Abstract
Recent research has provided mechanistic insight into the important contributions of the gut microbiota to vertebrate biology, but questions remain about the evolutionary processes that have shaped this symbiosis. In the present study, we showed in experiments with gnotobiotic mice that the evolution of Lactobacillus reuteri with rodents resulted in the emergence of host specialization. To identify genomic events marking adaptations to the murine host, we compared the genome of the rodent isolate L. reuteri 100-23 with that of the human isolate L. reuteri F275, and we identified hundreds of genes that were specific to each strain. In order to differentiate true host-specific genome content from strain-level differences, comparative genome hybridizations were performed to query 57 L. reuteri strains originating from six different vertebrate hosts in combination with genome sequence comparisons of nine strains encompassing five phylogenetic lineages of the species. This approach revealed that rodent strains, although showing a high degree of genomic plasticity, possessed a specific genome inventory that was rare or absent in strains from other vertebrate hosts. The distinct genome content of L. reuteri lineages reflected the niche characteristics in the gastrointestinal tracts of their respective hosts, and inactivation of seven out of eight representative rodent-specific genes in L. reuteri 100-23 resulted in impaired ecological performance in the gut of mice. The comparative genomic analyses suggested fundamentally different trends of genome evolution in rodent and human L. reuteri populations, with the former possessing a large and adaptable pan-genome while the latter being subjected to a process of reductive evolution. In conclusion, this study provided experimental evidence and a molecular basis for the evolution of host specificity in a vertebrate gut symbiont, and it identified genomic events that have shaped this process.
Author Summary
The gastrointestinal microbiota of vertebrates is important for nutrient utilization, resistance against pathogens, and immune maturation of its host, but little is known about the evolutionary relationships between vertebrates and individual bacterial members of these communities. Here we provide robust evidence that the evolution of the gut symbiont Lactobacillus reuteri with vertebrates resulted in the emergence of host specialization. Genomic approaches using a combination of genome sequence comparisons and microarray analysis were used to identify the host-specific genome content in rodent and human strains and the evolutionary events that resulted in host adaptation. The study revealed divergent patterns of genome evolution in rodent and human lineages and a distinct genome inventory in host-restricted sub-populations of L. reuteri that reflected the niche characteristics in the gut of their particular vertebrate hosts. The ecological significance of representative rodent-specific genes was demonstrated in gnotobiotic mice. In conclusion, this work provided evidence that the vertebrate gut symbiont Lactobacillus reuteri, despite the likelihood of horizontal transmission, has remained stably associated with related groups of vertebrate hosts over evolutionary time and has evolved a lifestyle specialized to these host animals.
Introduction
Vertebrates are associated with trillions of microbes, the majority of which inhabit the digestive tract [1]. Research has led to an appreciation of the importance of these microbial communities, revealing substantial roles in development and performance of the host [2], [3]. As vertebrates evolved, they did so in association with microbes, and these reciprocal interactions have shaped both the attributes of the microbiomes and the phenotypic complexity of the host species [4]. It is conceivable that the beneficial functions of the gut microbiota conferred important selective traits during vertebrate evolution [3], [5]. A joint evolutionary trajectory between host and microbes is evident in anatomical features of vertebrates (rumen, cecum) which allow bacterial fermentations that provide energy to the host and an intensive gut associated immune system that is in place to maintain beneficial microbial communities [6], [7]. These features serve as clear testimony that we cannot attempt to understand the evolution of vertebrates without considering their microbial partners [1], [8].
Comparative analysis of genomes of bacteria originating from human hosts, greatly facilitated through the Human Microbiome Project, provided important insight into the adaptations and ecological roles of different microbial species in the human gut [9]–[11]. Despite these advances, very little is known about the evolutionary strategies of vertebrate gut symbionts. It is often postulated that the evolution of gut microbes involved coevolution of individual lineages with their host species, which is supported by the presence of phylotypes that are specific to particular vertebrate species [3]. However, conclusive evidence for stable associations of specific lineages with vertebrate hosts over evolutionary time-scales has not been provided by 16S rRNA data. Patterns of community similarity provide evidence for codiversification of entire gut communities with their hosts, which suggests that there are host-specific evolutionary interactions between mammals and their microbiomes [4]. In addition, some gut microbes are highly host specific, such as Helicobacter pylori, which has been used to track human migrations over long-time spans [12]. However, many microbial lineages in the mammalian gut are shared across host species [4], implying that some members of the gut microbiota are generalists that pursue promiscuous lifestyles. Such an evolutionary strategy is exemplified by commensal Escherichia coli, which have a broad host range and alternate between niches within the environment and their vertebrate hosts [13], [14]. To date, there are very few vertebrate gut symbionts for which host specificity has been clearly established. Furthermore, little is known about the mechanisms by which gut microbes, for whom symbiotic life is facultative and which have ample opportunities for horizontal transmission, can evolve stable associations with their host species that would allow for reciprocal evolutionary interactions between bacterial lineages and host genotypes.
The Gram-positive bacterium Lactobacillus reuteri is an excellent model organism to study the evolutionary strategy of a vertebrate gut symbiont as this species inhabits the gastrointestinal tract (GIT) of mammals as diverse as humans, pigs, mice, and rats as well as different species of birds. In rodents, pigs, and chickens, it is one of the dominant species in the GIT and forms biofilm-like associations with the stratified squamous epithelial lining of the proximal regions of the digestive tract [15]–[19]. We recently observed that strains of L. reuteri from global sources comprised distinct phylogenetic clusters that can be detected with Multilocus Sequence Analysis (MLSA) and Amplified Fragment Length Polymorphism (AFLP), and these clades show significant association with host origin [20]. The population structure suggests a stable association of L. reuteri with particular vertebrates over evolutionary time and the emergence of host adapted subpopulations. In addition to the genotypic patterns, an adaptive evolutionary process is also reflected by the phenotypic characteristics of L. reuteri strains in terms of ecological performance in the gut and adhesion to epithelial cells [20]–[27]. However, the molecular basis for these host adaptations is still unknown, and it is unclear to what degree the lifestyle and evolution of L. reuteri have remained restricted to particular hosts.
Genomic approaches in combination with experiments in animal models offer mechanistic insight into the evolution and ecology of microbial symbionts of vertebrates. In this study, we used such an approach and showed that only rodent isolates of L. reuteri colonize the gut of reconstituted Lactobacillus-free (LF) mice in high numbers, while isolates from humans, swine, and chicken form either lower populations or fail to colonize. We determined the genome sequence of the rodent isolate L. reuteri 100-23 and performed a comparative genomic analysis with the genome of the human isolate F275. A microarray analysis using genes representative of both strains was used to probe 57 L. reuteri strains, revealing specific gene combinations in host-adapted lineages of L. reuteri. Further genomic comparisons of nine isolates across five MLSA lineages confirmed the microarray data and further allowed the identification of the evolutionary processes that resulted in host-specific genomic features. The ecological significance of rodent-specific genes was demonstrated in gnotobiotic mice, where perturbations in 7 out of 8 genes unique to the rodent lineage resulted in impaired ability to propagate in the murine host.
Results/Discussion
Evolution of a Narrow Host Range in a Vertebrate Gut Symbiont
We tested the ability of thirteen L. reuteri isolates originating from different vertebrate hosts (mouse, rat, human, chicken, and pig) to colonize the digestive tract of LF mice (Table 1). LF mice were previously derived from conventional mice by treatment with penicillin followed by reconstitution with cultures of microbes, non-cultivable microbes attached to epithelial cells, and cecal contents from mice treated with chloramphenicol [28]. These mice are maintained under gnotobiotic conditions and contain a gut microbiota functionally equivalent to conventional animals, but without any lactobacilli. Therefore, these mice are different from germ-free animals in that they allow the investigation of adaptations of Lactobacillus strains under more ecologically relevant conditions. The experiments revealed that only strains originating from mice and rats reached colonization levels after two weeks that were equivalent to those of Lactobacillus populations in conventional mice (Table 1). Isolates from other hosts formed either smaller populations (≤106 cells per gram in the forestomach, <105 cells per gram in the cecum) or were not detectable. These findings provide experimental evidence that the evolution of L. reuteri with rodents has resulted in host specialization. This reinforces previous findings which identified a population structure composed of subpopulations that were to a large degree host confined [20]. The colonization phenotypes in LF mice and the population structure further imply that host restriction does not occur at the host genus level, as L. reuteri isolates from rats showed excellent colonization of mice and group with isolates from mice in MLSA lineages [20].
Table 1. Log10 lactobacilli per gram of organ or fecal sample of ex-Lactobacillus-free mice inoculated with a single strain of Lactobacillus reuteri.
| Strain | Origin | Number of animals | Fecal sample (day 2) | Forestomach (day 14) | Cecum (day 14) |
| N2D | Rat | 7 | 7.5 | 8.7 (0.1)1 | 8.6 (0.3) |
| 100-23 | Rat | 5 | ND2 | 8.0 (0.2) | 7.4 (0.2) |
| 6799-JM1 | Mouse | 7 | 6.5 | 8.7 (0.1) | 7.8 (0.3) |
| #20 | Mouse | 6 | ND | 8.6 (0.1) | 7.7 (0.2) |
| JW2015 | Pig | 7 | 5.1 | 6.0 (0.8) | 4.7 (0.6) |
| LPA1 | Pig | 6 | 4.4 | <2.0 | <2.0 |
| DSM 20016T | Human | 5 | 4.2 | <2.0 | <2.0 |
| CF4-6G | Human | 7 | <2.0 | <2.0 | <2.0 |
| ATCC55730 | Human | 7 | 3.5 | <2.0 | <2.0 |
| M27U15 | Human | 6 | 2.8 | <2.0 | <2.0 |
| CF48-3A1 | Human | Females 3Males 4 | 4.33.4 | 5.5 (0.8)<2.0 | 4.0 (0.8)<2.0 |
| 1366 | Chicken | 6 | 4.3 | <2.0 | <2.0 |
| CSF8 | Chicken | 6 | <2.0 | <2.0 | <2.0 |
Mean (standard error of the mean).
ND = not done.
The human isolate L. reuteri strain CF48-3A1 was detectable in the forestomach and cecum of female mice, but not of male mice, 14 days after gavage. The numbers of CF48-3A1 detected in the gut of female mice were considerably lower than those attained by strains of rodent origin, supporting host specificity of L. reuteri, but the apparent influence of gender was nevertheless noteworthy. The molecular mechanism that supports this gender-specific persistence of lactobacilli in the gut is unknown. Speculatively, gender-specific effects might involve the relative frequencies of receptors for Lactobacillus adhesins on the forestomach epithelium of female compared to male mice. Other factors may include relative retention times of digesta in the large bowel coupled with coprophagy, along with differences in coprophagous habits. These possibilities, however, remain unexplored for the present.
Identification of Host-Specific Genome Content in L. reuteri Strains
Genome sequence of the rodent L. reuteri strain 100-23
The evolution of host specialization in bacteria, which is very well understood for symbionts of invertebrates, can follow diverse paths that can include large scale gene acquisitions and loss and more subtle modifications such as modifications of gene (protein) sequences and regulatory pathways [29]–[33]. The availability of L. reuteri isolates that differed in their ability to colonize the GIT of LF mice paved the way for a genomic analysis to identify the molecular basis of host specificity in a vertebrate symbiont. The sequence of the human isolate F275 was previously completed [34], and four human isolates (CF48-3a, MM4-1, MM2-3, ATCC 55730) were sequenced in the course of the Human Microbiome Project [35]. We produced a high quality genome sequence of L. reuteri 100-23, an isolate from the stomach of a rat that served as a model organism for mechanistic studies on gut microbial ecology, in vivo biofilm formation, and immunology in combination with LF mice [26], [36]–[41]. This strain groups with the rodent-associated MLSA lineage III [20]. Genome sequencing resulted in two scaffolds of 729,351 bp and 1,576,206 bp with a total of 2375 detected genes, consisting of 2269 protein-coding genes, and 106 RNA genes (including six rRNA operons). The general genome features are listed in Table S1. L. reuteri 100-23 harbors two indigenous plasmids, pGT231 (5,254 bp; GenBank accession no. GU108604) and pGT232 (5,123 bp, GenBank accession no. NC_001757, with pGT232 belonging to the pC194/pUB110 family of rolling-circle plasmids [42]). The significance of these plasmids for the biology of L. reuteri has not yet been determined, and the plasmid sequences were not included in the comparative genomic analysis below.
