Abstract
Background
The PII protein family comprises homotrimeric proteins which act as transducers of the cellular nitrogen and carbon status in prokaryotes and plants. In Herbaspirillum seropedicae, two PII-like proteins (GlnB and GlnK), encoded by the genes glnB and glnK, were identified. The glnB gene is monocistronic and its expression is constitutive, while glnK is located in the nlmAglnKamtB operon and is expressed under nitrogen-limiting conditions.
Results
In order to determine the involvement of the H. seropedicae glnB and glnK gene products in nitrogen fixation, a series of mutant strains were constructed and characterized. The glnK- mutants were deficient in nitrogen fixation and they were complemented by plasmids expressing the GlnK protein or an N-truncated form of NifA. The nitrogenase post-translational control by ammonium was studied and the results showed that the glnK mutant is partially defective in nitrogenase inactivation upon addition of ammonium while the glnB mutant has a wild-type phenotype.
Conclusions
Our results indicate that GlnK is mainly responsible for NifA activity regulation and ammonium-dependent post-translational regulation of nitrogenase in H. seropedicae.
Background
The PII family comprises homotrimeric proteins that have important roles in the control of the central metabolism in bacteria and plants, acting as transducers of the cellular nitrogen and carbon levels [1,2]. In many Proteobacteria studied there is a pair of PII proteins, usually called GlnB and GlnK, and their function is to sense the cellular levels of nitrogen, carbon and energy by binding the effectors 2-oxoglutarate, ATP and ADP [2,3]. These signals are then relayed to target proteins through conformational changes triggered by interaction with the effectors. The proteobacterial PII proteins also undergo a cycle of uridylylation/deuridylylation catalyzed by the bifunctional GlnD protein [1] in response to the intracellular levels of nitrogen. These conformational and covalent state changes stimulate or inhibit interactions of PII with different cellular protein targets involved in nitrogen and carbon metabolism [2].
PII proteins are key players in the regulation of nitrogen fixation in Proteobacteria. In Klebsiella pneumoniae and Azotobacter vinelandii, GlnK is required to regulate the activity of NifL, which inhibits NifA, the nif gene specific activator, under nitrogen-excess conditions [4-6]. In Azospirillum brasilense and Rhodospirillum rubrum GlnB is necessary for the activation of NifA under nitrogen-limiting conditions [7-9], whereas in Rhodobacter capsulatus both PII proteins are necessary for the NH4+-dependent regulation of NifA activity [10]. In addition, PII proteins are also involved in the post-translational control of nitrogenase activity in R. rubrum [11] and in A. brasilense through interaction with DraT, DraG and AmtB [12].
Herbaspirillum seropedicae is a nitrogen-fixing β-Proteobacterium isolated from the rhizosphere and tissues of several plants, including economically important species [13]. In this organism two PII-like coding genes were identified, glnB and glnK [14,15]. The glnB gene is monocistronic and its expression is constitutive [14], whereas glnK is apparently co-transcribed with amtB and orf1, which encode for an ammonium transporter and a membrane associated protein of unknown function, respectively [15]. Recently orf1 was named nlmA (nitrogen limitation membrane protein A) since its product was detected in membrane extracts of H. seropedicae grown under nitrogen-limitation conditions [16]. The expression of the nlmAglnKamtB operon is dramatically increased under nitrogen-limiting conditions and is dependent on NtrC [15]. As in other Proteobacteria, both PII proteins from H. seropedicae are targets of covalent modification by GlnD (uridylyl-transferase/uridylyl removing enzyme) in response to the levels of ammonium ions [17].
Results and Discussion
To analyze the role of GlnK and GlnB in the control of nitrogen fixation in H. seropedicae, glnB (LNglnB) and glnK (LNglnK) insertional mutants and a glnK in-frame deletion mutant strain (LNglnKdel) were constructed and their phenotypes analyzed under different physiological conditions. These mutant strains were able to grow using nitrate as sole nitrogen source (data not shown).
The effect of glnB and glnK disruption on the NtrC-dependent expression of the nlmAglnKamtB operon [15] was determined using chromosomal amtB::lacZ transcriptional fusions of strains LNamtBlacZ, LNglnBamtBlacZ and LNglnKamtBlacZ. These strains were grown under N-limiting (5 mmol/L glutamate or 2 mmol/L NH4Cl) or N-excess (20 mmol/L NH4Cl) conditions and assayed for β-galactosidase. The LNamtBlacZ strain grown under N-limiting conditions showed β-galactosidase activity 21 times higher than in high ammonium (Table 1), confirming that nlmAglnKamtB is highly expressed under N-limiting conditions [15]. Strains LNglnKamtBlacZ and LNglnBamtBlacZ revealed a similar pattern of amtB expression, indicating that the mutation of either glnK or glnB does not affect nlmAglnKamtB expression. Since nlmAglnKamtB transcription is NtrC-dependent, these results suggest that GlnB and GlnK can substitute for each other in control of the NtrC/NtrB system in H. seropedicae. In agreement with this suggestion, ntrC [18] and glnD (unpublished results) mutants strains of H. seropedicae are unable to grow on nitrate, whereas the glnB and glnK mutant strains can use nitrate as sole nitrogen source.
Table 1.
Effect of glnB and glnK mutations on nlmAglnKamtB expression
| Growth Conditions | β-galactosidase Activity [nmol o-nitrophenol/(min.mg protein)] | ||
|---|---|---|---|
| Strains | |||
| LNamtBlacZ (SmR1, amtB::lacZ) |
LNglnKamtBlacZ (ΔglnK, amtB::lacZ) |
LNglnBamtBlacZ (glnB-TcR, amtB::lacZ) |
|
| 5 mmol/L glutamate | (2.5 ± 0.2) × 103 | (2.4 ± 0.2) × 103 | (2.3 ± 0.2) × 103 |
| 2 mmol/L NH4Cl | (2.1 ± 0.1) × 103 | (2.29 ± 0.08) × 103 | (2.2 ± 0.1) × 103 |
| 20 mmol/L NH4Cl | (1.1 ± 0.2) × 102 | (1.4 ± 0.4) × 102 | (1.6 ± 0.3) × 102 |
Indicated strains of H. seropedicae were grown in the presence of glutamate or NH4Cl. β-galactosidase activity was determined as described. Values are the mean of at least three independent experiments ± standard deviation.
