Abstract
Background:
Identification of tumour-associated antigens (TAAs) that induce cytotoxic T lymphocytes (CTLs) specific to cancer cells is critical for the development of anticancer immunotherapy. In this study, we aimed at identifying a novel TAA of pancreatic cancer for immunotherapy.
Methods:
On the basis of the genome-wide cDNA microarray analysis, we focused on KIF20A (also known as RAB6KIFL/MKlp2) as a candidate TAA in pancreatic cancer cells. The HLA-A2 (A*02:01)-restricted CTL epitopes of KIF20A were identified using HLA-A2 transgenic mice (Tgm) and the peptides were examined to check whether they could generate human CTLs exhibiting cytotoxic responses against KIF20A+, HLA-A2+ tumour cells in vitro.
Results:
KIF20A was overexpressed in pancreatic cancer and in some other malignancies, but not in their non-cancerous counterparts and many normal adult tissues. We found that KIF20A-2 (p12–20, LLSDDDVVV), KIF20A-8 (p809–817, CIAEQYHTV), and KIF20A-28 (p284–293, AQPDTAPLPV) peptides could induce HLA-A2-restricted CTLs in HLA-A2 Tgm without causing autoimmunity. Peptide-reactive human CTLs were generated from peripheral blood mononuclear cells of HLA-A2+ healthy donors by in vitro stimulation with the three peptides, and those CTLs successfully exhibited cytotoxic responses to cancer cells expressing both KIF20A and HLA-A2.
Conclusion:
KIF20A is a novel promising candidate for anticancer immunotherapeutic target for pancreatic cancers.
Keywords: anticancer immunotherapy, tumour-associated antigen, CTL, KIF20A/RAB6KIFL/MKlp2, HLA-transgenic mouse
Pancreatic cancer is one of the highly lethal malignancies with an overall 5-year survival rate of ∼5% (Jemal et al, 2007). Although a surgical resection is the only treatment for long-term survival, patients with resectable pancreatic cancer are in the minority (9–22%) (Eloubeidi et al, 2006; Goonetilleke and Siriwardena, 2007). Furthermore, even the 5-year survival rate after a curative resection is reported to be ∼20% (Cleary et al, 2004; Smeenk et al, 2005; Moon et al, 2006). Therefore, there is a strong need for development of novel therapeutic modalities.
Anticancer immunotherapy is considered to be the candidate modality for pancreatic cancer. Recently, analyses of gene expression profiles of cancer and normal cells using cDNA microarray technologies have provided an effective approach for the identification of tumour-associated antigens (TAAs) (Nakatsura et al, 2004b; Uchida et al, 2004; Yoshitake et al, 2004; Watanabe et al, 2005; Komori et al, 2006; Suda et al, 2006; Harao et al, 2008; Imai et al, 2008). This study analysed the gene expression profiles of pancreatic cancer using a genome-wide cDNA microarray consisting of 27 648 genes, which revealed that KIF20A was overexpressed in pancreatic cancer tissues but not in many normal tissues.
In this study, we examined whether KIF20A could be a potential target for anticancer immunotherapy. To this aim, human KIF20A-derived and HLA-A2-restricted cytotoxic T lymphocyte (CTL) epitopes were identified using HLA-A2 transgenic mice (Tgm), and the ability of peptides to induce KIF20A-reactive human CTLs that kill cancer cells and the safety not to induce autoimmune responses in the mouse were investigated.
Materials and methods
cDNA microarray analysis
A data set of genome-wide cDNA microarray analyses using cancerous and adjacent normal tissues obtained by a laser microbeam dissection (Nakamura et al, 2004) was used in this study. The tissue samples were obtained from surgical specimens of pancreatic cancer patients. All patients provided their written informed consent to participate in this study.
Mice, cell lines, and HLA expression
The HLA-A2 Tgm, H-2Db−/− β2m−/− double knockout mice introduced with a monochain gene construct of human β2m-HLA-A2.1(α1, α2)-H-2Db(α3, transmembrane, and cytoplasmic) (Pascolo et al, 1997; Firat et al, 1999), were kindly provided by Dr FA Lemonnier. Mice were maintained and handled in accordance with the animal care guidelines of the Kumamoto University.
The human pancreatic cancer cell line PANC1, the colon cancer cell line CaCo-2, and a transporter associated with antigen processing (TAP)-deficient and HLA-A2 (A*02:01)-positive cell line T2 were purchased from the Riken Cell Bank (Tsukuba, Japan). The human liver cancer cell line SKHep1 and the human pancreatic cancer cell line PK9 were provided by Dr Kyogo Itoh (Kurume University, Kurume, Japan) and the Cell Resource Center for Biomedical Research Institute of Development, Aging and Cancer (Tohoku University, Sendai, Japan), respectively.
The expression of HLA-A2 was examined by flow cytometry with an anti-HLA-A2 monoclonal antibody (mAb), BB7.2 (One Lambda Inc., Canoga Park, CA, USA) to select HLA-A2-positive blood donors and target cancer cell lines.
Patients, blood samples, and tumour tissues
The clinical research using peripheral blood mononuclear cells (PBMCs) obtained from healthy donors was approved by the Institutional Review Board of the Kumamoto University. The cancer and adjacent non-cancerous tissues were obtained from 14 patients during routine diagnostic procedures from patients in the Kumamoto University Hospital. The tissues were subjected to either reverse transcription–PCR (RT–PCR) or immunohistochemical analyses as listed in Table 1. Blood and tissue samples were obtained from donors and patients, respectively, after receiving their written informed consent.
Table 1. Expression of the KIF20A gene or protein in pancreatic cancer tissues.
