Abstract
Fifteen variants in 10q26 are in strong linkage disequilibrium and are associated with an increased risk for age-related macular degeneration (AMD), a frequent cause of blindness in developed countries. These variants tag a single-risk haplotype encompassing the genes ARMS2 (age-related maculopathy susceptibility 2) and part of HTRA1 (HtrA serine peptidase 1). To define the true AMD susceptibility gene in 10q26, several studies have focused on the influence of risk alleles on the expression of ARMS2 and/or HTRA1, but the results have been inconsistent. By heterologous expression of genomic ARMS2 variants, we now show that ARMS2 mRNA levels transcribed from the risk haplotype are significantly reduced compared with non-risk mRNA isoforms. Analyzing variant ARMS2 constructs, this effect could specifically be assigned to the known insertion/deletion polymorphism (c.(*)372_815del443ins54) in the 3′-untranslated region of ARMS2. Reporter gene assays with HTRA1 promoter sequences demonstrated the presence of a Müller glia-specific cis-regulatory region further upstream of the transcription start site. However, AMD risk alleles had little or no effect on HTRA1 promoter activity in the retina. Analysis of a large series of human post-mortem retina/retinal pigment epithelial samples heterozygous for the risk haplotype confirmed the in vitro/ex vivo results and demonstrated that the risk haplotype affects ARMS2 but not HTRA1 mRNA expression. Furthermore, we provide in vivo evidence that a common non-risk-associated non-synonymous variant (rs2736911) also leads to decreased ARMS2 transcript levels. Consequently, our data suggest that pathogenic effects due to ARMS2 protein deficiency are unlikely to account for AMD pathology.
INTRODUCTION
Age-related macular degeneration (AMD) is a degenerative disorder of the central retina and a major cause of legal blindness in industrialized countries (1). The etiology of AMD is complex with environmental and genetic risk factors influencing susceptibility to disease (2). Therapeutic options to halt the onset or prevent progression of the disease are limited and benefit only a minority of patients (3).
Major advances in identifying the genetic variants associated with AMD have revealed a strong connection between the complement cascade, importantly the regulation of innate immunity and disease susceptibility (4). Independent risk alleles were reproducibly identified in complement factor H (CFH) on 1q32 (5–8), the CFH-related genes 1 and 3 (CFHR1/CFHR3) on 1q32 (9,10), complement factor B on 6p21.3 (11–14), complement factor 3 (C3) on 19p13 (15,16) and complement factor I on 4q25 (17,18). Average effect sizes for individual risk alleles vary greatly. Consistent replication has also firmly established the apolipoprotein E gene (APOE) at 19q13.2 as an AMD-associated gene, although with minor contribution to overall disease load and thus far unknown effects on disease pathology (10,19,20).
Successive refinement of a prominent linkage signal on chromosome 10q26 (21–23) identified common variants with large effect sizes near the ARMS2 (age-related maculopathy susceptibility 2) and HTRA1 (Htra serine peptidase 1) genes establishing this site as a second major AMD susceptibility locus (24,25). A total of 15 variants in strong linkage disequilibrium were shown to tag a single-risk haplotype leaving statistics-based analyses with insufficient power to discriminate between these variants (26). Instead, it became obvious that functional dissection of the effects of each risk-associated variant in 10q26 is required to determine which of the two genes, ARMS2 or HTRA1, is the true AMD susceptibility gene.
Subsequent studies suggested that a functional polymorphism (rs11200638) in the putative promoter of HTRA1 results in increased expression (27,28). Further analyses, however, could not replicate these findings and instead suggested ARMS2 as a mitochondria-associated protein involved in AMD susceptibility (29). Finally, identification of a previously undetected insertion/deletion (indel) polymorphism in the 3′-untranslated region (UTR) of ARMS2 (c.(*)372_815del443ins54) suggested a causal role for ARMS2, as this variant was shown to result in an instable ARMS2 transcript (26).
Additional studies further addressed a possible influence of the 10q26-associated risk variants on ARMS2 or HTRA1 expression. For example, analysis of mRNA and protein expression in cultured human retinal pigment epithelial (RPE) cells revealed a 2–3-fold increase in HTRA1 expression on the risk haplotype (30). Similar results were obtained by in vitro reporter assays and by expression analysis in human placenta, also suggesting a 2–3-fold increased transcription rate of the risk-associated HTRA1 isoform (31). In contrast, two independent studies showed no differences in expression between risk- and non-risk-associated HTRA1 transcripts based on data from human lymphocytes and retina (32,33). Contradictory results were also found for risk-associated ARMS2 expression. By semi-quantitative sequencing and protein analyses, our group reported haploinsufficency for ARMS2 in human placenta and retina from carriers heterozygous for the risk-associated variants (26). This was recently confirmed by analyzing human placenta (31). In contrast, two other reports failed to detect alterations of ARMS2 expression in human retina and blood lymphocytes (33,34).
Adding another level of ambiguity is the common non-synonymous ARMS2 variant rs2736911 which results in a premature stop (R38Stop) and which should lead to deficiency of the ARMS2 protein similar to the risk-associated indel isoform (26). The rs2736911 variant, however, is not risk-associated thus contradicting the role of ARMS2 deficiency in AMD pathogenesis (35). Recently published data on placental tissue demonstrated markedly reduced ARMS2 mRNA expression in association with rs2736911 heterozygosity (31). Interestingly, a subsequent study failed to detect an rs2736911-dependent alteration of ARMS2 transcript levels (33).
We have now performed a comprehensive study addressing allelic and haplotypic effects of the 15 AMD risk-associated polymorphisms in 10q26 on risk- and non-risk-associated expression of ARMS2 and HTRA1. In addition, we have also included the non-risk-associated variant rs2736911 in this analysis. Both, in vitro and in vivo data clearly demonstrate reduced ARMS2 mRNA expression associated with the risk haplotype specifically ascribable to the c.(*)372_815del443ins54 mutation. In contrast, an influence of the 10q26 risk alleles on HTRA1 expression could not be substantiated. Importantly, we also show reduced ARMS2 retinal RNA expression on a heterozygous rs2736911 background. Together, our data suggest that a mere deficiency of the ARMS2 protein is unlikely to account for AMD pathology.
RESULTS
In order to refine the extent of the AMD-associated risk haplotype in 10q26, we imputed missing genotypes and genotypes of over 100 additional single nucleotide polymorphisms (SNPs) using data from our previous study (26). The genetic markers spanned a region of 174 kb encompassing the complete gene loci of PLEKHA1, ARMS2 and HTRA1. Besides the known risk-associated SNPs rs10490924, rs3750848, rs3750847, c.(*)372_815del443ins54, rs11200638, rs3793917 and rs932275, no additional risk variants were identified. The defined 20 kb haplotype segment perfectly overlapped the previously delineated candidate region [(26) and Supplementary Material, Fig. 1]. In addition, haplotype analysis revealed no additional common haplotype (Supplementary Material, Table S1).
Heterologous expression of ARMS2 variants in vitro
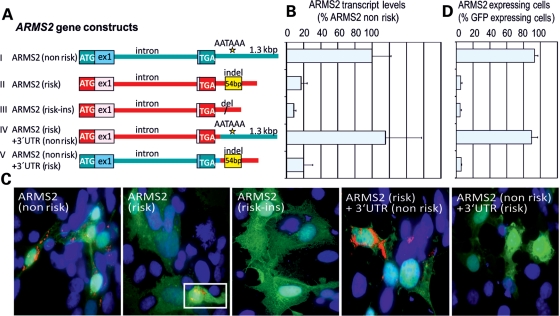
To assess risk- and non-risk-associated expression of ARMS2, we employed heterologous expression in COS-7 cells. Expression of ARMS2 from the risk haplotype H2 (Fig. 1A) and the major non-risk haplotype H1 (Fig. 1A) were analyzed by quantitative real-time (q) reverse transcriptase polymerase chain reaction (RT–PCR) and immunocytochemistry (Fig. 1 and Supplementary Material, Table S1). Compared with the non-risk isoform, the risk isoform carrying nine risk-associated variants revealed strongly reduced transcript (Fig. 1B) and protein expression (Fig. 1C and D), further supporting our previous findings (26).
Figure 1.