In Escherichia coli both GlnB and GlnK are involved in the regulation of NtrC phosphorylation by NtrB, although GlnB is more effective [19]. Although several attempts were made, we failed to construct a double glnBglnK mutant suggesting that an essential role is shared by these proteins in H. seropedicae.
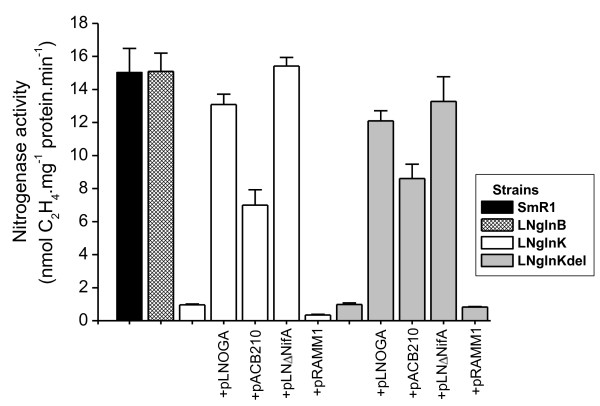
The effect of glnK or glnB mutation on nitrogenase activity of H. seropedicae was determined in cultures grown in NH4+-free semi-solid NFbHP medium (Figure 1). Nitrogenase activity was reduced by approximately 95% in both glnK strains (LNglnKdel and LNglnK) indicating that GlnK is required for nitrogenase activity in H. seropedicae. On the other hand, the glnB strain (LNglnB) showed activity similar to that of the wild-type. These results contrast with those reported by Benelli et al [14] who constructed a H. seropedicae glnB::Tn5-20B mutant (strain B12-27) that was unable to fix nitrogen. Immunoblot assays did not detect GlnK in the B12-27 strain [Additional file 1: Supplemental Figure S1], suggesting that a secondary recombination event may have happened in this strain resulting in loss of GlnK not observed by Benelli et al [14].
Figure 1.
Nitrogenase activity of H. seropedicae wild-type, glnB and glnK strains. Nitrogenase activity was determined as described using strains SmR1 (wild-type), LNglnB (glnB-TcR), LNglnK (glnK-KmR), LNglnKdel (ΔglnK) grown in semi-solid medium. The glnK mutants carrying plasmids pLNOGA, pACB210, pLNΔNifA or pRAMM1, which respectively express NmlA-GlnK-AmtB, GlnB, ΔN-NifA and NifA were also evaluated. Data represent the average of at least three independent experiments and bars indicate the standard deviations.
The nitrogenase phenotype of the glnK mutants was complemented by pLNOGA (nlmAglnKamtB) and also partially restored (about 50%) by a plasmid expressing glnB under control of its own promoter (pACB210) suggesting that a higher copy number of glnB can substitute for glnK under N-limitation. The lower nitrogenase activity of the glnK strains could be due to lack of nif expression or inhibition of nitrogenase. We therefore analyzed the effect of the glnK mutation on the NtrC-dependent nifA promoter [20] and on the NifA-dependent nifB promoter of H. seropedicae [21] by using plasmids carrying nifA::lacZ (pRW1) or nifB::lacZ (pEMS140) fusions (Table 2). The β-galactosidase activity was the same in both wild-type (SmR1) and glnK (LNglnK) strains containing nifA::lacZ, supporting the view that GlnK is not strictly necessary for NtrC regulation in H. seropedicae in the presence of a functional glnB gene. On the other hand, expression of the nifB::lacZ fusion was reduced 10-fold in the glnK mutant compared to the wild-type, indicating that GlnK is required for nifB expression in H. seropedicae, even in the presence of wild type glnB. These results indicate that the lower nitrogenase activity in the glnK mutants was the result of lack of nif expression, most likely due to impaired NifA activity.
Table 2.
Promoter activity of nifA::lacZ and nifB::lacZ fusions in H. seropedicae wild-type (SmR1) and glnK mutant (LNglnK) strains
| Strains | β-galactosidase Activity [nmol o-nitrophenol/(min.mg protein)] | |||
|---|---|---|---|---|
| Plasmids | ||||
| none | pPW452 (promoter-less lacZ vector) |
pRW1 (nifA::lacZ) |
pEMS140 (nifB::lacZ) |
|
| SmR1 | (3 ± 1) × 10 | (6 ± 2) × 10 | (7 ± 1) × 102 | (2.8 ± 0.1) × 103 |
| LNglnK | (2.0 ± 0.7) × 10 | (4 ± 2) × 10 | (6 ± 1) × 102 | (2.5 ± 0.3) × 102 |
H. seropedicae strains carrying the indicated plasmids were grown in NFbHP medium supplemented with 10 mmol/L of NH4Cl under air at 30°C. The cells were then centrifuged, resuspended in NFbHP (nitrogen-free) medium and de-repressed for 7 hours under 1.5% oxygen. β-galactosidase was determined as described. Values are averages of at least three independent experiments ± standard deviation
Previous results showed that the N-terminal domain of H. seropedicae NifA is required for controlling its activity in response to NH4+, and that an N-truncated form of NifA is transcriptionally active, but not responsive to NH4+ levels [22,23]. Thus, the nitrogenase activity was determined in the glnK mutants carrying pRAMM1 or pLNΔNifA which express a full NifA and an N-truncated form of NifA, respectively (Figure 1). The nitrogenase activity of the glnK mutants was restored only by the N-truncated-NifA protein, reinforcing the indication that the nitrogenase negative phenotype of glnK strain is due to the presence of an inactive NifA.
