Abstract
Acclimation of cyanobacteria to environmental changes includes major changes in the gene expression patterns partly orchestrated by the replacement of a particular σ subunit with another in the RNA polymerase holoenzyme. The cyanobacterium Synechocystis sp. strain PCC 6803 encodes nine σ factors, all belonging to the σ70 family. Cyanobacteria typically encode many group 2 σ factors that closely resemble the principal σ factor. We inactivated three out of the four group 2 σ factors of Synechocystis simultaneously in all possible combinations and found that all triple inactivation strains grow well under standard conditions. Unlike the other strains, the ΔsigBCD strain, which contains SigE as the only functional group 2 σ factor, did not grow faster under mixotrophic than under autotrophic conditions. The SigB and SigD factors were important in low-temperature acclimation, especially under diurnal light rhythm. The ΔsigBCD, ΔsigBCE, and ΔsigBDE strains were sensitive to high-light-induced photoinhibition, indicating a central role of the SigB factor in high-light tolerance. Furthermore, the ΔsigBCE strain (SigD is the only functional group 2 σ factor) appeared to be locked in the high-fluorescence state (state 1) and grew slowly in blue but not in orange or white light. Our results suggest that features of the triple inactivation strains can be categorized as (i) direct consequences of the inactivation of a particular σ factor(s) and (ii) effects resulting from the higher probability that the remaining group 2 σ factors associate with the RNA polymerase core.
Cyanobacteria are eubacteria capable of oxygen-producing photosynthesis, and the chloroplasts of plants and algae evolved from cyanobacteria (26). Synechocystis sp. strain PCC 6803 (here Synechocystis) is a unicellular cyanobacterium commonly used as a model organism (11). The glucose-tolerant strain of Synechocystis is especially popular, as genetic manipulation of this naturally competent strain is easy, and the cells can be grown under autotrophic, mixotrophic, photoheterotrophic, or light-activated heterotrophic growth conditions (1, 38).
Light is among the key environmental factors, and cyanobacteria are able to respond to both light quality and light quantity. Cyanobacteria acclimate to light intensity changes by adjusting the expression of many genes, including those coding for components of the photosynthetic apparatus (8-10). In addition, the orange carotenoid protein (OCP) (39, 40), the IsiA pigment protein complex (7, 41), high-light-induced proteins (6, 36), and the efficient photosystem II (PSII) repair cycle (33, 34) keep the photosynthetic machinery functional under high-light conditions. The most dramatic light quality effect is seen in those cyanobacteria that perform complementary chromatic adaptation (15, 17). They change color from blue-green to brick red when the cells change phycocyanin to phycoerythrin in phycobilisome antennae upon transfer from red light to green light (15, 17). Synechocystis is not able to perform complementary chromatic adaptation, but it can balance energy distribution between photosystem I (PSI) and PSII according to the light quality (14, 35). Phycobilisomes, which function as major light-harvesting antennae of PSII, efficiently collect orange light, while the chlorophyll (Chl) a antennae of PSI harvest mainly blue and red light. State transitions balance energy distribution between the photosystems according to the light quality: illumination with orange PSII light leads to state 2, in which energy is transferred more efficiently to PSI, and treatment with PSI light (blue light) leads to compensatory energy flow to PSII (state 1) (14, 35).
Acclimation to different environmental conditions is based on adjustments of gene expression. The promoter-recognizing σ subunit of the RNA polymerase (RNAP) holoenzyme has a central role in this process. Different σ factors compete for a limited number of catalytically active RNA polymerase cores, and switching between different σ factors is a major determinant of the overall gene expression pattern in eubacteria (19, 21). In cyanobacteria, all σ factors belong to the σ70 family, which is divided into three subgroups (18, 22). The group 1 σ factor is essential for cell viability, group 2 σ factors are very similar to the group 1 σ factor but are nonessential, and group 3 σ factors differ considerably from groups 1 and 2.
Cyanobacteria typically code for several group 2 σ factors, and recent studies have revealed that group 2 σ factors play crucial roles in acclimation to suboptimal conditions (for a review, see reference 22). Synechocystis encodes four group 2 σ factors. Studies with single inactivation strains have revealed that the SigB (13, 28, 30, 31) and SigC (5, 32) factors are involved in acclimation to high-temperature stress. The SigD factor, together with the SigB and SigE factors, is involved in light regulation both in light-dark transitions and upon light intensity changes (12, 13, 24, 25, 29, 42). The SigE factor has a role in sugar metabolism and is required for light-activated heterotrophic growth (23). All group 2 σ factors affect the acclimation of Synechocystis cells to osmotic stress conditions (25).
In the present study, we constructed strains in which three group 2 σ factor genes were interrupted simultaneously in all possible combinations, leaving only one group 2 σ factor gene active in each strain. Our results point to two distinct features of the inactivation strains: (i) direct consequences of the inactivation of a particular σ factor(s) and (ii) effects resulting from an overdose of the remaining group 2 σ factor.
MATERIALS AND METHODS
Construction of triple inactivation strains of group 2 σ factors in Synechocystis.
The glucose-tolerant strain Synechocystis sp. PCC 6803 (38) was used as a control strain (CS). To construct the triple inactivation strains, the sigC and sigE genes were amplified by PCR using the primers shown in Table 1. The PCR products were cloned into a pCR Blunt II-TOPO vector (Invitrogen). pCR Blunt II-TOPO-sigC was digested with SpeI and EcoRV, and the sigC fragment was ligated into an XbaI and SmaI double-digested pUC19. pCR Blunt II-TOPO-sigE was digested with PstI and EcoRI, and the sigE fragment was ligated into PstI and EcoRI double-digested pUC19. The fragment conferring chloramphenicol (Cm) resistance was obtained from pKRP10 (CVector) by BamHI digestion and subsequently ligated into BamHI-digested pUC19-sigC or pUC19-sigE. The resulting inactivation plasmid, pUC19-sigC-Cmr, was used to transform the ΔsigBD, ΔsigBE, and ΔsigDE strains (25) to obtain the ΔsigBCD, ΔsigBCE, and ΔsigCDE strains, respectively, and to obtain the ΔsigBDE strain, the ΔsigBD strain was transformed with pUC19-sigE-Cmr (see Fig. S1 in the supplemental material for constructs of the triple inactivation strains). Selective agar plates contained kanamycin (Kn) (50 μg/ml), streptomycin (Str) (20 μg/ml), spectinomycin (Spc) (10 μg/ml), and Cm (10 μg/ml).
TABLE 1.
