1117.6067 | K.SEVAHR.F | 796.384 | |
| 3103.4594 | −0.3399 | R.LKCASLQK.F | 1045.5627 | K.VGSKCCK.H | 936.4181 | S5 |
| 3110.5362 | −4.0087 | K.KYLYEIAR.R | 1153.6181 | K.HPEAKR.M | 835.4322 | S6 |
| 3175.6019 | −1.545 | K.KYLYEIAR.R | 1153.6181 | R.NLGKVGSK.C | 900.5027 | S7 |
| 3211.5202 | −2.7146 | K.KYLYEIAR.R | 1153.6181 | K.VGSKCCK.H | 936.4185 | S8 |
| 3242.4984 | −2.3921 | R.FKDLGEENFK.A | 1324.6314 | K.SEVAHR.F | 796.384 | S9 |
| 3263.7129 | −1.3881 | K.KQTALVELVK.H | 1226.7275 | K.ATKEQLK.A | 915.5045 | S10 |
| 3323.562 | −0.76 | R.VTKCCTESLVNR.R | 1564.7375 | R.YTKK.V | 637.3449 | |
| 3350.6923 | −3.8909 | -- | 1510.8378 | -- | 718.3679 | ‡ |
| 3351.6488 | −3.1718 | R.LAKTYETTLEK.C | 1394.7316 | K.HPEAKR.M | 835.4322 | |
| 3361.6492 | 0.4582 | R.LAKTYETTLEK.C | 1394.7316 | K.ASSAKQR.L | 845.4407 | |
| 3404.6133 | −3.2296 | R.NLGKVGSKCCK.H | 1447.6958 | K.HPEAKR.M | 835.4322 | S11 |
| 3416.7191 | −1.6877 | R.LAKTYETTLEK.C | 1394.7316 | R.NLGKVGSK.C | 900.5057 | S12 |
| 3452.6231 | 2.2849 | R.LAKTYETTLEK.C | 1394.7316 | K.VGSKCCK.H | 936.4189 | S13 |
| 3454.5845 | −1.2745 | K.ADDKETCFAEEGKK.T | 1725.7578 | K.HKPK.A | 607.3458 | S14 |
| 3534.6786 | −3.6771 | K.DVCKNYAEAK.D | 1295.5851 | R.AFKAWAVAR.L | 1117.6067 | S15 |
| 3561.7809 | −4.0461 | R.LAKTYETTLEK.C | 1394.7316 | R.LKCASLQK.F | 1045.5615 | S16 |
| 3674.7018 | 0.4412 | K.LDELRDEGKASSAK.Q | 1616.8069 | K.VGSKCCK.H | 936.4181 | S17 |
| 3691.7369 | −3.9582 | R.LKCASLQKFGER.A | 1633.8303 | K.VGSKCCK.H | 936.4185 | S18 |
| 3731.9886 | −4.8931 | R.QIKKQTALVELVK.H | 1694.9934 | K.ATKEQLK.A | 915.5045 | S19 |
| 3774.9603 | −3.8717 | K.KVPQVSTPTLVEVSR.N | 1737.9682 | K.ATKEQLK.A | 915.5039 | S20 |
| 3792.7552 | −1.2255 | -.DAHKSEVAHR | 1346.6395 | R.FKDLGEENFK.A | 1324.6345 | S21 |
| 3800.8674 | 1.1668 | R.LKCASLQKFGER.A | 1633.831 | R.LKCASLQK.F | 1045.5616 | |
| 3891.9012 | −0.3875 | K.LDELRDEGKASSAK.Q | 1616.8041 | K.KYLYEIAR.R | 1153.6181 | S22 |
| 3956.918 | 0.1151 | K.LDELRDEGKASSAKQR.L | 2000.0071 | K.HPEAKR.M | 835.4333 | S23 |
| 4057.8998 | −0.0591 | K.LDELRDEGKASSAKQR.L | 2000.0036 | K.VGSKCCK.H | 936.4181 | S24 |
| 4113.9543 | −2.063 | K.VGSKCCKHPEAK.R | 1597.7386 | R.LAKTYETTLEK.C | 1394.7316 | S25 |
| 4150.0499 | −3.3262 | R.LKCASLQKFGER.A | 1633.8303 | R.LAKTYETTLEK.C | 1394.7316 | S26 |
| 4167.0433 | 0.7967 | K.LDELRDEGKASSAKQR.L | 2000.0051 | R.LKCASLQK.F | 1045.5627 | S27 |
| 4203.0135 | −3.0743 | R.LKCASLQKFGER.A | 1633.8303 | R.NLGKVGSKCCK.H | 1447.6958 | S28 |
| 4266.1931 | −1.2954 | R.YTKKVPQVSTPTLVEVSR.N | 2229.2072 | K.ATKEQLK.A | 915.5039 | S29 |
| 4275.0975 | 0.4206 | K.LDELRDEGKASSAKQR.L | 2000.0028 | K.KYLYEIAR.R | 1153.6181 | S30 |
