The ZEB1/miR-200 feedback loop controls Notch signalling in cancer cells
The ZEB1 transcription factor and miRNA-200 repress each others' expression, with important consequences for tumour metastasis. Here, Notch pathway components are found as miR-200 targets, identifying a pathway whereby ZEB1 promotes Notch signalling in cancer cells. See also Vallejo et al in this issue.
Keywords: EMT, miR-200, Notch, stemness, ZEB1
Abstract
Notch signalling is important for development and tissue homeostasis and activated in many human cancers. Nevertheless, mutations in Notch pathway components are rare in solid tumours. ZEB1 is an activator of an epithelial–mesenchymal transition (EMT) and has crucial roles in tumour progression towards metastasis. ZEB1 and miR-200 family members repress expression of each other in a reciprocal feedback loop. Since miR-200 members target stem cell factors, ZEB1 indirectly induces stemness maintenance and associated drug resistance. Here, we link ZEB1 and its cancer promoting properties to Notch activation. We show that miR-200 members target Notch pathway components, such as Jagged1 (Jag1) and the mastermind-like coactivators Maml2 and Maml3, thereby mediating enhanced Notch activation by ZEB1. We further detected a coordinated upregulation of Jag1 and ZEB1, associated with reduced miR-200 expression in two aggressive types of human cancer, pancreatic adenocarcinoma and basal type of breast cancer. These findings explain increased Notch signalling in some types of cancers, where mutations in Notch pathway genes are rare. Moreover, they indicate an additional way how ZEB1 exerts its tumour progressing functions.
Introduction
Notch signalling is important for embryonic development and adult tissue homeostasis and if aberrantly activated has a crucial role in the development of many human cancers. The Notch pathway is activated by binding of Notch ligands (Jag1, Jag2, DLL1, DLL3, DLL4) to Notch receptors, which are subsequently cleaved extracellularly by ADAM-type proteases, and intracellularly by γ-secretase. The resulting Notch intracellular domain (NICD) translocates to the nucleus, where it builds up a transcription activator complex together with the transcription factor CSL and coactivators, such as mastermind-like factors (Maml 1,2,3). This complex activates the transcription of typical Notch target genes like Hes and Hey1 (Koch and Radtke, 2007). The Notch pathway controls central cellular processes including stemness, differentiation, proliferation and survival (Hurlbut et al, 2007; Roy et al, 2007). This may explain why aberrant Notch pathway activation is described for many human cancer types, including lung, colorectal and breast cancer, as well as leukaemias (Koch and Radtke, 2007; Rizzo et al, 2008). Its relevance was also shown in mouse models for pancreatic cancer, where inhibition of Notch signalling by a γ-secretase inhibitor (GSI) completely blocked tumour formation (Plentz et al, 2009).
Epithelial–mesenchymal transition (EMT) is a reversible embryonic program, which allows partial or complete transition between an epithelial and a mesenchymal phenotype. EMT is essential for embryonic processes like gastrulation, and if aberrantly activated is a trigger of tumour progression and metastasis. EMT is activated by key signalling pathways, including the TGFβ and FGF pathway, converging in the stimulation of EMT activators, a group of transcription factors repressing epithelial gene expression. These include members of the snail family, of the bHLH family and of the ZFH family (ZEB1 and ZEB2) (Thiery et al, 2009). It is now known that EMT activators not only activate cellular motility, but are also associated with the maintenance of stem cell properties and cell survival (Mani et al, 2008; Morel et al, 2008). We and others have shown that thereby the miR-200 family of microRNAs has a crucial role (Shimono et al, 2009; Wellner et al, 2009). ZEB1 suppresses the expression of all miR-200 family members (miR-141, -200a,b,c and -429), which in turn inhibit translation of ZEB1 mRNA, resulting in the double-negative ZEB/miR-200 feedback loop (Burk et al, 2008; Gregory et al, 2008; Korpal et al, 2008; Park et al, 2008; Brabletz and Brabletz, 2010). Since stem cell factors, such as Bmi1, are additional targets of miR-200, an overexpression of ZEB1 in tumour cells indirectly, through inhibition of miR-200 expression, leads to maintenance of stem cell properties, as shown for breast and pancreatic cancer (Shimono et al, 2009; Wellner et al, 2009).
Thus, it emerged that EMT and Notch signalling have many functional overlaps by controlling central processes such as stemness and cell survival. An obvious link is indicated by the finding that Notch signalling can also induce EMT in cell culture (Leong et al, 2007; Sahlgren et al, 2008). However, albeit the importance of Notch signalling for tumour formation and malignant progression, mutations in Notch pathway components are rarely detected in solid cancers, including human pancreatic adenocarcinomas, as analysed in a recent comprehensive survey of multiple genomes (Jones et al, 2008). Thus, other ways of aberrant Notch signalling activation must exist in cancer. We investigated if EMT activation can stimulate the Notch pathway and show here that the EMT inducer ZEB1 can trigger Notch signalling in cancer cells by stabilizing the expression of Notch pathway components, such as Jag1, Maml2 and Maml3, through inhibition of miR-200 expression.
Results
Knockdown of ZEB1 expression reduces Notch pathway activity
We have previously shown that, in addition to classical EMT-associated properties, the EMT activator ZEB1 also controls other central cellular processes and states, such as stemness and survival, by inhibiting expression of miR-200 family members (Wellner et al, 2009). Since the Notch signalling pathway is known to control these processes as well, we investigated if the EMT inducer ZEB1 can influence Notch signalling. Stable knockdown of ZEB1 decreased the endogenous activity of a Notch reporter construct (4xCSL-Luc) in the undifferentiated pancreatic cancer cell line Panc1 (Figure 1A). A mutated control reporter was not significantly affected by ZEB1. Also a transient, siRNA-mediated knockdown of ZEB1 led to a reduced Notch reporter activity (Supplementary Figure S1A). We re-analysed whole-genome gene expression arrays of stable ZEB1 knockdown clones performed earlier in our laboratory (Spaderna et al, 2008) for Notch pathway-related genes affected by ZEB1. Decrease of ZEB1 expression led to a downregulation of mRNAs for the Notch ligand Jag1, the Notch transcriptional coactivators Maml2 and Maml3, as well as the Notch target factor Hey1 in pancreatic and breast cancer cell lines (Figure 1B). Other Notch pathway components were not affected by ZEB1 in the investigated cell lines (not shown). We initially focused on Jag1, because it was already shown to be overexpressed in many cancer types, such as the basal type of breast cancer (Reedijk et al, 2008). Downregulation of Jag1 after both stable and transient knockdown of ZEB1 was confirmed by quantitative real-time PCR (Figure 1C–E). Moreover, also the expression of the Notch target Hey1 was reduced (Figure 1C and D). These data indicate that ZEB1 is enhancing Notch pathway activation, at least partially, by increasing the expression of the ligand Jag1.
Figure 1.
