Abstract
Protein production within the secretory pathway is accomplished by complex but organized processes. Here, we demonstrate that the growth factor midkine interacts with LDL receptor-related protein 1 (LRP1) at high affinity (Kd value, 2.7 nm) not only at the cell surface but also within the secretory pathway during biosynthesis. The latter premature ligand-receptor interaction resulted in aggregate formation and consequently suppressed midkine secretion and LRP1 maturation. We utilized an endoplasmic reticulum (ER) retrieval signal and an LRP1 fragment, which strongly bound to midkine and the LRP1-specialized chaperone receptor-associated protein (RAP), to construct an ER trapper. The ER trapper efficiently trapped midkine and RAP and mimicked the premature ligand-receptor interaction, i.e. suppressed maturation of the ligand and receptor. The ER trapper also diminished the inhibitory function of LRP1 on platelet-derived growth factor-mediated cell migration. Complementary to these results, an increased expression of RAP was closely associated with midkine expression in human colorectal carcinomas (33 of 39 cases examined). Our results suggest that the premature ligand-receptor interaction plays a role in protein production within the secretory pathway.
Keywords: Cancer Therapy, Chaperone Chaperonin, Endoplasmic Reticulum (ER), Growth Factors, Protein Folding, Protein Synthesis, Biosynthesis
Introduction
Protein production within the secretory pathway is tightly regulated by complex but organized processes, which include folding in the endoplasmic reticulum (ER),2 glycosylation, and vesicular transport to the cell surface through the ER and Golgi apparatus. Although these processes play important roles in quality control and maturation of proteins, it is not fully understood how production of proteins in the secretory pathway is regulated. This regulation mechanism is also important to understand some diseases, as disorganized production of growth factors can be causative events in some pathological conditions, such as cancer (1). A fundamental question yet to be carefully addressed is whether a ligand and its receptor come into contact during their biosyntheses. To address this, we focused on two molecules in the present study, the growth factor midkine (MK) and its receptor low density lipoprotein (LDL) receptor-related protein 1 (LRP1).
MK is a heparin-binding growth factor, which was originally discovered as the product of a retinoic acid-responsive gene (2, 3). MK and pleiotrophin/heparin-binding growth-associated molecule form a family distinct from other heparin-binding growth factor families (4–6). MK plays important biological roles mainly in four areas, i.e. cancer, inflammation, neuro- and myocardioprotection, and hypertension. To date, the biological functions in which MK has been implicated are all attributable to the secreted form of MK, and thus exogenous MK reverses phenotypes observed in MK-deficient mice. For example, MK-deficient mice are more resistant to vascular restenosis and hypertension associated with chronic kidney disease, whereas exogenous MK promotes these pathogeneses (7, 8). It is noteworthy that not only MK expression in carcinoma tissues but also the blood MK level is increased in cancer patients (9, 10). Both the MK expression level and MK blood level are closely related to clinical outcome (11–13). Furthermore, MK expression knockdown leads to tumor growth suppression (14, 15). These data suggest that the MK secreted by cancer cells plays a critical role in cancer progression.
LRP1 is a large endocytosis receptor belonging to the LDL receptor family (16). Because LRP1 recognizes many different ligands, the physiological roles of LRP1 govern a diverse array of biological functions, including metabolism of lipoproteins and proteinases, cellular entry of viruses and toxins, activation of lysosomal enzymes, and neurotransmission. Disruption of the LRP1 gene is embryonic-lethal, supporting the fundamental role of LRP1 in development (17). The receptor-associated protein (RAP) is a specialized ER chaperone for members of the LDL receptor family (18–20). Maturation of LRP1 is severely impaired in RAP-deficient mice, supporting an important role of RAP in the biosynthesis of LRP1 (20).
LRP1 is a receptor for MK and is required for MK-mediated cell survival (21, 22). Because MK and LRP1 are produced simultaneously in most cancer cell lines, this ligand and receptor are produced in the same secretory pathway. Therefore, this is a good model to investigate the regulatory mechanism of protein production in an autocrine secretory pathway. Here, we developed a new method, ER trapping, which simultaneously suppressed MK secretion and LRP1 maturation. Utilizing this and other techniques, we found that MK interacted with LRP1 not only on the cell surface but also in the secretory pathway during their biosyntheses. We named the latter interaction the premature ligand-receptor interaction. We found that the premature ligand-receptor interaction plays a negative regulatory role in the production of MK and LRP1 within the secretory pathway. Avoiding this premature interaction may allow cancer cells to overproduce a growth factor and may be a strategy of those cells for survival and expansion.
EXPERIMENTAL PROCEDURES
Cells, DNA Constructs, and Antibodies
CHO K1 cells were cultured in DMEM with 10% fetal bovine serum. TGW cells (a human neuroblastoma cell line) were cultured in RPMI 1640 medium with 10% fetal bovine serum. CHO-mini-LRP (mLRP)-I, -mLRP-II, -mLRP-III, -mLRP-IV, and -mLRP-IVm cells and a series of mLRP expression vectors were generated as described previously (23–25). The expression vector for mouse RAP tagged with the FLAG epitope at the C terminus was constructed in pIRES-EGFP (Clontech). The expression vector for human MK was constructed in pcDNA3.1 (Invitrogen). To generate an ER expression vector (ER-TRAP) for a peptide that contained SLRP2N-EGF (25) and the ER retention signal HNEL, the cDNA encoding this peptide was placed underneath the cytomegalovirus promoter of pcDNA3 (Invitrogen). Anti-HA and -FLAG antibodies were purchased from Roche Applied Science and Sigma, respectively. Anti-calnexin and anti-phosphotyrosine (Tyr(P)-100) antibodies were from Stressgen and Cell Signaling, respectively.
