Abstract
Human immunodeficiency virus (HIV), hepatitis B virus (HBV), and hepatitis C virus (HCV) are the most prevalent deadly chronic viral diseases. HIV is treated by small molecule inhibitors. HBV is treated by immunomodulation and small molecule inhibitors. HCV is currently treated primarily by immunomodulation but many small molecules are in clinical development. Although HIV is a retrovirus, HBV is a double-stranded DNA virus, and HCV is a single-stranded RNA virus, antiviral drug resistance complicates the development of drugs and the successful treatment of each of these viruses. Although their replication cycles, therapeutic targets, and evolutionary mechanisms are different, the fundamental approaches to identifying and characterizing HIV, HBV, and HCV drug resistance are similar. This review describes the evolution of HIV, HBV, and HCV within individuals and populations and the genetic mechanisms associated with drug resistance to each of the antiviral drug classes used for their treatment.
Keywords: HIV, HBV, HCV, antiviral therapy, drug resistance, evolution, quasispecies
1. Introduction
The human immunodeficiency virus (HIV), the hepatitis B virus (HBV), and the hepatitis C virus (HCV) each cause lifelong human infection and illness. HIV infects nearly 40 million persons and causes about two million deaths per year. HBV infects more than 400 million persons and causes nearly one million deaths per year. HCV infects nearly 200 million persons and is responsible for 50 to 75% of hepatocellular carcinomas in industrialized countries. Antiviral compounds targeting essential enzymes and other viral targets are either licensed or in advanced clinical development for each of these infections.
The development of drug resistance is the most compelling evidence that an antiviral drug acts by specifically inhibiting the virus rather than its cellular host. The genetic mechanisms of antiviral drug resistance are identified during the earliest stages of drug development by in vitro selection experiments and by ex vivo analysis of viruses obtained from individuals receiving antiviral therapy. This review describes the evolution of HIV, HBV, and HCV within individuals and populations and the genetic mechanisms associated with drug resistance to each of the antiviral drug classes (Table 1).
Table 1.
Human Immunodeficiency Virus (HIV), Hepatitis B Virus (HBV), and Hepatitis C Virus (HCV): Replication Characteristics and Antiviral Treatment.
| Virus | Genomic Classification | Intracellular Reservoir | Mutation Rate* | Plasma Levels† | Recombination | Antiviral Drug Classes§ |
|---|---|---|---|---|---|---|
| HIV | Retrovirus | Proviral DNA | 10−5 | 103 − 106 | Major contribution to virus evolution in individuals and populations | Nucleoside RT inhibitors Nonnucleoside RT inhibitors Protease inhibitors Integrase inhibitors Fusion inhibitors CCR5 inhibitors |
| HBV | DS DNA virus with obligate RNA intermediate | Nuclear covalently-closed circular DNA (cccDNA) | 10−5 | 105 − 109 | Possible contribution to virus evolution within individuals | Interferon Nucleoside RT inhibitors |
| HCV | Positive single-stranded RNA virus | None | 10−4 − 10−5 | 104 − 107 | Possible contribution to virus evolution within individuals | Interferon + Ribavirin Protease inhibitors Nucleoside inhibitors Nonnucleoside inhibitors NS5A inhibitors Cyclophilin inhibitors |
Mutation rates during a single round of replication have been estimated experimentally for HIV-1. For HBV and HCV these rates have been estimated from mathematical models and comparisons with other viruses.
RNA copies per mL for HIV-1 and DNA copies per mL for HBV. Range encompasses the majority of untreated individuals with ongoing replication.
HCV protease inhibitors are in Phase III clinical trials. HCV nucleoside, nonnucleoside, NS5A, and cyclophilin inhibitors are in Phase II clinical trials.
2. Viral Replication and Persistence
2.1. HIV
HIV enters CD4+ T lymphocytes in a three-step process: gp120 Env binds the CD4 receptor and induces a conformational change that enables it to also bind either the CCR5 or CXCR4 coreceptor. The formation of the gp120-CD4-coreceptor complex exposes the extended form of the transmembrane gp41 protein, which fuses the virus and host cell membranes. Following cell entry and viral disassembly, HIV RT converts two copies of single-stranded RNA into minus-strand DNA and then copies minus-strand DNA to create a DS DNA copy of the viral genome. Integrase (IN) catalyzes the cleavage of conserved dinucleotides from the 3’ ends of double-stranded HIV-1 DNA and remains bound to each of the 3’-ends as this circular pre-integration complex translocates to the nucleus. IN then catalyzes the strand transfer reaction, which leads to the integration of the HIV-1 genome into the host genome.
HIV integration is usually followed by viral transcription, translation, and maturation. The latter is characterized by the cleavage of Gag and Gag-Pol polypeptides by protease into the structural and enzymatic proteins of the newly created virus. However, in a certain proportion of infected cells, particularly in resting CD4+ T cells, HIV persists as an integrated proviral genome. Although many proviral DNA genomes are defective or irreversibly silenced by epigenetic mechanisms, many are also capable of reactivating particularly when the host cell undergoes immune stimulation. This proviral DNA reservoir decays slowly and is only minimally affected by antiretroviral therapy [1,2]. As a result, recurrent viremia and immunological decline ensue whenever therapy is discontinued regardless of the duration of previous virologic suppression.
2.2. HBV
In virions, the HBV genome is a relaxed circular DNA (RC-DNA) molecule that is only partially double stranded. After infection of a hepatocyte, RC-DNA is transported to the nucleus and converted by cellular enzymes into a covalently closed circular DNA molecule (cccDNA). cccDNA serves as the template for transcription and for pre-genomic RNA, which has two possible fates: (i) it can be encapsidated with HBV viral polymerase, serve as the template for minus-strand DNA and RC-DNA, and secreted extracellularly or (ii) it can be recycled back to the nucleus to amplify or replenish the cccDNA pool [3].
HBV cccDNA is highly stable. It can be eliminated by cell turnover, immune mechanisms, or possibly epigenetic silencing [4–7]. Nonetheless, most acutely infected adults clear their infection within six months coincident with the development of antibodies to the HBV envelope S protein (HBsAb seroconversion) and the disappearance of plasma HBsAg and viral DNA. In contrast, perinatally infected newborns, horizontally infected infants, and about 5 to 10% of immunocompetent adults develop persistent infection. Among individuals with persistent infection, spontaneous clearance is uncommon, occurring at a frequency of <1% per year [8].
2.3. HCV
HCV is a positive-sense, single-stranded enveloped virus with a genome of about 9.5 kb. The genome encodes a single large 9.0 kb open-reading frame flanked by conserved 5’- and 3’-untranslated regions. The 5’-untranslated region contains the internal ribosomal entry site (IRES), which is necessary for initiating translation. Viral replication occurs in a membrane-associated cytoplasmic replicase complex, consisting of the nonstructural proteins NS3, NS4A, NS4B, NS5A, and NS5B which directs the synthesis of a negative-strand copy of the genome. The resulting duplex RNA serves as a template for the synthesis of multiple copies of the positive-strand genome for protein production and packaging.
HCV persists in up to 70% of untreated infected persons [9–11]. There is a strong association of HCV clearance with genetic variation in the IL28B gene underscoring the importance of innate immunity in the host response to infection [12]. HCV’s life-long persistence in the majority of infected persons in the absence of treatment is a remarkable demonstration of its ability to evade the innate and adaptive immune system of its host. The absence of a stable intracellular reservoir (in contrast to HIV and HBV), however, makes viral eradication possible.
3. Virus Evolution in Individuals
Viral evolution within an infected host is determined by the number of viral replication cycles, the frequency of nucleotide incorporation errors, the potential for viral recombination, and host-mediated and antiviral selection pressures. Intra-host viral genetic diversity also depends on the time between initial infection and viral sampling and on whether the initial infection was clonal or composed of multiple heterogeneous clones.
Acute HIV infection has been shown to be clonal in the majority of infected patients and oligoclonal in the remaining patients [13,14]. Such data, however, are not generally available for HBV and HCV. Transmitted HIV drug resistance also occurs commonly in many parts of the world but is extremely rare for HBV and HCV.
Intra-host viral genetic diversity differs by genomic region with greater diversity occurring within genes encoding envelope proteins compared with structural or enzymatic proteins. Although synonymous mutation rates are usually greater than nonsynonymous mutation rates, synonymous mutations should not be assumed to be neutral because RNA viruses contain many functional genomic secondary structural elements and potentially structural constraints imposed by viral genome packaging [15,16].
HIV, HBV, and HCV are usually called quasispecies because they exist within individuals as highly heterogeneous virus populations that diversify during the course of infection. Although the extent of genomic diversity in these infections does not meet the original definition of a quasispecies, which requires an effectively infinite population size, population geneticists have nonetheless found quasispecies theory to be useful for finite viral populations with high mutation rates and have generally accepted the use of the term quasispecies when applied to HIV, HBV, and HCV infections [17,18].
3.1. HIV
HIV, like most lentiviruses, replicates throughout the course of infection. Plasma virus RNA levels range from 103 to 106 copies/mL in most untreated infected individuals. The plasma virus half-life is estimated to be about five hours and up to 1010 viruses are produced each day in untreated individuals [19].
Mutations occur at two stages of HIV replication: (i) when RT catalyzes the conversion of the two copies of single-stranded genomic RNA into DS DNA; and (ii) when host DNA-dependent RNA polymerase transcribes viral RNA from provirus. HIV mutation rates per replication cycle have been estimated using intracellular fidelity assays designed to detect either the inactivation of a reporter gene or the reversion of an inactivating mutation in a reporter gene [20]. Based on such studies, the HIV-1 nucleotide misincorporation rate has been estimated to be about 1 × 10−5, which is similar to that of other retroviruses [21,22]. However, not all nucleotide positions mutate at the same rate. Mutations occur at increased rates in homopolymeric regions [23].
Recombination is a feature shared among retroviruses. It occurs because RT switches between two co-packaged SS RNA genomes as it creates a single DNA copy. If the co-packaged SS RNA genomes were derived during infection of a single cell by viruses with different sequences, then recombination during the next cycle of replication produces mosaic viral sequences that may differ from the parental genomes at multiple nucleotide positions [24,25]. Recombination, therefore, has a high potential to shape HIV evolution [26], although its effect is limited by the requirement that different HIV variants infect the same cell and by the possibility that the recombinant progeny may not replicate as well as their non-recombinant precursors [27,28].
HIV is under constant selection pressure to avoid adaptive humoral and cellular host immune defenses, which results in a high frequency of mutation at HLA-compatible cytotoxic T lymphocyte (CTL) epitopes. In individuals infected with a single virus strain, genetic diversity usually increases progressively during the course of infection, by as much as 1% per year in the envelope gene [29]. In the absence of drug selection pressure, the rates of divergence are considerably lower in the enzymatic targets of therapy [30,31]. However, in patients receiving incompletely suppressive antiretroviral therapy, many mutations can develop within days to weeks or months.
APOBEC-mediated deamination of cytidines to uracil in negative-strand DNA molecules of retroviruses, retrotransposons, and hepatitis B is an innate host defense mechanism that results in a marked excess of G-to-A mutations (G→A hypermutation) in viral plus-strand DNA [32]. The antiretroviral effects of APOBEC3G (GG→AG) and APOBEC3F (GA→AA) are so significant that HIV is unable to replicate in the absence of Vif, a protein that neutralizes these enzymes. Although APOBEC-mediated G→A hypermutation of HIV usually results in nonviable viruses relegated to proviral DNA [33,34], the possibility that low-level APOBEC activity may have contributed to virus evolution is suggested by an increased frequency of nonsynonymous mutation at the dinucleotides typically targeted by APOBEC3G and APOBEC3F [35].
3.2. HBV
In the absence of therapy, plasma HBV DNA levels are often as high as 107 to 109 copies/mL and up to 1011 to 1013 virions per day may be produced within infected persons [36–38]. The half-life of HBV has been estimated to range from four hours for plasma viruses [39,40] to up to 24 hours for newly formed virions in the process of being released extracellularly [36,41].
HBV mutations accumulate within individuals at 10−4 to 10−5 substitutions per nucleotide per year [42–44]. On the basis of these data and comparisons with other hepadnaviruses, the mutation rate of HBV per round of replication is estimated to be about 10−5, a rate similar to that of HIV and other retroviruses [45,46]. Phylogenetic analysis of complete HBV genome sequences suggests that recombination has occurred at least several times during the virus’s evolutionary history [47]. However the mechanism by which HBV recombination occurs and the frequency with which it leads to the development of new variants within individuals are not known.
During acute infection, HBV faces selection pressure from the host’s innate and adaptive immune systems [48,49]. However, once infection is established, HBV often evolves to induce an immune tolerant state that may benefit the virus by allowing those infected perinatally to survive to adulthood and to transmit their infection to future generations.
Despite its high rate of mutation and replication, HBV’s evolution is constrained because nearly two-thirds of its genome encodes multiple proteins in overlapping reading frames [50,51]. Therefore, regardless of the rate at which mutations occur, the rate at which they become fixed is lower than that for HIV and HCV. Drug resistance, in particular, evolves much more slowly for HBV than for HIV or HCV because even in the presence of antiviral therapy, drug-susceptible viruses remain capable of producing intracellular virus. NRTI-resistant viruses however are more successful at replenishing the cccDNA pool and at infecting new hepatocytes—the two steps that require reverse transcription of pre-genomic RNA [52].
The mean number of nucleotide differences between plasma virus clones within antiviral-naïve infected persons is often less than 1% in the core and polymerase genes particularly during the immunotolerant stages of infection [53–55]. However, genetic diversity is higher during antiviral treatment failure [55–57] and possibly during acute infection and those stages of infection in which the virus is under immune selection pressure.
APOBEC mediated G→A hypermutation manifests differently in HBV than in HIV. First, it is not caused solely by APOBEC3G and APOBEC3F. Additional APOBEC enzymes—including APOBEC3C, which has no dinucleotide preference—appear to contribute to hypermutation [58]. Second, HBV does not appear to have a defense mechanism (such as Vif) against G→A hypermutation suggesting that APOBEC enzymes are not a critical threat to HBV replication. Third, hypermutated clones are detected at low levels (i.e., 0.1% to 5.0%) in most clinical plasma samples [55,59]. HBV G→A hypermutation is important to recognize because certain drug-resistance mutations (A181T and M204I) are unlikely to be clinically significant if they occur in hypermutated, nonfunctional genomes [60].
3.3. HCV
HCV plasma levels typically range from 104.5 to 106.5 IU units/mL where one IU is about 1 to 5 RNA copies depending on the commercial assay used for quantification [61]. HCV has an estimated half-life of about three hours and, in the absence of antiviral therapy, up to 1012 virions are produced daily [62–65].
Like other RNA-dependent RNA polymerases (RdRp), HCV’s polymerase has a high error rate. Studies of virus evolution during point source outbreaks and over short time spans have shown that HCV accumulates about 1 × 10−3 nucleotide changes per site per year [66,67]. Based on these data and comparisons with related viruses, it has been estimated that 10−4 to 10−5 substitutions occur per nucleotide per round of replication [68], a mutation rate typical of non-retroviral RNA viruses.
Recombination occurs through a process of template switching during replication in many families of positive-strand RNA viruses. However, intra-host recombination has rarely been observed [69] and there have only been several documented inter-genotypic or inter-subtype recombinants [70–72]. The paucity of recombination may reflect the lesser fitness of recombinants compared with their parental strains due to mutational incompatibilities.
Adaptive humoral [73,74] and cellular [75,76] immunity create ongoing antiviral selection pressure throughout HCV infection. HCV also has multiple defense mechanisms against innate intracellular antiviral responses [77], but it is uncertain whether innate immunity influences HCV evolution within individuals. HCV’s ability to respond to external selection pressure is demonstrated by the rapidity with which it can develop resistance to small molecule inhibitors in vitro and in vivo.
HCV quasispecies become increasingly complex during the course of infection. On average, the genetic distance among genomes can range from 5 to 10% in NS5A or to greater than 10% in hypervariable regions of the envelope [78–81].
4. Virus Evolution in Populations
4.1. HIV
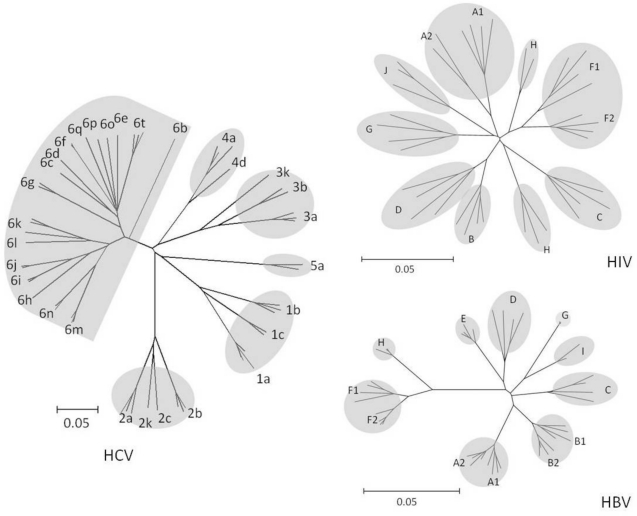
HIV-1 and HIV-2 are two of more than 15 primate lentivirus species that differ from one other by 40 to 60% of their amino acids. HIV-1 groups M and N represent cross-species transmissions from chimpanzees, whereas groups O and P represent cross-species transmission from chimpanzees or gorillas. HIV-1 group M is responsible for the worldwide HIV-1 pandemic; HIV-1 groups O, N, and P are extremely rare. Group M viruses began spreading among humans about 100 years ago and gave rise to multiple subtypes and well-characterized inter-subtype recombinants [82,83]. HIV-1 subtypes differ from each other by about 10 to 30% of their nucleotides throughout their genome. However, within the enzymatic targets of therapy, the inter-subtype diversity averages 10 to 12% at the nucleotide level and 5 to 6% at the amino acid level (Figure 1).
Figure 1.
Phylogenetic Trees Created from HIV-1 Group M RT, HBV RT, and HCV Polymerase Sequences. The trees demonstrate the greater distances separating the HCV genotypes compared with those separating the HIV-1 group M subtypes and the HBV genotypes. Distances were calculated using the HKY85 substitution model with rate variation conforming to a gamma distribution. Trees were constructed using the neighbor-joining algorithm.
The NRTIs, INIs, and—to a lesser extent—the PIs are active against HIV-2 strains in vitro and in vivo and are likely to be active against the rare non-M HIV-1 groups. In contrast, the NNRTIs and the fusion inhibitor enfuvirtide appear to be consistently active only against group M viruses. CCR5 inhibitors should theoretically be active against all HIV-1 strains that must bind the CCR5 receptor. There do not appear to be any consistent differences among group M subtypes in their susceptibility to the six antiretroviral drug classes [84]. However, there are several differences among the subtypes in their propensity to developing specific drug resistance mutations [85–95].
4.2. HBV
HBV infects humans and non-human primates. There are at least eight HBV genotypes, which differ from one other by approximately 8 to 10% of their nucleotides. However, because primate HBV sequences are very similar to non-primate HBV sequences, it is possible that multiple cross-species transmission events occurred and that current HBV strains in humans do not have a single common human virus ancestor (Figure 1).
With the exception of genotype G viruses, which contain a 36-bp insertion in the core gene and two pre-core stop codons and which usually occurs in combination with genotype A viruses [96], there are no proven biological differences among the genotypes. Although several studies have suggested that the HBV genotype may influence disease progression and response to Interferon therapy, few data suggests that genotype influences viral response to NRTI therapy [97–101].
4.3. HCV
There are six major genotypes that differ in their nucleotide sequence by 30% to 35%. Within genotypes, subtypes differ by 20% to 25% [102,103]. Although HCV shares the same basic genomic organization as other flaviviruses, they are only distantly related and the origin of HCV is uncertain (Figure 1). Although there appear to be no differences in clinical severity among the various genotypes, there are major differences in the response to IFN-based therapy [104–106]. Small molecule inhibitors have been targeted towards genotype 1 because this genotype is the most difficult to treat with IFN and Ribavirin and is the most prevalent genotype in the U.S. and Europe.
5. HIV Drug Resistance
Twenty-four antiretroviral drugs belonging to six mechanistic classes have been licensed for HIV-1 treatment: Seven nucleoside and one nucleotide RT inhibitors (NRTIs), nine protease inhibitors (PIs), four non-nucleoside RT inhibitors (NNRTIs), one fusion inhibitor, one IN inhibitor (INI), and one CCR5 inhibitor. In previously untreated individuals infected with drug susceptible HIV-1, combinations of three drugs from two drug classes leads to prolonged virus suppression and, in most patients, immune reconstitution. Once complete HIV-1 suppression is achieved, it usually persists indefinitely as long as therapy is not interrupted [107].
