Abstract
There are many examples within gene complexes of transcriptional enhancers interacting with only a subset of target promoters. A number of molecular mechanisms including promoter competition, insulators and chromatin looping are thought to play a role in regulating these interactions. At the Drosophila bithorax complex (BX-C), the IAB5 enhancer specifically drives gene expression only from the Abdominal-B (Abd-B) promoter, even though the enhancer and promoter are 55 kb apart and are separated by at least three insulators. In previous studies, we discovered that a 255 bp cis-regulatory module, the promoter tethering element (PTE), located 5′ of the Abd-B transcriptional start site is able to tether IAB5 to the Abd-B promoter in transgenic embryo assays. In this study we examine the functional role of the PTE at the endogenous BX-C using transposon-mediated mutagenesis. Disruption of the PTE by P element insertion results in a loss of enhancer-directed Abd-B expression during embryonic development and a homeotic transformation of abdominal segments. A partial deletion of the PTE and neighboring upstream genomic sequences by imprecise excision of the P element also results in a similar loss of Abd-B expression in embryos. These results demonstrate that the PTE is an essential component of the regulatory network at the BX-C and is required in vivo to mediate specific long-range enhancer-promoter interactions.
Introduction
To ensure a high fidelity of gene expression patterns in embryos a very strict functional requirement exists for the interaction of cis-regulatory modules (CRMs) in the genome of animals during development [1], [2], [3], [4]. Embryonic transcriptional enhancers, which direct specific spatio-temporal patterns of gene expression in the embryo, are not permitted to promiscuously activate transcription from non-target promoters. Chromatin structural organization, insulators and promoter competition are thought to play a role in regulating enhancer-promoter interactions ([5], [6], [7], [8], [9], for recent reviews see [10], [11], [12]). At the Drosophila bithorax complex (BX-C), an extensive network of CRMs located in over 300 kb of infraabdominal (iab) non-genic sequence is responsible for directing embryonic expression of just three homeotic genes; Ultrabithorax (Ubx), abdominal-A (abd-A) and Abdominal-B (Abd-B) (Fig. 1) [13], [14]. These three homeotic genes are critical for establishing cellular identities in the presumptive thoracic and abdominal segments during development [2], [15].
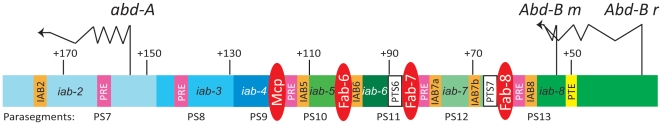
Figure 1. A network of interacting cis-regulatory modules control embryonic expression of abdominal-A and Abdominal-B in the Drosophila bithorax complex.
The abd-A and Abd-B morphogenetic (m) and regulatory (r) transcripts are indicated by leftward arrows. The regulatory regions iab-2, iab-3 and iab-4 (blue) interact with abd-A. The iab-5, iab-6, iab-7 and iab-8 regions (green) interact with Abd-B m. Each iab region is thought to contain at least one characterized enhancer (orange rectangles) capable of directing Hox gene expression in a specific embryonic parasegments (PS). The positions of the characterized Fab-6, Fab-7, Fab-8 and Mcp insulators (red ellipses), promoter targeting sequence (PTS) modules (white rectangles), promoter tethering element (PTE) (yellow rectangle) and polycomb response elements (PREs) (pink rectangles) are indicated. Numbers above line refer to kilobase positions in DNA sequence accession number: DM31961.
We recently identified a 255 bp promoter tethering element (PTE) located 5′ of the Abd-B transcriptional start site that is responsible for specifically recruiting the IAB5 enhancer from the BX-C to the Abd-B promoter in competition assays on transgenes [16]. Furthermore, the PTE also demonstrates anti-insulator activity by enabling the IAB5 enhancer to bypass an insulator from the BX-C to activate a target promoter on transgenes [17]. The ability of the PTE to facilitate specific enhancer-promoter interactions may explain how certain IAB enhancers, such as IAB5, IAB6, IAB7a and IAB7b [8], [18], [19], are able to bypass the Frontabdominal (Fab) insulators Fab-6, Fab-7 and Fab-8 [19], [20], [21], [22] and direct transcription from the Abd-B morphogenetic (m) promoter at the endogenous BX-C (Fig. 1). As such, the PTE may be part of the complex regulatory network of CRMs, including the insulators, Promoter Targeting Sequences (PTSs) [23], [24] and additional tethering sequences located 5′ of the Abd-B gene [25], [26], responsible for mediating enhancer-promoter interactions in the BX-C. In contrast to the PTSs, which are located adjacent to the Fab-7 and Fab-8 insulators, the PTE is adjacent to the Abd-B promoter (Fig. 1). The PTSs have been implicated in mediating insulator bypass for the IAB enhancers on transgenic constructs [23], [24]. However, deletion of either of the characterized PTSs from the endogenous BX-C did not result in a significant phenotype [8], [18], [19], suggesting that there might be other sequences capable of mediating the long-range enhancer-promoter specificity at the BX-C.
In this study, we demonstrate the functional importance of the PTE at the endogenous BX-C. Disruption of the PTE by a P element insertion in vivo results in a loss of IAB enhancer-directed Abd-B expression during development and a homeotic phenotype when placed in hemizygosity with an Abd-B null allele. Further molecular analysis of a mutant generated by imprecise excision of the P element reveals that a partial deletion of the PTE and neighboring upstream genomic sequences results in a similar loss of enhancer-directed Abd-B expression. These results demonstrate the critical functional role that the PTE plays in regulating promoter-enhancer interactions at the endogenous Drosophila BX-C.
Results
Insertions at the PTE disrupt Abd-B expression and result in homeotic transformation
Genetic studies were carried out to determine whether the activity of the PTE is important for in vivo IAB enhancer–Abd-B interactions in the context of the BX-C. The previously identified Abd-BT2N mutation [27] is derived from a P element insertion in the endogenous PTE sequence (−241 bp relative the Abd-B transcription start site). The original P element insertion line (Abd-BLac1) contains a 5.6 kb Abd-B promoter region which includes 4.3 kb of 5′ sequence harboring the PTE sequence, fused to a lacZ reporter gene. This P element also contains the Fab-8 insulator element, an IAB8 enhancer element and the rosy gene. In contrast, the derived Abd-BT2N line contains only a truncated Abd-B promoter-lacZ reporter gene, generated by imprecise excision of the 5′ region of the P element (Fig. 2a). In the Abd-BT2N line the P element is inserted within the endogenous 255 bp PTE sequence, while leaving the entire Abd-B morphogenetic (m) transcription unit intact [27]. The Abd-BT2N mutant is described as a Class I Abd-B mutant, capable of complementing mutations in the Abd-B regulatory (r) transcript, but affecting Abd-B m transcript expression [27]. In hemizygosis with the Abd-BM1 null allele, the Abd-BT2N insertion line shows a phenotypic transformation of abdominal segments 5–8 towards a more anterior abdominal segment identity. This is seen most clearly in the complete transformation of the seventh abdominal tergite to a more anterior identity in the Abd-BT2N insertion line (Fig. 2c) when compared to wild-type (WT) (Fig. 2b). The homeotic transformation is easiest to see in the cuticles of adult females due to their lighter abdominal pigmentation when compared to males (Fig. 2b and c), although it is apparent in both sexes. This phenotype is characteristic of Abd-B m mutations [27], [28], suggesting that the P element insertion into the PTE is disrupting expression from the adjacent Abd-B m promoter and is consistent with a reduction in IAB5-7 enhancer-Abd-B interactions.