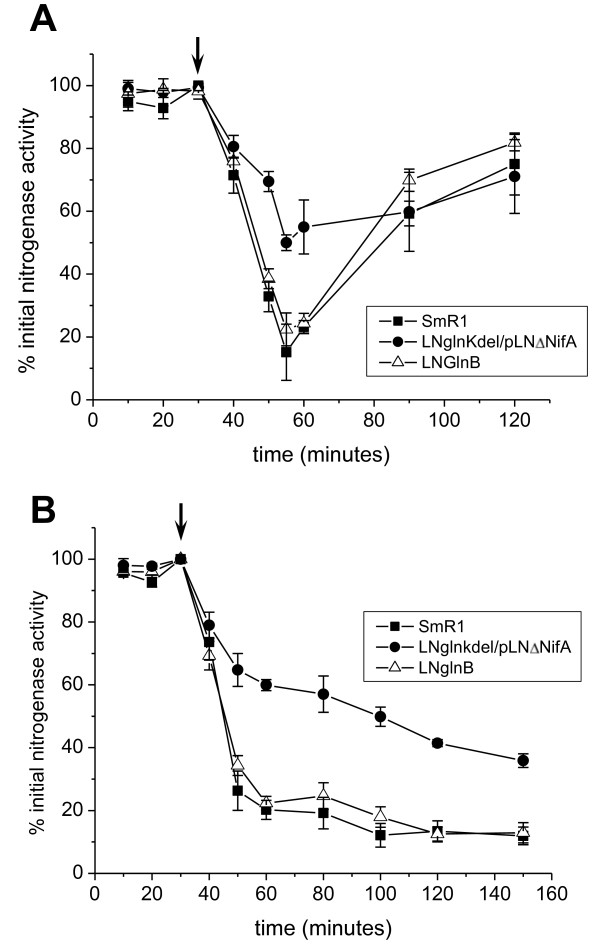
Nitrogenase activity is reversibly inhibited by addition of ammonium or energy depletion in several diazotrophs, a phenomenon called nitrogenase switch-off. The best studied process is the reversible NifH ADP-ribosylation carried out by the DraT and DraG enzymes whose activities are controlled by processes involving PII proteins at least in some diazotrophs [11,12,24,25]. A PII protein has also been implicated in the control of nitrogenase by direct interaction with NifH in the methanogenic archaeon Methanococcus maripaludis [26]. In H. seropedicae reversible ADP-ribosylation of NifH by the DraT/DraG does not occur since draTG genes are absent [27] [GenBank:CP002039]. Although the mechanism of NH4+-dependent nitrogenase control in this organism is not known, it is thought to be due to change in prevailing physiological conditions leading to nitrogenase inhibition. Since the glnK mutant is Nif-, we used strain LNglnKdel carrying plasmid pLNΔNifA for the switch-off experiments. Addition of low concentrations of NH4Cl (300 μmol/L) to derepressed cells caused an inactivation of nitrogenase (Figure 2A). Wild-type and glnB strains retained less than 20% of initial nitrogenase activity 25 minutes after ammonium addition, which was restored to 60-70% of initial activity 60 minutes after ammonium addition. This effect does not involve protein synthesis since the presence of chloramphenicol or tetracycline had no effect on this behavior [28]. Although nitrogenase of strain LNglnKdel/pLNΔNifA was partially inhibited by ammonium addition, the strain retained about 50% of its initial activity, indicating only a partial nitrogenase switch-off (Figure 2A). After addition of 1 mmol/L of NH4Cl (Figure 2B) the activity of the wild-type and glnB strains dropped sharply to less than 30% and did not recover even 120 minutes after ammonium addition. In contrast, 40% of the initial nitrogenase activity was still present in the glnK strain 120 minutes after ammonium addition and the decrease in nitrogenase activity was slower: 20 minutes after ammonium addition the wild-type had only 25% activity, whereas the glnK strain had about 65% of the original nitrogenase activity. These results indicate that GlnK is involved in the nitrogenase inactivation by NH4+ in H. seropedicae, and that GlnB cannot fully replace GlnK in triggering nitrogenase switch-off. It is interesting to note that there was also a delay in nitrogenase reactivation in the glnK mutant (Figure 2A), which may suggest that GlnK is involved in both nitrogenase inactivation by NH4+ and re-activation upon NH4+ depletion.
Figure 2.
Effect of ammonium ions on nitrogenase activity in H. seropedicae wild-type, glnB and glnK strains. Nitrogenase switch-off/on of H. seropedicae wild-type, glnB and glnK carrying plasmid pLNΔNifA was performed as described. Cells were grown under nitrogenase de-repressing conditions when NH4Cl was added (arrow). Samples were analyzed 10, 20 and 30 minutes after acetylene injection to confirm linear nitrogenase activity. Panel A: addition of NH4Cl (0.3 mmol.L-1). Panel B: addition of NH4Cl (1 mmol.L-1). The results represent the average of experiments with three independent cultures and bars indicate the standard deviation.