HIV-1 drug resistance may be acquired or transmitted. It is acquired in patients in whom ongoing virus replication occurs in the presence of suboptimal antiviral therapy. Although suboptimal antiviral therapy was once a consequence of an insufficient number of active drugs, it now usually results from treatment interruptions or incomplete adherence. Transmitted drug resistance accounts for about 15% of new infections in the U.S. [108], 10% in Europe [109], 5% in South and Central America, and less than 5% in most parts of Sub-Saharan Africa and South and Southeast Asia [110,111].
5.1. Nucleoside/Nucleotide RT Inhibitors (NRTIs)
The NRTIs are prodrugs that must be triphosphorylated—or in the case of the nucleotide Tenofovir (TDF) diphosphorylated—to their active form. This dependence on intracellular phosphorylation complicates the in vitro assessment of NRTI activity because phosphorylation occurs at different rates in different cell types and leads to discordances between in vitro and in vivo NRTI potency. Specifically, differences in the intracellular dNTP pools between the highly activated lymphocytes used for susceptibility testing and the wider variety of cells that are infected in vivo explain why NRTIs differ in their dynamic susceptibility ranges and in their clinically significant levels of in vitro resistance [112,113]. Clinical isolates from persons failing NRTI therapy may have several-hundred-fold reductions in susceptibility to Zidovudine (AZT), Lamivudine (3TC), and Emtricitabine (FTC), but will rarely have more than five-fold reductions in susceptibility to Didanosine (ddI), Stavudine (d4T), and TDF. However, even slight reductions in in vitro susceptibility to this second category of drugs are clinically significant [114].
There are two biochemical mechanisms of NRTI resistance that are caused predominantly by mutations in the N-terminal polymerase-coding region of HIV-1 RT. One mechanism is mediated by discriminatory mutations that reduce the affinity of RT for an NRTI, preventing its addition to the DNA chain [115]. Another mechanism is mediated by ‘primer-unblocking’ mutations that favor the hydrolytic removal of an NRTI that has been incorporated into the HIV-1 primer chain [112,116]. Because they are selected by the thymidine analog inhibitors AZT and d4T, primer-unblocking mutations are also referred to as thymidine analog mutations or ‘TAMs’.
All recommended first-line treatment regimens include one of the two cytosine analogues—3TC and FTC. Although highly potent, each has a low genetic barrier to resistance. A single mutation, M184V, confers a greater than 200-fold decrease in susceptibility to these drugs. Although M184V limits the effectiveness of 3TC and FTC for salvage therapy, both of these drugs retain some benefit even in the presence of this mutation—possibly as a result of the decreased replication capacity of viruses with M184V or of the fact that M184V increases HIV-1 susceptibility to AZT, d4T, and TDF, drugs that have frequently been used in combination with 3TC and FTC.
The most common TAMs include M41L, D67N, K70R, L210W, T215Y/F, and K219Q/E. A subset of these mutations—M41L, L210W, and T215Y—is particularly important for causing cross-resistance to ddI, Abacavir (ABC), and TDF [117–119]. In patients receiving regimens without thymidine analogs, K65R and L74V have replaced the TAMs as the mutations that occur most commonly in combination with M184V. K65R causes low-level resistance to d4T, intermediate resistance to 3TC and FTC, and high-level resistance to ABC, ddI, and TDF; however, it increases susceptibility to AZT [120,121].
T69SSS and Q151M are multi-NRTI resistance mutations. T69SSS is a double amino insertion at HIV-1 RT position 69. It nearly always occurs with multiple TAMs, where it causes intermediate resistance to 3TC and FTC and high-level resistance to the remaining NRTIs [122,123]. Q151M usually occurs in combination with several otherwise uncommon mutations (A62V, V75I, F77L, and F116Y). It causes intermediate resistance to TDF, 3TC, and FTC, and high-level resistance to the remaining NRTIs [124,125].
Many additional accessory NRTI-resistance mutations have been described, including mutations in the C-terminal regions of HIV-1 RT [126–128]. Most of these C-terminal mutations appear to facilitate primer unblocking by slowing primer/template translocation or RNAseH activity [129,130]. A detailed review of the role of C-terminal mutations in HIV-1 RT drug resistance is also included in this issue [131].
5.2. Nonnucleoside RT Inhibitor Resistance (NNRTIs)
The NNRTIs inhibit HIV-1 RT allosterically by binding to a hydrophobic pocket close to the enzyme’s active site. This binding pocket is less well conserved than the enzyme’s active dNTP-binding site. As a result, group M viruses have greater inter-isolate variability in their susceptibility to NNRTIs than to NRTIs [132]. Three NNRTIs are commonly used: Nevirapine (NVP), Efavirenz (EFV), and Etravirine (ETR).
Many single mutations in the NNRTI-binding pocket confer high-level NVP resistance; several also confer high-level EFV resistance (Table 2). Resistance emerges rapidly when NNRTIs are administered as monotherapy, or in the presence of incomplete virus suppression, which suggests that NNRTI resistance is caused by the selection of rare pre-existing populations of mutant viruses within an individual. The administration of a single dose of NPV to prevent mother-to-child HIV transmission routinely selects for NNRTI-resistant mutants that are detectable by standard sequencing for two months or longer [133,134].
Table 2.
Mechanisms of Resistance to Human Immunodeficiency Virus Type 1 (HIV-1) Inhibitors.
| Drug Class | Mechanism of Resistance | Mutations | Drug Resistance Mutations |
|---|---|---|---|
| Nucleoside/Nucleotide RT inhibitors (NRTIs): Abacavir (ABC) Didanosine (ddI) Emtricitabine (FTC) Lamivudine (3TC) Stavudine (d4T) Zidovudine (AZT) Tenofovir (TDF) |
RT mutations that enhance discrimination between NRTIs and natural nucleosides | K65R, L74V, Y115F, Q151M, M184V | K65R causes high-level resistance to ddI, ABC, and TDF, intermediate resistance to 3TC and FTC, low-level resistance to d4T, and increased susceptibility to AZT. L74V decreases susceptibility to ddI and ABC. Y115F decreases susceptibility to ABC and TDF. Q151M causes high-level resistance to AZT, d4T, ddI, and ABC, and intermediate resistance to TDF, 3TC, and FTC. M184V causes high-level resistance to 3TC and FTC and low-level resistance to ABC and ddI. Reviewed in [177,178]. |
| RT mutations that promote ATP-dependent hydrolytic removal of chain-terminating nucleotide monophosphates(also known as thymidine analog mutationsor TAMs). | M41L, D67N, K70R, L210W, T215F/Y, K219Q/E T69S_SS | M41L, D67N, K70R, L210W, T215FY, and K219QE develop in viruses from patients receiving AZTand d4T. The accumulation of several TAMs causescross-resistance to each of the other NRTIs except 3TC and FTC. T69S_SS is an uncommon amino acid insertion that confers resistance to each of the NRTIs when it occurs in combination with multiple TAMs. Reviewed in [177,178]. | |
| Non-nucleoside RT inhibitors (NNRTIs): Efavirenz (EFV) Etravirine (ETR) Nevirapine (NVP) |
Mutations in the HIV-1 RT NNRTI-binding pocket | L100I, K101E/P, K103N, V106A/M Y181C/I/V, Y188L G190A/S, M230L | These mutations cause high-level resistance to NVP and intermediate or high-level resistance to EFV. With the exception of K103N, V106A/M, and Y188L, each mutation is also associated with decreased ETR susceptibility. Reviewed in [136,141,177]. |
| Protease inhibitors (PIs): Atazanavir (ATV) Darunavir (DRV) Fosamprenavir (FPV) Indinavir (IDV) Lopinavir/r (LPV/r) Nelfinavir (NFV) Saquinavir (SQV) Tipranavir (TPV) |
Protease mutations interfere with inhibitor binding or compensate for the decreased replication associated with other mutations. | D30N, V32I, V47V/A, G48V, I50V/L, I54M/L/V/A/T, L76V, V82A/T/F/S/L, I84V/A, N88S, L90M | Positions 30, 32, 47, 48, 50, 82, and 84 are in the substrate cleft. Position 54 is in the flap and directly interacts with PIs as they enter the substrate cleft. The mutations at positions 76, 88, and 90 influence the shape of the substrate cleft indirectly. Reviewed in [142,177]. |
| These mutations are primarily compensatory | L10I/V/F, L24I, L33F, M46I/L F53L, A71V/T/I/L, Q58E, G73S/T/C/A, T74P, N83D, N88D, L89V |
L10I/V, L33F, M46I/L, and A71V/T are minimally polymorphic occurring in 0.5% to 5% of viruses from untreated persons depending on the subtype. Reviewed in [142,177]. | |
| Integrase inhibitors (INIs): Raltegravir (RAL) In Phase III trials: Elvitegravir (EVG) S/GSK1349572 (572) |
Mutations in residues surrounding the IN active site. | Q148H/R/K ± G140SA, N155H ± E92Q, Y143C/R, T66I/A/K, S147G | Q148H/R/K ± G140SA cause high-level RAL and EVG resistance and intermediate 572 resistance. N155H + E92Q causes high-level RAL and EVG resistance. Y143C/R + T97A causeshigh-level RAL resistance. T66I and S147G are selected in patients receiving EVG and decrease EVG susceptibility but do not appear to cause RAL cross-resistance. Reviewed in [179]. |
| Fusion inhibitors: Enfuvirtide (ENF) | Mutations in the first heptad repeat region (HR1) of the gp41 transmembrane protein interfere with the association of HR1 and HR2 required for virus cell fusion. | G36D/E/V/S, I37V, V38E/A/M/G, Q48H, N42T, N43D/K/S, L44M, L45M | G36D/E, V38E/A, Q40H, and N43D each reduce ENF susceptibility >10-fold [168,169]. Two mutations are usually sufficient to cause high-level ENF resistance. |
| CCR5 inhibitors: Maraviroc (MVC) | Virological failure and resistance is usually caused by expansion of pre-existing CXCR4-tropic variants that were not detected at the start of therapy. In vitro, and occasionally, in vivo resistance is caused by gp120 mutations that facilitate binding to an inhibitor bound CCR5 molecule. | Positively charged residues at positions 11 and 25 of the V3 loop of gp120 and many other combinations of mutations primarily but not exclusively within the V3 loop are associated with CXCR4 tropism [180]. No consistent pattern of gp120 mutations has been identified to be associated with virus binding to an inhibitor-bound CCR5 receptor [174–176]. | |
A minimum of two mutations is required to cause high-level ETR resistance [135,136]. ETR’s increased genetic barrier to resistance is a result of its ability to adopt multiple biding modes within the NNRTI-binding pocket [137]. The NRTIs and NNRTIs are often synergistic. Several NNRTI-resistance mutations increase susceptibility to certain NRTIs [138] and several NRTI-resistance mutations increase NNRTI susceptibility [139,140].
5.3. Protease Inhibitors (PIs)
More than 80 non-polymorphic PI-selected mutations have been reported [127]. Most of these contribute to decreased in vitro susceptibility to one or more PIs [141,142]. The mutations with the greatest impact on susceptibility—D30N, V32I, G48V, I50V/L, V82A/T/L/F/S, and I84V/A—occur in the substrate cleft [142] reducing the binding affinity between the PI and the protease. However, several mutations in the enzyme flap, such as I54M/L, and in the enzyme core, such as L76V, and N88S, can also markedly decrease PI susceptibility (Table 2). Mutations elsewhere in the enzyme either compensate for the decreased kinetics of enzymes with active site mutations; cause resistance by altering enzyme catalysis, dimer stability, and inhibitor binding kinetics; or by re-shaping the active site through long-range structural perturbations [143,144]. Mutations at several protease cleavage sites are also selected during PI treatment, improving the kinetics of protease enzymes with PI-resistance mutations [145–148].
Ritonavir-boosted PIs, particularly lopinavir/r and darunavir/r have the highest genetic barrier to resistance among all antiretrovirals. Multiple mutations are required to compromise their antiretroviral activity [149–153].
5.4. Integrase Inhibitors (INIs)
Although IN catalyzes both the 3’-processing and strand-transfer reactions, only those compounds that specifically inhibit strand transfer are effective INIs [154,155]. The FDA-licensed INI raltegravir and two additional INIs in advanced clinical development—elvitegravir and S/GSK1349572—bind the essential divalent metal cations Mg++ or Mn++ and a hydrophobic region within a cavity formed by IN and the 3’ HIV-1 DNA ends [156,157].
Mutations at nine positions (T66I/A/K, E92Q/V, F121Y, Y143C/R, P145S, Q146P, S147G, Q148H/R/Q, and N155H/S) are selected by raltegravir or elvitegravir and reduce susceptibility to either one or both of these drugs by more than five-fold [158–163]. A large number of secondary compensatory mutations have also been described. The most important of these are G140S/A/C and E138K/A, which increase the fitness of viruses with Q148H/R/K and lead to high-level resistance to all INIs, and T97A, which causes high-level resistance to raltegravir in the presence of Y143C/R [163–166].
5.5. Fusion Inhibitors
Enfuvirtide is a synthetic peptide that inhibits fusion by binding to gp41’s HR1 region and preventing it from folding back and binding to its HR2 region [167]. Enfuvirtide-resistant isolates contain either single or double mutations between positions 36 and 45 of gp41 HR1 [168,169]. Single mutants typically decrease enfuvirtide susceptibility about 10-fold whereas double mutations decrease susceptibility about 100-fold. Despite being one of the most potent antiretroviral drugs, the genetic barrier to enfuvirtide resistance is low and virological rebound emerges rapidly if Enfuvirtide is not administered with a sufficient number of other active inhibitors [170].
5.6. CCR5 Inhibitors
Maraviroc allosterically inhibits the binding of HIV-1 gp120 to the seven-transmembrane G protein-coupled CCR5 receptor [171]. CCR5 inhibitor resistance develops during in vitro passage experiments via gp120 mutations that enable HIV-1 to bind to the CCR5—CCR5-inhibitor complex [172]. Resistance via this mechanism, however, does not occur rapidly nor does it occur by a consistent pattern of gp120 mutations. In patients receiving CCR5 inhibitors, the most common mechanism of virological failure is the expansion of pre-existing CXCR4 tropic viruses that are intrinsically insensitive to CCR5 inhibitors [173]. Less commonly, virological failure emerges via mutations that allow the virus to bind to the CCR5—CCR5-inhibitor complex [174–176].
6. HBV Drug Resistance
There are two forms of Interferon and five nucleoside/nucleotide analogs (NRTIs) licensed for the treatment of chronic HBV infection. Alpha IFN was licensed in 1992 and pegylated alpha IFN 2a was licensed in 2005. The five NRTIs are 3TC (1998), Adefovir (ADV; 2002), Entecavir (ETV; 2005), Telbivudine (LdT; 2006), and TDF (2008). FTC, which is structurally similar to 3TC, is also active against HBV and is frequently used for HBV treatment because it is co-formulated with TDF to treat HIV. 3TC, FTC, and LdT are L-nucleoside analogs; ADV and TDF are acylic nucleotide analogs; and ETV is a deoxyguanosine analog.
3TC, FTC, ADV, and TDF were each originally identified as antiretroviral drugs used for HIV-1. ETV, which was originally reported to be inactive against HIV-1 in vitro, was subsequently shown to reduce plasma HIV-1 RNA levels and to select for the RT mutation M184V in HIV-1 co-infected patients [100].
6.1. Interferon (IFN)
Although NRTIs are used more commonly than α-IFN, pegylated α-IFN is an important option for HBV treatment because unlike the NRTIs, a 24 to 48 week course of therapy is associated with an increased likelihood of sustained virologic response and HBsAb+/HBsAg− seroconversion. Recent pilot studies have also suggested that a combination of NRTIs plus pegylated α-IFN may induce higher rates of sustained response and HBsAb+/HBsAg− seroconversion than pegylated α-IFN alone [181–184].
6.2. Nucleoside/Nucleotide RT Inhibitors (NRTIs)
The three-dimensional structure of HBV RT has not been solved because it has been difficult to obtain sufficient amounts of highly purified active protein. However, homology modeling with other polymerases, including HIV-1 RT, has shown that HBV RT contains regions similar to the fingers, palm, and thumb of HIV-1 and seven sub-domains that are conserved among many published polymerase enzyme sequences [185,186]. In 2001, a standardized numbering system for mutations was established for the RT part of the HBV pol gene [187].
3TC resistance during 3TC monotherapy develops in 15% to 30% of individuals treated for one year, 40% to 50% treated for three years, and 70% treated for five years [188–190]. High-level (>1,000 fold) 3TC resistance is caused by the mutations M204V/I, which are in the YMDD motif adjacent to two of the RT enzyme’s catalytic aspartates [191,192] and likely sterically inhibits HBV RT binding to 3TC [185]. M204 mutations are also frequently accompanied by compensatory mutations, particularly L180M and, less commonly, V173L and/or L80V/I [192–195] (Table 3). M204 mutations are also selected by LdT, albeit at a slower rate than 3TC: 11% versus 26% after two years of monotherapy [196].
Table 3.
Mechanisms of Resistance to Hepatitis B Virus (HBV) Inhibitors.
| Antiviral Agents | Mechanism of resistance | Mutations | Drug Resistance |
|---|---|---|---|
| Interferon | Unknown | Unknown | Unknown |
| Lamivudine (3TC) Telbivudine (LdT) Emtricitabine (FTC)* Entecavir (ETV) Adefovir (ADV) Tenofovir (TDF) |
RT mutations that interfere with nucleotide triphosphate binding. Whether any of these mutations also facilitate primer unblocking is not known. | M204V/I ± L180M ± L80I, V173L | M204V/I ± L180M and less commonly L80I and V173L emerge during 3TC treatment and confer cross-resistance to LdT and FTC; and partial cross-resistance to ETV. M204V/I also emerge during LdT therapy. Reviewed in [215,223,226]. |
| N236T | Selected by ADV and causes partial cross-resistance to TDF. Reviewed in [215,223,226]. | ||
| A181V/T | Selected by ADV and less commonly 3TC. May causes partial cross-resistance to TDF but not ETV. Reviewed in [215,223,226]. | ||
| I169T, T184S/A/G, S202G/I, M250V | Selected by ETV particularly in viruses with pre-existing 3TC-resistance mutations. Reviewed in [215,223,226]. |
FTC is not licensed for HBV treatment. However, it is frequently used in combination with Tenofovir for salvage therapy because there is a co-formulated version of TDF and FTC (Truvada) licensed for the treatment of HIV-1. Several mutations are not shown because they are either extremely rare (e.g., M204S, A181S) or because their association with resistance is controversial: e.g., A233V for Adefovir [227–229], and A194T for Tenofovir [230–231].
Although both HIV-1 and HBV develop 3TC resistance by the substitution of an I or V for an M in their RT’s YMDD motif, the slower development of HBV resistance and HBV’s frequent requirement for compensatory mutations in addition to M204V/I contrasts with the rapid development of 3TC resistance by M184V/I alone in HIV-1-infection.
High-level ETV resistance requires M204V/I + L180M and two or three of the following additional mutations I169T, T184S/A/G, S202G/I, or M250V [197–201]. Virological failure and ETV resistance are exceedingly uncommon when ETV is used to treat NRTI-naïve patients [200]. Although ETV retains considerable antiviral activity against 3TC-resistant variants [199,202], the risk of virological failure and high-level ETV resistance is considerable in 3TC-resistant patients.
ADV resistance emerges more slowly than 3TC resistance. It occurs in about 10% and 30% of individuals receiving ADV monotherapy for two and five years, respectively [189,203–205]. N236T and A181V/T, mutations close to the HBV active site, reduce ADV susceptibility by about 3 to 10-fold [204,206–209]. Although these reductions in susceptibility are much lower than the level of 3TC resistance conferred by M204V/I, they are associated with virologic breakthrough [206,207]. N236T causes partial cross-resistance to TDF but not to 3TC, LdT, or ETV [210].
ADV and TDF retain complete antiviral activity in vitro against viruses with the 3TC-resistance mutations: M204V/I, L180M, V173L, and L80I/V [208,211–213]. Indeed, these mutations may increase HBV susceptibility to ADV and TDF [214–217]. TDF has a high genetic barrier to resistance and the emergence of virological failure and TDF resistance is exceedingly uncommon when TDF is used to treat NRTI-naïve patients.