| 4336.0239 | 2.2791 | K.LDELRDEGKASSAK.Q | 1616.8081 | K.VGSKCCKHPEAK.R | 1597.7452 | |
| 4372.1111 | 0.3773 | R.LKCASLQKFGER.A | 1633.8303 | K.LDELRDEGKASSAK.Q | 1616.8041 | S31 |
| 4456.1652 | −3.3807 | K.QNCELFEQLGEYKFQNALLVR.Y | 2697.3309 | R.YTKK.V | 637.3451 | |
| 4616.1618 | 0.0076 | -- | 1758.8272 | -- | 1735.8567 | |
| 4766.3216 | −3.8064 | K.LDELRDEGKASSAKQR.L | 2000.0051 | K.LKECCEKPLLEK.S | 1644.8247 | S32 |
| 4847.3091 | −2.0059 | R.NLGKVGSKCCKHPEAK.R | 2109.0168 | K.LDELRDEGKASSAK.Q | 1616.8069 | |
| 4879.2598 | 2.1359 | R.HPYFYAPELLFFAKR.Y | 1997.0335 | R.YKAAFTECCQAADK.A | 1760.7564 | |
| 5690.664 | −2.0077 | K.LDELRDEGKASSAKQR.La | 3121.4811 | R.NLGKVGSKCCK.H | 1447.6958 | † |
| 5690.664 | −0.9572 | R.NLGKVGSKCCK.Ha | 2569.1792 | K.LDELRDEGKASSAKQR.L | 2000.0025 | † |
| 5746.7162 | −4.5562 | -- | 3230.4856 | R.LAKTYETTLEK.C | 1394.7316 |
Reactive amino acids are underlined, bold indicates peptide sequence identification through accurate mass and inspection of MS/MS spectra. After PIR-cleavage, the peptides retain a 99.032 Da residual modification mass. An asterisk (*) indicates the peptide was observed with the reporter mass still attached due to incomplete PIR fragmentation.
False positive after targeted analysis.
Same cross-link relationship. See text for details.
Figure 4.
Histogram of the distances between ε-amines of inter-cross-linked peptides. The distances were calculated from the crystal structure of HSA.
Validation of p values less than 0.05
The inter-cross-linked peptides identified using BLinks were validated in a follow-up analysis that isolated cross-linked precursor ions and analyzed them by collision induced dissociation (CID). The 104 inter-cross-linked precursor m/z values were targeted for CID using mass and time inclusion lists generated from the BLinks results. Thirty-two of the 46 inter-cross-linked peptides with p<0.05 were validated by CID fragmentation of the BRink cross-linker (Supplemental Figures 1-32). Thirteen of the 46 inter-cross-linked peptides were missed by CID selection, or produced spectra of poor quality. One of the 46 inter-cross-linked peptide relationships was shown to be incorrect. Incidentally, this relationship was identified during the column wash portion of the chromatography which suggests that false relationship observations can be minimized by avoiding analysis of the many peptides that co-elute during the wash step. Although true cross-linked relationships may elute during the wash portion of the chromatography, they would likely be better observed using different fractionation methods.
