Abstract
Non-coding RNA polymerase II transcripts are processed by the poly(A)-independent termination pathway that requires the Nrd1 complex. The Nrd1 complex includes two RNA-binding proteins, the nuclear polyadenylated RNA-binding (Nab) 3 and the nuclear pre-mRNA down-regulation (Nrd) 1 that bind their specific termination elements. Here we report the solution structure of the RNA-recognition motif (RRM) of Nab3 in complex with a UCUU oligonucleotide, representing the Nab3 termination element. The structure shows that the first three nucleotides of UCUU are accommodated on the β-sheet surface of Nab3 RRM, but reveals a sequence-specific recognition only for the central cytidine and uridine. The specific contacts we identified are important for binding affinity in vitro as well as for yeast viability. Furthermore, we show that both RNA-binding motifs of Nab3 and Nrd1 alone bind their termination elements with a weak affinity. Interestingly, when Nab3 and Nrd1 form a heterodimer, the affinity to RNA is significantly increased due to the cooperative binding. These findings are in accordance with the model of their function in the poly(A) independent termination, in which binding to the combined and/or repetitive termination elements elicits efficient termination.
Keywords: NMR, Protein Structure, RNA Processing, RNA-Protein Interaction, Transcription Termination
Introduction
RNA Polymerase II (RNA Pol II)6 transcribes messenger RNA (mRNA), but also a subset of small nuclear and small nucleolar RNAs (snRNAs/snoRNAs), micro-RNA precursors, and a class of intergenic and antisense RNAs (1). RNA Pol II uses two different mechanisms for transcription termination of these “coding” and “non-coding” RNAs. Although the RNA Pol II termination of mRNA requires a large multiprotein complex that recognizes the poly(A) signal in the nascent transcript (2), the termination of the non-coding RNAs requires no poly(A) signal (2–4).
In the poly(A)-independent mechanism, transcription termination requires a specific factor, the Nrd1 complex. This complex consists of three proteins: the nuclear pre-mRNA down-regulation (Nrd) 1 protein, the nuclear polyadenylated RNA-binding (Nab) 3 protein, and the putative RNA helicase Sen1 (5–7). The Nrd1 complex interacts with the exosome, a complex of 10–12 exoribonucleolytic and RNA-binding proteins (8) and the Trf4-Air2-Mtr4 polyadenylation (TRAMP) complex (9–11), which are involved in the 3′ end processing of non-coding RNA transcripts (3, 4, 7).
In yeast, transcription termination mediated by the Nrd1 complex requires binding to both the nascent RNA and the carboxyl-terminal domain of RNA Pol II, which consists of 26 repeats of the sequence Tyr1-Ser2-Pro3-Thr4-Ser5-Pro6-Ser7 (1, 12). Interestingly, the Nrd1 complex binds the carboxyl-terminal domain when it is phosphorylated at Ser5, a typical feature of the early elongation phase of the transcription cycle. The Ser5-phosphorylated carboxyl-terminal domain is recognized by the carboxyl-terminal domain-interacting domain of Nrd1 (13, 14). The RNA-binding subunits of the Nrd1 complex, Nrd1 and Nab3, recognize their specific RNA sequences (called terminator elements) in the nascent transcripts of RNA Pol II. It is believed that this specific binding of Nrd1 complex to the terminator elements is the initial step in the assembly of termination machinery.
A number of studies narrowed the sequence regions with terminator elements (5, 6, 15–17) that were subsequently identified as GUAR (where R stands for purine) and UCUU sequences (18). GUAR and UCUU terminator elements are recognized by Nrd1 and Nab3, respectively, via their fragments encompassing RNA recognition motifs (RRMs) (18). These terminator sequences are located downstream of snRNA and snoRNA genes (18) although their relative orientation and spacing are not highly conserved. In addition, it was demonstrated that Nrd1 and Nab3 form a stable heterodimer and bind to snoRNA terminators that contain multiple Nrd1- and Nab3-binding sequences (19).
Both Nrd1 and Nab3 contain RRM that likely mediates the binding to their specific RNA sequences. The RRM is the most abundant RNA-binding domain in higher vertebrates; e.g. the RRM is present in about 2% of human genes (20). It is a small protein domain of ∼90 amino acids with a typical βαββαβ topology that forms a four-stranded β-sheet packed against two α-helices (21–23). The structure of this domain is relatively well defined despite a little sequence conservation among various RRMs. The solved structures of RRM bound to RNA show the complexity of protein-RNA recognition mediated by the RRM, which often involves not only RRM-RNA interactions but also RRM-RRM and other RRM-protein interactions. The main protein surface of the RRM involved in the interaction with the RNA is the four-to-five-stranded β-sheet, which typically contacts two or three nucleotides. Frequently, RRM-containing proteins bind more than three nucleotides and recognize longer single-stranded RNA or even internal RNA loops by employing of β-strand loops and N- or C-terminal flanking regions of RRMs (21–23).
To better understand the structural basis behind the poly(A) independent transcription termination pathway, we initiated an NMR study of Saccharomyces cerevisiae Nab3. Here, we present the three-dimensional solution structure of the Nab3 RRM in free form and in complex with the 5′-UCUU-3′ RNA substrate. The structure of the complex reveals recognition of the YCU sequence (where Y stands for pyrimidine) by the Nab3 RRM. We confirmed the sequence-specific intermolecular contacts by site-directed mutagenesis and fluorescence anisotropy (FA) measurements, and their physiological role was also confirmed by yeast phenotypic analyses. Finally, we demonstrate that the weak RNA binding of the isolated RRMs of Nab3 and Nrd1 is greatly enhanced when Nab3 and Nrd1 form a heterodimer and bind the RNA cooperatively.
EXPERIMENTAL PROCEDURES
Cloning, Expression, and Purification of Proteins
The coding sequence corresponding to the RRM of the Nab3 gene from S. cerevisiae (961–1245) was amplified by polymerase chain reaction (PCR), and cloned into a pET22b expression vector (Novagen) via NdeI and XhoI restriction sites. The resulting C-terminal His6-tagged construct was verified by DNA sequencing. The protein was overexpressed in Escherichia coli BL21-Codon Plus (DE3)-RIPL (Stratagene), transformed with the pET22b-RRM Nab3 construct at 37 °C in M9 minimal medium, supplemented with 50 mg/liter of ampicillin. For isotope labeling, the medium was supplemented with 15NH4Cl and [U-13C6]glucose. Cells were grown at 37 °C to A600 ∼1 and induced with 1 mm isopropyl β-d-thiogalactoside. Cells were harvested by centrifugation (6000 × g for 10 min), resuspended in lysis buffer (50 mm sodium phosphate, 300 mm NaCl, 10 mm β-mercaptoethanol, pH 8), and disrupted by sonication. The cell debris was cleared by centrifugation (14,000 × g for 60 min). Soluble lysate was loaded on a nickel-nitrilotriacetic acid column (Qiagen), equilibrated with lysis buffer, washed with a high salt buffer (50 mm sodium phosphate, 500 mm NaCl, 10 mm β-mercaptoethanol, 5 mm imidazole, pH 8), and eluted with imidazole gradient (50–500 mm) of elution buffer (50 mm sodium phosphate, 300 mm NaCl, 10 mm β-mercaptoethanol, pH 8). The protein was subsequently loaded on a Superdex 75 gel filtration column (GE Healthcare), equilibrated with lysis buffer. The protein fractions from gel filtration were dialyzed against lysis buffer. The purified protein was 99% pure, as judged by Coomassie-stained SDS-PAGE. For NMR measurements the pure protein was concentrated to 2.5 mm in 550 μl of 50 mm sodium phosphate (pH 8.0), containing 300 mm NaCl, and 10 mm β-mercaptoethanol. The cloning, expression, and purification of Nrd1 RRM-(340–410) were carried out in the same way as for Nab3 RRM.
The expression and purification of the Nrd1-Nab3 heterodimer have been done in a similar manner as reported previously (19). To improve the yield of expression, we used E. coli BL21-Codon Plus (DE3)-RIPL (Stratagene). We used the following final buffer (50 mm Tris (pH 8.0), containing 150 mm NaCl, and 10 mm β-mercaptoethanol) to have the same conditions for all fluorescence anisotropy measurements. Prior to RNA titration, all proteins were tested for the residual RNase activity using RNaseAlert Lab Test (Ambion). RNA oligonucleotides were purchased from Thermo Fisher Scientific/Dharmacon and Sigma.