Recently results using a proteomic approach [16] showed that H. seropedicae GlnK is associated with the membrane at higher concentration immediately after addition of ammonium. This membrane association was shown to be AmtB-dependent, as shown in several bacteria [2,16]. Previous results also showed that an amtB mutant has a partial NH4+ switch off very similar to that shown by the glnK mutant[15]. These results allow us to propose a model for the regulation of nitrogen fixation in H. seropedicae. Under N-limiting conditions, NtrC-dependent promoters are activated leading to expression of nifA and nlmAglnKamtB genes. The status of fixed nitrogen is signaled to NtrC via the uridylylation state of either GlnB or GlnK. Under a low ammonium and oxygen condition, NifA activates the expression of nif genes in a process which requires GlnK, most probably in an uridylylated form. Thus, under N-limiting conditions the nitrogenase complex is active, AmtB is associated with the membrane, NlmA is most probably in the periplasm and GlnK is mainly located in the cytoplasm. When ammonium is added, deuridylylated GlnK rapidly associates with the cell membrane by interacting with AmtB to form the GlnK-AmtB complex which, in turn, signals to nitrogenase to switch-off by a yet unknown process.
Conclusions
In summary, our results show that both GlnB and GlnK proteins can regulate NtrC-dependent promoters in H. seropedicae. Under physiological conditions, GlnK is required for NifA activity control. GlnK also controls the nitrogenase switch-off in response to NH4+ by a mechanism which most probably involves the formation of a membrane-bound GlnK-AmtB complex.
Methods
Plasmids, Bacterial strains and Growth conditions
The H. seropedicae and E. coli strains and plasmids used in this work are listed in Table 3. E. coli strains were grown routinely in Luria medium (Luria broth or Luria agar) [29] at 37°C. H. seropedicae was grown at 30°C in NFbHP medium [30] supplemented with NH4Cl (20 mmol/L) or the indicated nitrogen source. The concentrations of the antibiotics used were as follows: ampicillin (250 μg/mL), tetracycline (10 μg/mL), kanamycin (100 μg/mL for E. coli, 1 mg/mL for H. seropedicae), streptomycin (80 μg/mL) and choramphenicol (30 μg/mL for E. coli, 100 μg/mL for H. seropedicae).
Table 3.
Herbaspirillum seropedicae strains and plasmids
| Strains | Phenotype/genotype | Reference |
|---|---|---|
| Herbaspirillum seropedicae | ||
| SmR1 | Wild type, Nif+, SmR | [38] |
| LNglnK | SmR1 containing glnK::sacB- KmR | this work |
| LNglnKdel | SmR1 containing ΔglnK | this work |
| LNglnB | SmR1 containing glnB::TcR | this work |
| LNamtBlacZ | SmR1 containing amtB::lacZ-KmR | this work |
| LNglnKamtBlacZ | LNglnKdel containing amtB::lacZ-KmR | this work |
| LNglnBamtBlacZ | LNglnB containing amtB::lacZ-KmR | this work |
| B12-27 | SmR1 containing glnB::Tn5-20B | [14] |
| Escherichia coli | ||
| DH10B | Smr; F' [proAB+ lacZΔM15] | Life Technologies |
| S17.1 | SmR, Tra+ pro thi recA hsdR (RP4-2 kan::Tn7 tet::Mu) | [39] |
| Plasmids | Relevant characteristics | Reference |
| pACB192 | 1.7 kb DNA fragment containing the glnB gene of H. seropedicae in pSUP202 | This work |
| pACB194 | glnB gene of H. seropedicae with a tetracycline resistance transposon EZ::TN™ < TET-1 > (Epicentre) in pSUP202 | this work |
| pACB210 | glnB gene of H. seropedicae in pLAFR3.18Cm | this work |
| pDK6 | Expression vector/tac promoter, KmR | [37] |
| pDK6nifACT | H. seropedicae nifA deleted of 606 bp in the 5'coding region cloned into pDK6 carrying the nifA promoter | this work |
| pDK6pnifA | nifA gene promoter region of H. seropedicae in pDK6 | this work |
| pEMS140 | nifB-lacZ transcriptional fusion of H. seropedicae in pPW452 | [21] |
| pEMS301 | 1.7 kb EcoRI fragment that contains the promoter region and part of the nifA gene of H. seropedicae in pTZ19R | [40] |
| pLAFR3.18Cm | TcR, CmR, IncP cosmid with the pTZ18R cloning nest | [15] |
| pLNΔNifA | Expresses ΔN-NifA of H. seropedicae with its own promoter in pLAFR3.18Cm | this work |
| pLNOGA | 5.1 kb fragment that contains the nlmAglnKamtB operon of H. seropedicae in pLAFR3.18Cm (former named pLARF3.18OGA) | [15] |
| pLNglnK | 0.9 kb BamHI/HindIII fragment that contains the 3' terminal of the nlmA gene, the complete glnK gene and 5' terminal of the amtB gene of H. seropedicae in pTZ18R | this work |
| pMH1701 | KmR, contains a sacB-KmR cassette | [35] |
| pPW452 | TcR, transcriptional lacZ gene fusion | [41] |
| pRAM2T7 | contains H. seropedicae nifA deleted of 606 bp in the 5'end, encoding an N-truncated form of NifA deleted of its N-terminal domain and Q-linker | this work |
| pRAMM1 | nifA of H. seropedicae in pLAFR3.18Cm | this work |
| pRW1 | nifA-lacZ transcriptional fusion of H. seropedicae in pPW452 | [20] |
| pSUP202 | ApR, CmR, TcR, Mob | [39] |
| pSUPamtBClacZ | Central region of the amtB gene with a lacZ-KmR cassette insertion in pSUP202 | [15] |
| pSUPglnK | 0.9 kb BamHI/HindIII fragment that contains the 3' terminal of the nlmA gene, the complete glnK gene and 5' terminal of the amtB gene of H. seropedicae in pSUP202 | this work |
| pSUPglnKdel | ΔglnK (192bp) gene of H. seropedicae in pSUP202 | this work |
| pSUPglnKdelsacB | contains ΔglnK and a sacB-KmR cassette (from pMH1701) cloned into the vector pSUP202 | this work |
| pSUPglnKsacB | 0.9 kb fragment spanning from the 3'end of nlmA to the 5'end of amtB with a sacB-KmR (from pMH1701) inserted into the glnK gene | this work |
| pTZ19R | ApR lacZ f1 IG | [42] |
| pUC18 | ApR, lacZ, f1 | Invitrogen |
| pUCG08del | 0.8 kb DNA fragment that contains the 3' terminal of the nlmA gene, the complete glnK gene and the 5' terminal of the amtB gene of H. seropedicae in pUC18. | this work |
| pUCglnKdel | ΔglnK gene of H. seropedicae in pUC18 | this work |
Enzyme assays
β-galactosidase activity was determined in cells carrying a lacZ fusion as described [31]. To study the amtB-lacZ-KmR chromosomal fusion expression, H. seropedicae strains carrying chromosomal transcriptional fusions were grown for 14 hours in NFbHP medium containing glutamate (5 mmol/L) or NH4Cl (2 mmol/L or 20 mmol/L), and assayed for β-galactosidase activity. To study the nifA and nifB expression, H. seropedicae strains carrying plasmid-borne transcriptional fusions nifA::lacZ or nifB::lacZ were grown for 14 hours in NFbHP medium containing NH4Cl (10 mmol/L) under air at 30°C. The cells were then centrifuged, resuspended in 3 mL of NFbHP medium (O.D.600 = 0.2) and incubated in 25 mL flasks, at 30°C for 7 hours under 1.5% oxygen. The results are reported as nmol of o-nitrophenol (NP) produced per min per mg protein. Protein concentration was determined by the Bradford method [32] using bovine serum albumin as standard.