The HBV RT mutations A181V/T are unique in that they confer resistance to both L-nucleosides and acyclic nucleoside phosphonates and have emerged in individuals receiving ADV and, less commonly, 3TC [208,218–221]. A181T is of particular interest because it causes a stop codon in the reading frame coding for the surface protein, potentially allowing for ongoing hepatocellular replication without accompanying viral load rebound [222]. N236T, and to a lesser extent, A181V/T confer partial cross-resistance to TDF [223–225] and ETV is recommended for patients with these mutations [216].
7. HCV Drug Resistance
Although the combination of Peginterferon-α and Ribavirin is currently the only licensed treatment for HCV [106], many HCV-specific inhibitors are in advanced clinical development. Two PIs, telaprevir (TVR, formerly VX-950) and boceprevir (BVR, formerly SCH-503034) are in Phase III trials and at least 20 additional compounds—PIs, nucleoside inhibitors (NIs), nonnucleoside inhibitors (NNIs), an NS5A inhibitor, and a cyclophilin inhibitor—are in Phase II trials [232–235]. Improved in vitro systems that support intra-cellular replication have been essential to identifying HCV inhibitors and the genetic mechanisms of antiviral drug resistance [236–239].
7.1. Interferon (IFN) and Ribavirin
Pegylated α-IFN plus Ribavirin for six to 12 months is the standard treatment for HCV. Viral factors as well as host factors influence the response to IFN therapy. First, HCV genotype 2 and 3 viruses are significantly more likely than genotype 1 viruses to respond to IFN (sustained virologic response rates are about 70% for types 2 and 3 versus 45% for type 1) [105,106,240,241]. Second, therapy is more successful in acutely infected persons possibly because they harbor less-complex mixtures of quasispecies than do chronically infected persons [242,243]. Indeed, the likelihood of response to IFN is usually inversely proportional to the complexity of its quasispecies [244–248]. Third, specific mutational patterns in a 40-amino-acid region of genotype 1b NS5A have been associated with IFN responsiveness in several studies [249–251]. However, no specific mutations have been shown to be selected by or cause resistance to either IFN or Ribavirin [252–254].
A dose-response relationship exists between the nucleoside analog ribarivin and the likelihood of virologic suppression. Ribavirin interferes with dNTP metabolism by inhibiting cellular inosine monophosphate dehydrogenase [255,256] but may also directly inhibit HCV RNA polymerase [257], increase HCV mutagenesis [258,259], or modulate the HCV T cell immune response [256].
7.2. Protease Inhibitors (PIs)
The NS3 serine protease comprises the 189 N-terminal amino acids of NS3. NS3 forms a heterodimer with the 54-amino-acid NS4A cofactor. The HCV protease cleaves four sites in the HCV polypeptide precursor to generate the N termini of NS4A, NS4B, NS5A, and NS5B. Typical of other members of the trypsin family of serine proteases, NS3/4A contains a catalytic triad composed by H57, D81, and S139. Multiple three-dimensional structures of NS3/4A with and without inhibitors have been determined [260].
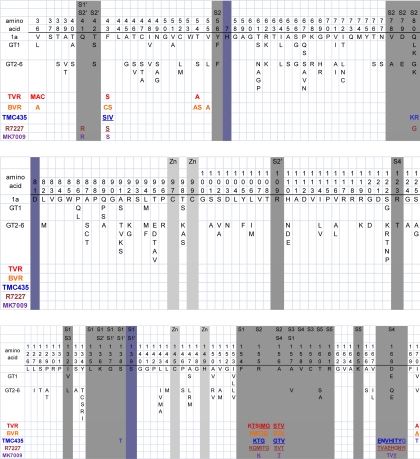
The HCV protease is a challenging drug target because it has a shallow substrate-binding pocket that normally binds a long peptide substrate with which it forms multiple weak interactions [261]. Sequence analysis of individual cleavage sites indicates that the intermolecular consensus sequence is relatively non-conserved: D/E-X-X-X-X-C/T↓A/S-X-X-X where X indicates multiple allowable residues [261,262]. Most HCV PI-resistance mutations occur within or near the substrate binding cleft particularly in the P1 to P4 binding pockets (S1 to S4 subsites; Figure 2). PI-resistance mutations selected in vitro have generally been predictive of those mutations selected in vivo [263].
Figure 2.
HCV NS3 Protease Variability and Protease Inhibitor (PI) Resistance Mutations. Alignment of NS3 residues 36 to 170 showing: (i) The consensus genotype 1a sequence and common variants in genotype 1 (GT1) and genotypes 2 to 6 (GT2–6) according to [277]; (ii) The active site residues are shaded blue-grey; (iii) The substrate binding site positions are shaded grey. The subsite numbering was derived from the following references: [278–282]; (iv) Mutations selected by specific PIs and/or associated with decreased PI susceptibility are indicated beneath the alignment. Underlined positions have been reported to decrease susceptibility >10-fold. PI abbreviations: Telaprevir (TVR), boceprevir (BVR), danoprevir (R7227), and vaniprevir (MK-7009). TMC435 does not have a generic name.
The PIs, TVR and BVR are linear peptidomimetics that bind covalently but reversibly to the active site serine. When combined with α-IFN plus ribavirin, TVR and BVR increase the frequency of sustained virologic response by about 25% compared with placebo [264–267].
PI resistance occurs commonly in those patients who do not achieve a sustained virologic response. The protease mutations associated with resistance to TVR and BVR are nearly completely overlapping. They include V36A/M/L/C, T54A/S, R155K/T, A156S/V/T, and V170A (Table 4) [233,263,268–270]. The emergence of two PI-resistance mutations is generally sufficient for high-level resistance and virological failure. Several macrocyclic PIs including TMC435 [271], vaniprevir (formerly MK-7009) [272], and danoprevir (formerly ITMN-191 and R7227) [273] that do not covalently bind the active site serine are also in Phase II clinical trials. In vitro selection and drug S1’susceptibility studies show that Q41R, F43S, R155K/T, A156S/V/T, and D168A/E/H/T/V/Y are the most important mutations for these inhibitors.
Table 4.
Mechanisms of resistance to Hepatitis C Virus (HCV) inhibitors.
| Antiviral Agents | Mechanism of resistance | Mutations | Drug Resistance | |
|---|---|---|---|---|
| Interferon-α | Unknown | Genotype 1 isolates respond less well than genotype 2 or 3 viruses but the molecular basis is not known. | ||
| Ribavirin | Unknown | Unknown | ||
| PIs: Telaprevir (TVR) Boceprevir (BVR) TMC435 Danoprevir Vaniprevir |
Mutations within or near the protease substrate cleft | V36A/M/C, Q41R, F43S/I/V, T54A/S, Q80K/R, R155K/T, A156S/V/T, D168A/E/I/N/T/V/Y, V170A/T | R155K/T and A156S/V/T decrease susceptibility to all PIs. V36A/M/C and T54A/S decrease susceptibility to the linear peptidomimetics TVR and BVR. V170A is selected by BVR but may cause cross-resistance to TVR. Q41R, F43S/I/V, and D168 mutations decrease susceptibility to TMC435, danoprevir, and vaniprevir. Q80K, a common polymorphism in genotype 1a, and V170T decrease TMC435 susceptibility about 5-fold. [233,263,268,271,272] | |
| NIs: NM203 (withdrawn) R1626 (withdrawn) R7128 MK-0608 |
Steric hindrance of nucleoside analog incorporation (S282T) | S282T | S282T in combination with compensatory mutations has been selected in vitroby 2’-C-methyl modified NIs including valopcitabine (NM203, an oral prodrug of the nucleoside analog 2’-C-methylcytidine) and R7128 (a pro-drug of PSI-6130). | |
| S96T ± N142T | S96T ± N142T are selected in vitro by R1626 (a prodrug of R1479, 4’-azidocytidine). These mutations are far from the HCV polymerase active site. R1626 has been withdrawn from clinical development [288]. | |||
| NNIs | Decreased binding to NNI I pocket (upper thumb) | P495S/A/L, P496S/A, V499A | NNI site 1 is about 30Å from the active site [306]. A series of benzimidazole 5-carboxamide compounds bind to this site [292,307,308]. GS9190, BI207127, and MK3281 are site 1 NNIs in clinical development [68,233]. Mutations at positions 495, 496, and 497 reduce susceptibility to site 1 NNIs [68,233]. | |
| Decreased binding to NNI site 2 (base of thumb) | L419V/M, M423T/V/I, I482L/V/T, V494A/I | NNI site 2 is a shallow hydrophobic pocket at the base of the thumb close to NNI site 1 and ∼35Å from the active site. Compounds that bind to this site such as filibuvir, VC-759, and VCH-796 have selected the mutations L419M, M423T/V/I, I482L, and V194A [68,233,294,309]. | ||
| Decreased binding to NNI site 3 (inner thumb / palm) | H95R, M414T, C451R, G554D, G558R, D559G | Benzothiadizine compounds that bind to this site have selected for M414T, C451R and G558R [310]. ANA598 is a site 3 NNI in Phase II trials [233]. Mutations associated with this drug have include M414T, G554D, and D559G [233]. M414T, which is polymorphic in genotypic 1 viruses, may play a role in resistance to both site 3 and site 4 NNIs. | ||
| Decreased binding to NNI site 4 (palm) | C316N/Y, S365T, L392F, M414T, Y448H | C316Y is selected rapidly in vitro by HCV-796 [311], a site 4 NNI that is no longer in clinical development. Other mutations that have been selected by HCV-796 include C316N, S365T/A, L392F, and M414T [233,311]. ABT-333 is a site 4 NNI that has selected for C316N/Y and Y448H [233]. | ||
| NS5A inhibitors: BMS-790052 | Unknown | M28T, Q30E/H/R, L31M/F/V, P32L, Y93C/H/N | In selection experiments with BMS-790052, M28T, Q30E/H/R, L31M/V, P32L, and Y93C/H/N have been selected in a genotype 1a replicon. L31F/V, P32L, and Y93H/N have been selected in vitro in a genotype 1b replicon. Two mutations are usually required for high-level resistance [302]. | |
| Cyclophilin inhibitors: Debio 025 | Unknown | Unknown | ||
All compounds other than IFN and Ribavirin are in clinical development.
Although the mutations associated with HCV PI resistance are for the most part conserved in genotype 1 viruses, sporadic mutations at these positions have been reported both as majority variants detectable by standard sequencing and as minority variants detected by deep sequencing methods [274,275]. In two studies of HCV protease sequences from more than 1,000 individuals with genotype I viruses, R155K was found in 0.7% of patients and V36M, T54A, D168E, and V170A were found in about 0.5% of patients [274,275]. Specific genotype-associated variants occur at accessory PI-resistance positions [276] and current PIs may have considerably decreased activity against viruses belonging to non-genotype 1 viruses [233].
7.3. Nucleoside (NI) and Non-nucleoside (NNI) Inhibitors
HCV RdRp is encoded by the 530 N-terminal amino acids of the NS5B gene. A C-terminal extension of NS5B anchors the catalytic domain to the endoplasmic reticulum as part of a larger viral replication complex that includes the NS3 RNA helicase. HCV RdRp, like other polymerases, contains palm, thumb, and finger subdomains that enclose the RNA template groove and a GDD catalytic triad [260]. HCV RdRp inhibitors include chain-terminating nucleoside analogs (NIs) and non-nucleoside analogs (NNIs) that target NS5B allosterically. Most HCV NIs differ from HIV-1 and HBV NRTIs in that chain termination is caused by steric hindrance rather than the absence of the 3’-hydroxyl group [68].
In contrast to the HCV PIs, NIs appear to be active against each of the HCV genotypes. The genetic barrier to NI resistance is higher than for the PIs and NNIs with prolonged in vitro passage required for the emergence of resistance [283]. Furthermore, the mutations associated with NI resistance generally reduce viral fitness to a greater extent than the NNIs and PIs [284,285]. Two non-cross-resistant mutational patterns associated with NI resistance have been described (Table 4): the active site mutation S282T has been selected in vitro by 2’-C-methyl modified nucleoside analogs including valopcitabine (NM203, an oral prodrug of the nucleoside analog 2’-C-methylcytidine), R7128 (a pro-drug of PSI-6130), and MK-0608 (2’-C-methyl-7-deaza-adenosine) [284,286]. S282T appears to sterically inhibit NIs containing 2-methyl-substituded nucleoside analogs [287]. S96T ± N142T, which are far from the active site, are selected in vitro by R1626 (a prodrug of R1479, 4’-azidocytidine) [288].
In a Phase II trial, the combination of R1626, Peginterferon alfa-2a, and Ribavirin led to complete virologic suppression by week four in nearly 75% of patients without selecting for S96T ± N142T, the R1626-resistance mutations [289]. In another dose-finding study of 32 patients receiving RG7128 monotherapy for two weeks and 85 patients receiving RG7128 for one month, the vast majority of patients experienced continuous virus load decline proportional to the RG7128 dose without evidence for the emergence of S282T, the R7128-resistance mutation. Similarly, in a dose-ranging study of MK-0608 administered intravenously and orally to chimpanzees for 37 days, plasma HCV levels displayed marked reductions in plasma HCV levels with the development of minority populations of S282T in two chimpanzees.
Although R1626 and valopcitabine have been withdrawn because of toxicity [232], the limited clinical experience with R1626 and the ongoing studies with RG7128 and MK-0608 demonstrates the potential of the HCV RdRp NI class. The absence of significant in vivo resistance to HCV NIs contrasts with the frequent resistance to the NRTIs used in the treatment of HIV and HBV. In a recent two-week clinical trial of the RG7128 in combination with the PI danoprevir (INFORM-1), subjects experienced a 3.7 to 5.2 log10 decrease in HCV IU/mL. Not only did RG7128 mutations fail to emerge, the combination also appeared to prevent the emergence of resistance to the PI danoprevir over the two week trial period [290].
Investigational non-nucleoside inhibitors (NNIs) targeting four allosteric binding sites are in early clinical development [232,291–293]. Two of these sites are in the thumb subdomain and two are in the palm subdomain. Mutations associated with resistance to each of the four allosteric sites have been selected in vitro and/or in vivo [233] (Table 4). Cross-resistance between NNIs and NIs has not been described. However, NNIs have uniformly displayed a low genetic barrier to resistance [294–296] and the activity of NNIs has often been variable even within the same genotype [295,297]. Moreover, several NNI- resistance mutations have been identified in previously untreated individuals either as dominant variants detected by standard sequencing or as minor variants detected by more sensitive methods [275,297,298].
7.4. NS5A Inhibitors
NS5A is a 447 amino acid membrane-associated phosphoprotein that is an essential part of the HCV replicase complex and an antagonist of endogenous IFN. The structure of the N-terminal domain of NS5A has been crystallized but how this domain and the complete protein functions are not known [299,300]. BMS-790052 was identified by a high throughput screening approach for targeting non-enzymatic HCV targets. It has an EC50 of below 10 picomoles in genotype 1a and 1b replicons. It decreased HCV plasma RNA levels about 1,000 fold within 24 hours in a randomized, double-blind, single ascending dose study [235].
The specificity of BMS-790052 for NS5A was demonstrated by the selection of mutations in the N-terminal domain that conferred high-level BMS-790052 resistance [235,301,302]. A combination of two BMS-790052 mutations is required to cause high-level BMS-790052 resistance [302].
7.5. Cyclophilin Inhibitors
Cyclophilin A is an important cellular cofactor for HCV replication. Although the role of cyclophilin A in HCV replication is not known, it appears to a binding partner of NS5A and possibly other HCV proteins [303]. Debio 025 is a non-immunosuppressive cyclosporine analog that potently inhibits the interaction of cyclophilin A and HCV in vitro [303] and in vivo [304]. The genetic barrier to Debio 025 resistance is high [305] and may require the selection of cyclophilin A-independent NS5A variants [303]. Drugs that block the interaction of cyclophilin A with HCV are somewhat analogous to HIV CCR5 inhibitors in that the primary target of therapy is a host protein.
8. Conclusions
Nearly 25 drugs belonging to six drug classes have been licensed for treating HIV-1. In previously untreated individuals infected with drug susceptible HIV-1, combinations of three drugs from two drug classes leads to prolonged virus suppression. However, because HIV cannot be eradicated from its proviral DNA reservoir lifelong therapy is necessary. Prolonged therapy carries the risk that periods of nonadherence will lead to the selection of drug-resistance variants. This risk is particularly high in low-income countries where interruptions in drug supply occur, the regimens used have lower genetic barriers to resistance than those used in high-income countries, and laboratory monitoring is less intensive. Continued research is therefore required to develop fixed-dose drug combinations with high genetic barriers to resistance that can be administered safely for long periods of time.
Although the fewest therapeutic options are available for treating HBV, it is the only virus in this review for which monotherapy—with either ETV or TDF—is capable of fully suppressing virus replication for many years in previously untreated persons. HBV has generally not been considered eradicatable because of its cccDNA form. However, eradication will be attempted by studies that combine pegylated IFN-α and other investigational IFN formulations with the two most potent NRTIs—ETV and TDF.
The first two small molecule HCV inhibitors may be licensed in 2011 and many more are likely to follow in the next decade. The introduction of new inhibitors will increase the frequency of virological cures and reduce HCV morbidity and mortality. It will also lead to widespread acquired drug resistance among the patients who do not achieve sustained virologic response. However, as the number of licensed new non-cross-resistant inhibitors increases, virological failure will decrease in frequency, salvage regimens will be available for patients with resistance to the first generations of small molecule inhibitors, and IFN-sparing regimens will be increasingly used.