Generation of Nab3 RRM Mutants
Site-specific mutagenesis was performed using the QuikChange site-directed mutagenesis kit (Stratagene) with complementary sense and antisense (AS) oligonucleotide primers as follows: R331A (S), 5′-gcacaatattcctccgaagtcagcattattcattggtaatttgccg-3′ and (AS), 5′-cggcaaattaccaatgaataatgctgacttcggaggaatattgtgc-3′; N361A (S), 5′-tccatacggtcatatcatgcaaatcgctatcaaaaatgcctttggattcatt-3′ and (AS), 5′-aatgaatccaaaggcatttttgatagcgatttgcatgatatgaccgtatgga-3′; E397A (S), 5′-gcaaaaagttgatcctggcagtttctagctcgaatgc-3′ and (AS), 5′-gcattcgagctagaaactgccaggatcaactttttgc-3′; E397K (S), 5′-tggcaaaaagttgatcctgaaagtttctagctcgaatgc-3′ and (AS) 5′-gcattcgagctagaaactttcaggatcaactttttgcca-3′; S399A (S), 5′-agttgatcctggaagttgctagctcgaatgctcgt-3′ and (AS), 5′-acgagcattcgagctagcaacttccaggatcaact-3′; S399K (S), 5′-actttggcaaaaagttgatcctggaagttaaaagctcgaatgctcgtcc-3′ and (AS), 5′-ggacgagcattcgagcttttaacttccaggatcaactttttgccaaagt-3′. All mutations were verified by DNA sequence analysis.
NMR Spectroscopy
All NMR spectra of 2.5 mm uniformly 15N,13C-labeled Nab3 RRM in 50 mm sodium phosphate buffer (pH 8.0), 300 mm NaCl, 10 mm β-mercaptoethanol (90% H2O/10% D2O) were recorded on Bruker AVANCE 600 and 900 MHz spectrometers equipped with a cryoprobe at a sample temperature of 30 °C. All spectra were processed with Topspin 2.1 (Bruker BioSpin) and analyzed with Sparky 3.0 (T. G. Goddard and D. G. Kneller, University of California, San Francisco). The 1H, 13C, and 15N chemical shifts of Nab3 RRM were assigned as described previously (24). All distance restraints were derived from the three-dimensional 15N- and 13C-edited NOESYs and two-dimensional 1H-1H-labeled NOESY (with mixing time of 150 ms) collected at 900 MHz spectrometer.
Akin to the free Nab3 RRM, the backbone resonance assignments of Nab3 RRM in the bound form were achieved using three-dimensional triple resonance experiments: HNCA, HNCACB, and CBCA(CO)NH (24–26). Resonances of the aliphatic side chains were assigned by a combination of three-dimensional HCCH TOCSY, three-dimensional HNHA, and three-dimensional 13C-edited NOESY spectra. Resonances of the aromatic side chains were assigned using two-dimensional homonuclear NOESY, three-dimensional 13Carom-edited NOESY, and two-dimensional (HB)CB(CGCD)HD spectra. All distance restraints were derived from the three-dimensional 15N- and 13C-edited NOESYs (with mixing time of 150 ms) collected at 900 MHz spectrometer. The RNA resonances in complex were assigned using a combination of standard through-space and through-bond experiments (27, 28). We could not assign all sugar resonances unambiguously due to a high resonance overlap in the spectra (only unlabeled RNA was used). The sugar pucker conformation for all nucleotides is C2′-endo, as identified in the two-dimensional homonuclear TOCSY spectrum (strong cross-peaks between the H1′ and H2′ resonances). Intermolecular distance constraints were obtained from the three-dimensional 13C F1-edited, F3-filtered NOESY-HSQC experiment (29), which was recorded in H2O (with WATERGATE water suppression) as a two-dimensional filter NOESY omitting 13C chemical shift evolution.
Structure Calculations
The preliminary structure determinations of the free and bound Nab3 RRM were performed with the automated NOE assignment module implemented in the CYANA program (30). This automated NOE assignment procedure is a re-implementation of the former CANDID algorithm (31) on the basis of a probabilistic treatment of the NOE assignment. CYANA carries out automated assignment and distance calibration of NOE intensities, removal of meaningless restraints, structure calculation with torsion angle dynamics, and automatic upper distance limit violation analysis. The resultant NOE cross-peak assignments were subsequently confirmed by visual inspection of the spectra. The predicted protein backbone φ and ψ torsion angle from the chemical shifts (32) for the secondary structure elements were also included in the calculations. In the next step, CYANA-generated restraints along with manually assigned protein-RNA intermolecular restraints were used for further refinement of the preliminary structures with AMBER 10.0 software (33). This calculations employed a modified version (AMBER ff99SB) of the force field described by Cornell et al. (34) along with a refinement protocol described in Padrta et al. (35), and an explicit solvent. From 40 refined structures, the 20 conformers with the lowest AMBER energy were selected to form the final ensemble of structures. Structural quality was assessed using PROCHECK (36) and WHAT IF (37). Molecular graphics were generated using MOLMOL (38) and PyMOL (57).
Fluorescence Anisotropy Measurements
The equilibrium binding of Nab3 RRM to different oligonucleotides was analyzed by fluorescence anisotropy. The RNA oligonucleotides were either 5′-labeled with TAMRA or fluorescein attached via a hexyl linker. The measurements were conducted on a FluoroMax-4 spectrofluorometer (Horiba Jobin-Yvon Edison, NJ). The instrument was equipped with a thermostatted cell holder with a Neslab RTE7 water bath (Thermo Scientific). The whole system was operated using FluorEssence software (version 2.5.3.0, Horiba Jobin-Yvon). The TAMRA fluorophore was excited at 561 nm and its emission was collected at 581 nm. The widths of both excitation and emission monochromatic slits were 8 nm and integration time was set to 3 s. The fluorescein fluorophore was excited at 488 nm and its emission was collected at 520 nm. For measurement with the individual domains (Nrd1-(340–410) and Nab3-(331–415)) the width of both excitation and emission monochromatic slits were 7 nm and the integration time was set to 3 s. 10 nm labeled oligonucleotide (volume 1.4 ml) was titrated with increasing amounts of the protein in 50 mm sodium phosphate buffer (pH 7.5), supplemented with 150 mm NaCl and 10 mm β-mercaptoethanol. For measurement with the heterodimer (Nab3-(191–565)–Nrd1-(1–548)), 1 nm fluorescein-labeled RNA was used. Both excitation and emission monochromatic slits were 14 nm, the integration time was set to 3 s.
In all measurements, an identical concentration of the oligonucleotide was included in the protein stock solution to prevent dilution of the RNA during titration. All experiments were carried out at 25 °C in a stirred 1.5-ml quartz cuvette. Protein aliquots were added stepwise until the cuvette was filled. After this point, a certain volume of the sample was always removed from the cuvette before addition of a protein aliquot of the same volume. A fixed delay of 30 s was set between each aliquot addition and start of the measurement to allow the reaction to reach equilibrium. This delay was sufficient, as no further change in anisotropy was observed. Each data point is an average of five measurements. Neither TAMRA nor fluorescein labels showed binding with any of the protein constructs.
The data were analyzed in SigmaPlot 11 software (Systat Software). The experimental isotherms were fit to a single-site binding model according to Heyduk and Lee (39) using non-linear least squares regression. The data were normalized for visualization purposes.
Yeast Strains and Plasmids
The plasmid for the expression of wild-type Nab3 in yeast contain 550 bp of NAB3 promoter upstream of the AUG and a 286-bp sequence downstream of the stop codon. SV320 contains the wild-type NAB3 inserted in pRS415 (a LEU2 CEN plasmid) (40). Plasmids SV321-SV326 contain point mutants R331A, N361A, E397A, E397K, S399A, and S399K, respectively (see above for the primers). Strain DLY889 containing the endogenous NAB3 under control of the GAL1 promoter (3) was transformed with plasmids containing either wild-type Nab3 or Nab3 RRM point mutants with a LEU2 selectable marker (SV320, SV321, SV322, SV323, SV324, SV325, and SV326, respectively). The resulting strains were used for growth tests and Western blot analyses.