Chemical Cross-linking
Radioiodination of MK was performed as described previously (22). The specific activity obtained was ∼5 × 107 cpm/μg. For chemical cross-linking, cells were washed with ice-cold phosphate-buffered saline containing calcium and magnesium (PBS+) as well as 0.1% bovine serum albumin (PBS-BSA), and supplied with 5 ml of ice-cold DMEM with 10% fetal bovine serum. Cells were then incubated with 125I-labeled MK (2 × 107 cpm) at 4 °C for 2 h by gentle rotation. After washing with PBS-BSA three times and PBS+ once, 5 ml of PBS+ was added. The cross-linking was begun by adding the water-soluble (BS3; Pierce) or -insoluble (DSS; Pierce) cross-linker at a final concentration of 0.27 mm and incubating at 4 °C for 15 min. The reaction was stopped with detaching buffer (10 mm Tris-HCl (pH 7.5), 10% glycerol, 1 mm calcium chloride, 1 mm magnesium chloride, and 1 mm PMSF). Cells were scraped in 500 μl of detaching buffer, collected by centrifuge, and lysed with 200 μl of lysis buffer (20 mm Tris-HCl (pH 7.5), 0.6% CHAPS, 150 mm sodium chloride, 1 mm calcium chloride, 1 mm magnesium chloride, and 1 mm PMSF). For in vitro cross-linking, 10 μl of protein solution dialyzed against appropriate buffers was mixed with 2 μl of 125I-labeled MK (∼10 ng) and was incubated at 20 °C for 1 h. Cross-linking was started by adding 0.2 μl of 20 mm DSS or BS3 and incubating at 4 °C for 10 min. The reaction was stopped by adding 1 μl of 1 m Tris-HCl (pH 7.4) and 4.5 μl of Laemmli 4 × sample buffer.
Sucrose Gradient Analysis
Cells grown in a 10-cm dish were lysed with 0.5 ml of 1% Triton in PBS+ and centrifuged at 15,000 × g at 4 °C for 10 min. The supernatant (0.3 ml) was layered on a stepwise sucrose gradient (60% 0.5 ml, 25% 1 ml, 20% 1 ml, 10% 1 ml, and 5% 1 ml in 0.1% Triton in PBS+) in a centrifuge tube (326819 tube; Beckman) and centrifuged at 130,000 × g at 17 °C for 19 h (SW50.1 rotor; Beckman). Fractionation was performed by collecting 7 drops/fraction from the bottom of the tube.
Sequential Immunoprecipitation
After the first immunoprecipitation, the immunoprecipitates were suspended in 100 μl of 1% SDS in the lysis buffer, boiled for 3 min, and diluted with 1 ml of the lysis buffer. This solution was then subjected to the second immunoprecipitation.
Generation and Purification of MK-TRAP-Fc
The expression vector for MK-TRAP-Fc was generated in pFUSE-Fc (Invivogen) and transfected into HEK293T cells. After 5 days, the conditioned media were collected and centrifuged to remove cellular debris. A 24-ml aliquot of the supernatants was loaded onto protein G-Sepharose (GE Healthcare) and washed with 30 ml of 20 mm phosphate buffer (pH 7.0). The bound fraction was eluted with 3.6 ml of 100 mm glycine HCl (pH 3.0) and neutralized immediately with 0.4 ml of 1.5 m Tris-HCl (pH 8.8). The purified protein was desalted with PD-10 (GE Healthcare) and concentrated using an Amicon Ultra-4 device (MWCO 3000; Millipore).
In Vitro Binding Assay of MK and MK-TRAP
A 100-μl aliquot of MK-TRAP-Fc solution was immobilized on a 96-well assay plate by incubating overnight at 4 °C. The wells were washed three times with PBS, and varying concentrations of AP-MK were applied. After 90 min at room temperature, unbound AP-MK was washed out by washing five times with HBHA buffer (20 mm HEPES, 0.1% BSA in Hanks' balanced salt solution). Bound AP-MK was estimated by enzymatic assay using p-nitrophenylphosphate as a substrate.
Sample Collection of Human Colorectal Carcinoma Tissues
The procedures and sources of sample collection under informed consent were as described previously (26).
Quantitative Real-time RT-PCR
Total RNA was isolated from colorectal tissues with ISOGEN (Nippon Gene). cDNA was obtained using a SuperScript III cDNA synthesis kit (Invitrogen). Quantitative real-time RT-PCR was performed with a TaqMan probe and ABI PRISM 7700 (Applied Biosystems) according to the manufacturer's instructions.
Cell Migration Assay
Chemotaxicells (8-μm pore; Kurabo, Osaka, Japan) were coated, unless otherwise indicated, on the underside surface with 1 μg/ml collagen I in PBS at 37 °C for 1 h. Harvested cells were washed with 10% FBS/DMEM once and 0.4% FBS/DMEM once, then suspended at 4 × 105 cells/ml in 0.4% FBS/DMEM. Collagen I-coated chambers were inserted into wells of a 24-well plate, where each well contained 600 μl of 0.2% BSA and 0.4% FBS/DMEM with or without 50 ng/ml PDGF-BB, and preincubated at room temperature for 10 min. Cells (100 μl) were added to each insert and incubated for 4 h in a CO2 incubator. Migrating cells were counted at 200 × magnification for five fields/chamber. Experiments were performed in triplicate or quadruplicate for each condition, with the numbers of migrating cells being presented as the average ± S.E.
Cell Survival Assay
SW620 cells were transfected with siRNA (siGENOME; Dharmacon) using Lipofectamine 2000 (Invitrogen). After 24 h, cells were seeded onto 96-well plates, and then the medium was switched to 0.5% FBS/F12-K. Viable cells were estimated with a Cell Counting Kit-8 (DOJINDO).