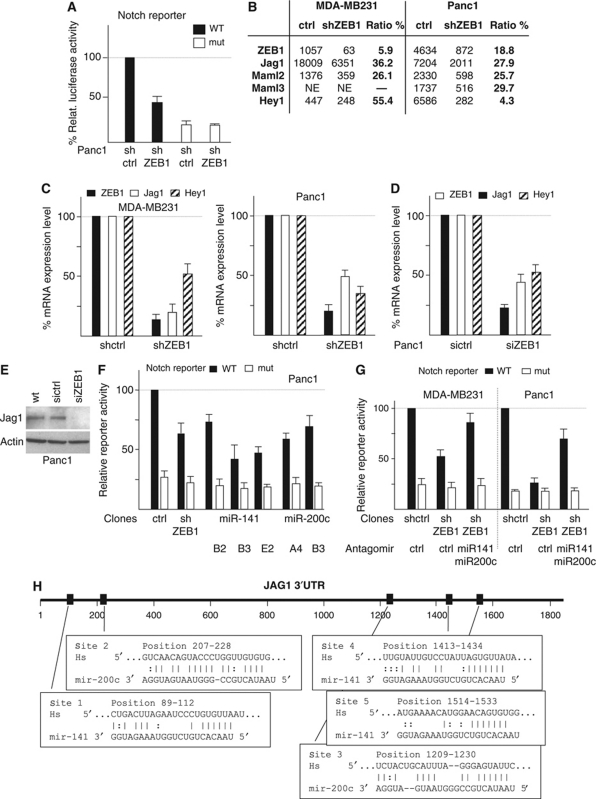
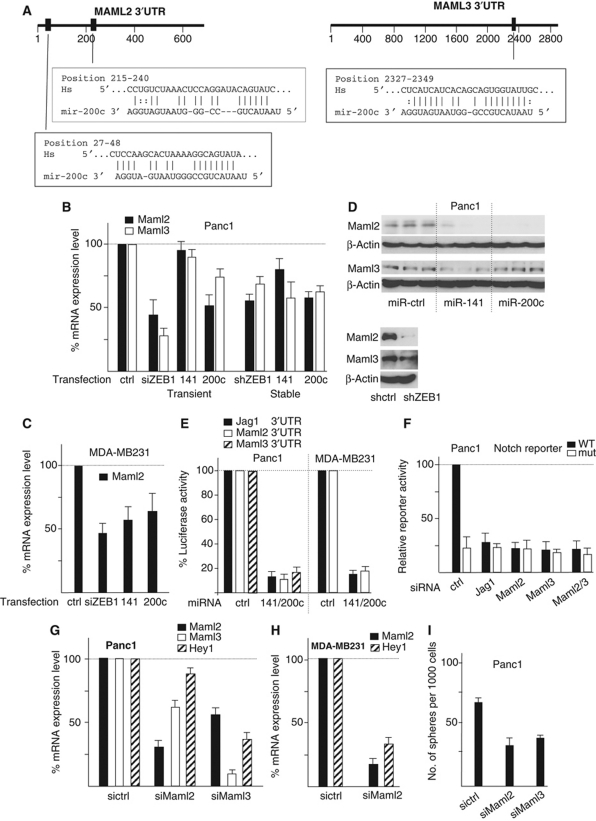
ZEB1 and miR-200 family members affect Notch signalling. (A) Stable knockdown of ZEB1 results in reduced Notch reporter activity (WT). There was no significant effect on a mutated (mut) reporter construct. Shown are mean values of each two independent clones. Control clones were set to 100%. (B) Expression values of ZEB1 and the indicated Notch pathway components deduced from a whole-genome expression screen comparing ZEB1 knockdown and control knockdown clones (NE= not expressed). (C, D) Quantitative RT–PCR after stable (C) and transient (D) knockdown of ZEB1. (E) Immunoblot showing reduced expression of Jag1 after transient knockdown of ZEB1. (F) Notch-luciferase reporter assay showing reduced activity in different clones after stable knockdown of ZEB1 or stable overexpression of miR-141 and miR-200c. (G) Antagomirs against miR-141 and miR-200c partially rescue Notch reporter inhibition by shZEB1. (H) Scheme showing five predicted target sites of miR-200 family members in the 3′UTR of Jag1.
Jag1 is a target of miR-200 family members
ZEB1-dependent expression of Notch pathway components is likely not due to direct activation by ZEB1, since it is mainly described as a transcriptional repressor (Vandewalle et al, 2009). Like previously shown for other genes indirectly activated by ZEB1, we also hypothesized that Jag1 is a miR-200 target and upregulated upon inhibition of miR-200 genes by ZEB1. We first analysed if miR-200 family members can affect Notch signalling. The miR-200 family members can be subdivided into two subgroups according to their seed sequences (subgroup I: miR-141 and miR-200a; subgroup II: miR-200b, c and miR-429), which indicate slight differences in their target gene sets (Peter, 2009). Both stable and transient overexpression of miR-141 and miR-200c, belonging to subgroups I and II, respectively, reduced the activity of the Notch reporter, but not of the control reporter (Figure 1F; Supplementary Figure S1A). Moreover, reduced Notch reporter activity after ZEB1 knockdown could be partially restored by inhibiting miR-141 and miR-200c using specific antagomirs (Figure 1G). By using the microRNA target prediction server mirecords (http://mirecords.biolead.org), we found that Jag1 is a predicted target of miR-200 family members (Figure 1H). Stable overexpression of miR-141 and miR-200c in the undifferentiated pancreatic cancer cell line Panc1, lacking endogenous miR-200 expression, led to a slightly reduced expression of Jag1 at mRNA level (Figure 2A). Accordingly, a transient overexpression of miR-141 and miR-200c in Panc1 cells slightly reduced Jag1 mRNA expression (Supplementary Figure S1B). Using immunoblots, we could show that knockdown of ZEB1 and overexpression of both miR-141 and miR-200c led to a reduction of Jag1 protein expression (Figure 2B). Vice versa, we used the differentiated pancreatic cancer cell line HPAF2 and breast cancer cell line MCF7, which endogenously express miR-200 members. Inhibition of their function by specific antagomirs led to an increase in Jag1 mRNA and in particular protein level (Figure 2C and D). We further constructed a luciferase reporter vector under the control of the Jag1 3′UTR to show direct effects of miR-200 members. Both miR-141 and miR-200c inhibited the Jag1 3′UTR driven reporter activity. The ZEB1 3′UTR reporter construct served as a known positive control, in particular for miR-200c (Figure 2E). Again, we used the differentiated line HPAF2 to confirm the inhibitory effect of endogenous miR-200 expression on the Jag1 3′UTR. Subsequent mutations of predicted miR-200 binding sites 1 to 5 steadily increased Jag1 3′UTR reporter activity, indicating that all five detected binding sites can confer to the inhibitory function of miR-200 (Figure 2F). Additionally, antagomirs against both miR-200c and miR-141 increased the activity of the Jag1 3′UTR reporter in HPAFII cells, and thus further confirmed that Jag1 is a direct target of miR-200 (Figure 2G).
Figure 2.
