Abstract
The impact of antiretroviral therapy (ART) on the genetics of simian immunodeficiency virus (SIV) or human immunodeficiency virus (HIV) populations has been incompletely characterized. We analyzed SIV genetic variation before, during, and after ART in a macaque model. Six pigtail macaques were infected with an SIV/HIV chimeric virus, RT-SHIVmne, in which SIV reverse transcriptase (RT) was replaced by HIV-1 RT. Three animals received a short course of efavirenz (EFV) monotherapy before combination ART was started. All macaques received 20 weeks of tenofovir, emtricitabine, and EFV. Plasma virus populations were analyzed by single-genome sequencing. Population diversity was measured by average pairwise difference, and changes in viral genetics were assessed by phylogenetic and panmixia analyses. After 20 weeks of ART, viral diversity was not different from pretherapy viral diversity despite more than 10,000-fold declines in viremia, indicating that, within this range, there is no relationship between diversity and plasma viremia. In two animals with consistent SIV RNA suppression to <15 copies/ml during ART, there was no evidence of viral evolution. In contrast, in the four macaques with viremias >15 copies/ml during therapy, there was divergence between pre- and during-ART virus populations. Drug resistance mutations emerged in two of these four animals, resulting in virologic failure in the animal with the highest level of pretherapy viremia. Taken together, these findings indicate that viral diversity does not decrease with suppressive ART, that ongoing replication occurs with viremias >15 copies/ml, and that in this macaque model of ART drug resistance likely emerges as a result of incomplete suppression and preexisting drug resistance mutations.
Studying the impact of antiretroviral therapy (ART) on human immunodeficiency virus type 1 (HIV-1) replication and genetic diversity in vivo is important for understanding the selection of drug resistance and mechanisms of viral persistence. It is likely that the success of combination ART is due to the low probability that multiple resistance mutations preexist on one viral genome and that complete or nearly complete suppression of viral replication is achieved with ART (8, 11, 14, 31, 40). It has been hypothesized that antiretroviral drug resistance mutations exist in patients at low levels prior to treatment (8). For multidrug-resistant variants to emerge during ART, either viral variants with multiple drug resistance mutations that exist prior to therapy must be selected during treatment or ongoing, low-level virus replication and recombination must allow the necessary multiple mutations to accumulate on single viral genomes. Many studies have demonstrated that suboptimal therapy results in the selection of drug-resistant variants as a consequence of incomplete viral suppression (15, 17, 21, 24, 32, 33, 42). For example, the widespread and rapid resistance to monotherapy and dual-drug combination therapy (6, 13, 16, 19, 25, 30, 35, 37, 41) and the emergence of nonnucleoside reverse transcriptase (RT) inhibitor (NNRTI) resistance mutations after single-dose nevirapine to prevent mother-to-child transmission of HIV during childbirth in resource-limited settings (12, 20, 26) support the hypothesis that preexisting resistance and ongoing replication during suboptimal treatment result in the selection of drug-resistant virus and virologic failure. Recent studies have attempted to determine whether the source of persistent viremia during successful combination ART also results from incomplete suppression of virus replication or if residual plasma virus is produced by long-lived, chronically infected cells. Some studies indicate that low-level viral replication may occur in specific anatomical compartments despite suppression of plasma HIV-1 RNA to less than 75 copies per ml by ART (5, 7, 9, 15, 28, 29, 36, 39), whereas others have found no evidence for ongoing HIV-1 replication during suppressive therapy (4, 7, 11, 23, 27, 34, 38, 40). For example, studies by Dinoso et al., McMahon et al., and Gandhi et al. showed no decrease in the level of persistent viremia in patients before, during, or after treatment intensification with several different classes of antiretroviral compounds by using an assay with single RNA copy sensitivity, suggesting a lack of ongoing new rounds of infection during effective ART (11, 14, 31). Bailey et al. investigated plasma and cellular viral sequences obtained by single-genome sequencing during suppressive ART and found that populations often include predominant plasma clones, indicating that viral diversity is lost during therapy (4). These data support the hypothesis that virus replication does not occur during suppressive ART and suggest that persistent viremia may be the result of virus release from long-lived cells.
To further investigate the impact of ART on viral replication, viral diversity, and the emergence of drug resistance, we examined HIV-1 pol genetics prior to, during, and after ART by using an RT-SHIVmne macaque model of ART (2, 3). This model allows for frequent sampling before, during, and after initiating ART and eliminates the impact of interhost diversity of transmitted virus and incomplete adherence to ART. RT-SHIV animal models have been demonstrated to be appropriate systems to investigate viral dynamics, persistent viremia, and emergence of drug resistance (2, 3, 10). Infection of pigtail macaques with RT-SHIVmne results in pretherapy RNA levels, dynamics of viral decay during ART, and levels of suppression of <50 copies/ml, similar to those in HIV-infected humans (3). Because of the widespread use of single-dose nevirapine to prevent mother-to-child transmission of HIV, we also studied the impact of short-course exposure to NNRTI monotherapy prior to initiating combination ART. The impact of ART on virus replication and evolution was investigated here by analyzing an average of 24 single-genome sequences obtained from frequent, longitudinal plasma samples (n = 55) from six animals infected with RT-SHIVmne. We found that complete suppression of virus replication was achieved in two animals with plasma viremia less than 15 copies/ml during ART, whereas evidence for viral evolution was detected in four macaques with viremia exceeding 15 copies/ml during ART. Drug resistance mutations emerged in two of the four animals with ongoing replication during ART, resulting in progressive increases in plasma simian immunodeficiency virus (SIV) RNA in the animal with the highest level of pretherapy viremia.
MATERIALS AND METHODS
Virus.
RT-SHIVmne is a pathogenic SIV/HIV chimeric virus in which SIVmne RT is replaced by HIV-1HXB2 RT (2). The challenge stock was grown on CEMx174 cells, and the titer was determined on TZM-bl cells. The virus was previously shown to be replication competent and pathogenic in pigtail macaques as a useful model to study antiretroviral suppression and emergence of drug resistance (3).
Animals.
Six pigtail macaques were housed at the Washington National Primate Research Center in accordance with American Association for Assessment and Accreditation of Laboratory Animal Care standards. Details of RT-SHIVmne challenge, treatment administration, blood draw/storage, and plasma virus load determination were reported previously (3). The limit of detection for RT-SHIVmne RNA in plasma is 15 copies/ml. To estimate virus population size before initiating ART, we summed all pretherapy SHIVmne RNA measurements (6 measurements per animal) (see Table 2).