Primers used for amplification of sig genes
| Primer | Sequence |
|---|---|
| sigB forward | 5′-ATGGTAACAGTGACAGTTAT-3′ |
| sigB reverse | 5′-TAGCTCTTGGCCATCGTTA-3′ |
| sigC forward | 5′-ATGACTAAACCAAGCAACGA-3′ |
| sigC reverse | 5′-AATCTAGCAAAATTTCCTGC-3′ |
| sigD forward | 5′-ATGACTGCCAGAACCAGCCC-3′ |
| sigD reverse | 5′-AGATCCAACTGTAGGGAGGC-3′ |
| sigE forward | 5′-ATGAGCGATATGTCTTCCCT-3′ |
| sigE reverse | 5′-CTATAACCAACCTTTGAGGC-3′ |
PCR analysis was used to verify the complete replacement of the native gene with the inactivated gene (Fig. 1 A). Genomic DNA was isolated according to the method of Williams (38), and the sigB, sigC, sigD, and sigE genes were amplified using gene-specific primers (Table 1).
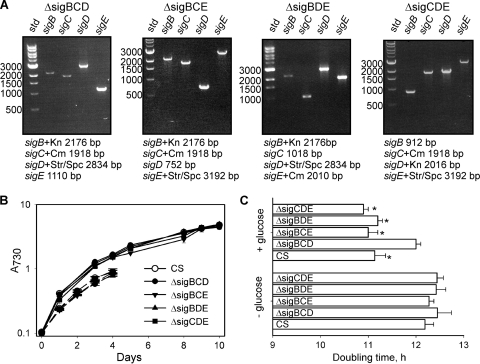
FIG. 1.
Triple inactivation strains of group 2 σ factors in Synechocystis. (A) PCR analysis of the ΔsigBCD, ΔsigBCE, ΔsigBDE, and ΔsigCDE inactivation strains. Genomic DNA was isolated from the inactivation strains, and each sig gene was amplified by PCR. The expected size of each PCR product is shown under each gel. std, standard. (B) Growth of strains. The A730 of the cultures was set to 0.1, and the cells were grown in BG-11 medium, pH 7.5, at a PPFD of 40 μmol m−2 s−1 in continuous light (solid lines) or in a 12-h light/12-h dark rhythm (dashed lines) at 32°C. Each data point represents the mean of at least three biological replicates with independent liquid cultures, and the error bars indicate standard errors (SE). CS, control strain. (C) Doubling times of the strains in the presence and absence of 5 mM glucose at a PPFD of 40 μmol m−2 s−1. The doubling times were calculated on the basis of growth during the first 24 h. Each data point represents the mean of at least four biological replicates with independent liquid cultures, and the error bars indicate SE. The asterisks denote significant differences between the growth rate in the absence and in the presence of glucose (P < 0.01; Student's t test).
Growth of Synechocystis.
Standard growth conditions for Synechocystis were BG-11 medium (25) supplemented with 20 mM HEPES-NaOH, pH 7.5; a continuous photosynthetic photon flux density (PPFD) of 40 μmol m−2 s−1; 32°C; and ambient CO2. The BG-11 agar plates of triple mutants were supplemented with Kn (50 μg/ml), Str (20 μg/ml), Spc (10 μg/ml), and Cm (10 μg/ml) and those of single mutants with Kn (50 μg/ml). For the measurements, cells were grown in liquid media without addition of antibiotics.
For growth measurements, the A730 of liquid cultures was set to 0.1 (30 ml cell culture in a 100-ml Erlenmeyer flask), and the A730 was measured every 24 h. Samples of dense cultures were diluted so that the A730 did not exceed 0.3, and the dilutions were taken into account when the final results were calculated. Similar relationships between the A730 and cell numbers can be expected in the control strain and in all inactivation strains because cells harvested on an A730 basis contain similar amounts of chlorophyll, RNA, and proteins and have the same photosynthetic activity in all strains. Furthermore, the whole-cell absorption profiles of all strains are virtually identical.
Cells were grown under standard conditions under continuous illumination or in a 12-h light/12-h dark rhythm. In addition, growth was measured at 22°C under continuous illumination (PPFD, 40 μmol m−2 s−1) or in a 12-h light/12-h dark rhythm. For light rhythm measurements, the experiments were started in the middle of the light period. Cells were also grown under continuous blue or orange light. Blue light was obtained by illuminating the cultures through a filter with a transmission peak at 425 nm (HT 119 Dark Blue; Lee Filters) and orange light by using filters 105 Orange (Lee Filters) and Rosco number 89; this combination has a transmission peak at 568 nm. Mixotrophic growth conditions were obtained by adding 5 mM glucose to the growth medium under otherwise standard conditions. The growth during the first day was used for the doubling-time calculations.
Determination of photosynthetic and PSII capacities.
Light-saturated photosynthetic activity in vivo was measured from 1 ml of cell suspension (10 μg Chl/ml) in BG-11 medium supplemented with 10 mM NaHCO3 under saturating light (500 μmol photons m−2 s−1) with a Clark-type oxygen electrode (Hansatech) at 32°C. For PSII capacity measurements, 0.7 mM 2,6-dichloro-p-benzoquinone (DCBQ) was used as an artificial electron acceptor, and 0.7 mM ferricyanide was added to keep the quinone in an oxidized form.
Photoinhibition treatments in vivo.
In the photoinhibition experiments, cell cultures containing 10 μg Chl/ml were illuminated at a PPFD of 1,500 μmol m−2 s−1 with a slide projector at 32°C. One-milliliter samples were drawn for PSII measurements from untreated cultures and after 15, 30, and 45 min of illumination.
Chl a fluorescence measurements.
For nonphotochemical quenching (NPQ) measurements, cells were grown under standard growth conditions and concentrated to an A730 of 2.5. Then, the cultures were further incubated for 1 h under standard growth conditions or illuminated with blue light (HT 119 Dark Blue filter) at 40 μmol photons m−2 s−1. The cell suspension (1 ml) was placed in a temperature-controlled cuvette, and chlorophyll a fluorescence was measured with a PAM-2000 fluorometer (Heinz Walz GmbH) at 32°C. The suspension was first incubated in the dark for 3 min, the initial fluorescence level (F0) was measured, and then the maximum fluorescence level (FM) was measured using a saturating flash of 0.4 s and a PPFD of 5,000 μmol m−2 s−1. Illumination with blue-green light (PPFD, 1,000 μmol m−2 s−1) was initiated, and the maximum fluorescence level in the light (FM′) was measured at 40-s intervals by firing a saturating flash. For state transition measurements at room temperature, the cells were taken from standard growth conditions and incubated in the dark for 3 min. Thereafter, the cell suspension was illuminated for 135 s with blue-green light (PPFD, 80 μmol m−2 s−1) obtained by filtering the output from a slide projector through 400-nm Corion long-pass and 500-nm Corion short-pass filters and thereafter for 195 s with orange light (PPFD, 20 μmol m−2 s−1), obtained through 600-nm Corion long-pass and 650-nm Corion short-pass filters. A saturating flash was fired every 30 s throughout the procedure to measure FM′.