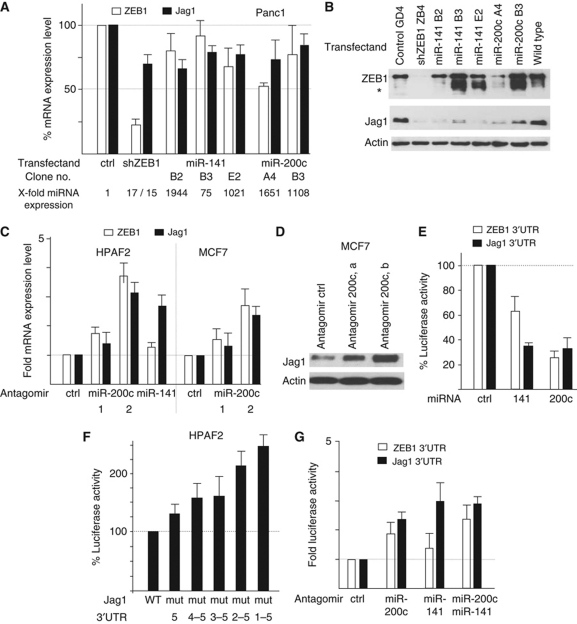
Jag1 is a target of miR-200 family members. (A, B) Quantitative RT–PCR (A) and immunoblot (B) for ZEB1 and Jag1 after stable knockdown of ZEB1 or overexpression of miR-141 and 200c in Panc1 cells (* is a degradation product of ZEB1). Also shown is the relative expression of the indicated microRNAs in the different clones. (C, D) Inhibition of endogenous miRs by the indicated specific antagomirs increase the mRNA levels of ZEB1 and Jag1 (C) and the protein level of Jag1 (D) in differentiated cancer cell lines. (E) Transient overexpression of miR-141 or miR-200c reduced the activity of Jag1 3′UTR reporter and a ZEB1 3′UTR control reporter in Panc1 cells. (F) Subsequent mutations of predicted miR-200 binding sites in the Jag1 3′UTR-luciferase construct increased reporter activity in HPAFII cells, endogenously expressing miR-200 family members. (G) Inhibition of endogenous miRs in HPAFII cells by the indicated specific antagomirs (antagomir miR-200c (b) was used) increased the activity of the Jag1 3′UTR reporter and a ZEB1 3′UTR control reporter.
Notch activity and Jag1 expression are important for essential properties of cancer cells
Next, we wanted to analyse if Notch signalling, and in particular expression of the Notch ligand Jag1, are important for crucial properties of cancer cells. Notch signalling was blocked by treatment of tumour cells with a γ-secretase inhibitor (GSI), and Jag1 expression was transiently knocked down by two different, specific siRNAs (Supplementary Figure S1B). By applying Notch reporter assays and by detection of activated (cleaved) Notch2, we could demonstrate that both GSI treatment and the knockdown of Jag1 inhibited Notch signalling (Figure 3A and B). Inhibition of Notch signalling by Jag1 knockdown was also confirmed by a reduced expression of the Notch target gene Hey1 (Supplementary Figure S1C and D). Moreover, we detected a reduced expression of ZEB1 after knockdown of Jag1, which confirms published data of ZEB1 expression being activated by Notch signalling (Wang et al, 2009). Both GSI treatment and knockdown of Jag1 also reduced the proliferative capacity of cancer cells (Figure 3C; Supplementary Figure S1E) and increased apoptosis after irradiation of cancer cells with 5 Gy (Figure 3D). Next, we analysed the effects on stemness-associated properties. We used the stem cell sphere assay to quantify the numbers of potential cancer stem cells in culture: if tumour cells are cultivated in suspension in a serum-free selection medium, only cancer cells with stem cell properties are able to grow out and form spheres, whereas all other cells undergo anoikis (Gou et al, 2007). The number of spheres formed per 1000 cultivated cells reflects the number of potential cancer stem cells. We have previously demonstrated that knockdown of ZEB1 or overexpression of miR-200c affects the sphere-formation capacity in breast and pancreatic cancer cells (Wellner et al, 2009). Here, we show that inhibition of Notch signalling by GSI treatment strongly inhibited the sphere-forming capacity of cancer cells (Figure 4A). This indicated that stem cell associated properties depend on Notch signalling, although the reduced sphere-forming capacity after GSI treatment might be an additive effect of reduced cancer stem cell numbers and reduced growth capacities. A transient knockdown of Jag1 had no effect on sphere formation in the first generation, however, led to a reduction of the sphere-forming capacity in the second-sphere generation (Figure 4B; Supplementary Figure S1F). This indicated that Jag1 expression is necessary for maintenance of stem cell properties, in particular for a self-renewal capacity.
Figure 3.
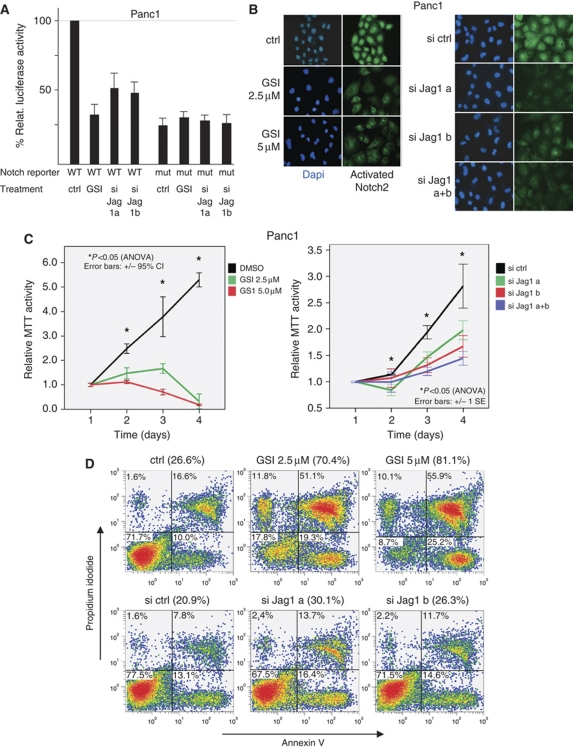
Notch signalling and Jag1 expression are important for cancer cell properties. (A) Treatment with 5 μM GSI or transient knockdown of Jag1 inhibits Notch reporter activity. (B) Immunofluorescence showing reduced nuclear expression of activated Notch2 after GSI treatment or transient Jag1 knockdown. (C) GSI treatment or transient knockdown of Jag1 inhibits proliferation measured in an MTT assay. X axis indicates days after cell seeding. Asterisks indicate significance of control versus treated cells. (D) GSI treatment or transient knockdown of Jag1 increases apoptosis of Panc1 cells irradiated with 5 Gy. The percentage of apoptotic cells is indicated, resulting from the addition of the two right, annexin V positive quadrants.
Figure 4.