Nitrogenase activity was determined using cells grown in semi-solid NFbHP medium containing glutamate (0.5 mmol/L). For nitrogenase switch-off/on assays cells were grown in liquid NFbHP medium with glutamate (4 mmol/L) at 30°C and 120 rpm [28]. Nitrogenase activity was determined by acetylene reduction [33,34].
Construction of the LNglnB mutant of H. seropedicae
Plasmid HS26-FP-00-000-021-E03 (Genopar consortium, http://www.genopar.org), which contains the H. seropedicae glnB gene in pUC18, was linearized with EcoRI and treated with T4DNA polymerase. It was then digested with HindIII to release a 1.7 kb fragment containing the glnB gene. This fragment was subcloned into the vector pSUP202 previously linearized with BamHI, treated with T4DNA polymerase and digested with HindIII to produce plasmid pACB192.
In vitro transposon mutagenesis of the glnB gene carried by plasmid pACB192 was performed using the EZ::TN™ <TET-1> Insertion Kit (Epicentre Technologies) following the manufacturer's instructions. A plasmid containing the transposon insertion in the glnB coding region was selected and named pACB194. This plasmid was introduced by conjugation to H. seropedicae SmR1 using E. coli strain S17.1 as the donor. Recombinant colonies were selected for tetracycline resistance and screened for the loss of chloramphenicol resistance (vector marker). Southern blot of restriction enzyme digested genomic DNA was used to confirm the presence of the transposon in the glnB gene (data not shown). This H. seropedicae glnB-TcR strain was named LNglnB.
Construction of the LNglnK mutant of H. seropedicae
To clone the glnK gene, chromosomal DNA of H. seropedicae was amplified using the primers glnKD (5'-GACTGAAAGGATCCGCGTGTCC-3', BamHI restriction site is underlined) and glnKR (5'-CGAGGGCAAAGCTTCTTCGGTGG-3', HindIII restriction site is underlined). The amplified fragment was then ligated into BamHI/HindIII-cut pTZ18R, generating the plasmid pLNglnK. This BamHI/HindIII fragment containing the glnK gene was then subcloned into pSUP202, yielding plasmid pSUPglnK. A sacB-KmR cassette excised with BamHI from pMH1701 [35] was inserted into the BglII site of the glnK gene. The resulting plasmid (pSUPglnKsacB) was transferred into H. seropedicae SmR1 by conjugation using E. coli strain S17.1 as the donor. Mutant colonies were selected for kanamycin resistance and screened for the loss of chloramphenicol resistance, as before. Hybridization of genomic DNA was used to confirm the presence of the cassette in the glnK gene (data not shown). This glnK-KmR mutant was named LNglnK.
Construction of the LNglnKdel mutant of H. seropedicae
To construct a mutant containing an in-frame 192 bp deletion of the glnK gene, plasmid pUCG08del containing the 3' terminus of the nlmA gene, the glnK gene and the 5' terminus of the amtB gene was used as a template in two distinct PCR reactions. Primers M13universal and GlnKdelR (5' AAGCCCTCGAGTTCAGTCACGGT 3', XhoI restriction site is underlined) were used to amplify a 180 bp region upstream of glnK and the first 107 bp of the glnK gene. The primers M13reverse and GlnKdelD (5' GGACCTGCTCGAGGTGATCCGT 3', XhoI restriction site is underlined) were used to amplify the last 58 bp of the glnK gene and the first 180 bp of amtB. The amplified fragments were joined by the XhoI sites. This fragment containing glnK deleted of 192 bp was then used as template for a PCR reaction with the primers M13reverse and M13universal. The resulting PCR product was digested with BamHI and PstI and inserted into pUC18 to give pUCglnKdel. This fragment was then subcloned into pSUP202, yielding the plasmid pSUPglnKdel. A sacB-KmR cassette excised with BamHI from pMH1701 [35] was inserted into the vector region of the BamHI-cut pSUPglnKdel plasmid. The resulting plasmid (pSUPglnKdelsacB) was conjugated into H. seropedicae SmR1 using E. coli strain S17.1 as the donor. Recombinant colonies were selected for kanamycin and chloramphenicol resistance. One mutant strain was selected, and grown overnight in liquid NFbHP medium supplemented with ammonium chloride (20 mmol/L) and 80 μg/mL streptomycin. One microliter of the culture was plated on solid NFbHP medium supplemented with 20 mmol/L NH4Cl, 5% sucrose and 80 μg/mL streptomycin. Sucrose is toxic to bacteria containing the sacB gene in the chromosome, therefore only strains that lost the sacB-KmR cassette by a second homologous recombination event would grow. The selected strains were analyzed by PCR with the primers GlnKF1 (5'TGTCCAAGACCTTCGACG3') and GlnKR1 (5'CATGCTCATTAGAGTTCC3') which were homologous to the glnK flanking 5'- and 3'- regions, confirming the deletion of the 192 bp glnK fragment (data not shown). This in-frame glnK strain (ΔglnK) was named LNglnKdel.
