Abstract
Sebaldella termitidis (Sebald 1962) Collins and Shah 1986, is the only species in the genus Sebaldella within the fusobacterial family ‘Leptotrichiaceae’. The sole and type strain of the species was first isolated about 50 years ago from intestinal content of Mediterranean termites. The species is of interest for its very isolated phylogenetic position within the phylum Fusobacteria in the tree of life, with no other species sharing more than 90% 16S rRNA sequence similarity. The 4,486,650 bp long genome with its 4,210 protein-coding and 54 RNA genes is part of the Genomic Encyclopedia of Bacteria and Archaea project.
Keywords: anaerobic, mesophile, nonmotile, non-sporeforming, Gram-negative, termite intestine, ‘Fusobacteria’, ‘Leptotrichiaceae’, GEBA
Introduction
Strain NCTC 11300T (= ATCC 33386TM = NCTC 11300) is the type strain of the species Sebaldella termitidis [1]. The strain was first isolated from posterior intestinal content of Reticulitermes lucifugus (Mediterranean termites) by the French microbiologist Madeleine Sebald [1,2], and was initially classified as Bacteroides termitidis [3]. The unusually low G+C content, as well as biochemical features which did not correspond to those known for the other members of the genus Bacteroides [4], and the subsequently described novel 16S rRNA sequences [5] made the position of B. termitidis within the genus Bacteroides appear controversial, and guided Collins and Shah in 1986 to reclassify B. termitidis as the type strain of the novel genus Sebaldella [1]. Here we present a summary classification and a set of features for S. termitidis NCTC 11300T, together with the description of the complete genomic sequencing and annotation.
Classification and features
NCTC 11300T represents an isolated species, with no other cultivated strain known in the literature belonging to the species. An uncultured clone with identical 16S rRNA sequence was identified in a mesophilic anaerobic digester that treats municipal wastewater sludge in Clos de Hilde, France [6], and another uncultured clone, PCD-1 (96.1% 16S rRNA sequence identity), was reported from the digestive tract of the ground beetle Poecilus chalcites [7]. The closest related type strains are those of the genus Leptotrichia, which share 85.9 to 89.96% 16S rRNA sequence similarity [8]. Neither environmental screenings nor metagenomic surveys provided any 16S rRNA sequence with significant sequence similarity to NCTC 11300T, indicating that members of the species S. termitidis and the genus Sebaldella are not very frequent in the environment (status February 2010).
Figure 1 shows the phylogenetic neighborhood of S. termitidis NCTC 11300T in a 16S rRNA based tree. The sequences of the four identical copies of the 16S rRNA gene in the genome do not differ from the previously published 16S rRNA sequence generated from ATCC 3386 (M58678), which is missing two nucleotides and contains 30 ambiguous base calls.
Figure 1.
Phylogenetic tree highlighting the position of S. termitidis NCTC 11300T relative to the other type strains within the family ‘Leptotrichiaceae’. The tree was inferred from 1,422 aligned characters [9,10] of the 16S rRNA gene sequence under the maximum likelihood criterion [11] and rooted in accordance with the current taxonomy. The branches are scaled in terms of the expected number of substitutions per site. Numbers above branches are support values from 1,000 bootstrap replicates if larger than 60%. Lineages with type strain genome sequencing projects registered in GOLD [12] are shown in blue, published genomes in bold, e.g. the recently published GEBA genomes from Leptotrichia buccalis [13], and Streptobacillus moniliformis [14].
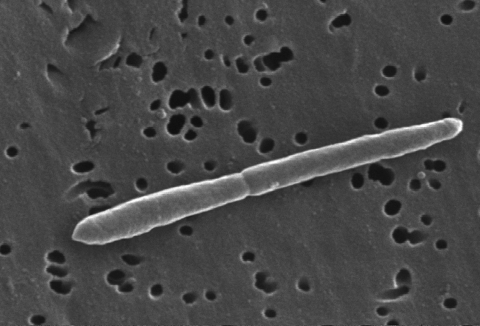
Cells of strain NCTC 11300T are Gram-negative, obligately anaerobic, nonmotile, nonspore-forming rods of 0.3 to 0.5 x 2 to 12 μm with central swellings (Figure 2 and Table 1) [1]. Cells occur singly, in pairs, as well as in filaments [1]. Colonies on surface are transparent to opaque, circular measuring 1-2 mm in diameter, whereas colonies in deep agar are non pigmented and lenticular [1].