Treatment regimen and dosing.
Three 200-mg doses of efavirenz (EFV) (Sustiva; Bristol Myers-Squibb, Princeton, NJ) were administered orally to three macaques (M03250, M04007, and M04008) at 13 weeks (days 91, 92, and 94) after infection (3). At week 17 after infection, all six animals were given daily doses of tenofovir (TNV; 20 mg/kg of body weight) and emtricitabine (FTC; 50 mg/kg) subcutaneously (Gilead Sciences, Foster City, CA) and EFV (200 mg) orally for up to 20 weeks (38). Drug therapy was discontinued for all animals at week 37.
Single-genome sequencing and genetic analyses.
Single-genome sequencing (SGS) of HIV-1 pol was performed as previously described (3). Plasma samples (n = 55) were collected from the six RT-SHIVmne-infected macaques during acute infection, prior to and after initiating short-course EFV monotherapy, prior to initiating ART and after viremia decreased 1,000- to 10,000-fold during ART, during virologic failure, if it occurred (rebound viremia >1,000 copies/ml), and after discontinuing ART (Table 1). An average of 24 single-genome sequences was obtained from each plasma sample for a total of 1,469 sequences. Table 1 shows all samples and the number of sequences obtained for analyses in detail.
TABLE 1.
Macaque plasma samples tested by SGS
| Animal identifier | Sample date (mo/day/yr) | No. of weeks postinfection | Viral load (copies/ml) | Time pointa | No. of RT sequences (no. with drug resistance) |
|---|---|---|---|---|---|
| M03430 | 12/6/2005 | 1 | 4,200,000 | Baseline | 21 (0) |
| 2/27/2006 | 13 | 570 | Pretreatment | 18 (0) | |
| 3/31/2006 | 17.5 | <15 | TNV/FTC/EFV | 2 (0) | |
| 5/2/2006 | 22 | <15 | TNV/FTC/EFV | 2 (0) | |
| 5/9/2006 | 23 | <15 | TNV/FTC/EFV | 1 (0) | |
| 9/5/2006 | 40 | 8,000 | Posttreatment | 18 (0) | |
| M04007 | 12/6/2005 | 1 | 1,500,000 | Baseline | 13 (0) |
| 2/27/2006 | 13 | 67,000 | Baseline prior to EFV only | 19 (0) | |
| 3/27/2006 | 17 | 39,000 | Baseline prior to ART | 39 (0) | |
| 3/31/2006 | 17.5 | 1,900 | TNV/FTC/EFV | 22 (0) | |
| 4/11/2006 | 19 | 140 | TNV/FTC/EFV | 19 (0) | |
| 5/2/2006 | 22 | 110 | TNV/FTC/EFV | 8 (0) | |
| 5/9/2006 | 23 | 120 | TNV/FTC/EFV | 2 (0) | |
| 9/5/2006 | 40 | 13,000 | Posttreatment | 32 (0) | |
| K02396 | 12/6/2005 | 1 | 3,600,000 | Baseline | 16 (0) |
| 2/27/2006 | 13 | 3,100,000 | Pretreatment | 20 (0) | |
| 3/27/2006 | 17 | 2,500,00 | Pretreatment | 23 (0) | |
| 3/13/2006 | 17.5 | 160,000 | TNV/FTC/EFV | 23 (0) | |
| 4/11/2006 | 19 | 17,000 | TNV/FTC/EFV | 24 (0) | |
| 5/2/2006 | 22 | 4,900 | TNV/FTC/EFV | 22 (0) | |
| 5/9/2006 | 23 | 2,100 | TNV/FTC/EFV | 24 (0) | |
| 6/6/2006 | 27 | 740 | TNV/FTC/EFV | 24 (0) | |
| 6/27/2006 | 30 | 430 | TNV/FTC/EFV | 2 (0) | |
| 7/25/2006 | 34 | 250 | TNV/FTC/EFV | 24 (0) | |
| 8/15/2006 | 37 | 750 | TNV/FTC/EFV | 34 (0) | |
| 8/22/2006 | 38 | 240,000 | Posttreatment | 40 (0) | |
| M04033 | 12/6/2005 | 1 | 4,400,000 | Baseline | 19 (0) |
| 2/27/2006 | 13 | 190,000 | Pretreatment | 23 (0) | |
| 3/31/2006 | 17.5 | 4,000 | TNV/FTC/EFV | 24 (0) | |
| 4/11/2006 | 19 | 150 | TNV/FTC/EFV | 10 (0) | |
| 5/2/2006 | 22 | 370 | TNV/FTC/EFV | 27 (0) | |
| 8/29/2006 | 39 | 11,000 | Posttreatment | 33 (0) | |
| M04008 | 12/6/2006 | 1 | 4,500,000 | Baseline | 33 (0) |
| 1/10/2006 | 6 | 1,400,000 | Pretreatment | 33 (0) | |
| 2/27/2006 | 13 | 87,000 | Baseline pre-EFV only | 23 (0) | |
| 3/37/2006 | 17 | 220,000 | Baseline prior to ART | 41 (5) | |
| 4/25/2006 | 21 | 530 | TNV/FTC/EFV | 3 (0) | |
| 5/2/2006 | 22 | 150 | TNV/FTC/EFV | 22 (1) | |
| 5/9/2006 | 23 | 100 | TNV/FTC/EFV | 20 (4) | |
| 6/27/2006 | 30 | 110 | TNV/FTC/EFV | 18 (0) | |
| 7/11/2006 | 32 | 40 | TNV/FTC/EFV | 17 (3) | |
| 8/1/2006 | 35 | 40 | TNV/FTC/EFV | 21 (4) | |
| 8/29/2006 | 39 | 440,000 | Posttreatment | 31 (11) | |
| M03250 | 12/6/2005 | 1 | 2,900,000 | Baseline | 24 (1) |
| 1/10/2006 | 6 | 1,700,000 | Pretreatment | 36 (1) | |
| 2/7/2006 | 10 | 1,500,000 | Pretreatment | 22 (1) | |
| 2/27/2006 | 13 | 1,600,000 | Baseline prior to EFV only | 44 (0) | |
| 3/27/2006 | 17 | 1,200,000 | Baseline prior to ART | 45 (9) | |
| 3/31/2006 | 17.5 | 64,000 | TNV/FTC/EFV | 44 (8) | |
| 4/11/2006 | 19 | 6,800 | TNV/FTC/EFV | 41 (8) | |
| 5/2/2006 | 22 | 4,900 | TNV/FTC/EFV | 37 (10) | |
| 5/9/2006 | 23 | 9,000 | TNV/FTC/EFV | 34 (18) | |
| 5/16/2006 | 24 | 69,000 | TNV/FTC/EFV | 34 (16) | |
| 5/23/2006 | 25 | 1,500,000 | TNV/FTC/EFV | 42 (40) | |
| 5/30/2006 | 26 | 540,000 | TNV/FTC/EFV | 35 (34) |
TNV/FTC/EFV refers to the time point during which TNV/FTC/EFV therapy occurred.