Construction of plasmid pLNΔNifA
An Eco47III/SacI DNA fragment containing the nifA gene promoter region of H. seropedicae was excised from the plasmid pEMS301[36] and sub-cloned into the SmaI/SacI-cut vector pDK6 [37], yielding plasmid pDK6pnifA. An XbaI DNA fragment encoding for the central and C-terminal region of NifA protein (ΔN-NifA) of H. seropedicae was excised from the plasmid pRAM2T7 and sub-cloned into the XbaI-cut pDK6pnifA, in the same orientation as the nifA promoter, yielding plasmid pDK6nifACT. Finally, a SacI/HindIII DNA fragment containing the nifA 5'-truncated gene was excised from pDK6nifACT and sub-cloned into pLAFR3.18Cm digested with SacI and HindIII. The generated plasmid was named pLNΔNifA and encodes for the central and C-terminal domains of NifA under control of the nifA promoter.
Construction of the plasmid pACB210
A 1.7 kb EcoRI-HindIII fragment containing the glnB gene with its promoter region was excised from the plasmid HS10-MP-00-000-014-E08 (Genopar consortium, http://www.genopar.org), and sub-cloned into the vector pLAFR3.18 digested with EcoRI-HindIII to yield plasmid pACB210.
Construction of chromosomal amtB::lacZ transcriptional fusions
To construct amtB-lacZ transcriptional fusions, the suicide plasmid pSUPamtBClacZ was introduced by conjugation, using E. coli strain S17.1 as the donor, into H. seropedicae strains SmR1, LNglnKdel and LNglnB resulting in the strains LNamtBlacZ, LNglnKamtBlacZ and LNglnBamtBlacZ, respectively. Genomic DNA hybridization confirmed the presence of the cassette lacZ-KmR in the amtB gene (data not shown).
List of Abbreviations
ApR: ampicillin resistance; CmR: chloramphenicol resistance, KmR: kanamycin resistance; TcR: tetracycline resistance; SmR: streptomycin resistance.
Authors' contributions
LN constructed plasmids and H. seropedicae mutants, carried out physiological experiments and helped to draft the manuscript; ACB constructed plasmids and carried out immunoassays; RAM constructed plasmids and designed some of the experiments; LN, RAM and LUR helped to draft the manuscript; FOP, EMS, MBRS and LSC conceived the study, participated in its design and in writing the manuscript, LSC also supervised the study. All authors read and approved the final manuscript.
Supplementary Material
Immunoblot analysis of H. seropedicae PII proteins.
Contributor Information
Lilian Noindorf, Email: noindorf@yahoo.com.br.
Ana C Bonatto, Email: anacbonatto@ufpr.br.
Rose A Monteiro, Email: rose.adele@terra.com.br.
Emanuel M Souza, Email: souzaem@ufpr.br.
Liu U Rigo, Email: lurigo@ufpr.br.
Fabio O Pedrosa, Email: fpedrosa@ufpr.br.
Maria BR Steffens, Email: steffens@ufpr.br.
Leda S Chubatsu, Email: chubatsu@ufpr.br.
Acknowledgements
We are grateful to the GENOPAR consortium for providing plasmids, and to Roseli Prado, Julieta Pie and Valter Baura for technical assistance. We are also grateful to Dr. Geoffrey Yates for reading the manuscript. This work was supported by INCT-FBN/CNPq/MCT, Institutos do Milênio, PRONEX, CAPES, CNPq and Fundação Araucária.