Acknowledgments
S. M. T. and R. W. S. were supported in part by NIH grant AI068581.
References and Notes
- 1.Han Y, Wind-Rotolo M, Yang HC, Siliciano JD, Siliciano RF. Experimental approaches to the study of HIV-1 latency. Nat Rev Microbiol. 2007;5:95–106. doi: 10.1038/nrmicro1580. [DOI] [PubMed] [Google Scholar]
- 2.Richman DD, Margolis DM, Delaney M, Greene WC, Hazuda D, Pomerantz RJ. The challenge of finding a cure for HIV infection. Science. 2009;323:1304–1307. doi: 10.1126/science.1165706. [DOI] [PubMed] [Google Scholar]
- 3.Kay A, Zoulim F. Hepatitis B virus genetic variability and evolution. Virus Res. 2007;127:164–176. doi: 10.1016/j.virusres.2007.02.021. [DOI] [PubMed] [Google Scholar]
- 4.Werle-Lapostolle B, Bowden S, Locarnini S, Wursthorn K, Petersen J, Lau G, Trepo C, Marcellin P, Goodman Z, Delaney WEt, et al. Persistence of cccDNA during the natural history of chronic hepatitis B and decline during adefovir dipivoxil therapy. Gastroenterology. 2004;126:1750–1758. doi: 10.1053/j.gastro.2004.03.018. [DOI] [PubMed] [Google Scholar]
- 5.Levrero M, Pollicino T, Petersen J, Belloni L, Raimondo G, Dandri M. Control of cccDNA function in hepatitis B virus infection. J Hepatol. 2009;51:581–592. doi: 10.1016/j.jhep.2009.05.022. [DOI] [PubMed] [Google Scholar]
- 6.Yang PL, Althage A, Chung J, Maier H, Wieland S, Isogawa M, Chisari FV. Immune effectors required for hepatitis B virus clearance. Proc Natl Acad Sci U S A. 2010;107:798–802. doi: 10.1073/pnas.0913498107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Guidotti LG, Rochford R, Chung J, Shapiro M, Purcell R, Chisari FV. Viral clearance without destruction of infected cells during acute HBV infection. Science. 1999;284:825–829. doi: 10.1126/science.284.5415.825. [DOI] [PubMed] [Google Scholar]
- 8.Ganem D, Prince AM. Hepatitis B virus infection--natural history and clinical consequences. N Engl J Med. 2004;350:1118–1129. doi: 10.1056/NEJMra031087. [DOI] [PubMed] [Google Scholar]
- 9.Kenny-Walsh E. Clinical outcomes after hepatitis C infection from contaminated anti-D immune globulin. Irish Hepatology Research Group. N Engl J Med. 1999;340:1228–1233. doi: 10.1056/NEJM199904223401602. [DOI] [PubMed] [Google Scholar]
- 10.Thomas DL, Astemborski J, Rai RM, Anania FA, Schaeffer M, Galai N, Nolt K, Nelson KE, Strathdee SA, Johnson L, et al. The natural history of hepatitis C virus infection: Host, viral, and environmental factors. JAMA. 2000;284:450–456. doi: 10.1001/jama.284.4.450. [DOI] [PubMed] [Google Scholar]
- 11.Seeff LB, Hollinger FB, Alter HJ, Wright EC, Cain CM, Buskell ZJ, Ishak KG, Iber FL, Toro D, Samanta A, et al. Long-term mortality and morbidity of transfusion-associated non-A, non-B, and type C hepatitis: A National Heart, Lung, and Blood Institute collaborative study. Hepatology. 2001;33:455–463. doi: 10.1053/jhep.2001.21905. [DOI] [PubMed] [Google Scholar]
- 12.Thompson AJ, Muir AJ, Sulkowski MS, Ge D, Fellay J, Shianna KV, Urban T, Afdhal NH, Jacobson IM, Esteban R, et al. Interleukin-28B polymorphism improves viral kinetics and is the strongest pretreatment predictor of sustained virologic response in genotype 1 hepatitis C virus. Gastroenterology. 2010;139:120–129. e118. doi: 10.1053/j.gastro.2010.04.013. [DOI] [PubMed] [Google Scholar]
- 13.Keele BF, Jones JH, Terio KA, Estes JD, Rudicell RS, Wilson ML, Li Y, Learn GH, Beasley TM, Schumacher-Stankey J, et al. Increased mortality and AIDS-like immunopathology in wild chimpanzees infected with SIVcpz. Nature. 2009;460:515–519. doi: 10.1038/nature08200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bar KJ, Li H, Chamberland A, Tremblay C, Routy JP, Grayson T, Sun C, Wang S, Learn GH, Morgan CJ, et al. Wide variation in the multiplicity of HIV-1 infection among injection drug users. J Virol. 2010;84:6241–6247. doi: 10.1128/JVI.00077-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Watts JM, Dang KK, Gorelick RJ, Leonard CW, Bess JW, Jr, Swanstrom R, Burch CL, Weeks KM. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature. 2009;460:711–716. doi: 10.1038/nature08237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Davis M, Sagan SM, Pezacki JP, Evans DJ, Simmonds P. Bioinformatic and physical characterizations of genome-scale ordered RNA structure in mammalian RNA viruses. J Virol. 2008;82:11824–11836. doi: 10.1128/JVI.01078-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jenkins GM, Worobey M, Woelk CH, Holmes EC. Evidence for the non-quasispecies evolution of RNA viruses [corrected] Mol Biol Evol. 2001;18:987–994. doi: 10.1093/oxfordjournals.molbev.a003900. [DOI] [PubMed] [Google Scholar]
- 18.Wilke CO. Quasispecies theory in the context of population genetics. BMC Evol Biol. 2005;5:44. doi: 10.1186/1471-2148-5-44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Perelson AS, Neumann AU, Markowitz M, Leonard JM, Ho DD. HIV-1 dynamics in vivo: Virion clearance rate, infected cell life-span, and viral generation time. Science. 1996;271:1582–1586. doi: 10.1126/science.271.5255.1582. [DOI] [PubMed] [Google Scholar]
- 20.Svarovskaia ES, Cheslock SR, Zhang WH, Hu WS, Pathak VK. Retroviral mutation rates and reverse transcriptase fidelity. Front Biosci. 2003;8:d117–d134. doi: 10.2741/957. [DOI] [PubMed] [Google Scholar]
- 21.Abram ME, Ferris AL, Shao W, Alvord WG, Hughes SH. Nature, position, and frequency of mutations made in a single cycle of HIV-1 replication. J Virol. 2010;84:9864–9878. doi: 10.1128/JVI.00915-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mansky LM. Forward mutation rate of human immunodeficiency virus type 1 in a T lymphoid cell line. AIDS Res Hum Retrovirus. 1996;12:307–314. doi: 10.1089/aid.1996.12.307. [DOI] [PubMed] [Google Scholar]
- 23.Coutsinos D, Invernizzi CF, Xu H, Moisi D, Oliveira M, Brenner BG, Wainberg MA. Template usage is responsible for the preferential acquisition of the K65R reverse transcriptase mutation in subtype C variants of human immunodeficiency virus type 1. J Virol. 2009;83:2029–2033. doi: 10.1128/JVI.01349-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hu WS, Temin HM. Genetic consequences of packaging two RNA genomes in one retroviral particle: Pseudodiploidy and high rate of genetic recombination. Proc Natl Acad Sci U S A. 1990;87:1556–1560. doi: 10.1073/pnas.87.4.1556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Levy DN, Aldrovandi GM, Kutsch O, Shaw GM. Dynamics of HIV-1 recombination in its natural target cells. Proc Natl Acad Sci U S A. 2004;101:4204–4209. doi: 10.1073/pnas.0306764101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ramirez BC, Simon-Loriere E, Galetto R, Negroni M. Implications of recombination for HIV diversity. Virus Res. 2008;134:64–73. doi: 10.1016/j.virusres.2008.01.007. [DOI] [PubMed] [Google Scholar]
- 27.Neher RA, Leitner T. Recombination rate and selection strength in HIV intra-patient evolution. PLoS Comput Biol. 2010;6:e1000660. doi: 10.1371/journal.pcbi.1000660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Galli A, Kearney M, Nikolaitchik OA, Yu S, Chin MP, Maldarelli F, Coffin JM, Pathak VK, Hu WS. Patterns of Human Immunodeficiency Virus type 1 recombination ex vivo provide evidence for coadaptation of distant sites, resulting in purifying selection for intersubtype recombinants during replication. J Virol. 2010;84:7651–7661. doi: 10.1128/JVI.00276-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shankarappa R, Margolick JB, Gange SJ, Rodrigo AG, Upchurch D, Farzadegan H, Gupta P, Rinaldo CR, Learn GH, He X, et al. Consistent viral evolutionary changes associated with the progression of human immunodeficiency virus type 1 infection. J Virol. 1999;73:10489–10502. doi: 10.1128/jvi.73.12.10489-10502.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Piantadosi A, Chohan B, Panteleeff D, Baeten JM, Mandaliya K, Ndinya-Achola JO, Overbaugh J. HIV-1 evolution in gag and env is highly correlated but exhibits different relationships with viral load and the immune response. AIDS. 2009;23:579–587. doi: 10.1097/QAD.0b013e328328f76e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kearney M, Maldarelli F, Shao W, Margolick JB, Daar ES, Mellors JW, Rao V, Coffin JM, Palmer S. Human immunodeficiency virus type 1 population genetics and adaptation in newly infected individuals. J Virol. 2009;83:2715–2727. doi: 10.1128/JVI.01960-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cullen BR. Role and mechanism of action of the APOBEC3 family of antiretroviral resistance factors. J Virol. 2006;80:1067–1076. doi: 10.1128/JVI.80.3.1067-1076.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kieffer TL, Kwon P, Nettles RE, Han Y, Ray SC, Siliciano RF. G→A hypermutation in protease and reverse transcriptase regions of human immunodeficiency virus type 1 residing in resting CD4+ T cells in vivo. J Virol. 2005;79:1975–1980. doi: 10.1128/JVI.79.3.1975-1980.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gifford RJ, Rhee SY, Eriksson N, Liu TF, Kiuchi M, Das AK, Shafer RW. Sequence editing by Apolipoprotein B RNA-editing catalytic component [corrected] and epidemiological surveillance of transmitted HIV-1 drug resistance. AIDS. 2008;22:717–725. doi: 10.1097/QAD.0b013e3282f5e07a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jern P, Russell RA, Pathak VK, Coffin JM. Likely role of APOBEC3G-mediated G-to-A mutations in HIV-1 evolution and drug resistance. PLoS Pathog. 2009;5:e1000367. doi: 10.1371/journal.ppat.1000367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nowak MA, Bonhoeffer S, Hill AM, Boehme R, Thomas HC, McDade H. Viral dynamics in hepatitis B virus infection. Proc Natl Acad Sci U S A. 1996;93:4398–4402. doi: 10.1073/pnas.93.9.4398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Whalley SA, Murray JM, Brown D, Webster GJ, Emery VC, Dusheiko GM, Perelson AS. Kinetics of acute hepatitis B virus infection in humans. J Exp Med. 2001;193:847–854. doi: 10.1084/jem.193.7.847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ho SK, Yam WC, Leung ET, Wong LP, Leung JK, Lai KN, Chan TM. Rapid quantification of hepatitis B virus DNA by real-time PCR using fluorescent hybridization probes. J Med Microbiol. 2003;52:397–402. doi: 10.1099/jmm.0.05071-0. [DOI] [PubMed] [Google Scholar]
- 39.Murray JM, Purcell RH, Wieland SF. The half-life of hepatitis B virions. Hepatology. 2006;44:1117–1121. doi: 10.1002/hep.21364. [DOI] [PubMed] [Google Scholar]
- 40.Dandri M, Murray JM, Lutgehetmann M, Volz T, Lohse AW, Petersen J. Virion half-life in chronic hepatitis B infection is strongly correlated with levels of viremia. Hepatology. 2008;48:1079–1086. doi: 10.1002/hep.22469. [DOI] [PubMed] [Google Scholar]
- 41.Herz AV, Bonhoeffer S, Anderson RM, May RM, Nowak MA. Viral dynamics in vivo: Limitations on estimates of intracellular delay and virus decay. Proc Natl Acad Sci U S A. 1996;93:7247–7251. doi: 10.1073/pnas.93.14.7247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hannoun C, Horal P, Lindh M. Long-term mutation rates in the hepatitis B virus genome. J Gen Virol. 2000;81:75–83. doi: 10.1099/0022-1317-81-1-75. [DOI] [PubMed] [Google Scholar]
- 43.Fares MA, Holmes EC. A revised evolutionary history of hepatitis B virus (HBV) J Mol Evol. 2002;54:807–814. doi: 10.1007/s00239-001-0084-z. [DOI] [PubMed] [Google Scholar]
- 44.Osiowy C, Giles E, Tanaka Y, Mizokami M, Minuk GY. Molecular evolution of hepatitis B virus over 25 years. J Virol. 2006;80:10307–10314. doi: 10.1128/JVI.00996-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Drake JW, Charlesworth B, Charlesworth D, Crow JF. Rates of spontaneous mutation. Genetics. 1998;148:1667–1686. doi: 10.1093/genetics/148.4.1667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Khudyakov Y. Coevolution and HBV drug resistance. Antivir Ther. 2010;15:505–515. doi: 10.3851/IMP1515. [DOI] [PubMed] [Google Scholar]
- 47.Simmonds P, Midgley S. Recombination in the genesis and evolution of hepatitis B virus genotypes. J Virol. 2005;79:15467–15476. doi: 10.1128/JVI.79.24.15467-15476.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gunther S. Genetic variation in HBV infection: Genotypes and mutants. J Clin Virol. 2006;36:S3–S11. doi: 10.1016/s1386-6532(06)80002-8. [DOI] [PubMed] [Google Scholar]
- 49.Guidotti LG, Chisari FV. Immunobiology and pathogenesis of viral hepatitis. Annu Rev Pathol. 2006;1:23–61. doi: 10.1146/annurev.pathol.1.110304.100230. [DOI] [PubMed] [Google Scholar]
- 50.Mizokami M, Orito E, Ohba K, Ikeo K, Lau JY, Gojobori T. Constrained evolution with respect to gene overlap of hepatitis B virus. J Mol Evol. 1997;44:S83–S90. doi: 10.1007/pl00000061. [DOI] [PubMed] [Google Scholar]
- 51.Zaaijer HL, van Hemert FJ, Koppelman MH, Lukashov VV. Independent evolution of overlapping polymerase and surface protein genes of hepatitis B virus. J Gen Virol. 2007;88:2137–2143. doi: 10.1099/vir.0.82906-0. [DOI] [PubMed] [Google Scholar]
- 52.Soriano V, Perelson AS, Zoulim F. Why are there different dynamics in the selection of drug resistance in HIV and hepatitis B and C viruses. J Antimicrob Chemother. 2008;62:1–4. doi: 10.1093/jac/dkn175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Lim SG, Cheng Y, Guindon S, Seet BL, Lee LY, Hu P, Wasser S, Peter FJ, Tan T, Goode M, et al. Viral quasi-species evolution during hepatitis Be antigen seroconversion. Gastroenterology. 2007;133:951–958. doi: 10.1053/j.gastro.2007.06.011. [DOI] [PubMed] [Google Scholar]
- 54.Chen L, Zhang Q, Yu DM, Wan MB, Zhang XX. Early changes of hepatitis B virus quasispecies during lamivudine treatment and the correlation with antiviral efficacy. J Hepatol. 2009;50:895–905. doi: 10.1016/j.jhep.2008.12.018. [DOI] [PubMed] [Google Scholar]
- 55.Margeridon-Thermet S, Shulman NS, Ahmed A, Shahriar R, Liu T, Wang C, Holmes SP, Babrzadeh F, Gharizadeh B, Hanczaruk B, et al. Ultra-deep pyrosequencing of hepatitis B virus quasispecies from nucleoside and nucleotide reverse-transcriptase inhibitor (NRTI)-treated patients and NRTI-naive patients. J Infect Dis. 2009;199:1275–1285. doi: 10.1086/597808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Pallier C, Rodriguez C, Brillet R, Nordmann P, Hezode C, Pawlotsky JM. Complex dynamics of hepatitis B Virus Res.istance to adefovir. Hepatology. 2009;49:50–59. doi: 10.1002/hep.22634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Pallier C, Castera L, Soulier A, Hezode C, Nordmann P, Dhumeaux D, Pawlotsky JM. Dynamics of hepatitis B Virus Res.istance to lamivudine. J Virol. 2006;80:643–653. doi: 10.1128/JVI.80.2.643-653.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Suspene R, Guetard D, Henry M, Sommer P, Wain-Hobson S, Vartanian JP. Extensive editing of both hepatitis B virus DNA strands by APOBEC3 cytidine deaminases in vitro and in vivo. Proc Natl Acad Sci U S A. 2005;102:8321–8326. doi: 10.1073/pnas.0408223102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Noguchi C, Ishino H, Tsuge M, Fujimoto Y, Imamura M, Takahashi S, Chayama K. G to A hypermutation of hepatitis B virus. Hepatology. 2005;41:626–633. doi: 10.1002/hep.20580. [DOI] [PubMed] [Google Scholar]
- 60.Reuman EC, Margeridon-Thermet S, Caudill HB, Liu T, Borroto-Esoda K, Svarovskaia ES, Holmes S, Shafer RW. A classification model for G-to-A hypermutation in hepatitis B virus ultra-deep pyrosequencing reads. Bioinformatics. 2010;26:2929–2932. doi: 10.1093/bioinformatics/btq570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Pawlotsky JM. Use and interpretation of virological tests for hepatitis C. Hepatology. 2002;36:S65–S73. doi: 10.1053/jhep.2002.36815. [DOI] [PubMed] [Google Scholar]
- 62.Neumann AU, Lam NP, Dahari H, Gretch DR, Wiley TE, Layden TJ, Perelson AS. Hepatitis C viral dynamics in vivo and the antiviral efficacy of interferon-alpha therapy. Science. 1998;282:103–107. doi: 10.1126/science.282.5386.103. [DOI] [PubMed] [Google Scholar]
- 63.Ramratnam B, Bonhoeffer S, Binley J, Hurley A, Zhang L, Mittler JE, Markowitz M, Moore JP, Perelson AS, Ho DD. Rapid production and clearance of HIV-1 and hepatitis C virus assessed by large volume plasma apheresis. Lancet. 1999;354:1782–1785. doi: 10.1016/S0140-6736(99)02035-8. [DOI] [PubMed] [Google Scholar]
- 64.Herrmann E, Neumann AU, Schmidt JM, Zeuzem S. Hepatitis C virus kinetics. Antivir Ther. 2000;5:85–90. [PubMed] [Google Scholar]
- 65.Guo JT, Bichko VV, Seeger C. Effect of alpha interferon on the hepatitis C virus replicon. J Virol. 2001;75:8516–8523. doi: 10.1128/JVI.75.18.8516-8523.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ogata N, Alter HJ, Miller RH, Purcell RH. Nucleotide sequence and mutation rate of the H strain of hepatitis C virus. Proc Natl Acad Sci U S A. 1991;88:3392–3396. doi: 10.1073/pnas.88.8.3392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Smith DB, Pathirana S, Davidson F, Lawlor E, Power J, Yap PL, Simmonds P. The origin of hepatitis C virus genotypes. J Gen Virol. 1997;78(Pt 2):321–328. doi: 10.1099/0022-1317-78-2-321. [DOI] [PubMed] [Google Scholar]
- 68.Powdrill M, Bernatchez J, Gotte M. Inhibitors of hepatitis C virus RNA-dependent RNA polymerase NS5B. Viruses. 2010;2:2169–2195. doi: 10.3390/v2102169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Simmonds P. Genetic diversity and evolution of hepatitis C virus--15 years on. J Gen Virol. 2004;85:3173–3188. doi: 10.1099/vir.0.80401-0. [DOI] [PubMed] [Google Scholar]
- 70.Kalinina O, Norder H, Mukomolov S, Magnius LO. A natural intergenotypic recombinant of hepatitis C virus identified in St. Petersburg. J Virol. 2002;76:4034–4043. doi: 10.1128/JVI.76.8.4034-4043.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Colina R, Casane D, Vasquez S, Garcia-Aguirre L, Chunga A, Romero H, Khan B, Cristina J. Evidence of intratypic recombination in natural populations of hepatitis C virus. J Gen Virol. 2004;85:31–37. doi: 10.1099/vir.0.19472-0. [DOI] [PubMed] [Google Scholar]
