Abstract
Tropomodulin is a tropomyosin-dependent actin filament capping protein involved in the structural formation of thin filaments and in the regulation of their lengths through its localization at the pointed ends of actin filaments. The disordered N-terminal domain of tropomodulin contains three functional sites: two tropomyosin-binding and one tropomyosin-dependent actin-capping sites. The C-terminal half of tropomodulin consists of one compact domain containing a tropomyosin-independent actin-capping site. Here we determined the structural properties of tropomodulin-1 that affect its roles in cardiomyocytes. To explore the significance of individual tropomyosin-binding sites, GFP-tropomodulin-1 with single mutations that destroy each tropomyosin-binding site was expressed in cardiomyocytes. We demonstrated that both sites are necessary for the optimal localization of tropomodulin-1 at thin filament pointed ends, with site 2 acting as the major determinant. To investigate the functional properties of the tropomodulin C-terminal domain, truncated versions of GFP-tropomodulin-1 were expressed in cardiomyocytes. We discovered that the leucine-rich repeat (LRR) fold and the C-terminal helix are required for its proper targeting to the pointed ends. To investigate the structural significance of the LRR fold, we generated three mutations within the C-terminal domain (V232D, F263D, and L313D). Our results show that these mutations affect both tropomyosin-independent actin-capping activity and pointed end localization, most likely by changing local conformations of either loops or side chains of the surfaces involved in the interactions of the LRR domain. Studying the influence of these mutations individually, we concluded that, in addition to the tropomyosin-independent actin-capping site, there appears to be another regulatory site within the tropomodulin C-terminal domain.
Keywords: Actin, Cardiac Muscle, Circular Dichroism (CD), Fluorescence, Protein Assembly, Cardiomyocytes, Leucine-rich Repeat Fold, Thin Filaments, Tropomodulin, Tropomyosin
Introduction
Formation of sarcomeres in striated muscle cells is a complex process that requires participation of both structural proteins for the formation of the sarcomeres and regulatory proteins for controlling the process of assembly. Tropomodulin (Tmod)4 is an actin filament pointed end capping protein involved in both the structural formation of thin filaments in sarcomeres and the regulation of thin filament lengths through its influence on actin dynamics at the pointed ends (for review, see Ref. 1). Tmod is also required for proper myofibril formation. Abundance, deficiency, and/or perturbation of Tmod1 results in improper thin filament lengths (2–5). Overexpression of Tmod1 in mice leads to dilated cardiomyopathy (3). Furthermore, absence of Tmod1 in differentiating embryonic stem cells leads to delayed myofibril assembly, and in mice, it leads to prominent heart defects, including aborted development of the myocardium as the primary defect, together with the inability to pump blood and fragility of primitive red blood cells, leading to embryonic lethality (6–9).
There are four known Tmod isoforms encoded by four genes, with Tmod1 being the most studied. Tmod1 is expressed mainly in the heart, skeletal muscles, and erythrocytes; it is also present in many other tissues but to a lesser extent (10). The C-terminal half of Tmod1 consists of one compact, cooperatively melting domain, whereas the N-terminal half has no cooperatively melting structure; rather, it is flexible and disordered (see Fig. 1A) (11–13). The crystal structure of Tmod1 depicts that the C-terminal domain is composed of alternate α-helices and β-strands, which are arranged in a solenoidal fold (14), characteristic for a leucine-rich repeat (LRR) motif. The solution structure of N-terminal Tmod1 (residues 1–92) was solved using NMR (15). Residues 24–35 are helical, whereas the rest of the peptide has no regular secondary structure. Despite the largely disordered structure, the N-terminal domain of Tmod1 is a major functional domain via its interaction with actin/tropomyosin (TM) filaments.
FIGURE 1.
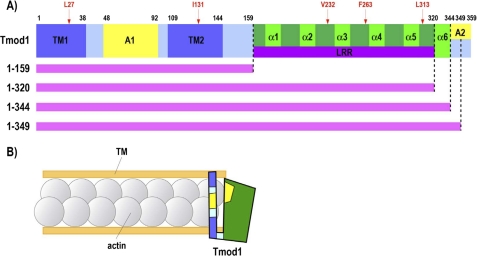
A, domain structure of Tmod1 and mutation sites generated. Tmod1 domain structure was determined based on biochemical, morphological, and structural analysis (for review, see Ref. 27). The Tmod1 molecule consists of two tropomyosin-binding sites (TM1 and TM2), two actin filament-capping regions (A1 and A2), and six α-helixes, including an LRR fold. Mutated amino acid residues and their locations are shown in red. Truncated fragments used in this study are indicated by pink lines (amino acids 1–159, 1–320, 1–344, and 1–349). B, model for Tmod1 actin filament pointed end capping based on previous in vitro experiments. One Tmod1 molecule cooperatively binds two molecules of TM and interacts with at least one or two actin molecules at the pointed end. As shown in A, two TM-binding domains and two actin filament-capping regions are shown in dark blue and yellow, respectively. The C-terminal half of Tmod1 consisting of the LRR is shown in green.
The N-terminal domain of Tmod1 contains three functional sites: two TM-binding sites for both short and long TM isoforms within residues 1–38 and 109–144 and a TM-dependent actin-capping site within residues 48–92 (see Fig. 1A) (15–17). Mutations in all of these sites were shown to cause loss of capping activity in in vitro experiments (17). A TM-independent actin-capping site is located near the C terminus of Tmod1, although the exact location is not known (18, 19). Previous studies have shown that removal of the most C-terminal 15 residues of Tmod1 destroys its capping ability in the absence of TM (19). In the context of sarcomeres in living myocytes, capping is a dynamic process, with actin, TM, and Tmod1 molecules continuously exchanging with free molecules at thin filament pointed ends (4, 20).
The influence of specific mutations in the known binding sites of Tmod1 is well studied in in vitro experiments via TM binding and pyrene actin polymerization assays. Results from these experiments are consistent with the hypothesis that Tmod1 contains structural elements that are not absolutely required for pointed end capping because destroying one of the two TM-binding sites or removal of the entire LRR domain does not affect in vitro TM-dependent actin-capping activity (17). In this work, we attempted to assess the structural properties of Tmod1 that affect its functional properties in neonatal rat cardiomyocytes. Interestingly, we discovered that the requirements for Tmod1 assembly at actin filament pointed ends in myocytes differ from those observed in vitro and that both TM-binding sites are required for its efficient assembly into sarcomeres at WT levels. We concluded that an unknown regulatory site that contributes to targeting Tmod1 to the pointed ends in cardiomyocytes likely exists within the C-terminal domain of Tmod1.