Heterologous expression of ARMS2 variants in COS-7 cells to assay the effects of AMD risk variants at 10q26 on ARMS2 transcript and protein expression. (A) Schematic overview of ARMS2 gene constructs. The ARMS2 expression vector simultaneously expresses GFP under the control of an independent promoter. (B) qRT–PCR analysis of ARMS2 cDNA expression. Results were normalized to GFP transcripts and calibrated to transcript levels of the non-risk-associated variant of ARMS2. Experiments were performed in triplicate wells in three independent experiments. The mean SD is given for each construct. (C) Immunocytochemistry of ARMS2 protein expression using an anti-ARMS2 antibody [(26) red staining]. GFP fluorescence (green) identified transfected cells. Nuclei were visualized by DAPI staining (blue). (D) Quantitation of immunocytochemical stainings showing the proportion of ARMS2 expressing cells relative to all transfected cells. Each 200 GFP expressing cells were analyzed for simultaneous ARMS2 staining.
As a likely candidate underlying this effect, we investigated the c.(*)372_815del443ins54 variant in more detail. In particular, we tested whether the influence of the 54 bp AU-rich insertion alone may explain the instability of the ARMS2 risk isoform. However, deletion of this element (Fig. 1A, construct III) failed to restore normal expression (Fig. 1B–D). In contrast, a chimeric construct replacing the 3′-UTR of the risk- with the non-risk-associated isoform (altering the c.(*)372_815del443ins54 risk variant but none of the remaining eight risk-associated ARMS2 variants) (Fig. 1A, construct IV) restored expression to normal levels (Fig. 1B–D). A reciprocal construct with the 3′-UTR of ARMS2 derived from the risk isoform, while exon1, intron 1 as well as exon 2 represent the non-risk isoform (Fig. 1A, construct V), again reveals strongly decreased expression similar to the risk-associated isoform (Fig. 1, construct II). Together, these data suggest that the indel polymorphism causes the decrease in ARMS2 mRNA expression, likely by the loss of the ARMS2 polyadenylation signal.
HTRA1 promoter activity in heterologous luciferase assays and murine retinal explants
Next, we performed reporter assays with HTRA1 promoter constructs in human RPE cell line ARPE-19 and rat Müller cell line rMC-1. Prior to generating the expression constructs, we determined the major retinal transcriptional start sites (TSSs) of HTRA1 by 5′-rapid amplification of complementary DNA (cDNA) ends (RACE) (Supplementary Material, Fig. S2). Three major TSSs were found at −128, −66 and +385 relative to the HTRA1 translational start site (TLS).
All promoter constructs were generated corresponding to the risk-(H2) and the major non-risk-associated (H1) haplotype (Supplementary Material, Table S1). To assess effects on regulatory elements around the TSSs and the start codon, we initially tested three luciferase expression constructs carrying different 3′-extensions of the genomic HTRA1 promoter (Fig. 2, constructs I–III). There was a strong decrease in promoter activity for promoter construct III with the 3′-end extending into the coding sequence up to +395 likely due to translation initiation from the upstream HTRA1 start codon instead of the luciferase start codon (36). The promoter fragment extending at the 3′-end to position +13 showed maximum activity and was used as template for further construct generation.
Figure 2.
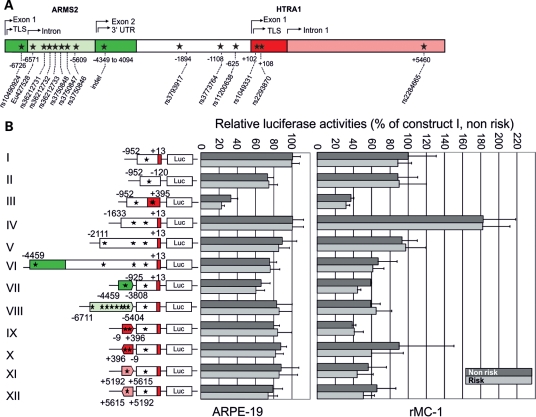
Heterologous luciferase assays assessing the effects of risk- and non-risk-associated variants on HTRA1 promoter activity in human-cultured RPE (ARPE-19) and rat-cultured Müller cells (rMC-1). (A) Schematic overview of the ARMS2/HTRA1 genomic region comprising the 15 AMD risk-associated variants and their position relative to the HTRA1 TLS. Asterisks display risk haplotype tagging polymorphisms, their relative positions are not to scale. (B) Luciferase activities of HTRA1 promoter constructs in ARPE-19 and rMC-1 cells. Each promoter construct (I–XII) was generated for the respective risk- and non-risk-associated variants. The pGL3-basic vector without insert served as a negative control (data not shown). A β-galactosidase expressing vector was cotransfected with each construct as an internal control for normalization. Normalized luciferase activity was measured in triplicate wells in three independent experiments. The mean SD is given for each construct.
We then aimed to analyze possible effects of each single-risk haplotype tagging variant in the HTRA1 promoter region and the ARMS2/HTRA1 intergenic region on HTRA1 promoter activity. Genomic HTRA1 promoter constructs with varying 5′-ends (Fig. 2, constructs I–VI) revealed comparable expression activities on the non-risk haplotype and no differences in any of the tested constructs compared with the respective risk-associated haplotype. Remarkably, even the longest genomic construct (Fig. 2, construct VI) which is largely similar to the one used in Yang et al. (31) failed to show risk-associated differences in promoter activity.
We also assessed risk-associated polymorphisms in 10q26 physically distant from the HTRA1 promoter. To this end, promoter construct I (Fig. 2B) was fused to genomic regions around the risk-associated variant rs10490924 in exon 1 of ARMS2 (Fig. 2B, construct VII), the seven variants EU427528, rs36212731, rs36212732, rs36212733, rs3750848, rs3750847 and rs3750846 in intron 1 of ARMS2 (Fig. 2B, construct VIII), the two variants rs1049331 and 2293870 in exon 1 of HTRA1 [Fig. 2B, constructs IX (forward orientation) and X (reverse orientation)], and the variant rs2284665 in intron 1 of HTRA1 [Fig. 2B, constructs XI (forward orientation) and XII (reverse orientation)]. None of the constructs revealed a significant effect on HTRA1 promoter activity. More importantly, when comparing risk- and non-risk-associated haplotypes, there was no measurable difference between the constructs analyzed.
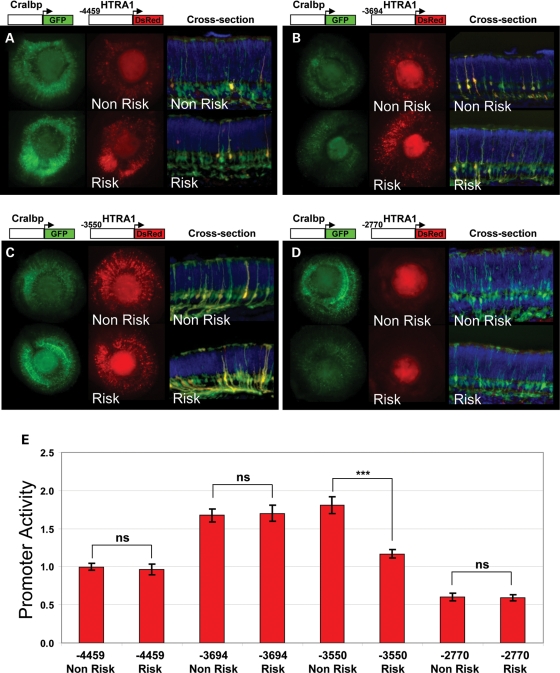
To investigate risk- and non-risk-associated HTRA1 promoter activity in the retina, HTRA1 promoter-DsRed reporter constructs were electroporated into newborn (P0) mouse retinas along with a loading control [the Müller glia-specific promoter from the cellular retinaldehyde binding protein, Cralbp, fused to green fluorescent protein (GFP)]. DsRed fluorescence was assayed at P8 and revealed HTRA1 promoter activity restricted to Müller glia (Fig. 3A–C). Two shorter promoter fragments (extending from −3694 to −5 and from −3550 to −5) exhibited a moderate increase in HTRA1 promoter activity relative to the full-length construct (−4459 to −5) (Fig. 3D). Further truncation resulted in a marked decrease in Müller glial expression (−2770 to −5) (Fig. 3D). These data suggest the presence of a Müller cell-specific cis-regulatory element between −3550 and −2770 bp upstream of the HTRA1 TLS.
Figure 3.