- 72.Moreau I, Hegarty S, Levis J, Sheehy P, Crosbie O, Kenny-Walsh E, Fanning LJ. Serendipitous identification of natural intergenotypic recombinants of hepatitis C in Ireland. Virol J. 2006;3:95. doi: 10.1186/1743-422X-3-95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Mondelli MU, Cerino A, Meola A, Nicosia A. Variability or conservation of hepatitis C virus hypervariable region 1? Implications for immune responses. J Biosci. 2003;28:305–310. doi: 10.1007/BF02970149. [DOI] [PubMed] [Google Scholar]
- 74.Sheridan I, Pybus OG, Holmes EC, Klenerman P. High-resolution phylogenetic analysis of hepatitis C virus adaptation and its relationship to disease progression. J Virol. 2004;78:3447–3454. doi: 10.1128/JVI.78.7.3447-3454.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Erickson AL, Kimura Y, Igarashi S, Eichelberger J, Houghton M, Sidney J, McKinney D, Sette A, Hughes AL, Walker CM. The outcome of hepatitis C virus infection is predicted by escape mutations in epitopes targeted by cytotoxic T lymphocytes. Immunity. 2001;15:883–895. doi: 10.1016/s1074-7613(01)00245-x. [DOI] [PubMed] [Google Scholar]
- 76.Soderholm J, Ahlen G, Kaul A, Frelin L, Alheim M, Barnfield C, Liljestrom P, Weiland O, Milich DR, Bartenschlager R, et al. Relation between viral fitness and immune escape within the hepatitis C virus protease. Gut. 2006;55:266–274. doi: 10.1136/gut.2005.072231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Gale M, Jr, Foy EM. Evasion of intracellular host defence by hepatitis C virus. Nature. 2005;436:939–945. doi: 10.1038/nature04078. [DOI] [PubMed] [Google Scholar]
- 78.Martell M, Esteban JI, Quer J, Genesca J, Weiner A, Esteban R, Guardia J, Gomez J. Hepatitis C virus (HCV) circulates as a population of different but closely related genomes: Quasispecies nature of HCV genome distribution. J Virol. 1992;66:3225–3229. doi: 10.1128/jvi.66.5.3225-3229.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Manzin A, Solforosi L, Petrelli E, Macarri G, Tosone G, Piazza M, Clementi M. Evolution of hypervariable region 1 of hepatitis C virus in primary infection. J Virol. 1998;72:6271–6276. doi: 10.1128/jvi.72.7.6271-6276.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Cristina J, del Pilar Moreno M, Moratorio G. Hepatitis C virus genetic variability in patients undergoing antiviral therapy. Virus Res. 2007;127:185–194. doi: 10.1016/j.virusres.2007.02.023. [DOI] [PubMed] [Google Scholar]
- 81.Fan X, Mao Q, Zhou D, Lu Y, Xing J, Xu Y, Ray SC, Di Bisceglie AM. High diversity of hepatitis C viral quasispecies is associated with early virological response in patients undergoing antiviral therapy. Hepatology. 2009;50:1765–1772. doi: 10.1002/hep.23290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Hemelaar J, Gouws E, Ghys PD, Osmanov S. Global and regional distribution of HIV-1 genetic subtypes and recombinants in 2004. AIDS. 2006;20:W13–W23. doi: 10.1097/01.aids.0000247564.73009.bc. [DOI] [PubMed] [Google Scholar]
- 83.McCutchan FE. Global epidemiology of HIV. J Med Virol. 2006;78:S7–S12. doi: 10.1002/jmv.20599. [DOI] [PubMed] [Google Scholar]
- 84.Parkin NT, Schapiro JM. Antiretroviral drug resistance in non-subtype B HIV-1, HIV-2 and SIV. Antivir Ther. 2004;9:3–12. [PubMed] [Google Scholar]
- 85.Kantor R, Katzenstein DA, Efron B, Carvalho AP, Wynhoven B, Cane P, Clarke J, Sirivichayakul S, Soares MA, Snoeck J, et al. Impact of HIV-1 subtype and antiretroviral therapy on protease and reverse transcriptase genotype: Results of a global collaboration. PLoS Med. 2005;2:e112. doi: 10.1371/journal.pmed.0020112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Cane PA, de Ruiter A, Rice P, Wiselka M, Fox R, Pillay D. Resistance-associated mutations in the human immunodeficiency virus type 1 subtype c protease gene from treated and untreated patients in the United Kingdom. J Clin Microbiol. 2001;39:2652–2654. doi: 10.1128/JCM.39.7.2652-2654.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Sugiura W, Matsuda Z, Yokomaku Y, Hertogs K, Larder B, Oishi T, Okano A, Shiino T, Tatsumi M, Matsuda M, et al. Interference between D30N and L90M in selection and development of protease inhibitor-resistant human immunodeficiency virus type 1. Antimicrob Agents Chemother. 2002;46:708–715. doi: 10.1128/AAC.46.3.708-715.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Brenner B, Turner D, Oliveira M, Moisi D, Detorio M, Carobene M, Marlink RG, Schapiro J, Roger M, Wainberg MA. A V106M mutation in HIV-1 clade C viruses exposed to efavirenz confers cross-resistance to non-nucleoside reverse transcriptase inhibitors. AIDS. 2003;17:F1–F5. doi: 10.1097/00002030-200301030-00001. [DOI] [PubMed] [Google Scholar]
- 89.Grossman Z, Istomin V, Averbuch D, Lorber M, Risenberg K, Levi I, Chowers M, Burke M, Bar Yaacov N, Schapiro JM. Genetic variation at NNRTI resistance-associated positions in patients infected with HIV-1 subtype C. AIDS. 2004;18:909–915. doi: 10.1097/00002030-200404090-00008. [DOI] [PubMed] [Google Scholar]
- 90.Grossman Z, Paxinos EE, Averbuch D, Maayan S, Parkin NT, Engelhard D, Lorber M, Istomin V, Shaked Y, Mendelson E, et al. Mutation D30N is not preferentially selected by human immunodeficiency virus type 1 subtype C in the development of resistance to nelfinavir. Antimicrob Agents Chemother. 2004;48:2159–2165. doi: 10.1128/AAC.48.6.2159-2165.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Camacho R, Godinho A, Gomes P, Abecasis A, Vandamme A-M, Palma C, Carvalho AP, Cabanas J, Goncalves J. Different substitutions under drug pressure at protease codon 82 in HIV-1 subtype G compared to subtype B infected individuals including a novel I82M resistance mutations [abstract 138] Antivir Ther. 2005;10:S151. [Google Scholar]
- 92.Martinez-Cajas JL, Pai NP, Klein MB, Wainberg MA. Differences in resistance mutations among HIV-1 non-subtype B infections: A systematic review of evidence 1996–2008. J Int AIDS Soc. 2009;12:11. doi: 10.1186/1758-2652-12-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Doualla-Bell F, Avalos A, Brenner B, Gaolathe T, Mine M, Gaseitsiwe S, Oliveira M, Moisi D, Ndwapi N, Moffat H, et al. High prevalence of the K65R mutation in human immunodeficiency virus type 1 subtype C isolates from infected patients in Botswana treated with didanosine-based regimens. Antimicrob Agents Chemother. 2006;50:4182–4185. doi: 10.1128/AAC.00714-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Hosseinipour MC, van Oosterhout JJ, Weigel R, Phiri S, Kamwendo D, Parkin N, Fiscus SA, Nelson JA, Eron JJ, Kumwenda J. The public health approach to identify antiretroviral therapy failure: High-level nucleoside reverse transcriptase inhibitor resistance among Malawians failing first-line antiretroviral therapy. AIDS. 2009;23:1127–1134. doi: 10.1097/QAD.0b013e32832ac34e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Invernizzi CF, Coutsinos D, Oliveira M, Moisi D, Brenner BG, Wainberg MA. Signature nucleotide polymorphisms at positions 64 and 65 in reverse transcriptase favor the selection of the K65R resistance mutation in HIV-1 subtype C. J Infect Dis. 2009;200:1202–1206. doi: 10.1086/605894. [DOI] [PubMed] [Google Scholar]
- 96.Stuyver L, De Gendt S, Van Geyt C, Zoulim F, Fried M, Schinazi RF, Rossau R. A new genotype of hepatitis B virus: Complete genome and phylogenetic relatedness. J Gen Virol. 2000;81:67–74. doi: 10.1099/0022-1317-81-1-67. [DOI] [PubMed] [Google Scholar]
- 97.Fung SK, Lok AS. Hepatitis B virus genotypes: Do they play a role in the outcome of HBV infection. Hepatology. 2004;40:790–792. doi: 10.1002/hep.1840400407. [DOI] [PubMed] [Google Scholar]
- 98.Liu CJ, Kao JH. Genetic variability of hepatitis B virus and response to antiviral therapy. Antivir Ther. 2008;13:613–624. [PubMed] [Google Scholar]
- 99.Moucari R, Martinot-Peignoux M, Mackiewicz V, Boyer N, Ripault MP, Castelnau C, Leclere L, Dauvergne A, Valla D, Vidaud M, et al. Influence of genotype on hepatitis B surface antigen kinetics in hepatitis B e antigen-negative patients treated with pegylated interferon-alpha2a. Antivir Ther. 2009;14:1183–1188. doi: 10.3851/IMP1458. [DOI] [PubMed] [Google Scholar]
- 100.McMahon BJ. The influence of hepatitis B virus genotype and subgenotype on the natural history of chronic hepatitis B. Hepatol Int. 2009;3:334–342. doi: 10.1007/s12072-008-9112-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Liaw Y, Brunetto M, Hadziyannis S. The natural history of chronic HBV infection and geographical differences. Antivir Ther. 2010;15:25–33. doi: 10.3851/IMP1621. [DOI] [PubMed] [Google Scholar]
- 102.Zein NN. Clinical significance of hepatitis C virus genotypes. Clin Microbiol Rev. 2000;13:223–235. doi: 10.1128/cmr.13.2.223-235.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Simmonds P, Bukh J, Combet C, Deleage G, Enomoto N, Feinstone S, Halfon P, Inchauspe G, Kuiken C, Maertens G, et al. Consensus proposals for a unified system of nomenclature of hepatitis C virus genotypes. Hepatology. 2005;42:962–973. doi: 10.1002/hep.20819. [DOI] [PubMed] [Google Scholar]
- 104.Simmonds P. The origin and evolution of hepatitis viruses in humans. J Gen Virol. 2001;82:693–712. doi: 10.1099/0022-1317-82-4-693. [DOI] [PubMed] [Google Scholar]
- 105.Legrand-Abravanel F, Colson P, Leguillou-Guillemette H, Alric L, Ravaux I, Lunel-Fabiani F, Bouviers-Alias M, Trimoulet P, Chaix ML, Hezode C, et al. Influence of the HCV subtype on the virological response to pegylated interferon and ribavirin therapy. J Med Virol. 2009;81:2029–2035. doi: 10.1002/jmv.21583. [DOI] [PubMed] [Google Scholar]
- 106.Ghany MG, Strader DB, Thomas DL, Seeff LB. Diagnosis, management, and treatment of hepatitis C: An update. Hepatology. 2009;49:1335–1374. doi: 10.1002/hep.22759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Phillips AN, Miller V, Sabin C, Cozzi Lepri A, Klauke S, Bickel M, Doerr HW, Hill A, Staszewski S. Durability of HIV-1 viral suppression over 3.3 years with multi-drug antiretroviral therapy in previously drug-naive individuals. AIDS. 2001;15:2379–2384. doi: 10.1097/00002030-200112070-00005. [DOI] [PubMed] [Google Scholar]
- 108.Bennett D, McCormick L, Kline R, Wheeler W, Hemmen M, Smith A, Zaidi I, Dondero TUS. surveillance of HIV drug resistance at diagnosis using HIV diagnostic sera [Abstract 674]. Proceeding of the 12th Conference on Retroviruses and Opportunistic Infections; Boston, MA, USA. 22–25 February 2005. [Google Scholar]
- 109.Vercauteren J, Wensing AM, van de Vijver DA, Albert J, Balotta C, Hamouda O, Kucherer C, Struck D, Schmit JC, Asjo B, et al. Transmission of drug-resistant HIV-1 is stabilizing in Europe. J Infect Dis. 2009;200:1503–1508. doi: 10.1086/644505. [DOI] [PubMed] [Google Scholar]
- 110.Geretti AM. Epidemiology of antiretroviral drug resistance in drug-naive persons. Curr Opin Infect Dis. 2007;20:22–32. doi: 10.1097/QCO.0b013e328013caff. [DOI] [PubMed] [Google Scholar]
- 111.Chan P, Kantor R. Transmitted drug resistance in nonsubtype B HIV-1 infection. HIV Therapy. 2009;3:447–465. doi: 10.2217/hiv.09.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Meyer PR, Matsuura SE, Schinazi RF, So AG, Scott WA. Differential removal of thymidine nucleotide analogues from blocked DNA chains by human immunodeficiency virus reverse transcriptase in the presence of physiological concentrations of 2’-deoxynucleoside triphosphates. Antimicrob Agents Chemother. 2000;44:3465–3472. doi: 10.1128/aac.44.12.3465-3472.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Lennerstrand J, Stammers DK, Larder BA. Biochemical mechanism of human immunodeficiency virus type 1 reverse transcriptase resistance to stavudine. Antimicrob Agents Chemother. 2001;45:2144–2146. doi: 10.1128/AAC.45.7.2144-2146.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Whitcomb JM, Parkin NT, Chappey C, Hellmann NS, Petropoulos CJ. Broad nucleoside reverse-transcriptase inhibitor cross-resistance in human immunodeficiency virus type 1 clinical isolates. J Infect Dis. 2003;188:992–1000. doi: 10.1086/378281. [DOI] [PubMed] [Google Scholar]
- 115.Sarafianos SG, Das K, Hughes SH, Arnold E. Taking aim at a moving target: Designing drugs to inhibit drug-resistant HIV-1 reverse transcriptases. Curr Opin Struct Biol. 2004;14:716–730. doi: 10.1016/j.sbi.2004.10.013. [DOI] [PubMed] [Google Scholar]
- 116.Arion D, Sluis-Cremer N, Parniak MA. Mechanism by which phosphonoformic acid resistance mutations restore 3’- azido-3’-deoxythymidine (AZT) sensitivity to AZT-resistant HIV-1 reverse transcriptase. J Biol Chem. 2000;275:9251–9255. doi: 10.1074/jbc.275.13.9251. [DOI] [PubMed] [Google Scholar]
- 117.Lanier ER, Ait-Khaled M, Scott J, Stone C, Melby T, Sturge G, St Clair M, Steel H, Hetherington S, Pearce G, et al. Antiviral efficacy of abacavir in antiretroviral therapy-experienced adults harbouring HIV-1 with specific patterns of resistance to nucleoside reverse transcriptase inhibitors. Antivir Ther. 2004;9:37–45. doi: 10.1177/135965350400900102. [DOI] [PubMed] [Google Scholar]
- 118.Molina JM, Marcelin AG, Pavie J, Heripret L, De Boever CM, Troccaz M, Leleu G, Calvez V. Didanosine in HIV-1-infected patients experiencing failure of antiretroviral therapy: A randomized placebo-controlled trial. J Infect Dis. 2005;191:840–847. doi: 10.1086/428094. [DOI] [PubMed] [Google Scholar]
- 119.Miller MD, Margot N, Lu B, Zhong L, Chen SS, Cheng A, Wulfsohn M. Genotypic and phenotypic predictors of the magnitude of response to tenofovir disoproxil fumarate treatment in antiretroviral-experienced patients. J Infect Dis. 2004;189:837–846. doi: 10.1086/381784. [DOI] [PubMed] [Google Scholar]
- 120.Parikh UM, Koontz DL, Chu CK, Schinazi RF, Mellors JW. In vitro activity of structurally diverse nucleoside analogs against human immunodeficiency virus type 1 with the K65R mutation in reverse transcriptase. Antimicrob Agents Chemother. 2005;49:1139–1144. doi: 10.1128/AAC.49.3.1139-1144.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Gallant JE, DeJesus E, Arribas JR, Pozniak AL, Gazzard B, Campo RE, Lu B, McColl D, Chuck S, Enejosa J, et al. Tenofovir DF, emtricitabine, and efavirenz vs. zidovudine, lamivudine, and efavirenz for HIV. N Engl J Med. 2006;354:251–260. doi: 10.1056/NEJMoa051871. [DOI] [PubMed] [Google Scholar]
- 122.Winters MA, Coolley KL, Girard YA, Levee DJ, Hamdan H, Shafer RW, Katzenstein DA, Merigan TC. A 6-basepair insert in the reverse transcriptase gene of human immunodeficiency virus type 1 confers resistance to multiple nucleoside inhibitors. J Clin Invest. 1998;102:1769–1775. doi: 10.1172/JCI4948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Masquelier B, Race E, Tamalet C, Descamps D, Izopet J, Buffet-Janvresse C, Ruffault A, Mohammed AS, Cottalorda J, Schmuck A, et al. Genotypic and phenotypic resistance patterns of human immunodeficiency virus type 1 variants with insertions or deletions in the reverse transcriptase (RT): Multicenter study of patients treated with RT inhibitors. Antimicrob Agents Chemother. 2001;45:1836–1842. doi: 10.1128/AAC.45.6.1836-1842.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Shafer RW, Kozal MJ, Winters MA, Iversen AK, Katzenstein DA, Ragni MV, Meyer WA, Gupta P, Rasheed S, Coombs R, et al. Combination therapy with zidovudine and didanosine selects for drug- resistant human immunodeficiency virus type 1 strains with unique patterns of pol gene mutations. J Infect Dis. 1994;169:722–729. doi: 10.1093/infdis/169.4.722. [DOI] [PubMed] [Google Scholar]
- 125.Shirasaka T, Kavlick MF, Ueno T, Gao WY, Kojima E, Alcaide ML, Chokekijchai S, Roy BM, Arnold E, Yarchoan R, et al. Emergence of human immunodeficiency virus type 1 variants with resistance to multiple dideoxynucleosides in patients receiving therapy with dideoxynucleosides. Proc Natl Acad Sci U S A. 1995;92:2398–2402. doi: 10.1073/pnas.92.6.2398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Ehteshami M, Gotte M. Effects of mutations in the connection and RNase H domains of HIV-1 reverse transcriptase on drug susceptibility. AIDS Rev. 2008;10:224–235. [PubMed] [Google Scholar]
- 127.Shahriar R, Rhee SY, Liu TF, Fessel WJ, Scarsella A, Towner W, Holmes SP, Zolopa AR, Shafer RW. Nonpolymorphic human immunodeficiency virus type 1 protease and reverse transcriptase treatment-selected mutations. Antimicrob Agents Chemother. 2009;53:4869–4878. doi: 10.1128/AAC.00592-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Gupta S, Fransen S, Paxinos EE, Stawiski E, Huang W, Petropoulos CJ. Combinations of mutations in the connection domain of human immunodeficiency virus type 1 reverse transcriptase: Assessing the impact on nucleoside and nonnucleoside reverse transcriptase inhibitor resistance. Antimicrob Agents Chemother. 2010;54:1973–1980. doi: 10.1128/AAC.00870-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Acosta-Hoyos AJ, Scott WA. The Role of Nucleotide Excision by Reverse Transcriptase in HIV Drug Resistance. Viruses. 2010;2:372–394. doi: 10.3390/v2020372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Nikolenko GN, Delviks-Frankenberry KA, Palmer S, Maldarelli F, Fivash MJ, Jr, Coffin JM, Pathak VK. Mutations in the connection domain of HIV-1 reverse transcriptase increase 3’-azido-3’-deoxythymidine resistance. Proc Natl Acad Sci U S A. 2007;104:317–322. doi: 10.1073/pnas.0609642104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Delviks-Frankenberry KA, Nikolenko GN, Pathak VK. The “connection” between HIV drug resistance and RNase H. Viruses. 2010;2:1476–1503. doi: 10.3390/v2071476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Parkin NT, Hellmann NS, Whitcomb JM, Kiss L, Chappey C, Petropoulos CJ. Natural variation of drug susceptibility in wild-type HIV-1. Antimicrob Agents Chemother. 2004;48:437–443. doi: 10.1128/AAC.48.2.437-443.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Jackson JB, Becker-Pergola G, Guay LA, Musoke P, Mracna M, Fowler MG, Mofenson LM, Mirochnick M, Mmiro F, Eshleman SH. Identification of the K103N resistance mutation in Ugandan women receiving nevirapine to prevent HIV-1 vertical transmission. AIDS. 2000;14:F111–F115. doi: 10.1097/00002030-200007280-00001. [DOI] [PubMed] [Google Scholar]