Comparison of the genomes of the rodent strain 100-23 and the human isolate F275
L. reuteri F275 is a fecal isolate from a healthy adult human and a member of the human-associated MLSA lineage II (Clonal complex CC-47) [20]. The strain is unable to colonize LF mice (Table 1), and its genome is around 270 kb smaller than 100-23, containing around 290 fewer genes (Table S1). F275 does not contain any plasmids, and in contrast to 100-23, it contains 35 pseudogenes. A whole genome BLASTP comparison revealed that L. reuteri 100-23 contains 633 genes with no orthologues in F275, while the latter has 352 genes without an orthologue in 100-23. A summary of the unique genes is given in Table S2. Both genomes contained more than a hundred genes annotated as transposases (most with homologies to described IS elements), integrases, and phage related proteins, many of which were strain specific. Genes with assigned functions that are unique to the two genomes included genes coding for cell wall and membrane bound proteins, transport proteins, regulatory proteins, enzymes, and glycosyltransferases. An auxiliary protein secretion system (SecA2 cluster) and a urease gene cluster were unique to 100-23, while only F275 contained the pdu-cbi-cob-hem cluster [34], [43].
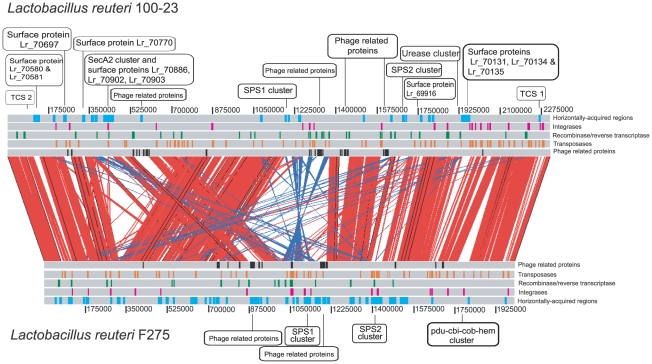
We used the Artemis Comparison Tool to localize strain specific genomic regions in L. reuteri 100-23 and F275 (Figure 1). The two genomes contained many regions of synteny, especially around the origin of replication. However, one major rearrangement and a major inversion were also present. The rearrangement was likely to have occurred in 100-23 as the F275 genome sequence shows greater synteny with the genome of the related species Lactobacillus fermentum (Figure S1), and it is therefore likely to reflect the ancestral structure. In addition, the inversion within the genome of 100-23 was rich in genetic elements (e.g. transposases), which may have caused the rearrangement through a recombination event (Figure 1). Many of the genes that were unique to 100-23 or F275 were clustered in genomic regions that were completely absent in the other strain. Several of these regions showed characteristics of genomic islands as they were associated with unusual sequence features such as low %GC content, atypical codon bias, mobile genetic elements (prophage related genes or putative IS elements/transposons), and they were predicted to be transferred by lateral gene transfer (LGT) using the software Alien_hunter. Several other regions were identified to be present in both genomes but differed significantly in terms of gene content. These regions coded for genes involved in the production of surface polysaccharides (SPS1 and SPS2) or contained putative prophages.
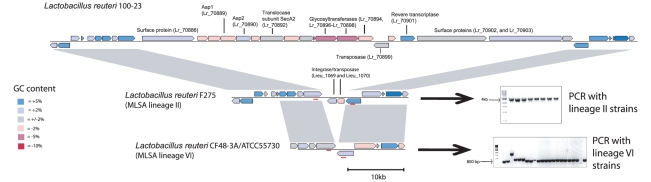
Figure 1. Pair-wise genomic comparisons between L. reuteri strains 100-23 and F275.
Linear genomic comparison of the chromosomes of 100-23 and F275 (using the sequence of JCM1112T). Both sequences are read left to right from the predicted origin of replication. Homologous regions within the two genomes identified by reciprocal BLASTN are indicated by red (same orientation) and blue (reverse orientation) bars. Putative horizontally acquired islands as identified by Alien_hunter (blue boxes), phage proteins (black boxes), transposases (orange boxes), and integrases (pink boxes) are indicated.
Genomic survey of 57 L. reuteri strains from different vertebrate hosts
To differentiate true host-specific gene content from strain-level differences, the genomes of 100-23 and F275 were used to design representative spotted genomic microarrays. These were used to interrogate the genome content of 55 additional L. reuteri strains (>99% homology on 16S rRNA sequence to the type strain F275) by comparative genomic hybridization (CGH). The strain collection was composed of isolates from six different vertebrate hosts that belong to five distinct MLSA lineages (Table S3). MLSA previously revealed that human isolates belong to two separate lineages (II and VI) [20], and representatives from both lineages were included in CGH. MLSA lineage II is mainly composed of strains isolated from human fecal samples, and strain F275, which falls within this cluster, has been reported to be detectable in fecal samples of the same individual for around six months [44]. The strains from MLSA lineage II are therefore likely autochthonous to the human digestive tract. In contrast, human isolates from lineage VI primarily originate from other body parts (vagina, mouth, breast-milk), and they group tightly with strains from poultry, indicating that they are allochthonous to the human GIT [20].
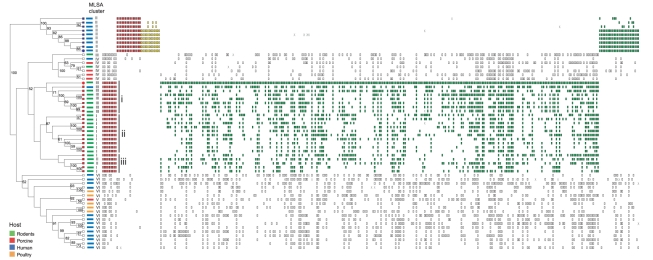
CGH patterns were analyzed using the MARKFIND program [45], which performs a cluster analysis based on genome polymorphisms by the unweighted pair-group method with arithmetic means (UPGMA). As shown in Figure 2, the phylogeny inferred from the genome polymorphisms reflected both host origin and MLSA typing. The rodent strains formed one cluster comprised of three sub-groups (i, ii, and iii), with i and ii corresponding to MLSA lineages III and I, respectively. The human and poultry isolates that belong to the MLSA lineage VI also formed one separate group with CGH. The human strains from MLSA cluster II formed an isolated cluster that grouped distantly from all other L. reuteri strains, which is indicative of markedly different genome content (Figure 2). Thus, although the topologies of the dendrograms inferred from gene polymorphisms and MLSA sequences were different, both methods resulted in trees with host-specific phylogenetic clusters that were congruent. This indicates that L. reuteri has diverged into genetically and ecologically cohesive subpopulations (ecotypes) whose gene content reflects particular host niches.
Figure 2. Analysis of genome content by reference microarray.
The dendrogram is derived from UPGMA analysis of binary data generated from 57 strains of Lactobacillus reuteri. Bootstrap scores, using 10,000 repetitions of a UPGMA search, are shown on nodes scoring >50%. Each strain is color coded by host, and affiliation to MLSA lineages of each strain is indicated. MARKFIND was used to identify gene polymorphisms specific to human (MLSA lineage II) and rodent strains (mostly MLSA lineages I and III), which clustered in the dendrogram. The leaves of human (lineage II) and rodent strains, which have been used for comparisons by MARKFIND, are colored blue and red, respectively. The two reference strains, 100-23 and F275, are labeled by a red and a blue circle, respectively. Vertical rectangles to the right depict polymorphisms present in a given strain sorted by the MARKFIND program. Those polymorphisms conserved within all members of a lineage and absent in the other lineage are colored red. Genes that are lineage-specific but non conserved are colored green (polyphyletic) or yellow (monophyletic). Polymorphisms that are not lineage specific are not shown.
Evolutionary genomics of L. reuteri
CGH with multiple-strain comparisons both within and across L. reuteri lineages allowed us to identify evolutionarily- and ecologically-relevant patterns of genome variation. We used MARKFIND to identify genes that were unique to rodent strains (including 100-23) when compared to strains of the human MLSA cluster II (including F275) and vice versa, and these polymorphisms are represented by red, green, and yellow rectangles in Figure 2. We focused this analysis on the differences of rodent and human lineage II strains as the spotted microarray was based on the genomes of strains that belong to these groups, assuring more reliable hybridizations (see Materials and Methods). MARKFIND identified eight genes that are conserved in all rodent strains but absent in human strains and 256 genes that were specific to rodent strains but non-conserved (Table S4). 15 genes were identified to be conserved among the human cluster II and absent in rodent strains, while 37 genes were identified to be specific to human strains but not conserved (Table S5). It is of note that all genes identified as ‘rodent-specific’ when compared to the human lineage II were also detected in at least some strains associated with pigs and poultry.
A summary of the host-specific genes detected by MARKFIND in comparisons between rodent and human MLSA lineage II strains is presented in Table 2. Many of these genes are mobile genetic elements. In addition, almost half of the rodent-specific genes encoded hypothetical proteins with unknown function that showed a very low conservation even among rodent strains. Only 10 genes with a functional annotation other than transposition were specific to the human lineage. Most of these genes were glycosyl-transferases from the SPS2 cluster and several enzymes (histidine decarboxylase, histidyl-tRNA synthetase, dextransucrase, two lipolytic proteins). Rodent strains possessed 93 host-specific genes with assigned functions other than DNA transposition. Most of these genes fell within the putative genomic islands identified above (Figure 1). The distribution of the genes within these islands among all strains included in the CGH analysis is shown in Figure 3. The urease cluster was the only feature that was both conserved across rodent strains and absent in isolates from other hosts. All other rodent-specific clusters showed different degrees of strain-to-strain variation. Genes encoding eleven large surface proteins and the Two-Component Regulatory System (TCS) TCS2 were rare in isolates from non-rodent hosts, while the xylose cluster and the asp3 gene of the SecA2 cluster were also detectable in isolates from pigs. The two SPS clusters, though to a large degree host-specific, showed a very high variability in gene composition among rodent strains and were also detectable in strains of lineages IV (pig) and VI (poultry/human). A second regulatory system (TCS1) and the Multidrug efflux cluster (ABC) were only detectable in a small number of rodent strains.
Table 2. Host-specific genes in rodent and human lineage II strains as identified by CGH.
| Genes | Rodent | Human |
| Transposases/Integrases | 21 | 2 |
| Phage-related proteins | 32 | 16 |
| DNA Binding and restriction endonucleases | 11 | 8 |
| Urease gene cluster | 1 | 0 |
| Cell Wall/Membrane bound proteins | 12 | 0 |
| Transport proteins (Ion, peptide, sugar) | 19 | 0 |
| Regulatory proteins | 11 | 1 |
| Enzymes (peptidases, hydrolases, dehydrogenases, kinases, amylases, reductases) | 25 | 5 |
| Glycosyl transferases and sugar isomerases/epimerases | 25 | 4 |
| Hypothetical and unknown proteins | 107 | 16 |
| Total | 264 | 52 |
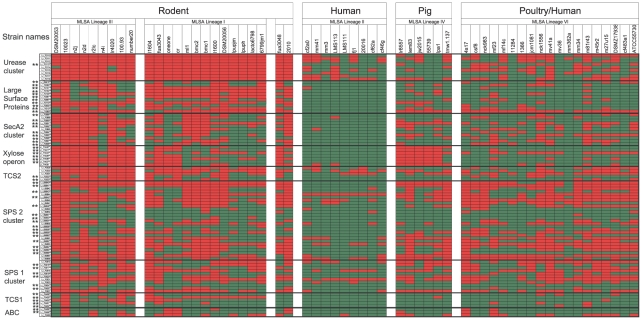
Figure 3. Distribution of genes that belong to putative genomic islands in 57 L. reuteri strains.