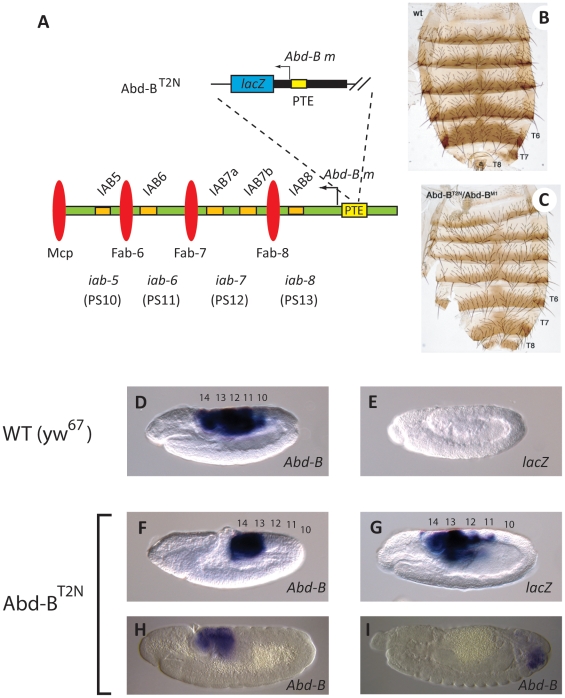
Figure 2. P element insertion into the endogenous PTE results in homeotic transformation and loss of Abd-B expression.
(A) Schematic diagram of the Abd-BT2N P element line, showing the insertion site −241 bp 5′ of the Abd-B m transcription start site in the endogenous PTE sequence. The same symbols and color scheme as in Fig. 1 indicate the neighboring cis-regulatory modules. (B) Dorsal cuticles were prepared from adult females. A normal abdominal pigmentation pattern is observed in wild-type (WT) adults. (C) In Abd-BT2N/Abd-BM1 hemizygotes the seventh abdominal tergite (T7) is fully developed, indicating a transformation toward a more anterior abdominal segment identity. The eighth abdominal tergite (T8) also shows a partial transformation toward a more anterior identity. Abd-BM1 is a Class III null allele for both the Abd-B m and r transcripts [15], [27]. Abd-B gene expression pattern in wild-type (WT) (D) and Abd-BT2N (F) germ-band elongation stage 9 embryos. The staining patterns for the Abd-BT2N line show reduced Abd-B expression in parasegments (PS) 10, 11 and 12 at stage 9 (F), stage 11 (H) and stage 13 (I) of development. The lacZ reporter gene is expressed in developing posterior regions of the Abd-BT2N embryos (G). Expression is strongest in PS13, although the pattern extends from PS 10–14 and is very similar to the endogenous Abd-B transcription pattern. No lacZ expression is detectable in WT embryos (E).
In WT D. melanogaster embryos, the expression pattern of the Abd-B m transcript extends from parasegment (PS) 10–13 during the germ-band elongation stage of development, while the r transcript is predominantly restricted to PS14 at this stage [29], [30]. In situ hybridization with an RNA probe that detects both the m and r transcripts in germ-band elongation stage embryos generated from crosses of heterozygous balanced Abd-BT2N adults (Fig. 2f) [27], demonstrates a loss of Abd-B expression in PS10–12 (presumptive abdominal segments 5–7) when compared with expression in WT embryos (Fig. 2d). These results indicate that the transposon insertion in the PTE impairs the ability of the IAB5–7 enhancers in the BX-C to drive expression from the endogenous Abd-B m promoter. The detectable expression of Abd-B in PS13 in Abd-BT2N embryos raises the question of why there is a homeotic transformation of the eighth abdominal segment in hemizygous Abd-BT2N/Abd-BM1 adults. While this phenotype is significant, it is only a partial transformation of A8 towards a more anterior identity (Fig. 2c), suggesting that there is not a complete loss of Abd-B patterning function. It is possible that there may be subtle modulation of the level of Abd-B gene expression in PS13 which, while responsible for the phenotypic transformation, is not readily detectable using RNA in situ hybridization. The expression of Abd-B persists in PS13 and 14 at least through stage 13 of development (Fig. 2h and i). However, it is also possible that transcription of Abd-B in late embryonic or larval stages may be modulated from the Abd-BT2N allele.
Knowing that expression from the endogenous Abd-B m promoter was perturbed in Abd-BT2N mutants, we examined whether the enhancers from the BX-C were re-directed to the intact ectopic Abd-B m promoter (containing the PTE sequence), which drives lacZ, on the P element insertion. In Abd-BT2N embryos, lacZ expression was detected in a pattern extending from PS10 to 14 (Fig. 2g). The lacZ expression appears strongest in PS13 (presumptive abdominal segment 8) and weaker in PS10 (Fig. 2g), presumably due to the proximity of the endogenous IAB8 regulatory sequences to the target promoter, as previously observed [27]. This pattern of lacZ expression is consistent with the Abd-B expression pattern detected in wild-type embryos (Fig. 2d) and indicates that the IAB enhancers from the endogenous BX-C are now being re-directed to the intact ectopic Abd-B promoter (containing the PTE) on the P element to drive expression of lacZ in Abd-BT2N embryos.