Growth Test Analyses
To test whether the mutated residues were essential for growth, the resulting transformants were grown in SC-LEU-HIS + 2% galactose at 30 °C to an A600 1.0. The cultures were then serially diluted in 96-well plates by a factor of 10, and spotted onto SC-LEU-HIS medium containing 2% glucose to repress the expression of the endogenous NAB3 or control medium (SC-HIS + 2% galactose). These plates were incubated at 24, 30, and 37 °C for 3 days.
Western Blot Analysis
Protein extracts were prepared from cultures grown either on galactose containing medium and cultures shifted to glucose containing medium (as described above). Proteins were resolved on a 12% SDS-PAGE gel, transferred to nitrocellulose membrane by a semi-dry electroblotter (Bio-Rad), and probed for the presence of Nab3p with the mAb 2F12 (41), or antibodies directed against the HA epitope (sc805, Santa Cruz Biotechnology) present on the endogenous Nab3. For loading control we used the antibodies against Air2 protein (42).
RESULTS
Structure of Nab3 RRM
The RRM of S. cerevisiae Nab3 was examined by NMR spectroscopy (Fig. 1, A and B). The 1H, 13C, and 15N chemical shift assignments were obtained as described previously (24). All NMR experiments were measured at a high salt concentration (300 mm NaCl, 50 mm sodium phosphate (pH 8.0), and 10 mm β-mercaptoethanol) to prevent protein precipitation. The solution structure determination of Nab3 RRM employed homonuclear and heteronuclear NMR techniques. The 15N-1H HSQC experiment shows a well dispersed spectrum (Fig. 1C), indicating a folded domain. There are a number of missing peaks in this spectrum that mainly correspond to the N- and C-terminal regions of the studied protein construct (outside of the RRM domain). These regions were included in the study as they often form additional structural elements (β-strand or α-helix) in RRMs or contribute to the RNA binding. In addition, several residues in the loops showed no NMR signals. These missing signals are likely a result from the relatively high pH used in the NMR study that was necessary to prevent the precipitation of Nab3 RRM. The three-dimensional structure of Nab3 RRM was determined by combined automated NOESY cross-peak assignment (30) and structure calculations with torsion angle dynamics implemented in the program CYANA 2.1 (43), followed by refinement in explicit solvent using AMBER 10 (33). An ensemble of the 20 lowest energy structures along with the best energy structure are shown in Fig. 1, D and E, respectively. These structures have an average backbone root mean square deviation of 0.57 ± 0.12 Å for the secondary structure elements. A full summary of structural statistics including the backbone φ-ψ angle distribution is given in Table 1.
FIGURE 1.
Overview of the RRM of Nab3 sequence, topology, NMR spectra, solution structure, and domain structure in Nab3 and Nrd1. A, amino acid sequence of the S. cerevisiae Nab3 RRM along with its secondary structure elements and general consensus of RNP1 and RNP2 motifs. B, a schematic drawing of the domain structure of Nab3 and Nrd1. C, two-dimensional 1H-15N HSQC spectrum of 2.5 mm uniformly 15N,13C-labeled Nab3 RRM in 50 mm sodium phosphate buffer (pH 8.0), 300 mm NaCl, and 10 mm β-mercaptoethanol (90% H2O, 10% D2O). The spectrum was acquired at 303 K on a Bruker Avance 600 MHz spectrometer. The assignments are labeled by the one-letter code of amino acids accompanied by a sequence number. The side chain resonances of asparagine and glutamine are connected by horizontal lines. D, stereo view of the 20 lowest energy structures of Nab3 RRM. The protein backbone is shown as a wire model. E, stereo view of the representative (the lowest energy) structure of Nab3 RRM shown as a ribbon diagram. The figure was generated with MOLMOL (38).
TABLE 1.
NMR and refinement statistics for Nab3 RRM and Nab3 RRM–UCUU complex
| NMR distance and dihedral angle restraints | Nab3 RRM | Nab3 RRM–UCUU complex |
|---|---|---|
| Distance restraints | ||
| Total NOEs | 857 | 852 |
| Intra-residue | 201 | 228 |
| Inter-residue | ||
| Sequential (|i–j| = 1) | 218 | 187 |
| Medium range (1 〈|i-j|〉5) | 164 | 116 |
| Long range (|i–j[bar ≥ 5) | 274 | 310 |
| Hydrogen bond restraints | 25 | 25 |
| Intermolecular | 11 | |
| Dihedral angle restraints | ||
| φ and ψ | 76 | 88a |
| Structure statisticsb | ||
| Residual NOE violations (mean ± S.D.) | ||
| Number > 0.20 Å | 1.5 (± 0.83) | 4 (± 2) |
| Maximum (Å) | 0.24 (± 0.03) | 0.46 (± 0.08) |
| Residual dihedral angle violations | ||
| Number > 10.0° | 0 | 0 |
| Maximum (°) | 0 | 0 |
| Ramachandran plot statisticsb,c,d | ||
| Residues in most favored regions (%) | 90.8 | 88.6 |
| Residues in additionally allowed regions (%) | 9.1 | 9.6 |
| Residues in generously allowed regions (%) | 0.1 | 1.7 |
| Residues in disallowed regions (%) | 0.0 | 0.1 |
| Deviations from idealized geometry | ||
| Bond length (Å) | 0.0011 ± 0.0001 | 0.0010 ± 0.0001 |
| Bond angles (Å) | 1.46 ± 0.02 | 1.48 ± 0.02 |
| Average root mean square deviation to mean structure (Å)b | ||
| Protein | ||
| Backbone atomsb | 0.57 ± 0.12 | 0.49 ± 0.10 |
| Heavy atomsb | 1.46 ± 0.16 | 1.24 ± 0.12 |
| RNA | ||
| All RNA heavy atomse | 0.98 ± 0.21 | |
| Complex | ||
| Proteinb and RNA heavy atomse | 1.27 ± 0.12 | |
| WHAT IFf structure Z-scoresd,g | ||
| Packing quality | −2.3 | −1.9 |
| Ramachandran plot appearance | −3.5 | −3.1 |
a Includes C2′-endo sugar pucker and anti conformation of the glycosidic bond was used for all nucleotides (56).
b Calculated for an ensemble of the 20 lowest energy structures.
c Based on PROCHECK analysis (32).
d Calculated for the structured part of the protein construct.
e Calculated for U1C2U3.
f Based on WHAT IF analysis (37).
The three-dimensional structure of Nab3 RRM adopts a compact fold with an β1α1β2β3α2β4 topology that is similar to the canonical fold of RRM family (21, 22). The fold is composed of two α-helices and a 310 helix that are packed along a face of a four-stranded antiparallel β-sheet. A central hydrophobic core composed of the residues shown in Fig. 1A stabilizes the fold of the domain. Nab3 RRM contains a well conserved signature of the RRM family, RNP1 and RNP2 sequences (44–46). These two conserved amino acid sequences found between Leu332–Leu337 and Asn364–Phe371 are located on the β3- and β1-strands, respectively. Their sequence compositions correspond to the general RNP2 and RNP1 consensus (ILV)-(FY)-(ILV)-X-N-L and (RK)-G-(FY)-(GA)-(FY)-(ILV)-X-(FY), respectively, except for the first two amino acids of the RNP1 (Fig. 1A). Nab3 RRM has asparagine and alanine in these positions (Fig. 1A). The presence of aromatic residues in RNP1 and RNP2 sequences, which usually mediates the stacking interaction with RNA bases, along with a number of basic and polar residues on the β-sheet surface, indicate a potential role of Nab3 RRM in RNA binding.