References
- Arcondeguy T, Jack R, Merrick M. P(II) signal transduction proteins, pivotal players in microbial nitrogen control. Microbiol Mol Biol Rev. 2001;65(1):80–105. doi: 10.1128/MMBR.65.1.80-105.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forchhammer K. P-II signal transducers: novel functional and structural insights. Trends Microbiol. 2008;16(2):65–72. doi: 10.1016/j.tim.2007.11.004. [DOI] [PubMed] [Google Scholar]
- Jiang P, Ninfa AJ. Escherichia coli PII signal transduction protein controlling nitrogen assimilation acts as a sensor of adenylate energy charge in vitro. Biochemistry. 2007;46(45):12979–12996. doi: 10.1021/bi701062t. [DOI] [PubMed] [Google Scholar]
- He LH, Soupene E, Ninfa A, Kustu S. Physiological role for the GlnK protein of enteric bacteria: Relief of NifL inhibition under nitrogen-limiting conditions. J Bacteriol. 1998;180(24):6661–6667. doi: 10.1128/jb.180.24.6661-6667.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jack R, De Zamaroczy M, Merrick M. The signal transduction protein GlnK is required for NifL-dependent nitrogen control of nif gene expression in Klebsiella pneumoniae. J Bacteriol. 1999;181(4):1156–1162. doi: 10.1128/jb.181.4.1156-1162.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Little R, Reyes-Ramirez F, Zhang Y, van Heeswijk WC, Dixon R. Signal transduction to the Azotobacter vinelandii NIFL-NIFA regulatory system is influenced directly by interaction with 2-oxoglutarate and the PII regulatory protein. Embo J. 2000;19(22):6041–6050. doi: 10.1093/emboj/19.22.6041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arsene F, Kaminski PA, Elmerich C. Modulation of NifA activity by PII in Azospirillum brasilense: evidence for a regulatory role of the NifA N-terminal domain. J Bacteriol. 1996;178(16):4830–4838. doi: 10.1128/jb.178.16.4830-4838.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Araujo LA, Monteiro RA, Souza EM, Steffens MBR, Rigo LU, Pedrosa FO, Chubatsu LS. GlnB is specifically required for Azospirillum brasilense NifA activity in Escherichia coli. Res Microbiol. 2004;155(6):491–495. doi: 10.1016/j.resmic.2004.03.002. [DOI] [PubMed] [Google Scholar]
- Zhang YP, Pohlmann EL, Roberts GP. Identification of critical residues in GlnB for its activation of NifA activity in the photosynthetic bacterium Rhodospirillum rubrum. Proc Natl Acad Sci USA. 2004;101(9):2782–2787. doi: 10.1073/pnas.0306763101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drepper T, Gross S, Yakunin AF, Hallenbeck PC, Masepohl B, Klipp W. Role of GlnB and GlnK in ammonium control of both nitrogenase systems in the phototrophic bacterium Rhodobacter capsulatus. Microbiology-Sgm. 2003;149:2203–2212. doi: 10.1099/mic.0.26235-0. [DOI] [PubMed] [Google Scholar]
- Zhang YP, Wolfe DM, Pohlmann EL, Conrad MC, Roberts GP. Effect of AmtB homologues on the post-translational regulation of nitrogenase activity in response to ammonium and energy signals in Rhodospirillum rubrum. Microbiology-Sgm. 2006;152:2075–2089. doi: 10.1099/mic.0.28903-0. [DOI] [PubMed] [Google Scholar]
- Huergo LF, Merrick M, Monteiro RA, Chubatsu LS, Steffens MBR, Pedrosa FO, Souza EM. In Vitro Interactions between the P-II Proteins and the Nitrogenase Regulatory Enzymes Dinitrogenase Reductase ADP-ribosyltransferase (DraT) and Dinitrogenase Reductase-activating Glycohydrolase (DraG) in Azospirillum brasilense. J Biol Chem. 2009;284(11):6674–6682. doi: 10.1074/jbc.M807378200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baldani JI, Baldani VLD, Seldin L, Dobereiner J. Characterization of Herbaspirillum seropedicae Gen-Nov, Sp-Nov, a Root-Associated Nitrogen-Fixing Bacterium. Int J Syst Bacteriol. 1986;36(1):86–93. doi: 10.1099/00207713-36-1-86. [DOI] [Google Scholar]
- Benelli EM, Souza EM, Funayama S, Rigo LU, Pedrosa FO. Evidence for two possible glnB-type genes in Herbaspirillum seropedicae. J Bacteriol. 1997;179(14):4623–4626. doi: 10.1128/jb.179.14.4623-4626.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noindorf L, Rego FGM, Baura VA, Monteiro RA, Wassem R, Cruz LM, Rigo LU, Souza EM, Steffens MBR, Pedrosa FO. et al. Characterization of the orf1glnKamtB operon of Herbaspirillum seropedicae. Arch Microbiol. 2006;185(1):55–62. doi: 10.1007/s00203-005-0066-4. [DOI] [PubMed] [Google Scholar]
- Huergo LF, Noindorf L, Gimenes C, Lemgruber RSP, Cordellini DF, Falarz LJ, Cruz LM, Monteiro RA, Pedrosa FO, Chubatsu LS. et al. Proteomic analysis of Herbaspirillum seropedicae reveals ammonium-induced AmtB-dependent membrane sequestration of P-II proteins. FEMS Microbiol Lett. 2010;308(1):40–47. doi: 10.1111/j.1574-6968.2010.01986.x. [DOI] [PubMed] [Google Scholar]
- Bonatto AC, Couto GH, Souza EM, Araujo LM, Pedrosa FO, Noindorf L, Benelli EM. Purification and characterization of the bifunctional uridylyltransferase and the signal transducing proteins GlnB and GlnK from Herbaspirillum seropedicae. Protein Expr Purif. 2007;55:293–299. doi: 10.1016/j.pep.2007.04.012. [DOI] [PubMed] [Google Scholar]
- Persuhn DC, Souza EM, Steffens MB, Pedrosa FO, Yates MG, Rigo LU. The transcriptional activator NtrC controls the expression and activity of glutamine synthetase in Herbaspirillum seropedicae. FEMS Microbiol Lett. 2000;192(2):217–221. doi: 10.1111/j.1574-6968.2000.tb09385.x. [DOI] [PubMed] [Google Scholar]
- Atkinson MR, Ninfa AJ. Role of the GlnK signal transduction protein in the regulation of nitrogen assimilation in Escherichia coli. Mol Microbiol. 1998;29(2):431–447. doi: 10.1046/j.1365-2958.1998.00932.x. [DOI] [PubMed] [Google Scholar]