EXPERIMENTAL PROCEDURES
Plasmid Construction
Site-directed mutagenesis of Tmod1 was performed using a QuikChange site-directed mutagenesis kit (Stratagene, La Jolla, CA). Plasmids encoding Tmod1 were amplified by PCR using PfuTurbo DNA polymerase and two complementary sets of oligonucleotides, which contain changed triplets, according to the manufacturer's instructions with one modification. For PCR, instead of mixing all components together, two solutions were made; each of them contained only one complementary oligonucleotide. After four cycles the solutions were mixed, and then 18 additional cycles were performed. To mutate Tmod1 for in vitro experiments, the plasmid encoding chicken Tmod1 (11) was used as a template. For transfection experiments, mouse Tmod1 (accession number NM_21883) was subcloned into pEGFP-C1 (Clontech, Mountain View, CA), and this plasmid was used as a template. After PCR, the original plasmid was digested using DpnI, and the mixture was used to transform Escherichia coli maximum-efficiency DH5α (Invitrogen). After plasmid purification, the presence of mutations was confirmed by DNA sequencing (for mutation sites, see Fig. 1A). Synthesis of all oligonucleotides and DNA sequencing were done at the University of Medicine and Dentistry of New Jersey DNA Synthesis and Sequencing Facility (Robert Wood Johnson Medical School, Piscataway, NJ).
Protein Expression and Purification
Tmod1 (WT and mutants) was overexpressed in Escherichia coli BL21(DE3)pLysS and purified (11). Chicken pectoral skeletal muscle G-actin was purified from acetone powder and labeled with pyrenyl-iodoacetamide (19). N-Acetylated striated muscle α-TM (stTM; TM1a9a) was a generous gift from Sarah Hitchcock-DeGregori (Robert Wood Johnson Medical School). Recombinant human gelsolin was a generous gift from John Hartwig (Brigham and Women's Hospital, Boston, MA). Protein concentration was determined using a BCA protein assay kit (Thermo Fisher Science) or by measuring difference spectra in 6 m guanidine HCl between pH 12.5 and 7.0 (21), using extinction coefficients of 2357 per tyrosine and 830 per tryptophan (22).
Fluorescence Measurements
Actin polymerization in the presence of stTM and Tmod1 was measured as the change in pyrene actin fluorescence using a PTI fluorometer (excitation, 366 nm; emission, 387 nm; with a 2-nm slit) as described (19).
CD Measurements
Far-UV CD spectra of Tmod1 (WT and mutants) were measured using an Aviv Model 400 spectropolarimeter in 1-mm cuvettes. Changes in helical content during urea and temperature denaturation were monitored at 222 nm.
Cell Culture and Transfection Procedure
Cardiomyocytes were isolated from 1–3-day-old neonatal rats and maintained as described (23). Isolated cells were plated in 35-mm tissue culture dishes containing 12-mm round glass coverslips for staining (∼1 × 106 cells/dish). Transfection was performed using Effectene (Qiagen, Valencia, CA) according to the manufacturer's instructions. Briefly, 24 h after plating, cultured myocytes were placed in 1.5 ml of prewarmed culture medium and returned to the incubator while DNA-Effectene lipid complexes were formed. 0.5 μg of plasmid in 1 μl of endotoxin-free TE buffer (10 mm Tris-HCl, 1 mm EDTA, pH 8) was mixed with 1.6 μl of enhancer in 50 μl of optimized buffer and incubated at room temperature for 5 min. Next, 5 μl of Effectene reagent was added, and the mixture was incubated at room temperature for 10 min. After the incubation, the mixture was brought to a 0.5-ml volume using prewarmed culture medium and then added dropwise to the culture dish.
Immunofluorescence Microscopy
Four days after transfection, myocytes were incubated in relaxing buffer (150 mm KCl, 5 mm MgCl2, 10 mm MOPS (pH 7.4), 1 mm EGTA, and 4 mm ATP) for 15 min and fixed with 2% paraformaldehyde in PBS for 15 min. Coverslips were washed and stored in PBS at 4 °C until staining. Transfected cardiac myocytes were essentially stained as described previously (24). Briefly, the fixed cells were permeabilized in 0.2% Triton X-100/PBS for 15 min. The cells were incubated in blocking solution (2% BSA plus 1% normal donkey serum in PBS) and then for 1 h with primary antibodies diluted in blocking solution. The primary antibodies included rabbit anti-GFP antibodies (1:2000) (ab290, Abcam, Cambridge, MA) and monoclonal anti-sarcomeric α-actinin antibodies (1:3000) (EA-53, Sigma-Aldrich). Texas Red-conjugated phalloidin (1:50) (T7471, Invitrogen) was used to stain F-actin. The cells were then incubated with secondary antibodies in PBS for 45 min. Secondary antibodies obtained from Invitrogen included Alexa Fluor 488-conjugated goat anti-rabbit IgG (1:1000) (A11034) and Alexa Fluor 350-conjugated goat anti-mouse IgG (1:350) (A21049). All coverslips were mounted on slides using Aqua Poly/Mount (Polysciences, Warren, PA) and subsequently analyzed on a Zeiss Axiovert microscope using a 100× (NA 1.3) objective, and micrographs were recorded as digital images on a Hamamatsu Photonics ORCA-ER digital camera. Images were processed for presentation using Photoshop® CS (Adobe Systems, San Jose, CA). GFP-positive cardiomyocytes were classified based upon the thin filament pointed end localization of GFP-Tmod1 and its mutants: “consistent,” clear and well defined striated staining with little or no cytoplasmic background; “inconsistent,” faint and partial striated staining with high levels of diffuse cytoplasmic staining; and “diffuse,” no striated staining. The data are presented in terms of percentage of the total number of GFP-positive cardiomyocytes in each group (mean ± S.D.). Data are from three cultures.
Using the AutoThreshold plug-in in the ImageJ 1.41 software (National Institutes of Health), the GFP intensity ratios (assembled GFP versus soluble GFP) were calculated from measurements of 10 consistent cells expressing either WT GFP-Tmod1 or GFP-Tmod1(L27E). Four algorithms (MaxEntropy, RenyiEntropy, Triangle, and Yen) were used to confirm the threshold ratio of the striation (assembled GFP) versus the background (soluble GFP) intensities. Statistical analysis was done using Student's t test (*, p < 0.01).
RESULTS AND DISCUSSION
TM-binding Sites Have Different Significance
Previously we proposed a novel model for Tmod actin filament pointed end capping based on in vitro experiments (Fig. 1B) (17, 25). According to this model, one Tmod molecule cooperatively binds two molecules of TM and interacts with at least one or two actin molecules at the pointed end. In this study, to determine the significance of individual TM-binding sites, GFP-Tmod1 containing mutations that specifically destroy its TM-binding sites were expressed in live neonatal rat cardiomyocytes. We evaluated the thin filament pointed end targeting of Tmod1 mutants by classifying the GFP-positive cardiomyocytes into three groups (consistent, inconsistent, and diffuse).