Electroporation of HTRA1 promoter constructs into murine retinal explants assaying the effect of the risk- and non-risk-associated variants in 10q26 on HTRA1 promoter activity. HTRA1 reporter constructs were generated for four promoter fragments extending from (A) −4459 to −5, (B) −3694 to −5, (C) −3550 to −5 and (D) −2770 to −5. Each construct was derived from a risk- and a non-risk-associated haplotype and cloned into a DsRed reporter construct. Each construct was co-electroporated with a Müller glia cell-specific Cralbp promoter driving GFP expression into P0 CD-1 mouse retinas and cultured as explants for 8 days. Retinas were fixed and imaged in both red and green channels as flat mounts, then embedded, sectioned and imaged in cross-section. All flat-mounts were exposed for the same length of time to permit comparison of strength of expression. (E) Quantitation of HTRA1 promoter activity. Each experiment was normalized to the HTRA1 −4459 to −5 non-risk construct; t-test performed for each non-risk, risk pair; ns, not significant; ***P < 0.05. Error bars indicate standard error.
Next, we quantified the DsRed fluorescence driven by both risk and non-risk versions of the HTRA1 promoter. We found no difference in expression between risk and non-risk variants for three of the four constructs (−4459, −3694 and −2770) (Fig. 3E). The −3550 construct showed a modest difference between risk and non-risk with the risk variant driving somewhat lower levels of expression. However, this difference was quite small. In summary, the in vivo and ex vivo data show little or no influence of the risk-associated alleles on HTRA1 promoter activity.
Endogenous retinal/RPE expression of ARMS2 and HTRA1
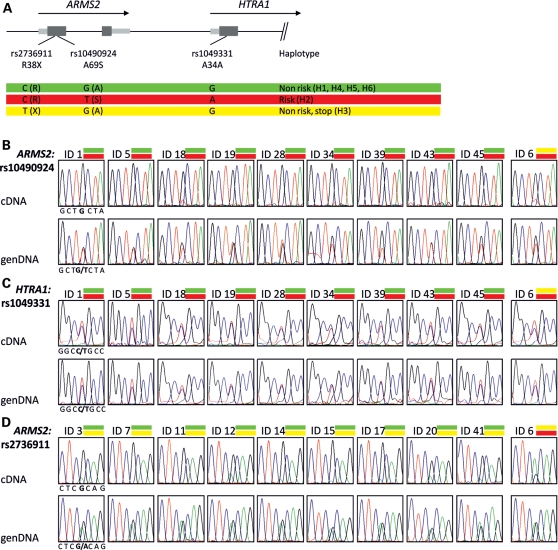
To analyze expression of ARMS2 and HTRA1 variants in vivo, we developed a semi-quantitative sequencing approach comparing genomic and cDNA sequences from human post-mortem retina/RPE samples (Supplementary Material, Fig. S3). Genomic sequencing identified 10 samples heterozygous for the risk-associated variants in ARMS2 (rs10490924, A69S) and HTRA1 (rs1049331, A34A) (Fig. 4A). Nine of these were also heterozygous for any one of the non-risk-associated haplotypes (H1, H4, H5 or H6) while sample ID6 was heterozygous for haplotype H3, carrying the stop variant at codon 38 (rs2736911). Moreover, we found an additional nine samples heterozygous for haplotype H3 and heterozygous for any one of the non-risk-associated haplotypes (H1, H4, H5 or H6). Together, these 19 samples facilitated tracing of heterozygous RNA expression of ARMS2 (Fig. 4B) and HTRA1 (Fig. 4C) originating from the risk-associated haplotype as well as ARMS2 expression from haplotype H3 harboring the 38Stop mutation (Fig. 4D). Analyzing ARMS2 expression, all cDNA samples revealed exclusively the presence of the non-risk-associated ARMS2 transcript (Fig. 4B). In contrast, HTRA1 transcript levels of risk- and non-risk-isoform were detectable in cDNA sequences at intensities similar to the genomic sequence profiles (Fig. 4C).
Figure 4.
Retinal/RPE RNA expression of ARMS2 and HTRA1 in samples heterozygous for haplotype tagging variants in 10q26. Relative expression of ARMS2 and HTRA1 isoforms was analyzed by semi-quantitative sequencing of allele-specific transcripts. (A) Schematic diagram of 10q26 haplotypes and relative position of variants used for identifying allele-specific ARMS2 or HTRA1 isoforms. (B) Heterozygous risk/non-risk-associated alleles at the ARMS2 gene locus as determined by genomic sequencing of variant rs10490924 (A69S). RT–PCR analyses of the same samples reveal exclusively the presence of the non-risk-associated ARMS2 allele. (C) Heterozygous risk-/non-risk-associated alleles at the HTRA1 gene locus as determined by genomic sequencing of variant rs1049331 (A34A). RT–PCR analyses of the same samples reveal the presence of both the risk- and non-risk-associated HTRA1 alleles. (D) Heterozygous non-risk-associated ARMS2 38Stop isoform (H3 haplotype) as determined by genomic sequencing of variant rs2736911 (R38Stop). RT–PCR analyses of the same samples reveal greatly reduced allele intensities of the H3 haplotype. Note that the H3 haplotype in ID6 reveals minor expression over the expression from the H2 risk haplotype.
To further assess ARMS2 transcript expression from the non-risk-associated haplotype H3, we sequenced human retina/RPE cDNA samples ID3, ID7, ID11, ID12, ID14, ID15, ID17, ID20, ID41 and ID6, all heterozygous at the genomic sequence for the H3 haplotype (rs2736911, Fig. 4D). Similar to the risk-associated ARMS2 isoform, the ARMS2 38Stop variant also revealed a significantly decreased expression although some samples appeared to show a less prominent signal reduction (ID12, ID20) (Fig. 4D). This was confirmed in ID6, a retina/RPE sample heterozygous for the ARMS2 risk and the 38Stop allele. This sample showed higher expression of the ARMS2 38Stop isoform than of the risk isoform (Fig. 4A and C) while no influence of the H3 haplotype was seen on HTRA1 expression (Fig. 4B).
As the semi-quantitative sequencing has a sensitivity of allelic discrimination between the non-risk (H1), the risk (H2) or the 38Stop (H3) cDNA ARMS2 variant between 1:12.5 and 1:15 (Supplementary Material, Fig. S3), our data suggest that the ARMS2 risk-associated transcript is reduced over the non-risk transcript by a ratio of 1:12.5 or less, whereas the transcript carrying the 38Stop variant shows a reduction in expression in the range of 2.5–15-fold. HTRA1 cDNA in retinas heterozygous for the risk- and non-risk-associated variants indicated similar expression of the two isoforms.
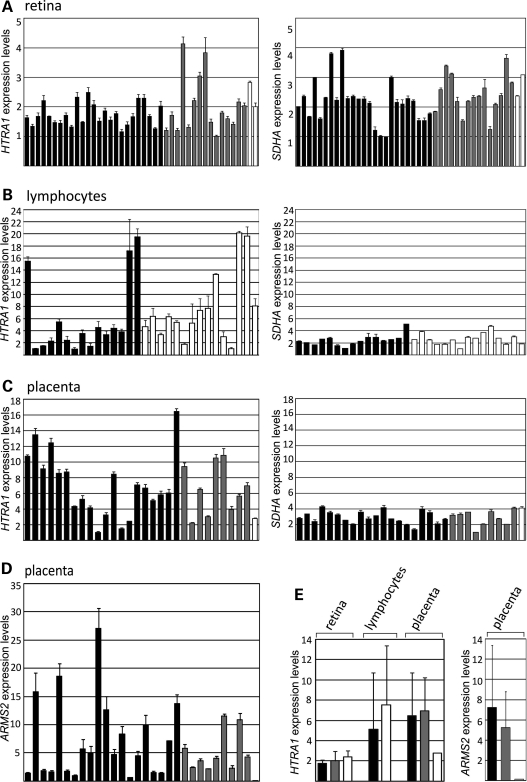
To further verify our findings of genotype-independent variation in HTRA1 expression, we performed qRT–PCR analyses with cDNAs from 45 retinas, 30 lymphocyte and 30 placental samples from human donors (Fig. 5A–C, left panels). For each sample, the haplotype status was determined. In retina, no statistically significant deviations of HTRA1 expression due to allelic differences were observed (Fig. 5A and E, left panels). In lymphocytes and placenta, we noticed striking inter-individual variability in HTRA1 expression independent of haplotype combinations. For example, expression in lymphocytes varied up to 20-fold in samples homozygous for the non-risk or the risk allele (Fig. 5B, left panel), and in placenta up to 16-fold in samples homozygous for the non-risk allele (Fig. 5C, left panel). Within this wide range of expression, there was no statistically significant deviation (Fig. 5E, left panel). To exclude methodological influences on the varying expression levels, we included qRT–PCR analysis of succinate dehydrogenase complex, subunit A, flavoprotein (SDHA), an additional independent housekeeper. SDHA expression was homogenous between individual samples with a normal variation of up to 4-fold differences (Fig. 5A–C, right panels). In addition, we reproduced the qRT–PCR results for HTRA1 in lymphocytes in a second independent cDNA sample preparations but using polyT primers instead of random primers and normalizing against SDHA instead of 18S rRNA (Supplementary Material, Fig. S4). Despite some minor differences in individual expression levels, the overall expression pattern was comparable among the two independent assays, indicating that the inter-individual variability in HTRA1 expression is not due to methodological inconsistencies.