Characterization of the Nab3-UCUU Interactions by NMR
To investigate the interaction and binding mode between Nab3 RRM and RNA, we carried out an NMR chemical shift perturbation study with a UCUU element, which has been shown to elicit transcription termination via the Nrd1 pathway. In the RNA titration experiment, we observed that the protein amide resonances moved upon RNA binding from their initial positions, corresponding to the free form, in a stepwise directional manner until they reach their final positions that correspond to the fully bound state, with stoichiometry of 1:1 (Fig. 2). Additional RNA aliquots resulting in excess RNA resulted in no further change of chemical shifts, confirming the 1:1 stoichiometry of the complex. These titration data suggest that protein amide resonances are in a fast exchange regime between their free and bound forms relative to NMR time scale. The binding of UCUU to the RRM of Nab3 induces chemical shift perturbation of the residues shown in Fig. 2.
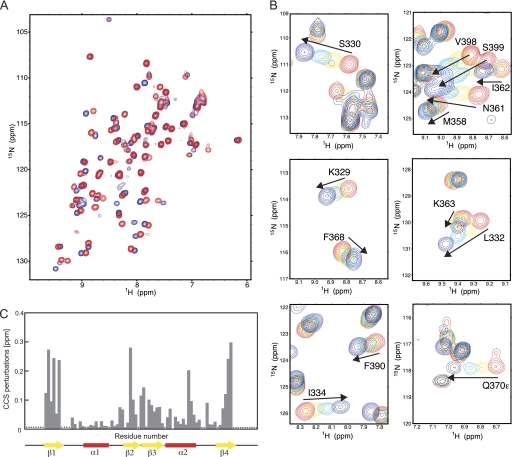
FIGURE 2.
NMR titration experiments of Nab3 RRM with UCUU RNA. A, 1H-15N HSQC spectra of Nab3 RRM alone (in red) and in the presence of 1 eq of 5′-UCUU-3′ (in blue) at 303 K. B, close-up views of the 1H-15N HSQC spectra, showing selected chemical shift changes during the titration. C, quantification of chemical shift perturbations of Nab3 RRM upon binding to UCUU RNA. The combined chemical shift perturbations ([ωHNΔδHN]2 + [ωNΔδN]2)1/2, where ωHN = 1 and ωN = 0.154 are weight factors of the nucleus (52), are plotted versus the amino acid residue number. Large changes occur on the β-sheet surface. The assignments of residues indicated by asterisks could not be obtained for neither the free nor bound protein, or indicates proline residues.
These chemical shift changes indicate that the above mentioned residues are involved in binding to the RNA, or alternatively, could undergo a conformational change upon RNA binding. Mapping the perturbed residues on the sequence of Nab3 RRM delineates that the Nab3 RRM binds the RNA through its β-sheet surface and also through the β2β3 loop (Fig. 2C).
Structure of Nab3 RRM in Complex with UCUU
When solving the structure of Nab3 RRM bound to RNA, we extensively tested different lengths of RNA, buffer conditions, and temperatures with the aim to optimize the NMR spectral quality of the complexes. Longer RNA substrates, a UCUU core motif with flanking sequences, resulted in the significant broadening of NMR signals of the complexes. Interestingly, we obtained the NMR spectra of better quality (for both protein and RNA in complex) with a four-nucleotide UCUU despite the fact that this RNA has lower affinity to Nab3 RRM compared with the longer substrates (see below). Similar improvement of the NMR spectral quality by using of a minimal specific RNA sequence has also been observed for other protein-RNA complexes investigated by NMR (47–49). Therefore, we pursued the structure determination of the Nab3 RRM-UCUU complex.
The RRM of Nab3 in complex with RNA display the canonical RRM-fold with an β1α1β2β3α2β4 topology and is similar to that of the unbound form (Figs. 1 and 3). Akin to the free form of Nab3 RRM, the N- and C-terminal regions as well as the long α2β4 loop are structurally undefined due to the lack of experimental data. The UCUU RNA adopts a single-stranded conformation and the first three nucleotides are positioned over the whole β-sheet surface in a canonical arrangement in which the 5′ end is located on the first half of the β-sheet (β4β1) and the 3′ end on the second half (β3β2) (21, 22) (Fig. 2). The overall position of RNA on the β-sheet coincides with the perturbed residues from the titration experiment (Figs. 2 and 3). All bases have an anti conformation of the glycosidic bond and C2′-endo conformation of the sugar pucker.
FIGURE 3.
Overview of the solution structure of the Nab3 RRM in complex with UCUU. A, stereo view of the 20 lowest energy structures of the Nab3 RRM-UCUU complex. The protein backbone is shown as a wire model in black. The RNA heavy atoms are shown as a wire model in red. B, stereo view of the representative (the lowest energy) structure of the Nab3 RRM-UCUU complex. The RNA is represented as a white stick model and the protein is shown as a ribbon model with residues that contact the RNA shown in yellow. Putative hydrogen bonds are shown by dotted magenta lines. C, scheme showing contacts between Nab3 RRM and the UCUU RNA. Protein residues that form putative hydrogen bonds to the RNA are shown in blue and the one having hydrophobic interactions are in yellow. A hypothetic recognition of U4 is labeled by a gray question mark. D, solvent-accessible surface representation of Nab3 RRM colored by electrostatic potential (blue, positive; red, negative) and stick representation for the RNA of the representative structure of the complex. Figures were generated with MOLMOL (38).
The NMR spectra provided a limited number of intermolecular NOEs (11 unambiguous intermolecular NOEs) that loosely define the position of U1C2U3 on the β-sheet surface of Nab3 RRM (Fig. 3A), but are sufficient to reveal the molecular basis of U1C2U3 recognition by Nab3 RRM (Fig. 3, B and C; the protein-RNA hydrogen bonds described below are inferred from the final ensemble of structures and thus they should be considered as putative hydrogen bonds). We could not define the position of U4 due to the lack of intermolecular NOEs. Based on NMR titration data, we speculate that U4 could be recognized by the asparagine side chain or a main chain of the β2–β3 loop that are in proximity to the base of U4, as displayed in the representative structure (Fig. 3B). In our NMR structure, C2 and U3 are involved in base stacking with the aromatic rings of Phe333 and Phe368, respectively. The Watson-Crick edge of C2 is recognized by the main chain carbonyl group of Val398 and the hydroxyl group of Ser399 that form hydrogen bonds with the amino and imino groups of C2, respectively. One-half of the 20 structures in the final ensemble has Arg331 in a position in which it contacts the O2 oxygen of C2 (for the importance of Arg331 see below). In addition, it is likely also that Ser400 could be involved in the recognition of C2 as the resonances of this residue broadened beyond detection upon RNA binding. Our structure also rules out the possibility that a purine could be accommodated in the C2 position due to a steric restriction imposed by the Glu397 side chain.
The recognition of U3 is mediated by the Arg331 and Asn361 side chains. The side chain NH2 group of Arg331 contacts the O4 carbonyl functional group of the base and the side chain carbonyl group of Asn361 forms a hydrogen bond with the imino proton of U3. Akin to C2, the position of U3 cannot be exchanged by a purine due to a steric hindrance of Arg331. The sugar of the U3 residue is further contacted by the aliphatic region of the Lys363 side chain.
In contrast to C2 and U3, the recognition of U1 is less evident from the structure. A single hydrogen bond is formed between the imino proton of U1 and the Oϵ of Glu397. There are also hydrophobic contacts between the sugar and the base of U1 and the side chain of Ile395. However, these contacts do not explain fully the sequence specificity of a uridine nucleotide. A cytidine nucleotide in this position could also form a similar interaction with the glutamate. Altogether, our NMR structure indicates that Nab3 RRM recognizes the YCU sequence (where Y stands for pyrimidine).
Nab3 RRM Binds the Nab3 Termination Element with Low Affinity
The FA measurements were carried out to further characterize the binding of Nab3 RRM to various RNA substrates. In FA measurements, formation of the protein-RNA complex is monitored directly from an increase of the FA value that occurs when the protein binds fluorescently labeled RNA. Binding curves were recorded in the course of titration experiments, where protein aliquots were added to 10 nm fluorescently labeled RNAs (Fig. 4).
FIGURE 4.