In previous in vitro studies, dissociation constants calculated for the binding of long muscle TM to each of the Tmod1 TM-binding sites (sites 1 and 2) are comparable: 1.1 ± 0.4 and 1.3 ± 0.3 μm, respectively (17, 26–27). On the basis of these data, we did not anticipate any significant differences in the contribution of each of the two TM-binding sites with respect to the propensity of Tmod1 to target to the pointed ends of thin filaments in live cardiomyocytes. We did, however, hypothesize that we would observe a loss of Tmod1 pointed end assembly when both binding sites were destroyed.
Indeed, when Leu-27 was changed to Glu (L27E) destroying site 1 and Ile-131 was changed to Asp (I131D) destroying site 2 in GFP-Tmod1, the staining pattern of this double mutant displayed no or very faint pointed end localization when expressed in cardiomyocytes (Figs. 2i and 3a). Note, these mutations were previously shown to cause a loss of in vitro TM binding in sites 1 and 2, respectively (15, 17). Unexpectedly, when GFP-Tmod1 containing either of the single mutations (L27E or I131D) was expressed in cardiomyocytes, there was a striking difference between the two with respect to their targeting properties. The pointed end localization of GFP-Tmod1(I131D) was dramatically perturbed (only 7.3 ± 2.5% consistent), whereas the localization of GFP-Tmod1(L27E) was reduced only ∼15% (75.0 ± 3.8% consistent), in comparison with that of WT GFP-Tmod1 (90.0 ± 2.0% consistent) (Figs. 2, e, m, and q, and 3a). The GFP intensity ratios (assembled GFP versus soluble GFP) between cells expressing WT and GFP-Tmod1(L27E) were also quantified using ImageJ software. The ratio of assembly/soluble GFP was significantly lower for GFP-Tmod1(L27E) in comparison with WT GFP-Tmod1 (Fig. 3b). Therefore, these data indicate that 1) both TM-binding sites are necessary for optimal assembly of Tmod1 at actin filament pointed ends and 2) the TM-binding site 2 is primarily responsible for determining the pointed end assembly of Tmod1 in cardiomyocytes.
FIGURE 2.
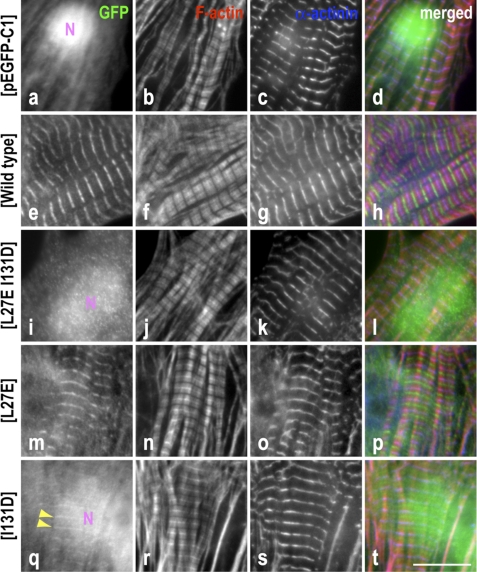
Both leucine 27 and isoleucine 131 residues in Tmod1 are required for thin filament pointed end localization. Neonatal rat cardiomyocytes expressing GFP-Tmod1 (WT or mutants) or GFP alone were stained for GFP (a, e, i, m, and q), F-actin using fluorescently conjugated phalloidin (b, f, j, n, and r), and sarcomeric α-actinin to stain the Z-discs (c, g, k, o, and s). d, h, l, p, and t show merged images of the triple staining (GFP, green; F-actin, red; α-actinin, blue). Clear and consistent striated localization of WT GFP-Tmod1 was observed at thin filament pointed ends (e). GFP-Tmod1(L27E) assembled like WT GFP-Tmod1 (m), but the percentage of the cells displaying striations was lower, and the soluble/unbound GFP molecule in cytoplasm was higher compared with those expressing WT GFP-Tmod1 (Fig. 3). In contrast, GFP-Tmod1(I131D) was rarely observed at the pointed ends (q, arrowheads), and GFP-Tmod1(L27E/I131D) was never observed in a striated pattern (i). GFP-Tmod1(I131D), GFP-Tmod1(L27E/I131D), and GFP alone were observed to be localized in the nucleus (N), all of which do not assemble at thin filament pointed ends. Scale bar = 10 μm.
FIGURE 3.
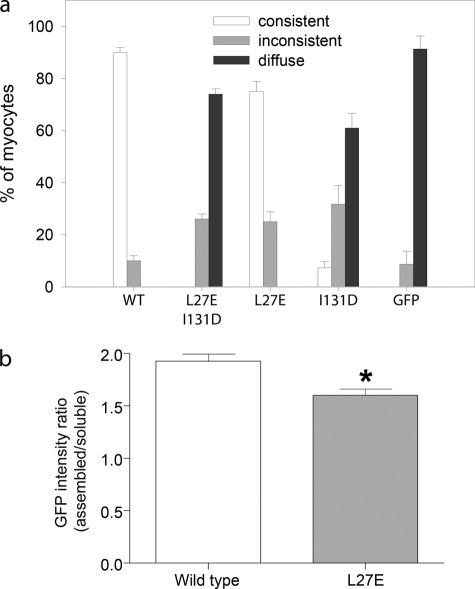
Tropomyosin-binding site 2 is primarily responsible for determining the pointed end localization of Tmod1 in cardiomyocytes. a, graph shows the percentage of myocytes demonstrating consistent, inconsistent, or diffuse thin filament pointed end localization (mean ± S.D.); categories are as described under “Experimental Procedures.” A single mutation within tropomyosin-binding site 2 (I131D) strikingly perturbed Tmod1 pointed end localization (7.3 ± 2.5% cells displayed consistent localization versus 90.0 ± 2.0% consistent localization in cells expressing WT GFP-Tmod1). The localization was completely abolished when an additional mutation within another tropomyosin-binding site (L27E, site 1) was added. However, the single mutation within site 1 (L27E) did not have a strong effect on pointed end localization (75.0 ± 3.8% cells displayed consistent localization). b, graph shows the GFP intensity ratio (assembled/soluble GFP) determined by ImageJ software (Student's t test; *, p < 0.01). Cells expressing GFP-Tmod1(L27E) displayed more diffuse/soluble GFP staining in the cytoplasm compared with cells expressing WT GFP-Tmod1.