The strains are arranged by host and MLSA lineage [20]. Red and green colors indicate presence or absence, respectively. Genes that have been identified as rodent-specific when compared to human lineage II strains by MARKFIND are indicated by asterisks (**).
Validation of Host-Specific Gene Content by Genome Comparisons and PCR
The species L. reuteri shows a significant degree of genetic variation, especially between strains from different MLSA lineages [20]. Sequence divergence can confound the CGH data as it impairs hybridizations. This was apparent because even though hybridizations were very reliable for the genomes of the reference strains 100-23 and F275 (>96% accuracy), the error rate was approximately 18.5% for strain CF48-3A of lineage VI. Therefore, to confirm the findings obtained with the CGH analysis and to gain further insight into the distribution of host-specific gene content throughout the entire L. reuteri population, we performed additional genomic comparisons in combination with PCR. First, we generated draft genome sequences (>15× coverage) of two additional rodent strains (lpuph1 and MLC3) and one pig strain (ATCC 53608). We then determined the presence of the host-specific genes identified by CGH and the pdu-cbi-cob-hem cluster in all available L. reuteri genomes (100-23, lpuph1, MLC3, ATCC 53608, F275, MM4-1a, MM2-3, ATCC55730, and CF48-3A). These genomes represent five MLSA lineages, lineages I and III (rodent), lineage II (human), lineage IV (pig), and lineage VI (poultry/human), and the genome characteristics are shown in Table S6. The average nucleotide identity (ANI) of a core set of genes within these L. reuteri genomes and L. vaginalis is shown in Table S7. An ANI of >95% was determined in all the L. reuteri genome comparisons, providing additional evidence that these strains, despite their considerable genomic differences, fall within what is currently considered to be one prokaryotic species [46].
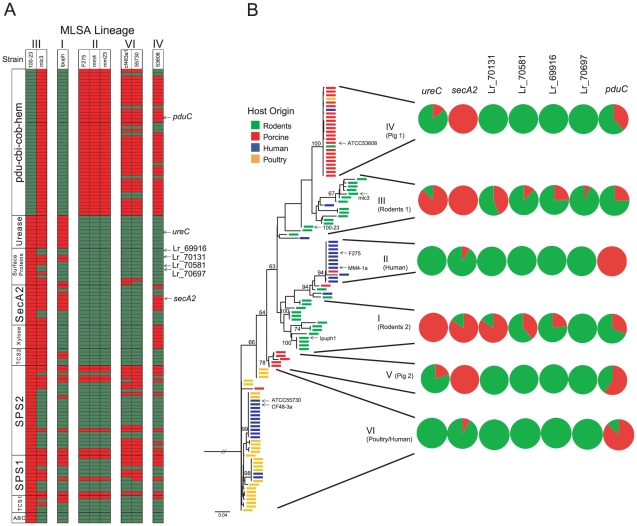
As shown in Figure 4A, the genomic comparisons confirmed the findings obtained with the CGH analysis. The pdu-cbi-cob-hem cluster was detected in all human isolates (MLSA lineage II and VI) and the pig isolate ATCC 53608 (MLSA lineage IV), but it was only present in one of the three rodent strains. The urease cluster was strictly conserved among the three rodent strains and absent in all other genomes, while the surface proteins and the TCS2 cluster were to a large degree specific to rodents but more variable. The SecA2 and xylose clusters were detectable in rodent and porcine strains but completely absent in strains from lineage II and VI. The SPS and TCS1 clusters showed a much higher variability among rodent strains and several of the genes were datable in the lineage VI and IV strains, while most of the genes were absent in human lineage II strains. Consistent with CGH, the ABC transporter was specific to strain 100-23. To study the distribution of host-specific genomic features throughout the L. reuteri population, PCR was used to determine the presence of genes encoding SecA2, several surface proteins (Lr_70131, Lr_70581, Lr_70697, Lr_69916), UreC (the urease alpha subunit), and PduC (diol/glycerol dehydratase encoded by the pdu-cbi-cob-hem cluster) in 88 L. reuteri strains (Table S3). The results are shown in Figure 4B in a phylogenetic context. This analysis confirmed that several of the key genetic determinants identified by CGH are to a large degree associated with specific MLSA lineages and vertebrate hosts.
Figure 4. Confirmation of host-specific gene content by genomic comparisons and PCR.
(A) A heatmap representation of BLASTN comparison of rodent-specific genes against sequenced L. reuteri genomes. Genes whose distribution throughout the species L. reuteri was determined by PCR are marked by arrows. (B) Distribution of host-specific genes throughout the phylogenetic spectrum of L. reuteri. A maximum likelihood tree of MLSA data from 116 L. reuteri strains is shown [20], and the strains for which genome sequences were included are marked by arrows. Pie charts showing the proportion of queried strains within lineages possessing the targeted genes (ureC, urease; Lr_70892, secA2, surface proteins Lr_70131, Lr_70581, Lr_70697, and pduC of the pdu-cbi-cob-hem cluster) as detected by PCR.
Genomic Features Associated with Host Origin
The urease cluster
This cluster (genes Lr_70110–Lr_70118 in the genome of 100-23) is highly conserved among rodent strains and highly host specific (Figure 3 and Figure 4). This is in accordance to previous phenotypic characterizations which showed that urease activity can be detected in rodent L. reuteri isolates, while the activity is rare in porcine isolates and absent in human and poultry isolates [5]. Genes for urease production are absent in all currently available genomes of other Lactobacillus species, but orthologs (42–75% amino acid identity) are present in Streptococcus species, suggesting that the cluster was acquired by L. reuteri through LGT. Urease has been shown to be an important component of survival in acidic conditions as well as in biofilm communities by ameliorating the buildup of acidic metabolic end-products [47], [48], by which it could contribute to the survival of L. reuteri in the forestomach of rodents.
Large surface proteins
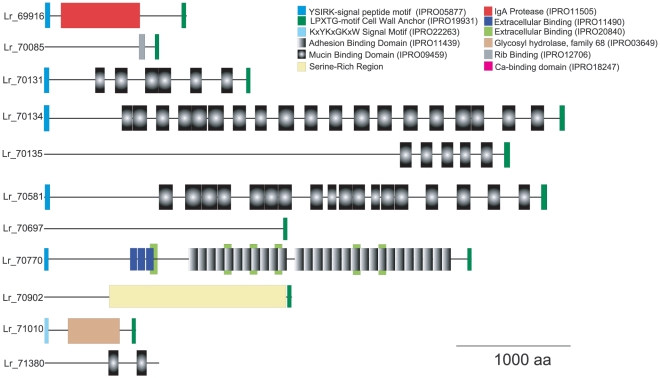
Eleven large (>750 aa) surface proteins were detected in rodent strains that were very rare in isolates of pigs and poultry and absent in human isolates of MLSA lineage II (Figure 3). The characteristics of these proteins are described in Table 3 and their schematic representation is shown in Figure 5. Most of the surface proteins are predicted to be involved in epithelial adhesion and biofilm formation. Six proteins (Lr_69656, Lr_70131, Lr_70134, Lr_70135, Lr_70581, Lr_71380) contained putative mucin-binding MucBP domains (Pfam PF06458) and other domains involved in extracellular matrix binding. Additional domains detected included a glycosyltransferase (family 68) domain in a predicted levansucrase (Lr_71010), and a putative IgA-specific protease (Lr_69916).
Table 3. Large surface proteins (>750 amino acids) specific to rodent strains.
| Protein (AA) | Similar proteins (name and/or GenBank entry) | Organisms (%Identify/no amino acids) | Features (Interpro) |
| Lr_70770 (3787) | NP_964984.1 | L. johnsonii NCC 533 (55/3971) | YSIRK signal peptide, LPXTG anchor, 28 DUF1542 motifs (Pfam07564), 3 Ebh (extracellular matrix-binding) motifs (Pfam07554) |
| Lr_70697 (2129) | ZP_04009761 | L. salivarius ATCC 11741 (27/1040) | LPXTG motif, signal peptide |
| Lr_70131 (1828) | Lr_70581Mlp | L. reuteri 100-23 (85/807),L. reuteri BR11 (78/802) | YSIRK signal peptide, LPXTG anchor, 6 MucBP motifs (Pfam06458) |
| Lr_70134 (4634) | YP_194248 | L. acidophilus NCFM (75/3130) | YSIRK signal peptide, LPXTG anchor, 19 MucBP motifs (Pfam06458) |
| Lr_70135 (4141) | LJ0484/NP_964510 | L. johnsonii NCC 533 (80/2495) | LPXTG motif, 6 MucBP motifs (Pfam06458) |
| Lr_70581 (4454) | Mlp/AAP41738YP_193900 | L. reuteri BR11 (73/2094)L. acidophilus NCFM (58/1166) | YSIRK signal peptide, LPXTG anchor, 18 MucBP motifs (Pfam06458) |
| Lr_70902 (2180) | LJ1711 | L. johnsonii NCC 533 | LPXTG motif, shares similarities (around 25%) with trophinin from mice and rat, a host protein involved in the |
| Lr_69916 (1235) | LJ1680 | L. johnsonii NCC 533 (54/1234) | LPXTG motif, IgA protease conserved among Streptococci |
| Lr_70085 (1034) | EEW53535.1 | L. antri DSM16041 (30/483) | Rib-binding domain, LPXTG anchor |
| Lr_71010 (796) | ZP_03974578 | L. reuteri CF483A1 (82/591)L. sanfranciscensis (51/451) | Levansucrase Ftf, KxYKxGKxW signal peptide, LPXTG anchor |
| Lr_71380 (1036) | ZP_03975092.1 | L. reuteri CF483A1 (90/516)L. gasseri JV-V03 (24/103) | 2 MucBP domains |
Figure 5. Rodent-specific large surface proteins of L. reuteri 100-23.
The architecture of large surface proteins (>750 aa) specific to rodent strains when compared to human isolates of the MLSA lineage II. Functional domains of each protein are shown as indicated and to scale.
The accessory Sec (SecA2) system
The SecA2 cluster was detected by PCR in most strains from rodents and pigs (MLSA lineages I, III, IV, and V), while it is rare in isolates from human and poultry hosts (MLSA lineages II and VI). This auxiliary protein secretion system is present in a limited number of gram-positive bacteria and mycobacteria in addition to the canonical SecA system [49]. Conservation of the SecA2 cluster with other members of the Class Bacilli and sparse distribution among different species of lactobacilli implies that this system was horizontally acquired by only a few Lactobacillus lineages. LGT of this cluster in L. reuteri is supported by the presence of mobile genetic elements (Lr_70899 and Lr_70901) within the cluster, a low GC content (Figure 6), and by analysis with Alien_hunter (Figure 1). As shown in Figure S2, gene content within the accessory Sec cluster is conserved in L. gasseri, Streptococcus gordonii, and L. reuteri 100-23. In streptococci, the accessory SecA2 system facilitates the selective export of glycosylated serine-rich proteins that often function as adhesins [49], [50]. Though we do not yet know which proteins are secreted through this pathway in L. reuteri, the surface proteins Lr_70886, Lr_70902, and Lr_70903 are adjacent to the cluster in the genome of 100-23. Of those, Lr_70902 is unusually serine rich (35% serine), and the serine residues may be glycosylated by glycosyltransferases associated with the SecA2 cluster (Lr_70896–Lr_70898) analogous to serine rich surface protein in streptococci, such as GspB [49]–[51].
Figure 6. Comparison of the genomic location that contains the accessory SecA2 cluster in L. reuteri 100-23.
Gene map of the accessory SecA2 cluster in Lactobacillus reuteri 100-23 and the same genomic region in strains F275 (MLSA lineage II) and CF48-3A and ATCC55730 (MLSA lineage VI). Genes are colored according to differences in GC content when compared to the genome background (39%). The PCR products shown were generated with primers that targeted conserved genes that flank the location of the SecA2 cluster (primer sites are shown by red bars).