Deletion of PTE and neighboring upstream sequences results in a loss of enhancer-directed Abd-B transcription
In the Abd-BT2N mutant endogenous PTE function is disrupted by P element insertion. In order to address the loss-of-functional activity of the PTE more directly we generated a mutant line in which the 5′ portion of the PTE and neighboring upstream genomic sequences are deleted from the endogenous BX-C locus (Fig. 3). This was accomplished by performing an imprecise excision of the P element insertion from the Abd-BLDN mutant (Fig. S1). The Abd-BLDN mutant was generated by a P element replacement strategy and carries an insertion containing the GAL4 and white genes in the identical location within the PTE (−241 bp relative to the Abd-B transcription start site) as the Abd-BT2N line, but does not contain an ectopic copy of the PTE (Fig. 3a) [31]. In the Abd-BLDN line, both the white and GAL4 genes are only expressed clearly in PS13 and very weakly in PS14 in embryos (data not shown), indicating that the distantly located IAB5–7 enhancers in the BX-C are not activating transcription of either reporter gene. The Abd-BLDN line was utilized to generate an imprecise excision mutant as it carries the readily detectable white reporter gene (Fig. 3a). As a result, adult flies in which P element excision had occurred were easily identified by loss of red eye color (Fig. S1). PCR-based screening with primers from the proximal Abd-B m promoter region and from 0.5 kb, 1 kb and 1.5 kb 5′ of the Abd-B transcription start site was used to identify a 1.2 kb deletion in the Abd-B ΔPTE-UP allele (Fig. 3a–c). The molecular nature of the deletion was characterized by sequencing and found to have removed 53 bp of the PTE sequence 5′ of the original P element insertion site, as well as 1134 bp of endogenous genomic sequence 5′ of the defined PTE. In this Abd-B ΔPTE-UP allele, a portion of the GAL4 gene from the original P element remains, along with 202 bp of the 3′ PTE sequence (Fig. 3b).
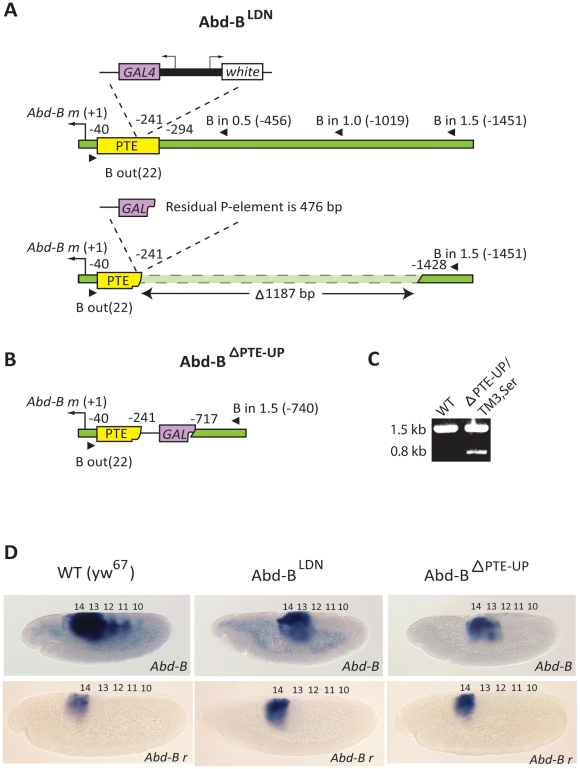
Figure 3. Deletion of the PTE and upstream sequences at the endogenous BX-C disrupts Abd-B expression.
(A) The Abd-BLDN mutant features a P element inserted within the PTE (yellow box) (−241 bp relative to the Abd-B transcription start site (black arrow)). The P element contains GAL4 (purple box) and white (white box) reporter genes. PCR primers (black triangles) designed against the Abd-B promoter region and 0.5 kb, 1 kb and 1.5 kb upstream of the Abd-B m transcription start site were used to characterize the approximately 1.2 kb deletion (dashed box) in the Abd-B ΔPTE-UP mutant generated by imprecise excision of the P element. (B) The resulting Abd-B ΔPTE-UP deletion removes 53 bp of the PTE sequence 5′ of the original insertion site and an additional 1134 bp of endogenous sequence 5′ of the PTE. A 476 bp portion of the GAL4 coding sequence (purple polygon) from the original P element remains, as well as 202 bp of the 3′ PTE sequence (yellow polygon). (C) Primers designed against the Abd-B promoter region and 1.5 kb upstream of the Abd-B transcription start site were used to genotype adult flies carrying the deletion allele by amplifying the 1.5 kb wild-type (WT) band and the 0.8 kb Abd-B ΔPTE-UP band in balanced Abd-B ΔPTE-UP (ΔPTE-UP/TM3,Ser) individuals. (D) In situ hybridization with an RNA probe that can detect both the Abd-B m and r transcripts (Abd-B) shows WT expression in parasegments (PS) 10–14 in yw67 embryos, while expression is found only in PS13–14 in a portion of embryos collected from crosses of balanced Abd-B ΔPTE-UP and Abd-B LDN lines. In situ hybridization with a probe specifically designed against the Abd-B r transcript (Abd-B r) detects identical patterns of expression in PS14 of germ-band elongation stage embryos collected from WT, mutant Abd-BLDN and Abd-BΔPTE-UP balanced lines.
Based on the Abd-B expression observed in the Abd-BT2N insertion line, we hypothesized that the expression pattern of the endogenous Abd-B gene from the Abd-B ΔPTE-UP allele may also be perturbed. In situ hybridization with an RNA probe that can detect both the m and r transcripts shows that Abd-B gene expression is lost specifically in PS10-12, but not in PS13–14 in germ-band elongation stage embryos generated from crosses of heterozygous balanced Abd-B ΔPTE-UP adults (Fig. 3d). Statistical analysis demonstrates that the number of embryos demonstrating this restricted Abd-B expression pattern from both the Abd-B ΔPTE-UP and Abd-BLDN alleles is highly significant (p<0.01) when compared to the number of embryos with the WT Abd-B expression pattern. This restricted pattern of Abd-B expression persists at least through stage 13 of development (data not shown). One possible explanation for the loss of Abd-B expression in PS10–12 is that the genomic region deleted in the Abd-B ΔPTE-UP allele harbors a CRM capable of driving transcription in these specific parasegments. However, when tested in a transgenic reporter gene assay the 1.2 kb region does not exhibit embryonic enhancer activity (data not shown).
To confirm that mutations in the PTE only affect expression from the Abd-B m promoter, but not the Abd-B r promoter, in situ hybridization with a RNA probe (BPP, [32]) that specifically detects only the Abd-B r transcript was performed on embryos collected from WT, Abd-BLDN and Abd-B ΔPTE-UP balanced lines. The expression pattern of the Abd-B r transcript was confirmed to be identical in the WT, Abd-BLDN and Abd-B ΔPTE-UP embryos, appearing only in PS14 of germ-band elongation stage embryos (Fig. 3d). These observations are consistent with the known pattern of expression from the Abd-B r promoter [29] and confirm that the disruption to the PTE in these mutants only affects enhancer-mediated transcription from the Abd-B m promoter (see discussion for more detail).