RESULTS
MK Binds to the Second and Fourth Ligand-binding Domains of LRP1 on the Cell Surface
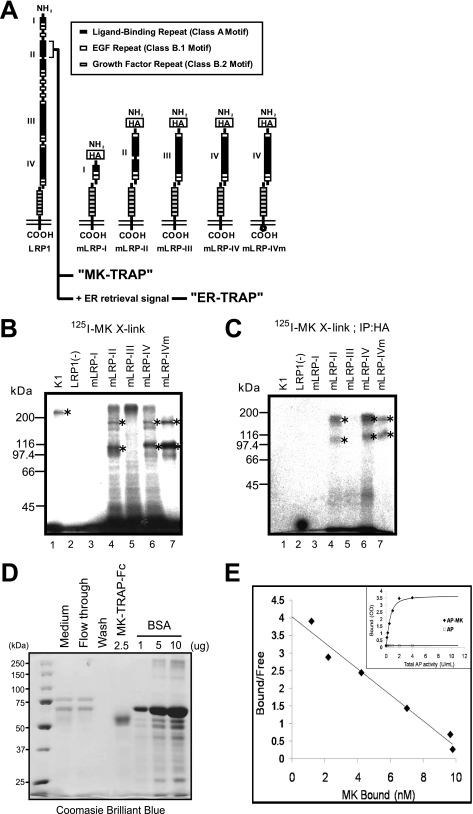
LRP1 consists of two heterodimeric subunits that are produced through furin-mediated processing in the trans-Golgi (Fig. 1A). The larger chain (α chain) is extracellular and contains four ligand-binding domains (clusters of ligand-binding repeats). The shorter chain (β chain) contains the transmembrane domain and the cytoplasmic tail. LRP1 was previously identified as a protein bound to the MK-affinity column (21). To clarify the physical association between these two molecules on the cell surface, here we used several membrane-bound mLRP constructs (Fig. 1A) (23, 24). The difference between mLRP-IV and mLRP-IVm is that the latter contains a mutation in the intracellular domain. Each mLRP protein carries an HA tag near its N terminus. Each construct was transfected into LRP1-deficient CHO cells. Cells were then incubated with 125I-labeled MK and treated with a water-insoluble chemical cross-linker, DSS.
FIGURE 1.
Exogenously added 125I-MK is chemically cross-linked to LRP1. A, schematic presentation of LRP1 and membrane-bound mini-LRP-I, -II, -III, -IV, and -IVmutant. mLRP-IVm has an intracellular domain mutation within its endocytosis signal, which results in cell surface accumulation of this mutant receptor. Biotinylated cell surface mLRP-IV was detected by anti-HA antibody after a pulldown assay with streptavidin-agarose. B and C, CHO cells cross-linked with 125I-MK (B) and immunoprecipitated with anti-HA antibody (C). K1 is the wild-type CHO cell line expressing endogenous LRP1. LRP1(−) is a mutant cell line deficient in LRP1. LRP-I, -II, -III, -IV, and -IVm are stable transfectants expressing these genes in the background of LRP1(−) cells. Asterisks indicate bands corresponding to various forms of LRP1 (see “Results”). D, purified MK-TRAP-Fc subjected to SDS-PAGE and stained with Coomassie Brilliant Blue. E, binding kinetics of MK and MK-TRAP-Fc. Scatchard plot analysis yielded a Kd value of 2.7 nm.
As shown in Fig. 1B, a discrete cross-linked band appeared in wild-type CHO cells (K1) but not in LRP1-deficient CHO cells (LRP1−/−). Notably, mLRP-II, -IV, and -IVm-expressing cells showed two cross-linked bands. The upper one represents the unprocessed protein that contains both α and β chains, whereas the lower band is the processed α chain. Because a small fraction of the unprocessed protein is localized on the cell surface,3 the cross-linked pattern shown in Fig. 1B suggests that MK binds to the cell surface α chain containing either the second or fourth ligand-binding domains. Immunoprecipitation with anti-HA antibody confirmed that the cross-linked bands indeed contained the HA-tagged α chain of each mLRP (Fig. 1C). A water-soluble cross-linker, BS3, showed the same results (data not shown). Therefore, these data collectively suggest that MK specifically binds to cell surface LRP1.
These data were also consistent with our previous finding in experiments using soluble LRP1 fragments: the N-terminal halves of the second and fourth ligand-binding domains in the LRP1 moiety show the highest affinity to MK (27). We further performed Scatchard plot analysis to reveal the affinity between MK and the N-terminal half of the second ligand-binding domain of LRP1 (formerly designated MK-TRAP; Fig. 1A). Indeed, MK and MK-TRAP showed a high affinity with a Kd value of 2.7 nm (Fig. 1, D and E).
LRP1 Is Required for Intracellular Retention of MK
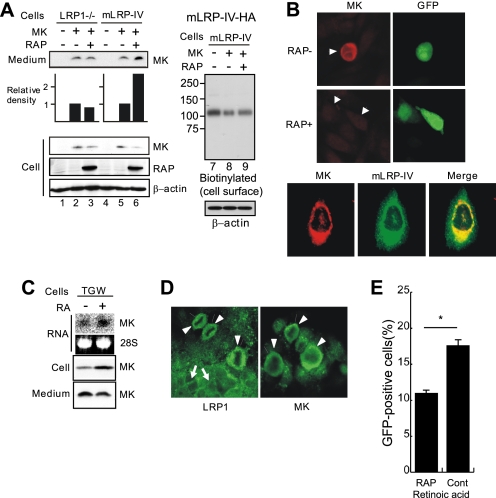
We next investigated the role of LRP1 in MK production in the secretory pathway by comparing LRP1-deficient and mLRP-IV-expressing cells. In mLRP-IV-expressing cells, co-expression of MK and the LRP1 chaperone RAP enhanced MK secretion into the medium compared with MK expression alone (Fig. 2A, lanes 5 and 6, Medium, MK). In contrast, the effect of RAP on MK secretion was not seen in LRP1-deficient cells (LRP1−/−) (Fig. 2A, lanes 2 and 3, Medium, MK). MK retention in the cell appeared to be reduced by co-expression with RAP (Fig. 2A, lanes 5 and 6, Cells, MK). Furthermore, translocation of mLRP-IV to the cell surface of MK-expressing cells was significantly reduced compared with that in vector-transfected cells (Fig. 2A, lanes 7 and 8). This phenomenon was also restored by co-expression of RAP (Fig. 2A, lanes 8 and 9).
FIGURE 2.
RAP enhances MK secretion. A, CHO LRP1−/− or CHO-mLRP-IV cells were transfected with a combination of MK, RAP, or control vector (−). Results of Western blot analyses are shown. Graphs represent the relative density of each MK band appearing in the Medium. B, CHO-mLRP-IV cells were transfected with MK and IRES-EGFP (RAP−) or MK and RAP-IRES-EGFP (RAP+) and then stained with anti-MK antibody. Arrows indicate transfected cells. In the bottom row, CHO-mLRP-IV cells were transfected with MK-vector. C, TGW cells, a human neuroblastoma cell line, were treated with 10−6 m retinoic acid for 24 h. RNA and protein expressions were examined by Northern and Western blot analyses, respectively. D, LRP1 and MK localization in TGW cells treated with 10−6 m retinoic acid is shown. E, TGW cells were co-transfected with GFP- and RAP- or control vector and then treated with 10−6 m retinoic acid for 48 h. The population of GFP-expressing cells, which represented RAP- or control vector-transfected cells, among detached (dying) cells was determined by FACS. The expression efficiency as estimated on the day after transfection was comparable (∼17%) between the RAP and control vectors. *, p = 0.015 (Student's t test).