Importance of the C-terminal Domain of Tmod1 for Localization at the Thin Filament Pointed Ends
Whereas two binding sites for TM and one binding site for actin are localized in the N-terminal domain of Tmod1, no binding sites have been localized to date in the LRR fold within the C-terminal domain of Tmod1. Tmod1 lacking its C-terminal domain can cap actin filaments in the presence of TM with an affinity close to that of the full-length protein in vitro (17). To investigate a role of the C-terminal domain of Tmod1 in live myocytes, truncated and specifically mutated GFP-Tmod1 constructs were expressed in neonatal rat cardiomyocytes.
In this study, the fragments that were analyzed included Tmod1(1–159) which lacks the entire C-terminal domain; Tmod1(1–320), which lacks the sixth α-helix that is not a part of the LRR fold; Tmod1(1–344), which contains the complete six α-helixes but lacks 15 C-terminal residues, which were previously shown to decrease TM-independent actin-capping activity (19); and Tmod1(1–349), which lacks the 10 C-terminal residues that distinguish Tmod1 from shorter other Tmod isoforms (Fig. 1A).
Interestingly, GFP-Tmod1(1–159) was not observed to assemble at thin filament pointed ends (Figs. 4a and 5a). Although GFP-Tmod1(1–320) also did not assemble well at the pointed ends of the filaments (Figs. 4e and 5a), more than half of the cells expressing GFP-Tmod1(1–320) demonstrated faint and inconsistent striations (67.3 ± 8.3%). In contrast, GFP-Tmod1(1–344) and GFP-Tmod1(1–349) consistently localized at thin filament pointed ends (Figs. 4, i and m, and 5a). The percentage of cells demonstrating consistent pointed end localization of GFP-Tmod1(1–349) was comparable with the cells expressing WT GFP-Tmod1 (97.2 ± 3.1% versus 90.0 ± 2.0%, respectively) (Fig. 5a), whereas GFP-Tmod1(1–344) was ∼15% less efficient at pointed end localization (77.1 ± 0.6%). These data indicate that the C-terminal domain of Tmod1 (specifically, residues 160–349 of Tmod1) is required for targeting Tmod1 to the pointed ends of the actin filaments in sarcomeres. Residues 321–349 increase Tmod1 pointed end localization, and lastly, the terminal 5 amino acids (345–349) appear to increase the affinity for Tmod1 for the pointed ends of the filaments to WT levels.
FIGURE 4.
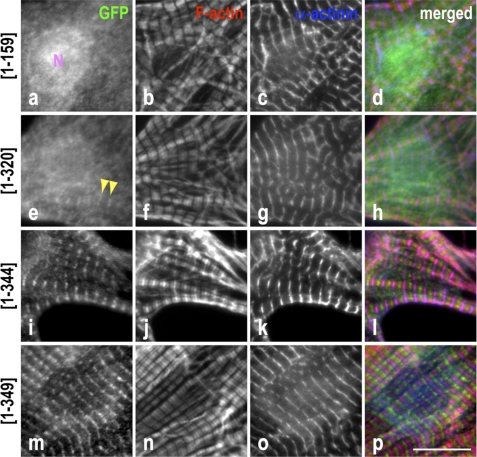
The LRR fold plus the C-terminal α-helix of Tmod1 are required for efficient localization to the pointed ends of thin filaments. Neonatal rat cardiomyocytes expressing GFP-Tmod1 fragments were stained for GFP (a, e, i, and m), F-actin using fluorescently conjugated phalloidin (b, f, j, and n), and α-actinin to stain the Z-discs (c, g, k, and o). d, h, l, and p show merged images of the triple staining (GFP, green; F-actin, red; α-actinin, blue). Clear and consistent striated pointed end localization of GFP-Tmod1(1–344) and GFP-Tmod1(1–349) (i and m) was observed, whereas GFP-Tmod1(1–159) (a) was diffuse in the cytoplasm with nuclear localization (N). Rare and faint (inconsistent) pointed end localization was observed for GFP-Tmod1(1–320) (e, arrowheads). Scale bar = 10 μm.
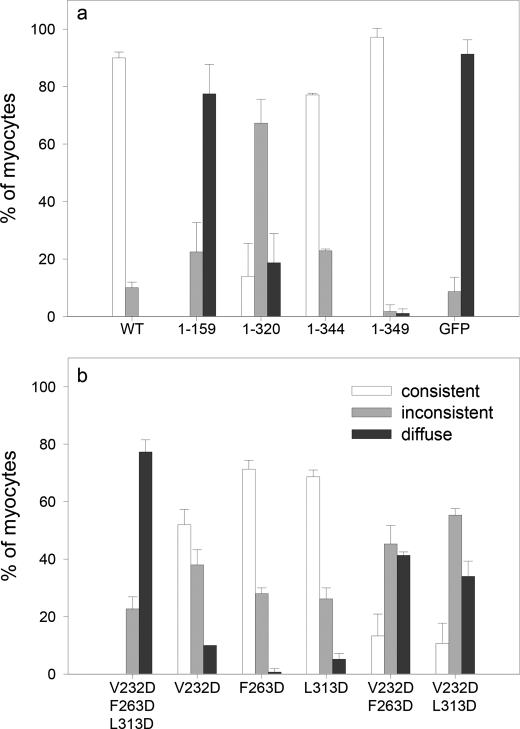
FIGURE 5.
Lack of the C-terminal domain or the addition of a V232D mutation within the LRR fold decreases Tmod1 pointed end localization. The graph shows the percentage of myocytes demonstrating consistent, inconsistent, or diffuse filament pointed end localization (mean ± S.D.); categories are as described under “Experimental Procedures.” a, C-terminal domain truncation. Like WT GFP-Tmod1 (90.2 ± 2.0%), the majority of cells expressing GFP-Tmod1(1–344) (77.1 ± 0.6%) and (1–349) (97.2 ± 3.1%) demonstrated consistent pointed end localization, whereas Tmod1(1–320), which lacks the 6th C-terminal α-helix but contains the entire LRR fold, remarkably perturbed its localization at the pointed ends (14.0 ± 11.5%). Loss of the entire C-terminal domain (GFP-Tmod1(1–159)) resulted in no detectable localization at the pointed ends. b, C-terminal LRR point mutation. Cells expressing GFP-Tmod1(F263D) and GFP-Tmod1(L313D) demonstrated consistent localization at the pointed ends (71.3 ± 3.1% and 68.7 ± 2.3%, respectively). The percentage was lower in the cells expressing GFP-Tmod1(V232D) (52.0 ± 5.3%). Triple and double GFP-Tmod1 mutants (V232D/F263D/L313D, V232D/F263D, and V232D/L313D) further perturbed Tmod1 pointed end localization (0%, 13.3 ± 7.6%, and 10.7 ± 7.0%, respectively).