Discussion
Parasegment-specific interactions between the IAB enhancers and Abd-B promoter in the BX-C
The absence of Abd-B expression in PS10–12 of Abd-BΔPTE-UP germ-band elongation stage embryos is consistent with a loss of IAB-enhancer directed expression from the Abd-B m promoter [29]. This suggests that deletion of the Abd-B promoter tethering sequence (PTE) and the neighboring 1.1 kb 5′ sequence in the Abd-BΔPTE-UP line leads to a disruption of the long-range interactions between the Abd-B m promoter and enhancers from the iab-5, iab-6 and iab-7 regions in PS10, 11 and 12, respectively. For example, in PS12 of WT embryos the tethering sequences upstream of the Abd-B m promoter enable the IAB7a and IAB7b embryonic enhancers to bypass the Fab-8 chromatin insulator and drive expression from the Abd-B m promoter (Fig. 4a). In Abd-BΔPTE-UP mutant embryos, removal of the tethering sequences appears to disrupt the ability of the IAB7a and IAB7b enhancers to activate the Abd-B m promoter, resulting in an absence of Abd-B expression in PS12 (Fig. 4a and 3d). Similarly, the IAB6 and IAB5 enhancers are unable to bypass intervening insulators to activate Abd-B m expression in PS11 and PS10 in Abd-BΔPTE-UP embryos. In contrast to PS10–12, the specific enhancer-promoter interactions at the BX-C in PS13 appear to be intact in Abd-BΔPTE-UP embryos. In WT embryos the IAB8 embryonic enhancer, located 3′ of the Abd-B gene, is solely responsible for directing expression from the Abd-B m promoter in PS13 [33]. In Abd-BΔPTE-UP mutant embryos the Abd-B m transcript remains strongly expressed in PS13 (Fig. 3d). This result indicates that the interaction between the IAB8 enhancer and the Abd-B m promoter may not require tethering activity, likely due to the physical proximity of the enhancer to the promoter and lack of an intervening chromatin insulator (Fig. 4a). However, given the partial transformation phenotype observed for PS13 in flies carrying a disruption of the PTE sequences (as in the case of the Abd-BT2N allele, Fig. 2c) it remains possible that loss of PTE activity may be responsible for subtle effects on transcription of Abd-B in PS13. In PS14, Abd-B expression in germ-band elongation stage embryos is not driven by the 3′ IAB enhancers, as it is initiated from the r transcriptional start site located 5′ of the PTE sequence (Fig. 1) [29], [34]. Consequently, expression of the r transcript in PS14 is not perturbed by the loss of tethering activity in Abd-BΔPTE-UP mutant embryos (Fig. 4a and 3d).
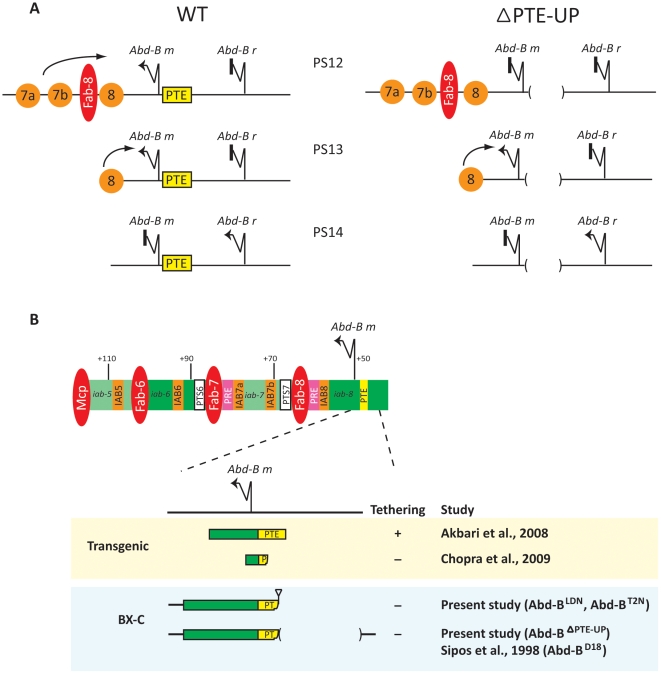
Figure 4. PTE sequence is required for specific promoter-enhancer tethering at the Abd-B gene.
(A) Abd-B transcription is disrupted in PTE mutants. In wild-type (WT) germ-band elongation stage embryos expression from the Abd-B m promoter is observed in parasegments (PS) 10–13 and from the Abd-B r promoter in PS14 [29]. The loss of Abd-B expression in PS10–12 in Abd-BΔPTE-UP (ΔPTE-UP) embryos is consistent with a disruption of expression from the Abd-B m promoter. In PS12 of WT embryos, the tethering sequences upstream of the Abd-B m promoter enable the IAB7a and IAB7b enhancers (7a and 7b, orange circles) to act over the long-range of intervening sequence and bypass the Fab-8 chromatin insulator (red ellipse) and drive expression from the Abd-B m promoter. In PS12 of Abd-BΔPTE-UP mutants the removal of the tethering sequences disrupts the ability of the IAB7 enhancers to activate the Abd-B m promoter across the intervening Fab-8 insulator. A similar disruption of promoter-enhancer tethering activity at Abd-BΔPTE-UP is also seen in PS11 (IAB6 enhancer) and PS10 (IAB5 enhancer). In PS13 of both WT and Abd-BΔPTE-UP embryos, IAB8 (8, orange circle) is able, due to its physical proximity and lack of intervening chromatin insulator, to drive expression from the Abd-B m promoter. In PS14 of both WT and Abd-BΔPTE-UP embryos, Abd-B r is transcribed independently of IAB embryonic enhancer activity and represses transcription from the Abd-B m promoter [28]. (B) Functional dissection of critical Abd-B tethering sequences. Schematic diagram of the regulatory region of the Abdominal-B gene shows enhancers (orange boxes) in the infraabdominal regions iab-5 to iab-8 (green) that are responsible for activating expression of the Abd-B gene in abdominal PS 10–13 during embryogenesis. Color scheme for other types of cis-regulatory modules is same as in Figure 1. Transgenic studies (highlighted in yellow box) show that a 1.4 kb of sequence from +1228 to −294, (including the 255 bp PTE sequence located 40 bp 5′ of the Abd-B m transcription start site), is sufficient for promoter-enhancer tethering [16]. Additional transgenic studies have shown that an approximately 200 bp sequence spanning the Abd-B m transcription start site (+100 to −100) is not sufficient for tethering activity [36]. P element insertion at the endogenous BX-C (as in the Abd-BLDN or Abd-BT2N mutations) demonstrates that the 242 bp sequence 5′ of the Abd-B m transcription site (from +1 to −241), including 202 bp of the identified PTE sequence, is not sufficient to tether the distally located IAB5, 6 and 7 enhancers in vivo. Deletion of an approximately 1.2 kb sequence (−241 to −1428) which include 53 bp of the 5′ region of the PTE and neighboring upstream sequences in the Abd-B ΔPTE-UP mutant also results in a loss of tethering. The 53 bp region of the PTE may therefore be important for functional tethering. A potential tethering role at the endogenous BX-C for the sequences 5′ of the PTE is also suggested by earlier genetic complementation studies in the Abd-BD18 allele [25].