Sequences were aligned using ClustalW. Population genetic diversity was calculated as average pairwise difference (APD) by using MEGA4 (http://www.megasoftware.net) and an in-house program (22). Shifts in population structure were calculated using a population subdivision test for panmixia with a significance level of P values <10−4 (1). The test used was derived from a geographic population structure test proposed by Hudson et al. (18). It compares the average pairwise distances in single-genome sequences obtained from samples taken at different times to distances calculated from imaginary populations containing the same sequences randomly reassigned to two groups. Random mixing of the populations to be compared, reassignment, and distance comparisons are performed many times, generating a P value for the probability that the randomized populations' structures are the same between sets of sequences. We used 10,000 relabelings/permutations to obtain the P values shown (see Fig. 3). Neighbor-joining phylogenetic analyses were done using MEGA4. Drug resistance mutations were identified using the Web-based genotypic resistance interpretation tool available at the Stanford HIV Drug Resistance Database 2009 (http://hivdb.stanford.edu/).
RESULTS
Impact of pretherapy viremia on response to ART.
To investigate the impact of the levels of pretherapy viremia on response to ART, we calculated the integrated pretherapy viremia (sum of SIV RNA levels during pretherapy weeks 1 to 13) for the six macaques and related this to the mean viremia during ART (weeks 17 to 37), the nadir of viremia during ART, and the probability of panmixia between populations prior to and during ART (Table 2). The panmixia test is a sensitive method for detecting population shift from sets of sequences (1). A P value (panmixia probability) of 1 denotes identical virus populations, while probabilities <1 imply a change in population structure. We used probabilities of <10−4 as evidence of significant population change, allowing for possible sampling error by SGS and for multiple nucleotide comparisons. Three groups of animals were identified by this analysis of pretherapy viremia versus response to ART (Fig. 1 and Table 2). Group 1 animals (M03430 and M04007) had the lowest integrated pretherapy viremia, the lowest mean plasma viral RNA during ART, and no evidence for viral evolution during therapy by the panmixia test (Fig. 1A and Table 2). The lack of population shifts in group 1 animals suggests that viral replication was fully suppressed in these animals. One of the animals in group 1 (M04007) was pretreated with short-course (1-week) EFV monotherapy (Fig. 1A) and did not show evidence for selection of NNRTI resistance mutations either by SGS (Table 1) or by allele-specific PCR (ASP) (data reported previously) (3). Failure to select for drug resistance during monotherapy may reflect the absence of preexisting NNRTI mutations in the replicating virus population in this animal, possibly due to relatively small virus population size prior to therapy. Group 2 animals (K02396 and M04033) had higher baseline viremia, higher mean plasma SIV RNA during ART, and evidence for population shift during ART by the panmixia test (Fig. 1B and Table 2). These data suggest that low-level, ongoing replication and viral evolution occurred in group 2 animals despite ART. Despite evidence for ongoing replication during ART in group 2 animals, no drug resistance mutations were identified by single-genome sequencing (Fig. 1B and Table 1) or by ASP (3). Group 2 animals were not treated with short-course EFV monotherapy prior to combination ART (Fig. 1B). Both group 3 animals (M04008 and M03250) were treated with short-course EFV monotherapy prior to combination ART and both developed resistance to EFV (K103N mutation in RT). These animals had high baseline viremia, high mean plasma SIV RNA levels during ART, and evidence of population shift during ART by the panmixia test (Fig. 1C and Tables 1 and 2). One animal (M03250) had a progressive rise in viremia associated with the emergence of multiple, linked drug resistance mutations 5 weeks after initiating ART (Fig. 1C and Table 1). Pretherapy drug resistance mutations were found in this animal (1/24 genomes at week 1 with K101E, 1/36 at week 6 with M184I, and 1/22 at week 10 with L74V) (Table 1). Together, these observations suggest that the level of suppression achieved by ART is likely related to the level of pretherapy viremia and that viral rebound results from both ongoing viral replication during ART and the emergence of preexisting drug-resistant variants.
TABLE 2.
Plasma viremia levels prior to and during ART
| Group | Animal identifiera | Viral load (no. of copies of RNA/ml) |
Lowest viral load (no. of samples with <15 copies/ml) | Probability of panmixiad | ||
|---|---|---|---|---|---|---|
| Integrated pretherapyb | Baseline (wk 17) | Mean during ARTc | ||||
| 1 | M03430 | 5.5 × 106 | 2.3 × 102 | 16 | <15 (18) | 0.9876 |
| M04007‡ | 4.7 × 106 | 3.9 × 104 | 88 | 15 (4) | 0.2648 | |
| 2 | K02396 | 2.8 × 107 | 2.5 × 106 | 4.8 × 103 | 40 (0) | <0.0001 |
| M04033 | 8.5 × 106 | 1.4 × 105 | 2.6 × 102 | <15 (1) | <0.0001 | |
| 3 | M04008‡ | 1.6 × 107 | 2.2 × 105 | 1.8 × 102 | <15 (1) | <0.0001 |
| M03250‡† | 1.4 × 107 | 1.2 × 106 | 2.4 × 105 | 2.9 × 103 (0) | <0.0001 | |
‡, animals that received EFV monotherapy at week 13; †, animal M03250 experienced virologic failure and was euthanized.
Sum of viremia at weeks 1 to 13.
Average of weeks 17 to 37.
Pretherapy (week 13) compared to 1,000- to 10,000-fold decline during ART.
FIG. 1.
Longitudinal changes in diversity of HIV-1 pol and plasma RNA level for animals in group 1 (A), group 2 (B), and group 3 (C). APDs are plotted for each animal (left y axis) as a function of time. Plasma viral RNA copies are plotted (right y axis) versus time for each animal. Samples with undetectable viral RNA (<15 copies/ml) are denoted by open symbols. Gray shading indicates EFV monotherapy (week 13) or ART (weeks 17 to 37). Animal M03250 experienced virologic failure and was euthanized (†). Circled points show samples included in Fig. 2.