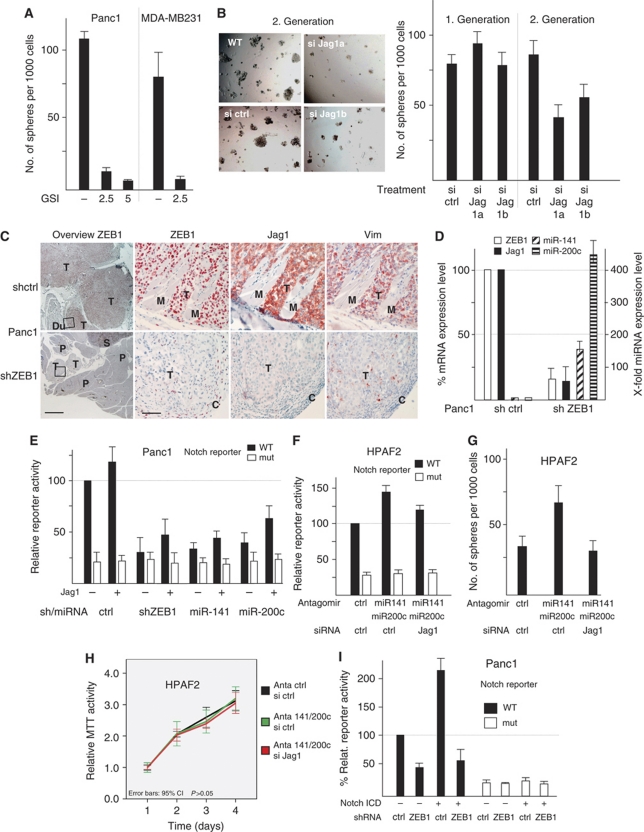
Jag1 alone is not sufficient for the effects of ZEB1 and miR-200 on Notch signalling. (A) GSI treatment inhibits the sphere-forming capacity. (B) Transient knockdown of Jag1 inhibits the sphere-forming capacity in the second, but not in the first generation of spheres in Panc1 cells. A representative picture of spheres from the second generation is shown. (C) Representative immunohistochemistry showing a huge control tumour (T) invading the duodenal wall (Du) and small, encapsulated, non-invasive tumour nodules of shZEB1 cells surrounded by pancreas (P) and spleen (S) tissue (overview). Squares indicate magnified areas showing a coordinated reduction of ZEB1, Jag1 and vimentin expression in tumours derived from ZEB1 knockdown cells, encapsulated by ZEB1 expressing fibroblasts (c) and high expression in control tumour cells invading the duodenal muscle layers (M). Size bar is 200 and 20 μm. (D) Quantitative RT–PCR after microdissection of orthotopic xenograft tumours showing decrease of Jag1 and increase of miR-141 and miR-200c after knockdown of ZEB1. Shown are the mean values of all grown tumours (six mice for control and four mice for ZEB1 knockdown Panc1 cell clones), control knockdown was set to 100% or 1. (E) Coexpression of Jag1 lacking the 3′UTR can only partially rescue Notch reporter inhibition by shZEB1 or miR-141 and miR-200c. (F) Inhibition of endogenous miR-141 and miR-200c in differentiated HPAF2 cells increases Notch reporter activity, which is only partially reversed by siRNA-mediated knockdown of Jag1. (G) Inhibition of endogenous miR-141 and miR-200c increases the second-generation sphere-forming capacity, which is reversed by siRNA-mediated knockdown of Jag1. (H) Proliferation of differentiated HPAF2 cells is not affected by antagomirs and Jag1 knockdown. (I) Cotransfection of the Notch ICD increases the Notch reporter activity in control clones, but not in stable ZEB1 knockdown clones. There was no significant effect on a mutated (mut) reporter construct. Shown are mean values of each two independent clones. Control clones were set to 100%.
We further analysed in vivo expression of ZEB1, miR-200 family members and Jag1 in orthotopic xenograft tumours derived from pancreatic cancer cells. We have previously shown that stable ZEB1 knockdown in the pancreatic cancer cell lines Panc1 and MiaPaCa2 affected important aspects of tumour formation. After ZEB1 knockdown, the cancer cells showed strong reduction in tumourigenicity and the few grown tumours were much smaller and completely lost their invasive and metastatic capacity (Wellner et al, 2009). By re-investigating these tumours, we detected a strongly decreased expression of Jag1 and the mesenchymal marker vimentin, and increased expression of miR-141 and miR-200c in tumours derived from ZEB1 knockdown clones. This correlated with their reduced tumourigenicity and aggressiveness (Figure 4C and D; Supplementary Figure S2A).
Jag1 alone cannot explain the effects of ZEB1 and miR-200 on Notch signalling
Next, we wanted to determine whether Jag1 is the major or only target mediating the inhibitory effect of ZEB1 knockdown or miR-200 overexpression on the Notch pathway. Overexpression of a Jag1 expression construct lacking the 3′UTR increased the Notch reporter activity, which was suppressed by knockdown of ZEB1 or overexpression of miR-141 and miR-200c. However, it could not fully rescue the suppressive effect (Figure 4E). Vice versa, inhibition of endogenous miR-141 and miR-200c in differentiated cancer cells by specific antagomirs increased Notch reporter activity, which again could be only partially repressed to the control level by siRNA-mediated knockdown of Jag1 (Figure 4F; Supplementary Figure S2B). In the same setting, the enhanced sphere-forming capacity after inhibition of miR-141 and miR-200c in HPAF2 cells was fully reduced by Jag1 knockdown (Figure 4G). Interestingly, reduction of Jag1 did not significantly affect the proliferative capacity of the differentiated lines HPAF2 and MCF7 (Figure 4H; Supplementary Figure S2C), in contrast to the undifferentiated lines Panc1 and MDA-MB231, which already expressed high endogenous levels of Jag1 (Figure 3C; Supplementary Figure S1E). Further work will address this difference. Altogether these data indicate that Jag1 alone is not the only target mediating the effects of ZEB1 and miR-200 on Notch signalling. To investigate if downstream Notch signalling is also directly affected by ZEB1 knockdown, we cotransfected an expression vector for the cleaved Notch1 ICD, a constitutive Notch signalling activator, which led to increased Notch reporter activation in Panc1 cells. NICD-induced activation was also decreased by stable knockdown of ZEB1 (Figure 4I), further indicating that ZEB1 and/or miR-200 influence Notch pathway activity not only by regulating the ligand Jag1 but also on downstream levels.
miR-200 targets the Notch coactivators Maml2 and Maml3
Besides Jag1, the only Notch pathway components detected in expression arrays to be affected by ZEB1 were the intracellular transcriptional coactivators Maml2 and Maml3 (Figure 1B), which could explain the reduced Notch reporter activity by ZEB1 knockdown also after cotransfection of NICD. Both coactivators are predicted miR-200 targets (Figure 5A). Knockdown of ZEB1 or overexpression of miR-200c and miR-141 inhibited expression of Maml2 and to a lesser extend Maml3 (Figure 5B–D). Moreover, 3′UTR reporter constructs of both factors were also affected by miR-200c/141 overexpression (Figure 5E). siRNA-mediated knockdown of Maml2 and Maml3 demonstrated their crucial role for Notch signalling, since it inhibited both Notch reporter activity (Figure 5F) and expression of the Notch target gene Hey1 (Figure 5G and H). Similar to a knockdown of Jag1 (see Figure 4F and Supplementary Figure S2B), a knockdown of Maml2 and Maml3 also led to a reduction of the sphere-forming capacity in the second-sphere generation of Panc1 cells (Figure 5I).
Figure 5.