77 K emission spectroscopy.
For state transition measurements with 77 K emission spectroscopy, cells (50 μg Chl/ml; 50-μl samples) were rapidly frozen with liquid nitrogen directly from growth conditions or cells were successively illuminated with blue light (450-nm Corion low-pass filter) at 40 μmol photons m−2 s−1 for 5 min, kept in the dark for 5 min, and illuminated again with blue light for 5 min and then frozen. Fluorescence emission spectra were measured at 77 K with an S2000 spectrometer (Ocean Optics) by exciting the sample with orange light obtained by filtering output from a slide projector through a 580-nm narrow-band filter (Corion). The spectra were corrected by subtracting a low background signal, smoothed with a moving median using a 2-nm window and normalized by dividing by the peak value of PSI emission at 723 nm.
In vivo absorption spectra.
In vivo absorption spectra were measured with a UV-3000 spectrophotometer (Shimadzu, Japan) from 350 nm to 800 nm.
Western blotting.
The cells (25-ml cell culture; A730 = 1) were harvested directly from growth light conditions after 1-h treatments at high light (PPFD, 1,000 μmol m−2 s−1) or blue light (HT 119 Dark Blue filter; 40 μmol photons m−2 s−1) by centrifugation at 10,000 × g for 5 min at 4°C. The cell pellets were washed with ice-cold STNE buffer (0.4 M sucrose, 10 mM Tris-HCl, pH 8.0, 10 mM NaCl, 20 mM Na-EDTA) and resuspended in 150 μl of STNE buffer. One-third volume of acid-washed glass beads (Sigma) was added, and the cells were broken by vortexing them three times for 2 min each time at 4°C; 40 μl of STNE buffer was added, and the glass beads and unbroken cells were removed by centrifugation at 4,000 × g for 4 min at 4°C. The centrifugation was repeated, and the collected supernatant was used as a total-protein sample. For thylakoid isolation, the supernatant was further centrifuged at 18,000 × g for 15 min, and the thylakoid pellet was resuspended in STNE buffer. Chlorophyll concentrations were measured in 80% acetone. Polypeptides were solubilized with Next Gel sample loading buffer (Amresco) for 5 min at 75°C, and samples containing 0.5 μg of Chl a were loaded and separated by 10% Next Gel SDS-PAGE according to the manufacturer's instructions (Amresco). After electrophoresis, the separated proteins were transferred to Immobilon-P membranes (Millipore). The membranes were stained with Ponceau S solution (Sigma) to verify equal loading and even transfer of the samples. The OCP was detected from total-protein samples using a 1:1,500 dilution of an OCP-specific antibody, a generous gift from Diana Kirilovsky. Commercial antibodies (Agrisera) were used to detect the PSII reaction center protein D1 (AS06 124A) and the PSI reaction center protein PsaA (AS06 172) from thylakoid samples. Both antibodies were used at 1:1,000 dilution. The goat anti-rabbit IgG (H+L) alkaline phosphatase conjugate (Zymed) and the CDP star chemiluminescence kit (New England Biolabs) were used for detection in Western blotting. The immunoblots were quantified with a FluorChem image analyzer (Alpha Innotech Corp.).
RESULTS
Triple inactivation strains of group 2 σ factors in Synechocystis.
We constructed all possible triple inactivation strains of group 2 σ factors in Synechocystis. The sigC and sigE genes were inactivated with a chloramphenicol resistance cassette, and transformation of the ΔsigBD, ΔsigBE, and ΔsigDE double inactivation strains (25) resulted in the ΔsigBCD, ΔsigBCE, ΔsigBDE, and ΔsigCDE triple inactivation strains. The complete segregation of the inactivated gene from the native gene was confirmed by PCR analysis (Fig. 1A). Two lines that were descendants of independently raised colonies on the first selection plate were used in measurements to minimize the possibility that secondary mutations would affect the results. However, the two lines behaved similarly, and the results are shown for one line only.
The growth curves of all triple inactivation strains (Fig. 1B) were similar to that of the control strain under standard growth conditions (BG-11 medium, pH 7.5; continuous light; PPFD of 40 μmol m−2 s−1; 32°C; and ambient CO2). Light-saturated photosynthetic and PSII activities were similar in all strains, as well (Table 2). These results, together with the complete segregation of the inactivation strains, show that three out of the four Synechocystis group 2 σ factors can be inactivated simultaneously in any combination without affecting the viability and growth of the cells under standard conditions. All triple inactivation strains grew well in a 12-h light/12-h dark rhythm (Fig. 1B).
TABLE 2.
Photosynthetic and PSII activities of triple inactivation strains of group 2 σ factors in Synechocystis sp. strain PCC 6803
| Strain | Light-saturated photosynthetic activity (μmol O2 mg chl−1 h−1) | Light-saturated PSII activity (μmol O2 mg chl−1 h−1) |
|---|---|---|
| Control | 125.8 ± 7.8 | 387.4 ± 7.6 |
| ΔsigBCD | 125.3 ± 3.0 | 413.1 ± 3.0 |
| ΔsigBCE | 124.6 ± 2.8 | 392.1 ± 9.6 |
| ΔsigBDE | 126.6 ± 1.4 | 387.0 ± 20.1 |
| ΔsigCDE | 132.2 ± 5.3 | 427.5 ± 13.2 |
In laboratory batch cultures of Synechocystis, a short exponential growth phase is followed by a phase of linear growth and finally a stationary phase (4, 25). The doubling times during the first day of growth were measured for autotrophically and mixotrophically growing cells under continuous illumination (PPFD, 40 μmol m−2 s−1) at 32°C. The control and the ΔsigBCE, ΔsigBDE, and ΔsigCDE strains grew faster under mixotrophic than under autotrophic growth conditions (Fig. 1C), while the ΔsigBCD strain grew similarly under autotrophic and mixotrophic conditions.
SigB and SigD factors are involved in acclimation to low-temperature stress.