Virus diversity before, during, and after ART.
To investigate the impact of ART on the genetic diversity of RT-SHIVmne (including the RT coding region targeted by the antiretrovirals used in this study), we compared the APDs of single-genome sequences obtained prior to, during, and after ART in each of the six animals. Plasma virus populations in all animals were nearly homogeneous 1 week after challenge (mean pol APD = 0.0008) (Fig. 1), similar to that of the inoculum (inoculum APD = 0.0009). Virus diversity increased slowly after infection at an average of 0.00002 nucleotide changes per sequence per day, similar to what has been observed in patients recently infected with HIV-1 (22). By week 13 after infection, prior to initiating ART, diversity had increased significantly from samples collected 1 week postinfection to a mean APD of 0.0023 (Fig. 1). Short-course EFV monotherapy and combination ART were initiated at weeks 13 and 17 postinfection, respectively. The impact of ART on HIV-1 RT genetics was determined by comparing the diversity of plasma virus RNA prior to ART (at week 13) to the diversity measured during ART (Fig. 2). After 20 weeks of ART and up to 10,000-fold decreases in plasma SIV RNA, the diversity of pol sequences during therapy (mean APD = 0.0034) was not lower than that at pretherapy (week 13, APD = 0.0023) for each of the six animals (Fig. 1 and 2). The genetic diversity of virus populations did not decrease even among group 1 animals with viral RNA suppression to <15 copies/ml (Fig. 1 and 2). The viral diversity increased slightly in group 2 and 3 animals, possibly resulting from ongoing viral replication in these animals despite ART. These observations show that there was no obvious, direct relationship between viral diversity and virus population size across the 10,000-fold range of viremia studied.
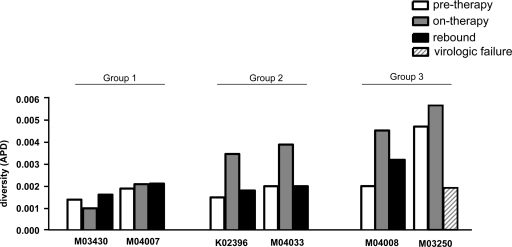
FIG. 2.
Effect of ART on RT-SHIVmne diversity in all animals. Genetic diversities were measured as APDs of sequences obtained from plasma samples collected from the six RT-SHIVmne-infected macaques before initiating treatment (week 13), after a 1,000- to 10,000-fold decline in plasma RNA during ART (weeks 22 to 37), and after stopping ART (weeks 38 to 40). Results are plotted for each time period for animals in group 1, group 2, and group 3.
To investigate the genetics of the rebounding virus population after cessation of ART, the diversity of viral sequences obtained after stopping therapy was compared to that prior to and during ART (Fig. 2). APDs of sequences obtained during viremia rebound from the six animals (mean pol APD = 0.0023) were not different from those of pretherapy sequences, further illustrating that 20 weeks of ART did not affect the diversity of HIV-1 pol. Although the overall diversity of virus populations was not significantly reduced at 20 weeks of ART, drug resistance mutations did emerge in group 3 animals (M03250 and M04008) (Table 1). K103N emerged following EFV monotherapy in both animals and K65R and M184I/V emerged in one animal (M03250) upon viral rebound (3) after about 5 weeks of ART. K013N, K65R, and M184I or -V were present on the same genomes. A genetic bottleneck in pol but not in env (data not shown) accompanied the selection of multidrug-resistant variants in this animal (APD = 0.0056 during ART, APD = 0.0013 at failure) (Fig. 2). This same animal had developed the greatest viral diversity in samples taken prior to the short-course EFV monotherapy (pol APD = 0.0058 by week 6) (Fig. 1) compared to that for all other animals (mean pol APD at week 13 = 0.0018). The highly diverse virus populations in this animal were largely due to frequent G-to-A changes (76% of pretherapy mutations at the peak of diversity at week 6 were G-to-A mutations, compared to 24% in other animals). Two sequences obtained from the week 6 plasma sample from animal M03250 were G-to-A hypermutants, and three sequences contained stop codons due to G-to-A mutations. These data suggest that the high viral diversity developing early after infection in this animal may have resulted, in part, from APOBEC3G or F-induced C-to-U deamination.
Intraanimal viral divergence before, during, and after ART.
To investigate viral evolution during ART, we evaluated population divergence within each animal by using the test for panmixia as well as phylogenetic analyses. Samples resulting in fewer than 10 sequences were excluded from the panmixia analysis because of the potential for sampling error. Single-genome sequences obtained from samples collected prior to, during, and after ART were evaluated for the probability that these populations were the same (panmixia). Panmixia probabilities of virus populations after 20 weeks of ART compared to pretherapy populations in animals from group 1 remained close to 1 (Fig. 3), indicating that significant divergence had not occurred. Phylogenetic analyses confirmed the lack of divergence during ART in these two animals (Fig. 4). Neighbor-joining trees showed that there was no change in population structure between sequences obtained prior to and during therapy. These combined results (panmixia and phylogenetic analyses) indicate that ongoing replication was unlikely during ART in group 1 animals.
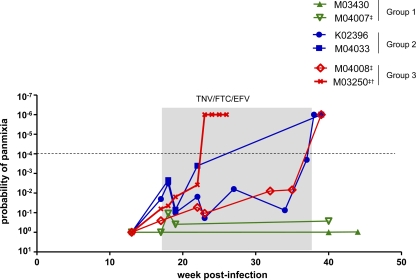
FIG. 3.
Effect of ART on RT-SHIVmne divergence. The significance of divergence was assessed by a test for panmixia of single-genome sequences from the plasma of six RT-SHIVmne-infected macaques before treatment (week 13), during combination ART (weeks 17.5 to 37), and after ART discontinuation (weeks 38 to 40). Shading indicates ART (weeks 17 to 37). The significance cutoff is shown by a dotted line at P values of <0.0001. Animals that received EFV monotherapy at week 13 are denoted by double daggers (‡). Animal M03250 experienced virologic failure and was euthanized at 26 weeks (†).
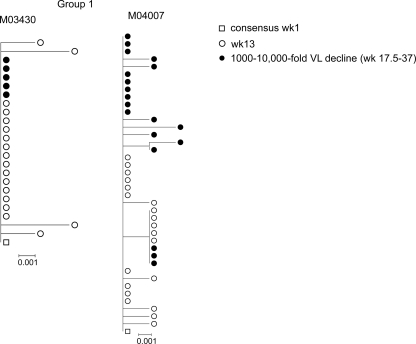
FIG. 4.