Inspection of p values greater than 0.05
Cross-linked PIR relationships that were found to have a p value greater than 0.05 were also validated using CID. This analysis was performed to confirm that p values derived from BLinks analyses can accurately discriminate between true and false discovery relationships. Of the 58 inter-cross-linked peptide relationships with p>0.05, sixteen were confirmed to be real cross-links (Table 2 and Supplemental Figures 33-48). Eleven of these sixteen relationships missed by the analysis with BLinks result from peptides that are involved in multiple co-eluting cross-linked relationships, causing a spike in peptide signal intensity. This spike is the cause of poor correlation between the two peptides or with the parent ion intensity, which suggests that the p value derived relationships represent a conservative subset of all relationships present in the sample.
Table 2.
Additional inter-cross-linked peptide relationships for HSA
| Parent Neutral Mass (Da) |
Mass Accuracy (ppm) |
Peptide #1 Sequence a | Peptide #1 Neutral Mass (Da) |
Peptide #2 Sequence a | Peptide #2 Neutral Mass (Da) |
Figure Key |
|---|---|---|---|---|---|---|
| 2366.1728 | −2.8711 | R.YTKK.V | 637.3455 | K.HKPK.A | 607.3458 | S33 |
| 2985.5536 | −2.0244 | K.KQTALVELVK.H | 1226.7275 | R.YTKK.V | 637.3444 | S34† |
| 3002.4768 | −2.4315 | R.LKCASLQK.F | 1045.5621 | K.HPEAKR.M | 835.4322 | S35 |
| 3012.4815 | −1.8369 | R.LKCASLQK.F | 1045.5627 | K.ASSAKQR.L | 845.4374 | S36† |
| 3084.5858 | −2.177 | K.KQTALVELVK.H | 1226.7275 | R.YTKK.V | 736.3761 | S37† |
| 3265.5539 | 1.5048 | K.KYLYEIAR.R | 1153.6181 | R.DEGKASSAK.Q | 990.4611 | S38† |
| 3284.6496 | −1.8369 | R.AFKAWAVAR.L | 1117.6067 | R.LKCASLQK.F | 1045.561 | S39† |
| 3600.7341 | 4.9315 | R.LKCASLQKFGER.A | 1633.8303 | K.ASSAKQR.L | 845.4381 | S40† |
| 3722.7855 | 2.432 | R.NLGKVGSKCCK.H | 1447.6958 | K.KYLYEIAR.R | 1153.6181 | S41† |
| 3963.9174 | −4.7753 | R.NLGKVGSKCCK.H | 1447.6943 | R.LAKTYETTLEK.C | 1394.7316 | S42† |
| 4073.9621 | −0.3636 | K.ADDKETCFAEEGKK.T | 1725.7553 | K.KQTALVELVK.H | 1226.7278 | S43† |
| 4080.9433 | 2.6783 | R.VTKCCTESLVNR.R | 1564.7417 | R.LAKTYETTLEK.C | 1394.7316 | S44 |
| 4150.0449 | −1.6752 | R.LKCASLQKFGER.A | 1633.8303 | R.LAKTYETTLEK.C | 1394.7316 | S45†,‡ |
| 4516.2116 | 0.0961 | K.LDELRDEGKASSAKQR.L | 2000.0024 | R.LAKTYETTLEK.C | 1394.7316 | S46† |
| 4569.1833 | −0.6595 | K.LDELRDEGKASSAKQR.L | 2000.0051 | R.NLGKVGSKCCK.H | 1447.698 | S47 |
| 4755.3174 | −1.754 | K.LDELRDEGKASSAKQR.L | 2000.0028 | R.LKCASLQKFGER.A | 1633.8303 | S48 |
Reactive amino acids are underlined, bold indicates peptide sequence identification through accurate mass and inspection of MS/MS spectra. After PIR-cleavage, the peptides retain a 99.032 Da residual modification mass.
Relationship also identified with BLinks with a chromatographically independent precursor ion of the same mass. The chromatographic separation is likely caused by chirality of the molecule.
Relationship showed poor correlation due to involvement of one or both peptides in another relationship.