When MK was co-expressed with GFP in mLRP-IV-expressing cells, the intracellular MK was densely stained in nearly detaching, round-shaped cells (Fig. 2B, top row, arrowhead), whereas MK staining became faint by RAP expression (Fig. 2B, middle row, arrowheads). Intracellular localization of MK overlapped that of mLRP-IV (Fig. 2B, bottom row). Together, these results suggest that LRP1 is essential for intracellular retention of MK during biosynthesis: this retention is probably due to intracellular interaction between MK and LRP1.
Because MK was originally discovered as a retinoic acid-responsive gene product (2), we next investigated endogenous MK protein production using this reagent. Retinoic acid induced MK mRNA expression and consequently intracellular MK protein production in TGW cells, a neuroblastoma cell line (Fig. 2C, RNA and Cell). However, surprisingly, it did not enhance MK protein secretion (Fig. 2C, Medium). After retinoic acid treatment, LRP1 was detected not only on the cell surface (Fig. 2D, arrows), but also as a densely stained intracellular body in nearly detaching, round-shaped cells (Fig. 2D, arrowheads). MK was similarly stained in round cells (Fig. 2D), with a pattern reminiscent of MK staining in CHO-mLRP-IV cells (Fig. 2B). The detaching, round-shaped cells were thought to be in a state of anoikis, cell death triggered by detachment from the matrix (28). We then tested the effect of RAP on retinoic acid-induced cell death. This chaperone was expected to relieve the ligand-receptor aggregation, based on the results of Fig. 2, A and B. Indeed, RAP overexpression significantly reduced cell death (Fig. 2E).
MK Interacts with Immature LRP1 and Forms Aggregates during Biosynthesis
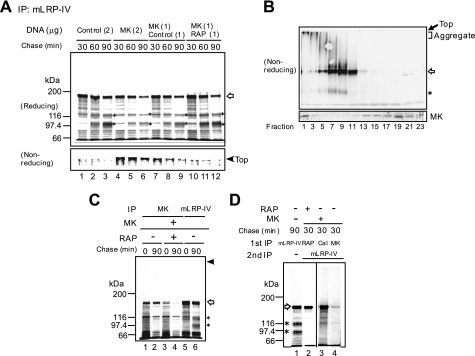
We next examined the effect of MK expression on LRP1 maturation. Pulse-chase metabolic labeling revealed that the unprocessed (immature) mLRP-IV appeared first (Fig. 3A, open arrow), and then the processed (mature) mLRP-IV (α and β chains) increased after 60–90-min chase (Fig. 3A, lanes 1–3, asterisks). This profile indicates that the furin-mediated cleavage of immature LRP1 gives rise to mature LRP1, which is consequently presented on the cell surface. A forced expression of MK strongly suppressed LRP1 maturation (Fig. 3A, lanes 4–6). However, if RAP was co-expressed with MK, LRP1 maturation was restored (Fig. 3A, lanes 7–12).
FIGURE 3.
MK binds to unprocessed LRP1 during biosynthesis. A, CHO-mLRP-IV cells were co-transfected with MK- and RAP-vector or control vector (−). Pulse-chase-labeled proteins were immunoprecipitated with an anti-HA antibody that detects mLRP-IV. Open arrow, unprocessed mLRP-IV; asterisks, processed mLRP-IV. B, sucrose gradient analysis was performed for CHO-mLRP-IV cells transfected with an MK-vector as described under “Experimental Procedures.” Western blot analysis was performed using anti-HA (upper panel, mLRP-IV) or anti-MK antibody (lower panel). The open arrow and asterisk indicate unprocessed and processed mLRP-IV, respectively. C, CHO-mLRP-IV cells were transfected with expression vectors as indicated. Immunoprecipitation was performed with anti-MK or anti-HA antibody. Open arrow, unprocessed mLRP-IV; asterisks, processed mLRP-IV. D, CHO-mLRP-IV cells were transfected with expression vectors as indicated. Pulse-chase labeling was performed, and then a sequential immunoprecipitation was performed with the indicated antibodies. Open arrow, unprocessed mLRP-IV; asterisks, processed mLRP-IV.
It is known that LRP1 aggregates are formed via intermolecular disulfide bonds and appear at the top of the separation gel under nonreducing conditions (29). In the present study, the aggregates increased in cells expressing MK alone but decreased in cells co-expressing MK and RAP (Fig. 3A, Nonreducing). Taken together, these data suggest that the intracellular MK-LRP1 interaction affects LRP1 maturation.
We next addressed the question of whether or not both MK and LRP1 were involved in aggregate formation. In a sucrose density gradient segregation of cellular proteins, bands at the top of the separation gel and smears underneath were detected in void fractions that contained higher molecular mass proteins under a nonreducing condition (Fig. 3B, fractions 1–5). It has previously been shown that the unprocessed and processed mLRP-IV bands migrate separately under a nonreducing condition (29), and in the present study these bands were detected in fractions 5–11 (open arrow and asterisk, respectively). MK was detected in the first fraction in addition to the later fractions under the reducing condition, suggesting that MK was also a component of the aggregates (Fig. 3B).
To examine the MK-LRP association, we performed immunoprecipitation (IP). When pulse-chase-labeled proteins were precipitated with anti-HA antibody (namely, mLRP-IV was precipitated), the unprocessed and processed mLRP-IV were detected at the 90-min chase (open arrow and asterisks, respectively in Fig. 3C, lane 6). In contrast, anti-MK antibody co-precipitated the unprocessed mLRP-IV but not the processed one at the 90-min chase (Fig. 3C, lane 2). Importantly, when RAP was co-expressed with MK, the MK-mLRP association was strongly suppressed (Fig. 3C, lanes 3 and 4). Fig. 3, B and C, suggests that MK and immature LRP1 formed a complex during biosynthesis.