Figure 2.
Scanning electron micrograph of S. termitidis NCTC 11300T. (J. Carr, CDC, Atlanta, Georgia). More EM photos of the organism can be found at http://phil.cdc.gov/phil.
Table 1. Classification and general features of S. termitidis NCTC 11300T according to the MIGS recommendations [15].
| MIGS ID | Property | Term | Evidence code |
|---|---|---|---|
| Current classification | Domain Bacteria | TAS [16] | |
| Phylum Fusobacteria | TAS [17] | ||
| Class ‘Fusobacteria’ | TAS [17] | ||
| Order ‘Fusobacteriales’ | TAS [17] | ||
| Family ‘Leptotrichiaceae’ | TAS [18] | ||
| Genus Sebaldella | TAS [1,19] | ||
| Species Sebaldella termitidis | TAS [1,19] | ||
| Type strain NCTC 11300 | TAS [1] | ||
| Gram stain | Gram negative | TAS [1] | |
| Cell shape | rod-shaped, with central swellings; occur singly, in pairs and in filaments |
TAS [1] | |
| Motility | nonmotile | TAS [1] | |
| Sporulation | nonsporulating | TAS [2] | |
| Temperature range | mesophile | NAS | |
| Optimum temperature | not determined | ||
| Salinity | not reported | ||
| MIGS-22 | Oxygen requirement | obligate anaerobic | TAS [1] |
| Carbon source | glucose and other sugars | TAS [1] | |
| Energy source | fermentation of glucose and other sugars | TAS [1] | |
| MIGS-6 | Habitat | bacterial flora of termite gastrointestinal tract |
TAS [1] |
| MIGS-15 | Biotic relationship | unknown | |
| MIGS-14 | Pathogenicity | none reported | NAS |
| Biosafety level | 2 | TAS [20] | |
| Isolation | posterior intestinal content of termites | TAS [2] | |
| MIGS-4 | Geographic location | unknown | |
| MIGS-5 | Sample collection time | 1962 or before | TAS [1,2] |
| MIGS-4.1 MIGS-4.2 |
Latitude Longitude |
not reported | |
| MIGS-4.3 | Depth | not reported | |
| MIGS-4.4 | Altitude | not reported |
Evidence codes - IDA: Inferred from Direct Assay (first time in publication); TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from of the Gene Ontology project [21]. If the evidence code is IDA, then the property was directly observed for a live isolate by one of the authors or an expert mentioned in the acknowledgements.
The major end products of the glucose metabolism by strain NCTC 11300T are acetic and lactic acids (with some formic acid) as opposed to succinic and acetic acids dominating in members of the genus Bacteroides [1]. Enzymes of the hexose-monophosphate-shunt are missing, while present in members of the genus Bacteroides [1,4]. A list of additional sugars and alcohols used or not-used for fermentation is provided by Collins and Shah [1].
Chemotaxonomy
The cell wall structure of strain NCTC 11300T has not yet been reported. Nonhydroxylated and 3-hydroxyated fatty acids were present [1]. The major long chain fatty acids are saturated and monounsaturated straight chain acids: C16:0 (37%) and C18:1 (41%), with methyl branched acids being absent [1], as opposed to straight-chain saturated, anteiso- and iso-methyl branched-chain acids in members of the genus Bacteroides, which are missing the monounsaturated acids [1]. Menaquinones were not detected, as opposed to members of the genus Bacteroides [1].
Genome sequencing and annotation
Genome project history
This organism was selected for sequencing on the basis of its phylogenetic position, and is part of the Genomic Encyclopedia of Bacteria and Archaea project [22]. The genome project is deposited in the Genome OnLine Database [12] and the complete genome sequence is deposited in GenBank. Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute (JGI). A summary of the project information is shown in Table 2.