Co-eluting inter-cross-links involving the same peptide sequence were observed for relationships above and below the p value cutoff of 0.05. This co-elution causes an irregular correlation graph that produces a low correlation score and a high p value. For example, even though a peptide may originate from the identified cross-linked product, this peptide may also be derived from other cross-linked products. If these products overlap chromatographically, misleadingly high p values will be derived from the correlation analysis. Although this co-elution may be expected to be only infrequently observed with in vivo PIR applications where each protein is cross-linked to a smaller extent than in purified protein experiments, Figure 5 illustrates an example. Here, BLinks was used to analyze XICs and interaction maps for each peptide arm to help visualize instances where co-elution of different cross-link relationships affects the same peptide. The peptide HKPK was found cross-linked to both QIKK in one relationship and ADDKETCFAEEGKK in another (Figure 5B). Despite the adverse affect on the correlation score, this relationship was still identified through BLinks analysis with a p value below 0.05. Again, the observation of co-eluting relationships involving the same peptide may be a result of heavily cross-linking a purified protein, and is less likely in a complex biological sample as discussed above. Nonetheless, these products present the most extreme challenges for informatics methods and PIR experiments and data here suggest these complications are surmountable by analyses using BLinks.
Figure 5.
(A) The correlation graph showing poor correlation between the intact PIR-linked ion and the second short arm in the relationship. (B) The relationship map for the second short arm shows that it is involved in two inter-crosslinked relationships, with an ion of mass 614.38 Da and an ion of mass 1725.76 Da. (C) The extracted ion chromatograms for the intact PIR-linked ion and the second short arm. The blue boxes in (B) and (C) indicate the region over which the correlation in (A) is made for the relationship. The contribution of the ions from the second inter-crosslinked peptide relationship are the cause of the poor correlation.
The abundance and proximity of available reactive sites in HSA increases the likelihood of inter-cross-linked peptides that contain multiple cross-linkers. An extreme case was observed in which two peptides, NLGKVGSKCCK and LDELRDEGKASSAKQR, each contained two reactive lysine residues bound by two cross-linkers (Table 1, † labeled rows). Despite this complexity, the peptide cross-link relationships could still be identified because incomplete cleavage of the PIR bonds resulted in dissociated peptide ions which still included a single reporter mass. Inter-cross-link relationships could be made that showed the incomplete cleavage on one or the other peptide. The observation of this doubly-linked relationship implies that cross-linking should also exist for single sites and indeed the simpler cross-linking of the subsequences VGSKCCK to LDELRDEGKASSAK was found. As shown in Tables 1 and 2, several short peptides are actually subsequences of larger peptides identified with multiple sites of cross-linker attachment.
From the peptide information, 30 unique sites of cross-linker attachment were identified. Twenty-seven of the sites were reactive lysine residues. Additionally, cross-links were identified involving a single serine residue, a tyrosine residue, and the protein N-terminus. Using the Swiss-PdbViewer19, the distances were mapped between inter-cross-linked amines. The shortest distance was computed to be 6.890 angstroms and the longest distances was 41.439 angstroms. These distances are consistent with the flexibility and estimated maximum length of 43 angstroms of the a similar Rink-based PIR cross-linker7. Many reactive residues were involved in multiple cross-link relationships. Inter-cross-links for which both peptides could be identified were compared to Rinner et al.20 where 10 unique pairs of lysine residues were found cross-linked with DSS (approximately 11 angstrom length). Seven of those residue pairs were also identified by this PIR method using BLinks.
Discussion
The in vivo application of PIR-cross-linking and the ease of peptide identification when using this technology offer great potential for the discovery and study of protein-protein interactions using mass spectrometry. Fundamental to the success of PIR-cross-linking is the mathematical assembly of product ion relationships after labile-bond breakage of the intact PIR precursor ion. Although this mathematical assembly is not difficult to perform for single cross-linked product analysis, its large-scale application presents many challenges. For complex biological samples, the number of PIR-cross-linked relationships can number in the thousands and existing software requires manual inspection of chromatographic profiles for validation. The BLinks software contains computational tools to dramatically reduce the data complexity and automate the validation of PIR-cross-linked peptides.