- 134.Jourdain G, Ngo-Giang-Huong N, Le Coeur S, Bowonwatanuwong C, Kantipong P, Leechanachai P, Ariyadej S, Leenasirimakul P, Hammer S, Lallemant M. Intrapartum exposure to nevirapine and subsequent maternal responses to nevirapine-based antiretroviral therapy. N Engl J Med. 2004;351:229–240. doi: 10.1056/NEJMoa041305. [DOI] [PubMed] [Google Scholar]
- 135.Vingerhoets J, Azijn H, Fransen E, De Baere I, Smeulders L, Jochmans D, Andries K, Pauwels R, de Bethune MP. TMC125 displays a high genetic barrier to the development of resistance: Evidence from in vitro selection experiments. J Virol. 2005;79:12773–12782. doi: 10.1128/JVI.79.20.12773-12782.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Vingerhoets J, Tambuyzer L, Azijn H, Hoogstoel A, Nijs S, Peeters M, de Bethune MP, De Smedt G, Woodfall B, Picchio G. Resistance profile of etravirine: Combined analysis of baseline genotypic and phenotypic data from the randomized, controlled Phase III clinical studies. AIDS. 2010;24:503–514. doi: 10.1097/QAD.0b013e32833677ac. [DOI] [PubMed] [Google Scholar]
- 137.Das K, Clark AD, Jr, Lewi PJ, Heeres J, De Jonge MR, Koymans LM, Vinkers HM, Daeyaert F, Ludovici DW, Kukla MJ, et al. Roles of conformational and positional adaptability in structure-based design of TMC125-R165335 (etravirine) and related non-nucleoside reverse transcriptase inhibitors that are highly potent and effective against wild-type and drug-resistant HIV-1 variants. J Med Chem. 2004;47:2550–2560. doi: 10.1021/jm030558s. [DOI] [PubMed] [Google Scholar]
- 138.Larder BA. Interactions between drug resistance mutations in human immunodeficiency virus type 1 reverse transcriptase. J Gen Virol. 1994;75:951–957. doi: 10.1099/0022-1317-75-5-951. [DOI] [PubMed] [Google Scholar]
- 139.Shulman N, Zolopa AR, Passaro D, Shafer RW, Huang W, Katzenstein D, Israelski DM, Hellmann N, Petropoulos C, Whitcomb J. Phenotypic hypersusceptibility to non-nucleoside reverse transcriptase inhibitors in treatment-experienced HIV-infected patients: Impact on virological response to efavirenz-based therapy. AIDS. 2001;15:1125–1132. doi: 10.1097/00002030-200106150-00007. [DOI] [PubMed] [Google Scholar]
- 140.Whitcomb JM, Huang W, Limoli K, Paxinos E, Wrin T, Skowron G, Deeks SG, Bates M, Hellmann NS, Petropoulos CJ. Hypersusceptibility to non-nucleoside reverse transcriptase inhibitors in HIV-1: Clinical, phenotypic and genotypic correlates. AIDS. 2002;16:F41–F47. doi: 10.1097/00002030-200210180-00002. [DOI] [PubMed] [Google Scholar]
- 141.Vermeiren H, Van Craenenbroeck E, Alen P, Bacheler L, Picchio G, Lecocq P. Prediction of HIV-1 drug susceptibility phenotype from the viral genotype using linear regression modeling. J Virol Methods. 2007;145:47–55. doi: 10.1016/j.jviromet.2007.05.009. [DOI] [PubMed] [Google Scholar]
- 142.Rhee SY, Taylor J, Fessel WJ, Kaufman D, Towner W, Troia P, Ruane P, Hellinger J, Shirvani V, Zolopa A, et al. HIV-1 protease mutations and protease inhibitor cross-resistance. Antimicrob Agents Chemother. 2010;54:4253–4261. doi: 10.1128/AAC.00574-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Erickson JW, Gulnik SV, Markowitz M. Protease inhibitors: Resistance, cross-resistance, fitness and the choice of initial and salvage therapies. AIDS. 1999;13:S189–S204. [PubMed] [Google Scholar]
- 144.Foulkes-Murzycki JE, Scott WR, Schiffer CA. Hydrophobic sliding: A possible mechanism for drug resistance in human immunodeficiency virus type 1 protease. Structure. 2007;15:225–233. doi: 10.1016/j.str.2007.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Cote HC, Brumme ZL, Harrigan PR. Human immunodeficiency virus type 1 protease cleavage site mutations associated with protease inhibitor cross-resistance selected by indinavir, ritonavir, and/or saquinavir. J Virol. 2001;75:589–594. doi: 10.1128/JVI.75.2.589-594.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Maguire MF, Guinea R, Griffin P, Macmanus S, Elston RC, Wolfram J, Richards N, Hanlon MH, Porter DJ, Wrin T, et al. Changes in human immunodeficiency virus type 1 Gag at positions L449 and P453 are linked to I50V protease mutants in vivo and cause reduction of sensitivity to amprenavir and improved viral fitnessin vitro. J Virol. 2002;76:7398–7406. doi: 10.1128/JVI.76.15.7398-7406.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Kolli M, Lastere S, Schiffer CA. Co-evolution of nelfinavir-resistant HIV-1 protease and the p1-p6 substrate. Virology. 2006;347:405–409. doi: 10.1016/j.virol.2005.11.049. [DOI] [PubMed] [Google Scholar]
- 148.Nijhuis M, van Maarseveen NM, Lastere S, Schipper P, Coakley E, Glass B, Rovenska M, de Jong D, Chappey C, Goedegebuure IW, et al. A Novel Substrate-Based HIV-1 Protease Inhibitor Drug Resistance Mechanism. PLoS Med. 2007;4:e36. doi: 10.1371/journal.pmed.0040036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Kempf DJ, Isaacson JD, King MS, Brun SC, Xu Y, Real K, Bernstein BM, Japour AJ, Sun E, Rode RA. Identification of genotypic changes in human immunodeficiency virus protease that correlate with reduced susceptibility to the protease inhibitor lopinavir among viral isolates from protease inhibitor-experienced patients. J Virol. 2001;75:7462–7469. doi: 10.1128/JVI.75.16.7462-7469.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.De Meyer S, Azijn H, Surleraux D, Jochmans D, Tahri A, Pauwels R, Wigerinck P, de Bethune MP. TMC114, a novel human immunodeficiency virus type 1 protease inhibitor active against protease inhibitor-resistant viruses, including a broad range of clinical isolates. Antimicrob Agents Chemother. 2005;49:2314–2321. doi: 10.1128/AAC.49.6.2314-2321.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Bierman WF, van Agtmael MA, Nijhuis M, Danner SA, Boucher CA. HIV monotherapy with ritonavir-boosted protease inhibitors: A systematic review. AIDS. 2009;23:279–291. doi: 10.1097/QAD.0b013e32831c54e5. [DOI] [PubMed] [Google Scholar]
- 152.Arribas JR, Horban A, Gerstoft J, Fatkenheuer G, Nelson M, Clumeck N, Pulido F, Hill A, van Delft Y, Stark T, et al. The MONET trial: Darunavir/ritonavir with or without nucleoside analogues, for patients with HIV RNA below 50 copies/mL. AIDS. 2010;24:223–230. doi: 10.1097/QAD.0b013e3283348944. [DOI] [PubMed] [Google Scholar]
- 153.Katlama C, Valantin MA, Algarte-Genin M, Duvivier C, Lambert-Niclot S, Girard PM, Molina JM, Hoen B, Pakianather S, Peytavin G, et al. Efficacy of darunavir/ritonavir maintenance monotherapy in patients with HIV-1 viral suppression: A randomized open-label, noninferiority trial, MONOI-ANRS 136. AIDS. 2010;24:2365–2374. doi: 10.1097/QAD.0b013e32833dec20. [DOI] [PubMed] [Google Scholar]
- 154.Hazuda DJ, Felock P, Witmer M, Wolfe A, Stillmock K, Grobler JA, Espeseth A, Gabryelski L, Schleif W, Blau C, et al. Inhibitors of strand transfer that prevent integration and inhibit HIV- 1 replication in cells. Science. 2000;287:646–650. doi: 10.1126/science.287.5453.646. [DOI] [PubMed] [Google Scholar]
- 155.Johnson AA, Santos W, Pais GC, Marchand C, Amin R, Burke TR, Jr, Verdine G, Pommier Y. Integration requires a specific interaction of the donor DNA terminal 5’-cytosine with glutamine 148 of the HIV-1 integrase flexible loop. J Biol Chem. 2006;281:461–467. doi: 10.1074/jbc.M511348200. [DOI] [PubMed] [Google Scholar]
- 156.McColl DJ, Chen X. Strand transfer inhibitors of HIV-1 integrase: Bringing IN a new era of antiretroviral therapy. Antivir Res. 2010;85:101–118. doi: 10.1016/j.antiviral.2009.11.004. [DOI] [PubMed] [Google Scholar]
- 157.Mouscadet JF, Delelis O, Marcelin AG, Tchertanov L. Resistance to HIV-1 integrase inhibitors: A structural perspective. Drug Resist Updat. 2010;13:139–150. doi: 10.1016/j.drup.2010.05.001. [DOI] [PubMed] [Google Scholar]
- 158.McColl D, Fransen S, Gupta S, Parking N, Margot N, Chuck S, Cheng A, Miller M. Resistance and cross-resistance to first generation integrase inhibitors: Insights from a phase II study of elvitegravir (GS-9137) [abstract 9] Antivir Ther. 2007;12:S11. [Google Scholar]
- 159.Shimura K, Kodama E, Sakagami Y, Matsuzaki Y, Watanabe W, Yamataka K, Watanabe Y, Ohata Y, Doi S, Sato M, et al. Broad antiretroviral activity and resistance profile of the novel human immunodeficiency virus integrase inhibitor elvitegravir (JTK-303/GS-9137) J Virol. 2008;82:764–774. doi: 10.1128/JVI.01534-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Kobayashi M, Nakahara K, Seki T, Miki S, Kawauchi S, Suyama A, Wakasa-Morimoto C, Kodama M, Endoh T, Oosugi E, et al. Selection of diverse and clinically relevant integrase inhibitor-resistant human immunodeficiency virus type 1 mutants. Antivir Res. 2008;80:213–222. doi: 10.1016/j.antiviral.2008.06.012. [DOI] [PubMed] [Google Scholar]
- 161.Goethals O, Clayton R, Van Ginderen M, Vereycken I, Wagemans E, Geluykens P, Dockx K, Strijbos R, Smits V, Vos A, et al. Resistance mutations in human immunodeficiency virus type 1 integrase selected with elvitegravir confer reduced susceptibility to a wide range of integrase inhibitors. J Virol. 2008;82:10366–10374. doi: 10.1128/JVI.00470-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Goethals O, Vos A, Van Ginderen M, Geluykens P, Smits V, Schols D, Hertogs K, Clayton R. Primary mutations selected in vitro with raltegravir confer large fold changes in susceptibility to first-generation integrase inhibitors, but minor fold changes to inhibitors with second-generation resistance profiles. Virology. 2010;402:338–346. doi: 10.1016/j.virol.2010.03.034. [DOI] [PubMed] [Google Scholar]
- 163.Fransen S, Gupta S, Danovich R, Hazuda D, Miller M, Witmer M, Petropoulos CJ, Huang W. Loss of raltegravir susceptibility by human immunodeficiency virus type 1 is conferred via multiple nonoverlapping genetic pathways. J Virol. 2009;83:11440–11446. doi: 10.1128/JVI.01168-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Delelis O, Malet I, Na L, Tchertanov L, Calvez V, Marcelin AG, Subra F, Deprez E, Mouscadet JF. The G140S mutation in HIV integrases from raltegravir-resistant patients rescues catalytic defect due to the resistance Q148H mutation. Nucl Acids Res. 2009;37:1193–1201. doi: 10.1093/nar/gkn1050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Reuman EC, Bachmann MH, Varghese V, Fessel WJ, Shafer RW. Panel of prototypical raltegravir-resistant infectious molecular clones in a novel integrase-deleted cloning vector. Antimicrob Agents Chemother. 2010;54:934–936. doi: 10.1128/AAC.01345-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Metifiot M, Maddali K, Naumova A, Zhang X, Marchand C, Pommier Y. Biochemical and pharmacological analyses of HIV-1 integrase flexible loop mutants resistant to raltegravir. Biochemistry. 2010;49:3715–3722. doi: 10.1021/bi100130f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Kilby JM, Eron JJ. Novel therapies based on mechanisms of HIV-1 cell entry. N Engl J Med. 2003;348:2228–2238. doi: 10.1056/NEJMra022812. [DOI] [PubMed] [Google Scholar]
- 168.Sista PR, Melby T, Davison D, Jin L, Mosier S, Mink M, Nelson EL, DeMasi R, Cammack N, Salgo MP, et al. Characterization of determinants of genotypic and phenotypic resistance to enfuvirtide in baseline and on-treatment HIV-1 isolates. AIDS. 2004;18:1787–1794. doi: 10.1097/00002030-200409030-00007. [DOI] [PubMed] [Google Scholar]
- 169.Mink M, Mosier SM, Janumpalli S, Davison D, Jin L, Melby T, Sista P, Erickson J, Lambert D, Stanfield-Oakley SA, et al. Impact of human immunodeficiency virus type 1 gp41 amino acid substitutions selected during enfuvirtide treatment on gp41 binding and antiviral potency of enfuvirtide in vitro. J Virol. 2005;79:12447–12454. doi: 10.1128/JVI.79.19.12447-12454.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Lu J, Deeks SG, Hoh R, Beatty G, Kuritzkes BA, Martin JN, Kuritzkes DR. Rapid emergence of enfuvirtide resistance in HIV-1-infected patients: Results of a clonal analysis. J Acquir Immune Defic Syndr. 2006;43:60–64. doi: 10.1097/01.qai.0000234083.34161.55. [DOI] [PubMed] [Google Scholar]
- 171.Hartley O, Klasse P, Sattentau Q, Moore JP. V3: HIV’s switch hitter. AIDS Res Hum Retrovirus. 2005;21:171–189. doi: 10.1089/aid.2005.21.171. [DOI] [PubMed] [Google Scholar]
- 172.Westby M, Smith-Burchnell C, Mori J, Lewis M, Mosley M, Stockdale M, Dorr P, Ciaramella G, Perros M. Reduced maximal inhibition in phenotypic susceptibility assays indicates that viral strains resistant to the CCR5 antagonist maraviroc utilize inhibitor-bound receptor for entry. J Virol. 2007;81:2359–2371. doi: 10.1128/JVI.02006-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Westby M, Lewis M, Whitcomb J, Youle M, Pozniak AL, James IT, Jenkins TM, Perros M, van der Ryst E. Emergence of CXCR4-using human immunodeficiency virus type 1 (HIV-1) variants in a minority of HIV-1-infected patients following treatment with the CCR5 antagonist maraviroc is from a pretreatment CXCR4-using Virus Res.ervoir. J Virol. 2006;80:4909–4920. doi: 10.1128/JVI.80.10.4909-4920.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Moore JP, Kuritzkes DR. A piece de resistance: How HIV-1 escapes small molecule CCR5 inhibitors. Curr Opin HIV AIDS. 2009;4:118–124. doi: 10.1097/COH.0b013e3283223d46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Soriano V, Perno CF, Kaiser R, Calvez V, Gatell JM, di Perri G, Pillay D, Rockstroh J, Geretti AM. When and how to use maraviroc in HIV-infected patients. AIDS. 2009;23:2377–2385. doi: 10.1097/QAD.0b013e328332d32d. [DOI] [PubMed] [Google Scholar]
- 176.Tilton JC, Doms RW. Entry inhibitors in the treatment of HIV-1 infection. Antivir Res. 2010;85:91–100. doi: 10.1016/j.antiviral.2009.07.022. [DOI] [PubMed] [Google Scholar]
- 177.Shafer RW, Schapiro JM. HIV-1 drug resistance mutations: An updated framework for the second decade of HAART. AIDS Rev. 2008;10:67–84. [PMC free article] [PubMed] [Google Scholar]
- 178.Singh K, Marchand B, Kirby KA, Michailidis E, Sarafianos SG. Structural Aspects of Drug Resistance and Inhibition of HIV-1 Reverse Transcriptase. Viruses. 2010;2:606–638. doi: 10.3390/v2020606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Metifiot M, Marchand C, Maddali K, Pommier Y. Resistance to integrase inhibitors. Viruses. 2010;2:1347–1366. doi: 10.3390/v2071347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Low AJ, Swenson LC, Harrigan PR. HIV coreceptor phenotyping in the clinical setting. AIDS Rev. 2008;10:143–151. [PubMed] [Google Scholar]
- 181.Piccolo P, Lenci I, Demelia L, Bandiera F, Piras MR, Antonucci G, Nosotti L, Mari T, De Santis A, Ponti ML, et al. A randomized controlled trial of pegylated interferon-alpha2a plus adefovir dipivoxil for hepatitis B e antigen-negative chronic hepatitis B. Antivir Ther. 2009;14:1165–1174. doi: 10.3851/IMP1466. [DOI] [PubMed] [Google Scholar]
- 182.Moucari R, Korevaar A, Lada O, Martinot-Peignoux M, Boyer N, Mackiewicz V, Dauvergne A, Cardoso AC, Asselah T, Nicolas-Chanoine MH, et al. High rates of HBsAg seroconversion in HBeAg-positive chronic hepatitis B patients responding to interferon: A long-term follow-up study. J Hepatol. 2009;50:1084–1092. doi: 10.1016/j.jhep.2009.01.016. [DOI] [PubMed] [Google Scholar]
- 183.Takkenberg B, Zaaijer H, Weegink C, Terpstra V, Dijkgraaf M, Jansen P, Janssen H, Beld M, Reesinkg H. Baseline HBsAg level predicts HBsAG loss in chronic hepatitis B patients treated with a combination of peginterferon alfa-2A and adefovir: An interim analysis. Proceeding of the European Association Liver Disease 44th Annual Meeting; Copenhagen, Denmark. April 2009. [Google Scholar]
- 184.Moucari R, Boyer N, Ripault MP, Castelnau C, Mackiewicz V, Dauvergne A, Valla D, Vidaud M, Chanoine MH, Marcellin P. Sequential therapy with adefovir dipivoxil and pegylated Interferon Alfa-2a for HBeAg-negative patients. J Viral Hepat. 2010 doi: 10.1111/j.1365-2893.2010.01332.x. [DOI] [PubMed] [Google Scholar]
- 185.Das K, Xiong X, Yang H, Westland CE, Gibbs CS, Sarafianos SG, Arnold E. Molecular modeling and biochemical characterization reveal the mechanism of hepatitis B virus polymerase resistance to lamivudine (3TC) and emtricitabine (FTC) J Virol. 2001;75:4771–4779. doi: 10.1128/JVI.75.10.4771-4779.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Bartholomeusz A, Tehan BG, Chalmers DK. Comparisons of the HBV and HIV polymerase, and antiviral resistance mutations. Antivir Ther. 2004;9:149–160. [PubMed] [Google Scholar]
- 187.Stuyver LJ, Locarnini SA, Lok A, Richman DD, Carman WF, Dienstag JL, Schinazi RF. Nomenclature for antiviral-resistant human hepatitis B virus mutations in the polymerase region. Hepatology. 2001;33:751–757. doi: 10.1053/jhep.2001.22166. [DOI] [PubMed] [Google Scholar]
- 188.Lai CL, Dienstag J, Schiff E, Leung NW, Atkins M, Hunt C, Brown N, Woessner M, Boehme R, Condreay L. Prevalence and clinical correlates of YMDD variants during lamivudine therapy for patients with chronic hepatitis B. Clin Infect Dis. 2003;36:687–696. doi: 10.1086/368083. [DOI] [PubMed] [Google Scholar]
- 189.Marcellin P, Lau GK, Bonino F, Farci P, Hadziyannis S, Jin R, Lu ZM, Piratvisuth T, Germanidis G, Yurdaydin C, et al. Peginterferon alfa-2a alone, lamivudine alone, and the two in combination in patients with HBeAg-negative chronic hepatitis B. N Engl J Med. 2004;351:1206–1217. doi: 10.1056/NEJMoa040431. [DOI] [PubMed] [Google Scholar]
- 190.Lok AS, Lai CL, Leung N, Yao GB, Cui ZY, Schiff ER, Dienstag JL, Heathcote EJ, Little NR, Griffiths DA, et al. Long-term safety of lamivudine treatment in patients with chronic hepatitis B. Gastroenterology. 2003;125:1714–1722. doi: 10.1053/j.gastro.2003.09.033. [DOI] [PubMed] [Google Scholar]
- 191.Tipples GA, Ma MM, Fischer KP, Bain VG, Kneteman NM, Tyrrell DL. Mutation in HBV RNA-dependent DNA polymerase confers resistance to lamivudine in vivo. Hepatology. 1996;24:714–717. doi: 10.1002/hep.510240340. [DOI] [PubMed] [Google Scholar]