One possible outcome of the disruption of IAB enhancer interactions with the Abd-B m promoter at the Abd-BΔPTE-UP allele may be the re-direction of those enhancers to the neighboring abd-A promoter in the BX-C. However, no change in the expression pattern of abd-A (extending from PS7 to PS13 in germ-band elongation stage embryos [35] could be detected from the Abd-BΔPTE-UP or Abd-B LDN alleles (data not shown).
Functional dissection of critical sequences at the PTE
The functional activity of the PTE at the endogenous BX-C prompts the question of exactly what sequences associated with the PTE are necessary to confer promoter-enhancer tethering. Our earlier studies demonstrated that a 255 bp region located between −40 and −294 bp 5′ of the Abd-B m transcription start site is sufficient to tether the IAB5 enhancer to an ectopic promoter in a transgene competition assay (Fig. 4b) [16]. In contrast, an approximately 200 bp region extending from −100 to +100 relative to the Abd-B m transcriptional start site is not sufficient to mediate promoter-enhancer tethering when tested in similar transgenic assays [36]. In agreement with this observation, the 202 bp of sequence from the 3′ end of the PTE remaining at the endogenous BX-C in the Abd-BΔPTE-UP allele is also not sufficient to mediate tethering of the IAB enhancers to the Abd-B m promoter (Fig. 4b). A previous study by Sipos and colleagues, utilizing an Abd-B mutant allele harboring a small deletion at the Abd-B m promoter region and a deletion at the iab-7 region carried on opposite chromosomes resulted in Abd-B transcriptional activity [25]. This indicates that a trans interaction can occur between the iab-7 regulatory region and the remaining Abd-B promoter region across chromosomes. The same genetic complementation test using the iab-7 mutant allele and the Abd-BD18 deletion allele (which removes approximately 8 kb of sequence 5′ of the Abd-B m transcription start site) on opposite chromosomes showes an increase in trans-mediated Abd-B activity [25]. These results indicate that additional sequences located 5′ of the 255 bp PTE may also contribute to tethering activity. A functional role for the sequences in the 5′ end of the PTE and neighboring upstream genomic regions is supported by the loss of enhancer-directed Abd-B m transcript expression at the Abd-BΔPTE-UP allele after deletion of the 5′ 53 bp region in the PTE and 1134 bp of sequence located 5′ of the defined PTE (Fig. 4b). The loss of Abd-B expression observed in the Abd-BΔPTE-UP mutant is consistent with the previously reported molecular phenotype of Abd-B m mutants [27] and confirms the role of the PTE and associated 5′ sequences in mediating specific IAB enhancer-directed expression of the Abd-B m transcript.
Promoter tethering as a general regulatory mechanism
Additional examples of genetic complexes in which long-range interactions between CRMs have been characterized include the human beta-globin locus [37], the Drosophila Antennapedia complex [38], [39] and mouse HoxD complex [3]. In the large genomes of vertebrates, where extensive global control regions have been identified, promoter-enhancer tethering is emerging as a critical general mechanism for regulation of gene expression [3], [37]. More recently, in Drosophila, other PTE sequences have been located in the even-skipped [40] and engrailed [41] loci that mediate enhancer-promoter communication. The long-range CRM communication in these critical developmental genes has been shown to be dynamic, changing through the course of development in some cases [40] and perhaps even capable of mediating the evolution of novel patterns of gene expression in different insect species [42].
Our current model for the molecular function of the PTE in the Drosophila BX-C is that regulatory interactions that enable the PTE to tether the IAB enhancers to the Abd-B m promoter in specific parasegments during embryonic development may be mediated by chromatin looping [17]. A number of studies have indicated that chromatin looping may facilitate promoter-enhancer tethering through the action of different transcription factors. For example, the sea urchin GCF1 protein is able to form higher order multimeric loop structures when added to target site oligonucleotides in vitro [43]. The abundant mammalian transcription factor Sp1 has also been shown to form multimers and to strongly facilitate in vivo activation of a promoter by distantly located enhancer CRMs [44], [45]. More recently, molecular studies have used high-magnification confocal imaging and 2D RNA fluorescence in situ hybridization (FISH) to visualize specific physical associations between distantly located cis-regulatory sequences in the nucleus at the human beta-globin locus [37].
Direct evidence for chromatin looping at the Drosophila BX-C comes from a study examining physical chromosomal interactions using probes designed against the IAB5 and IAB8 enhancers and a promoter-proximal region upstream of the Abd-B m transcription unit, containing the PTE sequence [46]. In a portion of nuclei taken from the eighth abdominal segment region of a germ-band elongation stage embryo, FISH signals from the Abd-B m promoter region co-localize with signals from the distal IAB5 enhancer, while the more proximal IAB8 enhancer remains distantly located and disassociated [46]. This result suggests that physical associations may indeed occur between the IAB5 enhancer and the 5′ upstream region of the Abd-B m promoter to facilitate expression of the Abd-B m transcript. These data support our model that tethering mediates the interaction of long-range enhancers (such as IAB5) to the Abd-B m promoter, but that it is not required for the functional interaction of the 3′ proximal IAB8 enhancer to the Abd-B m promoter.
In our current model the PTE may bind sequence-specific protein factors which interact with complementary factors bound near the IAB enhancers, allowing a molecular bridge or loop to form between the Abd-B m promoter and the IAB enhancers [17]. This model does not exclude the possibility that additional molecular interactions between known regulatory regions in the BX-C, including insulator and PTS sequences, may also mediate enhancer-promoter communication. In future studies other techniques, such as three-dimensional chromatin conformation capture (3C) [47] and Dam methylase identification [48], will be well suited to more fully elucidate the nature of such interactions and the molecular mechanisms of PTE function. In addition, molecular dissection of the functional sequences within the PTE and 5′ associated sequences in transgenic tethering assays and the biochemical identification of putative sequence-specific DNA-binding trans factors that may mediate the PTE activity will be essential.