For instruments with only a moderate duty cycle, a single PIR-cross-linked relationship may be observed dozens of times on an individual scan basis. Because hundreds of such relationships can exist, the resulting data from a complex sample is a dense web of interwoven PIR-cross-linked relationships interspersed with random relationships that occur as single scan events from noise and other spurious signals. By observing PIR-cross-linked relationships chromatographically rather than on an individual scan basis, significant data reduction is performed when using BLinks. Additionally, because noise and spurious ion-like signals do not persist from scan to scan, they are removed from the analysis and only chromatographically persistent peptide signals are used to compute PIR-cross-linked relationships. Thus, the likelihood of observing random PIR-cross-linked relationships is reduced when using BLinks.
The correlation of signal intensities across chromatographic profiles automates the validation of observed PIR-cross-linked relationships while minimizing manual interpretation of extracted ion chromatograms. A t-test is used to help interpret the results of Pearson’s correlation; Pearson’s correlation might not be an accurate indicator of a relationship when only a few data points are used. Similarly, a poor Pearson’s correlation might be observed for a valid relationship over many points, in which some of the signal intensity is explained by the contribution of a second relationship involving the same peptide. BLinks is used to perform these statistical tests and provide tools to map such overlapping relationships for cases in which manual interpretation is most prone to error.
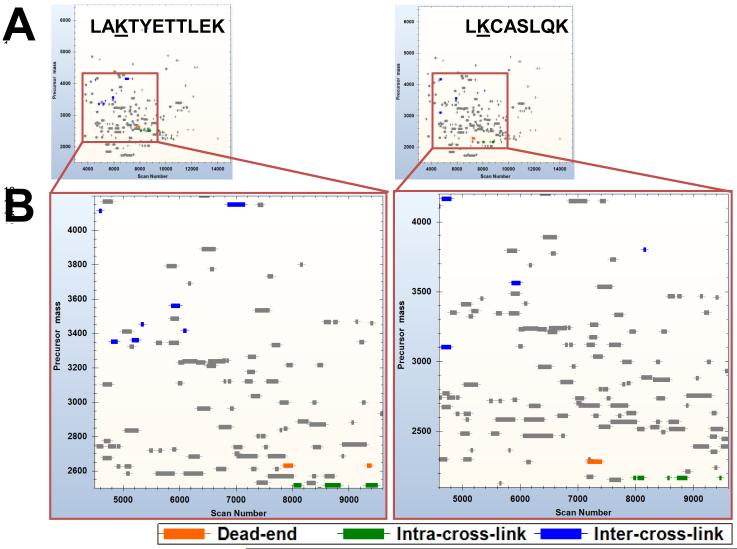
BLinks can be used to profile and classify all observed PIR cross-link relationships to optimize analysis of inter-cross-link relationships and facilitate topographical analysis. Figure 6 illustrates PIR cross-linked relationships graphed by mass and scan number (retention time), and color-coded to indicate dead-ends, intra-cross-links, and inter-cross-links. A large number of dead-ends and intra-cross-links were identified in the sample, which is expected given the possible reaction products of chemical cross-linking. This observation gives greater confidence to the analysis than if inter-cross-linked relationships were found without also identifying dead-end and intra-cross-linked relationships. Also, because of the physiochemical properties of the BRink PIR cross-linker, dead-ends, intra-cross-links, and inter-cross-links occupy separate regions of mass and retention time. These differences in mass and retention time can be exploited to focus more closely on inter-cross-linked relationships. For example, it is possible to incorporate multiple LC runs in the analysis that use additional SCX or SEC fractionation to comprehensively analyze inter-cross-link relationships. The visualization tools in BLinks can be used to focus on cross-linked peptides of interest. As shown in Figure 7, specific sites of interaction can be highlighted to show all the relationships involving two inter-cross-linked peptides. As expected, dead end relationships were observed for each peptide. Surprisingly, each peptide was indicated to be involved in intra-cross-linked relationships despite the existence of only a single lysine residue. Such relationships are possible if the precursor ion partially fragmented prior to analysis in the ICR during the alternating ISCID stage of data acquisition. This partial fragmentation results in precursor ions that contain only one labile bond linked to a peptide. Thus, a direct mass relationship between the precursor ion mass and a single peptide ion mass can be made like when identifying intra-cross-linked relationships. BLinks can be used to find these unusual cross-linker relationships, despite their misclassification. Because of this functionality, these unusual relationships can be targeted by MS/MS using the methods described above to obtain the correct classification. Figure 6 also shows that each peptide was involved in multiple inter-cross-linked relationships. With Blinks, complex PIR datasets can be uniquely filtered to graphically reveal complex cross-linking patterns of sites that, because of hyper-reactivity, importance in interactions, or both are linked in many different ways.