Equilibrium binding of Nab3 RRM, Nrd1 RRM, and Nrd1-Nab3 heterodimer with fluorescently labeled RNA monitored by fluorescence anisotropy. A, Nab3 RRM was titrated with UCUU and GUAA (each 10 nm), and their binding isotherms are shown as red circles and triangles, respectively. B, Nab3 RRM was titrated with snR47, snR13, and SL substrates (each 10 nm), and their binding isotherms are shown as red circles, squares, and triangles, respectively. C, Nrd1 RRM was titrated with GUAA and snR13 (each 10 nm) and their binding isotherms are shown as blue inverted triangles and circles, respectively. D, Nrd1-Nab3 heterodimer was titrated with snR13 (100 pm). E, summary of the association constants (Ka) for the RRM of Nrd1 (in blue) and Nab3 (in red) in their free forms as well as for the Nrd1-Nab3 heterodimer (in black). Logarithmic scale of Ka is shown to cover a wide range of affinities. F, RNA sequences used in the affinity measurements. The buffers contained the same ion strength and pH values for all proteins. Equilibrium dissociation constant (Kd) was calculated from the best fit to the data using a single-site binding isotherm. Error is denoted as S.E. The data were normalized for visualization purposes (A–C).
First, we assayed the binding affinity of Nab3 RRM to UCUU that has been reported as the minimal Nab3 terminator element. A tetranucleotide GUAA was used as a nonspecific control substrate. The comparison of the anisotropy data for specific and nonspecific four-nucleotide substrates along with determined equilibrium dissociation constants (Kd) are shown in Fig. 4A. Corresponding logarithmic values of Ka (Ka = 1/Kd) are shown in graph in Fig. 4E. Nab3 RRM binds the specific recognition sequence UCUU with more than 6-fold higher affinity compared with the nonspecific substrate GUAA.
As terminator elements often occur in multiple repeats, we tested a longer substrate with three UCUU repeats. The substrate sequence was derived from the snR47 that is terminated by the Nrd1 pathway (Fig. 4F). Nab3 RRM binds the three UCUU-containing substrate with a Kd of 48 ± 2 μm, 1 order of magnitude stronger than we observed for a single UCUU motif (Fig. 4B). Furthermore, we assayed snR13, which is another naturally occurring Nrd1-dependent terminator that contains two UCUU and one CCU motifs (Fig. 4F). As expected, Nab3 RRM binds this substrate with a Kd of 46 ± 1 μm, a similar binding affinity to that of snR47 (Fig. 4B). As a control, we used SL RNA, which has a similar size and its sequence lacks UCUU or even CU recognition motifs (Fig. 4F). For this nonspecific substrate, Nab3 RRM binding is reduced more than 30-fold compared with the snR13 and snR47 terminators (Fig. 4, B and E).
Nrd1 RRM Binds the Nrd1 Termination Element with Low Affinity
As Nab3 RRM binds its termination motif with a low affinity, we therefore decided to investigate the RNA-binding properties of Nrd1 RRM, which is the second RNA-binding domain occurring in the Nrd1-dependent termination complex (Fig. 1B). Akin to Nab3 RRM, we assayed the binding affinity of Nrd1 RRM to RNA using FA. We found that Nrd1 RRM-(340–410) binds to the minimal termination RNA sequence GUAA (GUAR is known as the Nrd1-termination element (where R stands for purine) (18) with a low affinity in the mid-micromolar range (Fig. 4C; Kd of 66 ± 1 μm). Next, we assayed a longer RNA substrate, snR13, which contains two GUAR motifs (Fig. 4F). The titration curve for snR13 yielded a Kd of 11 ± 1 μm (Fig. 4C).
A Complex of Nrd1-Nab3 Binds RNA with High Affinity
As the Nrd1 and Nab3 proteins form a heterodimer in vivo and in vitro (19), we assayed RNA binding of the co-expressed Nrd1-Nab3 heterodimer (19) using FA. As a substrate, we used RNA derived from the snR13 terminator that contains two copies of each recognition element (Fig. 4F). This RNA substrate binds the Nrd1-Nab3 heterodimer with a Kd of 2.4 ± 0.3 nm (Fig. 4D), about 4–5 orders of magnitude stronger compared with the individual RRMs of Nrd1 and Nab3 (Fig. 4E). Another snR13-derived substrate, ½ snR13, containing only one copy of each recognition element (Fig. 4F), binds the Nrd1-Nab3 heterodimer with only a slightly lower affinity (Kd of 5.3 ± 0.6 nm). However, when one of the recognition elements (either Nrd1 or Nab3) is removed from the ½ snR13 substrate, the affinity is reduced such that the dissociation constant cannot be determined (saturation of the binding curve cannot not be reached due to the low solubility limit of the Nrd1-Nab3 heterodimer; data not shown).
Impacts of Nab3 RRM Point Mutations on RNA Binding Affinity
It has been shown previously that mutations of C2 and U3 in the UCUU motif reduce the binding affinity to Nab3 (18) or to the Nrd1-Nab3 heterodimer (19). We performed the converse experiments in which we assessed Nab3 RRM mutants for their ability to bind snR47 RNA in a quantitative solution binding assay by fluorescence anisotropy titration experiments. We mutated the non-canonical amino acid residues on the β-sheet surface (R331A, N361A, S399A, and E397A) that specifically recognize the bases of the U1C2U3 sequence (Fig. 3, B and C). These residues surround the conserved residues of RNP1 and RNP2 consensuses. We found that mutants R331A and S399A showed a 3–4-fold decrease in binding affinity of that demonstrated by the wild-type protein (Fig. 5A). In contrast, mutants N361A and E397A showed binding affinity similar to the wild-type Nab3 RRM (Fig. 5A).
FIGURE 5.
The important residues of Nab3 RRM that are required for RNA binding and cell viability. A, equilibrium binding of Nab3 RRM mutants with fluorescently labeled RNA monitored by fluorescence anisotropy. The Nab3 RRM R331A, N361A, S399A, and E397A mutants along with the wild-type of Nab3 RRM were titrated with fluorescently labeled snR47 substrate. Equilibrium association constants (Ka) are shown for individual mutants with S.E. B, residues Arg331, Ser399, and Asn361 are required for yeast viability. The indicated Nab3 RRM mutants were expressed episomally from pRS415 plasmids in the yeast strain with the endogenous NAB3 driven by the GAL1 promoter. Mutant strains were spotted on plates containing 2% glucose and a control galactose plate and incubated for 3 days at the indicated temperatures. Growth on glucose-containing plates leads to the repression of GAL1-driven wild-type Nab3, and thus shows the functionality of the different Nab3 mutants. Vector is a control where the GAL1::NAB3 strain contains an empty pRS415 plasmid, wt is the wild-type NAB3. C, expression of Nab3 proteins from pRS415 in glucose-containing medium. Western blot analysis was perfomed with protein extracts from the original GAL1::NAB3 strain (DLY889) grown in galactose-containing medium and extracts from DLY889 transformed with plasmids carrying wild-type and mutant NAB3 grown for 20 h in glucose-containing medium. Air2 was used as a loading control.
Functional Significance of the Nab3 RRM Residues That Contact RNA
To address the importance of the specific contacts identified in the Nab3 RRM-UCUU complex for Nab3 function in vivo, single amino acid mutants (R331A, S399A, S399K, E397A, and E397K) were prepared in a yeast expression vector and introduced into a yeast strain in which the endogenous NAB3 promoter was replaced with the GAL1 promoter (3). To test whether the mutated residues were essential for growth, the resulting transformants were spotted onto glucose containing plates. The shift to glucose represses the expression of the GAL1-driven endogenous NAB3, which completely impairs cell viability (Fig. 5B). This lethality was rescued by wild-type Nab3 (Fig. 5B). Mutating the three residues involved in the recognition of C2 and U3 (Fig. 3), R331A, N361A and S399A, or S399K, caused lethality (Fig. 5B), providing further support for the functional significance of these contacts. In contrary, the Glu397 mutant displayed only slow growth at 24 and 37 °C.
To further confirm that mutant lethality was not due to impaired protein expression, we performed Western blot analysis with antibodies directed against Nab3 and antibodies against the HA tag present on the endogenous Nab3 only. We detected similar levels of Nab3 in all proteins expressed episomally, whereas no HA-tagged endogenous Nab3 was detected in yeast grown on glucose medium (Fig. 5C).