Table 2. Genome sequencing project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | Finished |
| MIGS-28 | Libraries used | One genomic 8kb pMCL200 library, one 454 pyrosequence library and one Illumina library |
| MIGS-29 | Sequencing platforms | Sanger, 454 Titanium, Illumina |
| MIGS-31.2 | Sequencing coverage | 9.2× Sanger; 30.3× 454 Titanium |
| MIGS-30 | Assemblers | Newbler, phrap |
| MIGS-32 | Gene calling method | Prodigal, GenePRIMP |
| INSDC ID | CP001739 (chromosome), CP001740, CP001741 (plasmids) |
|
| Genbank Date of Release | November 19, 2009 | |
| GOLD ID | Gc01144 | |
| NCBI project ID | 29539 | |
| Database: IMG-GEBA | 2501846314 | |
| MIGS-13 | Source material identifier | ATCC 33386 |
| Project relevance | Tree of Life, GEBA |
Growth conditions and DNA isolation
S. termitidis NCTC 11300T, ATCC 33386TM, was grown anaerobically in ATCC medium 1490 (Modified chopped meat medium) [23] at 37°C. DNA was isolated from cell paste using a basic CTAB extraction and then quality controlled according to JGI guidelines.
Genome sequencing and assembly
The genome was sequenced using a combination of Sanger and 454 sequencing platforms. All general aspects of library construction and sequencing can be found at http://www.jgi.doe.gov/. 454 Pyrosequencing reads were assembled using the Newbler assembler version 1.1.02.15 (Roche). Large Newbler contigs were broken into 4,966 overlapping fragments of 1,000 bp and entered into assembly as pseudo-reads. The sequences were assigned quality scores based on Newbler consensus q-scores with modifications to account for overlap redundancy and to adjust inflated q-scores. A hybrid 454/Sanger assembly was made using the parallel phrap assembler (High Performance Software, LLC). Possible mis-assemblies were corrected with Dupfinisher [24] or transposon bombing of bridging clones (Epicentre Biotechnologies, Madison, WI). Gaps between contigs were closed by editing in Consed, custom primer walk or PCR amplification. A total of 796 Sanger finishing reads were produced to close gaps, to resolve repetitive regions, and to raise the quality of the finished sequence. Illumina reads were used to improve the final consensus quality using an in-house developed tool (the Polisher, unpublished). The error rate of the completed genome sequence is less than 1 in 100,000. Together all sequence types provided 39.5× coverage of the genome. The final assembly contains 45,934 Sanger and 760,187 pyrosequence reads.
Genome annotation
Genes were identified using Prodigal [25] as part of the Oak Ridge National Laboratory genome annotation pipeline, followed by a round of manual curation using the JGI GenePRIMP pipeline [26]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) nonredundant database, UniProt, TIGRFam, Pfam, PRIAM, KEGG, COG, and InterPro databases. Additional gene prediction analysis and manual functional annotation was performed within the Integrated Microbial Genomes Expert Review (IMG-ER) platform [27].
Genome properties
The genome consists of a 4,418,842 bp long chromosome, and two plasmids with 54,160 bp and 13,648 bp length, respectively, with a 33.4% GC content (Table 3 and Figure 3). Of the 4,264 genes predicted, 4,210 were protein-coding genes, and 54 RNAs; 59 pseudogenes were identified. The majority of the protein-coding genes (60.4%) were assigned with a putative function while those remaining were annotated as hypothetical proteins. The distribution of genes into COGs functional categories is presented in Table 4.
Table 3. Genome Statistics.
| Attribute | Value | % of Total |
|---|---|---|
| Genome size (bp) | 4,486,650 | 100.00% |
| DNA coding region (bp) | 3,918,335 | 87.33% |
| DNA G+C content (bp) | 1,497,450 | 33.38% |
| Number of replicons | 3 | |
| Extrachromosomal elements | 2 | |
| Total genes | 4,264 | 100.00% |
| RNA genes | 54 | 1.27% |
| rRNA operons | 4 | |
| Protein-coding genes | 4,210 | 98.73% |
| Pseudogenes | 59 | 1.38% |
| Genes with function prediction | 2,576 | 60.41% |
| Genes in paralog clusters | 1,253 | 29.39% |
| Genes assigned to COGs | 2,299 | 60.95% |
| Genes assigned Pfam domains | 2,787 | 65.36% |
| Genes with signal peptides | 801 | 18.79% |
| Genes with transmembrane helices | 901 | 21.13% |
| CRISPR repeats | 1 |
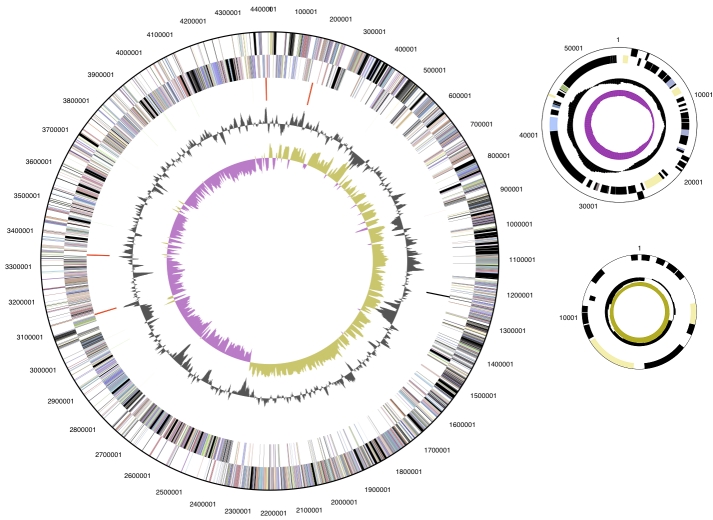
Figure 3.