Effect of ART on HIV-1 pol evolution analyzed by neighbor-joining phylogenetic analyses of animals with complete suppression of viremia and showing no evidence of a population shift during ART. Trees were rooted on the transmitted viral variant (open squares). Single-genome sequences were from plasma samples obtained before treatment (week 13; open circles) and after a 1,000- to 10,000-fold decline in viremia during ART (week 17.5 to 37; closed circles). M04007 received EFV monotherapy at weeks 13 to 14, but M03430 did not.
In contrast, the emergence of new virus populations was observed in animals in groups 2 and 3. Although these four animals had significant reductions in viremia after initiating ART, plasma virus levels were >15 copies/ml at most time points during therapy (Fig. 1). Both the tests for panmixia (Fig. 3) and phylogenetic analyses (Fig. 5 and 6) detected viral population shifts in all four of these animals during ART. Figure 3 shows the decreasing probability of panmixia in HIV-1 pol in these animals, consistent with ongoing replication and viral divergence. Population divergence in M03250 reached significance during therapy (P < 0.0001), with the emergence of multidrug-resistant variants and viremia rebound. Population shifts reached significant levels in the other three animals soon after discontinuing ART (Fig. 3), showing that rebound virus populations differed significantly from pretherapy populations but not in overall diversity.
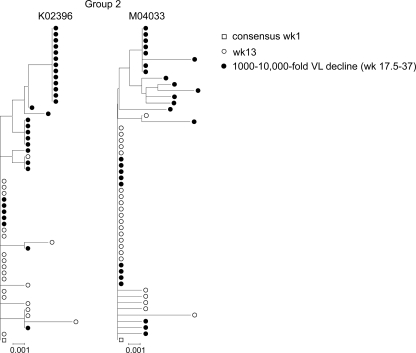
FIG. 5.
Effect of ART on HIV-1 pol evolution analyzed by neighbor-joining phylogenetic analyses of animals with incomplete plasma virus suppression with evidence of population shifts during ART. Neither of these animals received EFV monotherapy prior to ART. Symbols are as described in the legend to Fig. 4.
FIG. 6.
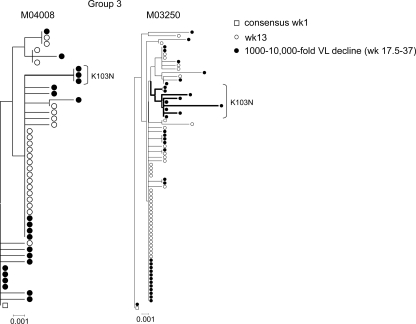
Effect of ART on HIV-1 pol evolution analyzed by neighbor-joining phylogenetic analyses of animals with incomplete plasma virus suppression and drug resistance arising following monotherapy during ART. Both of these animals received EFV monotherapy prior to ART. Animal M03250 experienced virological failure and was euthanized at week 26. Symbols are as described in the legend to Fig. 4.
Phylogenetic trees also showed clear evidence for population shifts during ART in the four incompletely suppressed animals, with distinct nodes of sequences appearing during therapy (Fig. 5 and 6). In group 3 animals, incomplete suppression allowed ongoing virus replication and divergence during ART, resulting in the emergence of variants carrying drug resistance mutations. Both group 3 animals received EFV monotherapy prior to ART, and variants carrying drug resistance mutations are labeled on phylogenetic trees in Fig. 6. Despite evidence for ongoing replication, neither the frequency nor the allele distribution of K103N changed in one of the group 3 animals (M04008) during ART, possibly because the replicating population was too small for a second resistance mutation (M184I/V) to appear, implying the absence of effective selection for the single mutant. These data demonstrate that viral divergence and ongoing replication occur during ART if viral suppression is incomplete but do not necessarily lead to the emergence of variants with resistance to more than one drug within a 20-week period.
Interanimal viral divergence before, during, and after ART.
Phylogenetic analyses were used to assess differences in virus population structures among the six animals before, during, and after ART. Consensus virus populations in all animals were indistinguishable from one another at week 1 postinfection (Fig. 7). Thirteen weeks after infection and prior to initiating therapy, consensus sequences from animals in group 3 clustered together phylogenetically, whereas sequences from the other four animals (groups 1 and 2) did not diverge from the original inoculating virus (Fig. 7). Sequences from group 3 animals were also significantly different from those of group 1 and 2 animals by a test for panmixia (P < 0.0001). Nonsynonymous mutations at codons 75 and 214 in RT primarily distinguished the pretherapy virus population in group 3 animals from that of group 1 and 2 animals. Neither mutation was detected in the virus challenge stock by SGS or by 454 sequencing (detection limit of 0.35%; data not shown). After 20 weeks of ART, virus populations in animals from groups 2 and 3 had diverged from one another and from the pretherapy populations (Fig. 7), further demonstrating that ongoing replication and viral divergence occurred during ART in these animals (test for panmixia, P < 0.0001). No divergence was found between animals in group 1 during ART (test for panmixia, P = 0.0341), again consistent with a lack of virus replication in group 1 animals during therapy. These data further show that viral replication and divergence persisted during ART in the four animals in groups 2 and 3, while full suppression was achieved in the two group 1 animals.
FIG. 7.
Interanimal evolution analyzed by neighbor-joining analyses of consensus sequences from week 1 and week 13, after a 1,000- to 10,000-fold decline in viremia during ART, and after discontinuing ART. Trees were rooted on the consensus sequence of virus in the cell culture supernatant used for the challenge.
DISCUSSION
We investigated HIV-1 pol populations in RT-SHIVmne-infected macaques before, during, and after ART to better understand the impact of therapy on viral genetic diversity and divergence. Longitudinal analysis of plasma virus populations in a well-characterized animal model following defined challenge stock (3) eliminated the effects of interhost diversity of transmitted virus, transmitted drug resistance, and medication nonadherence inherent in patient studies. Previous studies have shown that viral decay dynamics after ART initiation in RT-SHIVmne-infected macaques are similar to the dynamics of HIV-infected humans (3). However, the extent of suppression of viral replication in SHIVmne-infected macaques with ART has not been well defined. The goal of the current study was to analyze viral diversity and divergence to determine the degree to which viral replication was suppressed in SHIVmne-infected macaques with the ART regimen of tenofovir, emtricitabine, and efavirenz, which is highly effective in achieving sustained viral suppression in humans.