- 192.Allen MI, Deslauriers M, Andrews CW, Tipples GA, Walters KA, Tyrrell DL, Brown N, Condreay LD. Identification and characterization of mutations in hepatitis B Virus Res.istant to lamivudine. Lamivudine Clinical Investigation Group. Hepatology. 1998;27:1670–1677. doi: 10.1002/hep.510270628. [DOI] [PubMed] [Google Scholar]
- 193.Ono SK, Kato N, Shiratori Y, Kato J, Goto T, Schinazi RF, Carrilho FJ, Omata M. The polymerase L528M mutation cooperates with nucleotide binding-site mutations, increasing hepatitis B virus replication and drug resistance. J Clin Invest. 2001;107:449–455. doi: 10.1172/JCI11100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Delaney WEt, Yang H, Westland CE, Das K, Arnold E, Gibbs CS, Miller MD, Xiong S. The hepatitis B virus polymerase mutation rtV173L is selected during lamivudine therapy and enhances viral replication in vitro. J Virol. 2003;77:11833–11841. doi: 10.1128/JVI.77.21.11833-11841.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Warner N, Locarnini S, Kuiper M, Bartholomeusz A, Ayres A, Yuen L, Shaw T. The L80I substitution in the reverse transcriptase domain of the hepatitis B virus polymerase is associated with lamivudine resistance and enhanced viral replication in vitro. Antimicrob Agents Chemother. 2007;51:2285–2292. doi: 10.1128/AAC.01499-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Liaw YF, Gane E, Leung N, Zeuzem S, Wang Y, Lai CL, Heathcote EJ, Manns M, Bzowej N, Niu J, et al. 2-Year GLOBE trial results: Telbivudine Is superior to lamivudine in patients with chronic hepatitis B. Gastroenterology. 2009;136:486–495. doi: 10.1053/j.gastro.2008.10.026. [DOI] [PubMed] [Google Scholar]
- 197.Tenney DJ, Levine SM, Rose RE, Walsh AW, Weinheimer SP, Discotto L, Plym M, Pokornowski K, Yu CF, Angus P, et al. Clinical emergence of entecavir-resistant hepatitis B virus requires additional substitutions in virus already resistant to Lamivudine. Antimicrob Agents Chemother. 2004;48:3498–3507. doi: 10.1128/AAC.48.9.3498-3507.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Tenney DJ, Rose RE, Baldick CJ, Levine SM, Pokornowski KA, Walsh AW, Fang J, Yu CF, Zhang S, Mazzucco CE, et al. Two-year assessment of entecavir resistance in Lamivudine-refractory hepatitis B virus patients reveals different clinical outcomes depending on the resistance substitutions present. Antimicrob Agents Chemother. 2007;51:902–911. doi: 10.1128/AAC.00833-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Sherman M, Yurdaydin C, Simsek H, Silva M, Liaw YF, Rustgi VK, Sette H, Tsai N, Tenney DJ, Vaughan J, et al. Entecavir therapy for lamivudine-refractory chronic hepatitis B: Improved virologic, biochemical, and serology outcomes through 96 weeks. Hepatology. 2008;48:99–108. doi: 10.1002/hep.22323. [DOI] [PubMed] [Google Scholar]
- 200.Tenney DJ, Rose RE, Baldick CJ, Pokornowski KA, Eggers BJ, Fang J, Wichroski MJ, Xu D, Yang J, Wilber RB, et al. Long-term monitoring shows hepatitis B Virus Res.istance to entecavir in nucleoside-naive patients is rare through 5 years of therapy. Hepatology. 2009;49:1503–1514. doi: 10.1002/hep.22841. [DOI] [PubMed] [Google Scholar]
- 201.Choe WH, Hong SP, Kim BK, Ko SY, Jung YK, Kim JH, Yeon JE, Byun KS, Kim KH, Ji SI, et al. Evolution of hepatitis B virus mutation during entecavir rescue therapy in patients with antiviral resistance to lamivudine and adefovir. Antivir Ther. 2009;14:985–993. doi: 10.3851/IMP1417. [DOI] [PubMed] [Google Scholar]
- 202.Chang TT, Gish RG, Hadziyannis SJ, Cianciara J, Rizzetto M, Schiff ER, Pastore G, Bacon BR, Poynard T, Joshi S, et al. A dose-ranging study of the efficacy and tolerability of entecavir in Lamivudine-refractory chronic hepatitis B patients. Gastroenterology. 2005;129:1198–1209. doi: 10.1053/j.gastro.2005.06.055. [DOI] [PubMed] [Google Scholar]
- 203.Westland CE, Yang H, Delaney WEt, Gibbs CS, Miller MD, Wulfsohn M, Fry J, Brosgart CL, Xiong S. Week 48 resistance surveillance in two phase 3 clinical studies of adefovir dipivoxil for chronic hepatitis B. Hepatology. 2003;38:96–103. doi: 10.1053/jhep.2003.50288. [DOI] [PubMed] [Google Scholar]
- 204.Hadziyannis SJ, Tassopoulos NC, Heathcote EJ, Chang TT, Kitis G, Rizzetto M, Marcellin P, Lim SG, Goodman Z, Ma J, et al. Long-term therapy with adefovir dipivoxil for HBeAg-negative chronic hepatitis B. N Engl J Med. 2005;352:2673–2681. doi: 10.1056/NEJMoa042957. [DOI] [PubMed] [Google Scholar]
- 205.Hadziyannis SJ, Tassopoulos NC, Heathcote EJ, Chang TT, Kitis G, Rizzetto M, Marcellin P, Lim SG, Goodman Z, Ma J, et al. Long-term therapy with adefovir dipivoxil for HBeAg-negative chronic hepatitis B for up to 5 years. Gastroenterology. 2006;131:1743–1751. doi: 10.1053/j.gastro.2006.09.020. [DOI] [PubMed] [Google Scholar]
- 206.Angus P, Vaughan R, Xiong S, Yang H, Delaney W, Gibbs C, Brosgart C, Colledge D, Edwards R, Ayres A, et al. Resistance to adefovir dipivoxil therapy associated with the selection of a novel mutation in the HBV polymerase. Gastroenterology. 2003;125:292–297. doi: 10.1016/s0016-5085(03)00939-9. [DOI] [PubMed] [Google Scholar]
- 207.Fung SK, Chae HB, Fontana RJ, Conjeevaram H, Marrero J, Oberhelman K, Hussain M, Lok AS. Virologic response and resistance to adefovir in patients with chronic hepatitis B. J Hepatol. 2006;44:283–290. doi: 10.1016/j.jhep.2005.10.018. [DOI] [PubMed] [Google Scholar]
- 208.Borroto-Esoda K, Miller MD, Arterburn S. Pooled analysis of amino acid changes in the HBV polymerase in patients from four major adefovir dipivoxil clinical trials. J Hepatol. 2007;47:492–498. doi: 10.1016/j.jhep.2007.06.011. [DOI] [PubMed] [Google Scholar]
- 209.Santantonio T, Fasano M, Durantel S, Barraud L, Heichen M, Guastadisegni A, Pastore G, Zoulim F. Adefovir dipivoxil resistance patterns in patients with lamivudine-resistant chronic hepatitis B. Antivir Ther. 2009;14:557–565. [PubMed] [Google Scholar]
- 210.Yang H, Westland C, Xiong S, Delaney WEt. In vitro antiviral susceptibility of full-length clinical hepatitis B virus isolates cloned with a novel expression vector. Antivir Res. 2004;61:27–36. doi: 10.1016/j.antiviral.2003.07.003. [DOI] [PubMed] [Google Scholar]
- 211.Peters MG, Hann Hw H, Martin P, Heathcote EJ, Buggisch P, Rubin R, Bourliere M, Kowdley K, Trepo C, Gray Df, D, et al. Adefovir dipivoxil alone or in combination with lamivudine in patients with lamivudine-resistant chronic hepatitis B. Gastroenterology. 2004;126:91–101. doi: 10.1053/j.gastro.2003.10.051. [DOI] [PubMed] [Google Scholar]
- 212.Westland CE, Yang H, Delaney WEt, Wulfsohn M, Lama N, Gibbs CS, Miller MD, Fry J, Brosgart CL, Schiff ER, et al. Activity of adefovir dipivoxil against all patterns of lamivudine-resistant hepatitis B viruses in patients. J Viral Hepat. 2005;12:67–73. doi: 10.1111/j.1365-2893.2005.00578.x. [DOI] [PubMed] [Google Scholar]
- 213.Yang H, Qi X, Sabogal A, Miller M, Xiong S, Delaney WEt. Cross-resistance testing of next-generation nucleoside and nucleotide analogues against lamivudine-resistant HBV. Antivir Ther. 2005;10:625–633. [PubMed] [Google Scholar]
- 214.Rapti I, Dimou E, Mitsoula P, Hadziyannis SJ. Adding-on versus switching-to adefovir therapy in lamivudine-resistant HBeAg-negative chronic hepatitis B. Hepatology. 2007;45:307–313. doi: 10.1002/hep.21534. [DOI] [PubMed] [Google Scholar]
- 215.Lok AS, Zoulim F, Locarnini S, Bartholomeusz A, Ghany MG, Pawlotsky JM, Liaw YF, Mizokami M, Kuiken C. Antiviral drug-resistant HBV: Standardization of nomenclature and assays and recommendations for management. Hepatology. 2007;46:254–265. doi: 10.1002/hep.21698. [DOI] [PubMed] [Google Scholar]
- 216.Keeffe EB, Dieterich DT, Pawlotsky JM, Benhamou Y. Chronic hepatitis B: Preventing, detecting, and managing viral resistance. Clin Gastroenterol Hepatol. 2008;6:268–274. doi: 10.1016/j.cgh.2007.12.043. [DOI] [PubMed] [Google Scholar]
- 217.Lok AS, McMahon BJ. Chronic hepatitis B: Update 2009. Hepatology. 2009;50:661–662. doi: 10.1002/hep.23190. [DOI] [PubMed] [Google Scholar]
- 218.Yeh CT, Chien RN, Chu CM, Liaw YF. Clearance of the original hepatitis B virus YMDD-motif mutants with emergence of distinct lamivudine-resistant mutants during prolonged lamivudine therapy. Hepatology. 2000;31:1318–1326. doi: 10.1053/jhep.2000.7296. [DOI] [PubMed] [Google Scholar]
- 219.Yatsuji H, Noguchi C, Hiraga N, Mori N, Tsuge M, Imamura M, Takahashi S, Iwao E, Fujimoto Y, Ochi H, et al. Emergence of a novel lamivudine-resistant hepatitis B virus variant with a substitution outside the YMDD motif. Antimicrob Agents Chemother. 2006;50:3867–3874. doi: 10.1128/AAC.00239-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Gerolami R, Bourliere M, Colson P, Halfon P, Borentain P, Henry M, Botta D, Thibault V, Khiri H, Tamalet C. Unusual selection of rtA181V HBV mutants cross-resistant to adefovir following prolonged lamivudine monotherapy: Report of two cases. Antivir Ther. 2006;11:1103–1106. [PubMed] [Google Scholar]
- 221.Villet S, Pichoud C, Billioud G, Barraud L, Durantel S, Trepo C, Zoulim F. Impact of hepatitis B virus rtA181V/T mutants on hepatitis B treatment failure. J Hepatol. 2008;48:747–755. doi: 10.1016/j.jhep.2008.01.027. [DOI] [PubMed] [Google Scholar]
- 222.Warner N, Locarnini S. The antiviral drug selected hepatitis B virus rtA181T/sW172* mutant has a dominant negative secretion defect and alters the typical profile of viral rebound. Hepatology. 2008;48:88–98. doi: 10.1002/hep.22295. [DOI] [PubMed] [Google Scholar]
- 223.Zoulim F, Locarnini S. Hepatitis B Virus Res.istance to nucleos(t)ide analogues. Gastroenterology. 2009;137:1593–1608. e.1591–1592. doi: 10.1053/j.gastro.2009.08.063. [DOI] [PubMed] [Google Scholar]
- 224.Tan J, Degertekin B, Wong SN, Husain M, Oberhelman K, Lok AS. Tenofovir monotherapy is effective in hepatitis B patients with antiviral treatment failure to adefovir in the absence of adefovir-resistant mutations. J Hepatol. 2008;48:391–398. doi: 10.1016/j.jhep.2007.09.020. [DOI] [PubMed] [Google Scholar]
- 225.van Bommel F, de Man RA, Wedemeyer H, Deterding K, Petersen J, Buggisch P, Erhardt A, Huppe D, Stein K, Trojan J, et al. Long-term efficacy of tenofovir monotherapy for hepatitis B virus-monoinfected patients after failure of nucleoside/nucleotide analogues. Hepatology. 2010;51:73–80. doi: 10.1002/hep.23246. [DOI] [PubMed] [Google Scholar]
- 226.Shaw T, Bartholomeusz A, Locarnini S. HBV drug resistance: Mechanisms, detection and interpretation. J Hepatol. 2006;44:593–606. doi: 10.1016/j.jhep.2006.01.001. [DOI] [PubMed] [Google Scholar]
- 227.Schildgen O, Olotu C, Funk A, Zollner B, Helm M, Rockstroh JK, Sirma H. Selection and counterselection of the adefovir resistance mutation rtI233V during antiviral therapy. J Clin Microbiol. 2009 doi: 10.1128/JCM.01073-09.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Schildgen O, Sirma H, Funk A, Olotu C, Wend UC, Hartmann H, Helm M, Rockstroh JK, Willems WR, Will H, et al. Variant of hepatitis B virus with primary resistance to adefovir. N Engl J Med. 2006;354:1807–1812. doi: 10.1056/NEJMoa051214. [DOI] [PubMed] [Google Scholar]
- 229.Curtis M, Zhu Y, Borroto-Esoda K. Hepatitis B virus containing the I233V mutation in the polymerase reverse-transcriptase domain remains sensitive to inhibition by adefovir. J Infect Dis. 2007;196:1483–1486. doi: 10.1086/522521. [DOI] [PubMed] [Google Scholar]
- 230.Sheldon J, Camino N, Rodes B, Bartholomeusz A, Kuiper M, Tacke F, Nunez M, Mauss S, Lutz T, Klausen G, et al. Selection of hepatitis B virus polymerase mutations in HIV-coinfected patients treated with tenofovir. Antivir Ther. 2005;10:727–734. [PubMed] [Google Scholar]
- 231.Qi X, Xiong S, Yang H, Miller M, Delaney WEt. In vitro susceptibility of adefovir-associated hepatitis B virus polymerase mutations to other antiviral agents. Antivir Ther. 2007;12:355–362. [PubMed] [Google Scholar]
- 232.Burton JR, Jr, Everson GT. HCV NS5B polymerase inhibitors. Clin Liver Dis. 2009;13:453–465. doi: 10.1016/j.cld.2009.05.001. [DOI] [PubMed] [Google Scholar]
- 233.Sarrazin C, Zeuzem S. Resistance to direct antiviral agents in patients with hepatitis C virus infection. Gastroenterology. 2010;138:447–462. doi: 10.1053/j.gastro.2009.11.055. [DOI] [PubMed] [Google Scholar]
- 234.Naggie S, Patel K, McHutchison J. Hepatitis C virus directly acting antivirals: Current developments with NS3/4A HCV serine protease inhibitors. J Antimicrob Chemother. 2010;65:2063–2069. doi: 10.1093/jac/dkq284. [DOI] [PubMed] [Google Scholar]
- 235.Gao M, Nettles RE, Belema M, Snyder LB, Nguyen VN, Fridell RA, Serrano-Wu MH, Langley DR, Sun JH, O’Boyle DR, 2nd, et al. Chemical genetics strategy identifies an HCV NS5A inhibitor with a potent clinical effect. Nature. 2010;465:96–100. doi: 10.1038/nature08960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236.Lindenbach BD, Evans MJ, Syder AJ, Wolk B, Tellinghuisen TL, Liu CC, Maruyama T, Hynes RO, Burton DR, McKeating JA, et al. Complete replication of hepatitis C virus in cell culture. Science. 2005;309:623–626. doi: 10.1126/science.1114016. [DOI] [PubMed] [Google Scholar]
- 237.Zhong J, Gastaminza P, Cheng G, Kapadia S, Kato T, Burton DR, Wieland SF, Uprichard SL, Wakita T, Chisari FV. Robust hepatitis C virus infection in vitro. Proc Natl Acad Sci U S A. 2005;102:9294–9299. doi: 10.1073/pnas.0503596102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 238.Yi M, Villanueva RA, Thomas DL, Wakita T, Lemon SM. Production of infectious genotype 1a hepatitis C virus (Hutchinson strain) in cultured human hepatoma cells. Proc Natl Acad Sci U S A. 2006;103:2310–2315. doi: 10.1073/pnas.0510727103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Kato T, Matsumura T, Heller T, Saito S, Sapp RK, Murthy K, Wakita T, Liang TJ. Production of infectious hepatitis C virus of various genotypes in cell cultures. J Virol. 2007;81:4405–4411. doi: 10.1128/JVI.02334-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Pang PS, Planet PJ, Glenn JS. The evolution of the major hepatitis C genotypes correlates with clinical response to interferon therapy. PLoS One. 2009;4:e6579. doi: 10.1371/journal.pone.0006579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Asselah T, Estrabaud E, Bieche I, Lapalus M, De Muynck S, Vidaud M, Saadoun D, Soumelis V, Marcellin P. Hepatitis C: Viral and host factors associated with non-response to pegylated interferon plus ribavirin. Liver Int. 2010;30:1259–1269. doi: 10.1111/j.1478-3231.2010.02283.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Jaeckel E, Cornberg M, Wedemeyer H, Santantonio T, Mayer J, Zankel M, Pastore G, Dietrich M, Trautwein C, Manns MP. Treatment of acute hepatitis C with interferon alfa-2b. N Engl J Med. 2001;345:1452–1457. doi: 10.1056/NEJMoa011232. [DOI] [PubMed] [Google Scholar]
- 243.Gerlach JT, Diepolder HM, Zachoval R, Gruener NH, Jung MC, Ulsenheimer A, Schraut WW, Schirren CA, Waechtler M, Backmund M, et al. Acute hepatitis C: High rate of both spontaneous and treatment-induced viral clearance. Gastroenterology. 2003;125:80–88. doi: 10.1016/s0016-5085(03)00668-1. [DOI] [PubMed] [Google Scholar]
- 244.Farci P, Strazzera R, Alter HJ, Farci S, Degioannis D, Coiana A, Peddis G, Usai F, Serra G, Chessa L, et al. Early changes in hepatitis C viral quasispecies during interferon therapy predict the therapeutic outcome. Proc Natl Acad Sci U S A. 2002;99:3081–3086. doi: 10.1073/pnas.052712599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Morishima C, Polyak SJ, Ray R, Doherty MC, Di Bisceglie AM, Malet PF, Bonkovsky HL, Sullivan DG, Gretch DR, Rothman AL, et al. Hepatitis C virus-specific immune responses and quasi-species variability at baseline are associated with nonresponse to antiviral therapy during advanced hepatitis C. J Infect Dis. 2006;193:931–940. doi: 10.1086/500952. [DOI] [PubMed] [Google Scholar]
- 246.Veillon P, Payan C, Le Guillou-Guillemette H, Gaudy C, Lunel F. Quasispecies evolution in NS5A region of hepatitis C virus genotype 1b during interferon or combined interferon-ribavirin therapy. World J Gastroenterol. 2007;13:1195–1203. doi: 10.3748/wjg.v13.i8.1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Aurora R, Donlin MJ, Cannon NA, Tavis JE. Genome-wide hepatitis C virus amino acid covariance networks can predict response to antiviral therapy in humans. J Clin Invest. 2009;119:225–236. doi: 10.1172/JCI37085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Chary A, Winters MA, Kottilil S, Murphy AA, Polis MA, Holodniy M. Impact of Interferon-Ribavirin Treatment on Hepatitis C Virus (HCV) Protease Quasispecies Diversity in HIV- and HCV-Coinfected Patients. J Infect Dis. 2010;202:889–893. doi: 10.1086/655784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Enomoto N, Sakuma I, Asahina Y, Kurosaki M, Murakami T, Yamamoto C, Ogura Y, Izumi N, Marumo F, Sato C. Mutations in the nonstructural protein 5A gene and response to interferon in patients with chronic hepatitis C virus 1b infection. N Engl J Med. 1996;334:77–81. doi: 10.1056/NEJM199601113340203. [DOI] [PubMed] [Google Scholar]
- 250.Witherell GW, Beineke P. Statistical analysis of combined substitutions in nonstructural 5A region of hepatitis C virus and interferon response. J Med Virol. 2001;63:8–16. [PubMed] [Google Scholar]
- 251.Guo JT, Sohn JA, Zhu Q, Seeger C. Mechanism of the interferon alpha response against hepatitis C virus replicons. Virology. 2004;325:71–81. doi: 10.1016/j.virol.2004.04.031. [DOI] [PubMed] [Google Scholar]
- 252.Brillet R, Penin F, Hezode C, Chouteau P, Dhumeaux D, Pawlotsky JM. The nonstructural 5A protein of hepatitis C virus genotype 1b does not contain an interferon sensitivity-determining region. J Infect Dis. 2007;195:432–441. doi: 10.1086/510316. [DOI] [PubMed] [Google Scholar]