Figure 5.
Endogenous expression of HTRA1 and ARMS2 in human tissues assaying the effect of the risk- and non-risk-associated variants in 10q26. HTRA1 expression levels were determined by qRT–PCR in (A, left panel) retinas of 45 unrelated individuals, (B, left panel) lymphocytes of 30 unrelated individuals, and (C, left panel) placentas of 30 unrelated individuals. As a control for quantity and quality, SDHA expression was also determined by qRT–PCR in these tissues (right panels). (D) ARMS2 expression in placental samples. (E) Mean HTRA1 and ARMS2 expression in retina, placenta and lymphocytes with respect to 10q26 haplotypes. The mean +SD is given for each sample or genotype, respectively. All samples were normalized to 18S rRNA. Black bars: homozygous for non-risk-associated 10q26 haplotypes; grey bars: heterozygous for non-risk/risk-associated haplotypes; white bars: homozygous for risk-associated 10q26 haplotype.
We also analyzed ARMS2 expression in the different human tissues. No ARMS2 transcripts were detected in lymphocytes (data not shown). As noted by Rivera et al. in 2005 (25), ARMS2 expression in retinas is very weak, resulting in absent amplification or inconsistent cycle thresholds in qRT–PCR experiments (data not shown). In placental samples, ARMS2 expression was strong (25), but similar to HTRA1 expression, exhibited a large inter-individual variability rendering the data statistically insignificant (Fig. 5D and E, right panel). Although there was only a single sample available with homozygosity in the risk haplotype, it is of note that ARMS2 expression was strikingly reduced when compared with the remaining placental samples. This is in agreement with a greatly reduced stability of the ARMS2 risk variant (26).
DISCUSSION
A large series of studies has focused on the impact of AMD risk-associated polymorphisms on the expression of ARMS2 and/or HTRA1. The data are widely conflicting and often contradictory. Possible reasons for these discrepancies include: methodological differences, intrinsic inter-individual variability of gene expression and small numbers of experimental samples. Here, we have undertaken a comprehensive analysis of genotype-dependent expression of ARMS2 and HTRA1 and directly compared the expression profiles of the two genes in a number of complementary in vitro and in vivo experiments. We consistently show that the AMD risk-associated haplotype in 10q26 leads to significantly reduced RNA expression of ARMS2 but exerts little or no influence on HTRA1 expression. Also, we demonstrate that the non-risk-associated 38Stop variant leads to significantly reduced ARMS2 transcript levels and thus likely results in deficiency of ARMS2 protein similar to the haplotype H2-associated decline in mRNA expression. We therefore reason that ARMS2 protein deficiency is unlikely to be the pathogenic mechanism predisposing to AMD.
Supporting our initial data on the ARMS2 indel polymorphism (c.(*)372_815del443ins54) (26), we have further delineated the mechanism resulting in reduced ARMS2 transcript levels of the mutant isoform. We tested whether the AU-rich elements present in the indel variant known to enhance mRNA decay (37,38) are, in fact, responsible for the decline of mutant ARMS2 mRNA transcript levels. Surprisingly, the deletion of the destabilizing elements had no effect on mutant ARMS2 expression. Instead, the presence of the regular polyadenylation signal sequence proved crucial for mRNA stability.
Addressing differential ARMS2 expression associated with the 38Stop variant (rs2736911), one study reported reduced expression of the mutant transcript (31), while two additional independent studies could not detect haplotype-related differences in ARMS2 transcript levels in human retina and blood samples (33,34). Focusing on these discrepancies, we noticed that Kanda et al. (33) used an ARMS2 reverse primer (5′-TTGCTGCAGTGTGGATGAT-3′) targeted to exon 2 of ARMS2. We found that this primer shows moderate homology, particularly within its 3′-region, to exon 1 of ARMS2, a region immediately upstream of the ARMS2 TLS (5′-GAGCTGCAGTGTGGATTTT-3′ homologous nucleotides are given in bold). In our hands, only high stringency of the RT–PCR conditions resulted in a locus-specific amplification of the targeted 259 bp product from the correctly spliced ARMS2 cDNA, while the alternative PCR fragment could derive from genomic exon 1 of ARMS2 and would only be 166 bp in size. Consequently, PCR results in Kanda et al. (33) could reflect amplification from spurious genomic DNA contamination regularly present in RNA samples. Likewise, comparable methodological problems might account for the results of Wang et al. (34) who designed both PCR primers, forward and reverse, within exon 1 of ARMS2 facilitating amplification from genomic DNA as well as cDNA samples. In our hands, qRT–PCR assays in retinal samples with ARMS2 exon 1 and 2-specific primers yielded inconsistent qRT–PCR results due to the weak ARMS2 expression in this tissue and subsequent cycle thresholds beyond a regular range.
Our in vitro and in vivo data provide no evidence for genotype-related differences in HTRA1 gene expression. This is in agreement with qRT–PCR data analyzing HTRA1 expression in human lymphocytes and retina samples (32,33). In contrast, two further studies reported a risk-associated up-regulation of HTRA1 in placental or RPE samples, respectively (30,31). The discrepant results may be partly due to the fact that expression is strongly influenced by individual factors. For example, HTRA1 gene expression is strongly up-regulated during aging and cellular stress (39–41) or as a consequence of disease-related conditions such as cancer or articular cartilage degeneration (42–44). Thus, individual differences in tissue donor samples may have a noticeable impact on HTRA1 expression. Together with the issue of small sample sizes, this could lead to chance findings in HTRA1 gene expression. Our data derived from three human tissues are in line with these considerations revealing a high degree of inter-individual variability in HTRA1 expression independent of a given genotype at the 10q26 AMD risk locus.
HTRA1 promoter DsRed reporter assays in electroporated mouse retinas revealed the presence of a previously unreported Müller glia-specific cis-regulatory element several kilobases upstream of the translation start. This finding is in keeping with a recent single-cell transcriptome analysis of Müller glia which showed high levels of HTRA1 (alias PRSS11 in that study) in this cell type (45). Although one of the four reporter constructs did show a modest difference between risk and non-risk variants on reporter activity, the difference was quite small. Given the technical difficulties of quantifying fluorescence in rare populations of electroporated cells and the absence of any difference in the other constructs examined, this difference should be interpreted with caution.
In summary, this study provides a comprehensive and in-depth analysis of genotype-dependent mRNA expression of ARMS2 and HTRA1. We show that the AMD risk-associated variants in 10q26 exert a dramatic effect on ARMS2 but not HTRA1 gene expression. We further demonstrate that this effect on the ARMS2 transcript level is entirely accounted for by the previously identified indel polymorphism c.(*)372_815del443ins54 (26), more specifically by the concomitant removal of the regular polyadenylation signal sequence. We finally show that a second variant (rs2736911, R38Stop) not associated with AMD risk also results in a prominent reduction in ARMS2 mRNA transcripts strongly suggesting that a mere deficiency of ARMS2 is not likely to underlie AMD etiology. Nevertheless, could decreased expression of ARMS2 still be a risk factor for AMD? In 2010, Yang et al. (31) have proposed a dual causality model which assumes concomitant down-regulation of ARMS2 and up-regulation of HTRA1 on the AMD-associated haplotype but not the protective haplotype containing the 38Stop variant. However, our data do not support such a model as we fail to detect 10q26 risk-associated alterations in HTRA1 expression. Still, the dual causality model with a second risk-conferring effect associated with ARMS2 down-regulation on the 10q26 risk haplotype is conceivable, although such a ‘second’ effect is elusive at present. Taken together, our results suggest that currently unknown mechanisms mediate the pathogenic effects of the risk-associated variants at the 10q26 AMD locus. Many important issues remain to be resolved. For example, it is still unclear whether ARMS2 encodes a protein or rather may exert its cellular function as a non-coding RNA. Discrepancies in previously published efforts to trace the endogenous ARMS2 protein (26,29,46,47) could support the latter possibility. A more remote possibility which has so far not been widely considered in the literature is that the sequence variants at the 10q26 locus might affect expression of a more distant gene and thereby mediate long-range pathogenic effects.