DISCUSSION
RNA Recognition by Nab3 RRM and Its Comparison with Other RRMs
We have solved the structure of Nab3 RRM in free form and in complex with UCUU. In both forms, the fold of Nab3 RRM is very similar, with minor adjustments upon RNA binding, and resembles canonical RRM (21–23). However, it has shorter α-helices than observed in the structures of canonical RRMs, and an extra 310 helix between the α1 helix and β2 strand (22).
Nab3 RRM binds UCUU in a canonical manner as observed in other RRM-RNA complexes (21–23). Specifically, Nab3 RRM contains a well conserved signature of the RRM family, RNP1 and RNP2 sequences (44–46). Two conserved phenylalanine residues are used in RNP1 (Phe333) and RNP2 (Phe368) to mediate the stacking interaction with RNA bases C2 and U3, respectively. These aromatic residues are surrounded by basic and polar amino acid residues that mediate the sequence-specific recognition of C2U3 and in part of U1. The specifically recognized nucleotides are accommodated on the β-sheet only and neither loops nor N-/C-terminal regions to the RRM are involved in the recognition process.
Interestingly, the binding preference for CU has also been reported for the polypyrimidine tract-binding protein (PTB) (49). Unlike Nab3 RRM, PTB RRMs display additional topology elements to the canonical RRM involved in RNA recognition and significantly differ from RNP1 and RNP2 consensus (Fig. 6A). In particular, PTB RRMs lack aromatic residues in RNP1 and RNP2 that usually make extensive stacking interactions with the RNA bases and sugars as in the case of Nab3 RRM (Fig. 6). In addition, the structures of Nab3 and PTB RRMs revealed that these domains are sequentially unrelated on the entire RNA interaction surface, except for the last serine residue of β-strand 4 that is present in all PTB and Nab3 RRMs (Fig. 6A). The serine residue is involved in recognition of a cytosine in the structures of PTB (49) and Nab3 bound to RNA (Fig. 6). Furthermore, in the structures of PTB RRM1-RNA and Nab3 RRM-RNA, recognition of the 5′-end uridine is mediated in a similar way in which its imino proton is contacted by glutamine and glutamate, respectively (Fig. 6). In both structures, a cytidine could be tolerated instead of a uridine in this position. In contrast, recognition of the 3′-end uridine is mediated differently in these two structures. Whereas Nab3 RRM utilizes the side chains of arginine (in β1) and asparagine (in β2) to recognize the uridine, PTB RRM1 uses the main chain of leucine and lysine located in the C-terminal extension to the RRM to facilitate the uridine binding.
FIGURE 6.
Recognition of YCU by Nab3 and PTB RRMs. A, sequence alignment of PTB RRM1, -2, -3, and -4 and Nab3 RRM whose structures have been solved. The alignment was performed using ClustalW (53) and manually optimized using the three-dimensional structural information (49). For the RRMs of PTB, amino acids interacting with the RNA are shown in red boxes, residues in gray and black boxes are located in the β-sheet and residues in yellow and cyan boxes are in the α- or 310 helices, respectively (49). Residues in gray boxes form the hydrophobic core of the domains. For the RRM of Nab3, residues in red boxes are significantly perturbed upon RNA binding. B, comparison of Nab3 RRM (left: in yellow and red schematics) and PTB RRM1 (right: in cyan schematics) binding to UCU nucleotides (represented as a stick model). The protein residues that mediate the specific recognition are highlighted as a stick model. The C-terminal region of PTB RRM1 that mediates the recognition of U3 is shown in magenta.
The importance of the serine residue, which is discussed above, is demonstrated by our affinity measurements with the Nab3 RRM S399A mutant that decreases the binding affinity to its UCUU-containing substrate (Fig. 5A). Correspondingly, in vivo analyses of Nab3 S399A or S399K mutants show that the serine residue is essential for yeast viability (Fig. 5B). Similarly, the decreased affinity of the Nab3 RRM R331A mutant to the RNA is manifested by the lethal effect in yeast (Fig. 5). Interestingly, Asn361, which specifically recognizes U3 along with Arg331, shows lethality in vivo for the N361A mutant, whereas the affinity of this mutant is only slightly reduced compared with the wild-type (Fig. 5). Furthermore, the N361A mutant yeast strain shows accumulation of cryptic unstable transcripts when shifted to glucose,7 supporting the functional significance of this contact in the termination via the Nrd1 pathway. The Glu397 mutant does not have an impact on RNA binding in vitro and causes only slow growth at higher temperatures, indicating a minor role of this contact for the function of Nab3. Altogether, these functional data corroborate with our structural findings that Nab3 RRM specifically recognizes the YCU sequence.
Association between Nab3 and Nrd1 Increases the Affinity to the Termination Sequences
Our FA experiments showed that Nab3 RRM binds UCUU with the high micromolar range of equilibrium dissociation constant (Kd), yet the Nab3 RRM binds nonspecific four-nucleotide RNA with a Kd in the low millimolar range (Fig. 4). The observed affinity for the specific RNA substrate is weaker than the affinity usually observed for single canonical RRMs (21, 47). The affinity of Nab3 RRM binding to UCUU is lower by 2-fold compared with the affinity of PTB RRM1 to CUCU that also recognizes the YCU motif (Fig. 6) (47). Interestingly, the apparent Kd of Nab3 RRM binding to longer RNAs with multiple UCUU motifs (snR47 and snR13) are ∼50 μm, more than 1 order of magnitude stronger than we observed for a single UCUU motif (Fig. 4). The increased binding affinity likely originates from the presence of multiple binding sites and due to the presence of flanking sequences to the UCUU motif in snR47 and snR13 RNA substrates. The effect of flanking sequences is likely nonspecific, mediated by electrostatic interactions of additional phosphate groups. A similar increase in the binding affinity, when multiple binding motifs are present, has also been observed for the RRMs of PTB (47). The formation of multiple complexes between the snR47-derived RNA substrate and an RRM-containing Nab3 construct (Nab3-(277–565)) has also been observed previously using electrophoretic mobility shift assay (18). It has been also demonstrated that mutations in the 2nd, 3rd, and 4th position of the UCUU motif decrease the binding affinity to the Nab3-(277–565) (18). Furthermore, our data showed that Nrd1, the second RNA-binding subunit of the Nrd1 complex containing a single RRM, binds its termination element, GUAA, with a low affinity (Kd of 66 ± 1 μm; Fig. 4).
Therefore, it is very likely that Nrd1 and Nab3 bind RNA in a cooperative manner to achieve the nanomolar range of affinity to snR13 RNA, as previously estimated using electrophoretic mobility shift assay (EMSA) (19). Indeed, our FA measurements showed that the Nrd1-Nab3 heterodimer binds snR13 RNA with a Kd in the low nanomolar range, 4–5 orders of magnitude stronger compared with the individual RRMs of Nrd1 and Nab3 (Fig. 4E). The RRMs of Nrd1 and Nab3 do not bind each other (data not shown) and the regions that mediate the formation of the heterodimer are located near the N-terminal of each RRM (13).7 It is difficult to assay the binding affinity of full-length Nrd1 and Nab3 individually due to their instability (19); the Nrd1 alone rapidly aggregates.7 However, other regions outside of the RRMs of Nrd1 and Nab3 are not expected to contribute significantly to RNA binding as they do not contain an identifiable RNA-binding domain. In a similar way, the cooperative RNA binding of two RRM-containing proteins U2AF65 and U2AF35 is utilized to enhance the affinity and selectivity in the process of defining the site of spliceosomal assembly (50, 51).
Implication for Poly(A) Independent Transcription Termination
We have shown that the RRM of Nab3 binds specifically the YCU sequence (where Y stands for pyrimidine). This is in good agreement with previous functional data that led to the proposal of the UCUU sequence as the Nab3 termination element (6, 18). However, the first position of the UCUU motif is not fully conserved in some snRNA downstream sequences (18); it can be either U or C that perfectly matches our structural findings. Furthermore, the last position of UCUU is also not fully conserved but we cannot explain the recognition of another nucleotide due to the lack of experimental data for the recognition of this nucleotide in our structure. Considering the specificity only for the CU dinucleotide and relatively weak affinity of Nab3 RRM to YCU-like sequences, it is evident that Nab3 alone cannot recruit the Nrd1 complex to the correct termination sites. Indeed, we show the first quantitative evidence that the association of Nrd1 and Nab3 (each protein contains a single RRM) facilitates high affinity binding and sequence selectivity. It remains to be seen whether additional sequence elements are recognized, in addition to YCU and GUAR, upon the association of Nrd1-Nab3.