- 253.Cannon NA, Donlin MJ, Fan X, Aurora R, Tavis JE. Hepatitis C virus diversity and evolution in the full open-reading frame during antiviral therapy. PLoS One. 2008;3:e2123. doi: 10.1371/journal.pone.0002123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254.Cuevas JM, Torres-Puente M, Jimenez-Hernandez N, Bracho MA, Garcia-Robles I, Wrobel B, Carnicer F, del Olmo J, Ortega E, Moya A, et al. Genetic variability of hepatitis C virus before and after combined therapy of interferon plus ribavirin. PLoS One. 2008;3:e3058. doi: 10.1371/journal.pone.0003058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Lau JY, Tam RC, Liang TJ, Hong Z. Mechanism of action of ribavirin in the combination treatment of chronic HCV infection. Hepatology. 2002;35:1002–1009. doi: 10.1053/jhep.2002.32672. [DOI] [PubMed] [Google Scholar]
- 256.Shields WW, Pockros PJ. Ribavirin analogs. Clin Liver Dis. 2009;13:419–427. doi: 10.1016/j.cld.2009.05.006. [DOI] [PubMed] [Google Scholar]
- 257.Maag D, Castro C, Hong Z, Cameron CE. Hepatitis C virus RNA-dependent RNA polymerase (NS5B) as a mediator of the antiviral activity of ribavirin. J Biol Chem. 2001;276:46094–46098. doi: 10.1074/jbc.C100349200. [DOI] [PubMed] [Google Scholar]
- 258.Crotty S, Maag D, Arnold JJ, Zhong W, Lau JY, Hong Z, Andino R, Cameron CE. The broad-spectrum antiviral ribonucleoside ribavirin is an RNA virus mutagen. Nat Med. 2000;6:1375–1379. doi: 10.1038/82191. [DOI] [PubMed] [Google Scholar]
- 259.Chevaliez S, Brillet R, Lazaro E, Hezode C, Pawlotsky JM. Analysis of ribavirin mutagenicity in human hepatitis C virus infection. J Virol. 2007;81:7732–7741. doi: 10.1128/JVI.00382-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Penin F, Dubuisson J, Rey FA, Moradpour D, Pawlotsky JM. Structural biology of hepatitis C virus. Hepatology. 2004;39:5–19. doi: 10.1002/hep.20032. [DOI] [PubMed] [Google Scholar]
- 261.Kwong AD, Kim JL, Rao G, Lipovsek D, Raybuck SA. Hepatitis C virus NS3/4A protease. Antivir Res. 1999;41:67–84. [PubMed] [Google Scholar]
- 262.Landro JA, Raybuck SA, Luong YP, O’Malley ET, Harbeson SL, Morgenstern KA, Rao G, Livingston DJ. Mechanistic role of an NS4A peptide cofactor with the truncated NS3 protease of hepatitis C virus: Elucidation of the NS4A stimulatory effect via kinetic analysis and inhibitor mapping. Biochemistry. 1997;36:9340–9348. doi: 10.1021/bi963054n. [DOI] [PubMed] [Google Scholar]
- 263.Kieffer TL, Kwong AD, Picchio GR. Viral resistance to specifically targeted antiviral therapies for hepatitis C (STAT-Cs) J Antimicrob Chemother. 2010;65:202–212. doi: 10.1093/jac/dkp388. [DOI] [PubMed] [Google Scholar]
- 264.Hezode C, Forestier N, Dusheiko G, Ferenci P, Pol S, Goeser T, Bronowicki JP, Bourliere M, Gharakhanian S, Bengtsson L, et al. Telaprevir and peginterferon with or without ribavirin for chronic HCV infection. N Engl J Med. 2009;360:1839–1850. doi: 10.1056/NEJMoa0807650. [DOI] [PubMed] [Google Scholar]
- 265.McHutchison JG, Everson GT, Gordon SC, Jacobson IM, Sulkowski M, Kauffman R, McNair L, Alam J, Muir AJ. Telaprevir with peginterferon and ribavirin for chronic HCV genotype 1 infection. N Engl J Med. 2009;360:1827–1838. doi: 10.1056/NEJMoa0806104. [DOI] [PubMed] [Google Scholar]
- 266.McHutchison JG, Manns MP, Muir AJ, Terrault NA, Jacobson IM, Afdhal NH, Heathcote EJ, Zeuzem S, Reesink HW, Garg J, et al. Telaprevir for previously treated chronic HCV infection. N Engl J Med. 2010;362:1292–1303. doi: 10.1056/NEJMoa0908014. [DOI] [PubMed] [Google Scholar]
- 267.Berman K, Kwo PY. Boceprevir, an NS3 protease inhibitor of HCV. Clin Liver Dis. 2009;13:429–439. doi: 10.1016/j.cld.2009.05.008. [DOI] [PubMed] [Google Scholar]
- 268.Sarrazin C, Kieffer TL, Bartels D, Hanzelka B, Muh U, Welker M, Wincheringer D, Zhou Y, Chu HM, Lin C, et al. Dynamic hepatitis C virus genotypic and phenotypic changes in patients treated with the protease inhibitor telaprevir. Gastroenterology. 2007;132:1767–1777. doi: 10.1053/j.gastro.2007.02.037. [DOI] [PubMed] [Google Scholar]
- 269.Kieffer TL, Sarrazin C, Miller JS, Welker MW, Forestier N, Reesink HW, Kwong AD, Zeuzem S. Telaprevir and pegylated interferon-alpha-2a inhibit wild-type and resistant genotype 1 hepatitis C virus replication in patients. Hepatology. 2007;46:631–639. doi: 10.1002/hep.21781. [DOI] [PubMed] [Google Scholar]
- 270.Susser S, Welsch C, Wang Y, Zettler M, Domingues FS, Karey U, Hughes E, Ralston R, Tong X, Herrmann E, et al. Characterization of resistance to the protease inhibitor boceprevir in hepatitis C virus-infected patients. Hepatology. 2009;50:1709–1718. doi: 10.1002/hep.23192. [DOI] [PubMed] [Google Scholar]
- 271.Reesink HW, Fanning GC, Farha KA, Weegink C, Van Vliet A, Van ‘t Klooster G, Lenz O, Aharchi F, Marien K, Van Remoortere P, et al. Rapid HCV-RNA decline with once daily TMC435: A phase I study in healthy volunteers and hepatitis C patients. Gastroenterology. 2010;138:913–921. doi: 10.1053/j.gastro.2009.10.033. [DOI] [PubMed] [Google Scholar]
- 272.Liverton NJ, Carroll SS, Dimuzio J, Fandozzi C, Graham DJ, Hazuda D, Holloway MK, Ludmerer SW, McCauley JA, McIntyre CJ, et al. MK-7009, a potent and selective inhibitor of hepatitis C virus NS3/4A protease. Antimicrob Agents Chemother. 2010;54:305–311. doi: 10.1128/AAC.00677-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Rajagopalan R, Misialek S, Stevens SK, Myszka DG, Brandhuber BJ, Ballard JA, Andrews SW, Seiwert SD, Kossen K. Inhibition and binding kinetics of the hepatitis C virus NS3 protease inhibitor ITMN-191 reveals tight binding and slow dissociative behavior. Biochemistry. 2009;48:2559–2568. doi: 10.1021/bi900038p. [DOI] [PubMed] [Google Scholar]
- 274.Bartels DJ, Zhou Y, Zhang EZ, Marcial M, Byrn RA, Pfeiffer T, Tigges AM, Adiwijaya BS, Lin C, Kwong AD, et al. Natural prevalence of hepatitis C virus variants with decreased sensitivity to NS3.4A protease inhibitors in treatment-naive subjects. J Infect Dis. 2008;198:800–807. doi: 10.1086/591141. [DOI] [PubMed] [Google Scholar]
- 275.Kuntzen T, Timm J, Berical A, Lennon N, Berlin AM, Young SK, Lee B, Heckerman D, Carlson J, Reyor LL, et al. Naturally occurring dominant resistance mutations to hepatitis C virus protease and polymerase inhibitors in treatment-naive patients. Hepatology. 2008;48:1769–1778. doi: 10.1002/hep.22549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 276.Thibeault D, Bousquet C, Gingras R, Lagace L, Maurice R, White PW, Lamarre D. Sensitivity of NS3 serine proteases from hepatitis C virus genotypes 2 and 3 to the inhibitor BILN 2061. J Virol. 2004;78:7352–7359. doi: 10.1128/JVI.78.14.7352-7359.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 277.Lopez-Labrador FX, Moya A, Gonzalez-Candelas F. Mapping natural polymorphisms of hepatitis C virus NS3/4A protease and antiviral resistance to inhibitors in worldwide isolates. Antivir Ther. 2008;13:481–494. [PubMed] [Google Scholar]
- 278.Beyer BM, Zhang R, Hong Z, Madison V, Malcolm BA. Effect of naturally occurring active site mutations on hepatitis C virus NS3 protease specificity. Proteins. 2001;43:82–88. [PubMed] [Google Scholar]
- 279.Di Marco S, Rizzi M, Volpari C, Walsh MA, Narjes F, Colarusso S, De Francesco R, Matassa VG, Sollazzo M. Inhibition of the hepatitis C virus NS3/4A protease. The crystal structures of two protease-inhibitor complexes. J Biol Chem. 2000;275:7152–7157. doi: 10.1074/jbc.275.10.7152. [DOI] [PubMed] [Google Scholar]
- 280.Courcambeck J, Bouzidi M, Perbost R, Jouirou B, Amrani N, Cacoub P, Pepe G, Sabatier JM, Halfon P. Resistance of hepatitis C virus to NS3-4A protease inhibitors: Mechanisms of drug resistance induced by R155Q, A156T, D168A and D168V mutations. Antivir Ther. 2006;11:847–855. [PubMed] [Google Scholar]
- 281.Prongay AJ, Guo Z, Yao N, Pichardo J, Fischmann T, Strickland C, Myers J, Jr, Weber PC, Beyer BM, Ingram R, et al. Discovery of the HCV NS3/4A protease inhibitor (1R,5S)-N-[3-amino-1-(cyclobutylmethyl)-2,3-dioxopropyl]-3- [2(S)-[[[(1,1-dimethylethyl)amino]carbonyl]amino]-3,3-dimethyl-1-oxobutyl] - 6,6-dimethyl-3-azabicyclo[3.1.0]hexan-2(S)-carboxamide (Sch 503034) II. Key steps in structure-based optimization. J Med Chem. 2007;50:2310–2318. doi: 10.1021/jm060173k. [DOI] [PubMed] [Google Scholar]
- 282.Frecer V, Kabelac M, De Nardi P, Pricl S, Miertus S. Structure-based design of inhibitors of NS3 serine protease of hepatitis C virus. J Mol Graph Model. 2004;22:209–220. doi: 10.1016/S1093-3263(03)00161-X. [DOI] [PubMed] [Google Scholar]
- 283.McCown MF, Rajyaguru S, Le Pogam S, Ali S, Jiang WR, Kang H, Symons J, Cammack N, Najera I. The hepatitis C virus replicon presents a higher barrier to resistance to nucleoside analogs than to nonnucleoside polymerase or protease inhibitors. Antimicrob Agents Chemother. 2008;52:1604–1612. doi: 10.1128/AAC.01317-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 284.Ali S, Leveque V, Le Pogam S, Ma H, Philipp F, Inocencio N, Smith M, Alker A, Kang H, Najera I, et al. Selected replicon variants with low-level in vitro resistance to the hepatitis C virus NS5B polymerase inhibitor PSI-6130 lack cross-resistance with R1479. Antimicrob Agents Chemother. 2008;52:4356–4369. doi: 10.1128/AAC.00444-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 285.He Y, King MS, Kempf DJ, Lu L, Lim HB, Krishnan P, Kati W, Middleton T, Molla A. Relative replication capacity and selective advantage profiles of protease inhibitor-resistant hepatitis C virus (HCV) NS3 protease mutants in the HCV genotype 1b replicon system. Antimicrob Agents Chemother. 2008;52:1101–1110. doi: 10.1128/AAC.01149-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 286.Carroll SS, Ludmerer S, Handt L, Koeplinger K, Zhang NR, Graham D, Davies ME, MacCoss M, Hazuda D, Olsen DB. Robust antiviral efficacy upon administration of a nucleoside analog to hepatitis C virus-infected chimpanzees. Antimicrob Agents Chemother. 2009;53:926–934. doi: 10.1128/AAC.01032-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 287.Migliaccio G, Tomassini JE, Carroll SS, Tomei L, Altamura S, Bhat B, Bartholomew L, Bosserman MR, Ceccacci A, Colwell LF, et al. Characterization of resistance to non-obligate chain-terminating ribonucleoside analogs that inhibit hepatitis C virus replication in vitro. J Biol Chem. 2003;278:49164–49170. doi: 10.1074/jbc.M305041200. [DOI] [PubMed] [Google Scholar]
- 288.Le Pogam S, Jiang WR, Leveque V, Rajyaguru S, Ma H, Kang H, Jiang S, Singer M, Ali S, Klumpp K, et al. In vitro selected Con1 subgenomic replicons resistant to 2’-C-methyl-cytidine or to R1479 show lack of cross resistance. Virology. 2006;351:349–359. doi: 10.1016/j.virol.2006.03.045. [DOI] [PubMed] [Google Scholar]
- 289.Pockros PJ, Nelson D, Godofsky E, Rodriguez-Torres M, Everson GT, Fried MW, Ghalib R, Harrison S, Nyberg L, Shiffman ML, et al. R1626 plus peginterferon Alfa-2a provides potent suppression of hepatitis C virus RNA and significant antiviral synergy in combination with ribavirin. Hepatology. 2008;48:385–397. doi: 10.1002/hep.22357. [DOI] [PubMed] [Google Scholar]
- 290.Gane EJ, Roberts SK, Stedman CA, Angus PW, Ritchie B, Elston R, Ipe D, Morcos PN, Baher L, Najera I, et al. Oral combination therapy with a nucleoside polymerase inhibitor (RG7128) and danoprevir for chronic hepatitis C genotype 1 infection (INFORM-1): A randomised, double-blind, placebo-controlled, dose-escalation trial. Lancet. 2010;376:1467–1475. doi: 10.1016/S0140-6736(10)61384-0. [DOI] [PubMed] [Google Scholar]
- 291.Wang M, Ng KK, Cherney MM, Chan L, Yannopoulos CG, Bedard J, Morin N, Nguyen-Ba N, Alaoui-Ismaili MH, Bethell RC, et al. Non-nucleoside analogue inhibitors bind to an allosteric site on HCV NS5B polymerase. Crystal structures and mechanism of inhibition. J Biol Chem. 2003;278:9489–9495. doi: 10.1074/jbc.M209397200. [DOI] [PubMed] [Google Scholar]
- 292.Kukolj G, McGibbon GA, McKercher G, Marquis M, Lefebvre S, Thauvette L, Gauthier J, Goulet S, Poupart MA, Beaulieu PL. Binding site characterization and resistance to a class of non-nucleoside inhibitors of the hepatitis C virus NS5B polymerase. J Biol Chem. 2005;280:39260–39267. doi: 10.1074/jbc.M506407200. [DOI] [PubMed] [Google Scholar]
- 293.Hang JQ, Yang Y, Harris SF, Leveque V, Whittington HJ, Rajyaguru S, Ao-Ieong G, McCown MF, Wong A, Giannetti AM, et al. Slow binding inhibition and mechanism of resistance of non-nucleoside polymerase inhibitors of hepatitis C virus. J Biol Chem. 2009;284:15517–15529. doi: 10.1074/jbc.M808889200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 294.Le Pogam S, Kang H, Harris SF, Leveque V, Giannetti AM, Ali S, Jiang WR, Rajyaguru S, Tavares G, Oshiro C, et al. Selection and characterization of replicon variants dually resistant to thumb- and palm-binding nonnucleoside polymerase inhibitors of the hepatitis C virus. J Virol. 2006;80:6146–6154. doi: 10.1128/JVI.02628-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 295.Pauwels F, Mostmans W, Quirynen LM, van der Helm L, Boutton CW, Rueff AS, Cleiren E, Raboisson P, Surleraux D, Nyanguile O, et al. Binding-site identification and genotypic profiling of hepatitis C virus polymerase inhibitors. J Virol. 2007;81:6909–6919. doi: 10.1128/JVI.01543-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 296.Howe AY, Cheng H, Johann S, Mullen S, Chunduru SK, Young DC, Bard J, Chopra R, Krishnamurthy G, Mansour T, et al. Molecular mechanism of hepatitis C virus replicon variants with reduced susceptibility to a benzofuran inhibitor, HCV-796. Antimicrob Agents Chemother. 2008;52:3327–3338. doi: 10.1128/AAC.00238-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 297.Le Pogam S, Seshaadri A, Kosaka A, Chiu S, Kang H, Hu S, Rajyaguru S, Symons J, Cammack N, Najera I. Existence of hepatitis C virus NS5B variants naturally resistant to non-nucleoside, but not to nucleoside, polymerase inhibitors among untreated patients. J Antimicrob Chemother. 2008;61:1205–1216. doi: 10.1093/jac/dkn085. [DOI] [PubMed] [Google Scholar]
- 298.Gaudieri S, Rauch A, Pfafferott K, Barnes E, Cheng W, McCaughan G, Shackel N, Jeffrey GP, Mollison L, Baker R, et al. Hepatitis C virus drug resistance and immune-driven adaptations: Relevance to new antiviral therapy. Hepatology. 2009;49:1069–1082. doi: 10.1002/hep.22773. [DOI] [PubMed] [Google Scholar]
- 299.Tellinghuisen TL, Foss KL, Treadaway JC, Rice CM. Identification of residues required for RNA replication in domains II and III of the hepatitis C virus NS5A protein. J Virol. 2008;82:1073–1083. doi: 10.1128/JVI.00328-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 300.Tellinghuisen TL, Marcotrigiano J, Rice CM. Structure of the zinc-binding domain of an essential component of the hepatitis C virus replicase. Nature. 2005;435:374–379. doi: 10.1038/nature03580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 301.Lemm JA, O’Boyle D, 2nd, Liu M, Nower PT, Colonno R, Deshpande MS, Snyder LB, Martin SW, St Laurent DR, Serrano-Wu MH, et al. Identification of hepatitis C virus NS5A inhibitors. J Virol. 2010;84:482–491. doi: 10.1128/JVI.01360-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 302.Fridell RA, Qiu D, Wang C, Valera L, Gao M. Resistance analysis of the hepatitis C virus NS5A inhibitor BMS-790052 in an in vitro replicon system. Antimicrob Agents Chemother. 2010;54:3641–3650. doi: 10.1128/AAC.00556-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 303.Yang F, Robotham JM, Grise H, Frausto S, Madan V, Zayas M, Bartenschlager R, Robinson M, Greenstein AE, Nag A, et al. A major determinant of cyclophilin dependence and cyclosporine susceptibility of hepatitis C virus identified by a genetic approach. PLoS Pathog. 2010;6:e1001118. doi: 10.1371/journal.ppat.1001118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 304.Flisiak R, Feinman SV, Jablkowski M, Horban A, Kryczka W, Pawlowska M, Heathcote JE, Mazzella G, Vandelli C, Nicolas-Metral V, et al. The cyclophilin inhibitor Debio 025 combined with PEG IFNalpha2a significantly reduces viral load in treatment-naive hepatitis C patients. Hepatology. 2009;49:1460–1468. doi: 10.1002/hep.22835. [DOI] [PubMed] [Google Scholar]
- 305.Chatterji U, Lim P, Bobardt MD, Wieland S, Cordek DG, Vuagniaux G, Chisari F, Cameron CE, Targett-Adams P, Parkinson T, et al. HCV resistance to cyclosporin A does not correlate with a resistance of the NS5A-cyclophilin A interaction to cyclophilin inhibitors. J Hepatol. 2010;53:50–56. doi: 10.1016/j.jhep.2010.01.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 306.Di Marco V, Di Stefano R, Ferraro D, Almasio PL, Bonura C, Giglio M, Parisi P, Cappello M, Alaimo G, Craxi A. HBV-DNA suppression and disease course in HBV cirrhosis patients on long-term lamivudine therapy. Antivir Ther. 2005;10:431–439. [PubMed] [Google Scholar]
- 307.Erhardt A, Deterding K, Benhamou Y, Reiser M, Forns X, Pol S, Calleja JL, Ross S, Spangenberg HC, Garcia-Samaniego J, et al. Safety, pharmacokinetics and antiviral effect of BILB 1941, a novel hepatitis C virus RNA polymerase inhibitor, after 5 days oral treatment. Antivir Ther. 2009;14:23–32. [PubMed] [Google Scholar]
- 308.Hirashima S, Suzuki T, Ishida T, Noji S, Yata S, Ando I, Komatsu M, Ikeda S, Hashimoto H. Benzimidazole derivatives bearing substituted biphenyls as hepatitis C virus NS5B RNA-dependent RNA polymerase inhibitors: Structure-activity relationship studies and identification of a potent and highly selective inhibitor JTK-109. J Med Chem. 2006;49:4721–4736. doi: 10.1021/jm060269e. [DOI] [PubMed] [Google Scholar]
- 309.Cooper C, Lawitz EJ, Ghali P, Rodriguez-Torres M, Anderson FH, Lee SS, Bedard J, Chauret N, Thibert R, Boivin I, et al. Evaluation of VCH-759 monotherapy in hepatitis C infection. J Hepatol. 2009;51:39–46. doi: 10.1016/j.jhep.2009.03.015. [DOI] [PubMed] [Google Scholar]
- 310.Tomei L, Altamura S, Bartholomew L, Bisbocci M, Bailey C, Bosserman M, Cellucci A, Forte E, Incitti I, Orsatti L, et al. Characterization of the inhibition of hepatitis C virus RNA replication by nonnucleosides. J Virol. 2004;78:938–946. doi: 10.1128/JVI.78.2.938-946.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 311.McCown MF, Rajyaguru S, Kular S, Cammack N, Najera I. GT-1a or GT-1b subtype-specific resistance profiles for hepatitis C virus inhibitors telaprevir and HCV-796. Antimicrob Agents Chemother. 2009;53:2129–2132. doi: 10.1128/AAC.01598-08. [DOI] [PMC free article] [PubMed] [Google Scholar]