The xylose operon
A xylose operon is highly conserved in rodent (especially in lineage III) and porcine strains (MLSA lineage IV), while it was absent in all human and poultry strains (lineage II and VI). Xylose could be an important substrate for gut bacteria as it is a plant-derived sugar commonly found in straw and bran, and the xylA promoter of strain 100-23 was previously identified by in vivo expression technology to specifically induced in the gut of mice [38].
A two-component regulatory system
TCS2 was detected by CGH in more than half of the rodent strains, and the system contains a putative histidine kinase (Lr_70529), a response regulator of the LytR/AlgR family (Lr_70530), a bacteroicin-like peptide (Lr_70531), an ABC-type bacteriocin transporter (Lr_70532), and an ABC-type bacteriocin/lantibiotic exporter, containing an N-terminal double-glycine peptidase domain (Lr_70533). Lr_70532 showed high similarity (55%) to AbpT of Lactobacillus salivarius UCC118, which was shown to be important for bacteriocin activity [52]. The Lr_70531 peptide shares no common sequence homology to other proteins in the NCBI database, but contains a double-glycine motif. Cleavage at this site (at amino acid position 19) would produce a 35-aa peptide that shows characteristics described for extracellular bacterial signaling peptides [53]. Since strain 100-23 does not produce a bacteriocin, it is possible that this regulatory system is involved in quorum sensing (QS). The specificity of TCS2 to rodent L. reuteri strains suggests that it might affect the transcriptome facilitating host adaptation. In this respect, it is of note that a single two-component sensor kinase can alter the host range of Vibrio fischeri [29].
Genetic features specific to rodent strains that show high inter-strain variability
Several genes were identified by CGH to be rodent specific but were detected in only a small number of strains. These included a second two-component system (TCS1) that was comprised of a histidine kinase (Lr_70430), a LytR/AlgR family response regulator (Lr_70431), and a bacteriocin processing peptidase (Lr_70432). This system meets the criteria established by Sturme and coworkers for a peptide-based QS two-component regulatory system [54]. In addition, three genes (Lr_70458, Lr_70459, Lr_70460) that comprise a putative ABC-type Multidrug Efflux System were detected by CGH in three rodent strains. Also, several genes present in the two SPS clusters (SPS1 and SPS2), encoding predicted glycosyltransferses, epimerases, and capsular polysaccharide biosynthesis proteins (Figure S3), were identified by MARKFIND to be host specific. These clusters showed a very high variability among rodent strains (Figure 3).
Host-specific gene content in human strains
Around half of the genes specific to the human MLSA lineage II were related to mobile elements (transposases/integrases, phage proteins, restriction endonucleases) and hypothetical proteins with unknown functions (Table 2). The pdu-cbi-cob-hem cluster was conserved within human strains, and the cluster was absent in rodent strains of the CGH sub-groups i and ii, while it was present in 4 out of the 5 strains in the rodent sub-lineage iii (Figure 2). This cluster codes for cobalamin (vitamin B12) biosynthesis, glycerol utilization, propanediol fermentation, and production of the antimicrobial compound reuterin [34], [43], [55], [56].
Host-Specific Gene Content Reflects Niche Characteristics in Different Hosts
The functions of the genetic features associated with L. reuteri ecotypes are reflective of their lifestyle in respective hosts. In rodents, L. reuteri adheres directly to the stratified squamous epithelium present in the murine forestomach and forms thick cell layers that show characteristics of biofilms [25], [26], [36], [39]. Accordingly, several of the rodent-specific surface proteins are predicted to function as adhesins or mediators of biofilm formation, and the SecA2 system is likely involved in the secretion of some of these proteins (e.g. Lr_70902). Other factors, such as the TCS2, fructosyltransferase (Ftf), IgA specific metallopeptidase, and the urease cluster are likely to play roles in biofilm formation, cell aggregation, and the mitigation of low pH and exposure to IgA, respectively. It is striking that several of the genes identified as rodent-specific by CGH were also detectable in at least some strains that originate from pigs and poultry (Figure 2 and Figure 3), reflecting the similar lifestyle of rodent, porcine, and poultry lactobacilli which all form biofilm-like associations with epithelial surfaces in the proximal GIT [18], [57], [58].
The genome content of strains within the human MLSA lineage II is strikingly different when compared to other L. reuteri lineages. The absence of many genetic features involved in biofilm formation and adhesion reflects the lifestyle of L. reuteri in the human gut. Squamous stratified epithelia are absent, and epithelial cell layers rich in lactobacilli equivalent to those found in animals have not been described in the human GIT [19]. The genome content of strain F275 suggests a planktonic lifestyle in more distal regions of the human gut and limited, if any, interactions with the gut epithelium. This lifestyle would require fast multiplication rates, which could explain the absence of the large surface proteins in lineage II strains, which are likely to be a significant energetic burden. In addition, easily accessible nutrients are in low supply in the human colon having been absorbed in the small intestine, and the ability of L. reuteri to use 1,2-propanediol as an energy source through the pdu-cbi-cob-hem cluster might therefore constitute an important colonization factor in the human gut. The production of reuterin, which is also conferred by this cluster, might contribute to the fitness of L. reuteri in the human gut through inhibition of competitors in the same niche (as reviewed in [5]). Enzymes involved in 1,2-propanediol utilization and reuterin formation require Vitamin B12 as a co-factor [43], [55]. The synthesis of Vitamin B12 is also encoded by the pdu-cbi-cob-hem cluster, and it appears to be an important colonization factor for colonic bacteria, as demonstrated for Bacteroides thetaiotaomicron [59].
Rodent-Specific Genes Contribute to Fitness in the Mouse GIT
Although it is striking how gene content of L. reuteri lineages reflects niche characteristics in particular hosts, differences in gene frequencies within populations can arise not only through natural selection but also random genetic drift. In order to test whether the rodent specific genes were of ecological significance in the GIT of mice, we investigated the fitness of isogenic mutants of strain 100-23C in the gut of LF mice in competition with the parental strain. Eight genes representing major groups of genetic functions among the lineage-specific genes were selected for these experiments: Lr_70902 (serine-rich surface protein), Lr_70770 (putative adhesin), Lr_70892 (SecA2 translocase), Lr_70890 (Asp2, involved in SecA2 transport system), Lr_70894 (SecY2, involved in SecA2 transport system), Lr_70430 (two-component system histidine kinase), Lr_70458 (ABC-type multidrug transport system), Lr_70532 (ABC-type transporter of TCS2). This selection included sets of genes with high (Lr_70902, Lr_70770, Lr_70892, Lr_70890, Lr_70894, Lr_70532) and low conservation (Lr_70430, Lr_70458) among rodent strains. Further, it included genes with a variety of functions, such as adherence, secretion of surface proteins, and environmental sensing. As shown in Figure 7, when the parental strains and their mutant derivatives were introduced into LF mice, seven out of the eight mutants had impaired ecological fitness. The most significant defect in competitive fitness was caused through the inactivation of Lr_70890, Lr_70894, and Lr_70902, which are all associated with the secA2 operon. The only gene that did not contribute to ecological performance was Lr_70770, which encoded a putative adhesin. Given the large number of putative adhesins in the genome of L. reuteri 100-23 (Table 3), it is possible that redundancy exists in mechanisms that confer adherence.
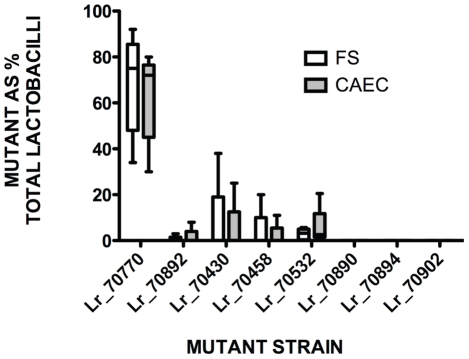
Figure 7. Ecological significance of rodent-specific genes for fitness of L. reuteri 100-23C in the mouse GIT.
Competition between wild-type and mutant L. reuteri 100-23C strains in Lactobacillus-free mice. Mixtures of wild-type and mutant (1∶1) were used to inoculate mice, and the percentage of mutants in the total Lactobacillus population was determined after 7 days. The whiskers show the range of values obtained with different animals (n = 5–6), with the box indicating the 25th and 75th percentile. The bars in the boxes indicate median values. Mutants are listed by gene names; Lr_70770 (surface protein), Lr_70892 (SecA2 translocase), Lr_70430 (two-component system histidine kinase), Lr_70458 (ABC-type multidrug transport system), Lr_70532 (ABC-type transporter of TCS2), Lr_70890 (Asp2, involved in SecA2 transport system), Lr_70894 (SecY2), Lr_70902 (serine-rich surface protein).
Identification of Genetic Mechanisms That Led to Differences in L. reuteri Genomes
The genetic architecture reflected in the genomes of the rodent and human-adapted L. reuteri strains 100-23 and F275 provides insight into the evolutionary processes that underlie host specialization. First, it is clear that that LGT played an important role in the evolution of L. reuteri, as many of the host-specific functions were found to be encoded on putative genomic islands or on regions with lost synteny between the two related strains (Figure 1). In addition, the pdu-cbi-cob-hem cluster, which is absent in most rodent strains, has previously been identified to be a horizontal acquisition of L. reuteri [34], [60]. Therefore, the acquisition of novel genetic material could have led to phenotypic innovations in L. reuteri and might have allowed lineages to become associated with vertebrates, radiate among vertebrate hosts, or to switch hosts during evolution.
However, closer scrutiny of the gene organizations at the loci of genomic difference between L. reuteri strains 100-23 and F275 suggested an additional mechanism of genome evolution. As shown in Figure 6 and Figures S3, S4, and S5, the pdu-cbi-cob-hem, SecA2, urease, and SPS clusters as well as the xylose operon and most of the surface proteins (Lr_70770, Lr_70131–Lr_70137 cluster, Lr_69916, Lr_70580/Lr_70581 cluster, and Lr_71380) are all replaced or interrupted by mobile genetic elements (e.g. putative IS elements and phage related genes) in the genomes of strains 100-23 and F275, respectively. These findings indicate that most of the lineage-specific genes in rodent and human lineage II strains were ancestral and appeared to be jettisoned after divergence of the two lineages. This means that genome evolution of L. reuteri strains is, in many cases, a process associated with gene deletions, possibly caused by mobile genetic elements that mediated rearrangements through recombination. Functional gene loss is a common mechanism that underlies host specialization in both pathogenic and symbiotic bacteria from various phylogenetic groups [30]–[32], [61]. Our findings indicate that it also plays an important role for host specialization in L. reuteri, especially in the human lineage II.
Important Genomic Events in the Evolutionary History of L. reuteri
Given the long time periods involved and the lack of intermediate steps, it is currently difficult to reconstruct the evolutionary processes that have shaped L. reuteri subpopulations. However, the genomic comparisons of strains spanning several MLSA lineages allowed us to pinpoint some specific key events in the evolution of the species. The pdu-cbi-cob-hem cluster appears to be an ancient acquisition of L. reuteri as it is distributed through the entire phylogenetic spectrum of the species (Figure 4). This is in accordance with conclusions based on codon adaptation index and GC content [60]. The cluster is absent in most rodent strains, and the analysis of the loci in strain 100-23 indicated that the cluster was deleted through the action of mobile elements (Figure S4A). It is one of only very few examples of gene loss exclusive to this lineage, making it interesting to speculate as to why its function may be obsolete for the success of L. reuteri in the rodent forestomach.
The SecA2 cluster, which is highly conserved in rodent and porcine strains (Figure 4), appears to be a later acquisition of L. reuteri, as all but one strain from the lineage VI lack this cluster. As shown in Figure 6, there is no evidence for deletion of the cluster in lineage VI strains, while strains of MLSA lineage II showed evidence for deletion through mobile genetic elements. This indicates that the cluster was acquired after diversification of more recent lineages from lineage VI. The acquisition of the SecA2 cluster might have been a pivotal innovation of L. reuteri strains to colonize the gut of mammals. The biological significance of the SecA2 cluster for life in the rodent gut was clearly demonstrated in our competition experiments in LF mice, in which inactivation of four different genes in strain 100-23C associated with this cluster (Lr_70890, Lr_70892, Lr_70894, and the surface protein Lr_70902) had the most detrimental effects when compared to the other mutants tested (Figure 7).