After comparing sequences from all animals, we conclude that 20 weeks of ART producing 10,000-fold declines in viremia had no impact on the total virus diversity. This finding indicates that ART inhibits replication of all variants in the virus population similarly. Viral diversity persisted at very low levels of viremia, even in animals with no evidence of ongoing replication during ART. This result implies that both long- and short-lived cells are infected with similarly diverse virus populations. The possibility of emergence of clonal populations, as observed in HIV-infected patients after long-term ART (4), could not be addressed in our study because of the relatively short duration of infection and treatment.
In addition to investigating the impact of ART on total virus diversity, we analyzed sequences for evidence of virus divergence to address the question of ongoing replication during ART. We found that both the level of suppression and the extent of divergence on therapy varied among animals. In two animals (group 1), we saw no evidence for viral divergence or emergence of drug resistance during ART, indicating that there was little, if any, ongoing replication, as reflected by viral sequences in plasma. Both of these animals had the lowest pretherapy viremia and achieved the lowest mean viral RNA levels during treatment, suggesting that the level of viremia prior to initiating therapy is related to the level of viremia during therapy. A study in humans found a similar relationship between pretherapy RNA copy number and the level of persistent viremia during ART (34).
Evidence of ongoing virus replication during ART was found in four animals (groups 2 and 3) despite up to 10,000-fold declines in plasma RNA levels during therapy. Incomplete virus suppression in animals in groups 2 and 3 indicates that the combination ART administered was not sufficiently potent to fully inhibit virus replication in animals with larger pretherapy replicating population sizes. It is likely that a more potent regimen is needed to fully suppress RT-SHIVmne replication in all pigtail macaques. This finding may have broad implications for interpreting results from simian models of ART. Specifically, an ART regimen that is highly effective in suppressing viral replication in humans may be less effective in animal models because of differences in drug dosing and pharmacokinetics.
Despite evidence of ongoing viral replication in four animals, drug-resistant variants emerged in only two animals (group 3). Failure to observe drug resistance in group 2 animals despite incomplete suppression of replication is likely a consequence of reducing the replicating population size to a point lower than the probability that multiple drug resistance mutations will arise on the same genome. Drug resistance did emerge in group 3 animals following short-course EFV monotherapy prior to combination ART. In one of these animals, M03250, viremia progressively rebounded during ART and was primarily associated with two linked mutations (K103N and M184I). However, in the other animal, M04008, viremia did not rebound over the 20-week treatment period despite having equivalent levels of detectable K103N before ART (3). Animal M04008 had pretherapy RNA levels 18-fold lower than those of M03250, a 2.5-fold lower APD, and did not have detectable preexisting drug resistance mutations at week 13, which likely explains the different outcomes during ART between these animals.
The results of this study highlight the importance of pretherapy virus population size, viral diversity, preexisting drug-resistant variants, and the suppressive ART regimen administered on the extent of response to ART in a simian model. Similar studies are in progress to assess the impact of these key variables on the outcome of ART in patients.
Acknowledgments
We thank Ann Wiegand, Valerie Boltz, and Helene Mens for helpful discussions, Robert Stephens for bioinformatics support, and Connie Kinna, Susan Jordan, and Susan Toms for administrative support.
Funding for this research was provided by the National Cancer Institute's intramural Center for Cancer Research, by NIH R01 grant AI080290 (Z.A.), by SAIC contract 25XS119 (J.W.M.), and in part by federal funds from the National Cancer Institute, NIH, under contract HHSN261200800001E. J.M.C. was a recipient of a Research Professorship from the American Cancer Society with support from the George Kirby Foundation.
The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. government.
Footnotes
Published ahead of print on 17 November 2010.
REFERENCES
- 1.Achaz, G., S. Palmer, M. Kearney, F. Maldarelli, J. W. Mellors, J. M. Coffin, and J. Wakeley. 2004. A robust measure of HIV-1 population turnover within chronically infected individuals. Mol. Biol. Evol. 21:1902-1912. [DOI] [PubMed] [Google Scholar]
- 2.Ambrose, Z., V. Boltz, S. Palmer, J. M. Coffin, S. H. Hughes, and V. N. Kewalramani. 2004. In vitro characterization of a simian immunodeficiency virus-human immunodeficiency virus (HIV) chimera expressing HIV type 1 reverse transcriptase to study antiviral resistance in pigtail macaques. J. Virol. 78:13553-13561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ambrose, Z., S. Palmer, V. F. Boltz, M. Kearney, K. Larsen, P. Polacino, L. Flanary, K. Oswald, M. Piatak, Jr., J. Smedley, W. Shao, N. Bischofberger, F. Maldarelli, J. T. Kimata, J. W. Mellors, S. L. Hu, J. M. Coffin, J. D. Lifson, and V. N. KewalRamani. 2007. Suppression of viremia and evolution of human immunodeficiency virus type 1 drug resistance in a macaque model for antiretroviral therapy. J. Virol. 81:12145-12155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bailey, J. R., A. R. Sedaghat, T. Kieffer, T. Brennan, P. K. Lee, M. Wind-Rotolo, C. M. Haggerty, A. R. Kamireddi, Y. Liu, J. Lee, D. Persaud, J. E. Gallant, J. Cofrancesco, Jr., T. C. Quinn, C. O. Wilke, S. C. Ray, J. D. Siliciano, R. E. Nettles, and R. F. Siliciano. 2006. Residual human immunodeficiency virus type 1 viremia in some patients on antiretroviral therapy is dominated by a small number of invariant clones rarely found in circulating CD4+ T cells. J. Virol. 80:6441-6457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Benito, J. M., M. Lopez, S. Lozano, P. Martinez, J. Gonzalez-Lahoz, and V. Soriano. 2004. CD38 expression on CD8 T lymphocytes as a marker of residual virus replication in chronically HIV-infected patients receiving antiretroviral therapy. AIDS Res. Hum. Retroviruses 20:227-233. [DOI] [PubMed] [Google Scholar]