Because a previous in vitro study demonstrated that the C-terminal domain of Tmod1 is not necessary for capping actin filaments in the presence of TM (17), it appears that this domain has an essential yet unknown regulatory role in cells. Tmod1(1–320) lacks 39 C-terminal residues that include the C-terminal α-helix and disordered C terminus. From our previous studies, it is known that the C terminus is important for the TM-independent actin filament-capping site, although the precise location of this site is not known (19). On the basis of these data, we suggest that both the LRR fold (residues 160–320) and residues 321–349 are important for regulating Tmod pointed end assembly.
Mapping Potential Interaction Surfaces within the LRR Fold
In the previous in vitro experiments, the C-terminal domain (residues 160–359) was not required for Tmod to cap actin/TM filaments; however, this study shows that without this domain Tmod1 is unable to localize to thin filament pointed ends in myocytes (see above). To start deciphering the role of the LRR fold, we decided to identify amino acid residues that may be involved in functionally important interactions and therefore may affect targeting of Tmod1 to the pointed ends. For analysis, a set of homologous Tmod sequences was selected, including the four isoforms of chicken Tmod and two crystal structures of the Tmod1 LRR domain from Gallus gallus and Caenorhabditis elegans (Protein Data Bank codes 1IO0 and 1PGV, correspondingly). Multiple sequence alignments of the selected proteins were mapped onto the crystal three-dimensional structure of the C-terminal half of Tmod (14). This allowed us to identify conserved amino acid residues that are located on the protein surface and are therefore available for intermolecular interactions.
To distinguish the surface side chains, we calculated the fractional accessible surface area for each residue of the structure using Insight II (28). In addition to the conserved residues, we also took into consideration surface residues that, despite some variability, preserve their nonpolar property. The mapping of these residues revealed that they form three clusters on the surface of the Tmod structure, representing potential interaction sites (Figs. 1A and 6). To test the engagement of these clusters in functional interactions, we substituted three nonpolar residues (Val-232, Phe-263, and Leu-313), each located in the center of a corresponding cluster, with charged residues (i.e. aspartates). The logic behind the design of these mutants was that because the selected nonpolar residues are on the surface, they should not affect the structure of Tmod. However, because of drastic changes of the physicochemical properties of the amino acids, these mutations should impair existing functional interactions.
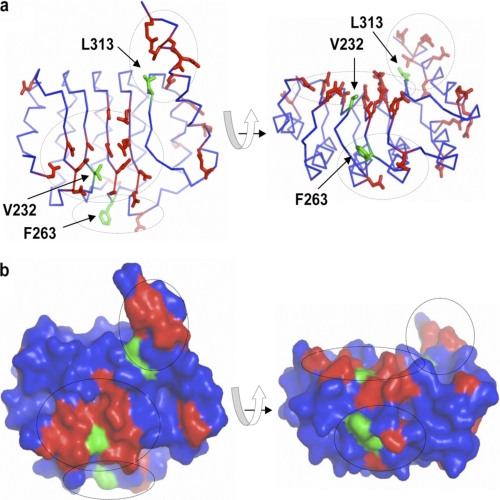
FIGURE 6.
Three-dimensional mapping identifies three potential interaction sites on the surface of Tmod1. The images indicate two projections of the crystal structure of the Tmod1 C-terminal domain (Protein Data Bank code 1IO0). a and b, Cα trace and surface of the protein, correspondingly. Only conserved residues located on the surface of the molecule are shown. Three clusters of these residues are denoted by dotted circles. Green side chains are in the center of corresponding clusters.
Effect of Mutations in the Tmod1 LRR Fold in Vitro and in Cardiac Myocytes
The Tmod1(V232D/F263D/L313D) mutant was expressed in E. coli and purified, and its TM-binding and actin filament-capping properties were studied using the pyrene actin fluorescence assay and compared with WT Tmod1. As expected, no difference in actin-capping abilities was found between Tmod1(V232D/F263D/L313D) and WT Tmod1; both proteins inhibited polymerization of actin in the presence of TM (Fig. 7a). When GFP-Tmod1(V232D/F263D/L313D) was expressed in cardiomyocytes, it was diffusely distributed in the cytoplasm with strong nuclear localization, and no detectable clear, consistent striations were observed (Figs. 5b and 8a). Thus, the triple mutant Tmod1(V232D/F263D/L313D) did not change the ability of Tmod to cap actin filaments in vitro, but prevented its proper targeting to the pointed end of the thin filament in myocytes.
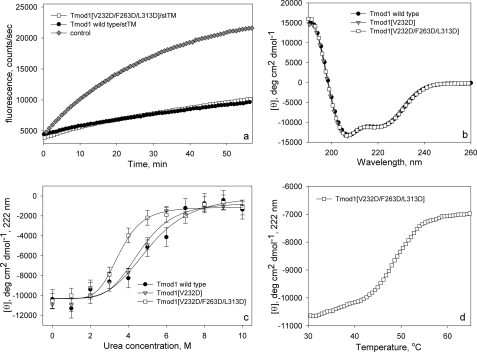
FIGURE 7.
Comparison of structural and functional properties of WT Tmod1 and Tmod1 with mutations in the LRR domain. a, pointed end elongation of gelsolin-capped actin filaments in the presence of stTM and Tmod1. Diamonds, control, no TM or Tmod; circles, with 1 μm stTM and 50 nm WT Tmod1; squares, with 1 μm stTM and 50 nm Tmod1(V232D/F263D/L313D). b, CD spectra of WT Tmod1 and Tmod1 mutants. c, titration of Tmod proteins by urea. d, temperature denaturation of Tmod1(V232D/F263D/L313D) in the presence of 1.5 m urea.
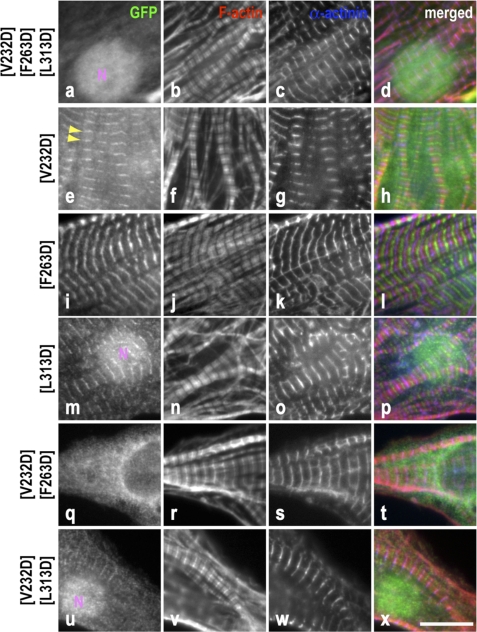
FIGURE 8.