The comparison of the genomes of L. reuteri 100-23 and F275 revealed evidence for only one event of LGT since the split of the two lineages. The surface protein Lr_70697 is arranged in an island with two transposases and two phage integrases next to a transfer RNA gene (tRNA-Val) in the genome of 100-23. This locus is intact in the genomes of F275, CF48-3a, and ATCC55730. Therefore, this gene cluster was likely acquired by a recent ancestor of 100-23 and inserted into a tRNA-Val gene, as described for islands in meseorhizobia and several pathogenic bacteria [62]. As with mesorhizobia, insertion of the cluster in L. reuteri left the entire tRNA gene (a Thr-tRNA) intact upon integration, whereas a small part (22 nucleotides in L. reuteri) became duplicated as a direct repeat (see Figure S5B). Both CGH (Figure 3) and PCR (Figure 4B) analyses showed that Lr_70697 was to a large degree specific to strain 100-23, supporting the hypothesis that this cluster was a recent genomic acquisition.
A Rodent-Specific Accessory Genome of L. reuteri
A recent study on the genomes of human L. reuteri strains revealed a closed pan-genome, with individual strains contributing to a very small number of new genes [35]. Our CGH analysis supported these observations, showing similar genome content and little genetic diversity among strains belonging to the human MLSA II lineage (Figure 2). However, strains from other hosts, and especially rodents, possessed a more variable gene content, and the majority of the rodent-specific genes detected by CGH were not conserved among rodent strains (Figure 2). Comparisons of the genomes of the three rodent L. reuteri strains 100-23, MLC3, and lpuph1 confirmed that rodent strains possess a larger pan-genome with a gene repertoire that extends beyond that of individual strains. Open pan-genomes have been described for many bacterial species, and they consist of a ‘core genome’ (genes present in all strains) and an ‘accessory’ genome (genes variable among strains) [14], [63], [64].
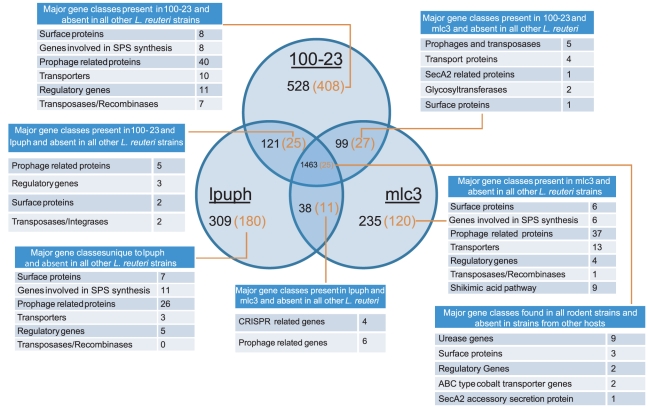
As shown in Figure 8, the three rodent strains shared around 1463 of the predicted protein coding genes. Of this core genome, only 25 genes were unique to rodent strains (Figure 8), confirming the CGH analysis in that only a small number of rodent specific genes are conserved among strains. Each strain possessed a significant proportion of genes that were absent in the other strains (528 proteins in 100-23; 235 in MLC3; and 309 in lpuph1), confirming the more variable gene pool among rodent strains. Of note, a large proportion of these genes were not found in the genomes of L. reuteri strains from non-rodent hosts (Figure 8). This rodent-specific accessory genome was comprised, apart from a large portion of mobile genetic elements, of the same functional groups as the genes identified by the CGH analysis to be rodent-specific (Figure 8 and Table S8). Thus, many of the rodent-specific surface proteins, glycosyltransferases involved in SPS synthesis, transport proteins, and regulatory proteins that are present in the genome of strain 100-23 are substituted by genes that are predicted to perform similar functions in strains MLC3 and lpuph1. The genomic comparisons revealed only one group of rodent-specific genes that were absent in the genome of 100-23 and were therefore not detected by CGH. These genes were all CRISPR-related and are likely to be involved in phage resistance.
Figure 8. The pan-genome of L. reuteri strains originating from rodents.
Venn diagram showing gene distribution and overlap between the genomes of three rodent L. reuteri strains (100-23, MLC3, and lpuph1). Shown in parentheses are the numbers of genes after subtracting genes found in the genomes of L. reuteri strains originating from non-rodent hosts (F275, MM2-3, MM4-1a, CF48-3a, ATCC55730, and ATCC53608). Major functional classes represented among these genes are listed (genes without functional annotations are not shown).
It is important to point out that the seven rodent-specific genes that contributed to ecological fitness in colonization experiments in LF mice (Figure 7) were not conserved among rodent strains. A key conclusion of this study is therefore that adaptive traits that allow life in the murine gut are encoded by a rodent-specific accessory genome and that different combinations of these genes promote successful colonization. This of course begs the question of why plasticity is favored in the rodent L. reuteri population but not the human lineage II. It has been suggested that bacterial accessory genomes encode special ecological adaptations in genes that remain unbounded and can be more rapidly incorporated where and when they become advantageous [65], [66]. Thus, the larger gene pool within in the rodent L. reuteri population might be sampled by individual cells through LGT to form the basis for adaption to environmental fluctuations. The population genetic structure of L. reuteri {Oh, 2010 #380} and the colonization phenotypes in LF mice imply that lineages maintained a broader host range and evolved with at least two diverse host genera (Mus and Rattus), and probably many species (around 40% of the world's mammalian species are rodents). Such an evolutionary strategy would require individual cells to adapt not only to physiological and immunological differences of individual animals but different host genera, and the larger accessory genome of the rodent L. reuteri population might reflect a higher diversity among the host population.
Reductive Evolution and a Population Bottleneck in Human L. reuteri Strains
The ecological forces that have shaped the autochthonous L. reuteri population in the human GIT appear fundamentally different than those in other hosts. Strains within the human-specific MLST lineage II, although obtained from world-wide locations, are highly conserved genetically and are clonally related [5], [20], suggesting a recent population bottleneck, founder effect, or clonal expansion. The genomic comparison of strain 100-23 and F275 further revealed that human strains underwent a process of reductive genome evolution. These evolutionary patterns resemble to some degree those found for genetically monomorphic pathogens, such as Yersinia pestis and Mycobacterium leprae [67]–[69], which show high clonality and genome evolution characterized by functional gene loss.
We can only speculate on what caused the specific genetic features of the human L. reuteri population. It has been suggested that the evolution of monomorphic pathogens was influenced by an expansion of the human population within the last 10,000–20,000 years, which possibly led to a significant increase of the available niche and a restriction to the human host [67]. The population bottleneck might also have been caused through altered transmission dynamics and changes in the human environment, which could have reduced the effective population size [5]. Low population sizes favor genetic drift and can lead to both decreased genetic variability [70] and the loss of genes (even if slightly beneficial) [71]. Alternatively, L. reuteri might have been acquired by humans more recently. Restriction to particular hosts or host changes have both been accompanied with a clonal population structure and functional gene loss, especially those associated with the cell envelope [72]–[75]. As described above, the genome of F275 shows clear evidence for pseudogene formation, gene deletions, and genome reduction, and although we do not yet know the causes of these patterns, the dramatic removal of surface proteins L. reuteri F275 suggests a process by which to bypass deleterious responses from the human immune system.
Concluding Remarks
The gut of vertebrates provides a multitude of nutrient rich habitats inhabited by complex microbial communities, whose composition is remarkably host specific and stable [4], [76]. These communities are important for normal development and growth of the host, but must be acquired during each generation as most vertebrates are essentially germ-free at birth. This process is poorly understood but relevant as benefits to the host are increased by the correct selection of true mutualists and their stable maintenance over evolutionary time [5], [29]. This study clearly established host specificity within the species L. reuteri through a combination of animal experiments and evolutionary genomics, and it revealed a first insight into the genomic changes that underlie host adaptation. Host specificity of L. reuteri in the mouse gut appears to be mediated to a large degree by specific adhesins. However, other factors are likely to contribute to host specificity and include adaptations to the environmental conditions (the urease cluster, Ig-A protease, factors for biofilm formation) and their regulation (possibly through TCS involved in quorum sensing).
In the last decade, our understanding of genome evolution in host-associated bacteria has advanced dramatically due to the availability of hundreds of sequenced genomes [77]. Common trends have been identified and range from those observed in obligate bacterial symbionts, who show extensive reductive genome evolution, to those of facultative symbionts with free-living stages, who have expanded genomes and high levels of LGT [77]–[79]. Genome evolution of L. reuteri shares some patterns that have been observed in other host associated bacteria, and the findings suggest an evolutionary intermediate transitioning from a facultative to an obligate, mutualisitic lifestyle, which concurs with the observed degree of host specialization. Accordingly, the high amount of mobile elements (e.g. IS elements) in L. reuteri genomes is a characteristic that is often associated with recent obligate host associations in bacteria [31]. Although mobile elements are common in all L. reuteri genomes, there are distinct trends of genome evolution in the rodent and human lineages, with the former possessing a large and adaptable pan-genome while the latter being subjected to a process of reductive evolution. These distinctions are likely related to differences in the microbe's host range and the ecology and genetic diversity of the host population.
Taken together, the results of this study revealed host adapted subpopulations among the species L. reuteri whose genome content reflected niche characteristics in their respective hosts. Although physiological and immunological differences of vertebrates were likely to constitute important selective forces that drove this specialization, the distinct patterns of genome evolution in rodent and human lineages suggest that the evolutionary trajectories of a vertebrate gut symbiont are not only determined by microbial competition but also by the ecology and evolutionary history of the host.
Materials and Methods
Ethics Statement
All animal experiments were approved by the Otago University Animal Ethics Committee (approval number 2/09).
Strains, Media, and Growth Conditions
Lactobacillus reuteri strains used in this study are listed in Table S3 and were grown anaerobically on MRS (Difco) plus 5g/L Fructose and 10g/L Maltose at 37°C or 45°C (where indicated). Escherichia coli EC1000, which was used for cloning vectors for gene inactivation in L. reuteri, was grown aerobically in LB media at 37°C. Erythromycin (200 µg/mL for E. coli, 5 µg/mL for Lactobacilli), kanamycin (40 µg/mL for E. coli), and chloramphenicol (7.5 µg/mL for lactobacilli) were used for the propagation of recombinant strains. L. reuteri 100-23C, which is a plasmid-free derivative of strain 100-23, was used to test the ecological relevance of selected genes (see below).
Determination of Colonization Phenotype in LF mice
LF mice were raised under gnotobiotic conditions, and the absence of lactobacilli was regularly tested by anaerobic culture on Rogosa SL agar for 48 hours. Mice (around 6 weeks of age) were inoculated by gavage on a single occasion with ∼106 Lactobacillus cells that had been cultured anaerobically in MRS medium overnight. Cell numbers of lactobacilli in fecal samples, the forestomach, and the cecum were determined by quantitative culture on Rogosa SL agar as described previously [39].
Genome Sequencing
Sequencing of L. reuteri 100-23 (rodent isolate) and DSM20016T (human isolate F275) genomes were accomplished through the Community Sequencing Program of the Joint Genome Institute (Walnut Creek, CA), using a combination of whole-genome shotgun sequencing of three libraries with 3-Kb, 8-Kb, and 40-Kb DNA inserts. The genomes were further sequenced using a Roche Genome Sequencer (FLX-GS) to reduce the amount of contigs, and gaps were closed manually by sequencing PCR products generated from the ends of contigs. This process resulted in a circular genome for DSM20016T and two scaffolds for 100-23 (729,351 bp and 1,576,206 bp). PCR reactions to amplify the DNA between these scaffolds failed on several attempts, probably due to the highly repetitive nature of the termini. Genomes were annotated using the JGI annotation pipeline, and the genome sequences have been deposited in GenBank under the accession numbers NC_009513 (strain DSM20016T) and NZ_AAPZ00000000 (strain 100-23).