To confirm the MK-immature LRP association further, pulse-chase-labeled proteins were sequentially immunoprecipitated by two different antibodies. Fig. 3D shows that RAP formed a complex with immature mLRP-IV but not mature mLRP-IV (Fig. 3D, lane 2). Similarly, the general ER chaperone calnexin as well as MK alone was associated with immature mLRP-IV (Fig. 3D, lanes 3 and 4).
ER-resident LRP1 Fragment Suppresses MK Secretion and LRP1 Maturation
As shown in Fig. 1E, the N-terminal half of the second ligand-binding domain of LRP1 (MK-TRAP) has a high affinity for MK. MK-TRAP traps extracellular MK, blocks MK binding to cell surface LRP1, and consequently suppresses MK function (27). We next utilized this fragment to investigate the biological role of the immature ligand-receptor interaction during biosynthesis. To this end, MK-TRAP was connected with an ER-retrieval signal found in RAP (the amino acid sequence is HNEL), so that MK-TRAP was retained in the ER. We named this construct ER-TRAP (Fig. 1A). Because RAP also shows high affinity for the LRP fragment corresponding to MK-TRAP (25), ER-TRAP was expected to trap both MK and RAP in the ER, leading to the suppression of MK secretion and LRP1 maturation.
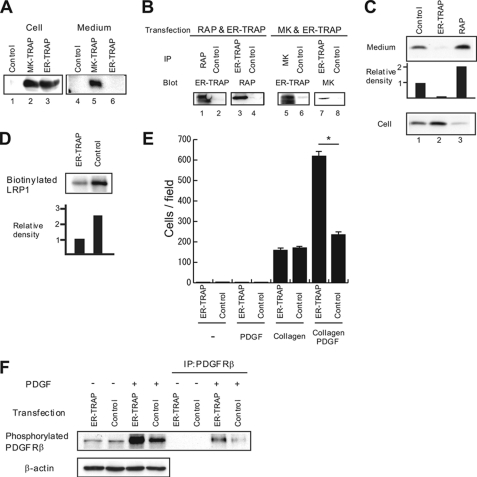
As shown in Fig. 4A, MK-TRAP could be detected in both the cell and the medium, whereas ER-TRAP was only detected in the cell, indicating that ER-TRAP was properly produced as expected. Immunoprecipitation revealed that ER-TRAP bound to both MK and RAP (Fig. 4B). ER-TRAP almost completely trapped MK in the cell, leading to cessation of MK secretion, whereas RAP reversed this phenomenon (Fig. 4C). ER-TRAP also significantly suppressed endogenous MK secretion in the osteosarcoma cell line, UMR106 (data not shown).
FIGURE 4.
ER-TRAP traps MK in the cell. A, the indicated expression vectors were transfected into CHO K1 cells. Proteins from cell lysates or media were subjected to Western blot analysis with anti-HA antibody that recognizes MK-TRAP and ER-TRAP. B, the indicated expression vectors were transfected into CHO K1 cells. Anti-HA and anti-FLAG antibodies were used to detect ER-TRAP and RAP, respectively. C, an MK-vector was co-transfected with the indicated expression vectors into CHO-mLRP-IV cells. Proteins were then detected by Western blot analysis. D, NIH3T3 cells were transfected with ER-TRAP or control vector. Cell surface proteins were labeled with biotin, immunoprecipitated (IP) with anti-LRP1 antibody, and separated under 5% SDS-PAGE. Biotinylated LRP1 was detected with HRP-conjugated streptavidin. E, NIH3T3 cells were transfected with ER-TRAP or control vector. Cell migration analysis was then performed. *, p < 0.001 (Student's t test). F, NIH3T3 cells transfected with ER-TRAP or control vector were incubated in 0.2% BSA in DMEM for 3 h and stimulated with 50 ng/ml PDGF-BB for 10 min. Phosphorylation of PDGF receptor was detected with anti-phosphotyrosine antibody (Tyr(P)-100).
We next examined the effect of ER-TRAP on endogenous LRP1 production. NIH3T3 cells are known to express endogenous LRP1 but not MK. ER-TRAP strongly suppressed the cell surface LRP1 production, as estimated by immunoprecipitation after cell surface biotinylation (Fig. 4D). To evaluate the biological significance of this phenomenon, we next examined PDGF-mediated migration because LRP1 is known to suppress PDGF receptor β function, as revealed by a study in which vascular smooth muscle cell-specific knock-out of LRP1 induced increased phosphorylation of PDGF receptor β and enhanced atherosclerosis (30). Although PDGF-BB did not significantly induce migration of NIH3T3 cells when administered alone, it did so in the presence of coated collagen I (Fig. 4E). ER-TRAP-transfected cells showed significantly enhanced migration compared with vector-transfected cells, whereas no difference was observed in collagen I-mediated migration (Fig. 4E). Moreover, PDGF receptor β phosphorylation in response to exogenous PDGF-BB was induced to a larger extent in ER-TRAP-transfected cells compared with vector-transfected cells (Fig. 4F). These results suggest that reduction of cell surface LRP1 reverses LRP1-mediated suppression of PDGF receptor β function.