MATERIALS AND METHODS
Imputation
Imputation was performed with the MACH1 web interface of the HapMap project (http://hapmap.ncbi.nlm.nih.gov/) based on the genotype data established previously (26) and the data of the CEU HapMap data release #22 (The International HapMap Consortium 2007). The best-guess genotypes obtained were filtered for SNPs with Rsq ≥ 0.3 (a measure for the estimate of the squared correlation between imputed and true genotypes) and analyzed with Haploview 4.2 to estimate haplotype blocks (48), frequencies and associations (49).
Genotyping of human tissues and sequencing of ARMS2 and HTRA1 transcripts
Genotyping of human tissues was done for AMD-associated variants at 10q26 from genomic DNA or cDNA with primer combinations as given in Supplementary Material, Table S2. For sequencing, internal oligonucleotide primers were used (Supplementary Material, Table S2).
For analyzing ARMS2 expression in heterozygous retinas, genomic and cDNA first-strand PCR amplification was achieved with primers Loc-ex1-F2 and Loc-Ex2-R, followed by sequencing with primer rs10490924R for variant R38Stop and Loc-Ex1-F for variant A69S (Supplementary Material, Table S2). Genomic PCR amplification of HTRA1 was achieved with primers HTRA1-Ex1-F0 and HTRA1-Ex1-R2, while expression in the heterozygous retina was analyzed with primers HTRA1-Ex1-F0 and HTRA1-Ex2-R. Sequencing of genomic and first-strand cDNA PCR products was performed with primer HTRA1-Ex1-F (Supplementary Material, Table S2).
Sensitivity to discriminate heterozygous signals in ARMS2 and HTRA1 by semi-quantitative sequencing was determined by a titration approach of ARMS2 and HTRA1 full-length cDNAs of different haplotypes, ligated into the pGEM T vector (Promega, Madison, WI, USA).
Cell culture
ARPE-19 and rMC-1 cell lines were maintained in the DMEM/HamsF12 medium containing 10% FCS and 100 U/ml penicillin/streptomycin (PAA, Pasching, Austria). COS-7 cells were cultivated in the DMEM high-glucose medium containing 10% FCS and 100U/ml penicillin/streptomycin (PAA). All cell lines were grown in a 37°C incubator with a 5% CO2 environment.
ARMS2 constructs
For heterologous expression in COS-7 cells, ARMS2 was cloned into the pBUDCE4.1 vector (Invitrogen, Carlsbad, CA, USA), allowing the simultaneous expression of two genes under the control of two independent promoters in mammalian cell lines.
The GFP open reading frame was cloned into the NotI/XhoI site of the EF-1α promoter. GFP-fluorescence served as control for transfection efficiency which was consistently >50%, but also as selection marker for purifying transfected cells by fluorescence activated cell sorting. Two ARMS2 isoforms were PCR amplified from DNA of placental tissue and genotyped as described above. These included the most frequent non-risk-associated haplotype H1 and the risk-associated haplotype H2 (Supplementary Material, Table S1). Primers for amplification were constructed to contain a KpnI site 5′ at the ARMS2 start codon (Loc-F-KpnI) and a BamHI site at the 3′-end to the ARMS2 gene (PRSS11-SNP5-R2-BamHI).
Deletion of the 54 bp insertion of the indel polymorphism was achieved by PCR with primer Loc-F-KpnI and a reverse primer flanking the 5′-end of the insertion (ARMS2-EcoRV-R), and by a second PCR with a forward primer flanking the 3′-end of the insertion (ARMS2-F-3′-UTR-EcoRV-F) and PRSS11-SNP5-R2-BamHI. Before cloning into the vector, the fragments were ligated via their EcoRV sites. Synthesis of the chimeric ARMS2 constructs was performed by PCRs with primers Loc-F-KpnI and ARMS2-BamHI-R, and with ARMS2-BamHI-F and PRSS11-SNP5-R2-BamHI. Before cloning into the vector, these fragments were ligated via their BamHI sites.
For enforcing ARMS2 protein synthesis, the QBI SP163 translational enhancer of the pcDNA4/HisMax vector (Invitrogen) was modified by adding a HindIII site to its 5′-end and a KpnI site to its 3′-end. The QBI SP163 translational enhancer was ligated to the ARMS2 gene via the KpnI site and the fused fragments were ligated with the cytomegalovirus (CMV) promoter of the pBUD CE4.1 vector via the HindIII and BamHI sites. Primer sequences are given in Supplementary Material, Table S3. For verification, all cloned constructs were directly sequenced by capillary sequencing (Applied Biosystems, Darmstadt, Germany).
Expression of ARMS2 in COS-7 cells
COS-7 cells were seeded overnight into cell culture dishes with a diameter of 10 cm. At a confluence of 70%, the cells were transfected with 10 µg of the ARMS2 expression constructs containing the risk, the non-risk or the chimeric variants of ARMS2 and 30 µl TransIT-LT1 Transfection Reagent (Mirus Bio LLC, Madison, WI, USA) following the manufacturer's instructions. After 48h, cells were harvested for RNA isolation, or used in immunocytochemistry.
Immunocytochemistry
Immunocytochemistry was performed as described previously (50), staining with the polyclonal anti-ARMS2 antibody [1:100, (26)] and an Alexa-conjugated secondary antibody (1:1000, Molecular Probes, Eugene, OR, USA).
RNA isolation and reverse transcription
Total RNA was extracted from tissues or cell lines using the RNeasy Protect Mini Kit followed by DNAse I treatment (QIAGEN, Hilden, Germany) according to the manufacturer's instructions. Purity and integrity of the RNA was assessed on the Agilent 2100 bioanalyzer with the RNA 6000 Nano LabChip reagent set (Agilent Technologies, Boeblingen, Germany). The RNA was quantified spectrophotometrically and then stored at −80°C. First-strand cDNA synthesis was performed using RevertAid M-MuLV Reverse Transcriptase (Fermentas, St Leon-Rot, Germany) and random or a poly(dT) primers according to the manufacturer's instructions.
Determination of the major retinal/RPE-specific TSS of HTRA1
To determine the major retinal/RPE-specific TSS of HTRA1, 5′-RACE experiments were performed with RNA from retinal/RPE samples heterozygous for the risk haplotype H2. RNA was isolated as mentioned above. 5′-RACE was conducted with the FirstChoice RLM-RACE Kit (Applied Biosystems/Ambion, Austin, TX, USA) according to the manufacturer's instructions. Reverse primers for first- and second-round PCRs were positioned in HTRA1 exon 8 and HTRA1 exon 3, respectively (Supplementary Material, Table S4). Amplified RACE products were ligated into the pGEM-T vector (Promega). Ten single-positive clones for each ligation were sequenced using M13 forward and reverse primers. To confirm the 5′-RACE results, we performed additional PCR reactions with retinal cDNA. Forward primers were starting upstream (−142 to −128, HTRA1-pre-TSS-F) and downstream of the most distal transcription start sites (−117 to −102, HTRA1-TSS-F). The reverse primer was positioned in HTRA1 exon 2. As expected, no fragment was obtained with the HTRA1-pre-TSS-F, while a 641 bp fragment was obtained with forward primer HTRA1-TSS-F. Sequencing confirmed the latter PCR product to be the correctly spliced HTRA1 fragment (Supplementary Material, Fig. S2B).