Figure 6.
Profile of all PIR relationships identified with BLinks. Inter-cross-links form a distinct cluster from dead-ends and intra-cross-links when plotted by retention time and mass.
Figure 7.
Complete PIR relationship profiles for two inter-cross-linked peptides. The specific peptides involved in a single inter-cross-linked relationship are highlighted to show their involvement in other cross-linked relationships.
The use of chromatographic profiles for each inter-cross-linked relationship also allows for expansion of the statistical tests and types of analyses to be performed. Because peptide intensity values are tracked over the entire elution profile for each PIR-linked peptide pair, it is possible to quantify and compare inter-cross-linked relationships between multiple samples. This capability has application when comparing differences in protein-protein interaction between different individuals or under different conditions when using PIR-based analysis methods. Thus, use of BLinks has the potential to expand the capabilities of PIR-cross-linking technology to the exciting areas of quantitative protein interaction and topology measurements.
Conclusion
The BLinks software provides new tools to facilitate identification of PIR-linked relationships when performing cross-linking analysis, extending the capabilities to analyze PIR-linked proteins in complex samples8,9. The use of extracted ion chromatograms to perform downstream analysis dramatically reduces the complexity and redundancy observed when using existing software tools. Spurious PIR relationships are eliminated from the analysis through the use of persistent ion signals. Additionally, the chromatographic profiles of the ions can be exploited to compute the correlation of fragmented PIR-linked peptides to their intact precursor ions, and thus automate the validation process for identified PIR-linked relationships. Finally, Blinks allows a global view of all detected cross-linked relationships with an entire LC/MS run or potentially, a complete set of LC/MS runs. This allows visualization of all cross-linked species and filtering to enable detection of all cross-linked species with any peptide of interest. These capabilities greatly accelerate the analysis of complex PIR datasets and allow unparalleled detection of cross-linked peptides which will significantly increase in vivo PIR applications.
Synopsis.
Protein interaction reporter (PIR) technology can enable identification of in vivo protein interactions using specialized chemical cross-linkers, liquid chromatography, and high-resolution mass spectrometry. Presented here is a software tool, BLinks, for use in the analysis of PIR-cross-linked proteins. BLinks was used to analyze human serum albumin and track peptides involved in multiple relationships that make accurate identification of protein interactions difficult. BLinks was also used to globally map all inter-peptide relationships from the data analysis and customize subsequent analysis to target specific peptides of interest, thus making it a useful tool for both discovery of protein interactions and mapping protein topology.
Supplementary Material
Acknowledgements
This research was supported by the National Institutes of Health through grants R01GM086688 and R01RR023334 and through the University of Washington Proteomics Resource (UWPR95794).