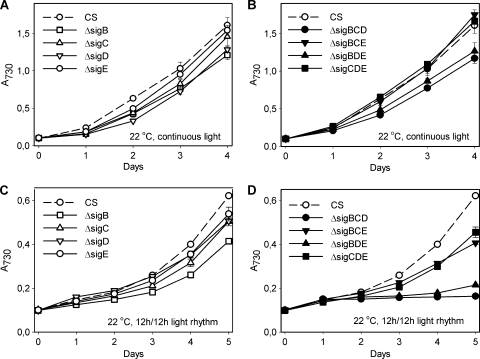
The roles of σ factors in acclimation to low-temperature stress were not tested previously in cyanobacteria. We used 22°C; at this temperature, cells of the control strain grow, but slowly, with a doubling time of circa 19 h. We first tested the single inactivation strains and found that none of them grew as well as the control strain at 22°C, and especially, the ΔsigB and ΔsigD strains grew slowly (Fig. 2 A). In accordance with the results obtained with single inactivation strains, the triple inactivation strains containing only SigC (ΔsigBDE) or only SigE (ΔsigBCD) grew poorly at 22°C, whereas those strains containing only SigD (ΔsigBCE) or only SigB (ΔsigCDE) grew like the control strain (Fig. 2B). These results point to the importance of the SigB and SigD factors for cold acclimation.
FIG. 2.
Growth of single and triple inactivation strains at low temperature. The cell cultures were grown at 22°C in continuous light (A and B) or in a 12-h light/12-h dark rhythm (C and D) at a PPFD of 40 μmol m−2 s−1. Each data point represents the mean of at least three biological replicates with independent liquid cultures, and the error bars denote SE.
Next, low-temperature stress was combined with a light/dark rhythm. All single mutants grew more slowly at low temperature in a light rhythm than the control strain; the ΔsigB strain was the most sensitive (Fig. 2C). The importance of the SigB and SigD factors was obvious under light rhythm at 22°C, as strains that contained only SigC (ΔsigBDE) or SigE (ΔsigBCD) hardly grew at all (Fig. 2D), while strains with only SigB (ΔsigCDE) or only SigD (ΔsigBCE) grew only somewhat more slowly than the control strain, at a rate similar to that of the ΔsigB strain. The SigB and SigD factors showed the highest mutual similarity among all group 2 σ factors (25), and these results suggest that they have functional redundancy under low-temperature conditions. We have also noticed redundancy in their functions under high light, under high-temperature stress, and when very dilute cell cultures are grown (25).
High-light acclimation is disturbed in triple inactivation strains.
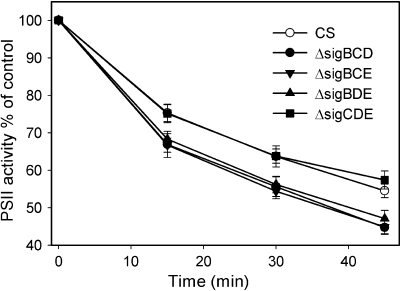
The sensitivities of the strains to high light were studied by subjecting the cells to 1,500 μmol photons m−2 s−1. Photoinhibition was determined as loss of PSII capacity compared to untreated samples. The ΔsigBCD, ΔsigBCE, and ΔsigBDE strains lost their PSII activities 20% faster than the control strain, but the ΔsigCDE strain behaved like the control strain (Fig. 3). These results indicate that the SigB factor is important in bright-light responses in Synechocystis.
FIG. 3.
Photoinhibition of the triple inactivation strains. Cell cultures were illuminated at a PPFD of 1,500 μmol m−2 s−1, and light-saturated PSII activity was measured after 0, 15, 30, and 45 min with a Clark-type oxygen electrode at 32°C using 0.7 mM 2.6-dichlorobentzoquinone as an artificial electron acceptor. PSII activity is expressed as a percentage of the activity measured from untreated control samples. Each data point represents an average of five independent experiments, and the error bars denote SE. After 45 min of illumination, the difference between the control strain and the ΔsigBCD, ΔsigBCE, and ΔsigBDE strains was statistically significant (P < 0.05; Student's t test).
The ΔsigBCE strain has defects in light quality acclimation.
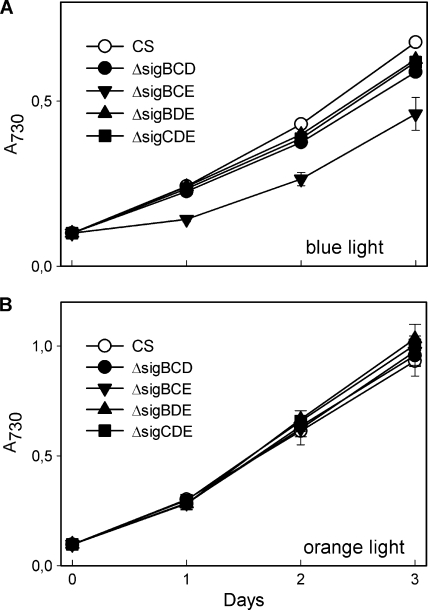
The light quality acclimation capacities of the triple inactivation strains were tested by growing the cells under continuous orange or blue light. The PPFD of blue and orange light was adjusted to 40 μmol photons m−2 s−1, the same as the PPFD of the white light of standard growth conditions. The doubling times of the control strain were 12 h, 15 h, and 19 h under white, orange, and blue light, respectively. These growth rates are in agreement with higher in vivo photosynthetic activity of Synechocystis cells under orange than under blue light (34). The ΔsigBCE strain grew more slowly than the control or the other triple inactivation strains in blue light, with a doubling time of 47 h (Fig. 4 A). In orange light, however, all triple mutants grew like the control strain (Fig. 4B).
FIG. 4.
Growth of the control strain and the triple inactivation strains in blue light, preferentially absorbed by PSI (A), or in orange light, preferentially absorbed by PSII (B). The PPFD was 40 μmol photons m−2 s−1. Each data point represents the mean of three independent cultures, and the error bars denote SE.
In cyanobacteria, the OCP is essential for phycobilisome-associated NPQ, a safety mechanism that reduces energy transfer from the phycobilisomes to the photosystems (39, 40). As NPQ in cyanobacteria is induced by blue-green light (3) and slow growth of the ΔsigBCE strain is a blue-light-specific phenomenon, we compared the OCP content and NPQ in the control and ΔsigBCE strains. Western blot analysis revealed that the amounts of the OCP were similar in the control and ΔsigBCE strains under standard growth conditions but that after a 1-h treatment in high (1,000 μmol photons m−2 s−1) or blue (40 μmol photons m−2 s−1) light, the OCP contents of the ΔsigBCE strain were 108% and 107%, respectively, of that measured for the control strain under the same conditions (Fig. 5 A). For NPQ measurements, cells were first illuminated for 1 h with blue light (40 μmol photons m−2 s−1), F0 and FM were measured with a PAM-2000 fluorometer after 3 min of incubation in the dark, and then the cells were exposed to bright blue-green light (1,000 μmol photons m−2 s−1) under the probe of the fluorometer. Saturating flashes were fired every 40 s to measure FM′. No differences were detected between cells of the different strains in the blue-green-light-induced nonphotochemical fluorescence quenching (Fig. 5B and C). The control and ΔsigBCE strains also behaved similarly when NPQ was measured for cells grown under standard conditions (data not shown).