The Notch coactivators Maml2 and Maml3 are targets of miR-200. (A) Scheme showing predicted target sites of miR-200 family members in the 3′UTR of Maml2 and Maml3. (B–D) Knockdown of ZEB1 or overexpression of miR-141 and miR-200c decrease the expression of Maml2 and Maml3 on mRNA (B, C) and protein (D) level. Overexpression of miR-200 in Panc1 was done by lentiviral transduction, shown in (D) are each three independent lines. (E) Overexpression of miR-141 and miR-200c decreases activity of the indicated 3′UTR reporter constructs. (F) Knockdown of Jag1, Maml2 or Maml3 decreases Notch reporter activity. (G, H) Knockdown of Maml2 or Maml3 decreases expression of the Notch target gene Hey1 in Panc1 (G) and MDA-MB231 (H). Maml3 is not expressed in MDA-MB231 (see Figure 1B). (I) Knockdown of Maml2 or Maml3 decreases the sphere-forming capacity of Panc1 in the second-sphere generation.
ZEB1, miR-200 and Notch signalling in human cancer
Finally, we analysed the expression of ZEB1, miR-200 family members and Notch pathway activation in human breast and pancreatic cancers. Distinct subtypes of these tumours are known to be associated with high Notch signalling activity. Typical human breast cancers are of the ductal invasive type, which express steroid hormone receptors and show a more differentiated phenotype. In contrast, the basal type of breast cancer is undifferentiated, does not express steroid hormone receptors (triple negative), and has a highly aggressive phenotype with poor clinical prognosis. The basal type of breast cancer is characterized by a Notch activation signature (Lee et al, 2008), and strikingly it was shown that this type is associated with a high expression of Jag1 (Reedijk et al, 2008). We observed that a high expression level of ZEB1 correlated with significantly lower miR-141 and miR-200c levels in the basal type, compared with the common ductal invasive type of breast cancer (Figure 6A). ZEB1 expression in basal type of breast cancers was heterogeneous within individual tumours. High ZEB1 expression levels, particularly in invasive tumour regions, correlated with expression of Jag1 and activation of Notch signalling, as indicated by expression of activated Notch1 and Notch2 (Figure 6B; Supplementary Figure S2D). Pancreatic cancer is also associated with increased Notch signalling activity (Miyamoto et al, 2003). In differentiated pancreatic adenocarcinomas only few tumour cells expressed ZEB1 (not shown). In contrast, a high proportion of the investigated undifferentiated pancreatic adenocarcinomas (classified with gradings G3 and G4) showed areas with strong expression of ZEB1 in tumour cells, again particularly in invasive regions. These tumour cells have undergone an EMT, as demonstrated by the expression of the mesenchymal marker vimentin. ZEB1 expression and EMT were strongly associated with expression of Jag1, mainly in invasive tumour regions (Figure 6C; Supplementary Figure S2D). Moreover, also undifferentiated, ZEB1 expressing pancreatic cancers expressed significantly lower miR-141 and miR-200c levels compared with differentiated cases (Figure 6A). These correlative data indicate that the experimentally validated link of ZEB1 expression, reduction of miR-200 family members and expression of Notch pathway components might be causal to increased Notch signalling in some types of human cancer.
Figure 6.
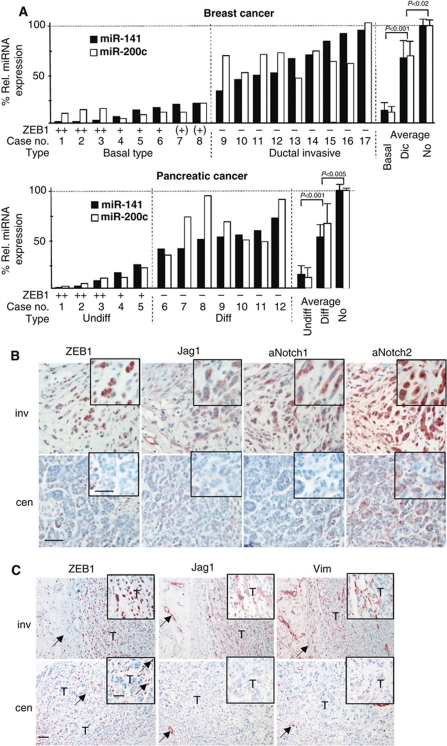
Correlation of ZEB1 and Notch activity in human cancers. (A) qPCR for miR-141 and miR-200c after microdissection of tumour areas from the indicated types of human primary tumours, showing a significant reduction in basal versus ductal invasive type (dic) of breast cancers, as well as in undifferentiated versus differentiated pancreatic adenocarcinomas. The statistical significance for the mean values of all included cases is shown. miRNA expression in normal (no) breast or pancreatic epithelial tissue was set to 100%. Shown are also the ZEB1 expression levels in tumour cells scored by immunohistochemistry. (B) Immunohistochemistry of a typical basal type of breast cancer showing correlated expression of ZEB1, Jag1, activated Notch1 and activated Notch2 in cancer cells of invasive (inv) tumour areas. In central tumour areas (cen), a correlated reduction was often detectable. (Inserts show magnifications, specific staining in red, nuclear counter staining in blue). Size bar is 20 and 12 μm for inserts. (C) Immunohistochemistry of an undifferentiated pancreatic adenocarcinoma showing correlated expression of ZEB1, Jag1 and the mesenchymal marker Vimentin in cancer cells of invasive (inv) tumour areas. In central tumour areas (cen) often a correlated reduction was detectable (squares show magnifications). Arrows show positive staining of fibroblasts for ZEB1 and Vimentin, and blood vessels for Jag1. T=tumour cells. Size bar is 20 and 12 μm for inserts.
Discussion
We have shown that the EMT activator ZEB1 increases Notch signalling in cancer cells via repression of miR-200 family members. Members of this microRNA family inhibit expression of Notch pathway components, as demonstrated for the Notch ligand Jag1 and the intracellular Notch coactivators Maml2 and Maml3. Notch signalling is known to be important for key cancer cell properties, including proliferation, survival and stemness (Hurlbut et al, 2007; Koch and Radtke, 2007; Rizzo et al, 2008), which could be confirmed in this study, in particular for Jag1. A correlated overexpression of ZEB1 and Jag1, associated with reduced miR-200 expression was detected in poorly differentiated pancreatic adenocarcinomas as well as in the highly aggressive basal type of breast cancer, particularly in invasive tumour cells.
Although the importance of Notch signalling for tumour formation and malignant progression is documented for many cancer types, mutations in Notch pathway components are detected very rarely in solid tumours. Our results describe an additional way to activate and stabilize Notch signalling by linking EMT and aberrant Notch signalling in cancer cells. This is of particular relevance since many of the properties already known to be regulated by the Notch pathway were recently shown to be also controlled by EMT activators. For instance, it was shown that factors like Snail, Twist and in particular ZEB1 not only control EMT-associated processes like cell motility, but also stemness and survival (Thiery et al, 2009). A link of EMT and Notch activation at molecular level can explain such overlaps in the control of crucial cellular processes. Moreover, a reverse link was already described recently, by showing that Notch signalling can induce EMT in cell culture (Leong et al, 2007; Sahlgren et al, 2008). For instance, an inhibition of Notch signalling resulted in the reversal of EMT and the associated drug resistance in pancreatic cancer cell lines by decreasing expression of ZEB1, Snail1 and NFκB (Wang et al, 2010). This is supported by our data, showing that a knockdown of Jag1 also affected the expression of ZEB1 in cancer cells (see Supplementary Figure S1B).