Materials and Methods
Genetic insertion lines
The Abd-BT2N and Abd-BLDN P element fly lines inserted −241 bp 5′ of the Abd-B m transcription start site were provided by Ernesto Sanchez-Herrero [27], [31].
P element imprecise excision from the Abd-BLDN line
Abd-BLDN flies were crossed with a transposase expressing line (Δ2–3) carrying a Stubble (Sb) dominant phenotypic marker (BL Stock 1798) in Cross 1 (Fig. S1). Cross 1 male progeny were screened for Sb (marking the presence of Δ2–3 transposase) and variegated red eye color (rather than white eye color). The selected Cross 1 males were crossed with female with D, a dominant marker, and a TM3 balancer chromosome carrying Serrate (Ser), a dominant phenotypic marker (BL Stock 7198) in Cross 2 (Fig. S1). The male Cross 2 progeny were screened for Ser (marking the presence of the TM3 balancer), white eyes (indicating excision of the Abd-B-GAL4LDN P element construct), and the absence of Sb (indicating loss of the Δ2–3 transposase) as well as against D. The selected Cross 2 male flies were crossed with BL Stock 7198 flies again in Cross 3 (Fig. S1). After a few days, the male parents were recovered from Cross 3 vials. PCR amplification of the Abd-B region with primers located 0.5 kb, 1 kb, and 1.5 kb upstream of the Abd-B transcription start site on the genomic DNA prepared from approximately 100 selected Cross 2 male flies was used to detect deletions in the PTE sequence.
Primers used:
Abd-B promoter out: 5′-CGA CAA CAT ATC CAC ATC GCT-3′
Abd-B promoter 0.5 kb in: 5′-AAG TGC GAT ACC ATC TTT-3′
Abd-B promoter 1 kb in: 5′-TGC CTT TGG AAG TGA GAC AA-3′
Abd-B promoter 1.5 kb in: 5′-GGA AAT AGA TTG CGG CAG TTA A-3′
The PCR products were ligated into pGEM-T Easy vector (Promega) and sequenced using T3 and SP6 sequencing primers. Progeny from Cross 3 vials seeded with a male parent exhibiting a disruption of the PTE were screened against D and for Ser and self-crossed to generate a balanced mutant line.
In situ analysis of abd-A and Abd-B expression
In situ hybridization probes to detect transcription of Abd-B and abd-A were PCR-amplified using D. melanogaster yw67 adult genomic DNA as a template. The previously described DNA sequences for the Bexon region (exon 8 of the D. melanogaster Abd-B gene), BPP (specific to the r transcript) and Aexon region [32] were PCR amplified and cloned into pGEMT-Easy (Promega). PCR primer sequences were as follows:
Bexon s: 5′-GAACAAGAAGAACTCACAGC-3′;
Bexon as: 5′-TAGGCATAGGTGTAGGTGTAGG-3′;
BPP s: 5′-TATTATTCGTCTCCAGTCGC-3′;
BPP as: 5′-CTCAGATTGATGGTGGTGGTGG-3′;
Aexon s: 5′- CACCAACAGCAGCAACAACAGC-3′ (173566);
Aexon as: 5′- CATTGTATTCAAGCGTTGGC-3′ (174756);
Antisense RNA probes (relative to the direction of Abd-B and abd-A transcription) were prepared using a digoxigenin (DIG) RNA-labeling kit (Roche, Gipf-Oberfrick, Switzerland). Embryos from each of the wild-type D. melanogaster, and mutant Abd-BT2N, Abd-BLDN and Abd-BΔPTE-UP lines were collected, fixed and hybridized with the appropriate probes as previously described [32]. Anti-sense Bexon RNA probes enable the detection of both the m and r transcript and anti-sense BPP RNA probes enable the specific detection of only the Abd-B r transcript [32].
Supporting Information
Crosses to generate Abd-BΔPTE-UP mutant. (A) Schematic diagram of the Abd-BLDN P element insertion line, showing the insertion site −241 bp 5′ of the Abd-B m transcription start site in the endogenous PTE sequence. The P element insertion in the Abd-BLDN line contains GAL4 (purple box) and white (white box) reporter genes. The same symbols and color scheme as shown in Fig. 1 are used to show the cis-regulatory modules. (B) Abd-BLDN flies were crossed with a transposase expressing line (Δ2–3) carrying Stubble (Sb) (BL Stock 1798) (Cross 1). Cross 1 male progeny were screened for Sb and variegated red eye color (rather than white eye color). (C) The selected Cross 1 males were crossed with a female with D, a dominant marker, and a TM3 balancer chromosome carrying Serrate (Ser), a dominant phenotypic marker (BL Stock 7198) (Cross 2). Cross 2 male progeny were screened for Ser, white eyes (indicating excision of the GAL4 LDN P element construct), and the absence of Sb and D. (D) The selected Cross 2 male flies were crossed with BL Stock 7198 flies again (Cross 3). After a few days, the male parents were recovered from the Cross 3 vials. PCR amplification of the Abd-B promoter region with primers (black triangles) located 0.5 kb, 1 kb, and 1.5 kb upstream of the Abd-B transcription start site on the genomic DNA prepared from these selected Cross 2 male flies was used to detect deletions in the PTE sequence. Progeny originating from Cross 3 vials seeded with a male parent exhibiting a disruption of the PTE (ΔPTE) were recovered and screened against D and for Ser. (E) These selected Cross 3 progeny were then self-crossed to generate the Abd-B ΔPTE-UP/TM3,Ser balanced line.