Increased RAP Expression Is Closely Associated with MK Expression in Human Colorectal Carcinomas
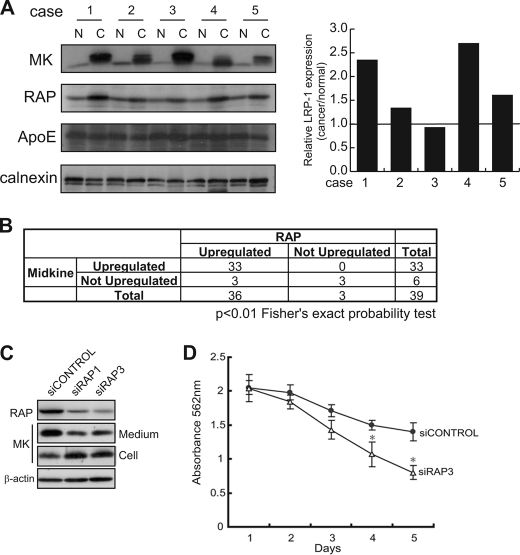
Human carcinomas frequently overproduce MK, and serum MK levels are elevated in cancer patients (9, 10, 13, 26). Based on our findings, we hypothesized that a mechanism preventing MK-LRP1 intracellular aggregation is involved in carcinogenesis. We examined the protein expression of MK and RAP in human colorectal carcinomas and corresponding adjacent normal tissues. In most of the cases examined (33 of 39), MK and RAP were strongly expressed in carcinoma tissues (Fig. 5, A left, and B). As controls for MK (a ligand of LRP1) and RAP (an ER chaperone), apolipoprotein E (apoE) and calnexin, respectively, were also examined. These control molecules did not show any difference of expression between carcinoma and adjacent normal tissues (Fig. 5A, left). Real-time RT-PCR examination of the mRNA expression of LRP1 in carcinoma tissues revealed that there were few cases in which the LRP1 expression was reduced compared with the adjacent normal tissue. Thus, among the 20 cases examined, there were 12 cases of up-regulation, 6 cases of no change, and 2 cases of down-regulation (the data of 5 cases corresponding to those in Fig. 5 left are shown in Fig. 5 right). These results indicate that the MK-LRP1 autocrine loop was activated, and the enhanced expressions of RAP and MK are closely associated in human colorectal carcinomas. To confirm the idea that RAP plays a pivotal role in tumor propagation by preventing premature interaction between MK and LRP1, we performed knockdown experiments for RAP of the human colorectal adenocarcinoma cell line SW620. Successful knockdown of RAP resulted in intracellular retention of MK and suppression of MK secretion and consequently in the suppression of cell survival under low serum conditions (Fig. 5D).
FIGURE 5.
Productions of MK and RAP are closely associated in human colorectal carcinomas. A, human colorectal carcinomas and adjacent normal tissues were subjected to Western blot analysis for MK, RAP, apoE, and calnexin protein expression and quantitative real-time RT-PCR for LRP1 mRNA expression. Five of 39 cases examined are shown here. C, carcinoma tissue; N, adjacent normal tissue. B, statistical analysis. C, SW620 cells were transfected with the indicated siRNAs. After 48 h, cells were harvested, and proteins were analyzed by immunoblotting. D, SW620 cells transfected with the indicated siRNAs were seeded, and the cell numbers were monitored by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. *, p < 0.01 (t test). siRAP1 also gave similar results with siRAP3 (data not shown).
DISCUSSION
Since the initial proposal of the concept of “autocrine secretion” by Sporn and Todaro (31), an increasing body of evidence has confirmed its biological significance. In the late 1980s, a biologically productive premature interaction, i.e. signal transduction via intracellular ligand-receptor binding, was reported for artificially ER-retained interleukin-3 and PDGF (32–34). These were the first demonstration that the premature interaction might have a biological role. However, our present data appear to be in conflict with these reports. The importance of our findings is that a growth factor and its receptor come to contact during biosynthesis, and this premature interaction negatively regulates their production within the secretory pathway. Avoiding this premature interaction may allow cells to overproduce a growth factor and thus to acquire a growth advantage. Indeed, here, we found that the LRP chaperone RAP blocks the premature MK-LRP1 interaction/aggregate formation and that increased RAP expression is closely associated with MK expression in human colorectal carcinomas. Consistent with this, serum MK levels have been shown to be elevated in human colorectal carcinoma patients (10). Thus, our findings highlight the biological significance of a premature interaction in the autocrine secretory pathway.
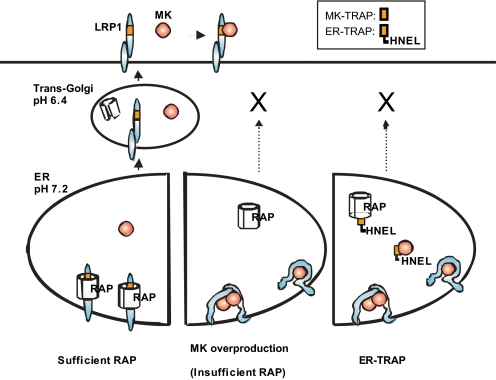
LRP1 is the fastest endocytosis receptor among the LDL receptor family members (35), and its endocytosis-dependent function has been intensively studied. For example, it has been determined that LRP1-mediated endocytosis of MK leads to nuclear targeting by endocytosed MK, which is, at least in part, responsible for MK-induced cell survival (22). The production and clearance of amyloid-β (Aβ), a pathogenic peptide of Alzheimer disease, are also dependent on the LRP1-mediated endocytosis. Thus, amyloid precursor protein (APP), a membrane protein, is intracellularly bridged with LRP1 by FE65 (36, 37). APP also binds to LRP1 extracellularly (38). The consequent APP-LRP1 complex is endocytosed and is further processed to produce Aβ. In regard to clearance of Aβ, extracellular Aβ binds directly to LRP1 and also binds indirectly to LRP1 through Aβ-binding molecules, including apoE (39–41); after these interactions, the Aβ is degraded intracellularly. In this context, our study has highlighted another function of LRP1, i.e. regulation of ligand biosynthesis. To the best of our knowledge, this is the first demonstration of a receptor playing an essential role in the biosynthesis of its ligand. We demonstrated that MK overexpression leads to the formation of aggregates between MK and LRP1, thereby suppressing MK secretion. Endogenous MK induced by retinoic acid treatment is also co-localized with LRP1 within the cell and thus is not secreted. The premature MK-LRP interaction may occur under both physiological and pathological conditions. It is conceivable that MK secretion may parallel its mRNA and protein expression until RAP is capable of blocking the premature MK-LRP1 interaction, but it may be impaired once MK protein expression overcomes RAP capacity (Fig. 6).
FIGURE 6.