- 6.Caliendo, A. M., and M. S. Hirsch. 1994. Combination therapy for infection due to human immunodeficiency virus type 1. Clin. Infect. Dis. 18:516-524. [DOI] [PubMed] [Google Scholar]
- 7.Chun, T. W., D. C. Nickle, J. S. Justement, J. H. Meyers, G. Roby, C. W. Hallahan, S. Kottilil, S. Moir, J. M. Mican, J. I. Mullins, D. J. Ward, J. A. Kovacs, P. J. Mannon, and A. S. Fauci. 2008. Persistence of HIV in gut-associated lymphoid tissue despite long-term antiretroviral therapy. J. Infect. Dis. 197:714-720. [DOI] [PubMed] [Google Scholar]
- 8.Coffin, J. M. 1995. HIV population dynamics in vivo: implications for genetic variation, pathogenesis, and therapy. Science 267:483-489. [DOI] [PubMed] [Google Scholar]
- 9.Cohen Stuart, J. W., M. D. Hazebergh, D. Hamann, S. A. Otto, J. C. Borleffs, F. Miedema, C. A. Boucher, and R. J. de Boer. 2000. The dominant source of CD4+ and CD8+ T-cell activation in HIV infection is antigenic stimulation. J. Acquir. Immune Defic. Syndr. 25:203-211. [DOI] [PubMed] [Google Scholar]
- 10.Deere, J. D., J. Higgins, E. Cannavo, A. Villalobos, L. Adamson, E. Fromentin, R. F. Schinazi, P. A. Luciw, and T. W. North. 2010. Viral decay kinetics in the highly active antiretroviral therapy-treated rhesus macaque model of AIDS. PLoS One 5:e11640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dinoso, J. B., S. Y. Kim, A. M. Wiegand, S. E. Palmer, S. J. Gange, L. Cranmer, A. O'Shea, M. Callender, A. Spivak, T. Brennan, M. F. Kearney, M. A. Proschan, J. M. Mican, C. A. Rehm, J. M. Coffin, J. W. Mellors, R. F. Siliciano, and F. Maldarelli. 2009. Treatment intensification does not reduce residual HIV-1 viremia in patients on highly active antiretroviral therapy. Proc. Natl. Acad. Sci. U. S. A. 106:9403-9408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Eshleman, S. H., S. E. Hudelson, A. Gupta, R. Bollinger, A. D. Divekar, R. R. Gangakhedkar, S. S. Kulkarni, M. R. Thakar, R. S. Paranjape, and S. Tripathy. 2005. Limited evolution in the HIV type 1 pol region among acute seroconverters in Pune, India. AIDS Res. Hum. Retroviruses 21:93-97. [DOI] [PubMed] [Google Scholar]
- 13.Fitzgibbon, J. E., A. E. Farnham, S. J. Sperber, H. Kim, and D. T. Dubin. 1993. Human immunodeficiency virus type 1 pol gene mutations in an AIDS patient treated with multiple antiretroviral drugs. J. Virol. 67:7271-7275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gandhi, R. T., L. Zheng, R. J. Bosch, E. S. Chan, D. M. Margolis, S. Read, B. Kallungal, S. Palmer, K. Medvik, M. M. Lederman, N. Alatrakchi, J. M. Jacobson, A. Wiegand, M. Kearney, J. M. Coffin, J. W. Mellors, and J. J. Eron. 2010. The effect of raltegravir intensification on low-level residual viremia in HIV-infected patients on antiretroviral therapy: a randomized controlled trial. PLoS Med. 7:e1000321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gunthard, H. F., J. K. Wong, C. C. Ignacio, J. C. Guatelli, N. L. Riggs, D. V. Havlir, and D. D. Richman. 1998. Human immunodeficiency virus replication and genotypic resistance in blood and lymph nodes after a year of potent antiretroviral therapy. J. Virol. 72:2422-2428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hammer, S. M., H. A. Kessler, and M. S. Saag. 1994. Issues in combination antiretroviral therapy: a review. J. Acquir. Immune Defic. Syndr. 7(Suppl. 2):S24-S35; discussion, S35-S37. [PubMed] [Google Scholar]
- 17.Hatano, H., P. Hunt, J. Weidler, E. Coakley, R. Hoh, T. Liegler, J. N. Martin, and S. G. Deeks. 2006. Rate of viral evolution and risk of losing future drug options in heavily pretreated, HIV-infected patients who continue to receive a stable, partially suppressive treatment regimen. Clin. Infect. Dis. 43:1329-1336. [DOI] [PubMed] [Google Scholar]
- 18.Hudson, R. R., D. D. Boos, and N. L. Kaplan. 1992. A statistical test for detecting geographic subdivision. Mol. Biol. Evol. 9:138-151. [DOI] [PubMed] [Google Scholar]
- 19.Jablonowski, H. 1995. Studies of zidovudine in combination with didanosine and zalcitabine. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 10(Suppl. 1):S52-S56. [PubMed] [Google Scholar]
- 20.Jourdain, G., N. Ngo-Giang-Huong, S. Le Coeur, C. Bowonwatanuwong, P. Kantipong, P. Leechanachai, S. Ariyadej, P. Leenasirimakul, S. Hammer, and M. Lallemant. 2004. Intrapartum exposure to nevirapine and subsequent maternal responses to nevirapine-based antiretroviral therapy. N. Engl. J. Med. 351:229-240. [DOI] [PubMed] [Google Scholar]
- 21.Kantor, R., R. W. Shafer, S. Follansbee, J. Taylor, D. Shilane, L. Hurley, D. P. Nguyen, D. Katzenstein, and W. J. Fessel. 2004. Evolution of resistance to drugs in HIV-1-infected patients failing antiretroviral therapy. AIDS 18:1503-1511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kearney, M., F. Maldarelli, W. Shao, J. B. Margolick, E. S. Daar, J. W. Mellors, V. Rao, J. M. Coffin, and S. Palmer. 2009. Human immunodeficiency virus type 1 population genetics and adaptation in newly infected individuals. J. Virol. 83:2715-2727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kieffer, T. L., M. M. Finucane, R. E. Nettles, T. C. Quinn, K. W. Broman, S. C. Ray, D. Persaud, and R. F. Siliciano. 2004. Genotypic analysis of HIV-1 drug resistance at the limit of detection: virus production without evolution in treated adults with undetectable HIV loads. J. Infect. Dis. 189:1452-1465. [DOI] [PubMed] [Google Scholar]