RT–PCR
Reverse transcription–PCR analyses were performed as described previously (Nakatsura et al, 2004a). The primers used were: KIF20A, sense 5′-CTACAAGCACCCAAGGACTCT-3′ (788-808, the 4th exon) and antisense 5′-AGATGGAGAAGCGAATGTTT-3′ (1400-1381, the 8th exon) and ACTB, sense 5′-CATCCACGAAACTACCTTCAACT-3′ (903-925, the 5th exon) and antisense 5′-TCTCCTTAGAGAGAAGTGGGGTG-3′ (1535-1513, the 6th exon). Primer positions are shown according to cDNA sequences presented in the Gene Bank Accession numbers of NM_005733 for human KIF20A and of NM_001101 for human ACTB. The products were 612 bp long for KIF20A and 632 bp long for ACTB. After normalisation by the intensity for ACTB mRNA, the expression levels of KIF20A mRNA were compared among the tissues and cell lines.
Western blot analysis and immunohistochemical examination
Western blot analysis and immunohistochemical staining of KIF20A using rabbit polyclonal anti-KIF20A antibody (category no. A300-879A) of Bethyl Laboratories (Montgomery, TX, USA) were performed as described previously (Nakatsura et al, 2001; Yoshitake et al, 2004). Immunohistochemical staining of CD4 or CD8 in tissue specimens of HLA-A2 Tgm immunised with the KIF20A-8809–817 peptide was performed as described previously (Matsuyoshi et al, 2004).
Lentiviral gene transfer
KIF20A cDNA was transduced into SKHep1 cells by a lentiviral vector-mediated gene transfer as described previously (Tahara-Hanaoka et al, 2002; Irie et al, 2006). The expression of KIF20A was confirmed by western blot analysis.
Induction and response of KIF20A-reactive mouse CTLs
A total of 36 human KIF20A-derived peptides (purity >95%) with predicted high binding scores for HLA-A2 (A*02:01) by BIMAS software (NIH, Bethesda, MD, USA; http://www-bimas.cit.nih.gov/) were synthesised by the American Peptide Company (Sunnyvale, CA, USA; Supplementary Table 1). The HLA-A2 Tgm were immunised intraperitoneally twice on days 0 and 7 with bone marrow-derived dendritic cells (BM-DCs) pulsed with 12 sets of a mixture of 3 kinds of the 36 synthesised peptides. Seven days after the last immunisation, CD4-depleted spleen cells (CD4− spleen cells) from Tgm using CD4 microbeads (Miltenyi Biotec, Auburn, CA, USA) were stimulated in vitro with BM-DCs pulsed with each peptide. The CTL responses to the peptides were tested by the ELISPOT assay (human INF-γ ELISPOT kit, BD Biosciences, Franklin Lakes, NJ, USA).
Induction of KIF20A-reactive human CTLs
Peripheral monocyte-derived DCs were generated from CD14+ cells isolated from PBMCs of HLA-A2-positive healthy donors using CD14 microbeads (Miltenyi Biotec), with stimulation of 100 ng ml−1 granulocyte/macrophage colony-stimulating factor, 10 ng ml−1 interleukin (IL)-4, and Streptococcal OK-432 (Picibanil, Chugai Pharmaceutical Co., Ltd., Tokyo, Japan) (Naito et al, 2006) as described previously (Harao et al, 2008). The cells were morphologically changed to express many dendrites, and their expression levels of MHC class-II and CD80 were upregulated after the stimulation with OK-432. The DCs were pulsed with 20 μg ml−1 candidate peptides in the presence of 4 μg ml−1 β2-microglobulin (Sigma-Aldrich, St Louis, MO, USA) for 2 h at 37°C in AIM-V medium (Invitrogen, Carlsbad, CA, USA) containing 2% autologous plasma. The DCs were then irradiated (40 Gy) and incubated with CD8+ cells isolated from the same PBMCs using CD8 microbeads (Miltenyi Biotec). These cultures were set up in 24-well plates; each well contained 1 × 105 peptide-pulsed DCs, 2 × 106 CD8+ T cells, and 5 ng ml−1 human IL-7 (Wako, Osaka, Japan), in 2 ml AIM-V with 2% autologous plasma. After 2 days, these cultures were supplemented with human IL-2 (PeproTec, Rocky Hill, NJ, USA) to a final concentration of 20 IU ml−1. Two additional stimulations with peptide-loaded autologous DCs were carried out on days 7 and 14. Six days after the last stimulation, antigen-specific CTL responses were investigated by the ELISPOT assay and the 51Cr release assay as described previously (Komori et al, 2006).
CTL responses against cancer cell lines
Human CTLs were co-cultured with cancer cells or peptide-pulsed T2 cells as a target (5 × 103 per well) at the indicated effector/target ratio, and a standard 51Cr release assay was performed (Komori et al, 2006). An HLA-A2 (A*02:01)-binding HIV-derived peptide, SLYNTVATL, was used as an irrelevant control peptide (Parker et al, 1992). Blocking of HLA-class I or HLA-DR in the ELISPOT assay was performed as follows: the target cancer cells were incubated with 10 μg ml−1 anti-HLA-class I mAb, W6/32, or 10 μg ml−1 anti-HLA-DR mAb, H-DR-1 (kindly provided by Dr Kyogo Itoh (Nakao et al, 1995)) for 1 h before co-culture with CTLs, and the inhibitory effects of mAbs on the production of IFN-γ by CTLs were monitored (Gomi et al, 1999).