Effect of point mutations in the LRR domain on Tmod1 pointed end localization. Neonatal rat cardiomyocytes expressing GFP-Tmod1 mutants V232D/F263D/L313D, V232D, F263D, L313D, V232D/F263D, and V232D/L313D were stained for GFP (a, e, i, m, q, and u), F-actin (b, f, j, n, r, and v), and α-actinin (c, g, k, o, s, and w). d, h, l, p, t, and x show merged images of the triple staining (GFP, green; F-actin, red; α-actinin, blue). No localization was observed for GFP-Tmod1 triple mutant V232D/F263D/L313D (a). Consistent localization of GFP-Tmod1(V232D) was only observed in 52.0 ± 5.3% of cells (e, arrowheads), whereas 70% of cells expressing GFP-Tmod1(F263D) and GFP-Tmod1(L313D) displayed consistent pointed end localization (71.3 ± 3.1% and 68.7 ± 2.3%, respectively) (i and m). Cells displaying consistent localization were strikingly decreased when the double mutation was expressed (13.3 ± 7.6% and 10.7 ± 7.0, respectively) (q and u). Nuclear localization (N) was observed in the cells expressing GFP-Tmod1 mutants V232D/F263D/L313D, V232D/L313D, and L313D, suggesting that leucine 313 is responsible for cytoplasmic localization of Tmod1. Scale bar = 10 μm.
The secondary structure and stability of WT and the triple-mutated Tmod1 were then compared using CD. These mutations caused no change in the CD spectrum and therefore in the secondary structure (Fig. 7b). The Tmod1 LRR domain represents a single cooperatively melting unit; however, it denatures irreversibly at physiological pH values because of aggregation (13). We next studied Tmod1 stability using denaturation by urea instead of by temperature. Dependence of mean residue ellipticity on urea concentration for WT Tmod1 and Tmod1(V232D/F263D/L313D) is shown in Fig. 7c. The midpoint of the two-state transition corresponded to 4 m urea for Tmod1(V232D/F263D/L313D) unlike 5 m for WT Tmod1. The decrease of the urea concentration necessary for Tmod1 unfolding indicates that there is a change in the tertiary structure, making mutated Tmod1 less stable. To determine whether the LRR domain of mutated Tmod1 is stable at physiological temperatures, we melted Tmod1(V232D/F263D/L313D) in the presence of 1.5 m urea to prevent aggregation (Fig. 7d). The melting temperature, which was determined from the denaturation curve, was 48 °C; therefore, the LRR domain is folded at physiological temperatures.
Next, the influence of the individual mutations on the pointed end localization of full-length Tmod1 was studied (Fig. 8). Each mutation caused a decrease in cells displaying consistent Tmod1 localization. Only 52.0 ± 5.3% of cells expressing V232D displayed consistent striations (Figs. 5b and 8e), whereas the frequency of the cells displaying consistent striations was higher in cells expressing F263D and L313D single mutants (Figs. 5b and 8, i and m) (71.3 ± 3.1% and 68.7 ± 2.3%, respectively). We also studied the structural properties of Tmod1 with the V232D mutation because this mutation had a more pronounced effect on its localization. The CD spectrum of Tmod1(V232D) was the same as for WT Tmod1 (Fig. 7b). Furthermore, unlike Tmod1(V232D/F263D/L313D), Tmod1(V232D) had the same stability as WT Tmod1 (Fig. 7c). Therefore, the effect of the V232D mutation on Tmod1 localization should not be a result of changes in its structure.
We also studied the influence of double mutations (V232D/F263D and V232D/L313D) on the distribution of Tmod1 (Fig. 8, q and u). With both sets of mutations, there was a drastic decrease in cells displaying consistent localization (13.3 ± 7.6% and 10.7 ± 7.0, respectively) (Fig. 5b).
Interestingly, nuclear localization was observed in the myocytes expressing GFP-Tmod1(L313D), but not with the other single mutants (Fig. 8m). Therefore, Leu-313 may be an important residue contributing to the cytoplasmic localization of Tmod1.
Effect of Mutations in the Tmod1 LRR Fold on TM-independent Actin-capping Activity
The influence of the mutations in Tmod1 that we engineered on its pointed end localization may be explained by the fact that although the three mutations V232D/F263D/L313D do not affect its secondary structure, they affect the tertiary structure of the LRR domain so it does not function properly. This potential effect may also happen in the case of the double mutants. Introducing additional charges may drastically change the position of other charged surface residues, causing disruption of the tertiary structure. However, in the case of the single Val-232 mutation, which had the most pronounced effect on the pointed end distribution in cells, the secondary and tertiary structures were not affected (see above). Therefore, there should be another explanation for the effect. A possible explanation is that the V232D mutation perturbed the TM-independent actin-capping site that is located at the C-terminal part of Tmod1, and it somehow affects Tmod1 assembly. A second possible explanation is that Val-232 is a crucial residue in the formation of a binding site for an unknown factor that regulates Tmod1 localization to thin filament pointed ends.
To determine the influence mutations in the LRR fold had on TM-independent capping, WT Tmod1, Tmod1(V232D/F263D/L313D), Tmod1(V232D), and Tmod1(L313D) were tested in a pyrene actin fluorescence assay in the absence of TM (Fig. 9). All mutants decreased the TM-independent capping activity, whereas the triple mutant decreased the capping activity the most; the capping ability of the triple mutant at 1.5 μm is 60% of the WT capping ability (Fig. 9b). Analysis of the two Tmod1 proteins with single mutations demonstrated that the L313D mutation had a bigger effect on the TM-independent actin-capping ability; at 1.5 μm concentration, the ability of Tmod1(V232D) to cap the pointed ends reaches the level of WT, whereas that of Tmod1(L313D) is only ∼80% of the WT capping ability (Fig. 9b). Interestingly, whereas all these mutations decrease TM-independent actin-capping activity, there appears to be no strict correlation between the decrease of actin-capping activity and perturbation in assembly of Tmod1 at the pointed ends. Our results cannot be explained by disruption of the TM-independent actin-capping function only. We believe that the Val-232 mutation is located in a site responsible for binding unknown regulatory factors.
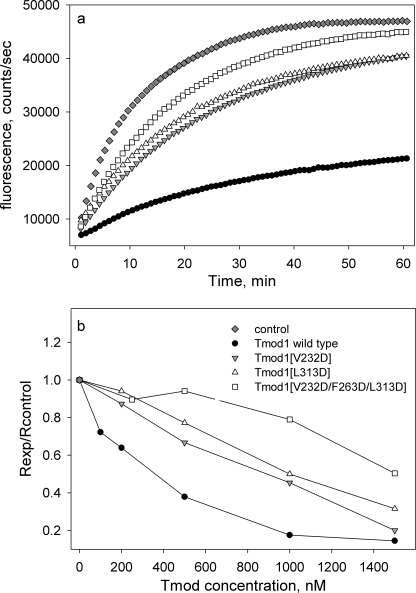
FIGURE 9.