FIG. 5.
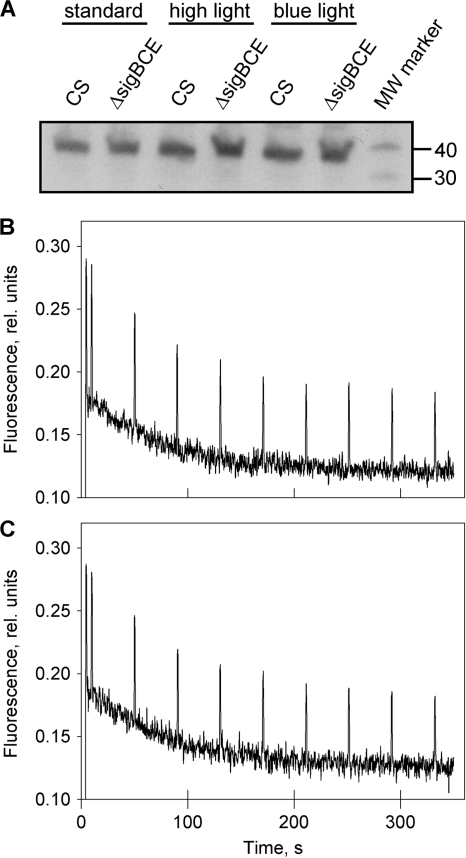
Orange carotenoid protein content and nonphotochemical quenching in the control and ΔsigBCE strains. (A) Western blot showing the amounts of the orange carotenoid protein under standard growth conditions (standard), after 1 h of illumination at a PPFD of 1,000 μmol m−2 s−1 (high light), and after 1 h of illumination with blue light at 40 μmol photons m−2 s−1 (blue light). (B and C) Chlorophyll a fluorescence yield measured with the PAM fluorometer from dark-adapted cells of the control strain (B) or the ΔsigBCE strain (C) during illumination with blue-green light (PPFD, 1,000 μmol m−2 s−1). Each peak corresponds to a saturating flash. rel, relative.
As no differences in NPQ between the ΔsigBCE and control strains were detected, we continued by studying the state transitions that function to balance energy distribution between PSI and PSII. To follow state transitions, fluorescence emission spectra at 77 K were measured from the control and ΔsigBCE strains. For the measurements, cells taken from standard growth conditions or cells that had passed a treatment consisting of 5 min in blue light, 5 min in the dark, and 5 min in blue light again were rapidly frozen in liquid nitrogen. In the dark, cells of the control strain showed a low PSII fluorescence state (state 2), and treatment of the cells with blue light induced a typical high PSII fluorescence state (state 1) (Fig. 6 A). In contrast, the blue-dark-blue treatment did not induce clear state transitions in the ΔsigBCE strain (Fig. 6B). In all measured spectra, the PSII fluorescence peaks at 685 and 695 nm were higher in the ΔsigBCE strain than in the control strain (Fig. 6A and B). Different PSI/PSII ratios in the mutant and control strains could explain this observation. However, in vivo whole-cell absorption spectra were identical in the control and ΔsigBCE strains, indicating similar relative absorption of chlorophyll a and phycobilisomes in both strains (Fig. 6C). Furthermore, the levels of the PSII reaction center protein D1 and the PSI reaction center protein PsaA (Fig. 6D) were virtually identical in the control and ΔsigBCE cells, suggesting similar photosystem stoichiometries in the control and ΔsigBCE strains under standard conditions. Thus, the photosynthetic machinery of the ΔsigBCE strain appears to be locked in state 1.
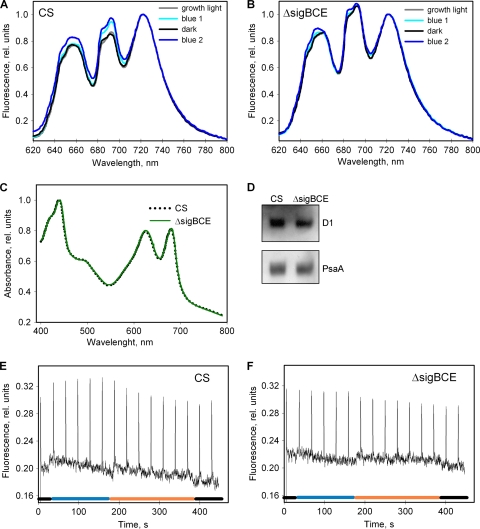
FIG. 6.
State transitions and PSII/PSI ratios in the control and ΔsigBCE strains. Emission spectra were measured at 77 K from control (A) and ΔsigBCE (B) cells taken directly from growth light conditions or from cells illuminated first for 5 min with blue light (blue 1), then incubated for 5 min in the dark, and finally illuminated for 5 min with blue light (blue 2). Light at 590 nm was used for excitation, and the data were normalized by dividing by the height of the photosystem I emission peak at 723 nm. Each spectrum represents an average of four independent experiments. (C) Absorption spectra of control and ΔsigBCE cells grown under standard conditions. The spectra were normalized to the Chl a absorption peak at 440 nm. (D) Amounts of PSII reaction center protein D1 and PSI reaction center protein PsaA. Proteins of isolated thylakoids (0.5 μg of Chl a) were separated by SDS-PAGE, and the amounts of the reaction center proteins were determined by Western blotting using D1 and PsaA protein-specific antibodies. (E and F) Fluorescence yields from dark-adapted control (E) and ΔsigBCE (F) cells were measured with a PAM-2000 fluorometer. The illumination protocol consisted of darkness, blue-green light (PPFD, 80 μmol m−2 s−1), orange light (PPFD, 20 μmol m−2 s−1), and finally darkness, as indicated. A saturating pulse was fired every 30 s to measure FM′.
The state transitions were further studied at room temperature, using a PAM fluorometer. In these measurements, the control strain showed a clear increase of FM′ in blue-green light and a decrease of FM′ in orange light (Fig. 6E). The blue-light-induced increase in FM′ did not occur in the ΔsigBCE strain, and only a very small decrease of FM′ occurred during illumination with orange light (Fig. 6F), confirming that the ΔsigBCE strain is not able to perform normal state transitions.