Results
Identification of the KIF20A gene upregulated in pancreatic cancer and various malignancies based on cDNA microarray analyses
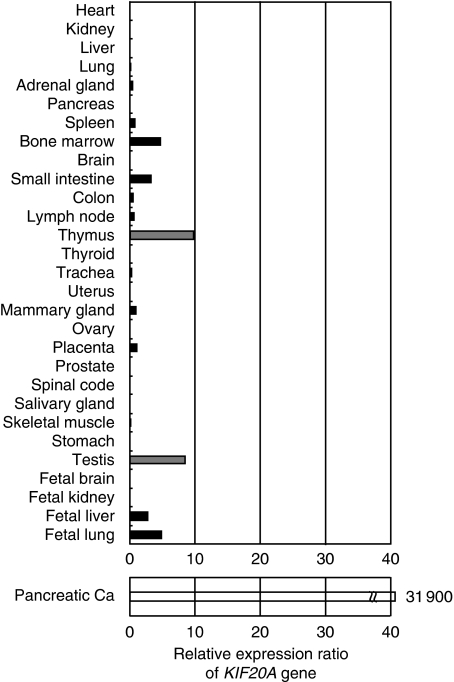
Using genome-wide cDNA microarray analyses, it turned out that six genes, namely CDH3, KIF20A, MICAL2, TRIM29, ABHD, and EPHA4, were overexpressed in the six pancreatic cancer tissues in comparison with their adjacent normal counterparts, and we reported that CDH3/P-cadherin is a new TAA for immunotherapy of pancreatic, gastric, and colorectal cancers (Imai et al, 2008). In this study, we focused on KIF20A as another pancreatic cancer-specific TAA for immunotherapeutic target. The cDNA microarray analyses revealed that expression of the KIF20A gene in pancreatic cancer tissues was markedly enhanced in all six patients investigated (the average of the relative expression ratio: 31 900, ranging 15–72 000), whereas the KIF20A gene was faintly expressed only in the testis and thymus among normal tissues (Figure 1). In addition, overexpression of the KIF20A gene was also observed in other malignancies, such as lung and bladder cancers (Table 2) (Kitahara et al, 2001; Hasegawa et al, 2002; Kikuchi et al, 2003; Nakamura et al, 2004; Obama et al, 2005) .
Figure 1.
The relative expression ratio of KIF20A mRNA in pancreatic cancer tissues (Pancreatic Ca) and in various normal tissues based on a cDNA microarray analysis. The KIF20A gene was overexpressed in all six pancreatic cancer tissues investigated, but barely expressed in many normal tissues, except for in the testis and thymus. The ratio of relative expression levels of KIF20A mRNA in six pancreatic cancer tissues to that of disease-free counterparts was calculated. For normal tissues, the average expression level of KIF20A mRNA in all tissues was assigned to be 1.0, and the relative expression level of KIF20A mRNA in each tissue was calculated. For pancreatic cancer, the expression levels of KIF20A mRNA in adjacent normal pancreatic tissues were assigned to be 1.0, and the relative expression levels of KIF20A mRNA in pancreatic cancers were calculated using specimens obtained from six patients with pancreatic cancer.
Table 2. Expression of the KIF20A gene in pancreatic cancer and various malignancies investigated by cDNA microarray analysesa.
| N | Positive ratea (%) | Average of relative expression ratio | |
|---|---|---|---|
| Pancreatic cancer | 6/6 | 100 | 31 900 |
| Small cell lung cancer | 15/15 | 100 | 22 |
| Bladder cancer | 30/31 | 97 | 20 500 |
| Non-small cell lung cancer | 20/22 | 91 | 25 800 |
| Cholangiocellular carcinoma | 7/11 | 64 | 3 780 |
| Breast cancer | 29/61 | 44 | 322 |
| Prostate cancer | 11/36 | 31 | 4 |
| Renal cell carcinoma | 3/11 | 27 | 5 |
| Oesophageal cancer | 2/13 | 15 | 3 |
| Colorectal cancer | 2/31 | 3 | 2 |
| Gastric cancer | 0/4 | 0 | 0 |
Data are obtained from our previous studies (Nakao et al, 1995; Uchida et al, 2004; Yoshitake et al, 2004; Taniuchi et al, 2005; Watanabe et al, 2005).
The relative expression ratio was calculated by dividing the value of the expression of KIF20A mRNA in cancer cells by that in the normal counterpart, and a relative expression ratio (cancer/normal tissue) of >5 was considered to be positive.
Expression of KIF20A mRNA and protein in normal organs, cancer cell lines, and pancreatic cancer tissues
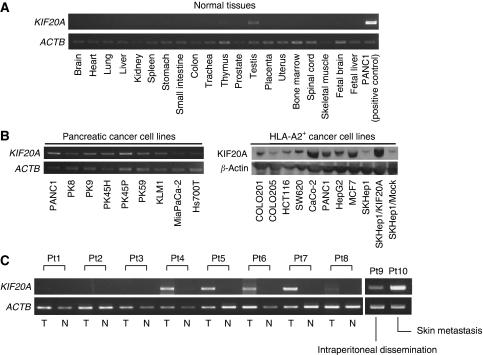
The expression of the KIF20A gene in normal tissues was analysed by RT–PCR analyses, which revealed its exclusive expression in the testis and thymus (Figure 2A). On the other hand, the expression of the KIF20A gene was detected in almost all pancreatic and other HLA-A2-positive cancer cell lines tested (Figure 2B, left). Those observations essentially coincided with data obtained by cDNA microarray analyses.
Figure 2.