(TIF)
Acknowledgments
The authors would like to thank Ernesto Sanchez-Herrero for providing the Abd-BT2N and Abd-BLDN lines. We would also like to thank Karl G. Johnson and Kathleen M. Beckingham for their helpful advice on the imprecise P element excision.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: The research in this paper was supported by funding to R.A.D. from the National Institutes of Health (NIH-HD54977) and the National Science Foundation (IOS-0845103) and a Howard Hughes Medical Institute Undergraduate Science Education Program grant (520051213) to the Biology department at Harvey Mudd College. M.C.W.H. received support from the Merck-American Association for the Advancement of Science (AAAS) Undergraduate Science Research Program. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Caplan AI, Ordahl CP. Irreversible gene repression model for control of development. Science. 1978;201:120–130. doi: 10.1126/science.351805. [DOI] [PubMed] [Google Scholar]
- 2.Lewis EB. A gene complex controlling segmentation in Drosophila. Nature. 1978;276:565–570. doi: 10.1038/276565a0. [DOI] [PubMed] [Google Scholar]
- 3.Spitz F, Gonzalez F, Duboule D. A Global Control Region Defines a Chromosomal Regulatory Landscape Containing the HoxD Cluster. Cell. 2003;113:405–417. doi: 10.1016/s0092-8674(03)00310-6. [DOI] [PubMed] [Google Scholar]
- 4.Osborne CS, Chakalova L, Brown KE, Carter D, Horton A, et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat Genet. 2004;36:1065–1071. doi: 10.1038/ng1423. [DOI] [PubMed] [Google Scholar]
- 5.Dorsett D. Distant liaisons: long-range enhancer-promoter interactions in Drosophila. Curr Opin Genet Dev. 1999;9:505–514. doi: 10.1016/s0959-437x(99)00002-7. [DOI] [PubMed] [Google Scholar]
- 6.Kellum R, Elgin SC. Chromatin boundaries: punctuating the genome. Curr Biol. 1998;8:R521–524. doi: 10.1016/s0960-9822(07)00337-5. [DOI] [PubMed] [Google Scholar]
- 7.Mihaly J, Hogga I, Barges S, Galloni M, Mishra RK, et al. Chromatin domain boundaries in the Bithorax complex. Cell Mol Life Sci. 1998;54:60–70. doi: 10.1007/s000180050125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ohtsuki S, Levine M, Cai HN. Different core promoters possess distinct regulatory activities in the Drosophila embryo. Genes & Development. 1998;12:547–556. doi: 10.1101/gad.12.4.547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bell AC, Felsenfeld G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature. 2000;405:482–485. doi: 10.1038/35013100. [DOI] [PubMed] [Google Scholar]
- 10.Celniker S, Drewell RA. Chromatin looping mediates boundary element promoter interactions. BioEssays. 2007;29:7–10. doi: 10.1002/bies.20520. [DOI] [PubMed] [Google Scholar]
- 11.Bushey AM, Dorman ER, Corces VG. Chromatin insulators: regulatory mechanisms and epigenetic inheritance. Molecular Cell. 2008;10:1–9. doi: 10.1016/j.molcel.2008.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wallace JA, Felsenfeld G. We gather together: insulators and genome organization. Curr Opin Genet Dev. 2007;17:400–407. doi: 10.1016/j.gde.2007.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bender W, Akam M, Karch F, Beachy PA, Peifer M, et al. Molecular Genetics of the Bithorax Complex in Drosophila melanogaster. Science. 1983;221:23–29. doi: 10.1126/science.221.4605.23. [DOI] [PubMed] [Google Scholar]
- 14.Martin CH, Mayeda CA, Davis CA, Ericsson CL, Knafels JD, et al. Complete sequence of the bithorax complex of Drosophila. Proceedings of the National Academy of Sciences of the United States of America. 1995;92:8398–8402. doi: 10.1073/pnas.92.18.8398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Karch F, Weiffenbach B, Peifer M, Bender W, Duncan I, et al. The abdominal region of the bithorax complex. Cell. 1985;43:81–96. doi: 10.1016/0092-8674(85)90014-5. [DOI] [PubMed] [Google Scholar]
- 16.Akbari OS, Bae E, Johnsen H, Villaluz A, Wong D, et al. A novel promoter-tethering element regulates enhancer-driven gene expression at the bithorax complex in the Drosophila embryo. Development. 2008;135:123–131. doi: 10.1242/dev.010744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Akbari OS, Schiller BJ, Goetz SG, Ho MCW, Bae E, et al. The Abdominal-B promoter tethering element mediates promoter-enhancer specificity at the Drosophila bithorax complex. Fly. 2007;1:337–339. doi: 10.4161/fly.5607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mihaly J, Barges S, Sipos L, Maeda R, Cleard F, et al. Dissecting the regulatory landscape of the Abd-B gene of the bithorax complex. Development. 2006;133:2983–2993. doi: 10.1242/dev.02451. [DOI] [PubMed] [Google Scholar]
- 19.Zhou J, Ashe H, Burks C, Levine M. Characterization of the transvection mediating region of the abdominal-B locus in Drosophila. Development. 1999;126:3057–3065. doi: 10.1242/dev.126.14.3057. [DOI] [PubMed] [Google Scholar]
- 20.Karch F, Galloni M, Sipos L, Gausz J, Gyurkovics H, et al. Mcp and Fab-7: molecular analysis of putative boundaries of cis-regulatory domains in the bithorax complex of Drosophila melanogaster. Nucleic Acids Research. 1994;22:3138–3146. doi: 10.1093/nar/22.15.3138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pérez-Lluch S, Cuartero S, Azorín F, Espinàs ML. Characterization of new regulatory elements within the Drosophila bithorax complex. Nucleic Acids Res. 2008;36:6926–6933. doi: 10.1093/nar/gkn818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gyurkovics H, Gausz J, Kummer J, Karch F. A new homeotic mutation in the Drosophila bithorax complex removes a boundary separating two domains of regulation. The Embo Journal. 1990;9:2579–2585. doi: 10.1002/j.1460-2075.1990.tb07439.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lin Q, Wu D, Zhou J. The promoter targeting sequence facilitates and restricts a distant enhancer to a single promoter in the Drosophila embryo. Development. 2003;130:519–526. doi: 10.1242/dev.00227. [DOI] [PubMed] [Google Scholar]
- 24.Lin Q, Chen Q, Lin L, Smith S, Zhou J. Promoter targeting sequence mediates enhancer interference in the Drosophila embryo. Proc Natl Acad Sci U S A. 2007;104:3237–3242. doi: 10.1073/pnas.0605730104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sipos L, Mihaly J, Karch F, Schedl P, Gausz J, et al. Transvection in the Drosophila Abd-B domain: extensive upstream sequences are involved in anchoring distant cis-regulatory regions to the promoter. Genetics. 1998;149:1031–1050. doi: 10.1093/genetics/149.2.1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sipos L, Gyurkovics H. Long-distance interactions between enhancers and promoters. The case of the Abd-B domain of the Drosophila bithorax complex. FEBS Journal. 2005;272:3253–3259. doi: 10.1111/j.1742-4658.2005.04757.x. [DOI] [PubMed] [Google Scholar]