- Wassem R, Pedrosa FO, Yates MG, Rego FG, Chubatsu LS, Rigo LU, Souza EM. Control of autogenous activation of Herbaspirillum seropedicae nifA promoter by the IHF protein. FEMS Microbiol Lett. 2002;212(2):177–182. doi: 10.1111/j.1574-6968.2002.tb11263.x. [DOI] [PubMed] [Google Scholar]
- Rego FG, Pedrosa FO, Chubatsu LS, Yates MG, Wassem R, Steffens MB, Rigo LU, Souza EM. The expression of nifB gene from Herbaspirillum seropedicae is dependent upon the NifA and RpoN proteins. Can J Microbiol. 2006;52(12):1199–1207. doi: 10.1139/W06-085. [DOI] [PubMed] [Google Scholar]
- Souza EM, Pedrosa FO, Drummond M, Rigo LU, Yates MG. Control of Herbaspirillum seropedicae NifA activity by ammonium ions and oxygen. J Bacteriol. 1999;181(2):681–684. doi: 10.1128/jb.181.2.681-684.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monteiro RA, Souza EM, Funayama S, Yates MG, Pedrosa FO, Chubatsu LS. Expression and functional analysis of an N-truncated NifA protein of Herbaspirillum seropedicae. FEBS Lett. 1999;447(2-3):283–286. doi: 10.1016/S0014-5793(99)00314-2. [DOI] [PubMed] [Google Scholar]
- Wang H, Franke CC, Nordlund S, Noren A. Reversible membrane association of dinitrogenase reductase activating glycohydrolase in the regulation of nitrogenase activity in Rhodospirillum rubrum; dependence on GlnJ and AmtB1. FEMS Microbiol Lett. 2005;253(2):273–279. doi: 10.1016/j.femsle.2005.09.049. [DOI] [PubMed] [Google Scholar]
- Tremblay PL, Hallenbeck PC. Ammonia-induced formation of an AmtB-GlnK complex is not sufficient for nitrogenase regulation in the photosynthetic bacterium Rhodobacter capsulatus. J Bacteriol. 2008;190(5):1588–1594. doi: 10.1128/JB.01643-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodsworth JA, Leigh JA. Regulation of nitrogenase by 2-oxoglutarate-reversible, direct binding of a PII-like nitrogen sensor protein to dinitrogenase. Proc Natl Acad Sci USA. 2006;103(26):9779–9784. doi: 10.1073/pnas.0602278103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu H, Burris RH. Ammonium Inhibition of Nitrogenase Activity in Herbaspirillum seropedicae. J Bacteriol. 1989;171(6):3168–3175. doi: 10.1128/jb.171.6.3168-3175.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klassen G, Pedrosa FO, Souza EM, Funayama S, Rigo LU. Effect of nitrogen compounds on nitrogenase activity in Herbaspirillum seropedicae SMR1. Can J Microbiol. 1997;43(9):887–891. doi: 10.1139/m97-129. [DOI] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning - a laboratory manual. second. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- Pedrosa FO, Yates MG. Regulation of Nitrogen-Fixation (nif) Genes of Azospirillum brasilense by NifA and Ntr (Gln) Type Gene-Products. FEMS Microbiol Lett. 1984;23(1):95–101. doi: 10.1111/j.1574-6968.1984.tb01042.x. [DOI] [Google Scholar]
- Miller JH. Experiments in Molecular Genetics. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press; 1972. [Google Scholar]
- Bradford MM. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Dilworth MJ. Acetylene Reduction by Nitrogen-Fixing Preparations from Clostridium Pasteurianum. Biochim Biophys Acta. 1966;127(2):285–294. doi: 10.1016/0304-4165(66)90383-7. [DOI] [PubMed] [Google Scholar]
- Schöllhorn R, Burris RH. Acetylene as a Competitive Inhibitor of N2 Fixation. Proc Natl Acad Sci USA. 1967;58(1):213–216. doi: 10.1073/pnas.58.1.213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hynes MF, Quandt J, Oconnell MP, Puhler A. Direct Selection for Curing and Deletion of Rhizobium Plasmids Using Transposons Carrying the Bacillus subtilis sacB Gene. Gene. 1989;78(1):111–120. doi: 10.1016/0378-1119(89)90319-3. [DOI] [PubMed] [Google Scholar]
- Souza EM, Funayama S, Rigo LU, Yates MG, Pedrosa FO. Sequence and Structural Organization of a nifA-Like Gene and Part of a nifB-Like Gene of Herbaspirillum seropedicae Strain Z78. J Gen Microbiol. 1991;137:1511–1522. doi: 10.1099/00221287-137-7-1511. [DOI] [PubMed] [Google Scholar]
- Kleiner D, Paul W, Merrick MJ. Construction of Multicopy Expression Vectors for Regulated over-Production of Proteins in Klebsiella pneumoniae and Other Enteric Bacteria. J Gen Microbiol. 1988;134:1779–1784. doi: 10.1099/00221287-134-7-1779. [DOI] [PubMed] [Google Scholar]
- Souza EM, Pedrosa FO, Rigo LU, Machado HB, Yates MG. Expression of the nifA gene of Herbaspirillum seropedicae: role of the NtrC and NifA binding sites and of the-24/-12 promoter element. Microbiology-Sgm. 2000;146:1407–1418. doi: 10.1099/00221287-146-6-1407. [DOI] [PubMed] [Google Scholar]
- Simon R, Priefer U, Puhler A. A Broad Host Range Mobilization System for Invivo Genetic-Engineering - Transposon Mutagenesis in Gram-Negative Bacteria. Bio-Technology. 1983;1(9):784–791. [Google Scholar]
- Souza EM, Funayama S, Rigo LU, Pedrosa FO. Cloning and Characterization of the nifA gene from Herbaspirillum seropedicae Strain Z78. Can J Microbiol. 1991;37(6):425–429. doi: 10.1139/m91-069. [DOI] [PubMed] [Google Scholar]
- Woodley P, Buck M, Kennedy C. Identification of sequences important for recognition of vnf genes by the VnfA transcriptional activator in Azotobacter vinelandii. FEMS Microbiol Lett. 1996;135(2-3):213–221. doi: 10.1111/j.1574-6968.1996.tb07992.x. [DOI] [PubMed] [Google Scholar]
- Mead DA, Szczesna-Skorupa E, Kemper B. Single-stranded DNA 'blue' T7 promoter plasmids: a versatile tandem promoter system for cloning and protein engineering. Protein Eng. 1986;1(1):67–74. doi: 10.1093/protein/1.1.67. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Immunoblot analysis of H. seropedicae PII proteins.