Mechanisms of biosynthetic regulation for MK and LRP1. Under physiological conditions, RAP promotes folding of LRP1 and prevents premature interaction between LRP1 and MK. If RAP becomes insufficient under pathological conditions, such as overproduction of MK during carcinogenesis, aggregate formation involving LRP1 and MK takes place. ER-TRAP may competitively occupy RAP, leading to aggregate formation of LRP1. ER-TRAP also directly binds and traps MK in the ER. Consequently, ER-TRAP suppresses both LRP1 maturation and MK secretion. MK-TRAP is the N-terminal half of the second ligand-binding domain of LRP1, which corresponds to SLRP2N-EGF (see “Experimental Procedures”). ER-TRAP is the MK-TRAP plus HNEL (an ER-retrieving signal).
MK and apoE are LRP1 ligands. Interestingly, both MK and apoE can bind to Aβ, and MK, apoE, and LRP1 are found in senile plaques (39, 41–44). In addition, MK and apoE also show similar characteristics from the point of view of premature interaction with LRP1: overexpression of apoE or MK suppresses LRP1 maturation and induces LRP1 aggregation, which is restored by RAP expression (20; this study). In contrast, we found that the associated protein expressions are observed for RAP and MK, but not apoE, in human colorectal carcinomas. This indicates that MK and apoE have distinct biological significance and suggests that the biosyntheses of MK and apoE are differentially regulated, at least in these cancer cells.
It should be noted that other LRP1 ligands, such as tPA, urokinase, and plasminogen activator inhibitor-1, do not induce LRP1 aggregation (20). As discussed previously (20), formation of ligand-receptor aggregation may require polymerization of the ligand itself and/or ability of the ligand to bind to multiple ligand-binding domains of the receptor. A previous study reported that MK forms a homodimer (45), and our present data indicate that MK binds to the second and fourth ligand-binding domains of LRP1. It is known that apoE can form small multivalent lipoprotein particles and binds to three ligand-binding domains of LRP1. In contrast, tPA and PAI-1 may bind to a single site of LRP1 (46). This could be a reason why MK forms an aggregate with LRP1 within the secretory pathway. However, the precise mechanisms underlying the premature ligand-LRP1 interaction remain to be elucidated.
The results of ER-TRAP not only support the significance of the premature interaction in the biosynthesis of MK and LRP1, but also provide an intriguing strategy by which simultaneous repression of a growth factor and its receptor can be achieved. This strategy allows efficient trapping of a growth factor and a chaperone specialized for its receptor within the ER (Fig. 6). It may be applicable to certain disease treatments aimed at the suppression of ligand and/or receptor expression.
Acknowledgments
We thank Sachie Shimada, Miho Okada, Motoki Sato, and Junna Yamaguchi for excellent technical assistance in the initial stage of this study.
This work was supported, in whole or in part, by National Institutes of Health Grant R01 AG027924 (to G. B.). This work was also supported by Grant-in-aid 20390092 from Ministry of Education, Culture, Sports, Science and Technology, Japan (to K. K.) and by the Global Center of Excellence program from Ministry of Education, Culture, Sports, Science and Technology, Japan (Nagoya University).
G. Bu, unpublished data.
- ER
- endoplasmic reticulum
- APP
- amyloid precursor protein
- LRP1
- LDL receptor-related protein 1
- MK
- midkine
- mLRP
- mini-LRP
- RAP
- receptor-associated protein.
REFERENCES
- 1. Sporn M. B., Roberts A. B. (1992) Ann. Intern. Med. 117, 408–414 [DOI] [PubMed] [Google Scholar]
- 2. Kadomatsu K., Tomomura M., Muramatsu T. (1988) Biochem. Biophys. Res. Commun. 151, 1312–1318 [DOI] [PubMed] [Google Scholar]
- 3. Tomomura M., Kadomatsu K., Matsubara S., Muramatsu T. (1990) J. Biol. Chem. 265, 10765–10770 [PubMed] [Google Scholar]
- 4. Rauvala H., Huttunen H. J., Fages C., Kaksonen M., Kinnunen T., Imai S., Raulo E., Kilpeläinen I. (2000) Matrix Biol. 19, 377–387 [DOI] [PubMed] [Google Scholar]
- 5. Deuel T. F., Zhang N., Yeh H. J., Silos-Santiago I., Wang Z. Y. (2002) Arch. Biochem. Biophys. 397, 162–171 [DOI] [PubMed] [Google Scholar]
- 6. Kadomatsu K., Muramatsu T. (2004) Cancer Lett. 204, 127–143 [DOI] [PubMed] [Google Scholar]
- 7. Horiba M., Kadomatsu K., Nakamura E., Muramatsu H., Ikematsu S., Sakuma S., Hayashi K., Yuzawa Y., Matsuo S., Kuzuya M., Kaname T., Hirai M., Saito H., Muramatsu T. (2000) J. Clin. Invest. 105, 489–495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Hobo A., Yuzawa Y., Kosugi T., Kato N., Asai N., Sato W., Maruyama S., Ito Y., Kobori H., Ikematsu S., Nishiyama A., Matsuo S., Kadomatsu K. (2009) J. Clin. Invest. 119, 1616–1625 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Tsutsui J., Kadomatsu K., Matsubara S., Nakagawara A., Hamanoue M., Takao S., Shimazu H., Ohi Y., Muramatsu T. (1993) Cancer Res. 53, 1281–1285 [PubMed] [Google Scholar]
- 10. Ikematsu S., Yano A., Aridome K., Kikuchi M., Kumai H., Nagano H., Okamoto K., Oda M., Sakuma S., Aikou T., Muramatsu H., Kadomatsu K., Muramatsu T. (2000) Br. J. Cancer 83, 701–706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Nakagawara A., Milbrandt J., Muramatsu T., Deuel T. F., Zhao H., Cnaan A., Brodeur G. M. (1995) Cancer Res. 55, 1792–1797 [PubMed] [Google Scholar]