DISCUSSION
To our knowledge, this is the first time that a complete series of inactivation strains with only one group 2 σ factor left active has been constructed and examined in a cyanobacterium. Summerfield and Sherman (29) constructed a ΔsigBDE triple inactivation strain in Synechocystis and, similar to our results, found no differences in growth under standard conditions between the control and inactivation strains. Double inactivation strains of Synechocystis (25, 29), Anabaena sp. strain PCC 7120 (16), and Synechococcus elongatus PCC 7942 (20) have also been shown to grow well under standard conditions.
Previous studies with the sigE gene inactivation strain revealed that the SigE factor is important for light-activated heterotrophic growth, and SigE has been shown to activate some genes involved in sugar catabolic pathways (23). The ΔsigBCD strain, containing SigE as the only functional group 2 σ factor, was not able to utilize glucose in the light like the other strains (Fig. 1). As all σ factors compete for binding to the same RNA polymerase core, inactivation of three group 2 σ factors could actually promote the binding of the remaining group 2 σ factor. Thus, our finding may suggest that inactivation of the other three group 2 σ factors leads to an overdose of the SigE factor in the RNA polymerase holoenzyme, and this overdose disturbs the utilization of glucose under continuous light. The data strengthen the idea that the SigE factor is an important regulator of sugar metabolism.
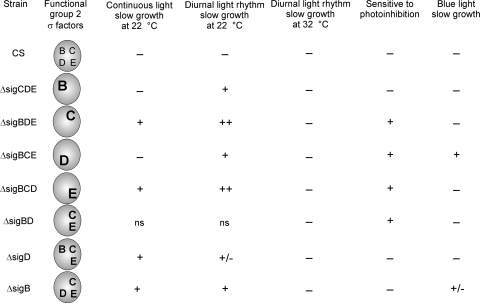
According to the data presented here, group 2 σ factors, particularly SigB and SigD, are important for acclimation to cold and light stresses in Synechocystis. The results are summarized in Fig. 7. Maximal growth of the triple inactivation strains containing either SigB or SigD as the only group 2 σ factor and the slightly retarded growth of the ΔsigB and ΔsigD strains suggest that either SigB or SigD is sufficient to ensure optimal growth at low temperatures if the σ factor does not need to compete with other group 2 σ factors for a limited pool of RNAP core enzymes. In Escherichia coli K-12, the only group 2 σ factor, RpoS, is involved in acclimation to low temperatures. Recent microarray analysis revealed that 7% of the genes in E. coli K-12 are upregulated under cold stress conditions, and 40% of those genes have been shown to be controlled by RpoS under at least one stress condition or during the stationary growth phase (37).
FIG. 7.
Summary of light and low-temperature acclimation properties of different inactivation strains. The results concerning photoinhibition of the ΔsigBD, ΔsigD, and ΔsigB strains were published previously (24); other data are from this study. Symbols indicate no phenotype (−) or a slight (+/−), clear (+), or prominent (++) phenotype. ns, not studied.
Cyanobacteria are unique among eubacteria in having endogenous circadian rhythms (2). This phenomenon has been investigated in detail in the cyanobacterium S. elongatus PCC 7942. Inactivation of any of the four group 2 σ factor genes individually or in pairs or overproduction of the group 2 σ factors altered the circadian expression of different reporter genes in S. elongatus PCC 7942, indicating that group 2 σ factors are involved in circadian regulation but that none of them alone is responsible (20). In Synechocystis, all triple inactivation strains grow well under a diurnal light rhythm at optimal temperature, but they are more sensitive to low temperatures under a light rhythm than under continuous light, suggesting that group 2 sigma factors might also be involved in circadian regulation in Synechocystis, although none of them alone is crucial (Fig. 7).
For light quantity acclimation, the SigB and SigD factors seem to be the most important. The sigB and sigD genes are upregulated upon transfer of Synechocystis cells to high-light conditions (9, 10). A close homolog of SigD, RpoD3 of S. elongatus PCC 7942, is upregulated in high light, and inactivation of the rpoD3 gene results in a light-sensitive phenotype (27). In Synechocystis, the ΔsigBD and ΔsigD strains were not able to enhance growth like the control strain upon doubling of the growth light intensity (24).
Under high-light conditions (PPFD, 1,500 μmol m−2 s−1), the half-life of the D1 protein is only 20 min (33) and the survival of the cells is crucially dependent on efficient repair of photoinhibited PSII. We have previously shown that the double inactivation strain ΔsigBD is sensitive to high light (24). The PSII repair cycle of the ΔsigBD strain does not function efficiently, because transcription of the psbA genes encoding the PSII reaction center protein D1 is not upregulated normally (24). The results of this study show that the SigB factor alone is sufficient to provide normal tolerance for high-light-induced photoinhibition. However, the ΔsigB strain can tolerate high-light treatment as well as the control strain (24), suggesting that, together with the SigC and SigE factors, the SigD factor is able to substitute for the SigB factor under high-light stress.
Another aspect of light acclimation is acclimation to different qualities of light. The slow growth of the ΔsigBCE strain in blue light (with a doubling time 2.5 times longer than in the control strain) might be due to an increased probability of having the SigD factor in the RNA polymerase holoenzyme because the other group 2 σ factors are missing. For a more comprehensive picture of the blue-light phenomenon, growth of the ΔsigBD, ΔsigB, and ΔsigD strains was also measured in blue light. The growth of the ΔsigB strain was slightly reduced (doubling time, 1.14 times longer than in the control strain). In the ΔsigBCE strain, SigD is the only group 2 σ factor, and in the ΔsigB strain, the absence of the SigB factor might promote binding of the other group 2 σ factors, including the SigD factor, to the RNA polymerase core (Fig. 7). In S. elongatus PCC 7942, overproduction of group 2 σ factors under an IPTG (isopropyl-β-d-thiogalactopyranoside)-induced promoter has been shown to affect circadian regulation of reporter genes (20).
The ΔsigBCE strain lacks the capacity for normal state transitions induced by blue light (Fig. 6). Obviously, the absence of state transitions leads to an inability to balance the functions of the two photosystems in blue light, although the ΔsigBCE strain appears to be locked in state 1, which is the state normally induced by blue light. On the other hand, another light-balancing system, the OCP-related NPQ, appears to function normally in the ΔsigBCE strain (Fig. 5). Thus, our results with the ΔsigBCE strain fully agree with the conclusion, drawn from experiments with the ΔOCP strain and phycobilisome and PSII mutants (39, 40), that state transitions and NPQ are separate phenomena in cyanobacteria.