Effect of mutations in the Tmod1 C-terminal domain on TM-independent actin capping. a, pyrene fluorescence that measures actin polymerization is plotted versus time. Initial protein concentrations were 6 nm gelsolin-capped filaments, 1.1 μm G-actin, and 1 μm WT or mutant Tmod1. b, dependence of Tmod1 concentration on inhibition of actin polymerization. The polymerization data were fitted to a three-parameter exponential equation using SigmaPlot and initial polymerization rates (Rexp) were calculated as the first derivative at time 0. Rexp values were then normalized by the initial rate obtained for polymerization of actin on gelsolin-capped actin seeds (Rcontrol).
Nuclear Localization of Tmod1 Mutants
The Tmod1 mutants studied in our experiments differ with respect to their nuclear localization, although it should be noted that we did not observe nuclear localization of WT GFP-Tmod1. Specifically, Tmod1 (L27E), Tmod1(V232D), Tmod1(F263D), Tmod1(V232D/F263D), Tmod1(1–320), and Tmod1(1–344) have cytoplasmic perinuclear localization. Tmod1(1–159), Tmod1(I131D), Tmod1(L27E/I131D), Tmod1(L313D), Tmod1(V232D/L313D), Tmod1(V232D/F263D/L313D), and GFP alone showed nuclear localization.
Previously, Kong and Kedes (29) reported that mutations at residues 134 and 135 of Tmod1 caused nuclear localization of Tmod1 in fibroblasts and C2 myogenic cells, which was explained by the disruption of a likely nuclear export signal motif located in this region. The authors also stated that amino acids 340–343 (RKRR) represent a pattern 4 nuclear localization signal, and Tmod1 with K341A and R342G mutations was excluded from nuclei. Our data are not fully consistent with these conclusions. We observed that Tmod1 with the I131D mutation was localized in nuclei (Fig. 2q); this mutation was in the region determined by Kong and Kedes as the nuclear export signal. However, this region was not impaired in Tmod1(1–159); moreover, this mutant did not contain the region determined by Kong and Kedes as the nuclear localization signal, but Tmod1 (1–159) was localized in nuclei (Fig. 4a).
Furthermore, we did not find a clear correlation between the ability of Tmod1 to localize at thin filament pointed ends and within the nucleus (Figs. 3a and 5, a and b). Also, our data cannot simply be explained by nuclear localization of GFP itself. For example, the double mutants Tmod1(V232D/F263D) and Tmod1(V232D/L313D) similarly have a decreased ability to localize at the pointed ends; however, only one of them appears in the nucleus (Tmod1(V232D/L313D)) (Fig. 8u). Thus, their localization depends on the presence of the Leu-313 mutation. Therefore, we believe that there are other explanations for this phenomenon that still need to be clarified.
CONCLUSIONS
We found that the requirements for Tmod1 localization at actin filament pointed ends in vivo differ from those observed in in vitro. Two novel and important conclusions can be generated from our data. 1) Both TM-binding sites are necessary for optimal assembly of Tmod1 at the pointed end of the thin filaments, and 2) the LRR domain is necessary for pointed end assembly of Tmod1. Removal of the LRR domain, as well as alteration of its structure by three point mutations, resulted in complete loss of the pointed end localization of Tmod1 in cardiac myocytes.
The fact that both TM-binding sites are necessary for the localization of Tmod1 at thin filament pointed ends confirmed our previous model where one Tmod molecule binds both TM molecules at the pointed end (Fig. 1b) (17). Of the two TM-binding sites, site 2 (residues 109–144) is primarily responsible for determining the pointed end localization of Tmod in sarcomeres. Because the position of the last two actin molecules at the pointed end is not symmetrical, there are various possible ways for Tmod to interact with the terminal actin subunits. Although it is important to note that it is still unclear how Tmod1 is localized with respect to the terminal actin molecules at the pointed ends or how the correct orientation of the molecules is determined, we predict based on the data obtained in this study that the differences we determined between the two TM-binding sites may guide Tmod1 to the pointed end with correct orientation (Fig. 10b).
FIGURE 10.
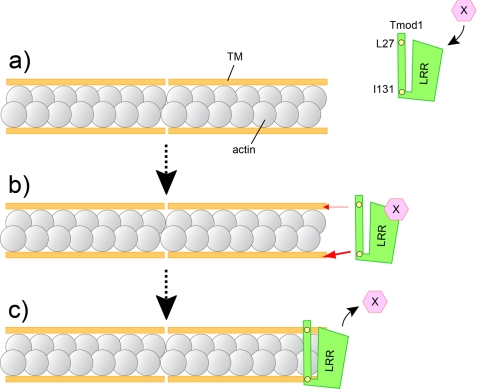
Possible in vivo model of the Tmod1 capping at thin filament pointed ends in cardiomyocytes. a, an unidentified regulatory factor (X) binds to the LRR within Tmod1; the interaction assists Tmod1 to be targeted to thin filament pointed ends in cardiomyocytes. b, both TM-binding sites 1 and 2 are required for proper assembly of Tmod1 to the thin filament pointed ends in cardiac myocytes. Yellow circles are critical amino acid residues for TM binding site 1 (Leu-27) and site 2 (Ile-131), respectively. TM-binding site 2 contributes to the pointed end distribution of Tmod1 more than TM-binding site 1 (the thickness of the red arrow indicates the degree of contribution). The differences presumably guide Tmod1 to thin filament pointed ends with correct orientation. c, model for Tmod1 actin filament pointed end capping based on previous in vitro observation (17, 26) and present results. One Tmod1 molecule cooperatively binds two molecules of TM and interacts with at least one or two actin molecules at the pointed end.
A previous study showed that the Tmod1 C-terminal domain is not necessary for Tmod1 TM-dependent capping function in vitro (17), but it is clearly required in the context of sarcomeres in living cardiomyocytes. Although TM-independent actin filament-capping activity was drastically reduced by mutations within LRR, we did not identify any correlation between a decrease in Tmod1 pointed end targeting in cardiomyocytes and a decrease (loss) in TM-independent actin-capping activity in in vitro experiments.