The genomes of L. reuteri lpuph1 and MLC3 (rodent isolates) were sequenced to draft status at the Core for Applied Genomics and Ecology (CAGE, University of Nebraska, Lincoln, USA) with a standard shotgun library prep kit of the Roche GS FLX Titanium series. The genome of L. reuteri ATCC53608 (pig isolate) was sequenced at the Biotechnology and Biological Research Council's TGAC (The Genome Analysis Centre, Norwich Research Park, UK). Sequencing resulted in 185,905 (lpuph1), 115,542 (MLC3), and 617,241 (ATCC53608) reads that were assembled de novo using the gsAssembler (Newbler) module of the GS-FLX Off-Instrument Sofware Suite. This resulted in draft sequences of 127, 126, and 142 contigs, for lpuph1, MLC3, and ATCC53608 respectively. The draft sequencing resulted in a final coverage of around 30 fold (lpuph1), 20 fold (MLC3), and 100 fold (ATCC53608). The genome characteristics are listed in Table S6. Genome sequences for mlc3 and lpuph are available at DDBJ/EMBL/GenBank under the accession numbers AEAW00000000 and AEAX00000000, respectively. Genome sequences for ATCC 53608 are available at EMBL under the accession numbers CACS01000001 to CACS01000142.
Genome Analysis and Comparison
L. reuteri F275 was isolated in the 1960s and later deposited in both the Japanese and German culture collections. The genome sequence of the strain deposited in the Japan Collection of Microorganisms (JCM1112T) was recently published [34]. In the present study, the strain deposited in the Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSM20016T) was sequenced. As described by Morita and coworkers, strain DSM20016T has undergone genomic modifications of its genome during in vitro propagation [34], and as a consequence, JCM1112T contains two additional regions (a total of 40 kb) compared to the genome of DSM20016T. For the analysis and comparisons of gene content in 100-23 and F275, we used the genome annotations of strains 100-23 and DSM20016T, as they were both done with the JGI annotation pipeline. The genes that were encoded by the extra sequence identified in JCM112T were added to the gene set of DSM20016T and considered in all comparisons.
The Integrated Microbial Genomes (IMG) system of the JGI was used to analyze genome characteristics and compare genomes [80]. Unique and conserved genes between strains 100-23 and F275 were determined by the BLASTP algorithm implemented in the IMG Phylogenetic Profiler with a maximum E value of 1e−5 and a minimum amino acid identity of 70%. Whole genome comparisons were completed using the Artemis Comparison Tool (ACT) [81]. For this analysis, the two remaining scaffolds of the 100-23 genome were combined and the chromosome replication initiation site was identified. Visual genome comparisons of the genomes of strains 100-23 and JCM1112T were prepared by using ACT (BLASTN with a score cutoff of 1900). Alien_hunter was used to identify areas affected by LGT [82]. This program utilizes interpolated variable order motifs to identify regions of the genome with atypical sequence composition and thus integrates codon, and nucleotide compositional changes into its predictions.
BLASTP was used to identify homologous genes (>70% identity, >70% coverage) found in all L. reuteri strains and the closely-related L. vaginalis. Nucleotide sequences for these 169 orthologous genes were individually aligned in MUSCLE and concatenated and used to calculate the average nucleotide identity (ANI) as described by Konstantinidis and Tiedje [83]. The same BLASTP criteria were applied to determine the core and accessory genomes of L. reuteri strains (Figure 8).
Comparative Genome Hybridization Using Spotted Microarrays
Spotted microarrays were designed to contain probes representing all detected open reading frames (ORFs) of the rodent strain 100-23 and ORF unique to strain F275 when compared to 100-23. The phylogenetic profiler tool of the IMG platform was used to identify unique genes of F275 (using the sequence of strain DSM20016T) with a maximum E value of 1e−10 and an amino acid percentage of less than 90%. This analysis revealed 403 unique genes for F275. Probes (60 bp) were designed for all ORFs of sufficient size by using Oligo Array 2.1 [84]. Multiple probes were designed for genes of 100-23 larger than 4.5 kb (3 per gene). In total, the probe set comprised 2192 probes representing 2170 genes of strain 100-23 and 320 probes representing 320 genes of F275. Oligomers were synthesized by Invitrogen (Carlsbad, CA USA) and spotted in duplicate using an Omnigrid arrayer (Gene Machines, San Carlos, California).
L. reuteri strains used in microarray typing are listed in Table S3, which include 24 isolates from humans (including DSM20016T), 24 from rodents (including 100-23), 5 pig isolates, and 5 chicken isolates. Chromosomal DNA of bacteria was prepared as described by Oh and coworkers [20]. DNA of strains 100-23 and DSM20016T was mixed at a 1∶1 ratio, and 2 µg was amplified by random priming using Cy5 dye-labeled nucleotides and the BioPrime DNA labeling kit (Life Technologies, Rockville, Md.) to generate the reference DNA. Test DNA was generated by random priming PCR from all strains with Cy3 dye-labeled nucleotides. Concentrated labeled products from each reference test pair were hybridized in formamide-containing buffer (Array Hyb Low Temp; Sigma, St. Louis, Mo.) for 4 h at 47°C. Slides were washed once each in 1× SSC (0.15 M NaCl plus 0.015 M sodium citrate)−0.03% sodium dodecyl sulfate, 0.2× SSC, and finally 0.05× SSC. Fluorescence intensities of the array addresses were determined using a GenePix4000 multicolor microarray scanner and GenePix software (Axon Instruments, Union City, CA USA).
Genome content comparisons were performed using MARKFIND, as described by Zhang et al 2003 [45]. MARKFIND performs a cluster analysis based on genome polymorphisms implementing the unweighted pair-group method with arithmetic means (UPGMA). The program also uses an algorithm for sorting polymorphic characters in the binary strings relative to user-specified groups of taxa. For those genes being represented by three probes (i.e. large surface proteins), the gene was marked as present if at least two probes showed hybridization.
The accuracy of the microarray analysis was tested by comparing the results obtained by hybridizations with whole genome BLASTN comparisons. BLASTN was performed by comparing all gene sequences that are represented on the microarray slide with the genome sequences of L. reuteri 100-23 (rodent III cluster), lpuph1 (rodent I), F275 (human cluster II), and CF48-3A1 (human/chicken cluster IV). Genes were considered present if BLASTN resulted in alignments with more than 70% identity and at least 50% coverage to the query sequence. This analysis revealed that the microarray analysis had a very high accuracy for the two reference strains, showing >96.7% and 96.3% accuracy for 100-23 and DSM20016T, respectively. The accuracy dropped for strain lpuph1 to 92.5%, and it was lowest for strain CF48-3a (81.5%). So as expected, the accuracy of the microarray analysis decreased as gene divergence between the test and reference strains increased (see Table S7 for ANIs).
Confirmation of Rodent-Specific Gene Polymorphisms by PCR
Eighty-eight L. reuteri strains from all known MLSA lineages of the species (Table S3) were tested by PCR for the presence of representative rodent-specific genes: surface proteins (Lr_70131, Lr_70581, Lr_70697, Lr_69916), secA2 (Lr_70892), pduC (encoding a subunit of diol/glycerol dehydratase, the first enzyme in the propanediol fermentation/reuterin formation pathway), and ureC (encoding the urease alpha subunit). Primers were constructed based on the sequences of all strains that possessed the gene to first amplify an internal region of the gene, and second, to target the flanking genes and amplify the loci in which the gene was located in strain 100-23. The PCRs were carried out in 25 µl volumes containing 20 pmol of each primer and 0.5 units of Taq polymerase (Takara). After an initial denaturation for 3 min at 94°C, the reaction mixtures were cycled 30 times at 94°C for 30 s, 30 s at appropriate annealing temp, and 72°C for 3 min, followed by a 7-min extension at 72°C. Primer sequences and annealing temperatures are listed in Table S9.
Determination of the Ecological Relevance of Genes in L. reuteri 100-23C
The contribution of genes for ecological performance was determined as described previously [37]. Briefly, genes were inactivated in strain 100-23C by insertional mutagenesis by inserting the plasmid pORI28 into the target sites, which renders the mutant erythromycin-resistant. 1∶1 mixtures of mutant and wild type were administered by intragastric gavage to anesthetized LF mice. The mice were killed 7 days after inoculation, and lactobacilli were cultured quantitatively from the forestomach and cecum. To determine the proportion of the mutant strain, lactobacilli were quantified on agar plates with and without erythromycin.
Supporting Information
Genome comparisons between L. reuteri strains and Lactobacillus fermentum. Comparisons of the genomes of Lactobacillus reuteri 100-23 (top) and F275 (bottom) with that of Lactobacillus fermentum IFO3956 (center). The BLASTN comparison using the Artemis Comparison Tool revealed an inversion in 100-23 when compared to the other two genomes.
(3.36 MB PDF)
Accessory SecA2 cluster in L. reuteri and related bacteria. SecA2 cluster in L. reuteri 100-23, Lactobacillus gasseri JV-V03, and Streptococcus agalactiae A909. Conserved regions are labeled with grey boxes.
(0.31 MB PDF)
Gene maps of SPS clusters in L. reuteri 100-23 and F275. Genomic regions that encode for proteins involved in surface polysaccharide synthesis in L. reuteri 100-23 and F275. Flanking regions that are conserved in both genomes are indicated by boxes.
(0.34 MB PDF)
Comparison of genomic locations that contain host-specific genes. (A) Genome region with the pdu-cbi-cob-hem cluster in F275 and the same genomic region in 100-23. (B) Genome schematic showing the urease cluster in strain 100-23, and the same loci in F275. (C) Genomic region containing the xylose operon in 100-23 and the same loci, without xylose operon, in F275.
(0.38 MB PDF)
Comparison of the genomic locations that contain large surface proteins in L. reuteri 100-23. Comparison of the sites in strains 100-23 and F275 that contain rodent-specific large surface proteins. (A) Lr_70770, (B) Lr_70697, (C) Lr_70131, Lr_70134, and Lr_70135, (D) Lr_70581, and (E) Lr_71380.
(0.43 MB PDF)
General genome features of Lactobacillus reuteri strains 100-23 and F275.
(0.01 MB DOCX)
Unique genes in L. reuteri 100-23 and F275 in pair-wise comparisons.
(0.01 MB DOCX)
Strains used in this study.
(0.02 MB DOCX)
Rodent-specific genes, as detected by MARKFIND.
(0.03 MB XLSX)
Genes specific to the human MLSA lineage II.
(0.01 MB XLSX)
Characteristics of the nine Lactobacillus reuteri genomes included in this study.
(0.01 MB DOCX)
Average Nucleotide Identity (ANI) between genomes.
(0.01 MB DOCX)
Unique genes in the genomes of the rodent strains 100-23, mlc3, and lpuph.
(0.01 MB DOCX)
PCR primers used in this study to confirm host-specificity of genes.
(0.01 MB DOCX)
Footnotes
The authors have declared that no competing interests exist.