Acknowledgments
We thank to Dr. D. Libri for the GAL1::Nab3 strain and Dr. J. L. Corden for the generous gift of the Nab3 2F12 antibodies and the pST39 plasmid, containing Nab3-(191–565) and Nrd1-(1–548). We are grateful to Massimo Lucci for assistance with NMR measurements at CERM Florence, Italy. The NOESY spectra including the intermolecular filter experiments were obtained at the CERM NMR facility supported by European Union-NMR program Grant RII3–026145.
This work was supported in part by a Howard Hughes Medical Institute/European Molecular Biology Organization start-up grant, a Human Frontier Science Program Career Development Award, Ministry of Education of the Czech Republic Grants MSM0021622413, MSM0021622415, and Ingo LA08008, Czech Science Foundation Grants 204/08/1212 and 305/10/1490, Grant Agency of the Academy of Sciences of the Czech Republic IAA401630903, Wellcome Trust Grant 084316/Z/07/Z, and EMBO Installation Grant 1642.
The atomic coordinates and structure factors (codes 2KVI and 2L41) have been deposited in the Protein Data Bank, Research Collaboratory for Structural Bioinformatics, Rutgers University, New Brunswick, NJ (http://www.rcsb.org/).
F. Hobor, D. Hrossova, S. Vanacova, and R. Stefl, unpublished data.
- RNA Pol II
- RNA polymerase II
- FA
- fluorescence anisotropy
- Nab3
- nuclear polyadenylated RNA-binding 3
- Nrd1
- nuclear pre-mRNA down-regulation 1
- RRM
- RNA-recognition motif
- snoRNAs
- small nucleoar RNAs
- HSQC
- heteronuclear single quantum coherence
- RNP
- ribonucleoprotein
- TAMRA
- N,N,N′,N′-tetramethyl-6-carboxyrhodamine
- PTB
- polypyrimidine tract-binding protein.
REFERENCES
- 1. Richard P., Manley J. L. (2009) Genes Dev. 23, 1247–1269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Bentley D. (2002) Curr. Opin. Cell Biol. 14, 336–342 [DOI] [PubMed] [Google Scholar]
- 3. Thiebaut M., Kisseleva-Romanova E., Rougemaille M., Boulay J., Libri D. (2006) Mol. Cell 23, 853–864 [DOI] [PubMed] [Google Scholar]
- 4. Arigo J. T., Eyler D. E., Carroll K. L., Corden J. L. (2006) Mol. Cell 23, 841–851 [DOI] [PubMed] [Google Scholar]
- 5. Steinmetz E. J., Brow D. A. (1996) Mol. Cell. Biol. 16, 6993–7003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Steinmetz E. J., Conrad N. K., Brow D. A., Corden J. L. (2001) Nature 413, 327–331 [DOI] [PubMed] [Google Scholar]
- 7. Vasiljeva L., Buratowski S. (2006) Mol. Cell 21, 239–248 [DOI] [PubMed] [Google Scholar]
- 8. Mitchell P., Petfalski E., Shevchenko A., Mann M., Tollervey D. (1997) Cell 91, 457–466 [DOI] [PubMed] [Google Scholar]
- 9. Vanácová S., Wolf J., Martin G., Blank D., Dettwiler S., Friedlein A., Langen H., Keith G., Keller W. (2005) PLoS Biol. 3, e189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. LaCava J., Houseley J., Saveanu C., Petfalski E., Thompson E., Jacquier A., Tollervey D. (2005) Cell 121, 713–724 [DOI] [PubMed] [Google Scholar]
- 11. Wyers F., Rougemaille M., Badis G., Rousselle J. C., Dufour M. E., Boulay J., Régnault B., Devaux F., Namane A., Séraphin B., Libri D., Jacquier A. (2005) Cell 121, 725–737 [DOI] [PubMed] [Google Scholar]
- 12. Proudfoot N. J., Furger A., Dye M. J. (2002) Cell 108, 501–512 [DOI] [PubMed] [Google Scholar]
- 13. Vasiljeva L., Kim M., Mutschler H., Buratowski S., Meinhart A. (2008) Nat. Struct. Mol. Biol. 15, 795–804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Gudipati R. K., Villa T., Boulay J., Libri D. (2008) Nat. Struct. Mol. Biol. 15, 786–794 [DOI] [PubMed] [Google Scholar]
- 15. Steinmetz E. J., Brow D. A. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 6699–6704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Conrad N. K., Wilson S. M., Steinmetz E. J., Patturajan M., Brow D. A., Swanson M. S., Corden J. L. (2000) Genetics 154, 557–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Morlando M., Greco P., Dichtl B., Fatica A., Keller W., Bozzoni I. (2002) Mol. Cell Biol. 22, 1379–1389 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Carroll K. L., Pradhan D. A., Granek J. A., Clarke N. D., Corden J. L. (2004) Mol. Cell Biol. 24, 6241–6252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Carroll K. L., Ghirlando R., Ames J. M., Corden J. L. (2007) RNA 13, 361–373 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Venter J. C., Adams M. D., Myers E. W., Li P. W., Mural R. J., Sutton G. G., Smith H. O., Yandell M., Evans C. A., Holt R. A., Gocayne J. D., Amanatides P., Ballew R. M., Huson D. H., Wortman J. R., Zhang Q., Kodira C. D., Zheng X. H., Chen L., Skupski M., Subramanian G., Thomas P. D., Zhang J., Miklos G. L., Nelson C., Broder S., Clark A. G., Nadeau J., McKusick V. A., Zinder N., Levine A. J., Roberts R. J., Simon M., Slayman C., Hunkapiller M., Bolanos R., Delcher A., Dew I., Fasulo D., Flanigan M., Florea L., Halpern A., Hannenhalli S., Kravitz S., Levy S., Mobarry C., Reinert K., Remington K., Abu-Threideh J., Beasley E., Biddick K., Bonazzi V., Brandon R., Cargill M., Chandramouliswaran I., Charlab R., Chaturvedi K., Deng Z., Di Francesco V., Dunn P., Eilbeck K., Evangelista C., Gabrielian A. E., Gan W., Ge W., Gong F., Gu Z., Guan P., Heiman T. J., Higgins M. E., Ji R. R., Ke Z., Ketchum K. A., Lai Z., Lei Y., Li Z., Li J., Liang Y., Lin X., Lu F., Merkulov G. V., Milshina N., Moore H. M., Naik A. K., Narayan V. A., Neelam B., Nusskern D., Rusch D. B., Salzberg S., Shao W., Shue B., Sun J., Wang Z., Wang A., Wang X., Wang J., Wei M., Wides R., Xiao C., Yan C., Yao A., Ye J., Zhan M., Zhang W., Zhang H., Zhao Q., Zheng L., Zhong F., Zhong W., Zhu S., Zhao S., Gilbert D., Baumhueter S., Spier G., Carter C., Cravchik A., Woodage T., Ali F., An H., Awe A., Baldwin D., Baden H., Barnstead M., Barrow I., Beeson K., Busam D., Carver A., Center A., Cheng M. L., Curry L., Danaher S., Davenport L., Desilets R., Dietz S., Dodson K., Doup L., Ferriera S., Garg N., Gluecksmann A., Hart B., Haynes J., Haynes C., Heiner C., Hladun S., Hostin D., Houck J., Howland T., Ibegwam C., Johnson J., Kalush F., Kline L., Koduru S., Love A., Mann F., May D., McCawley S., McIntosh T., McMullen I., Moy M., Moy L., Murphy B., Nelson K., Pfannkoch C., Pratts E., Puri V., Qureshi H., Reardon M., Rodriguez R., Rogers Y. H., Romblad D., Ruhfel B., Scott R., Sitter C., Smallwood M., Stewart E., Strong R., Suh E., Thomas