Expression of KIF20A mRNA and protein in human normal tissues, cancer cell lines, and pancreatic cancer tissues. (A) RT–PCR analysis of KIF20A mRNA expression in various normal tissues. KIF20A mRNA was not detected except for faint expression observed in the testis. (B) The expression of KIF20A mRNA and protein in various cancer cell lines investigated by RT–PCR (left) and western blot analyses (right). KIF20A mRNA was detected in all pancreatic cancer cell lines examined. The major bands at ∼100 kDa, which corresponds to the calculated molecular weight for KIF20A, are shown. It must be noted that SKHep1 and SKHep1/Mock expressed only a trace amount of the KIF20A protein, whereas KIF20A cDNA-transfected SKHep1, SKHep1/KIF20A, relatively expressed a large amount of the protein. (C) RT–PCR analyses of the KIF20A expression in pancreatic cancer tissues (T), and their normal counterparts (N). The expression of the KIF20A mRNA was detected in five of eight pancreatic tumour tissues, but little expression was detected in their normal counterparts. It must be noted that KIF20A mRNA expression was also detected in the peritoneum (patient 9 (Pt9)) and metastatic foci of the skin (patient 10 (Pt10)).
We then checked the expression of the KIF20A gene in surgically resected pancreatic cancer tissues and their adjacent normal counterparts by RT–PCR analyses. The KIF20A gene was detected in five of eight pancreatic cancer tissues, whereas virtually no expression was observed in their normal counterparts (Figure 2C). It is noteworthy that KIF20A was also detected in the metastatic foci of the skin and peritoneum.
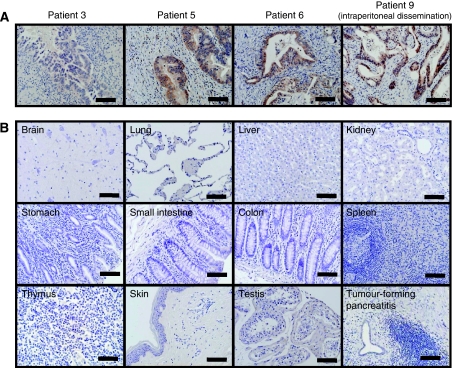
Western blot analysis revealed expression of the KIF20A protein in various HLA-A2+ cancer cell lines tested, except for SKHep1 (Figure 2B, right). To confirm the tumour-associated overexpression of the KIF20A protein, various paraffin-embedded normal tissue specimens and pancreatic cancer specimens were examined by immunohistochemical analyses. We investigated nine samples of pancreatic cancer (Table 1), and a strong staining of KIF20A was mainly observed in the cytoplasm and nuclei of cancer cells in six cases, whereas a very weak staining was observed in acinar cells and the normal ductal epithelium of their normal adjacent pancreatic tissues (Figure 3A). Little staining was detected in tumour-forming pancreatitis. KIF20A was not detected in the normal brain, lung, liver, kidney, stomach, small intestine, colon, spleen, skeletal muscle, skin, thymus, and skin (Figure 3B). Therefore, the pattern of KIF20A protein expression essentially paralleled with its gene expression profile.
Figure 3.
Immunohistochemical analyses of the KIF20A protein in (A) pancreatic cancer and in (B) various normal tissues. (Panel A) A strong staining of KIF20A was mainly observed at the cytoplasm and nuclei of cancer cells in six of nine cases (representative patients 5, 6, and 9 are shown), whereas a very weak staining was observed in acinar cells and in the normal ductal epithelium of their normal adjacent pancreatic tissues. A pancreatic cancer tissue without KIF20A expression is also shown (Patient 3). A similar strong staining was observed in the metastatic foci of the peritoneum (Patient 9). (Panel B) KIF20A was not detected in the normal brain, lung, liver, kidney, stomach, small intestine, colon, spleen, skeletal muscle, skin, thymus, and testis. Little staining was detected in tumour-forming pancreatitis. Positive staining signals are seen as brown. The scale bars represent 100 μm.
Identification of KIF20A-derived and HLA-A2-restricted mouse CTL epitopes using HLA-A2 Tgm
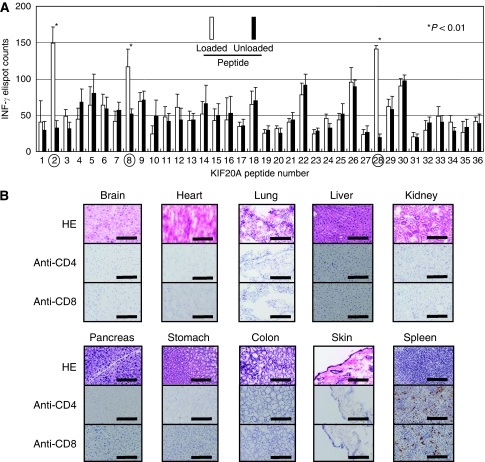
A total of 36 peptides with high binding scores to HLA-A2 (A*02:01) by BIMAS (Supplementary Table 1) were immunised to HLA-A2 Tgm, and CD4− spleen cells were stimulated in vitro with BM-DCs pulsed with each peptide. The CD4− spleen cells stimulated with KIF20A-2 (p12–20, LLSDDDVVV), KIF20A-8 (p809–817, CIAEQYHTV), and KIF20A-28 (p284–293, AQPDTAPLPV) peptides produced a significant amount of INF-γ in a peptide-specific manner in the ELISPOT assay (Figure 4A). Those CD4− spleen cells (2 × 104) showed 149.0±22.2, 117.2±23.4, and 141.2±5.5 spot counts per well in response to BM-DCs pulsed with the KIF20A-2, KIF20A-8, and KIF20A-28 peptides, respectively, whereas they showed 32.6±9.9, 51.4±7.8, and 19.2±5.2 spot counts per well, respectively, without peptide loading (P<0.01). No significant peptide-specific response was observed with the other peptides. These results suggest that the KIF20A-2, KIF20A-8, and KIF20A-28 peptides could be the HLA-A2-restricted CTL epitope peptides in HLA-A2 Tgm and those peptides were expected to be epitopes for human CTLs.
Figure 4.