This study was funded by seed grants from the University of Nebraska, BioGaia, USDA NIFA Hatch (Acc No 0212027), and the Biotechnology and Biological Research Council UK. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Ley RE, Peterson DA, Gordon JI. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell. 2006;124:837–848. doi: 10.1016/j.cell.2006.02.017. [DOI] [PubMed] [Google Scholar]
- 2.Cerutti A, Rescigno M. The biology of intestinal immunoglobulin A responses. Immunity. 2008;28:740–750. doi: 10.1016/j.immuni.2008.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dethlefsen L, McFall-Ngai M, Relman DA. An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature. 2007;449:811–818. doi: 10.1038/nature06245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, et al. Evolution of mammals and their gut microbes. Science. 2008;320:1647–1651. doi: 10.1126/science.1155725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Walter J, Britton RA, Roos S. Microbes and Health Sackler Colloquium: Host-microbial symbiosis in the vertebrate gastrointestinal tract and the Lactobacillus reuteri paradigm. Proc Natl Acad Sci U S A. 2010 doi: 10.1073/pnas.1000099107. Epub ahead of print. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.McFall-Ngai M. Adaptive immunity: care for the community. Nature. 2007;445:153. doi: 10.1038/445153a. [DOI] [PubMed] [Google Scholar]
- 7.Stevens CE, Hume ID. Contributions of microbes in vertebrate gastrointestinal tract to production and conservation of nutrients. Physiol Rev. 1998;78:393–427. doi: 10.1152/physrev.1998.78.2.393. [DOI] [PubMed] [Google Scholar]
- 8.Ley RE, Lozupone CA, Hamady M, Knight R, Gordon JI. Worlds within worlds: evolution of the vertebrate gut microbiota. Nat Rev Microbiol. 2008;6:776–788. doi: 10.1038/nrmicro1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sela DA, Chapman J, Adeuya A, Kim JH, Chen F, et al. The genome sequence of Bifidobacterium longum subsp. infantis reveals adaptations for milk utilization within the infant microbiome. Proc Natl Acad Sci U S A. 2008;105:18964–18969. doi: 10.1073/pnas.0809584105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Xu J, Mahowald MA, Ley RE, Lozupone CA, Hamady M, et al. Evolution of symbiotic bacteria in the distal human intestine. PLoS Biol. 2007;5:e156. doi: 10.1371/journal.pbio.0050156. 10.1371/journal.pbio.0050156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, et al. The human microbiome project. Nature. 2007;449:804–810. doi: 10.1038/nature06244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Linz B, Balloux F, Moodley Y, Manica A, Liu H, et al. An African origin for the intimate association between humans and Helicobacter pylori. Nature. 2007;445:915–918. doi: 10.1038/nature05562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tenaillon O, Skurnik D, Picard B, Denamur E. The population genetics of commensal Escherichia coli. Nat Rev Microbiol. 2010;8:207–217. doi: 10.1038/nrmicro2298. [DOI] [PubMed] [Google Scholar]
- 14.Touchon M, Hoede C, Tenaillon O, Barbe V, Baeriswyl S, et al. Organised genome dynamics in the Escherichia coli species results in highly diverse adaptive paths. PLoS Genet. 2009;5:e1000344. doi: 10.1371/journal.pgen.1000344. 10.1371/journal.pgen.1000344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Brooks SP, McAllister M, Sandoz M, Kalmokoff ML. Culture-independent phylogenetic analysis of the faecal flora of the rat. Can J Microbiol. 2003;49:589–601. doi: 10.1139/w03-075. [DOI] [PubMed] [Google Scholar]
- 16.Leser TD, Amenuvor JZ, Jensen TK, Lindecrona RH, Boye M, et al. Culture-independent analysis of gut bacteria: the pig gastrointestinal tract microbiota revisited. Appl Environ Microbiol. 2002;68:673–690. doi: 10.1128/AEM.68.2.673-690.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Salzman NH, de Jong H, Paterson Y, Harmsen HJ, Welling GW, et al. Analysis of 16S libraries of mouse gastrointestinal microflora reveals a large new group of mouse intestinal bacteria. Microbiology. 2002;148:3651–3660. doi: 10.1099/00221287-148-11-3651. [DOI] [PubMed] [Google Scholar]
- 18.Tannock GW. Lactic microbiota of pigs, mice and rats. In: Wood BJB, editor. The Lactic Acid Bacteria in Health and Disease. London: Elsevier Applied Science; 1992. pp. 21–48. [Google Scholar]
- 19.Walter J. Ecological role of lactobacilli in the gastrointestinal tract: implications for fundamental and biomedical research. Appl Environ Microbiol. 2008;74:4985–4996. doi: 10.1128/AEM.00753-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Oh PL, Benson AK, Peterson DA, Patil PB, Moriyama EN, et al. Diversification of the gut symbiont Lactobacillus reuteri as a result of host-driven evolution. ISME J. 2010;4:377–387. doi: 10.1038/ismej.2009.123. [DOI] [PubMed] [Google Scholar]
- 21.Carbajal N, Casas IA, Dobrogosz WJ. Effect of host-specific Lactobacillus reuteri on ileal tissue development in gnotobiotic BALB/c mice. Microbial Ecol Health Dis. 1999;11 (Abst.):184. [Google Scholar]
- 22.Casas IA, Dobrogosz WJ. Validation of the Probiotic Concept: Lactobacillus reuteri Confers Broad-spectrum Protection against Disease in Humans and Animals. Microb Ecol Health Dis. 2000;12:247–285. [Google Scholar]
- 23.Moller PL, Paerregaard A, Gad M, Kristensen NN, Claesson MH. Colitic scid mice fed Lactobacillus spp. show an ameliorated gut histopathology and an altered cytokine profile by local T cells. Inflamm Bowel Dis. 2005;11:814–819. doi: 10.1097/01.mib.0000175906.77340.15. [DOI] [PubMed] [Google Scholar]
- 24.Schreiber O, Petersson J, Phillipson M, Perry M, Roos S, et al. Lactobacillus reuteri prevents colitis by reducing P-selectin-associated leukocyte- and platelet-endothelial cell interactions. Am J Physiol Gastrointest Liver Physiol. 2009;296:G534–542. doi: 10.1152/ajpgi.90470.2008. [DOI] [PubMed] [Google Scholar]
- 25.Lin JH-C, Savage DC. Host specificity of the colonization of murine gastric epithelium by lactobacilli. FEMS Microbiol Letters. 1984;24:67–71. [Google Scholar]
- 26.Wesney E, Tannock GW. Association of rat, pig, and fowl biotypes of lactobacilli with the stomach of gnotobiotic mice. Microb Ecol. 1979;5:35–42. doi: 10.1007/BF02010576. [DOI] [PubMed] [Google Scholar]
- 27.Suegara N, Morotomi M, Watanabe T, Kawal Y, Mutai M. Behavior of microflora in the rat stomach: adhesion of lactobacilli to the keratinized epithelial cells of the rat stomach in vitro. Infect Immun. 1975;12:173–179. doi: 10.1128/iai.12.1.173-179.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tannock GW, Crichton C, Welling GW, Koopman JP, Midtvedt T. Reconstitution of the gastrointestinal microflora of Lactobacillus-free mice. Appl Environ Microbiol. 1988;54:2971–2975. doi: 10.1128/aem.54.12.2971-2975.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mandel MJ, Wollenberg MS, Stabb EV, Visick KL, Ruby EG. A single regulatory gene is sufficient to alter bacterial host range. Nature. 2009;458:215–218. doi: 10.1038/nature07660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Moran NA. Symbiosis as an adaptive process and source of phenotypic complexity. Proc Natl Acad Sci U S A. 2007;104(Suppl 1):8627–8633. doi: 10.1073/pnas.0611659104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Moran NA, Plague GR. Genomic changes following host restriction in bacteria. Curr Opin Genet Dev. 2004;14:627–633. doi: 10.1016/j.gde.2004.09.003. [DOI] [PubMed] [Google Scholar]
- 32.Moya A, Pereto J, Gil R, Latorre A. Learning how to live together: genomic insights into prokaryote-animal symbioses. Nat Rev Genet. 2008;9:218–229. doi: 10.1038/nrg2319. [DOI] [PubMed] [Google Scholar]
- 33.Dale C, Moran NA. Molecular interactions between bacterial symbionts and their hosts. Cell. 2006;126:453–465. doi: 10.1016/j.cell.2006.07.014. [DOI] [PubMed] [Google Scholar]
- 34.Morita H, Toh H, Fukuda S, Horikawa H, Oshima K, et al. Comparative genome analysis of Lactobacillus reuteri and Lactobacillus fermentum reveal a genomic island for reuterin and cobalamin production. DNA Res. 2008;15:151–161. doi: 10.1093/dnares/dsn009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nelson KE, Weinstock GM, Highlander SK, Worley KC, Creasy HH, et al. A catalog of reference genomes from the human microbiome. Science. 2010;328:994–999. doi: 10.1126/science.1183605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tannock GW, Ghazally S, Walter J, Loach D, Brooks H, et al. Ecological behavior of Lactobacillus reuteri 100-23 is affected by mutation of the luxS gene. Appl Environ Microbiol. 2005;71:8419–8425. doi: 10.1128/AEM.71.12.8419-8425.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Walter J, Chagnaud P, Tannock GW, Loach DM, Dal Bello F, et al. A high-molecular-mass surface protein (Lsp) and methionine sulfoxide reductase B (MsrB) contribute to the ecological performance of Lactobacillus reuteri in the murine gut. Appl Environ Microbiol. 2005;71:979–986. doi: 10.1128/AEM.71.2.979-986.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Walter J, Heng NC, Hammes WP, Loach DM, Tannock GW, et al. Identification of Lactobacillus reuteri genes specifically induced in the mouse gastrointestinal tract. Appl Environ Microbiol. 2003;69:2044–2051. doi: 10.1128/AEM.69.4.2044-2051.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Walter J, Loach DM, Alqumber M, Rockel C, Hermann C, et al. D-alanyl ester depletion of teichoic acids in Lactobacillus reuteri 100-23 results in impaired colonization of the mouse gastrointestinal tract. Environ Microbiol. 2007;9:1750–1760. doi: 10.1111/j.1462-2920.2007.01292.x. [DOI] [PubMed] [Google Scholar]
- 40.Hoffmann M, Rath E, Holzlwimmer G, Quintanilla-Martinez L, Loach D, et al. Lactobacillus reuteri 100-23 transiently activates intestinal epithelial cells of mice that have a complex microbiota during early stages of colonization. J Nutr. 2008;138:1684–1691. doi: 10.1093/jn/138.9.1684. [DOI] [PubMed] [Google Scholar]
- 41.Livingston M, Loach D, Wilson M, Tannock GW, Baird M. Gut commensal Lactobacillus reuteri 100-23 stimulates an immunoregulatory response. Immunol Cell Biol. 2010;88:99–102. doi: 10.1038/icb.2009.71. [DOI] [PubMed] [Google Scholar]
- 42.Heng NC, Bateup JM, Loach DM, Wu X, Jenkinson HF, et al. Influence of different functional elements of plasmid pGT232 on maintenance of recombinant plasmids in Lactobacillus reuteri populations in vitro and in vivo. Appl Environ Microbiol. 1999;65:5378–5385. doi: 10.1128/aem.65.12.5378-5385.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sriramulu DD, Liang M, Hernandez-Romero D, Raux-Deery E, Lunsdorf H, et al. Lactobacillus reuteri DSM 20016 produces cobalamin-dependent diol dehydratase in metabolosomes and metabolizes 1,2-propanediol by disproportionation. J Bacteriol. 2008;190:4559–4567. doi: 10.1128/JB.01535-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Reuter G. The Lactobacillus and Bifidobacterium microflora of the human intestine: composition and succession. Curr Issues Intest Microbiol. 2001;2:43–53. [PubMed] [Google Scholar]
- 45.Zhang C, Zhang M, Ju J, Nietfeldt J, Wise J, et al. Genome diversification in phylogenetic lineages I and II of Listeria monocytogenes: identification of segments unique to lineage II populations. J Bacteriol. 2003;185:5573–5584. doi: 10.1128/JB.185.18.5573-5584.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007;57:81–91. doi: 10.1099/ijs.0.64483-0. [DOI] [PubMed] [Google Scholar]