- 27.Estrada B, Casares F, Busturia A, Sanchez-Herrero E. Genetic and molecular characterization of a novel iab-8 regulatory domain in the Abdominal-B gene of Drosophila melanogaster. Development. 2002;129:5195–5204. doi: 10.1242/dev.129.22.5195. [DOI] [PubMed] [Google Scholar]
- 28.Casanova J, Sanchez-Herrero E, Morata G. Identification and characterization of a parasegment specific regulatory element of the abdominal-B gene of drosophila. Cell. 1986;47:627–636. doi: 10.1016/0092-8674(86)90627-6. [DOI] [PubMed] [Google Scholar]
- 29.Sánchez-Herrero E, Crosby MA. The Abdominal-B gene of Drosophila melanogaster: overlapping transcripts exhibit two different spatial distributions. EMBO J. 1988;7:2163–2173. doi: 10.1002/j.1460-2075.1988.tb03055.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Delorenzi M, Bienz M. Expression of Abdominal-B homeoproteins in Drosophila embryos. Development. 1990;108:323–329. doi: 10.1242/dev.108.2.323. [DOI] [PubMed] [Google Scholar]
- 32.Bae E, Calhoun VC, Levine M, Lewis EB, Drewell RA. Characterization of the intergenic RNA profile at abdominal-A and Abdominal-B in the Drosophila bithorax complex. PNAS. 2002;99:16847–16852. doi: 10.1073/pnas.222671299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ho MC, Johnsen H, Goetz SE, Schiller BJ, Bae E, et al. Functional evolution of cis-regulatory modules at a homeotic gene in Drosophila. PLoS Genetics. 2009;5:e1000709. doi: 10.1371/journal.pgen.1000709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Boulet AM, Lloyd A, Sakonju S. Molecular definition of the morphogenetic and regulatory functions and the cis-regulatory elements of the Drosophila Abd-B homeotic gene. Development. 1991;111:393–405. doi: 10.1242/dev.111.2.393. [DOI] [PubMed] [Google Scholar]
- 35.Macias A, Casanova J, Morata G. Expression and regulation of the abd-A gene of Drosophila. Development. 1990;110:1197–1207. doi: 10.1242/dev.110.4.1197. [DOI] [PubMed] [Google Scholar]
- 36.Chopra VS, Cande J, Hong JW, Levine M. Stalled Hox promoters as chromosomal boundaries. Genes and Development. 2009;23:1505–1509. doi: 10.1101/gad.1807309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Patrinos GP, de Krom M, de Boer E, Langeveld A, Imam AMA, et al. Multiple interactions between regulatory regions are required to stabilize an active chromatin hub. Genes Dev. 2004;18:1495–1509. doi: 10.1101/gad.289704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Calhoun VC, Stathopoulos A, Levine M. Promoter-proximal tethering elements regulate enhancer-promoter specificity in the Drosophila Antennapedia complex. Proceedings of the National Academy of Sciences of the United States of America. 2002;99:9243–9247. doi: 10.1073/pnas.142291299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Calhoun VC, Levine M. Long-range enhancer-promoter interactions in the Scr-Antp interval of the Drosophila Antennapedia complex. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:9878–9883. doi: 10.1073/pnas.1233791100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Fujioka M, Wu X, Jaynes JB. A chromatin insulator mediates transgene homing and very long-range enhancer-promoter communication. Development. 2009;136:3077–3087. doi: 10.1242/dev.036467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kwon D, Mucci D, Langlais K, Americo JL, DeVido SK, et al. Enhancer-promoter communication at the Drosophila engrailed locus. Development. 2009;136:3067–3075. doi: 10.1242/dev.036426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cande J, Chopra VS, Levine M. Evolving enhancer-promoter interactions within the tinman complex of the flour beetle, Tribolium castaneum. Development. 2009;136:3153–3160. doi: 10.1242/dev.038034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zeller RW, Griffith JD, Moore JG, Kirchhamer CV, Britten RJ, et al. A multimerizing transcription factor of sea urchin embryos capable of looping DNA. PNAS. 1995;92:2989–2993. doi: 10.1073/pnas.92.7.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Su W, Jackson S, Tjian R, Echols H. DNA looping between sites for transcriptional activation: self-association of DNA-bound Sp1. Genes & Development. 1991;5:820–826. doi: 10.1101/gad.5.5.820. [DOI] [PubMed] [Google Scholar]
- 45.Mastrangelo IA, Courey AJ, Wall JS, Jackson SP, Hough PV. DNA looping and Sp1 multimer links: a mechanism for transcriptional synergism and enhancement. Proceedings of the National Academy of Sciences of the United States of America. 1991;88:5670–5674. doi: 10.1073/pnas.88.13.5670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ronshaugen M, Levine M. Visualization of trans-homolog enhancer-promoter interactions at the Abd-B Hox locus in the Drosophila embryo. Dev Cell. 2004;7:925–932. doi: 10.1016/j.devcel.2004.11.001. [DOI] [PubMed] [Google Scholar]
- 47.Hagège H, Klous P, Braem C, Splinter E, Dekker J, et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nat Protoc. 2007;2:1722–1733. doi: 10.1038/nprot.2007.243. [DOI] [PubMed] [Google Scholar]
- 48.Cleard F, Moshkin Y, Karch F, Maeda RK. Probing long-distance regulatory interactions in the Drosophila melanogaster bithorax complex using Dam identification. Nature Genetics. 2006;38:931–935. doi: 10.1038/ng1833. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crosses to generate Abd-BΔPTE-UP mutant. (A) Schematic diagram of the Abd-BLDN P element insertion line, showing the insertion site −241 bp 5′ of the Abd-B m transcription start site in the endogenous PTE sequence. The P element insertion in the Abd-BLDN line contains GAL4 (purple box) and white (white box) reporter genes. The same symbols and color scheme as shown in Fig. 1 are used to show the cis-regulatory modules. (B) Abd-BLDN flies were crossed with a transposase expressing line (Δ2–3) carrying Stubble (Sb) (BL Stock 1798) (Cross 1). Cross 1 male progeny were screened for Sb and variegated red eye color (rather than white eye color). (C) The selected Cross 1 males were crossed with a female with D, a dominant marker, and a TM3 balancer chromosome carrying Serrate (Ser), a dominant phenotypic marker (BL Stock 7198) (Cross 2). Cross 2 male progeny were screened for Ser, white eyes (indicating excision of the GAL4 LDN P element construct), and the absence of Sb and D. (D) The selected Cross 2 male flies were crossed with BL Stock 7198 flies again (Cross 3). After a few days, the male parents were recovered from the Cross 3 vials. PCR amplification of the Abd-B promoter region with primers (black triangles) located 0.5 kb, 1 kb, and 1.5 kb upstream of the Abd-B transcription start site on the genomic DNA prepared from these selected Cross 2 male flies was used to detect deletions in the PTE sequence. Progeny originating from Cross 3 vials seeded with a male parent exhibiting a disruption of the PTE (ΔPTE) were recovered and screened against D and for Ser. (E) These selected Cross 3 progeny were then self-crossed to generate the Abd-B ΔPTE-UP/TM3,Ser balanced line.
(TIF)