Identification of HLA-A2-restricted mouse CTL epitopes of human KIF20A using HLA-A2 Tgm. (A) HLA-A2 Tgm were immunised with syngeneic BM-DCs pulsed with the peptide mixtures as described in the ‘Materials and Methods’ section, and CD4− spleen cells were stimulated with BM-DCs pulsed with or without each peptide for 6 days. INF-γ-producing CTLs were detected by an ELISPOT assay. KIF20A-2 (LLSDDDVVV), KIF20A-8 (CIAEQYHTV), and KIF20A-28 (AQPDTAPLPV) peptides indicated by the circles were shown to induce peptide-reactive CTLs. These assays were performed twice with similar results. (B) Immunohistochemical staining with anti-CD4 or anti-CD8 mAb in tissue specimens of HLA-A2 Tgm immunised with the KIF20A-8 peptide. After two-times vaccinations, these specimens were removed and analysed.
No autoimmune phenomenon induced by immunisation with KIF20A-8 in HLA-A2 Tgm
Whether immunisation with KIF20A peptides induces autoimmune reactions is of great importance. The immunohistochemical analyses of several vital organs with anti-CD4 and anti-CD8 mAbs were performed in HLA-A2 Tgm after two-times vaccination with the KIF20A-8 peptide, as its amino-acid sequence was identical between human and mouse KIF20A. As shown in Figure 4B, no pathological change that suggests autoimmunity, such as lymphocyte infiltration or tissue destruction, was observed, indicating that lymphocytes stimulated with the KIF20A-8 peptide were safe at least in HLA-A2 Tgm.
Induction of KIF20A-reactive human CTLs from PBMCs of HLA-A2-positive healthy donors
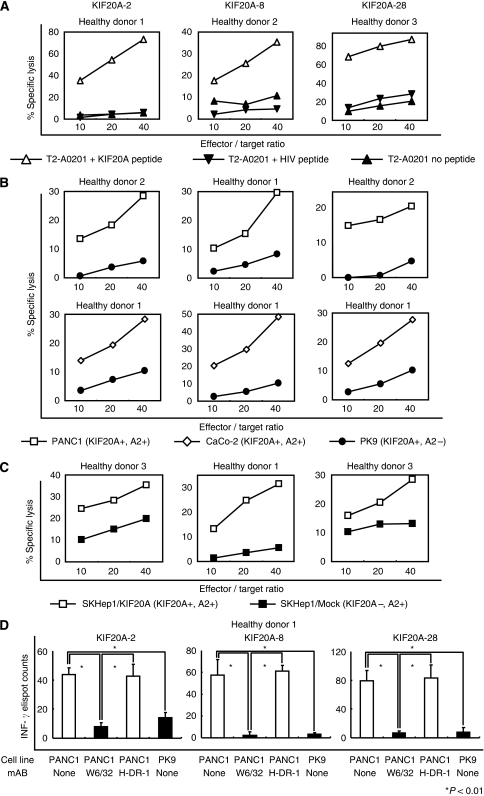
To investigate whether the KIF20A-2, KIF20A-8, and KIF20A-28 peptides could generate KIF20A-specific human CTLs, CD8+ T cells sorted from PBMCs of HLA-A2-positive healthy donors were incubated with the autologous CD14+ cell-derived DCs pulsed with each peptide. After three-times stimulations, the cytotoxic activities of CD8+ T cells against peptide-pulsed T2 cells were examined by the 51Cr release assay (Figure 5A). Each CTL killed the T2 cells pulsed with KIF20A-2 (left), KIF20A-8 (middle), or KIF20A-28 (right) peptides, but not the T2 cells pulsed with irrelevant HLA-A2-binding HIV peptide or those without peptide loading. The data indicate that KIF20A peptides successfully induced human CTLs with peptide-specific cytotoxicity.
Figure 5.
Induction of KIF20A-specific human CTLs from PBMCs of three HLA-A2-positive healthy donors. (A) KIF20A peptide-reactive CTLs generated from PBMCs of three HLA-A2-positive healthy donors effectively killed T2 cells (HLA-A2+, TAP deficient) pulsed with each peptide but not those unpulsed or pulsed with irrelevant and HLA-A2-restricted CTL epitopes of HIV. (B) Human CTLs exhibited cytotoxicity to the KIF20A+, HLA-A2+ human pancreatic cancer cell line PANC1 and to the colon cancer cell line CaCo-2, but not to KIF20A+, HLA-A2− human pancreatic cancer cell line PK9. (C) The cytotoxicity of human CTLs was KIF20A specific. Those CTLs killed SKHep1/KIF20A, but not mock-transfected SKHep1 cells. Representative data are shown. (D) Human CTL responses were inhibited by anti-HLA-class I mAb (W6/32, IgG2a) but not by anti-HLA-DR mAb (H-DR-1, IgG2a). The target cells used were PANC-1cell (KIF20A+, HLA-A2+) and PK9 cell (KIF20A+, HLA-A2−). (Panels A–D) Representative data from one of the three donors with similar results are shown.
Next, the capacity of these CTLs to kill human cancer cell lines expressing both KIF20A and HLA-A2 was examined. As shown in Figure 5B, KIF20A-reactive CTLs stimulated with KIF20A-2 (left), KIF20A-8 (middle), or KIF20A-28 (right) peptides exhibited effective cytotoxicity to PANC1 (KIF20A+, HLA-A2+) and CaCo-2 (KIF20A+, HLA-A2+), but not to PK9 (KIF20A+, HLA-A2−). Furthermore, all CTLs exhibited cytotoxicity against SKHep1/KIF20A (KIF20A+, HLA-A2+), SKHep1 cells (KIF20A−, HLA-A2+) transfected with the KIF20A gene (Figure 2B, right), but not against SKHep1/Mock, SKHep1 cells transfected with an empty vector (Figure 5C). Those results suggested that CTL responses were specific to KIF20A expression and that the epitope peptides were naturally processed and expressed on the surface of cancer cells in the context of HLA-A2 molecules.