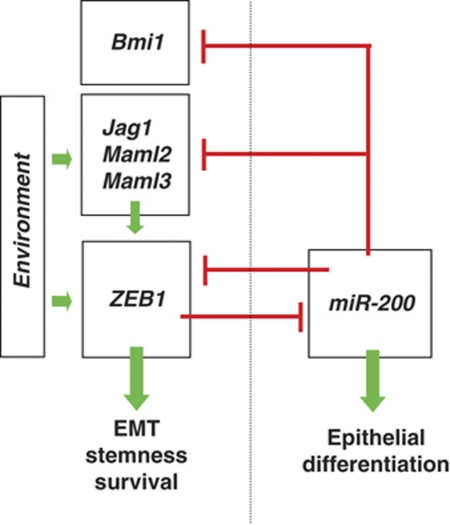
Our results indicate the existence of reciprocal activation of Notch signalling and EMT in the sense of a positive feedback loop. We therefore would like to propose the following model (Figure 7): aberrant expression of ZEB1, for example induced by environmental signals and hypoxia at the tumour invasion front, was shown to induce stemness and EMT in cancers cells through suppression of stemness-inhibiting microRNAs, including miR-200 family members (Shimono et al, 2009; Wellner et al, 2009). Targets of miR-200 are not only ZEB1 itself and stem cell factors, such as Bmi1 (Brabletz and Brabletz, 2010), but also Notch pathway components like Jag1, Maml2 and Maml3. Accordingly, indirect stabilization of Notch pathway components by aberrant ZEB1 re-enforces a Notch signalling loop, with all its known effects. In addition, it was already shown (Wang et al, 2009) and also confirmed in this study (see Supplementary Figure S1B), that Notch signalling stimulates ZEB1 expression, which might further stabilize motility, survival and a stemness phenotype in cancer cells, particularly at the invasive front. We have validated our experimental findings by analysing the expression of ZEB1 and Jag1 and the activation of Notch signalling in subtypes of human pancreatic and breast cancers, which were selected, because of their strong association with aberrant Notch signalling. In particular, the highly aggressive basal type of breast cancer is characterized by high expression of Jag1 (Reedijk et al, 2008). Here, we found an associated expression of Jag1 and ZEB1, as well as Notch activation in tumour cells of this type of breast cancer, supporting the proposed link of high ZEB1, low miR-200 and Notch activation at molecular level. However, in the more differentiated ductal invasive type of breast cancer, we rarely found ZEB1 expressing tumour cells, although Jag1 was expressed in some tumour areas (not shown). This indicates that additional ways of aberrant Notch activation might exist in different cancer subtypes. A second example is pancreatic cancer, which is also characterized by abundant activation of Notch signalling (Miyamoto et al, 2003). We already described a high frequency of aberrant ZEB1 expression in poorly differentiated pancreatic adenocarcinomas, in particular in invasive regions (Wellner et al, 2009). As described for basal type of breast cancers (Burk et al, 2008; Gregory et al, 2008), we show here that also undifferentiated pancreatic cancers express significantly lower levels of miR-200 family members compared with differentiated cases. We further detected a correlated expression of ZEB1, the mesenchymal marker vimentin and Jag1 expression in invasive regions of such cancers, supporting the existence of the proposed molecular link also in pancreatic adenocarcinomas. Future work will show if this link between ZEB1, miR-200 and Notch activation is also active in other cancer types and is controlling physiological processes in embryonic development and adult tissue homeostasis as well.
Figure 7.
Proposed model for a molecular link between ZEB1, miR-200 and Notch signalling (explanation see Discussion section).
In summary, we here described that ZEB1 can indirectly activate Notch signalling by inhibiting miR-200 expression, leading to a stabilization of Notch pathway components and Notch signalling. Thus, activation of Notch signalling is an additional way how ZEB1 exerts its established tumour progressing function. This link could explain how aberrant EMT activation is molecularly connected to crucial properties driving cancer initiation and progression, such as maintenance of cancer stem cells, as well as proliferation and drug resistance of cancer cells. It could further explain how Notch signalling is amplified in some types of poorly differentiated cancers, where mutations in Notch pathway genes are rarely detected.
Materials and methods
DNA constructs
For construction of miR-200c and miR-141 expression constructs, mature sequences of miR-141 and miR-200c were cut out of pUC57 miR-141 or pUC57 miR-200c (GenScript, Piscataway, NJ) and cloned in pRetroSuper (Brummelkamp et al, 2002). For construction of the Jagged1 3′UTR reporter plasmid, nucleotides +4164 to +5966 of human Jagged 1 cDNA, for construction of the Maml2 and Maml3 3′UTR reporter plasmids, nucleotides +4759 to +5419 of human Maml2 cDNA and nucleotides +5564 to +7064 of human Maml2 cDNA were amplified and cloned into the multiple cloning site downstream of the luciferase gene in the pMIR-REPORT vector (Ambion, Austin, TX). For construction of the Jag1 expression vector lacking the 3′UTR nucleotides +517 to + 4174 of human Jag1 cDNA were amplified and cloned into pCI-neo (Promega, Madison, WI). The Notch-CSL-1 reporters, 4xwt-CSL-Luc, which contains four tandem repeats of the consensus CSL DNA-binding sequence, GTGGGAA and the mutant (4xmt-CSL-luc) were gifts from Diane Hayward (Johns Hopkins University School of Medicine, Baltimore, MD). The plasmid pcDNA-FLAG-Notch-IC, expressing the Notch1 ICD, was a gift from Tilman Borggrefe (MPI for Immunobiology, Freiburg, Germany). For mutagenesis of the indicated miRNA seed sequence binding sites for mir-141, mir-200a and mir-200c in the Jagged1 3′UTR, PCRs were performed using the Pfu Ultra Hotstart 2 × Master Mix (Stratagene, Santa Clara) followed by DpnI (Fermentas, St Leon-Rot, Germany) digestion for 1 h at 37°C. Mutagenesis was verified by sequencing.
Cell culture and various assays
All cell lines were purchased from ATCC. Stable knockdown clones for ZEB1 and control clones were constructed as previously described (Spaderna et al, 2008). For generation of stable overexpression Panc1 clones for miRNA-141 or miRNA-200c, cells were transfected with sequence verified pRetroSuper-miRNA constructs. Cell lines were selected and cultivated under standard conditions in DMEM + 10% fetal bovine serum+2 μg/ml puromycin. Transient transfections, reporter assays, immunoblots and transient siRNA-mediated knockdown were done as previously described (Spaderna et al, 2006, 2008). All experiments were done at least three times. For reporter assays, the firefly luciferase values were normalized against the values of a cotransfected pCMV-Renilla-Luciferase construct, in order to correct differences in transfection efficiencies. RNA was isolated using RNeasy Plus Mini Kit (Qiagen), mirVana™ miRNA Isolation Kit (Ambion) for microRNAs and RecoverAll™ RNA Isolation Kit for formalin-fixed, paraffin-embedded tissues (Ambion). mRNA expression values were measured in triplicates on a Roche LightCycler 480 and normalized to β-actin expression as control. Specific quantitative real-time PCR experiments for miRNAs were carried out using TaqMan MicroRNA Assays (Applied Biosystems). To block Notch signalling, cells were incubated for 24 h in the presence of GSI type I (Z-Leu-Leu-Nle-CHO; Calbiochem #565750, San Diego, CA) in concentrations of 5 and 2.5 μM. Lentiviral-based miRNA expression vectors were used for generating control GFP (pCDH-CMV-MCS-EF1-copGFP, System Biosciences, Mountain View, CA), miR-141 + GFP (pMIRH141-PA-1, System Biosciences) and miR-200c + GFP (pMIRH200c-PA-1, System Biosciences) overexpressing cells. Lentiviral particles were produced by cotransfection of HEK293T cells with the appropriate transfer and lentiviral helper plasmids (pCMVDR8.74 packaging vector and pMD2.VSVG envelope vector) using Lipofectamine with Plus reagent (Invitrogen). The medium was exchanged 3 h after transfection and lentiviral supernatant was collected 36 and 72 h later. The lentiviral supernatants were concentrated by ultracentrifugation at 20°C for 2.5 h at 19 500 r.p.m. Infection of Panc1 cells was performed in 12-well plates in the presence of 6 μg/ml Polybrene (Hexadimethrinbromid, Sigma-Aldrich). Infection with different amounts of viruses was tested on Panc1 cells and the infection efficiency determined by GFP expression and quantitative real-time PCR for microRNAs.