In conclusion, three out of four group 2 σ factors can be inactivated simultaneously in any combination without affecting growth or photosynthetic activity under our standard growth conditions. The presence of either the SigB or the SigD factor is important for low-temperature acclimation, particularly under a diurnal light rhythm. The results obtained with the full series of single, double, and triple inactivation strains suggest that competition of group 2 σ factors for binding to the RNA polymerase core might have an important role. The lack of the other three group 2 σ factors in the ΔsigBCD strain might promote association of the SigE factor with the RNA polymerase core and lead to abnormalities in glucose utilization. Similarly, the competition advantage of the SigD factor in the ΔsigBCE strain leads to slow growth in blue light, apparently because state transitions do not occur normally in the ΔsigBCE strain.
Supplementary Material
Acknowledgments
This study was financially supported by the Academy of Finland and the Finnish Cultural Foundation.
We thank Diana Kirilovsky for the generous gift of the OCP antibody and Jussi Meriluoto for help with the absorption spectra measurements. We thank Paula Mulo for critical reading of the manuscript.
Footnotes
Published ahead of print on 22 October 2010.
Supplemental material for this article may be found at http://jb.asm.org/.
REFERENCES
- 1.Anderson, S. L., and L. McIntosh. 1991. Light-activated heterotrophic growth of the cyanobacterium Synechocystis sp. strain PCC 6803: a blue-light-requiring process. J. Bacteriol. 173:2761-2767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Dong, G., and S. S. Golden. 2008. How a cyanobacterium tells time. Curr. Opin. Microbiol. 11:541-546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.El Bissati, K., E. Delphin, N. Murata, A. Etienne, and D. Kirilovsky. 2000. Photosystem II fluorescence quenching in the cyanobacterium Synechocystis PCC 6803: involvement of two different mechanisms. Biochim. Biophys. Acta 1457:229-242. [DOI] [PubMed] [Google Scholar]
- 4.Foster, J. S., A. K. Singh, L. J. Rothschild, and L. A. Sherman. 2007. Growth-phase dependent differential gene expression in Synechocystis sp. strain PCC 6803 and regulation by a group 2 sigma factor. Arch. Microbiol. 187:265-279. [DOI] [PubMed] [Google Scholar]
- 5.Gunnelius, L., I. Tuominen, S. Rantamäki, M. Pollari, V. Ruotsalainen, E. Tyystjärvi, and T. Tyystjärvi. 2010. SigC sigma factor is involved in acclimation to low inorganic carbon at high temperature in Synechocystis sp. PCC 6803. Microbiology 156:220-229. [DOI] [PubMed] [Google Scholar]
- 6.Havaux, M., G. Guedeney, Q. He, and A. R. Grossman. 2003. Elimination of high-light-inducible polypeptides related to eukaryotic chlorophyll a/b-binding proteins results in aberrant photoacclimation in Synechocystis PCC6803. Biochim. Biophys. Acta 1557:21-33. [DOI] [PubMed] [Google Scholar]
- 7.Havaux, M., G. Guedeney, M. Hagemann, N. Yeremenko, H. C. P. Matthijs, and R. Jeanjean. 2005. The chlorophyll-binding protein IsiA is inducible by high light and protects the cyanobacterium Synechocystis PCC6803 from photooxidative stress. FEBS Lett. 579:2289-2293. [DOI] [PubMed] [Google Scholar]
- 8.Herranen, M., T. Tyystjärvi, and E. M. Aro. 2005. Regulation of photosystem I reaction center genes in Synechocystis sp. strain PCC 6803 during light acclimation. Plant Cell Physiol. 46:1484-1493. [DOI] [PubMed] [Google Scholar]
- 9.Hihara, Y., A. Kamei, M. Kanehisa, A. Kaplan, and M. Ikeuchi. 2001. DNA microarray analysis of cyanobacterial gene expression during acclimation to high light. Plant Cell 13:793-806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Huang, L., M. P. McCluskey, H. Ni, and R. A. LaRossa. 2002. Global gene expression profiles of the cyanobacterium Synechocystis sp. strain PCC 6803 in response to irradiation with UV-B and white light. J. Bacteriol. 184:6845-6858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ikeuchi, M., and S. Tabata. 2001. Synechocystis sp. PCC 6803—a useful tool in the study of the genetics of cyanobacteria. Photosynth. Res. 70:73-83. [DOI] [PubMed] [Google Scholar]
- 12.Imamura, S., M. Asayama, H. Takahashi, K. Tanaka, H. Takahashi, and M. Shirai. 2003. Antagonistic dark/light-induced SigB/SigD, group 2 sigma factors, expression through redox potential and their roles in cyanobacteria. FEBS Lett. 554:357-362. [DOI] [PubMed] [Google Scholar]
- 13.Imamura, S., S. Yoshihara, S. Nakano, N. Shiozaki, A. Yamada, K. Tanaka, H. Takahashi, M. Asayama, and M. Shirai. 2003. Purification, characterization, and gene expression of all sigma factors of RNA polymerase in a cyanobacterium. J. Mol. Biol. 325:857-872. [DOI] [PubMed] [Google Scholar]
- 14.Joshua, S., and C. W. Mullineaux. 2004. Phycobilisome diffusion is required for light-state transitions in cyanobacteria. Plant Physiol. 135:2112-2119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kehoe, D. M., and A. Gutu. 2006. Responding to color: the regulation of complementary chromatic adaptation. Annu. Rev. Plant Biol. 57:127-150. [DOI] [PubMed] [Google Scholar]
- 16.Khudyakov, I. Y., and J. W. Golden. 2001. Identification and inactivation of three group 2 sigma factor genes in Anabaena sp. strain PCC 7120. J. Bacteriol. 183:6667-6675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li, L., R. M. Alvey, R. P. Bezy, and D. M. Kehoe. 2008. Inverse transcriptional activities during complementary chromatic adaptation are controlled by the response regulator RcaC binding to red and green light-responsive promoters. Mol. Microbiol. 68:286-297. [DOI] [PubMed] [Google Scholar]