- 47.Li YH, Chen YY, Burne RA. Regulation of urease gene expression by Streptococcus salivarius growing in biofilms. Environ Microbiol. 2000;2:169–177. doi: 10.1046/j.1462-2920.2000.00088.x. [DOI] [PubMed] [Google Scholar]
- 48.Sjostrom JE, Larsson H. Factors affecting growth and antibiotic susceptibility of Helicobacter pylori: effect of pH and urea on the survival of a wild-type strain and a urease-deficient mutant. J Med Microbiol. 1996;44:425–433. doi: 10.1099/00222615-44-6-425. [DOI] [PubMed] [Google Scholar]
- 49.Rigel NW, Braunstein M. A new twist on an old pathway–accessory Sec [corrected] systems. Mol Microbiol. 2008;69:291–302. doi: 10.1111/j.1365-2958.2008.06294.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bensing BA, Sullam PM. Characterization of Streptococcus gordonii SecA2 as a paralogue of SecA. J Bacteriol. 2009;191:3482–3491. doi: 10.1128/JB.00365-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Bensing BA, Sullam PM. An accessory sec locus of Streptococcus gordonii is required for export of the surface protein GspB and for normal levels of binding to human platelets. Mol Microbiol. 2002;44:1081–1094. doi: 10.1046/j.1365-2958.2002.02949.x. [DOI] [PubMed] [Google Scholar]
- 52.Flynn S, van Sinderen D, Thornton GM, Holo H, Nes IF, et al. Characterization of the genetic locus responsible for the production of ABP-118, a novel bacteriocin produced by the probiotic bacterium Lactobacillus salivarius subsp. salivarius UCC118. Microbiology. 2002;148:973–984. doi: 10.1099/00221287-148-4-973. [DOI] [PubMed] [Google Scholar]
- 53.Kleerebezem M, Quadri LE, Kuipers OP, de Vos WM. Quorum sensing by peptide pheromones and two-component signal-transduction systems in Gram-positive bacteria. Mol Microbiol. 1997;24:895–904. doi: 10.1046/j.1365-2958.1997.4251782.x. [DOI] [PubMed] [Google Scholar]
- 54.Sturme MH, Francke C, Siezen RJ, de Vos WM, Kleerebezem M. Making sense of quorum sensing in lactobacilli: a special focus on Lactobacillus plantarum WCFS1. Microbiology. 2007;153:3939–3947. doi: 10.1099/mic.0.2007/012831-0. [DOI] [PubMed] [Google Scholar]
- 55.Talarico TL, Axelsson LT, Novotny J, Fiuzat M, Dobrogosz WJ. Utilization of Glycerol as a Hydrogen Acceptor by Lactobacillus reuteri: Purification of 1,3-Propanediol:NAD Oxidoreductase. Appl Environ Microbiol. 1990;56:943–948. doi: 10.1128/aem.56.4.943-948.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Talarico TL, Casas IA, Chung TC, Dobrogosz WJ. Production and isolation of reuterin, a growth inhibitor produced by Lactobacillus reuteri. Antimicrob Agents Chemother. 1988;32:1854–1858. doi: 10.1128/aac.32.12.1854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Fuller R, Barrow PA, Brooker BE. Bacteria associated with the gastric epithelium of neonatal pigs. Appl Environ Microbiol. 1978;35:582–591. doi: 10.1128/aem.35.3.582-591.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Fuller R, Brooker BE. Lactobacilli which attach to the crop epithelium of the fowl. Am J Clin Nutr. 1974;27:1305–1312. doi: 10.1093/ajcn/27.11.1305. [DOI] [PubMed] [Google Scholar]
- 59.Goodman AL, McNulty NP, Zhao Y, Leip D, Mitra RD, et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe. 2009;6:279–289. doi: 10.1016/j.chom.2009.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Santos F, Vera JL, van der Heijden R, Valdez G, de Vos WM, et al. The complete coenzyme B12 biosynthesis gene cluster of Lactobacillus reuteri CRL1098. Microbiology. 2008;154:81–93. doi: 10.1099/mic.0.2007/011569-0. [DOI] [PubMed] [Google Scholar]
- 61.Kikuchi Y, Hosokawa T, Nikoh N, Meng XY, Kamagata Y, et al. Host-symbiont co-speciation and reductive genome evolution in gut symbiotic bacteria of acanthosomatid stinkbugs. BMC Biol. 2009;7:2. doi: 10.1186/1741-7007-7-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Sullivan JT, Ronson CW. Evolution of rhizobia by acquisition of a 500-kb symbiosis island that integrates into a phe-tRNA gene. Proc Natl Acad Sci U S A. 1998;95:5145–5149. doi: 10.1073/pnas.95.9.5145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R. The microbial pan-genome. Curr Opin Genet Dev. 2005;15:589–594. doi: 10.1016/j.gde.2005.09.006. [DOI] [PubMed] [Google Scholar]
- 64.Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”. Proc Natl Acad Sci U S A. 2005;102:13950–13955. doi: 10.1073/pnas.0506758102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Reno ML, Held NL, Fields CJ, Burke PV, Whitaker RJ. Biogeography of the Sulfolobus islandicus pan-genome. Proc Natl Acad Sci U S A. 2009;106:8605–8610. doi: 10.1073/pnas.0808945106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Prosser JI, Bohannan BJ, Curtis TP, Ellis RJ, Firestone MK, et al. The role of ecological theory in microbial ecology. Nat Rev Microbiol. 2007;5:384–392. doi: 10.1038/nrmicro1643. [DOI] [PubMed] [Google Scholar]
- 67.Achtman M. Evolution, population structure, and phylogeography of genetically monomorphic bacterial pathogens. Annu Rev Microbiol. 2008;62:53–70. doi: 10.1146/annurev.micro.62.081307.162832. [DOI] [PubMed] [Google Scholar]
- 68.Holt KE, Parkhill J, Mazzoni CJ, Roumagnac P, Weill FX, et al. High-throughput sequencing provides insights into genome variation and evolution in Salmonella Typhi. Nat Genet. 2008;40:987–993. doi: 10.1038/ng.195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Chain PS, Carniel E, Larimer FW, Lamerdin J, Stoutland PO, et al. Insights into the evolution of Yersinia pestis through whole-genome comparison with Yersinia pseudotuberculosis. Proc Natl Acad Sci U S A. 2004;101:13826–13831. doi: 10.1073/pnas.0404012101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Bright M, Bulgheresi S. A complex journey: transmission of microbial symbionts. Nat Rev Microbiol. 8:218–230. doi: 10.1038/nrmicro2262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Moran NA. Tracing the evolution of gene loss in obligate bacterial symbionts. Curr Opin Microbiol. 2003;6:512–518. doi: 10.1016/j.mib.2003.08.001. [DOI] [PubMed] [Google Scholar]
- 72.Eppinger M, Baar C, Linz B, Raddatz G, Lanz C, et al. Who ate whom? Adaptive Helicobacter genomic changes that accompanied a host jump from early humans to large felines. PLoS Genet. 2006;2:e120. doi: 10.1371/journal.pgen.0020120. 10.1371/journal.pgen.0020120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Holden MT, Heather Z, Paillot R, Steward KF, Webb K, et al. Genomic evidence for the evolution of Streptococcus equi: host restriction, increased virulence, and genetic exchange with human pathogens. PLoS Pathog. 2009;5:e1000346. doi: 10.1371/journal.ppat.1000346. 10.1371/journal.ppat.1000346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Parkhill J, Sebaihia M, Preston A, Murphy LD, Thomson N, et al. Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica. Nat Genet. 2003;35:32–40. doi: 10.1038/ng1227. [DOI] [PubMed] [Google Scholar]
- 75.Lowder BV, Guinane CM, Ben Zakour NL, Weinert LA, Conway-Morris A, et al. Recent human-to-poultry host jump, adaptation, and pandemic spread of Staphylococcus aureus. Proc Natl Acad Sci U S A. 2009;106:19545–19550. doi: 10.1073/pnas.0909285106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, et al. Bacterial Community Variation in Human Body Habitats Across Space and Time. Science. 2009 doi: 10.1126/science.1177486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Medina M, Sachs JL. Symbiont genomics, our new tangled bank. Genomics. 2010;95:129–137. doi: 10.1016/j.ygeno.2009.12.004. [DOI] [PubMed] [Google Scholar]
- 78.Ruby EG, Urbanowski M, Campbell J, Dunn A, Faini M, et al. Complete genome sequence of Vibrio fischeri: a symbiotic bacterium with pathogenic congeners. Proc Natl Acad Sci U S A. 2005;102:3004–3009. doi: 10.1073/pnas.0409900102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Normand P, Lapierre P, Tisa LS, Gogarten JP, Alloisio N, et al. Genome characteristics of facultatively symbiotic Frankia sp. strains reflect host range and host plant biogeography. Genome Res. 2007;17:7–15. doi: 10.1101/gr.5798407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Markowitz VM, Szeto E, Palaniappan K, Grechkin Y, Chu K, et al. The integrated microbial genomes (IMG) system in 2007: data content and analysis tool extensions. Nucleic Acids Res. 2008;36:D528–533. doi: 10.1093/nar/gkm846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Carver TJ, Rutherford KM, Berriman M, Rajandream MA, Barrell BG, et al. ACT: the Artemis Comparison Tool. Bioinformatics. 2005;21:3422–3423. doi: 10.1093/bioinformatics/bti553. [DOI] [PubMed] [Google Scholar]
- 82.Vernikos GS, Parkhill J. Interpolated variable order motifs for identification of horizontally acquired DNA: revisiting the Salmonella pathogenicity islands. Bioinformatics. 2006;22:2196–2203. doi: 10.1093/bioinformatics/btl369. [DOI] [PubMed] [Google Scholar]
- 83.Konstantinidis KT, Tiedje JM. Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci U S A. 2005;102:2567–2572. doi: 10.1073/pnas.0409727102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Rouillard JM, Zuker M, Gulari E. OligoArray 2.0: design of oligonucleotide probes for DNA microarrays using a thermodynamic approach. Nucleic Acids Res. 2003;31:3057–3062. doi: 10.1093/nar/gkg426. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Genome comparisons between L. reuteri strains and Lactobacillus fermentum. Comparisons of the genomes of Lactobacillus reuteri 100-23 (top) and F275 (bottom) with that of Lactobacillus fermentum IFO3956 (center). The BLASTN comparison using the Artemis Comparison Tool revealed an inversion in 100-23 when compared to the other two genomes.
(3.36 MB PDF)
Accessory SecA2 cluster in L. reuteri and related bacteria. SecA2 cluster in L. reuteri 100-23, Lactobacillus gasseri JV-V03, and Streptococcus agalactiae A909. Conserved regions are labeled with grey boxes.
(0.31 MB PDF)
Gene maps of SPS clusters in L. reuteri 100-23 and F275. Genomic regions that encode for proteins involved in surface polysaccharide synthesis in L. reuteri 100-23 and F275. Flanking regions that are conserved in both genomes are indicated by boxes.
(0.34 MB PDF)
Comparison of genomic locations that contain host-specific genes. (A) Genome region with the pdu-cbi-cob-hem cluster in F275 and the same genomic region in 100-23. (B) Genome schematic showing the urease cluster in strain 100-23, and the same loci in F275. (C) Genomic region containing the xylose operon in 100-23 and the same loci, without xylose operon, in F275.
(0.38 MB PDF)
Comparison of the genomic locations that contain large surface proteins in L. reuteri 100-23. Comparison of the sites in strains 100-23 and F275 that contain rodent-specific large surface proteins. (A) Lr_70770, (B) Lr_70697, (C) Lr_70131, Lr_70134, and Lr_70135, (D) Lr_70581, and (E) Lr_71380.
(0.43 MB PDF)
General genome features of Lactobacillus reuteri strains 100-23 and F275.
(0.01 MB DOCX)
Unique genes in L. reuteri 100-23 and F275 in pair-wise comparisons.
(0.01 MB DOCX)
Strains used in this study.
(0.02 MB DOCX)
Rodent-specific genes, as detected by MARKFIND.
(0.03 MB XLSX)
Genes specific to the human MLSA lineage II.
(0.01 MB XLSX)
Characteristics of the nine Lactobacillus reuteri genomes included in this study.
(0.01 MB DOCX)
Average Nucleotide Identity (ANI) between genomes.
(0.01 MB DOCX)
Unique genes in the genomes of the rodent strains 100-23, mlc3, and lpuph.
(0.01 MB DOCX)
PCR primers used in this study to confirm host-specificity of genes.
(0.01 MB DOCX)