HTRA1 promoter constructs
For luciferase assays, HTRA1 promoter fragments were cloned into the SacI/XhoI site of the luciferase vector pGL4.10 (Promega). All fragments were generated by PCR from human genomic DNA of non-risk- and risk-associated haplotypes. Primer sequences are given in Supplementary Material, Table S5. Continuous fragments were generated between –4459 and +397 (nucleotide positions are given relative to the HTRA1 TLS) by using SacI and XhoI flanked oligonucleotide primers (Supplementary Material, Table S5). For testing putative enhancer/silencer effects of risk-associated variants farther down- or upstream of the HTRA1 promoter, fragments around these polymorphisms were amplified with SacI-flanked primers and cloned into the SacI site at the 5′-end of the HTRA1 (−800 to +119) promoter construct.
For assaying HTRA1 promoter activity in ex vivo murine retinal and RPE explants, two HTRA1 promoter constructs were amplified from the existing luciferase constructs by using Htra-LucA-2-F (position −952 to −921 relative to the HTRA1 TLS) and Delins-LucA-F (position −4459 to −4160) primers and Htra-5-KpnI_R1 (position −30 to −5). Additional fragments, extending from −3694, −3550 and −2770 to −5, respectively, were generated from the longer fragment (position −4459 to −5) by using internal restriction sites. All fragments were ligated into the pGEM T easy vector (Promega) adding EcoRI restriction sites to the 5′- and 3′-ends of the promoter fragments. Subsequently, the inserts were cloned into the EcoRI site of a promoterless DsRED reporter vector. All constructs were directly sequenced by capillary sequencing (Applied Biosystems).
Luciferase assays
For transfection, ∼106 ARPE-19 or rMC-1 cells were seeded overnight into the six-well plates. They were then transfected with the HTRA1 promoter constructs using TransIT-LT1 Transfection Reagent (Mirus Bio LLC). A cotransfection with a vector containing the β-galactosidase open reading frame under the control of a CMV promoter was performed to normalize HTRA1 promoter activities. After 48 h, cells were harvested and lyzed in Reporter Lysis Buffer (Promega). For luciferase assays, 10 µl of cytosolic extract and 100 µl of Luciferase Assay Reagent (Promega) were used. Light emission was measured with a FluoStar Optima (BMG LABTECH GmbH, Offenburg, Germany). For β-galactosidase assay, 150 µl of cytosolic extract was mixed with 150 µl of a buffer consisting of 200 mm NaPO4, pH 7.3, 2 mm MgCl2, 100 mm mercaptoethanol and 1.33 mg/ml o-Nitrophenyl-β-d-galactopyranoside, and incubated for 30 min at 37°C. The reaction was stopped by the addition of 500 µl 1 m Na2CO3. Relative β-galactosidase levels were evaluated by measuring the optical density at 420 nm. Luciferase activities were normalized to the relative β-galactosidase levels. Each experiment was performed at least three times in triplicate wells.
Mouse retinal explant electroporation and quantification
In vitro electroporation and explant cultures of mouse retinas were carried out as described previously (51). In brief, eyes were enucleated from P0 CD1 mice under sterile conditions. The retinas were dissected, subjected to electroporation, and cultured as explants for 8 days. Subsequently, they were washed, fixed and analysed as flat-mounts and sections as described (51) with the following modifications. P0 CD-1 retinas were co-electroporated with an HTRA1-DsRed and the Müller glia-specific Cralbp promoter driving GFP expression as a loading control (52). Upon harvesting, the retinas were briefly fixed with 4% paraformaldehyde and whole-mount images were captured with an Olympus BX 51 equipped with a DP70 camera using DP Controller software. Retinas were then embedded in OCT, and 14 μm thick sections were collected and imaged.
In order to quantify DsRed fluorescence within retinal cross-sections, four areas were imaged from each electroporated retina. Images were imported into Image J (NIH, Bethesda, MD, USA) where 15 regions of interest (containing electroporated cells) and three background control regions (containing no electroporated cells) were analyzed per image. A minimum of three separate electroporations with three retinas each were analyzed for each construct. The data from each experiment were normalized to the expression level of the full-length (−4459) HTRA1 non-risk reporter construct, and the normalized data were used to generate frequency tables and distribution plots in Microsoft Excel. A t-test (assuming unequal variances) was used to determine whether there was a significant difference in the expression level between the risk and non-risk constructs.
Quantitative real-time RT–PCR
Amplifications were performed in triplicates with 50 ng cDNA in an ABI7900HT (Applied Biosystems) in 10 μl reaction mixtures containing 1 × TaqMan Universal PCR Master Mix (Applied Biosystems), 200 nm of primers, and 0.25 μl dual-labelled probe (Roche ProbeLibrary, Roche Applied Science, Mannheim, Germany). The reaction parameters included 2 min hold at 50°C, 30 min hold at 60°C and 5 min hold at 95°C, followed by 45 cycles of 20 s, 94°C melt and 1 min anneal/extension at 60°C. Results were analyzed with the ΔΔCt method for relative quantification. 18S rRNA was used for normalization in human tissues, and SDHA was used for normalization in cell culture. Primer sequences and Roche Library Probe numbers are listed in Supplementary Material, Table S6. For qRT–PCR of the 18S gene, the TaqMan Ready-Developed Assay Reagent for human 18S rRNA (Applied Biosystems) was used.
WEB RESOURCES
The URL for data presented herein is as follows: International HapMap interface/Impute data, http://hapmap.ncbi.nlm.nih.gov/.
SUPPLEMENTARY MATERIAL
Supplementary material is available at HMG online.
FUNDING
This work was supported in parts by grants from the National Health Institute (NIH) (EY018826 to J.C.), The American Health Assistance Foundation (to J.C.), the Deutsche Forschungsgemeinschaft (DFG) (WE1259/18-1, WE1259/19-1 to B.H.F.W.), The Ruth and Milton Steinbach Foundation New York (to B.H.F.W.) and the Alcon Research Institute (to B.H.F.W.).
ACKNOWLEDGEMENTS
We would like to thank Christina Sendlmeier for help with generating reporter constructs and Dr G. Huber (St Hedwig Clinics, Regensburg, Germany) for providing placental samples.
Conflict of Interest statement. None declared.