As a result of these observations, we hypothesize that the C-terminal domain of Tmod1 binds an unidentified regulatory factor(s) whose interaction allows/assists Tmod1 to be targeted to thin filament pointed ends, therefore regulating the ability of Tmod to cap actin filaments in vivo (Fig. 10a). The LRR fold is generally regarded as a ligand-interacting domain (30). The three conserved clusters we mapped in this study are located on different sides of the LRR domain. According to the docking model proposed in Fig. 1B and Ref. 14, the C-terminal α-helix, as well as the hydrophobic armpit of this helix formed by Ile-283, Leu-313, Leu-338, and Val-339, interacts with F-actin; in this model, it is less plausible that the βα-surface of the LRR domain is involved in the interface with F-actin. However, the conserved cluster that contains Val-232 is located at the βα-surface, and the addition of the V232D mutation has the biggest effect on pointed end distribution; therefore, it appears that this cluster may provide a binding site for this unidentified regulator, and mutations in this site may destroy this functional interaction.
The drastic decrease in TM-independent activity observed in in vitro actin polymerization experiments and absence of the pointed end localization in cells when the three mutations V232D/F263D/L313D were introduced into the Tmod1 LRR domain may be explained by affecting the local conformations of either loops or side chains of the surfaces involved in the interactions of the LRR domain. Proper folding of the LRR domain is important for both TM-independent activity and the pointed end distribution. We suggest that unlike TM-binding sites in the N-terminal half of Tmod1, the binding site for the potential unknown regulatory factor, as well as the TM-independent actin-capping site, is formed by tertiary structure elements of the LRR domain rather than by secondary structure elements.
Acknowledgments
We thank Ellen Taylor and Marcos DeMarco for technical assistance.
This work was supported, in whole or in part, by National Institutes of Health Grant HL083146 (to C. C. G.), National Institutes of Health IMSD Fellowship 5R25GM62584 (to S. M. N.), and National Institutes of Health Grant GM081688 (to A. S. K.). This work was also supported by American Heart Association Grant 0825870G to (T. T.) and University of Medicine and Dentistry of New Jersey Foundation grants (to A. S. K.).
- Tmod
- tropomodulin
- LRR
- leucine-rich repeat
- TM
- tropomyosin
- stTM
- striated muscle α-TM.
REFERENCES
- 1. Littlefield R. S., Fowler V. M. (2008) Semin. Cell Dev. Biol. 19, 511–519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Gregorio C. C., Weber A., Bondad M., Pennise C. R., Fowler V. M. (1995) Nature 377, 83–86 [DOI] [PubMed] [Google Scholar]
- 3. Sussman M. A., Welch S., Cambon N., Klevitsky R., Hewett T. E., Price R., Witt S. A., Kimball T. R. (1998) J. Clin. Invest. 101, 51–61 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Littlefield R., Almenar-Queralt A., Fowler V. M. (2001) Nat. Cell Biol. 3, 544–551 [DOI] [PubMed] [Google Scholar]
- 5. Mudry R. E., Perry C. N., Richards M., Fowler V. M., Gregorio C. C. (2003) J. Cell Biol. 162, 1057–1068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Ono Y., Schwach C., Antin P. B., Gregorio C. C. (2005) Dev. Biol. 282, 336–348 [DOI] [PubMed] [Google Scholar]
- 7. Fritz-Six K. L., Cox P. R., Fischer R. S., Xu B., Gregorio C. C., Zoghbi H. Y., Fowler V. M. (2003) J. Cell Biol. 163, 1033–1044 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. McKeown C. R., Nowak R. B., Moyer J., Sussman M. A., Fowler V. M. (2008) Circ. Res. 103, 1241–1248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Chu X., Chen J., Reedy M. C., Vera C., Sung K. L., Sung L. A. (2003) Am. J. Physiol. Heart Circ. Physiol. 284, H1827–H1838 [DOI] [PubMed] [Google Scholar]
- 10. Conley C. A., Fritz-Six K. L., Almenar-Queralt A., Fowler V. M. (2001) Genomics 73, 127–139 [DOI] [PubMed] [Google Scholar]
- 11. Kostyukova A., Maeda K., Yamauchi E., Krieger I., Maéda Y. (2000) Eur. J. Biochem. 267, 6470–6475 [DOI] [PubMed] [Google Scholar]
- 12. Fujisawa T., Kostyukova A., Maéda Y. (2001) FEBS Lett. 498, 67–71 [DOI] [PubMed] [Google Scholar]
- 13. Kostyukova A. S., Tiktopulo E. I., Maéda Y. (2001) Biophys. J. 81, 345–351 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Krieger I., Kostyukova A., Yamashita A., Nitanai Y., Maéda Y. (2002) Biophys. J. 83, 2716–2725 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Greenfield N. J., Kostyukova A. S., Hitchcock-DeGregori S. E. (2005) Biophys. J. 88, 372–383 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kostyukova A. S., Rapp B. A., Choy A., Greenfield N. J., Hitchcock-DeGregori S. E. (2005) Biochemistry 44, 4905–4910 [DOI] [PubMed] [Google Scholar]
- 17. Kostyukova A. S., Choy A., Rapp B. A. (2006) Biochemistry 45, 12068–12075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Fowler V. M., Greenfield N. J., Moyer J. (2003) J. Biol. Chem. 278, 40000–40009 [DOI] [PubMed] [Google Scholar]
- 19. Kostyukova A. S., Hitchcock-DeGregori S. E. (2004) J. Biol. Chem. 279, 5066–5071 [DOI] [PubMed] [Google Scholar]
- 20. Pappas C. T., Krieg P. A., Gregorio C. C. (2010) J. Cell Biol. 189, 859–870 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Edelhoch H. (1967) Biochemistry 6, 1948–1954 [DOI] [PubMed] [Google Scholar]
- 22. Fasman G. D. (1989) Practical Handbook of Biochemistry and Molecular Biology, p. 84, CRC Press, Boca Raton, FL [Google Scholar]
- 23. Gustafson T. A., Bahl J. J., Markham B. E., Roeske W. R., Morkin E. (1987) J. Biol. Chem. 262, 13316–13322 [PubMed] [Google Scholar]
- 24. Pappas C. T., Bhattacharya N., Cooper J. A., Gregorio C. C. (2008) Mol. Biol. Cell 19, 1837–1847 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kostyukova A. S., Hitchcock-Degregori S. E., Greenfield N. J. (2007) J. Mol. Biol. 372, 608–618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Greenfield N. J., Fowler V. M. (2002) Biophys. J. 82, 2580–2591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Kostyukova A. S. (2008) Cell Mol. Life Sci. 65, 563–569 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Dayringer H. E., Tramontano A., Sprang S. R., Fletterick R. J. (1986) J. Mol. Graph. Model. 4, 82–87 [Google Scholar]
- 29. Kong K. Y., Kedes L. (2004) J. Biol. Chem. 279, 30856–30864 [DOI] [PubMed] [Google Scholar]
- 30. Kobe B., Kajava A. V. (2001) Curr. Opin. Struct. Biol. 11, 725–732 [DOI] [PubMed] [Google Scholar]