- 12. Mashour G. A., Ratner N., Khan G. A., Wang H. L., Martuza R. L., Kurtz A. (2001) Oncogene 20, 97–105 [DOI] [PubMed] [Google Scholar]
- 13. Ikematsu S., Nakagawara A., Nakamura Y., Sakuma S., Wakai K., Muramatsu T., Kadomatsu K. (2003) Br. J. Cancer 88, 1522–1526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Takei Y., Kadomatsu K., Matsuo S., Itoh H., Nakazawa K., Kubota S., Muramatsu T. (2001) Cancer Res. 61, 8486–8491 [PubMed] [Google Scholar]
- 15. Takei Y., Kadomatsu K., Goto T., Muramatsu T. (2006) Cancer 107, 864–873 [DOI] [PubMed] [Google Scholar]
- 16. Herz J., Strickland D. K. (2001) J. Clin. Invest. 108, 779–784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Herz J., Clouthier D. E., Hammer R. E. (1992) Cell 71, 411–421 [DOI] [PubMed] [Google Scholar]
- 18. Bu G., Geuze H. J., Strous G. J., Schwartz A. L. (1995) EMBO J. 14, 2269–2280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Bu G., Rennke S. (1996) J. Biol. Chem. 271, 22218–22224 [DOI] [PubMed] [Google Scholar]
- 20. Willnow T. E., Rohlmann A., Horton J., Otani H., Braun J. R., Hammer R. E., Herz J. (1996) EMBO J. 15, 2632–2639 [PMC free article] [PubMed] [Google Scholar]
- 21. Muramatsu H., Zou K., Sakaguchi N., Ikematsu S., Sakuma S., Muramatsu T. (2000) Biochem. Biophys. Res. Commun. 270, 936–941 [DOI] [PubMed] [Google Scholar]
- 22. Shibata Y., Muramatsu T., Hirai M., Inui T., Kimura T., Saito H., McCormick L. M., Bu G., Kadomatsu K. (2002) Mol. Cell. Biol. 22, 6788–6796 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Li Y., Marzolo M. P., van Kerkhof P., Strous G. J., Bu G. (2000) J. Biol. Chem. 275, 17187–17194 [DOI] [PubMed] [Google Scholar]
- 24. Obermoeller-McCormick L. M., Li Y., Osaka H., FitzGerald D. J., Schwartz A. L., Bu G. (2001) J. Cell Sci. 114, 899–908 [DOI] [PubMed] [Google Scholar]
- 25. Obermoeller L. M., Warshawsky I., Wardell M. R., Bu G. (1997) J. Biol. Chem. 272, 10761–10768 [DOI] [PubMed] [Google Scholar]
- 26. Ye C., Qi M., Fan Q. W., Ito K., Akiyama S., Kasai Y., Matsuyama M., Muramatsu T., Kadomatsu K. (1999) Br. J. Cancer 79, 179–184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Chen S., Bu G., Takei Y., Sakamoto K., Ikematsu S., Muramatsu T., Kadomatsu K. (2007) J. Cell Sci. 120, 4009–4015 [DOI] [PubMed] [Google Scholar]
- 28. Frisch S. M., Screaton R. A. (2001) Curr. Opin. Cell Biol. 13, 555–562 [DOI] [PubMed] [Google Scholar]
- 29. Obermoeller L. M., Chen Z., Schwartz A. L., Bu G. (1998) J. Biol. Chem. 273, 22374–22381 [DOI] [PubMed] [Google Scholar]
- 30. Boucher P., Gotthardt M., Li W. P., Anderson R. G., Herz J. (2003) Science 300, 329–332 [DOI] [PubMed] [Google Scholar]
- 31. Sporn M. B., Todaro G. J. (1980) N. Engl. J. Med. 303, 878–880 [DOI] [PubMed] [Google Scholar]
- 32. Keating M. T., Williams L. T. (1988) Science 239, 914–916 [DOI] [PubMed] [Google Scholar]
- 33. Dunbar C. E., Browder T. M., Abrams J. S., Nienhuis A. W. (1989) Science 245, 1493–1496 [DOI] [PubMed] [Google Scholar]
- 34. Bejcek B. E., Li D. Y., Deuel T. F. (1989) Science 245, 1496–1499 [DOI] [PubMed] [Google Scholar]
- 35. Li Y., Lu W., Marzolo M. P., Bu G. (2001) J. Biol. Chem. 276, 18000–18006 [DOI] [PubMed] [Google Scholar]
- 36. Trommsdorff M., Borg J. P., Margolis B., Herz J. (1998) J. Biol. Chem. 273, 33556–33560 [DOI] [PubMed] [Google Scholar]
- 37. Pietrzik C. U., Yoon I. S., Jaeger S., Busse T., Weggen S., Koo E. H. (2004) J. Neurosci. 24, 4259–4265 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Kounnas M. Z., Moir R. D., Rebeck G. W., Bush A. I., Argraves W. S., Tanzi R. E., Hyman B. T., Strickland D. K. (1995) Cell 82, 331–340 [DOI] [PubMed] [Google Scholar]
- 39. Strittmatter W. J., Weisgraber K. H., Huang D. Y., Dong L. M., Salvesen G. S., Pericak-Vance M., Schmechel D., Saunders A. M., Goldgaber D., Roses A. D. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 8098–8102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Deane R., Wu Z., Sagare A., Davis J., Du Yan S., Hamm K., Xu F., Parisi M., LaRue B., Hu H. W., Spijkers P., Guo H., Song X., Lenting P. J., Van Nostrand W. E., Zlokovic B. V. (2004) Neuron 43, 333–344 [DOI] [PubMed] [Google Scholar]
- 41. Tamamizu-Kato S., Cohen J. K., Drake C. B., Kosaraju M. G., Drury J., Narayanaswami V. (2008) Biochemistry 47, 5225–5234 [DOI] [PubMed] [Google Scholar]
- 42. Yasuhara O., Muramatsu H., Kim S. U., Muramatsu T., Maruta H., McGeer P. L. (1993) Biochem. Biophys. Res. Commun. 192, 246–251 [DOI] [PubMed] [Google Scholar]
- 43. Strittmatter W. J., Saunders A. M., Schmechel D., Pericak-Vance M., Enghild J., Salvesen G. S., Roses A. D. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 1977–1981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Rebeck G. W., Reiter J. S., Strickland D. K., Hyman B. T. (1993) Neuron 11, 575–580 [DOI] [PubMed] [Google Scholar]
- 45. Kojima S., Muramatsu H., Amanuma H., Muramatsu T. (1995) J. Biol. Chem. 270, 9590–9596 [DOI] [PubMed] [Google Scholar]
- 46. Willnow T. E., Orth K., Herz J. (1994) J. Biol. Chem. 269, 15827–15832 [PubMed] [Google Scholar]