Cell proliferation assay
Cells were seeded in 96-well plates at 3000 cells/well. After 24 h, cells were transfected with siRNAs or treated with GSI. At indicated time points, MTT (methylthiazolyldiphenyl-tetrazolium bromide; Sigma #M5655) was added (5 mg/ml) to the medium. After 4 h incubation, the medium was removed and the crystals were diluted in 200 μl acidified isopropanol (0.04 N HCl). Absorption was measured at 570 nm with 650 nm as a reference wavelength using a Tecan Infinite M200 Reader to determine MTT activity, which is indicated in relation to activity measured at day 1 after seeding (set to 1).
Cancer stem cell spheroid assay
To induce sphere formation, cells were dissociated to single cells by 0.05% trypsin-EDTA solution (Invitrogen) and plated at 20 000 cells/ml in serum-free medium (SFM) into Poly(2-hydroxyethyl metacrylate) (Polyhema, Sigma)-coated six-well plates to prevent the cells from attaching to the surface. The SFM consists of DMEM-F12 (Invitrogen) supplemented with 20 ng/ml epidermal growth factor (R&D Systems), 0.4% bovine serum albumin (Sigma). For pancreatic cell lines, it additionally contains 10 ng/ml fibroblast growth factor (BD Bioscience) and 10 μg/ml Insulin, 10 μg/ml Transferrin, 10 ng/ml Sodium-Selenite Mix (Sigma), whereas the SFM for breast cancer cells is further supplemented with B27 Supplement 1:50 (Invitrogen) and 4 μg/ml Insulin (Invitrogen). For in vitro propagation, the spheres were dissociated to single cells after 5 days and again seeded as described to form the next generation of spheres. For quantification of the sphere-formation capacity, 1000 cells/well of the dissociated single cells were seeded in SFM containing 1% methylcellulose (Sigma) into polyhema-coated 96-well plates. After 7 days, Panc1 cells colony numbers >75 μm were counted, for MDA-MB231 cell line all colonies >5 cells were counted. For the treatment of HPAFII cells with antagomirs against miR-141 and miR-200c and subsequent siRNA-mediated knockdown of Jag1, tumour cells were first transfected with siRNAs against Jag1 using Oligofectamine (Invitrogen). At 4 h after transfection, medium was changed and 500 nM antagomirs added to the cell culture medium. After 24 h, cells were trypsinized and seeded for the spheroid assay in the presence of 500 nM antagomirs.
Apoptosis assay
Exponentially growing cells were transfected with the indicated siRNAs or treated with GSI. At 8 h after transfection or 3 h after GSI treatment, cells were irradiated with 5 Gy using a Gammacell 40 137Cs laboratory irradiator. After 3 days, cells were stained for Annexin V and with propidium iodide using an Annexin V-FITC Kit from Milteniy Biotec. Apoptosis was measured by flow cytometry on a Cytomics FC 500 instrument from Beckman Coulter.
miRNA target gene search
For identification of potential mircoRNA target genes, the miRecords website, which integrates different microRNA target prediction tools, was used (http://mirecords.biolead.org).
MicroRNA overexpression
In all, 3 × 104 cells/well were seeded in 12-well plates. After 24 h, cells were transfected with 30 pmol miRNA oligonucleotides for hsa-miR-141 or hsa-miR-200c (Ambion). The Ambion Pre-miR™ miRNA Precursor-Negative Control #1 and #2 was used as control. Oligofectamine™ Reagent (Invitrogen) was used as transfection reagent. After 3 to 6 days of incubation, cells were used for several assays.
Specific inhibition of miRNAs using antagomirs
Antagomirs (Dharmacon) were designed as described (Krutzfeldt et al, 2007). In all, 500 nM of antagomirs were added to the normal cell culture medium right after seeding in 12-well plates. In all, 4–5 days after seeding, the cells were harvested for specific assays.
Tissue specimens and immunohistochemistry
Immunohistochemistry on formalin-fixed, paraffin-embedded samples of breast and pancreatic carcinomas from patients who underwent surgery or of nude mice xenografts was done as previously described (Brabletz et al, 2004; Wellner et al, 2009). For quantification of miRNA expression by qPCR, tumour areas were separated by microdissection under microscope, total RNAs including small RNAs were isolated by RecoverAll RNA Isolation Kit for formalin-fixed, paraffin-embedded tissues (Ambion).
Statistics
For comparison of rational variables, the ANOVA algorithm of SPSS 17.0 (SPSS Inc., Chicago) with the significance level set to P=0.05 was used. Relative MTT absorption levels were plotted with error bars as indicated (confidence interval, standard error). Association of dichotomous variables was analyzed using two-sided Fisher's exact test of SPSS Version 17.0 (SPSS Inc., Chicago), with a significance level of P<0.05.
Supplementary Material
Acknowledgments
For expert technical assistance we thank Bettina Schuler, Jessica Pfannstiel, Kerstin Meyer, Anja Schmitt and Stephanie Mewes. For helpful discussions, we are grateful to Tilman Borggrefe (MPI for Immunobiology, Freiburg, Germany). For generous gift of reagents, we thank Tilman Borggrefe, Maria Dominguez (Instituto de Neurosciencas, Alicante, Spain) and Diane Hayward (Johns Hopkins University School of Medicine, Baltimore, MD). This work was supported by grants to TB from the EU (MCSC contract no. 037297), and the DFG (no. BR 1399/6-1 and the SFB 850, B2), and to GN from the Clotten Foundation.
Author contributions: SB designed the study, performed the experiments and analysed the data; KB, SM, UB, GN and EF performed the experiments and analysed the data; UW performed statistical analyses; AD and GF investigated and analysed human cancers; JS designed the parts of the study, performed the experiments and analysed the data; TB designed the study, analysed the data and wrote the manuscript.
Footnotes
The authors declare that they have no conflict of interest.