Blocking mAb specific to HLA-class I (W6/32) markedly reduced the number of INF-γ-producing CTLs generated by stimulation with KIF20A-2 (left), KIF20A-8 (middle), or KIF20A-28 (right) peptides by co-culture with PANC1 cells with statistical significance (Figure 5D, P<0.01), whereas anti-HLA-DR mAb (H-DR-1) had no effect on CTL responses. The data verified that CTLs recognised KIF20A-expressing target cells in an HLA-class I-restricted manner.
Discussion
In this study, whether KIF20A is applicable to a target of anticancer immunotherapy was carefully investigated. The KIF20A gene was overexpressed in pancreatic cancer cells but barely expressed in their normal counterparts and in many normal adult tissues as revealed by the cDNA microarray analysis, RT–PCR analysis, and immunohistochemical analyses. As a weak expression of KIF20A in the normal thymus in addition to the testis was observed in the cDNA microarray and RT–PCR analyses, KIF20A is not a perfect cancer/testis antigen but a TAA overexpressed in pancreatic cancer, as well as in bladder cancer, non-small cell lung cancer, and cholangiocellular carcinoma. We have reported that CDH3/P-cadherin is another TAA that is overexpressed in pancreatic cancer (Imai et al, 2008); however, it has been often observed that using a single TAA as a target of anticancer immunotherapy does not yield satisfactory therapeutic outcomes in animal models. Using multiple TAAs in combination as targets drastically improved the outcome of anticancer immunotherapy (Fukushima et al, 2009), and therefore KIF20A could be not only a versatile tumour marker but also a TAA useful as a target of anticancer immunotherapy for pancreatic cancer by itself or in combination with other TAA(s). Furthermore, the involvement of KIF20A in pancreatic carcinogenesis suggests that KIF20A would be a promising immunotherapeutic target for pancreatic cancer as described below.
The KIF20A protein, also known as RAB6KIFL/MKlp2, was first identified to localise to Golgi apparatus and to have an important role in the dynamics in this organelle by an interaction with the GTP-bound form of Rab6 (Echard et al, 1998). KIF20A belongs to a large family of motor proteins that accumulate in mitotic cells, and microinjection of anti-KIF20A antibodies resulted in the failure of cytokinesis (Hill et al, 2000). The knockdown of KIF20A gene expression in pancreatic cancer cell lines by small-interfering RNA drastically inhibited the growth of those cells (Taniuchi et al, 2005).
The potential CTL target epitopes of KIF20A were listed among the HLA-A2 (A*02:01) binders predicted by the BIMAS software and surveyed using HLA-A2 Tgm. All three candidate epitope peptides identified could stimulate the generation of human CTLs that showed KIF20A-specific cytotoxicity in an HLA-A2-restricted manner in all three individuals examined. Importantly, immunisation of the KIF20A-8 peptide, of which the amino-acid sequence is conserved between human and mouse KIF20A, did not induce any signs of autoimmune phenomenon, suggesting the safety of this peptide for using anticancer immunotherapy targeting KIF20A. In this sense, the HLA-A2 is not only a powerful tool for rapid identification of HLA-A2-restricted CTL epitopes but also for evaluating the in vivo outcomes of immunotherapy using epitope peptides.
In conclusion, our data showed that KIF20A is a novel TAA and a potential target for anticancer immunotherapy for cancer cells expressing KIF20A at least in an HLA-A2-restricted situation. As KIF20A is highly expressed in a wider range of human malignancies, KIF20A is therefore a promising target for peptide-based immunotherapy for the treatment of malignancies, especially pancreatic cancer. Further investigation of the capability for induction of KIF20A-specific CTLs in pancreatic cancer patients thus remains an issue of great importance for clinical application.
Acknowledgments
We thank Dr Hideyuki Saya (Keio University, Tokyo, Japan), the Cell Resource Center for Biomedical Research Institute of Development, Aging and Cancer, Tohoku University, and the Health Science Research Resources Bank for providing the cell lines, and Dr Kyogo Itoh for providing the SKHep1 cell line and the anti-HLA-DR mAb, H-DR-1. We also thank Dr FA Lemonnier and Dr Hiroyuki Miyoshi (Riken BioResource Center) for providing HLA-A2.1 Tgm and the lentiviral vector, respectively. Grant-in-aids (nos. 17015035, 18014023 and 22133005) were received from the Ministry of Education, Culture, Sports, Science, and Technology, Japan, a Research Grant for Health Sciences from the Ministry of Health, Labor and Welfare, Japan; funding was from the Onco Therapy Science Co., The Sagawa Foundation for the Promotion of Cancer Research, and the Foundation for the Promotion of Cancer Research in Japan.