REFERENCES
- 1.Friedman D.S., O'Colmain B.J., Munoz B., Tomany S.C., McCarty C., de Jong P.T., Nemesure B., Mitchell P., Kempen J. Prevalence of age-related macular degeneration in the United States. Arch. Ophthalmol. 2004;122:564–572. doi: 10.1001/archopht.122.4.564. doi:10.1001/archopht.122.4.564. [DOI] [PubMed] [Google Scholar]
- 2.Swaroop A., Chew E.Y., Rickman C.B., Abecasis G.R. Unraveling a multifactorial late-onset disease: from genetic susceptibility to disease mechanisms for age-related macular degeneration. Annu. Rev. Genomics Hum. Genet. 2009;10:19–43. doi: 10.1146/annurev.genom.9.081307.164350. doi:10.1146/annurev.genom.9.081307.164350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Folk J.C., Stone E.M. Ranibizumab therapy for neovascular age-related macular degeneration. N. Engl. J. Med. 2010;363:1648–1655. doi: 10.1056/NEJMct1000495. doi:10.1056/NEJMct1000495. [DOI] [PubMed] [Google Scholar]
- 4.Gehrs K.M., Jackson J.R., Brown E.N., Allikmets R., Hageman G.S. Complement, age-related macular degeneration and a vision of the future. Arch. Ophthalmol. 2010;128:349–358. doi: 10.1001/archophthalmol.2010.18. doi:10.1001/archophthalmol.2010.18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Edwards A.O., Ritter R., 3rd, Abel K.J., Manning A., Panhuysen C., Farrer L.A. Complement factor H polymorphism and age-related macular degeneration. Science. 2005;308:421–424. doi: 10.1126/science.1110189. doi:10.1126/science.1110189. [DOI] [PubMed] [Google Scholar]
- 6.Hageman G.S., Anderson D.H., Johnson L.V., Hancox L.S., Taiber A.J., Hardisty L.I., Hageman J.L., Stockman H.A., Borchardt J.D., Gehrs K.M., et al. A common haplotype in the complement regulatory gene factor H (HF1/CFH) predisposes individuals to age-related macular degeneration. Proc. Natl Acad. Sci. USA. 2005;102:7227–7232. doi: 10.1073/pnas.0501536102. doi:10.1073/pnas.0501536102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Haines J.L., Hauser M.A., Schmidt S., Scott W.K., Olson L.M., Gallins P., Spencer K.L., Kwan S.Y., Noureddine M., Gilbert J.R., et al. Complement factor H variant increases the risk of age-related macular degeneration. Science. 2005;308:419–421. doi: 10.1126/science.1110359. doi:10.1126/science.1110359. [DOI] [PubMed] [Google Scholar]
- 8.Klein R.J., Zeiss C., Chew E.Y., Tsai J.Y., Sackler R.S., Haynes C., Henning A.K., SanGiovanni J.P., Mane S.M., Mayne S.T., et al. Complement factor H polymorphism in age-related macular degeneration. Science. 2005;308:385–389. doi: 10.1126/science.1109557. doi:10.1126/science.1109557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hughes A.E., Orr N., Esfandiary H., Diaz-Torres M., Goodship T., Chakravarthy U. A common CFH haplotype, with deletion of CFHR1 and CFHR3, is associated with lower risk of age-related macular degeneration. Nat. Genet. 2006;38:1173–1177. doi: 10.1038/ng1890. doi:10.1038/ng1890. [DOI] [PubMed] [Google Scholar]
- 10.Fritsche L.G., Lauer N., Hartmann A., Stippa S., Keilhauer C.N., Oppermann M., Pandey M.K., Kohl J., Zipfel P.F., Weber B.H., et al. An imbalance of human complement regulatory proteins CFHR1, CFHR3 and factor H influences risk for age-related macular degeneration (AMD) Hum. Mol. Genet. 2010;19:4694–4704. doi: 10.1093/hmg/ddq399. doi:10.1093/hmg/ddq399. [DOI] [PubMed] [Google Scholar]
- 11.Gold B., Merriam J.E., Zernant J., Hancox L.S., Taiber A.J., Gehrs K., Cramer K., Neel J., Bergeron J., Barile G.R., et al. Variation in factor B (BF) and complement component 2 (C2) genes is associated with age-related macular degeneration. Nat. Genet. 2006;38:458–462. doi: 10.1038/ng1750. doi:10.1038/ng1750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jakobsdottir J., Conley Y.P., Weeks D.E., Ferrell R.E., Gorin M.B. C2 and CFB genes in age-related maculopathy and joint action with CFH and LOC387715 genes. PloS ONE. 2008;3:e2199. doi: 10.1371/journal.pone.0002199. doi:10.1371/journal.pone.0002199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Maller J., George S., Purcell S., Fagerness J., Altshuler D., Daly M.J., Seddon J.M. Common variation in three genes, including a noncoding variant in CFH, strongly influences risk of age-related macular degeneration. Nat. Genet. 2006;38:1055–1059. doi: 10.1038/ng1873. doi:10.1038/ng1873. [DOI] [PubMed] [Google Scholar]
- 14.Spencer K.L., Hauser M.A., Olson L.M., Schmidt S., Scott W.K., Gallins P., Agarwal A., Postel E.A., Pericak-Vance M.A., Haines J.L. Protective effect of complement factor B and complement component 2 variants in age-related macular degeneration. Hum. Mol. Genet. 2007;16:1986–1992. doi: 10.1093/hmg/ddm146. doi:10.1093/hmg/ddm146. [DOI] [PubMed] [Google Scholar]
- 15.Maller J.B., Fagerness J.A., Reynolds R.C., Neale B.M., Daly M.J., Seddon J.M. Variation in complement factor 3 is associated with risk of age-related macular degeneration. Nat. Genet. 2007;39:1200–1201. doi: 10.1038/ng2131. doi:10.1038/ng2131. [DOI] [PubMed] [Google Scholar]
- 16.Yates J.R., Sepp T., Matharu B.K., Khan J.C., Thurlby D.A., Shahid H., Clayton D.G., Hayward C., Morgan J., Wright A.F., et al. Complement C3 variant and the risk of age-related macular degeneration. N. Engl. J. Med. 2007;357:553–561. doi: 10.1056/NEJMoa072618. doi:10.1056/NEJMoa072618. [DOI] [PubMed] [Google Scholar]
- 17.Fagerness J.A., Maller J.B., Neale B.M., Reynolds R.C., Daly M.J., Seddon J.M. Variation near complement factor I is associated with risk of advanced AMD. Eur. J. Hum. Genet. 2009;17:100–104. doi: 10.1038/ejhg.2008.140. doi:10.1038/ejhg.2008.140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chen W., Stambolian D., Edwards A.O., Branham K.E., Othman M., Jakobsdottir J., Tosakulwong N., Pericak-Vance M.A., Campochiaro P.A., Klein M.L., et al. Genetic variants near TIMP3 and high-density lipoprotein-associated loci influence susceptibility to age-related macular degeneration. Proc. Natl Acad. Sci. USA. 2010;107:7401–7406. doi: 10.1073/pnas.0912702107. doi:10.1073/pnas.0912702107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Klaver C.C., Kliffen M., van Duijn C.M., Hofman A., Cruts M., Grobbee D.E., van Broeckhoven C., de Jong P.T. Genetic association of apolipoprotein E with age-related macular degeneration. Am. J. Hum. Genet. 1998;63:200–206. doi: 10.1086/301901. doi:10.1086/301901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fritsche L.G., Freitag-Wolf S., Bettecken T., Meitinger T., Keilhauer C.N., Krawczak M., Weber B.H. Age-related macular degeneration and functional promoter and coding variants of the apolipoprotein E gene. Hum. Mutat. 2009;30:1048–1053. doi: 10.1002/humu.20957. doi:10.1002/humu.20957. [DOI] [PubMed] [Google Scholar]
- 21.Majewski J., Schultz D.W., Weleber R.G., Schain M.B., Edwards A.O., Matise T.C., Acott T.S., Ott J., Klein M.L. Age-related macular degeneration—a genome scan in extended families. Am. J. Hum. Genet. 2003;73:540–550. doi: 10.1086/377701. doi:10.1086/377701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Seddon J.M., Santangelo S.L., Book K., Chong S., Cote J. A genomewide scan for age-related macular degeneration provides evidence for linkage to several chromosomal regions. Am. J. Hum. Genet. 2003;73:780–790. doi: 10.1086/378505. doi:10.1086/378505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kenealy S.J., Schmidt S., Agarwal A., Postel E.A., De La Paz M.A., Pericak-Vance M.A., Haines J.L. Linkage analysis for age-related macular degeneration supports a gene on chromosome 10q26. Mol. Vis. 2004;10:57–61. [PubMed] [Google Scholar]
- 24.Jakobsdottir J., Conley Y.P., Weeks D.E., Mah T.S., Ferrell R.E., Gorin M.B. Susceptibility genes for age-related maculopathy on chromosome 10q26. Am. J. Hum. Genet. 2005;77:389–407. doi: 10.1086/444437. doi:10.1086/444437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rivera A., Fisher S.A., Fritsche L.G., Keilhauer C.N., Lichtner P., Meitinger T., Weber B.H. Hypothetical LOC387715 is a second major susceptibility gene for age-related macular degeneration, contributing independently of complement factor H to disease risk. Hum. Mol. Genet. 2005;14:3227–3236. doi: 10.1093/hmg/ddi353. doi:10.1093/hmg/ddi353. [DOI] [PubMed] [Google Scholar]