Graphical circular maps of the chromosome and the two plasmids. From outside to the center: Genes on forward strand (color by COG categories), Genes on reverse strand (color by COG categories), RNA genes (tRNAs green, rRNAs red, other RNAs black), GC content, GC skew.
Table 4. Number of genes associated with the general COG functional categories.
| Code | value | %age | Description |
|---|---|---|---|
| J | 152 | 3.6 | Translation, ribosomal structure and biogenesis |
| A | 0 | 0.0 | RNA processing and modification |
| K | 265 | 6.3 | Transcription |
| L | 130 | 3.1 | Replication, recombination and repair |
| B | 0 | 0.0 | Chromatin structure and dynamics |
| D | 22 | 0.5 | Cell cycle control, cell division, chromosome partitioning |
| Y | 0 | 0.0 | Nuclear structure |
| V | 47 | 1.1 | Defense mechanisms |
| T | 96 | 2.3 | Signal transduction mechanisms |
| M | 155 | 3.7 | Cell wall/membrane biogenesis |
| N | 17 | 0.4 | Cell motility |
| Z | 0 | 0.0 | Cytoskeleton |
| W | 0 | 0.0 | Extracellular structures |
| U | 41 | 1.0 | Intracellular trafficking, secretion and vesicular transport |
| O | 71 | 1.7 | Posttranslational modification, protein turnover, chaperones |
| C | 128 | 3.0 | Energy production and conversion |
| G | 468 | 11.1 | Carbohydrate transport and metabolism |
| E | 219 | 5.2 | Amino acid transport and metabolism |
| F | 93 | 2.2 | Nucleotide transport and metabolism |
| H | 106 | 2.5 | Coenzyme transport and metabolism |
| I | 59 | 1.4 | Lipid transport and metabolism |
| P | 105 | 2.5 | Inorganic ion transport and metabolism |
| Q | 32 | 0.8 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 403 | 9.6 | General function prediction only |
| S | 241 | 5.7 | Function unknown |
| - | 1,665 | 39.5 | Not in COGs |
Acknowledgements
We would like to gratefully acknowledge the help of Janice Carr (Centers of Disease Control, Atlanta, Georgia) for providing the EM photo of S. thermitidis NCTC 11300T. This work was performed under the auspices of the US Department of Energy's Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, Los Alamos National Laboratory under contract No. DE-AC02-06NA25396, and Oak Ridge National Laboratory under contract DE-AC05-00OR22725
References
- 1.Collins MD, Shah HN. Reclassification of Bacteroides termitidis Sebald (Holdeman and Moore) in a new genus Sebaldella termitidis comb. nov. Int J Syst Bacteriol 1986; 36:349-350 10.1099/00207713-36-2-349 [DOI] [Google Scholar]
- 2.Sebald M. Etude sur les bacteries anaérobes gram-négatives asporulées. Thèse de l'Université Paris. Imprimerie Barnéoud S.A. Laval, France, 1962. [Google Scholar]
- 3.Holdeman LV, Kelly RW, Moore WEC. Genus Bacteroides, p. 604-631. In: Krieg NR, Holt JG (eds) Bergey's manual of systematic bacteriology, Vol. 1. The Williams & Wilkins Co., Baltimore. 1984 [Google Scholar]
- 4.Shah HN, Collins MD. Genus Bacteroides: a chemotaxonomical perspective. J Appl Bacteriol 1983; 55:403-416 [DOI] [PubMed] [Google Scholar]
- 5.Paster BJ, Ludwig W, Weisburg WG, Stackebrandt E, Hespell RB, Hahn CM, Reichenbach H, Stetter KO, Woese CR. A phylogenetic grouping of the Bacteroides, Cytophagas, and certain Flavobacteria. Syst Appl Microbiol 1985; 6:34-42 [Google Scholar]
- 6.Rivière D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J, Li T, Camacho P, Sghir A. Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 2009; 3:700-714 10.1038/ismej.2009.2 [DOI] [PubMed] [Google Scholar]