References
- Brabletz S, Brabletz T (2010) The ZEB/miR-200 feedback loop[mdash]a motor of cellular plasticity in development and cancer? EMBO Rep 11: 670–677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brabletz T, Spaderna S, Kolb J, Hlubek F, Faller G, Bruns CJ, Jung A, Nentwich J, Duluc I, Domon-Dell C, Kirchner T, Freund JN (2004) Down-regulation of the homeodomain factor Cdx2 in colorectal cancer by collagen type I: an active role for the tumor environment in malignant tumor progression. Cancer Res 64: 6973–6977 [DOI] [PubMed] [Google Scholar]
- Brummelkamp TR, Bernards R, Agami R (2002) Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell 2: 243–247 [DOI] [PubMed] [Google Scholar]
- Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S, Brabletz T (2008) A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 9: 582–589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gou S, Liu T, Wang C, Yin T, Li K, Yang M, Zhou J (2007) Establishment of clonal colony-forming assay for propagation of pancreatic cancer cells with stem cell properties. Pancreas 34: 429–435 [DOI] [PubMed] [Google Scholar]
- Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y, Goodall GJ (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10: 593–601 [DOI] [PubMed] [Google Scholar]
- Hurlbut GD, Kankel MW, Lake RJ, Artavanis-Tsakonas S (2007) Crossing paths with Notch in the hyper-network. Curr Opin Cell Biol 19: 166–175 [DOI] [PubMed] [Google Scholar]
- Jones S, Zhang X, Parsons DW, Lin JC-H, Leary RJ, Angenendt P, Mankoo P, Carter H, Kamiyama H, Jimeno A, Hong S-M, Fu B, Lin M-T, Calhoun ES, Kamiyama M, Walter K, Nikolskaya T, Nikolsky Y, Hartigan J, Smith DR et al. (2008) Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321: 1801–1806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koch U, Radtke F (2007) Notch and cancer: a double-edged sword. Cell Mol Life Sci 64: 2746–2762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korpal M, Lee ES, Hu G, Kang Y (2008) The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 283: 14910–14914 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krutzfeldt J, Kuwajima S, Braich R, Rajeev KG, Pena J, Tuschl T, Manoharan M, Stoffel M (2007) Specificity, duplex degradation and subcellular localization of antagomirs. Nucleic Acids Res 35: 2885–2892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee C, Simin K, Liu Q, Plescia J, Guha M, Khan A, Hsieh C-C, Altieri D (2008) A functional Notch-survivin gene signature in basal breast cancer. Breast Cancer Res 10: R97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leong KG, Niessen K, Kulic I, Raouf A, Eaves C, Pollet I, Karsan A (2007) Jagged1-mediated Notch activation induces epithelial-to-mesenchymal transition through Slug-induced repression of E-cadherin. J Exp Med 204: 2935–2948 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mani SA, Guo W, Liao M-J, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M, Campbell LL, Polyak K, Brisken C, Yang J, Weinberg RA (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133: 704–715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyamoto Y, Maitra A, Ghosh B, Zechner U, Argani P, Iacobuzio-Donahue CA, Sriuranpong V, Iso T, Meszoely IM, Wolfe MS, Hruban RH, Ball DW, Schmid RM, Leach SD (2003) Notch mediates TGF alpha-induced changes in epithelial differentiation during pancreatic tumorigenesis. Cancer Cell 3: 565–576 [DOI] [PubMed] [Google Scholar]
- Morel AP, Lievre M, Thomas C, Hinkal G, Ansieau S, Puisieux A (2008) Generation of breast cancer stem cells through epithelial-mesenchymal transition. PLoS ONE 3: e2888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park SM, Gaur AB, Lengyel E, Peter ME (2008) The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22: 894–907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peter ME (2009) Let-7 and miR-200 microRNAs: guardians against pluripotency and cancer progression. Cell Cycle 8: 843–852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plentz R, Park JS, Rhim AD, Abravanel D, Hezel AF, Sharma SV, Gurumurthy S, Deshpande V, Kenific C, Settleman J, Majumder PK, Stanger BZ, Bardeesy N (2009) Inhibition of gamma-secretase activity inhibits tumor progression in a mouse model of pancreatic ductal adenocarcinoma. Gastroenterology 136: 1741–1749, e1746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reedijk M, Pinnaduwage D, Dickson BC, Mulligan AM, Zhang H, Bull SB, O'Malley FP, Egan SE, Andrulis IL (2008) JAG1 expression is associated with a basal phenotype and recurrence in lymph node-negative breast cancer. Breast Cancer Res Treat 111: 439–448 [DOI] [PubMed] [Google Scholar]
- Rizzo P, Osipo C, Foreman K, Golde T, Osborne B, Miele L (2008) Rational targeting of Notch signaling in cancer. Oncogene 27: 5124–5131 [DOI] [PubMed] [Google Scholar]
- Roy M, Pear WS, Aster JC (2007) The multifaceted role of Notch in cancer. Curr Opin Genet Dev 17: 52–59 [DOI] [PubMed] [Google Scholar]
- Sahlgren C, Gustafsson MV, Jin S, Poellinger L, Lendahl U (2008) Notch signaling mediates hypoxia-induced tumor cell migration and invasion. Proc Natl Acad Sci 105: 6392–6397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimono Y, Zabala M, Cho RW, Lobo N, Dalerba P, Qian D, Diehn M, Liu H, Panula SP, Chiao E, Dirbas FM, Somlo G, Pera RAR, Lao K, Clarke MF (2009) Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 138: 592–603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spaderna S, Schmalhofer O, Hlubek F, Berx G, Eger A, Merkel S, Jung A, Kirchner T, Brabletz T (2006) A transient, EMT-linked loss of basement membranes indicates metastasis and poor survival in colorectal cancer. Gastroenterology 131: 830–840 [DOI] [PubMed] [Google Scholar]
- Spaderna S, Schmalhofer O, Wahlbuhl M, Dimmler A, Bauer K, Sultan A, Hlubek F, Jung A, Strand D, Eger A, Kirchner T, Behrens J, Brabletz T (2008) The transcriptional repressor ZEB1 promotes metastasis and loss of cell polarity in cancer. Cancer Res 68: 537–544 [DOI] [PubMed] [Google Scholar]
- Thiery JP, Acloque H, Huang RY, Nieto MA (2009) Epithelial-mesenchymal transitions in development and disease. Cell 139: 871–890 [DOI] [PubMed] [Google Scholar]
- Vandewalle C, Van Roy F, Berx G (2009) The role of the ZEB family of transcription factors in development and disease. Cell Mol Life Sci 66: 773–787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Li Y, Kong D, Ahmad A, Banerjee S, Sarkar FH (2010) Cross-talk between miRNA and Notch signaling pathways in tumor development and progression. Cancer Lett 292: 141–148 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Li Y, Kong D, Banerjee S, Ahmad A, Azmi AS, Ali S, Abbruzzese JL, Gallick GE, Sarkar FH (2009) Acquisition of epithelial-mesenchymal transition phenotype of gemcitabine-resistant pancreatic cancer cells is linked with activation of the Notch signaling pathway. Cancer Res 69: 2400–2407 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A, Waldvogel B, Vannier C, Darling D, zur Hausen A, Brunton VG, Morton J, Sansom O, Schuler J, Stemmler MP, Herzberger C, Hopt U, Keck T, Brabletz S, Brabletz T (2009) The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 11: 1487–1495 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.