R., Tint N. N., Tse S., Vech C., Wang G., Wetter J., Williams S., Williams M., Windsor S., Winn-Deen E., Wolfe K., Zaveri J., Zaveri K., Abril J. F., Guigó R., Campbell M. J., Sjolander K. V., Karlak B., Kejariwal A., Mi H., Lazareva B., Hatton T., Narechania A., Diemer K., Muruganujan A., Guo N., Sato S., Bafna V., Istrail S., Lippert R., Schwartz R., Walenz B., Yooseph S., Allen D., Basu A., Baxendale J., Blick L., Caminha M., Carnes-Stine J., Caulk P., Chiang Y. H., Coyne M., Dahlke C., Mays A., Dombroski M., Donnelly M., Ely D., Esparham S., Fosler C., Gire H., Glanowski S., Glasser K., Glodek A., Gorokhov M., Graham K., Gropman B., Harris M., Heil J., Henderson S., Hoover J., Jennings D., Jordan C., Jordan J., Kasha J., Kagan L., Kraft C., Levitsky A., Lewis M., Liu X., Lopez J., Ma D., Majoros W., McDaniel J., Murphy S., Newman M., Nguyen T., Nguyen N., Nodell M., Pan S., Peck J., Peterson M., Rowe W., Sanders R., Scott J., Simpson M., Smith T., Sprague A., Stockwell T., Turner R., Venter E., Wang M., Wen M., Wu D., Wu M., Xia A., Zandieh A., Zhu X. (2001) Science 291, 1304–1351 [DOI] [PubMed] [Google Scholar]
- 21. Maris C., Dominguez C., Allain F. H. (2005) FEBS J. 272, 2118–2131 [DOI] [PubMed] [Google Scholar]
- 22. Stefl R., Skrisovska L., Allain F. H. (2005) EMBO Rep. 6, 33–38 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Cléry A., Blatter M., Allain F. H. (2008) Curr. Opin. Struct. Biol. 18, 290–298 [DOI] [PubMed] [Google Scholar]
- 24. Pergoli R., Kubicek K., Hobor F., Pasulka J., Stefl R. (2010) Biomol. NMR Assign. 4, 119–121 [DOI] [PubMed] [Google Scholar]
- 25. Bax A., Grzesiek S. (1993) Acc. Chem. Res. 26, 131–138 [Google Scholar]
- 26. Sattler M., Schleucher J., Griesinger C. (1999) Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 [Google Scholar]
- 27. Varani G., Aboulela F., Allain F. H. (1996) Prog. Nucl. Magn. Reson. Spectrosc. 29, 51–127 [Google Scholar]
- 28. Peterson R. D., Theimer C. A., Wu H., Feigon J. (2004) J. Biomol. NMR 28, 59–67 [DOI] [PubMed] [Google Scholar]
- 29. Zwahlen C., Legault P., Vincent S. J., Greenblatt J., Konrat R., Kay L. E. (1997) J. Am. Chem. Soc. 119, 6711–6721 [Google Scholar]
- 30. Güntert P. (2004) Methods Mol. Biol. 278, 353–378 [DOI] [PubMed] [Google Scholar]
- 31. Herrmann T., Güntert P., Wüthrich K. (2002) J. Mol. Biol. 319, 209–227 [DOI] [PubMed] [Google Scholar]
- 32. Shen Y., Delaglio F., Cornilescu G., Bax A. (2009) J. Biomol. NMR 44, 213–223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Case D. A., Darden T. A., Cheatham T. E., 3rd, Simmerling C. L., Wang J., Duke R. E., Luo R., Crowley M., Walker R. C., Zhang W., Merz K. M., Wang B., Hayik S., Roitberg A., Seabra G., Kolossváry I., Wong K. F., Paesani F., Vanicek J., Wu X., Brozell S. R., Steinbrecher T., Gohlke H., Yang L., Tan C., Mongan J., Hornak V., Cui G., Mathews D. H., Seetin M. G., Sagui C., Babin V., Kollman P. A. (2008) Amber 10, University of California, San Francisco, CA [Google Scholar]
- 34. Cornell W. D., Cieplak P., Bayly C. I., Gould I. R., Merz K. M., Ferguson D. M., Spellmeyer D. C., Fox T., Caldwell J. W., Kollman P. A. (1995) J. Am. Chem. Soc. 117, 5179–5197 [Google Scholar]
- 35. Padrta P., Stefl R., Králík L., Zídek L., Sklenár V. (2002) J. Biomol. NMR 24, 1–14 [DOI] [PubMed] [Google Scholar]
- 36. Laskowski R. A., Rullmannn J. A., MacArthur M. W., Kaptein R., Thornton J. M. (1996) J. Biomol. NMR 8, 477–486 [DOI] [PubMed] [Google Scholar]
- 37. Vriend G. (1990) J. Mol. Graph. 8, 52–56, 29 [DOI] [PubMed] [Google Scholar]
- 38. Koradi R., Billeter M., Wuthrich K. (1996) J. Mol. Graph. 14, 51–55, 29–32 [DOI] [PubMed] [Google Scholar]
- 39. Heyduk T., Lee J. C. (1990) Proc. Natl. Acad. Sci. U.S.A. 87, 1744–1748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Sikorski R. S., Hieter P. (1989) Genetics 122, 19–27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Wilson S. M., Datar K. V., Paddy M. R., Swedlow J. R., Swanson M. S. (1994) J. Cell Biol. 127, 1173–1184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. San Paolo S., Vanacova S., Schenk L., Scherrer T., Blank D., Keller W., Gerber A. P. (2009) PLoS Genet. 5, e1000555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Güntert P., Mumenthaler C., Wüthrich K. (1997) J. Mol. Biol. 273, 283–298 [DOI] [PubMed] [Google Scholar]
- 44. Swanson M. S., Nakagawa T. Y., LeVan K., Dreyfuss G. (1987) Mol. Cell. Biol. 7, 1731–1739 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Adam S. A., Nakagawa T., Swanson M. S., Woodruff T. K., Dreyfuss G. (1986) Mol. Cell. Biol. 6, 2932–2943 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Dreyfuss G., Swanson M. S., Piñol-Roma S. (1988) Trends Biochem. Sci. 13, 86–91 [DOI] [PubMed] [Google Scholar]
- 47. Auweter S. D., Oberstrass F. C., Allain F. H. (2007) J. Mol. Biol. 367, 174–186 [DOI] [PubMed] [Google Scholar]
- 48. Hargous Y., Hautbergue G. M., Tintaru A. M., Skrisovska L., Golovanov A. P., Stevenin J., Lian L. Y., Wilson S. A., Allain F. H. (2006) EMBO J. 25, 5126–5137 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Oberstrass F. C., Auweter S. D., Erat M., Hargous Y., Henning A., Wenter P., Reymond L., Amir-Ahmady B., Pitsch S., Black D. L., Allain F. H. (2005) Science 309, 2054–2057 [DOI] [PubMed] [Google Scholar]
- 50. Kielkopf C. L., Rodionova N. A., Green M. R., Burley S. K. (2001) Cell 106, 595–605 [DOI] [PubMed] [Google Scholar]
- 51. Singh R., Valcárcel J., Green M. R. (1995) Science 268, 1173–1176 [DOI] [PubMed] [Google Scholar]
- 52. Mulder F. A., Schipper D., Bott R., Boelens R. (1999) J. Mol. Biol. 292, 111–123 [DOI] [PubMed] [Google Scholar]
- 53. Chenna R., Sugawara H., Koike T., Lopez R., Gibson T. J., Higgins D. G., Thompson J. D. (2003) Nucleic Acids Res. 31, 3497–3500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Spronk C. A. E. M., Nabuurs S. B., Krieger E., Vriend G., Vuister G. W. (2004) Prog. Nucleic Magn. Res. Spectro. 45, 315–337 [Google Scholar]
- 55. Hooft R. W., Sander C., Vriend G. (1997) Comput. Appl. Biosci. 13, 425–430 [DOI] [PubMed] [Google Scholar]
- 56. Trantírek L., Stefl R., Masse J. E., Feigon J., Sklenár V. (2002) J. Biomol. NMR 23, 1–12 [DOI] [PubMed] [Google Scholar]
- 57. DeLano W. L. (2002) The PyMOL Molecular Graphics System, DeLano Scientific, Palo Alto, CA [Google Scholar]