- 26.Fritsche L.G., Loenhardt T., Janssen A., Fisher S.A., Rivera A., Keilhauer C.N., Weber B.H. Age-related macular degeneration is associated with an unstable ARMS2 (LOC387715) mRNA. Nat. Genet. 2008;40:892–896. doi: 10.1038/ng.170. doi:10.1038/ng.170. [DOI] [PubMed] [Google Scholar]
- 27.Yang Z., Camp N.J., Sun H., Tong Z., Gibbs D., Cameron D.J., Chen H., Zhao Y., Pearson E., Li X., et al. A variant of the HTRA1 gene increases susceptibility to age-related macular degeneration. Science. 2006;314:992–993. doi: 10.1126/science.1133811. doi:10.1126/science.1133811. [DOI] [PubMed] [Google Scholar]
- 28.Dewan A., Liu M., Hartman S., Zhang S.S., Liu D.T., Zhao C., Tam P.O., Chan W.M., Lam D.S., Snyder M., et al. HTRA1 promoter polymorphism in wet age-related macular degeneration. Science. 2006;314:989–992. doi: 10.1126/science.1133807. doi:10.1126/science.1133807. [DOI] [PubMed] [Google Scholar]
- 29.Kanda A., Chen W., Othman M., Branham K.E., Brooks M., Khanna R., He S., Lyons R., Abecasis G.R., Swaroop A. A variant of mitochondrial protein LOC387715/ARMS2, not HTRA1, is strongly associated with age-related macular degeneration. Proc. Natl Acad. Sci. USA. 2007;104:16227–16232. doi: 10.1073/pnas.0703933104. doi:10.1073/pnas.0703933104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.An E., Sen S., Park S.K., Gordish-Dressman H., Hathout Y. Identification of novel substrates for the serine protease HTRA1 in the human RPE secretome. Invest. Ophthalmol. Vis. Sci. 2010;51:3379–3386. doi: 10.1167/iovs.09-4853. doi:10.1167/iovs.09-4853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yang Z., Tong Z., Chen Y., Zeng J., Lu F., Sun X., Zhao C., Wang K., Davey L., Chen H., et al. Genetic and functional dissection of HTRA1 and LOC387715 in age-related macular degeneration. PLoS Genet. 2010;6:e1000836. doi: 10.1371/journal.pgen.1000836. doi:10.1371/journal.pgen.1000836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chowers I., Meir T., Lederman M., Goldenberg-Cohen N., Cohen Y., Banin E., Averbukh E., Hemo I., Pollack A., Axer-Siegel R., et al. Sequence variants in HTRA1 and LOC387715/ARMS2 and phenotype and response to photodynamic therapy in neovascular age-related macular degeneration in populations from Israel. Mol. Vis. 2008;14:2263–2271. [PMC free article] [PubMed] [Google Scholar]
- 33.Kanda A., Stambolian D., Chen W., Curcio C.A., Abecasis G.R., Swaroop A. Age-related macular degeneration-associated variants at chromosome 10q26 do not significantly alter ARMS2 and HTRA1 transcript levels in the human retina. Mol. Vis. 2010;16:1317–1323. [PMC free article] [PubMed] [Google Scholar]
- 34.Wang G., Spencer K.L., Scott W.K., Whitehead P., Court B.L., Ayala-Haedo J., Mayo P., Schwartz S.G., Kovach J.L., Gallins P., et al. Analysis of the indel at the ARMS2 3'UTR in age-related macular degeneration. Hum. Genet. 2010;127:595–602. doi: 10.1007/s00439-010-0805-8. doi:10.1007/s00439-010-0805-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Allikmets R., Dean M. Bringing age-related macular degeneration into focus. Nat. Genet. 2008;40:820–821. doi: 10.1038/ng0708-820. doi:10.1038/ng0708-820. [DOI] [PubMed] [Google Scholar]
- 36.Kozak M. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene. 2005;361:13–37. doi: 10.1016/j.gene.2005.06.037. doi:10.1016/j.gene.2005.06.037. [DOI] [PubMed] [Google Scholar]
- 37.Barreau C., Paillard L., Osborne H.B. AU-rich elements and associated factors: are there unifying principles. Nucleic Acids Res. 2005;33:7138–7150. doi: 10.1093/nar/gki1012. doi:10.1093/nar/gki1012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Garneau N.L., Wilusz J., Wilusz C.J. The highways and byways of mRNA decay. Nat. Rev. Mol. Cell Biol. 2007;8:113–126. doi: 10.1038/nrm2104. doi:10.1038/nrm2104. [DOI] [PubMed] [Google Scholar]
- 39.Ly D.H., Lockhart D.J., Lerner R.A., Schultz P.G. Mitotic misregulation and human aging. Science. 2000;287:2486–2492. doi: 10.1126/science.287.5462.2486. doi:10.1126/science.287.5462.2486. [DOI] [PubMed] [Google Scholar]
- 40.De Luca A., De Falco M., De Luca L., Penta R., Shridhar V., Baldi F., Campioni M., Paggi M.G., Baldi A. Pattern of expression of HtrA1 during mouse development. J. Histochem. Cytochem. 2004;52:1609–1617. doi: 10.1369/jhc.4A6330.2004. doi:10.1369/jhc.4A6330.2004. [DOI] [PubMed] [Google Scholar]
- 41.Chien J., Staub J., Hu S.I., Erickson-Johnson M.R., Couch F.J., Smith D.I., Crowl R.M., Kaufmann S.H., Shridhar V. A candidate tumor suppressor HtrA1 is downregulated in ovarian cancer. Oncogene. 2004;23:1636–1644. doi: 10.1038/sj.onc.1207271. doi:10.1038/sj.onc.1207271. [DOI] [PubMed] [Google Scholar]
- 42.Campioni M., Severino A., Manente L., Tuduce I.L., Toldo S., Caraglia M., Crispi S., Ehrmann M., He X., Maguire J., et al. The serine protease HtrA1 specifically interacts and degrades the tuberous sclerosis complex 2 protein. Mol. Cancer Res. 2010;8:1248–1260. doi: 10.1158/1541-7786.MCR-09-0473. doi:10.1158/1541-7786.MCR-09-0473. [DOI] [PubMed] [Google Scholar]
- 43.Chien J., Campioni M., Shridhar V., Baldi A. HtrA serine proteases as potential therapeutic targets in cancer. Curr. Cancer Drug Targets. 2009;9:451–468. doi: 10.2174/156800909788486704. doi:10.2174/156800909788486704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Polur I., Lee P.L., Servais J.M., Xu L., Li Y. Role of HTRA1, a serine protease, in the progression of articular cartilage degeneration. Histol. Histopathol. 2010;25:599–608. doi: 10.14670/hh-25.599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Roesch K., Jadhav A.P., Trimarchi J.M., Stadler M.B., Roska B., Sun B.B., Cepko C.L. The transcriptome of retinal Muller glial cells. J. Comp. Neurol. 2008;509:225–238. doi: 10.1002/cne.21730. doi:10.1002/cne.21730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wang G., Spencer K.L., Court B.L., Olson L.M., Scott W.K., Haines J.L., Pericak-Vance M.A. Localization of age-related macular degeneration-associated ARMS2 in cytosol, not mitochondria. Invest. Ophthalmol. Vis. Sci. 2009;50:3084–3090. doi: 10.1167/iovs.08-3240. doi:10.1167/iovs.08-3240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kortvely E., Hauck S.M., Duetsch G., Gloeckner C.J., Kremmer E., Alge-Priglinger C.S., Deeg C.A., Ueffing M. ARMS2 is a constituent of the extracellular matrix providing a link between familial and sporadic age-related macular degenerations. Invest. Ophthalmol. Vis. Sci. 2010;51:79–88. doi: 10.1167/iovs.09-3850. doi:10.1167/iovs.09-3850. [DOI] [PubMed] [Google Scholar]
- 48.Gabriel S.B., Schaffner S.F., Nguyen H., Moore J.M., Roy J., Blumenstiel B., Higgins J., DeFelice M., Lochner A., Faggart M., et al. The structure of haplotype blocks in the human genome. Science. 2002;296:2225–2229. doi: 10.1126/science.1069424. doi:10.1126/science.1069424. [DOI] [PubMed] [Google Scholar]
- 49.Barrett J.C., Fry B., Maller J., Daly M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–265. doi: 10.1093/bioinformatics/bth457. doi:10.1093/bioinformatics/bth457. [DOI] [PubMed] [Google Scholar]
- 50.Karlstetter M., Walczak Y., Weigelt K., Ebert S., Van den Brulle J., Schwer H., Fuchshofer R., Langmann T. The novel activated microglia/macrophage WAP domain protein, AMWAP, acts as a counter-regulator of proinflammatory response. J. Immunol. 2010;185:3379–3390. doi: 10.4049/jimmunol.0903300. doi:10.4049/jimmunol.0903300. [DOI] [PubMed] [Google Scholar]
- 51.Lee J., Myers C.A., Williams N., Abdelaziz M., Corbo J.C. Quantitative fine-tuning of photoreceptor cis-regulatory elements through affinity modulation of transcription factor binding sites. Gene Ther. 2010;17:1390–1399. doi: 10.1038/gt.2010.77. doi:10.1038/gt.2010.77. [DOI] [PubMed] [Google Scholar]
- 52.Matsuda T., Cepko C.L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA. 2004;101:16–22. doi: 10.1073/pnas.2235688100. doi:10.1073/pnas.2235688100. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.