Footnotes
Supplementary Information accompanies the paper on British Journal of Cancer website (http://www.nature.com/bjc)
Supplementary Material
References
- Cleary SP, Gryfe R, Guindi M, Greig P, Smith L, Mackenzie R, Strasberg S, Hanna S, Taylor B, Langer B, Gallinger S (2004) Prognostic factors in resected pancreatic adenocarcinoma: analysis of actual 5-year survivors. J Am Coll Surg 198: 722–731 [DOI] [PubMed] [Google Scholar]
- Echard A, Jollivet F, Martinez O, Lacapere JJ, Rousselet A, Janoueix-Lerosey I, Goud B (1998) Interaction of a Golgi-associated kinesin-like protein with Rab6. Science 279: 580–585 [DOI] [PubMed] [Google Scholar]
- Eloubeidi MA, Desmond RA, Wilcox CM, Wilson RJ, Manchikalapati P, Fouad MM, Eltoum I, Vickers SM (2006) Prognostic factors for survival in pancreatic cancer: a population-based study. Am J Surg 192: 322–329 [DOI] [PubMed] [Google Scholar]
- Firat H, Garcia-Pons F, Tourdot S, Pascolo S, Scardino A, Garcia Z, Michel ML, Jack RW, Jung G, Kosmatopoulos K, Mateo L, Suhrbier A, Lemonnier FA, Langlade-Demoyen P (1999) H-2 class I knockout, HLA-A2.1-transgenic mice: a versatile animal model for preclinical evaluation of antitumor immunotherapeutic strategies. Eur J Immunol 29: 3112–3121 [DOI] [PubMed] [Google Scholar]
- Fukushima S, Hirata S, Motomura Y, Fukuma D, Matsunaga Y, Ikuta Y, Ikeda T, Kageshita T, Ihn H, Nishimura Y, Senju S (2009) Multiple antigen-targeted immunotherapy with alpha-galactosylceramide-loaded and genetically engineered dendritic cells derived from embryonic stem cells. J Immunother 32: 219–231 [DOI] [PubMed] [Google Scholar]
- Gomi S, Nakao M, Niiya F, Imamura Y, Kawano K, Nishizaka S, Hayashi A, Sobao Y, Oizumi K, Itoh K (1999) A cyclophilin B gene encodes antigenic epitopes recognized by HLA-A24-restricted and tumor-specific CTLs. J Immunol 163: 4994–5004 [PubMed] [Google Scholar]
- Goonetilleke KS, Siriwardena AK (2007) Nationwide questionnaire survey of the contemporary surgical management of pancreatic cancer in the United Kingdom & Ireland. Int J Surg 5: 147–151 [DOI] [PubMed] [Google Scholar]
- Harao M, Hirata S, Irie A, Senju S, Nakatsura T, Komori H, Ikuta Y, Yokomine K, Imai K, Inoue M, Harada K, Mori T, Tsunoda T, Nakatsuru S, Daigo Y, Nomori H, Nakamura Y, Baba H, Nishimura Y (2008) HLA-A2-restricted CTL epitopes of a novel lung cancer-associated cancer testis antigen, cell division cycle associated 1, can induce tumor-reactive CTL. Int J Cancer 123: 2616–2625 [DOI] [PubMed] [Google Scholar]
- Hasegawa S, Furukawa Y, Li M, Satoh S, Kato T, Watanabe T, Katagiri T, Tsunoda T, Yamaoka Y, Nakamura Y (2002) Genome-wide analysis of gene expression in intestinal-type gastric cancers using a complementary DNA microarray representing 23 040 genes. Cancer Res 62: 7012–7017 [PubMed] [Google Scholar]
- Hill E, Clarke M, Barr FA (2000) The Rab6-binding kinesin, Rab6-KIFL, is required for cytokinesis. EMBO J 19: 5711–5719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai K, Hirata S, Irie A, Senju S, Ikuta Y, Yokomine K, Harao M, Inoue M, Tsunoda T, Nakatsuru S, Nakagawa H, Nakamura Y, Baba H, Nishimura Y (2008) Identification of a novel tumor-associated antigen, cadherin 3/P-cadherin, as a possible target for immunotherapy of pancreatic, gastric, and colorectal cancers. Clin Cancer Res 14: 6487–6495 [DOI] [PubMed] [Google Scholar]
- Irie A, Harada K, Tukamoto H, Kim JR, Araki N, Nishimura Y (2006) Protein kinase D2 contributes to either IL-2 promoter regulation or induction of cell death upon TCR stimulation depending on its activity in Jurkat cells. Int Immunol 18: 1737–1747 [DOI] [PubMed] [Google Scholar]
- Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ (2007) Cancer statistics, 2007. CA Cancer J Clin 57: 43–66 [DOI] [PubMed] [Google Scholar]
- Kikuchi T, Daigo Y, Katagiri T, Tsunoda T, Okada K, Kakiuchi S, Zembutsu H, Furukawa Y, Kawamura M, Kobayashi K, Imai K, Nakamura Y (2003) Expression profiles of non-small cell lung cancers on cDNA microarrays: identification of genes for prediction of lymph-node metastasis and sensitivity to anti-cancer drugs. Oncogene 22: 2192–2205 [DOI] [PubMed] [Google Scholar]
- Kitahara O, Furukawa Y, Tanaka T, Kihara C, Ono K, Yanagawa R, Nita ME, Takagi T, Nakamura Y, Tsunoda T (2001) Alterations of gene expression during colorectal carcinogenesis revealed by cDNA microarrays after laser-capture microdissection of tumor tissues and normal epithelia. Cancer Res 61: 3544–3549 [PubMed] [Google Scholar]
- Komori H, Nakatsura T, Senju S, Yoshitake Y, Motomura Y, Ikuta Y, Fukuma D, Yokomine K, Harao M, Beppu T, Matsui M, Torigoe T, Sato N, Baba H, Nishimura Y (2006) Identification of HLA-A2- or HLA-A24-restricted CTL epitopes possibly useful for glypican-3-specific immunotherapy of hepatocellular carcinoma. Clin Cancer Res 12: 2689–2697 [DOI] [PubMed] [Google Scholar]