References
- 1.Fields S, Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989;340:245–246. doi: 10.1038/340245a0. [DOI] [PubMed] [Google Scholar]
- 2.Ho Y, Gruhler A, Heilbut A, Bader GD, Moore L, Adams SL, Millar A, Taylor P, Bennett K, Boutilier K, Yang L, Wolting C, Donaldson I, Schandorff S, Shewnarane J, Vo M, Taggart J, Goudreault M, Muskat B, Alfarano C, Dewar D, Lin Z, Michalickova K, Willems AR, Sassi H, Nielsen PA, Rasmussen KJ, Andersen JR, Johansen LE, Hansen LH, Jespersen H, Podtelejnikov A, Nielsen E, Crawford J, Poulsen V, Sorensen BD, Matthiesen J, Hendrickson RC, Gleeson F, Pawson T, Moran MF, Durocher D, Mann M, Hogue CW, Figeys D, Tyers M. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. 2002;415:180–183. doi: 10.1038/415180a. [DOI] [PubMed] [Google Scholar]
- 3.Rigaut G, Shevchenko A, Rutz B, Wilm M, Mann M, Seraphin B. A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol. 1999;17:1030–1032. doi: 10.1038/13732. [DOI] [PubMed] [Google Scholar]
- 4.Ramachandran N, Hainsworth E, Bhullar B, Eisenstein S, Rosen B, Lau AY, Walter JC, LaBaer J. Self-assembling protein microarrays. Science. 2004;305:86–90. doi: 10.1126/science.1097639. [DOI] [PubMed] [Google Scholar]
- 5.Zhu H, Snyder M. Protein chip technology. Curr Opin Chem Biol. 2003;7:55–63. doi: 10.1016/s1367-5931(02)00005-4. [DOI] [PubMed] [Google Scholar]
- 6.Sinz A. Investigation of protein-protein interactions in living cells by chemical crosslinking and mass spectrometry. Anal Bioanal Chem. 2010 doi: 10.1007/s00216-009-3405-5. Epub ahead of print. [DOI] [PubMed] [Google Scholar]
- 7.Tang X, Munske GR, Siems WF, Bruce JE. Mass spectrometry identifiable cross-linking strategy for studying protein-protein interactions. Anal Chem. 2005;77:311–318. doi: 10.1021/ac0488762. [DOI] [PubMed] [Google Scholar]
- 8.Zhang H, Tang X, Munske GR, Tolic N, Anderson GA, Bruce JE. Identification of protein-protein interactions and topologies in living cells with chemical cross-linking and mass spectrometry. Mol Cell Proteomics. 2009;8:409–420. doi: 10.1074/mcp.M800232-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Anderson GA, Tolic N, Tang X, Zheng C, Bruce JE. Informatics strategies for large-scale novel cross-linking analysis. J Proteome Res. 2007;6:3412–3421. doi: 10.1021/pr070035z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Leitner A, Walzthoeni T, Kahraman A, Herzog F, Rinner O, Beck M, Aebersold R. Probing native protein structures by chemical cross-linking, mass spectrometry and bioinformatics. Mol Cell Proteomics. doi: 10.1074/mcp.R000001-MCP201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Katritzky AR, Yang B, Qiu G, Zhang Z. A Convenient Trifluoroacetylation Reagent: N-(Trifluoroacetyl)succinimide. Synthesis. 1999;1:55–57. [Google Scholar]
- 12.Wisniewski JR, Zougman A, Mann M. Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J Proteome Res. 2009;8:5674–5678. doi: 10.1021/pr900748n. [DOI] [PubMed] [Google Scholar]
- 13.Hoopmann MR, Finney GL, MacCoss MJ. High-speed data reduction, feature detection, and MS/MS spectrum quality assessment of shotgun proteomics data sets using high-resolution mass spectrometry. Anal Chem. 2007;79:5620–5632. doi: 10.1021/ac0700833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Perkins DN, Pappin DJ, Creasy DM, Cottrell JS. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis. 1999;20:3551–3567. doi: 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 15.Rahman NA. A course in theoretical statistics for sixth forms, technical colleges, colleges of education, universities. Griffin; London: 1968. [Google Scholar]
- 16.Press WH, Numerical Recipes Software (Firm) Numerical recipes in C. Cambridge University Press; [Cambridge, England]; [New York, N.Y.]: 1993. [Google Scholar]
- 17.Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proc Natl Acad Sci U S A. 2003;100:9440–9445. doi: 10.1073/pnas.1530509100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Petyuk VA, Qian WJ, Chin MH, Wang H, Livesay EA, Monroe ME, Adkins JN, Jaitly N, Anderson DJ, Camp DG, 2nd, Smith DJ, Smith RD. Spatial mapping of protein abundances in the mouse brain by voxelation integrated with high-throughput liquid chromatography-mass spectrometry. Genome Res. 2007;17:328–336. doi: 10.1101/gr.5799207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 1997;18:2714–2723. doi: 10.1002/elps.1150181505. [DOI] [PubMed] [Google Scholar]
- 20.Rinner O, Seebacher J, Walzthoeni T, Mueller LN, Beck M, Schmidt A, Mueller M, Aebersold R. Identification of cross-linked peptides from large sequence databases. Nat Methods. 2008;5:315–318. doi: 10.1038/nmeth.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.