- 7.Lehman RM, Lundgren JG, Petzke LM. Bacterial communities associated with the digestive tract of the predatory ground beetle, Poecilus chalcites, and their modifications by laboratory rearing ans antibiotic treatment. Microb Ecol 2009; 57:349-358 10.1007/s00248-008-9415-6 [DOI] [PubMed] [Google Scholar]
- 8.Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW. EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 2007; 57:2259-2261 10.1099/ijs.0.64915-0 [DOI] [PubMed] [Google Scholar]
- 9.Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 2000; 17:540-552 [DOI] [PubMed] [Google Scholar]
- 10.Lee C, Grasso C, Sharlow MF. Multiple sequence alignment using partial order graphs. Bioinformatics 2002; 18:452-464 10.1093/bioinformatics/18.3.452 [DOI] [PubMed] [Google Scholar]
- 11.Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 2008; 57:758-771 10.1080/10635150802429642 [DOI] [PubMed] [Google Scholar]
- 12.Liolios K, Chen IM, Mavromatis K, Tavernarakis N, Hugenholtz P, Markowitz VM, Kyrpides NC. The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 2010; 38:D346-D354 10.1093/nar/gkp848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ivanova N, Gronow S, Lapidus A, Copeland A, Glavina Del Rio T, Nolan M, Lucas S, Cheng F, Tice H, Cheng JF, et al. Leptotrichia buccalis type strain (C-1013-bT). Stand Genomic Sci 2009; 1:126-132 10.4056/sigs.1854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Nolan M, Gronow S, Lapidus A, Ivanova N, Copeland A, Lucas S, Glavina Del Rio T, Chen F, Tice H, Pitluck S, et al. Streptobacillus moniliformis type strain (9901T). Stand Genomic Sci 2009; 1:300-397 10.4056/sigs.48727 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Garrity GM, Holt JG. Taxonomic Outline of the Archaea and Bacteria. In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 155-166. [Google Scholar]
- 18.Ludwig W, Euzeby J, Whitman WG. Draft taxonomic outline of the Bacteroidetes, Planctomycetes, Chlamydiae, Spirochaetes, Fibrobacteres, Fusobacteria, Acidobacteria, Verrucomicrobia, Dictyoglomi, and Gemmatimonadetes http://www.bergeys.org/outlines/Bergeys_Vol_4_Outline.pdf Taxonomic Outline 2008.
- 19.Skerman VBD, McGowan V, Sneath PHA. Approved Lists of Bacterial Names. Int J Syst Bacteriol 1980; 30:225-420 10.1099/00207713-30-1-225 [DOI] [PubMed] [Google Scholar]
- 20.CDC's Office of Health and Safety www.cdc.gov/od/ohs/biosfty/bmbl5/bmbl5toc.htm
- 21.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene Ontology: tool for the unification of biology. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, et al. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea. Nature 2009; 462:1056-1060 10.1038/nature08656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Growth media used at ATCC: www.atcc.org/Attachments/2718.pdf
- 24.Sims D, Brettin T, Detter J, Han C, Lapidus A, Copeland A, Glavina Del Rio T, Nolan M, Chen F, Lucas S, et al. Complete genome sequence of Kytococcus sedentarius type strain (541T). Stand Genomic Sci 2009; 1:12-20 10.4056/sigs.761 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: Prokaryotic Dynamic Programming Genefinding Algorithm. BMC Bioinformatics 2010; 11:119 10.1186/1471-2105-11-119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pati A, Ivanova N, Mikhailova N, Ovchinikova G, Hooper SD, Lykidis A, Kyrpides NC. GenePRIMP: A Gene Prediction Improvement Pipeline for microbial genomes. Nat Methods (In press). [DOI] [PubMed] [Google Scholar]
- 27.Markowitz VM, Ivanova NN, Chen IMA, Chu K, Kyrpides NC. IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 2009; 25:2271-2278 10.1093/bioinformatics/btp393 [DOI] [PubMed] [Google Scholar]