- Matsuyoshi H, Senju S, Hirata S, Yoshitake Y, Uemura Y, Nishimura Y (2004) Enhanced priming of antigen-specific CTLs in vivo by embryonic stem cell-derived dendritic cells expressing chemokine along with antigenic protein: application to antitumor vaccination. J Immunol 172: 776–786 [DOI] [PubMed] [Google Scholar]
- Moon HJ, An JY, Heo JS, Choi SH, Joh JW, Kim YI (2006) Predicting survival after surgical resection for pancreatic ductal adenocarcinoma. Pancreas 32: 37–43 [DOI] [PubMed] [Google Scholar]
- Naito K, Ueda Y, Itoh T, Fuji N, Shimizu K, Yano Y, Yamamoto Y, Imura K, Kohara J, Iwamoto A, Shiozaki A, Tamai H, Shimizu T, Mazda O, Yamagishi H (2006) Mature dendritic cells generated from patient-derived peripheral blood monocytes in one-step culture using streptococcal preparation OK-432 exert an enhanced antigen-presenting capacity. Int J Oncol 28: 1481–1489 [PubMed] [Google Scholar]
- Nakamura T, Furukawa Y, Nakagawa H, Tsunoda T, Ohigashi H, Murata K, Ishikawa O, Ohgaki K, Kashimura N, Miyamoto M, Hirano S, Kondo S, Katoh H, Nakamura Y, Katagiri T (2004) Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene 23: 2385–2400 [DOI] [PubMed] [Google Scholar]
- Nakao M, Yamana H, Imai Y, Toh Y, Toh U, Kimura A, Yanoma S, Kakegawa T, Itoh K (1995) HLA A2601-restricted CTLs recognize a peptide antigen expressed on squamous cell carcinoma. Cancer Res 55: 4248–4252 [PubMed] [Google Scholar]
- Nakatsura T, Kageshita T, Ito S, Wakamatsu K, Monji M, Ikuta Y, Senju S, Ono T, Nishimura Y (2004a) Identification of glypican-3 as a novel tumor marker for melanoma. Clin Cancer Res 10: 6612–6621 [DOI] [PubMed] [Google Scholar]
- Nakatsura T, Komori H, Kubo T, Yoshitake Y, Senju S, Katagiri T, Furukawa Y, Ogawa M, Nakamura Y, Nishimura Y (2004b) Mouse homologue of a novel human oncofetal antigen, glypican-3, evokes T-cell-mediated tumor rejection without autoimmune reactions in mice. Clin Cancer Res 10: 8630–8640 [DOI] [PubMed] [Google Scholar]
- Nakatsura T, Senju S, Yamada K, Jotsuka T, Ogawa M, Nishimura Y (2001) Gene cloning of immunogenic antigens overexpressed in pancreatic cancer. Biochem Biophys Res Commun 281: 936–944 [DOI] [PubMed] [Google Scholar]
- Obama K, Ura K, Li M, Katagiri T, Tsunoda T, Nomura A, Satoh S, Nakamura Y, Furukawa Y (2005) Genome-wide analysis of gene expression in human intrahepatic cholangiocarcinoma. Hepatology 41: 1339–1348 [DOI] [PubMed] [Google Scholar]
- Parker KC, Bednarek MA, Hull LK, Utz U, Cunningham B, Zweerink HJ, Biddison WE, Coligan JE (1992) Sequence motifs important for peptide binding to the human MHC class I molecule, HLA-A2. J Immunol 149: 3580–3587 [PubMed] [Google Scholar]
- Pascolo S, Bervas N, Ure JM, Smith AG, Lemonnier FA, Perarnau B (1997) HLA-A2.1-restricted education and cytolytic activity of CD8(+) T lymphocytes from beta2 microglobulin (beta2m) HLA-A2.1 monochain transgenic H-2Db beta2m double knockout mice. J Exp Med 185: 2043–2051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smeenk HG, Tran TC, Erdmann J, van Eijck CH, Jeekel J (2005) Survival after surgical management of pancreatic adenocarcinoma: does curative and radical surgery truly exist? Langenbecks Arch Surg 390: 94–103 [DOI] [PubMed] [Google Scholar]
- Suda T, Tsunoda T, Uchida N, Watanabe T, Hasegawa S, Satoh S, Ohgi S, Furukawa Y, Nakamura Y, Tahara H (2006) Identification of secernin 1 as a novel immunotherapy target for gastric cancer using the expression profiles of cDNA microarray. Cancer Sci 97: 411–419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tahara-Hanaoka S, Sudo K, Ema H, Miyoshi H, Nakauchi H (2002) Lentiviral vector-mediated transduction of murine CD34(−) hematopoietic stem cells. Exp Hematol 30: 11–17 [DOI] [PubMed] [Google Scholar]
- Taniuchi K, Nakagawa H, Nakamura T, Eguchi H, Ohigashi H, Ishikawa O, Katagiri T, Nakamura Y (2005) Down-regulation of RAB6KIFL/KIF20A, a kinesin involved with membrane trafficking of discs large homologue 5, can attenuate growth of pancreatic cancer cell. Cancer Res 65: 105–112 [PubMed] [Google Scholar]
- Uchida N, Tsunoda T, Wada S, Furukawa Y, Nakamura Y, Tahara H (2004) Ring finger protein 43 as a new target for cancer immunotherapy. Clin Cancer Res 10: 8577–8586 [DOI] [PubMed] [Google Scholar]
- Watanabe T, Suda T, Tsunoda T, Uchida N, Ura K, Kato T, Hasegawa S, Satoh S, Ohgi S, Tahara H, Furukawa Y, Nakamura Y (2005) Identification of immunoglobulin superfamily 11 (IGSF11) as a novel target for cancer immunotherapy of gastrointestinal and hepatocellular carcinomas. Cancer Sci 96: 498–506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshitake Y, Nakatsura T, Monji M, Senju S, Matsuyoshi H, Tsukamoto H, Hosaka S, Komori H, Fukuma D, Ikuta Y, Katagiri T, Furukawa Y, Ito H, Shinohara M, Nakamura Y, Nishimura Y (2004) Proliferation potential-related protein, an ideal esophageal cancer antigen for immunotherapy, identified using complementary DNA microarray analysis. Clin Cancer Res 10: 6437–6448 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.