- 24.Kristiansen, T. B., A. G. Pedersen, J. Eugen-Olsen, T. L. Katzenstein, and J. D. Lundgren. 2005. Genetic evolution of HIV in patients remaining on a stable HAART regimen despite insufficient viral suppression. Scand. J. Infect. Dis. 37:890-901. [DOI] [PubMed] [Google Scholar]
- 25.Larder, B. A. 1995. Viral resistance and the selection of antiretroviral combinations. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 10(Suppl. 1):S28-S33. [PubMed] [Google Scholar]
- 26.Lee, E. J., R. Kantor, L. Zijenah, W. Sheldon, L. Emel, P. Mateta, E. Johnston, J. Wells, A. K. Shetty, H. Coovadia, Y. Maldonado, S. A. Jones, L. M. Mofenson, C. H. Contag, M. Bassett, and D. A. Katzenstein. 2005. Breast-milk shedding of drug-resistant HIV-1 subtype C in women exposed to single-dose nevirapine. J. Infect. Dis. 192:1260-1264. [DOI] [PubMed] [Google Scholar]
- 27.Maldarelli, F., S. Palmer, M. S. King, A. Wiegand, M. A. Polis, J. Mican, J. A. Kovacs, R. T. Davey, D. Rock-Kress, R. Dewar, S. Liu, J. A. Metcalf, C. Rehm, S. C. Brun, G. J. Hanna, D. J. Kempf, J. M. Coffin, and J. W. Mellors. 2007. ART suppresses plasma HIV-1 RNA to a stable set point predicted by pretherapy viremia. PLoS Pathog. 3:e46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Martinez, E., M. Arnedo, V. Giner, C. Gil, M. Caballero, L. Alos, F. Garcia, C. Holtzer, J. Mallolas, J. M. Miro, T. Pumarola, and J. M. Gatell. 2001. Lymphoid tissue viral burden and duration of viral suppression in plasma. AIDS 15:1477-1482. [DOI] [PubMed] [Google Scholar]
- 29.Martinez, M. A., M. Cabana, A. Ibanez, B. Clotet, A. Arno, and L. Ruiz. 1999. Human immunodeficiency virus type 1 genetic evolution in patients with prolonged suppression of plasma viremia. Virology 256:180-187. [DOI] [PubMed] [Google Scholar]
- 30.Mayers, D. L., A. J. Japour, J. M. Arduino, S. M. Hammer, R. Reichman, K. F. Wagner, R. Chung, J. Lane, C. S. Crumpacker, G. X. McLeod, et al. 1994. Dideoxynucleoside resistance emerges with prolonged zidovudine monotherapy. The RV43 Study Group. Antimicrob. Agents Chemother. 38:307-314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McMahon, D., J. Jones, A. Wiegand, S. J. Gange, M. Kearney, S. Palmer, S. McNulty, J. A. Metcalf, E. Acosta, C. Rehm, J. M. Coffin, J. W. Mellors, and F. Maldarelli. 2010. Short-course raltegravir intensification does not reduce persistent low-level viremia in patients with HIV-1 suppression during receipt of combination antiretroviral therapy. Clin. Infect. Dis. 50:912-919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Napravnik, S., D. Edwards, P. Stewart, B. Stalzer, E. Matteson, and J. J. Eron, Jr. 2005. HIV-1 drug resistance evolution among patients on potent combination antiretroviral therapy with detectable viremia. J. Acquir. Immune Defic. Syndr. 40:34-40. [DOI] [PubMed] [Google Scholar]
- 33.Nettles, R. E., T. L. Kieffer, R. P. Simmons, J. Cofrancesco, Jr., R. D. Moore, J. E. Gallant, D. Persaud, and R. F. Siliciano. 2004. Genotypic resistance in HIV-1-infected patients with persistently detectable low-level viremia while receiving highly active antiretroviral therapy. Clin. Infect. Dis. 39:1030-1037. [DOI] [PubMed] [Google Scholar]
- 34.Palmer, S., F. Maldarelli, A. Wiegand, B. Bernstein, G. J. Hanna, S. C. Brun, D. J. Kempf, J. W. Mellors, J. M. Coffin, and M. S. King. 2008. Low-level viremia persists for at least 7 years in patients on suppressive antiretroviral therapy. Proc. Natl. Acad. Sci. U. S. A. 105:3879-3884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Raboud, J. M., S. Rae, R. Woods, M. Harris, and J. S. Montaner. 2002. Consecutive rebounds in plasma viral load are associated with virological failure at 52 weeks among HIV-infected patients. AIDS 16:1627-1632. [DOI] [PubMed] [Google Scholar]
- 36.Ruiz, L., J. van Lunzen, A. Arno, H. J. Stellbrink, C. Schneider, M. Rull, E. Castella, I. Ojanguren, D. D. Richman, B. Clotet, K. Tenner-Racz, and P. Racz. 1999. Protease inhibitor-containing regimens compared with nucleoside analogues alone in the suppression of persistent HIV-1 replication in lymphoid tissue. AIDS 13:F1-F8. [DOI] [PubMed] [Google Scholar]
- 37.Shafer, R. W., A. K. Iversen, M. A. Winters, E. Aguiniga, D. A. Katzenstein, and T. C. Merigan. 1995. Drug resistance and heterogeneous long-term virologic responses of human immunodeficiency virus type 1-infected subjects to zidovudine and didanosine combination therapy. The AIDS Clinical Trials Group 143 Virology Team. J. Infect. Dis. 172:70-78. [DOI] [PubMed] [Google Scholar]
- 38.Shen, L., and R. F. Siliciano. 2008. Viral reservoirs, residual viremia, and the potential of highly active antiretroviral therapy to eradicate HIV infection. J. Allergy Clin. Immunol. 122:22-28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Shiu, C., C. K. Cunningham, T. Greenough, P. Muresan, V. Sanchez-Merino, V. Carey, J. B. Jackson, C. Ziemniak, L. Fox, M. Belzer, S. C. Ray, K. Luzuriaga, and D. Persaud. 2009. Identification of ongoing HIV-1 replication in residual viremia during recombinant HIV-1 poxvirus immunizations in patients with clinically undetectable viral loads on durable suppressive HAART. J. Virol. 83:9731-9742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Siliciano, R. F. 2005. Scientific rationale for antiretroviral therapy in 2005: viral reservoirs and resistance evolution. Top. HIV Med. 13:96-100. [PubMed] [Google Scholar]
- 41.Skowron, G. 1992. Treatment of HIV infection: the antiretroviral nucleoside analogues. Nucleoside analogues: monotherapy. Hosp. Pract. (Off. Ed.) 27(Suppl. 2):5-13. [DOI] [PubMed] [Google Scholar]
- 42.Sungkanuparph, S., R. K. Groger, E. T. Overton, V. J. Fraser, and W. G. Powderly. 2006. Persistent low-level viraemia and virological failure in HIV-1-infected patients treated with highly active antiretroviral therapy. HIV Med. 7:437-441. [DOI] [PubMed] [Google Scholar]