- 18.Lonetto, M., M. Gribskov, and C. A. Gross. 1992. The σ70: sequence conservation and evolutionary relationships. J. Bacteriol. 174:3843-3849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Maeda, H., N. Fujita, and A. Ishihama. 2000. Competition among seven Escherichia coli σ subunits: relative binding affinities to the core RNA polymerase. Nucleic Acids Res. 28:3497-3503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nair, U., J. L. Ditty, H. Min, and S. S. Golden. 2002. Roles for sigma factors in global circadian regulation of the cyanobacterial genome. J. Bacteriol. 184:3530-3538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nyström, T. 2004. Growth versus maintenance: a trade-off dictated by RNA polymerase availability and sigma factor competition? Mol. Microbiol. 54:855-862. [DOI] [PubMed] [Google Scholar]
- 22.Osanai, T., M. Ikeuchi, and K. Tanaka. 2008. Group 2 sigma factors in cyanobacteria. Physiol. Plant 133:490-506. [DOI] [PubMed] [Google Scholar]
- 23.Osanai, T., Y. Kanesaki, T. Nakano, H. Takahashi, M. Asayama, M. Shirai, M. Kanehisa, I. Suzuki, N. Murata, and K. Tanaka. 2005. Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 sigma factor SigE. J. Biol. Chem. 280:30653-30659. [DOI] [PubMed] [Google Scholar]
- 24.Pollari, M., V. Ruotsalainen, S. Rantamäki, E. Tyystjärvi, and T. Tyystjärvi. 2009. Simultaneous inactivation of sigma factors B and D interferes with light acclimation of the cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriol. 191:3992-4001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pollari, M., L. Gunnelius, I. Tuominen, V. Ruotsalainen, E. Tyystjärvi, T. Salminen, and T. Tyystjärvi. 2008. Characterization of single and double inactivation strains reveals new physiological roles for group 2 σ factors in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 147:1994-2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rodríguez-Ezpeleta, N., H. Brinkmann, S. C. Burey, B. Roure, G. Burger, W. Löffelhardt, H. J. Bohnert, H. Philippe, and B. F. Lang. 2005. Monophyly of primary photosynthetic eukaryotes: green plants, red algae, and glaucophytes. Curr. Biol. 15:1325-1330. [DOI] [PubMed] [Google Scholar]
- 27.Seki, A., M. Hanaoka, Y. Akimoto, S. Masuda, H. Iwasaki, and K. Tanaka. 2007. Induction of a group 2 σ factor, RPOD3, by high light and the underlying mechanism in Synechococcus elongatus PCC 7942. J. Biol. Chem. 282:36887-36894. [DOI] [PubMed] [Google Scholar]
- 28.Singh, A. K., T. C. Summerfield, H. Li, and L. A. Sherman. 2006. The heat shock response in the cyanobacterium Synechocystis sp. strain PCC 6803 and regulation of gene expression by HrcA and SigB. Arch. Microbiol. 186:273-286. [DOI] [PubMed] [Google Scholar]
- 29.Summerfield, T. C., and L. A. Sherman. 2007. Role of sigma factors in controlling global gene expression in light/dark transitions in the cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriol. 189:7829-7840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tuominen, I., E. Tyystjärvi, and T. Tyystjärvi. 2003. Expression of primary sigma factor (PSF) and PSF-like sigma factors in the cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriol. 185:1116-1119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tuominen, I., M. Pollari, E. Tyystjärvi, and T. Tyystjärvi. 2006. The SigB σ factor mediates high-temperature responses in the cyanobacterium Synechocystis sp. PCC6803. FEBS Lett. 580:319-323. [DOI] [PubMed] [Google Scholar]
- 32.Tuominen, I., M. Pollari, E. A. von Wobeser, E. Tyystjärvi, B. W. Ibelings, H. C. P. Matthijs, and T. Tyystjärvi. 2008. Sigma factor SigC is required for heat acclimation of the cyanobacterium Synechocystis sp. strain PCC 6803. FEBS Lett. 582:346-350. [DOI] [PubMed] [Google Scholar]
- 33.Tyystjärvi, T., E. M. Aro, C. Jansson, and P. Mäenpää. 1994. Changes of amino acid sequence in PEST-like area and QEEET motif affect degradation rate of D1 polypeptide in photosystem II. Plant Mol. Biol. 25:517-526. [DOI] [PubMed] [Google Scholar]
- 34.Tyystjärvi, T., I. Tuominen, M. Herranen, E. M. Aro, and E. Tyystjärvi. 2002. Action spectrum of psbA gene transcription is similar to that of photoinhibition in Synechocystis sp. PCC 6803. FEBS Lett. 516:167-171. [DOI] [PubMed] [Google Scholar]
- 35.van Thor, J. J., C. W. Mullineaux, and H. C. P. Matthijs. 1998. Light harvesting and state transitions in cyanobacteria. Botanica Acta 111:430-443. [Google Scholar]
- 36.Wang, Q., S. Jantaro, B. Lu, W. Majeed, M. Bailey, and Q. He. 2008. The high light-inducible polypeptides stabilize trimeric photosystem I complex under high light conditions in Synechocystis PCC 6803. Plant Physiol. 147:1239-1250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.White-Ziegler, C. A., S. Um, N. M. Perez, A. L. Berns, A. J. Malhowski, and S. Young. 2008. Low temperature (23°C) increases expression of biofilm-, cold-shock- and RpoS-dependent genes in Escherichia coli K-12. Microbiology 154:148-166. [DOI] [PubMed] [Google Scholar]
- 38.Williams, J. G. K. 1988. Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol. 167:766-778. [Google Scholar]
- 39.Wilson, A., G. Ajlani, J. M. Verbavatz, I. Vass, C. A. Kerfeld, and D. Kirilovsky. 2006. A soluble carotenoid protein involved in phycobilisome-related energy dissipation in cyanobacteria. Plant Cell 18:992-1007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wilson, A., C. Punginelli, A. Gall, C. Bonetti, M. Alexandre, J. M. Routaboul, C. A. Kerfeld, R. van Grondelle, B. Robert, J. T. Kennis, and D. Kirilovsky. 2008. A photoactive carotenoid protein acting as light intensity sensor. Proc. Natl. Acad. Sci. U. S. A. 105:12075-12080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Yeremenko, N., R. Kouřil, J. A. Ihalainen, S. D'Haene, N. van Oosterwijk, E. G. Andrizhiyevskaya, W. Keegstra, H. L. Dekker, M. Hagemann, E. J. Boekema, H. C. Matthijs, and J. P. Dekker. 2004. Supramolecular organization and dual function of the IsiA chlorophyll-binding protein in cyanobacteria. Biochemistry 43:10308-10313. [DOI] [PubMed] [Google Scholar]
- 42.Yoshimura, T., S. Imamura, K. Tanaka, M. Shirai, and M. Asayama. 2007. Cooperation of group 2 σ factors, SigD and SigE for light-induced transcription in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett. 581:1495-1500. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.